Abstract
This Perspective explores how consideration of hydrogen bonding can be used to both predict and better understand partition coefficients. It is shown how polarity of both compounds and substructures can be estimated from measured alkane/water partition coefficients. When polarity is defined in this manner, hydrogen bond donors are typically less polar than hydrogen bond acceptors. Analysis of alkane/water partition coefficients in conjunction with molecular electrostatic potential calculations suggests that aromatic chloro substituents may be less lipophilic than is generally believed and that some of the effect of chloro-substitution stems from making the aromatic π-cloud less available to hydrogen bond donors. Relationships between polarity and calculated hydrogen bond basicity are derived for aromatic nitrogen and carbonyl oxygen. Aligned hydrogen bond acceptors appear to present special challenges for prediction of alkane/water partition coefficients and this may reflect ‘frustration’ of solvation resulting from overlapping hydration spheres. It is also shown how calculated hydrogen bond basicity can be used to model the effect of aromatic aza-substitution on octanol/water partition coefficients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lipophilicity is the most important physicochemical property in drug discovery and a key design parameter in medicinal chemistry [1–4]. Lipophilicity has traditionally been linked [5–8] to permeability although it has long been recognized that high lipophilicity is associated with poor aqueous solubility [9] and is an undesirable feature in compounds intended to be drugs [10]. Lipophilicity considerations feature prominently in the well-known ‘Rule of 5’ (Ro5) [11] which is essentially a statement of physicochemical property distributions for compounds that had been taken into Phase II clinical studies at some point before the publication of the original study. Although invoked frequently, and occasionally outside its applicability domain of oral absorption, Ro5 provides no guidance as to how compliant compounds should be optimized. It is also unclear why the high polarity limit for Ro5 is specified in terms of hydrogen bonding while the low polarity limit is defined by lipophilicity. The wide acceptance of Ro5, and the popularity of approaches to data presentation that hide or mask variation, tend to blind drug discovery scientists to the possibility that lipophilicity may be less predictive of outcomes, such as pharmacological promiscuity, than is commonly believed [12].
Lipophilicity is usually quantified as a partition coefficient (P) and the nature of solute partitioning between immiscible solvents has been understood for many years [13]. The distribution coefficient, D, of compound X may be defined as the ratio of concentrations of X in solvents S1 and S2, where [Xi](S1) and [Xi](S2) are the concentrations of form i of the compound in solvents S1 and S2 respectively:
The partition coefficient P is usually defined as the ratio of the concentrations of the neutral form of X in the two solvents:
Partition coefficients in drug discovery are conventionally defined with S1 as the organic solvent and S2 as water which means that the partitioning system may be specified by the organic solvent (e.g., Poct for octanol/water; Pchx for cyclohexane/water; Phxd for hexadecane/water). Partition coefficients are usually quoted as their base 10 logarithms and, in this study, we will use the abbreviations ‘logP’ and ‘logD’ for the base 10 logarithms of P and D with subscripts to indicate the organic phase (e.g., logPoct). The most commonly used organic solvent for lipophilicity measurement is octanol [14, 15] and a number of methods exist for prediction of logPoct [16]. The aqueous phase is typically buffered (e.g., pH 7.4) for lipophilicity measurements and it is D (as opposed to P) that is actually measured. The distribution coefficient, which is a function of pH, and P are identical for compounds that are not significantly ionized at the measurement pH. Making the assumption that only neutral forms of compounds partition into the organic phase, D can be written as a function of P and the fraction, Fneut, of compound present as neutral form in the aqueous phase [17]:
In some cases [18, 19], ionized forms of compounds do partition into the organic phase and, in these situations, D also depends on the nature and concentration of counter ion(s). If required, logP can be obtained from the logD-pH profile or by applying Eq. (3) with a measured pKa value. However, neither of these approaches is routinely used in drug discovery programs and the logP values quoted for compounds that are significantly ionized are usually calculated rather than measured.
Like molecular size, lipophilicity can be regarded as a risk factor in drug discovery and the most direct way to monitor it during the course of a lead optimization project is to plot the response of potency to lipophilicity [20]. Provided that it is not simply a reflection of a narrow range in the data, a weak correlation between potency and lipophilicity is actually desirable because it indicates that the discovery project team has room to maneuver. When potency and lipophilicity are more strongly correlated, the response of the former to the latter should be as steep as possible and this consideration can also be used to assess different structural series within a project. It is also useful to model the response of pIC50 to logP (or logD) because this allows potency to be ‘normalized’ with respect to risk factor and the residuals quantify the extent to which the activity of a compound beats (or is beaten by) the trend in the data [20]. Andrews et al. used residuals in an analogous manner in their 1984 study of functional group contributions to drug-receptor interactions [21]. Subtraction of logD [22] or logP [23] from pIC50 was suggested for normalization of activity with respect to lipophilicity and the difference between pIC50 and logP (or logD) subsequently became known as ligand lipophilicity efficiency or lipophilic ligand efficiency (LLE) and lipophilic efficiency (LiPE) [24]. The difference between potency and logP can be interpreted as a measure of the ease of transferring the neutral form of a compound from an organic solvent (usually octanol) to its binding site although this interpretation is no longer valid when compounds bind to targets in ionized forms [20]. LLE/LiPE will appear to decrease with lipophilicity if the gradient of a linear response of potency to lipophilicity is less than unity and this should generally be considered as a characteristic of the structural series rather than interpreted in terms of ‘quality’ of individual compounds.
