Abstract
The Drug Design Data Resource (D3R) Grand Challenges are blind contests organized to assess the state-of-the-art methods accuracy in predicting binding modes and relative binding free energies of experimentally validated ligands for a given target. The second stage of the D3R Grand Challenge 2 (GC2) was focused on ranking 102 compounds according to their predicted affinity for Farnesoid X Receptor. In this task, our workflow was ranked 5th out of the 77 submissions in the structure-based category. Our strategy consisted in (1) a combination of molecular docking using AutoDock 4.2 and manual edition of available structures for binding poses generation using SeeSAR, (2) the use of HYDE scoring for pose selection, and (3) a hierarchical ranking using HYDE and MM/GBSA. In this report, we detail our pose generation and ligands ranking protocols and provide guidelines to be used in a prospective computer aided drug design program.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Drug Design Data Resource (D3R) organizes each year a blind prediction challenge that represents a unique opportunity to evaluate and validate computer-aided drug design workflows [1]. The D3R community provides a set of ligands as input and asks participants to blindly predict their relative binding affinities for a given target, and finally compares predicted ranking to experimental data.
For its second edition, the Grand Challenge 2 (GC2), the D3R community focused on the Farnesoïd X receptor (FXR) and provided a set of 102 ligands [2]. For Stage 1, participants were asked to classify the 102 ligands based on their predicted affinity for FXR, and to predict the binding pose of a subset of 36 ligands (S1 set). By the end of Stage 1, co-crystallized X-ray structures of the S1 set were released, allowing participants to refine their pose prediction protocol and to focus on the ligands relative affinities ranking for the GC2 Stage 2. The 66 ligands for which no co-crystal structures were available are subsequently referred to as the S2 set. In the present report, we describe our participation to Stage 2 using the S1 set structural data provided at the end of Stage 1.
In GC2, a key issue was to account for the flexibility of the FXR ligand binding site (LBS) [3]. Several methods are used to apprehend flexibility in docking procedures, starting either from a single or from an ensemble of protein conformations. The former case comprises induced-fit docking that typically allows only restricted motions of LBS side chains. The ensemble approach uses PDB structures or molecular dynamics (MD) simulation snapshots to either perform docking on each receptor conformation, or merge the energetic contribution of individual protein conformations into a potential grid used as receptor for the docking. While the former method could explore extensively LBS local motions, the ensemble approach is likely to better account for large ligand-induced conformational changes in the LBS, such as loop motion. In the present report, we took advantage of the availability of diverse FXR structures co-crystallized with ligands of the same chemical series as the D3R ligands to combine two approaches to account for FXR LBS flexibility: (1) using the available FXR X-ray structures, we manually generated poses for the S2 set ligands based on S1 set co-crystallized ligands from the same chemical series; (2) using an ensemble docking approach with Autodock 4.2 [4] based on six FXR structures sampling the flexibility of the FXR LBS. We merged the poses generated by both protocols, and retained a single pose for each ligand of the S2 set based on HYDE [5] scores. The selected poses for S2 set ligands and X-ray structures of S1 set ligands were pooled and ranked according to HYDE. The ex-aequo compounds were re-ranked according to their binding free energy estimated by MM/GBSA (Fig. 1).
The classification we submitted to the GC2 Stage 2 ranked our protocol 5th out of the 77 submissions in the structure-based category. In the present report, we describe our poses generation and ligands ranking protocol, and discuss potential improvements to be routinely applied in a computer aided drug design pipeline.
Methods
Ligand preparation
The 102 small molecules provided by the D3R community in 2D SDF format were converted to 3D and protonated at pH 7.4 using Open Babel 2.3.2 [6]. Gasteiger partial charges were attributed using AutoDockTools 1.5.6 [4].
Manual edition using SeeSAR
Among the 102 ligands of the GC2, 96 are derived from four series (benzimidazoles, isoxazoles, spiros and sulfonamides) and six are miscellaneous. Most of the ligands from S2 set are very similar to the ligands from S1 set, whose co-crystals were released by the end of Stage 1. We selected the structurally closest compound from the S1 set for each compound of the S2 set, and used the SeeSAR software [7] to modify the corresponding ligand structure in the co-crystal. The energy of the generated poses was minimized using a two-step scheme: (1) the H-bond network within the protein and at the protein–ligand interface was optimized using Protoss [8]; (2) the ligand geometry was optimized using HYDE [5] as implemented in SeeSAR [7].