The octanol/water partitioning system is arbitrary and it has been suggested [17] that its adoption may reflect misinterpretation of work by Collander [25] who was aware of the relevance of the hydrogen bonding characteristics of the organic phase to partitioning. Octanol can form hydrogen bonds with solutes on account of the hydroxyl group in its molecular structure and its high water content at saturation (2.5 M; equivalent to mole fraction of 0.29) [26] is greater than that of cyclohexane (0.003 M) [27] or hexadecane (0.002 M) [28]. It has been argued [29, 30] that a hydrocarbon solvent is a more appropriate model for the lipid bilayer core. The alkane/water partition coefficient (logPalk) provides a more direct measure of aqueous solvation energy [17, 31–34] than its octanol/water counterpart (logPoct) while being more amenable to measurement than gas to water transfer free energy [35]. It has also been suggested that a solvent lacking hydrogen bonding capacity would represent the most appropriate reference state for normalizing potency with respect to lipophilicity [20]. Alkane/water partitioning systems are also more sensitive than octanol/water to changes in polarity resulting from conformational biasing and intramolecular hydrogen bonding [36]. Cyclohexane [37], and other hydrocarbon solvents such as hexadecane [28, 29] have been used for logP measurement for many years [38–59]. The difference between logPoct and logPalk provides a measure of solute hydrogen bonding capacity and is of considerable interest in its own right [39, 41, 44–46, 51, 53, 54]. It is usually given the symbol ΔlogP and it is effectively an octanol/alkane partition coefficient where both phases are saturated with water.
Although cyclohexane and hexadecane are the most commonly encountered organic solvents in alkane/water partitioning studies, other hydrocarbon solvents are also used and it is typically necessary to aggregate measurements for different alkanes (and different experimental protocols) for modelling studies [17]. The term ‘alkane/water partition coefficient’ (logPalk) is used both as a generic description of measurements made using very similar partitioning systems and to acknowledge that data has been aggregated for analysis. Compounds of interest to medicinal chemists tend to be poorly soluble in saturated hydrocarbons and this presents challenges for measurement of alkane/water partition coefficients. Self-association is more of a concern for logPalk measurement than for logPoct. Just as ionization in the aqueous phase makes compounds appear to be less lipophilic than they actually are, self-association in the organic phase effectively masks polarity and results in an increase in apparent lipophilicity. Furthermore, differences in spectral characteristics (e.g. dimer absorbs more strongly than monomer) have the potential to exaggerate effects of self-association. However, partitioning of ions into the organic phase is less likely for hydrocarbon solvents than for octanol and the low solubility of water in the former reduces the likelihood of interactions with other solutes that can lead to ‘water-dragging’ [56]. Measurement [34, 48, 57–59] and prediction [17, 33, 49–52, 60–62] of logPalk are both areas of active research.
The presence of hydrogen bond (HB) acceptors and donors in the molecular structure of a solute favors aqueous solvation and tends to make the solute less lipophilic. The less polar the organic phase, the greater the sensitivity of logP to solute hydrogen bonding capacity although it should be noted that contact between polar and non-polar molecular surfaces is not inherently repulsive [63]. HB acidity and basicity are usually quantified as association constants for 1:1 complexes in low-polarity solvents such as carbon tetrachloride or 1,1,1-trichloroethane and a large body of measured data (mainly HB basicity) is available [64–68]. Calculated molecular electrostatic potential (MEP) is an effective predictor of both HB acidity [69] and HB basicity [63, 70–72]. Minimized MEP (Vmin) reflects the electronic distribution within atoms and is arguably more relevant to intermolecular interactions than atomic charges which describe the electronic distribution between atoms [63]. MEP minima cannot, in general, be reproduced by atom-centered (or bond-centered) multipoles [63]. Vmin can be thought of as a ‘lone pair’ descriptor that is capable of explaining why pyrazine can accept a hydrogen bond despite lacking a permanent dipole moment. When relating measured HB acidity/basicity (1:1 complex) to solvation behavior, it is important to be aware that HB donors and acceptors of solute interact simultaneously with a number of solvent molecules (1:N complex) [63, 68]. HB acidity/basicity measured for a polyfunctional compound with non-equivalent HB donors/acceptors is not generally meaningful [63, 68] unless individual contributions to the overall formation constant can be determined [73]. Despite these limitations, HB acidity and basicity considerations can provide insight into partitioning phenomena just as partition coefficient measurements can provide insight into the nature and strength of hydrogen bonding. Taken together, formation constants of 1:1 hydrogen bonded complexes and partition coefficients complement views [74, 75] of molecular recognition that are more based on analysis of X-ray crystal structures.
In this perspective we first show how analysis of logPalk measurements can be used to quantify polarity of both compounds and substructures. We then illustrate the connection between polarity defined in this manner and hydrogen bonding by using examples of polar atom types (e.g. HB donors; aromatic nitrogen) and substructures (e.g. aromatic rings).
Computational details
ADD_CENTRE [63] and MEP2HB were created with the OEChem [76] toolkit which was also used with the OESpicoli toolkit [77] to create ClogPalk [17]. Each of the ADD_CENTRE, MEP2HB and ClogPalk programs uses the OpenEye [78] implementation of SMARTS [79, 80] to specify substructures. Source code and documentation for these three programs and READ_GAUSS_FILE is provided as supplementary material.
Molecular structures were encoded as isomeric SMILES [81, 82] strings and Omega [83, 84] was used to generate a single conformation for each. Molecular geometries were energy-minimized in gas phase (MMFF94S) [85] using the Szybki [86] molecular mechanics program. Molecular surface area (MSA) was calculated from atomic coordinates and Bondi [87] radii using ClogPalk with a probe radius of 1.4 Å. Minimized molecular electrostatic potential [63, 70, 71] was calculated with Gaussian 09 [88] using the Hartree–Fock [89], B3LYP [90, 91] or MP2 [89, 92, 93] theoretical models with 6-31G** or 6-311 + G** basis sets [94–96]. The ADD_CENTRE software was used to calculate starting coordinates for MEP minimization by placing points on conventional ‘lone pair’ axes at distances that were typically between 1.3 and 1.5 Å from the relevant nucleus. The version of ADD_CENTRE (1.1) used in this study differs from the version (1.0) used previously [63] in that it provides additional functionality to probe-systems and handle nitroso oxygen. Starting points for MEP minimization with π-systems were generated by placing points on normals to the plane of symmetry that either pass through atomic nuclei or bond centroids at distances in the range 1.5 to 2.0 Å. MEP minima are typically more difficult to locate for aromatic rings than for heteroatoms and ADD_CENTRE has a feature that allows a normal passing through a bond centroid to be rotated around the bond axis. Vmin values were extracted from Gaussian 09 output using READ_GAUSS_FILE and HB basicity (pKBHX) values were calculated for these using MEP2HB which applies models derived in a previous study [63]. For each atom type, Vmin was calculated at the level of theory corresponding to the most predictive model for pKBHX. An updated file of models for prediction of pKBHX from Vmin is provided as supplemental material.
Measured alkane/water partition coefficients were taken from the literature and classified as CHX (cyclohexane), HXD (hexadecane) or ALK (alkane other than cyclohexane or hexadecane) according to the organic solvent. Unless otherwise stated, data in these three categories were aggregated for analysis and a file of 1144 values measured for 812 compounds is provided as supplementary information with links to their respective literature sources. Files of 453 measured HB basicity (pKBHX) values and 63 measured pKa values are also made available as supplementary information. Octanol/water partition coefficients were taken from a published compilation [97] or, in the case of 1,5-naphthyridine, from primary literature [51].
ClogPalk [17] was used to calculate reference logPalk values from MSA. The reference values used in the analyses for heteroaromatic nitrogen and carbonyl oxygen accounted for polarity of benzylic substituents by subtraction of the following correction factors: benzyl (1.07), 3-chlorobenzyl (1.09) or 4-phenylbenzyl (1.78). Correction was only made the presence of benzylic substituents for one data point in the analysis for aromatic nitrogen and three data points in the analysis for carbonyl oxygen. The version of ClogPalk used in the current study differs from that described previously [17] in the way that SMARTS patterns are matched. Previously, the parameter associated with a SMARTS pattern was only assigned to the atom mapping onto the first atom of the SMARTS string. In the current version (1.1), the parameter associated with a SMARTS pattern is assigned to all atoms that map onto that SMARTS pattern. Updated parameter files for the ClogPalk model that are compatible with the current version of the software are provided as supplemental material.
MUDO [98] was used for Matched Molecular Pair Analysis (MMPA) [99–104] and all statistical analysis was performed with JMP [105]. The predictive models for pKBHX and polarity used in this study (M01 to M16) are provided in Table 1.