Structures selection
All FXR structures available in the PDB [9], consisting of 25 agonist-bound and 1 antagonist-bound structures (PDB Set), were retrieved and added to the 36 FXR structures provided after Stage 1 (S1 Set). We performed a hierarchical clustering of the PDB and the S1 Sets based on their pairwise LBS RMSD to select a subset of structures that samples experimentally observed FXR LBS flexibility. The LBS was defined as the residues with at least one atom within 5 Å of a co-crystallized ligand. Pairwise RMSD were computed using Pymol [10]. The structures were clustered according to hclust in R 3.2.3 [11]. We selected the representative structures based on three criteria: (1) we discarded isoxazoles and miscellaneous bound structures since they were not represented in the S2 set, (2) we selected at least one structure for each cluster and (3) the co-crystalized ligand shared a common scaffold with at least one of the 66 Stage 2 ligands. Selected FXR structures were protonated at pH 7.4 with Chimera [12] using the ProPKA [13] module and Gasteiger partial charges were attributed using AutoDockTools 1.5.6 [4]. Water molecules were not considered.
AutoDock 4.2
We ran AutoDock 4.2 [4] docking with default parameters (250 GA runs) to sample near-native poses and clustered the generated poses based on a 2 Å RMSD criteria. We considered cluster representatives, i.e. the lowest score pose of each cluster, for further scoring. S1 ligands predicted poses versus S1 ligands binding mode symmetry-corrected RMSD were computed using RDKit [14]. Ligand #33 was not included into the dataset for the docking protocol since the co-crystal showed an inconsistency in the ligand structure. A latter crystal structure provided by Roche allowed the completion of the dataset and was further included for the affinity ranking step.
Pose selection
For the ligands of the S1 set, we retained the crystallographic binding poses. For the ligands of the S2 set, refined poses of manually edited structures and cluster representatives of the predicted binding modes using AutoDock 4.2 were scored with HYDE; top score pose of each ligand was retained for further analysis.
HYDE
The HYdrogen bond and DEhydration (HYDE) empirical scoring function subsequently describes the energy balance between unfavorable hydrophilic dehydration and favorable hydrogen bonding during the binding process [5]. The essential feature of HYDE is the integrated use of log P-derived atomic increments for the prediction of free dehydration energy and hydrogen bonding energy. Taking the dehydration of atoms within the interface into account shows that some atoms contribute favorably to the overall score, while others contribute unfavorably [15].
MM/GBSA approach
MM/GBSA estimates the binding-free energy of a ligand to a target as the sum of the classical enthalpic contributions (bound, Van der Waals and electrostatic energies), the solvation free energies, and the entropic contribution [16]. Ligands parameters were generated with Antechamber [17, 18]. Up to 100 steps of conjugate gradient minimization of the complex were performed using NAMD [19] and the GAFF/ff99SB force field [18, 20]. The OBC1 Generalized Born (GB) model parameters [21] and the LCPO method [22] were used to compute the polar contribution to the solvation energy and the solvent-accessible surface area (SASA), respectively. The entropic term was omitted.
Results
Pose generation
Manual edition
Among the 102 of the challenge, the co-crystals of the 36 S1 set molecules were made available after GC2 Stage 1. Since the 66 S2 set molecules share high structural similarity with the S1 set ligands (average MACCS fingerprint Tanimoto = 0.94), we generated poses for the S2 ligands by editing closely related co-crystallized ligands from the S1 set. We edited the poses with the SeeSAR ligand editor. Poses were then submitted to a two-step minimization protocol using ProToss and HYDE as implemented in SeeSAR. The S2 ligands and their corresponding S1 co-crystallized ligands are presented in Supplementary Table S1.
This pose generation procedure was not possible for the compounds FXR #45 and #90 since steric clashes occurred in the binding site. Hence, only the docking procedure was used for these two ligands.
Docking
In order to cover the experimentally observed conformational variability of the FXR structures available in the PDB and the S1 set, we performed a hierarchical clustering based on their pairwise LBS RMSD. Overall, 1 structure from the PDB (PDB ID: 3OLF), and 5 structures from the S1 set (HQMF, YFJN, SJPR, KJYP and HVIH) were retained (Fig. 2).