Plot of measured logPalk against molecular surface area for saturated hydrocarbons (grey circles; model M01 in Table 1), saturated alcohols/diols (red diamonds; models M02/M03 in Table 1) and saturated ethers (green diamonds; model M04 in Table 1). Other than saturated carbon, only the atoms defining each functional group are present in molecular structures for this data set
Plot of polarity against measured HB basicity for structurally prototypical compounds with only a single HB acceptor or two symmetrically equivalent HB acceptors in their molecular structures. Data has been plotted on a per-HB acceptor basis by normalizing both Q and pKBHX by the number (n) of HB acceptor atoms and details for the least squares line of fit (model M05) are given in Table 1
Minimized Molecular Electrostatic Potential (MEP) for aromatic systems. The Vmin calculations were performed for MMFF94S energy-minimized structures using the MP2/6-311G** protocol and the atomic units (au) are Hartree per elemental charge. a Plot of HB basicity (pKBHX) against minimized MEP (Vmin) for non-fused aromatic compounds lacking conventional HB acceptors (model M06, Table 1). b Plot of polarity, Q, against (Vmin) for N-methylpyrrole, methyl-substituted benzenes and chloro-substituted benzenes. The logPalk value for 1-methylpyrrole has been estimated from the logPalk value measured for pyrrole, making the assumption that the effect of N-methylation will be the same for pyrrole as for indole. c Vmin/au values calculated for π-systems of 1-methylpyrrole, o-xylene and chlorobenzene showing the positions and magnitudes of the MEP minima
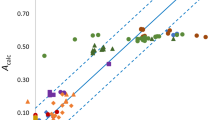
Relationship between corrected polarity, Qcorr, and pKBHX calculated for heteroaromatic nitrogen using model M08 (Table 1). A curve (model M12, Table 1) was fit to the filled black circles which correspond to compounds in which heteroaromatic rings are either non-fused or which have a doubly-connected nitrogen atom in each ring (e.g. 1,5-napthyridine). Halogen-substituted pyridines (red circles) and fused heterocycles (red and green diamonds) are shown for reference but were excluded from training set. For furan, logPalk was approximated by logPoct (ΔlogP ∼ 0 for weak HB acceptors) and the measured pKBHX value was used
Predicted logPalk for a selection of heteroaromatic compounds. Substructural polarity (qaromN) values associated with aromatic nitrogen atoms were calculated (M12, Table 1) from Vmin (M12, Table 1). A correction factor (+ 2.0) has been applied for presence of adjacent hydrogen bond acceptors in 10 and 12 (but not 7) and the uncorrected predictions are shown in parentheses for these compounds
Relationship between corrected polarity, Qcorr, and pKBHX calculated for carbonyl oxygen using model M09 (Table 1). The curve (model M12, Table 1) was fit to the black circles which correspond to molecular structures with either one carbonyl group (e.g. cyclopentanone) or two symmetrically equivalent carbonyl groups (benzoquinone). The red diamonds correspond to sulfoxides which were not included in the fitting and the pKBHX values for these compounds were predicted using model M10 (Table 1)
Estimation polarity from measured partition coefficients
The general framework used in this study for relating partition coefficients to hydrogen bond capacity can be summarized as:
In this framework, logP(expt) is the logP value measured for a compound and logP(ref) is logP for a physically meaningful reference state which may either be a measured or calculated value. The HB donor and acceptor capacities for the compound are represented by α and β respectively and these are vectors because, in general, molecular structures have multiple HB donors and acceptors. Equation (4) treats HB donors and acceptors as perturbations of the reference state and exploiting Eq. (4) requires that both reference state and function, f, be defined explicitly. Equation (4) can be used either to estimate HB capacity from logP measurements or to predict logP from calculated HB capacity.
The polarity of a compound may be estimated from measured logPalk by making use of the strong linear relationship (M01, Table 1) between logPalk and MSA that is observed for saturated hydrocarbons. The reference state is a hypothetical saturated hydrocarbon with identical MSA to the compound of interest for which logPalk can be calculated reliably using M01 (Table 1). The polarity, Q, of the compound is defined as the difference between the value of logPalk calculated for this reference state and the measured value:
Q can be treated as a sum of contributions (qi) from polar substructures where ni is the number of instances of substructure i in the molecular structure of the compound:
Equations (5) and (6) form the basis of the ClogPalk model [17] which associates qi values with substructures defined using SMARTS [79, 80] notation and is illustrated graphically in Fig. 1. Equations (5) and (6) can either be used with measured logPalk data to estimate qi or with calculated qi values to predict logPalk. A strong correlation between logPalk and molecular volume was also observed for saturated hydrocarbons and analogous analysis based on that relationship has been reported [45]. If measured logPalk values are available for a number of compounds with only the substructure of interest and saturated carbon present in their molecular structures then the mean value of Q provides a direct estimate of qi for that substructure. This is the preferred approach for estimation of substructural polarity from measured logPalk although its applicability may be limited by data availability. Once qi has been determined directly for substructure i (e.g. benzyl substituent), it can then be used to estimate qj from measured logPalk for compounds with only saturated carbon and substructures i and j in their molecular structures. This approach was used in the parameterization of the ClogPalk model [17] and estimation of substructural polarity in this manner may be termed ‘indirect’. When modelling the response of polarity to HB capacity, it can be useful to correct Q for presence of other HB acceptors and donors since this enables exploitation of more measured data than would otherwise be possible. A corrected value of Q may be defined as follows where n is the number of instances of the HB acceptor (or donor) of interest, and qcorr,i and ncorr,i are, respectively the polarity and number of instances of a substructure i:
In this study, Qcorr values were used to model the responses of polarity to calculated pKBHX for heteroaromatic nitrogen and carbonyl oxygen although correction factors were only defined for three substructures: benzyl (1.07), 3-chlorobenzyl (1.09) and 4-phenylbenzyl (1.78).