Predicted binding modes of the S1 and S2 ligands into the six selected conformations of FXR were generated using AutoDock 4.2. All generated poses were clustered according to their relative RMSD and only the cluster representatives, i.e. the lowest score member of each cluster, were retained. For each ligand, up to 55 poses were retained. Since experimental binding modes were available for the ligands of the S1 set, the docking accuracy of our protocol was evaluated. As presented in Table 1 and Fig. 3, for single structure docking, the best-predicted poses of the S1 ligands displayed average RMSDs from 2.63 to 3.84 Å. For the ensemble docking, the best predicted binding modes of the S1 ligands displayed an average RMSD of 1.62 Å. Near native poses (RMSD <2 Å) could be retrieved for single structure and ensemble docking for respectively 19 and 28 out of the 36 S1 ligands (Fig. 4).
According to the chemotype classification of the ligands of the S1 set provided by the D3R, the ensemble docking approach displayed a better performance on the benzimidazoles, spiros and sulfonamides compared to the isoxazoles and miscellaneous compounds (Fig. 4).
Pose selection
S1 set
Since the experimental binding modes were provided by the D3R for the S1 ligands, the performance of the HYDE scoring function as implemented in SeeSAR in retrieving their experimental binding mode among the retained docking poses generated with AutoDock 4.2 was evaluated.
For 27 out of the 35 S1 ligands, the top score provided by the HYDE scoring function corresponded to the experimental binding mode. For the remaining S1 ligands, one near native pose (RMSD <2 Å) was retrieved as a top score.
For the next step, experimental binding modes were retained.
S2 set
For the ligands of the S2 set, the poses generated using the manual edition and the docking protocols were scored using HYDE; the top score poses were selected. The poses generated using the docking approach (FXR ligands #39, 40, 41, 43–45, 50, 53, 55, 56, 60–64, 66, 68, 72, 73, 80, 90, 93 and 98) and using the manual edition approach (remaining ligands) were selected for the next step.
Relative affinity prediction
The S1 and S2 retained ligands were pooled and ranked according to HYDE. In case of similar scores, notably for the predicted high affinity ligands, the ex-aequo compounds were re-ranked according to their binding free energy estimated by MM/GBSA.
The final ranking of the pooled S1 and S2 compounds was submitted as our prediction for the D3R GC2 under the alias dh2du.
Discussion
Chemical series dependent sampling variations
As presented in Fig. 4, AutoDock 4.2. allowed to retrieve near-native poses for the benzimidazoles, sulfonamides and spiros compounds using the ensemble docking approach. This observation is probably due to the presence of these chemotypes in the structures selected in the ensemble docking approach. For the miscellaneous class of compounds (for which no co-crystal was selected in the ensemble), no near-native poses could be retrieved. Since the FXR LBS conformation depends on the bound ligand (Fig. 2, helix H2 and H6), it should be taken into consideration in the selection of structures for docking studies.
Scoring
Relative affinity prediction
By the end of GC2 Stage 2, the D3R disclosed the experimental IC50 of the ligands of the S1 and the S2 sets. The Kendall correlation factor τ between the ranking of the S1 and the S2 ligands set according to (1) their experimental IC50 (experimental ranking) and (2) their predicted relative affinities was computed to evaluate the performance of the different GC2 participants protocols in terms of ranking accuracy. The ranking we submitted to the D3R GC2 led to Kendall τ coefficients of 0.35 and 0.41, respectively, on the S1 ligands set and the S1 + S2 ligands set. Our ranking was classified as the 5th best predictive result out of the 77 submissions in the structure-based scoring category; the best Kendall τ coefficient of this category for the S1 + S2 ligands set being 0.46, and the best Kendall τ of the ligand-based category being 0.38.
To complete our observations, we evaluated the individual performance of the HYDE, MM/GBSA and the combinaison of HYDE and MM/GBSA ranking schemes by assessing the Kendall τ correlation coefficients between their associated rankings on either the S1 set or the S1 + S2 set, and the experimental ranking (Table 2). On the S1 set, the MM/GBSA and HYDE ranking schemes displayed similar performance (τ = 0.38). On the S1 + S2 set, the HYDE and HYDE–MM/GBSA ranking schemes displayed a better correlation with the experimental data than the MM/GBSA ranking scheme with respective τ of 0.42, 0.41 and 0.35 (Table 2).
It is to note that the maximum predicted affinity for a given ligand with HYDE is <10−3 nM, which led to ex-aequo ranks for several ligands of the S1 and the S2 set that strongly impacted the Kendall τ correlation coefficients.
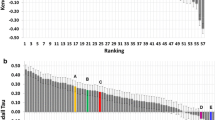
We then plotted the predicted ranking against the experimental IC50 values for the compounds of the S1 + S2 sets (Fig. 5). According to the binding free energies obtained with MM/GBSA, the spiros compounds were ranked before the benzimidazole compounds (Fig. 5a) whereas the scores obtained with HYDE resulted in the opposite trend (Fig. 5b), which is in better accordance with experimental data.
Considering the ranking of the most populated chemotypes, benzimidazoles and spiros were predicted with respective Kendall factors τ of 0.38 and 0.41 by MM/GBSA (Table 2). MM/GBSA outperformed HYDE score for the spiros family (τ = 0.41 vs. τ = 0.16, Table 2). Benzimidazoles rankings were better predicted by HYDE (τ= 0.46). Sulfonamides rankings were predicted by both methods with τ of −0.1 for HYDE and −0.09 for MM/GBSA. Other chemotypes (isoxazoles and miscellaneous compounds) were not sufficiently populated for a significant interpretation.
The MM/GBSA approach is widely used to approximate ligand–protein binding free energy [16]. In the present work, we used an unique MM/GBSA calculation on each individual pose. Better results could be obtained using MM/GBSA free energies averaged over a set of MD simulations snapshots or an ensemble of X-ray crystal structures since (1) the dynamics of the complex influences the binding free energy and (2) the accuracy of the results can be statistically assessed [16]. However, these benefits come with a significantly higher computational cost hardly compatible with a high-throughput pipeline. We only applied few minimization steps to reduce the steric clashes and computed the binding free energies from a single ligand–protein structure, implying a strong dependence between the predicted binding free energies and the predicted binding modes.
Protocol automation
For the D3R GC2, a lot of structural data were available since many X-ray structures were (1) provided by the organizers and (2) available in the PDB. Since structural data were not always available for the chemotypes that are structurally close to the compounds studied in a drug discovery program, we performed a retrospective evaluation of a fully automated procedure using AutoDock 4.2 [4] and AutoDock VINA [23] for the binding mode prediction step (Fig. 6) and HYDE for the pose selection and ranking steps.
We used the ensemble docking protocol we applied in the D3R GC2 with the 36 ligands from the S1 set. AutoDock VINA displayed a slightly better performance than AutoDock 4.2 in retrieving near-native poses (RMSD <2 Å) (30 and 28 out of 36 ligands, respectively) with respective average RMSD of 1.32 and 1.62 Å (Fig. 6). HYDE score allowed to retrieve most of the AutoDock VINA generated near-native poses (23 out of 30) which was not the case with the AutoDock 4.2 generated near-native poses (14 out of 28).
In terms of ranking, the automated protocol using AutoDock VINA and HYDE score displayed similar results compared to the strategy we used in the D3R GC2, with respective τ values of 0.44 and 0.40 on the S1 + S2 set and the S1 set, respectively. This automated protocol should be favored in a prospective CADD program.
Conclusion
In the present report, we described our structure-based hierarchical ranking protocol that was ranked 5th out of the 77 submissions in the structure-based category of the D3R GC2. This protocol strongly relied on experimental data since we used a manual edition of co-crystallized ligands and numerous target structures through an ensemble docking approach. However, we also described a fully automated approach to overcome the manual edition step that displays similar results (Kendall τ of 0.41 and 0.44, respectively). Scoring and ranking using the HYDE scoring function is adapted for high to low affinity ligands and can be routinely applied for medium size compounds collections. HYDE displayed a good performance in estimating the relative affinity of the FXR ligands in the datasets provided by the D3R. However, since ties can be obtained with ligands displaying similarly theoretical high affinities, a strict ranking cannot always be performed. Advanced rescoring at the cost of computational time using methods such as polarizable force field and molecular mechanics could bring a higher degree of accuracy for binding free energy prediction.