MMPA [99–104] can be used to estimate polarity differences between substructures. A matched molecular pair consists of two compounds that are linked by a specific structural transformation (e.g. carboxyl to tetrazole) that may be regarded as a perturbation of either structure. For example, the effect on logPalk of N-methylation of a secondary amide group may be estimated by averaging the difference in logPalk between secondary amides and their N-methylated analogs:
In general, the structural transformations that define matched molecular pairs result in changes in MSA and this must be accounted for when using MMPA to estimate polarity differences between substructures. For example, the difference in the polarity of substructures 1 and 2 can be written as:
The advantage of MMPA is that it allows measured data for compounds with non-equivalent HB donors and acceptors in their molecular structures to be exploited for estimation of polarity.
One advantage of defining polarity in terms of a difference between partition coefficients is that Q is invariant with respect to standard state. Partition coefficients are usually defined in terms of molar concentration units although mole fraction can also be used. Any model for partitioning (or binding) must be able to accommodate a change in standard state definition in order to be considered to have a valid thermodynamic basis. While prediction of partition coefficients is the main focus of this Perspective, measures of substructural polarity derived from logPalk are also of interest for modelling molecular recognition in aqueous media [55]. One of the objectives of this study is to evaluate calculated HB basicity as a predictor of substructural polarity and it is instructive to examine the relationship between Q and measured pKBHX that is illustrated in Fig. 2. The compounds in this data set were selected to have either a single HB acceptor (e.g. cyclohexanone) or two equivalent HB acceptors (e.g. dioxane) which means that Q can be associated with the HB acceptor of each compound. The results shown in Fig. 2 suggest that development of a model for logPalk that is based entirely (i.e. without substructural parameterization) on measures of HB acidity and basicity derived from formation constants of 1:1 hydrogen complexes is unlikely be feasible.
Polarity of hydrogen bond donors
The octanol/water system is relatively insensitive to the presence of HB donors in molecular structures and logPoct is of practically no value in assessing HB acidity [12, 46, 54]. Consequently, it is necessary to use alkane/water systems to study hydrogen bond donors with partition coefficient measurements. Polarity estimates for a number of common HB donors are presented in Table 2. Generally, the presence of an HB donor in a molecular structure implies that at least one HB acceptor is also present and this means that the HB donor contribution to polarity cannot be estimated directly using equations (5) and (6). Most of the values in Table 2 were derived from MMPA using Eq. (9). Availability of data made it possible to estimate polarity for hydroxyl, thiol and carboxylic acid HB donors by using equations (5) and (6) indirectly (e.g. as polarity difference between alcohols and ethers). Polarity was also estimated for the primary sulfonamide HB donors using equations (5) and (6) although this reflects lack of data for matched molecular pairs. One question that arises from this analysis concerns the extent to which alkylation of nitrogen or oxygen perturbs HB basicity although it is likely that donation of an HB to water will affect HB basicity in a similar manner.
The polarity estimates in Table 2 suggest that hydrogen atoms interact more strongly with water when bonded to oxygen than when bonded to nitrogen. This is broadly consistent with logKα values typically observed [66] for amides, phenols and carboxylic acids although it is important to be aware that HB donation by hydroxyl is likely to result in an increase in HB basicity of oxygen [51]. The interactions of the HB donors of benzamides and anilides with water appear particularly weak, suggesting that methylation of these nitrogen atoms favors conformations in which the amide carbonyl oxygen atom can form more effective interactions with water. Analogous observations have been made for chromatographically-measured lipophilicity [106].
Although polarity differences can be discerned between the different types of HB donor, it is more instructive to compare them with polarity estimates for compounds with a single HB acceptor nitrogen or oxygen. The values of Q (in parentheses) for acetonitrile (3.5), 1-methylimidazole (5.5), 1-methylpiperidine (3.8), tetrahydrofuran (3.1), acetone (3.8), dimethylformamide (5.7), N-acetylpyrrolidine (6.8) and dimethylsulfoxide (7.0) suggest that the HB acceptors in these compounds are typically more polar than any of the HB donors in Table 1. Defining polarity in terms of logPalk enables HB donors and acceptors to be brought onto the same scale in a way that is not possible with measures of HB acidity and basicity derived from association constants for 1:1 hydrogen bonded complexes. These observations point to a general tendency for water to interact more strongly with HB acceptors than with HB donors and are consistent with the view that anions interact more strongly than cations with water [107–109]. One question raised by the hydration imbalance between HB donors and acceptors concerns the extent to which it can be explained by the molecular (as opposed to the solvent) structure of water. The hydration imbalance between the HB donor and acceptor of the amide group should be considered when modelling protein folding and intramolecular hydrogen bonding of cyclic peptides.