References
Gathiaka S, Liu S, Chiu M, Yang H, Stuckey JA, Kang YN et al (2016) D3R Grand Challenge 2015: evaluation of protein–ligand pose and affinity predictions. J Comput Aided Mol Des 30:651–668
Gaieb Z, Liu S, Chui M, Yang H, Shao C, Gathiaka S et al (2016) Drug Design Data Resource Grand Challenge 2 Dataset: Farnesoid X Receptor. Drug Design Data Resource. University of California, San Diego
Jin L, Feng X, Rong H, Pan Z, Inaba Y, Qiu L et al (2013) The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism. Nat Commun 4:1937
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS et al (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791
Schneider N, Lange G, Hindle S, Klein R, Rarey M (2013) A consistent description of HYdrogen bond and DEhydration energies in protein–ligand complexes: methods behind the HYDE scoring function. J Comput Aided Mol Des 27:15–29
O’Boyle NM, Banck M, James CA, Morley C, Vandermeersch T, Hutchison GR (2011) Open Babel: an open chemical toolbox. J Cheminf 3:33
SeeSAR v5.5, BioSolveIT GmbH, St.Augustin, Germany. http://www.biosolveit.de/SeeSAR
Bietz S, Urbaczek S, Schulz B, Rarey M (2014) Protoss: a holistic approach to predict tautomers and protonation states in protein–ligand complexes. J Cheminf 6:12
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H et al (2000) The Protein Data Bank. Nucleic Acids Res 28:235–242
De Lano W (2002) The PyMOL molecular graphics system. DeLano Scientific, Palo Alto. http://www.pymol.org
Dessau RB, Pipper CB (2008) [‘‘R"--project for statistical computing]. Ugeskr Laeger 170:328–330
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC et al (2004) UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Olsson MH, Sondergaard CR, Rostkowski M, Jensen JH (2011) PROPKA3: Consistent treatment of internal and surface residues in empirical pKa predictions. J Chem Theory Comput 7:525–537
RDKit, Open-source cheminformatics. http://www.rdkit.org
Reulecke I, Lange G, Albrecht J, Klein R, Rarey M (2008) Towards an integrated description of hydrogen bonding and dehydration: decreasing false positives in virtual screening with the HYDE scoring function. ChemMedChem 3:885–897
Genheden S, Ryde U (2015) The MM/PBSA and MM/GBSA methods to estimate ligand-binding affinities. Expert Opin Drug Discov 10:449–461
Wang J, Wang W, Kollman PA, Case DA (2006) Automatic atom type and bond type perception in molecular mechanical calculations. J Mol Graph Model 25:247–260
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA (2004) Development and testing of a general amber force field. J Comput Chem 25:1157–1174
Phillips JC, Braun R, Wang W, Gumbart J, Tajkhorshid E, Villa E et al (2005) Scalable molecular dynamics with NAMD. J Comput Chem 26:1781–1802
Hornak V, Abel R, Okur A, Strockbine B, Roitberg A, Simmerling C (2006) Comparison of multiple Amber force fields and development of improved protein backbone parameters. Proteins 65:712–725
Onufriev A, Bashford D, Case DA (2004) Exploring protein native states and large-scale conformational changes with a modified generalized born model. Proteins 55:383–394
Weiser J, Shenkin P, Still WC (1999) Approximate atomic surfaces from linear combinations of pairwise overlaps (LCPO). J Comput Chem 20:217–230
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461
Acknowledgements
MR is recipient of a MNESR fellowship. We thank Dr. Marcus Gastreich and BioSolveIT GmBH for providing SeeSAR.
Author information
Authors and Affiliations
Corresponding author
Additional information
Manon Réau and Florent Langenfeld have contributed equally to this study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Réau, M., Langenfeld, F., Zagury, JF. et al. Predicting the affinity of Farnesoid X Receptor ligands through a hierarchical ranking protocol: a D3R Grand Challenge 2 case study. J Comput Aided Mol Des 32, 231–238 (2018). https://doi.org/10.1007/s10822-017-0063-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-017-0063-0