Aromatic π-systems
Unlike other substructures used as illustrative examples, the HB capacity of π−systems cannot be linked to individual atoms. Aromatic hydrocarbons are more polar than saturated hydrocarbons and water is an order of magnitude more soluble in benzene than cyclohexane at temperatures ranging from 10 to 40 °C [27]. A Q value of 1.0 can be calculated for benzene using equations (5) and (6), indicating polarity comparable with the HB donor of an amide. An increase in the extent of the π-system typically leads to an increase in polarity although the Q values for phenanthrene (1.5) and pyrene (1.4) suggest that the trend is not particularly strong. The Q value for N-methylindole (2.5) indicates that this heterocycle is particularly polar and this is a factor that may need to be specifically accounted for when modelling interactions of tryptophan residues. The π-systems of aromatic rings function as HB acceptors and pKBHX values have been measured [68] for benzene (− 0.49) and 1-methylpyrrole (0.23). Figure 3a illustrates the relationship (M06, Table 1) between pKBHX and Vmin which can be used to predict pKBHX for the aromatic rings of chlorobenzene (−1.0) and 1,3-dichlorobenzene (−1.6). While it is well-established that aromatic π-systems can interact with HB donors, the key question in pharmaceutical design is whether the π-system of an aromatic ring interacts more or less strongly with its binding partner than with water.
The Vmin values associated with π-systems provide a measure of potential for interaction with HB donors and could be used as physicochemical descriptors of aromatic character [110]. Two pairs of MEP minima were observed for the π-system of indole and these are associated with the C4-C5 bond (Vmin = −0.035 au; calculated pKBHX = −0.06) and the C2–C3 bond (Vmin = −0.032 au; calculated pKBHX = −0.18). The MEP minima associated with the C4–C5 bond lie closer to C5 than C4 and it is significant that 5-azaindole is most basic of the azaindoles [111]. MEP calculations can be used to compare the effects of substitution and ring-fusion. For example, the Vmin value (−0.0005 au; predicted pKBHX = −2.15) calculated for buckminsterfullerene suggests that its π-system accepts hydrogen bonds even less readily (on a per-bond basis) than 1,3,5-trichlorobenzene (−0.0021 au; calculated pKBHX = −2.01). Two challenges for using pKBHX (or Vmin) to model aqueous solvation of π-systems are that numbers of interacting water molecules are not generally known and that HB basicity derived from data for 1:1 complexes is not directly relevant when a π-system accepts more than a single HB.
A plot of Q against Vmin is shown in Fig. 3b for a selection of non-fused aromatic compounds and a line (M07, Table 1) has been fit to the data for 1-methylpyrrole, benzene and the methylated benzenes. The chlorinated benzenes all lie above the reference line indicating that they are more polar than would be expected from Vmin values calculated for their π-systems. These results are consistent with a view that some of the lipophilicity increase associated with chloro-substitution is the result of a reduction in the HB basicity of the ring which would imply that chloro substituents on aromatic rings are less lipophilic than is commonly assumed [112, 113]. Additional support for this view comes from MMPA [99–104] which shows that replacement of chloro with methyl for primary alkyl chlorides leads, on average, to a 1.4 unit increase in logPhxd (Table 3). In contrast, replacement of a chloro substituent on a benzene ring with a methyl group tends to result in a small decrease in logPhxd. MEP calculations suggest that the chlorine atoms of chlorobenzene (Vmin = −0.019 au) and dichloromethane (Vmin = −0.020 au) are of similar polarity. A single MEP minimum (Vmin = −0.024 au) was found for 1,2-dichlorobenzene and this indicates that, in contrast with dichloromethane, through-space interactions between the chlorine atoms are more important than through-bond interactions.
Aromatic nitrogen
Aromatic nitrogen is an important molecular recognition element in medicinal chemistry and the pKBHX and logKβ values measured [66–68] for it span a wide range, indicating that this atom type is relatively sensitive to substructural context. This makes it more difficult to parameterize polarity by substructure and therefore increases the potential impact of a polarity model based on MEP. The relationship between Qcorr and calculated pKBHX (Model M08, Table 1) is shown in Fig. 4 for compounds with aromatic nitrogen HB acceptors. In this analysis, a substructural correction (for benzyl) was applied for a single measured logPalk value although two other values of Qcorr reflect scaling by the number of heteroaromatic nitrogen atoms. For modelling, the dataset has been restricted to molecular structures with one or more nitrogen atoms present in each aromatic ring and that are either unsubstituted or alkyl-substituted (e.g. 4-methylpyridine and 1,5-naphthyridine but not quinoxaline). The underlying assumption is polarity of an aza-substituted aromatic ring is dominated by the nitrogen so that the contribution of the π-cloud may be neglected. Making this assumption allows Qcorr to be equated to the substructural polarity, qaromN, of aromatic nitrogen for the training set compounds. 1-Benzylimidazole was included in the training set because the contribution to polarity of the benzyl group can be corrected for. An exponential function (Model M12, Table 1) was fitted to the data which allows qaromN to be calculated from Vmin. The rationale for fitting an exponential function is that the contribution of an HB acceptor to logPalk tends asymptotically to zero as the HB basicity becomes very weak. Values of Qcorr were also plotted in Fig. 4 for a number of compounds that had been excluded from the training set because of uncertainty about the contributions to polarity from substructures other than aromatic nitrogen. Fused five-membered heteroaromatic rings all lie above the fitted curve, indicating that other factors (e.g. presence of oxygen in ring; π-cloud polarity of carbocyclic ring) need to be considered when interpreting polarity for these compounds. The data for quinoline, isoquinoline and quinoxaline were not used for fitting M12 (Table 1), on account of the carbocyclic rings in their molecular structures. However, all lie close to the fitted curve which suggests that the carbocylic rings of these compounds make only small contributions to polarity. The pKBHX values calculated (Model M06, Table 1) for the carbocyclic rings of 1-methylbenzimidazole (−0.2), quinoline (−0.8), isoquinoline (−1.2) and quinoxaline (−1.3) may explain why the largest positive residual was observed for the first compound. Positive residuals were also observed for the halogenated species and this suggests that the polarity of the halogen atoms cannot be neglected. The pKBHX values calculated for the nitrogen (0.4) and each fluorine (−0.6) atom of 2,6-difluoropyridine suggest that the fluoro substituents significantly influence the polarity of this compound.
Equations (5) and (6) were used with calculated qaromN (M12, Table 1) to predict logPalk for a number of compounds for which the only substructures with HB capacity were aromatic nitrogen atoms (Fig. 5). Predicted and measured logPalk values were compared for five compounds with two or more non-equivalent heteroaromatic HB acceptors. The largest discrepancies between measurement and prediction were observed for 2 and 4 and, in each of these cases, the predicted value is less than the measured value which indicates that HB acceptor capacity has been over-estimated in the context of alkane/water partitioning. It is well known [63, 66–68, 71, 114] that heteroaromatic compounds such as pyridazine (12) with adjacent nitrogen atoms are better HB acceptors than their proton basicity would suggest and this can be considered as a manifestation of the α effect [115] or thought of in terms of secondary electrostatic interactions [116]. While only 1:1 complexes are typically observed in the measurement of HB acidity or basicity, the HB donors and acceptors present in a molecular structure can all simultaneously form hydrogen bonds with water molecules in aqueous solution. HB donation to one of the nitrogen atoms of pyridazine would be expected to make it more difficult for the other nitrogen atom to accept an HB for a number of reasons. Firstly, accepting an HB makes nitrogen more electronegative and this will tend to draw electron density away from the other nitrogen atom. Secondly, simultaneous HB donation to both nitrogen atoms of pyridazine would result in an electrostatically repulsive orientation of water molecules that is enthalpically unfavorable. Thirdly, the orientation of two water molecules would increase the degree of constraint in the system and is therefore expected to be entropically unfavorable. It is noteworthy that the logPalk value calculated for 1 is very similar to the measured value and this observation is consistent with N3, which is predicted to be a significantly stronger HB acceptor than N2, dominating the solvation of this triazole.
The view that adjacency of HB acceptors compromises solvation has implications for molecular design and it can be conjectured that similar considerations apply to adjacent HB donors. The entropic costs of solvating adjacent polar atoms can also be thought of in terms of molecular complexity [117] and solvation can be described as ‘frustrated’ [118] when hydration spheres of polar atoms overlap to a significant extent. This implies that the presence of adjacent HB donors or acceptors in a concave region of a protein molecular surface should be viewed as a design opportunity [119]. It has also been suggested that molecular structures capable of presenting arrangements of hydrogen bonding groups that cannot easily be mimicked by clusters of water molecules represent a molecular recognition theme [63] that might be exploited in fragment design [120]. Measurement of logPalk for structurally prototypical compounds would allow frustrated hydration to be studied systematically.
Predictions for a number of heteroaromatic compounds for which experimental values have not been reported are also presented in Fig. 5 and the values calculated for 10 and 12 (but not 7) have been corrected (+2.0) for the presence of adjacent HB acceptors. The HB acceptors of 9 and 15 are predicted to be the weakest for the structures shown in Fig. 5 and measured logPalk values for these would be particularly informative for refining the model illustrated in Fig. 4. The cyclohexane/water partition coefficient component of the SAMPL5 challenge [33, 34] features compounds of higher molecular complexity than the structurally prototypical compounds typically encountered in the logPalk literature and this is certainly appropriate for testing prediction methods. Nevertheless, a case can be made for inclusion of structural prototypes that are likely to present specific challenges for solvation models. The prediction difficulties presented by strong HB acceptors that are aligned point to compounds of potential interest in initiatives like SAMPL5 [33, 34] and the HB acceptor characteristics of 1,8-naphthyridine and 1,2,3-triazine have already been highlighted in this context [63]. Prediction in drug design frequently focuses [99–104] on differences between values of properties (e.g. decrease in solubility resulting from chloro-substitution) and this is a theme that could be explored in challenges such as SAMPL5 [33, 34]. For example, measured logPalk for pairs of compounds of identical molecular shape, but differing in their hydrogen bonding characteristics (e.g. 1-butyltetrazole and 2-butyltetrazole), would enable comparison of different solvation models with respect to their treatment of electrostatics.
Carbonyl oxygen
As is the case for aromatic nitrogen, the HB basicity of carbonyl oxygen is very sensitive to substructural context and it is therefore difficult to parameterize polarity for this atom type using substructural definitions. Oxygen atoms are typically associated with two HB acceptor sites that are not in general equivalent although this does not present special difficulties for modelling HB basicity because the experiments are designed so that only 1:1 complexes are observed [67, 68]. The situation is very different in solvents with HB donor capacity because an oxygen atom can simultaneously accept two hydrogen bonds and using one HB acceptor site is likely to result in a decrease in the HB basicity of the remaining site [51]. The situation is analogous to that of aligned HB acceptors of 2 and 4 discussed in the previous section, although each HB acceptor site is likely to be even more sensitive to the environment of the other. The approach used in this study was to model polarity using the greater of the two pKBHX values predicted for each carbonyl oxygen atom in cases where the two values differ and HB basicity of carbonyl oxygen has been treated in an analogous manner for prediction of ΔlogP [51]. As was the case for aromatic nitrogen, the training set was restricted to compounds for which the polarity of the carbonyl oxygen could be estimated from measured logPalk. In cases where the carbonyl group is part of an extended, non-fused, π-system (e.g. tertiary amides and benzoquinone but not naphthoquinone) substructural polarity is assumed to be due to the carbonyl oxygen. Three quinolones with benzylic substituents on nitrogen were also included in the training set because their inclusion improves coverage of chemical space and the polarity of substituents can be accounted for. The relationship between Qcorr and pKBHX predicted using M09 (Table 1) is shown in Fig. 6 for compounds with carbonyl or sulfoxide oxygen as the only atoms with HB capacity in their molecular structures. The data points for the sulfoxides were not used for modelling and all lie below the curve (M13, Table 1) that has been fit which suggests that predicted pKBHX exaggerates the polarity of sulfoxide oxygen.
Equations (5) and (6) were used to predict logPalk using models M12 (aromatic nitrogen), M13 (carbonyl oxygen), M14 (hydroxyl donating intramolecular HB) and the qHBD values in Table 2 (HB donors). The results are shown in Fig. 7 and agreement between predicted and measured values is poorer than what might be expected from the root mean square error (RMSE) for model M13 (Table 1) which highlights the difficulties in extrapolating from structural prototypes to situations where HB acceptors and/or donors are in close proximity. The predictions tend to exaggerate the polarity of these compounds and predicted logPalk values are typically lower than the measured values. As noted in the previous section, simultaneous solvation of adjacent hydrogen bonding sites is likely to incur, at very least, an entropic cost and Vmin does not capture the polarization of a solute that accepts hydrogen bonds from one or more water molecules. The discrepancies between predicted and measured logPalk are particularly extreme for 21, 22, 24 and 25 which may reflect a structural feature (carbonyl group adjacent to doubly-connected nitrogen) that is shared by these compounds. However, a more subtle factor may also be exerting its influence here. The carbonyl oxygen atoms for the compounds in the training set typically have HB acceptor sites for which the calculated pKBHX values are either identical or, at least, very similar. In contrast, the pKBHX values calculated for the HB acceptor sites of 22 (3.4 and 2.2) differ by 1.2. This raises a more general question for quantitative structure activity/property relationship (QSAR/QSPR) modelling. Suppose two descriptors X1 and X2 are strongly correlated for the training set compounds. Should a set of compounds for which X1 and X2 are weakly correlated be considered to be within the same region of chemical space as the training set simply because the values of all descriptors used in the model lie within the ranges of training set values?
Calculated logPalk for compounds with carbonyl oxygen HB acceptors. Substructural polarity associated with atoms were calculated from Vmin using models M12, M13 or M14 (Table 1) or taken from Table 2. Each logPalk value is calculated by subtracting substructural polarity values from a reference logPalk value calculated (model M01, Table 1) for a saturated hydrocarbon with the same molecular surface area. Hydroxyl groups forming intramolecular hydrogen bonds were modelled (M14, Table 1) as ethers and the participating HB donors were treated as non-polar
Values of logPalk have been calculated for three compounds for which intramolecular hydrogen bonding is likely to influence partitioning characteristics. The calculated logPalk values are all lower (by 0.3 to 1.1 unit) than the measured values which indicates that the polarity of the compounds has been over-estimated. Formation of an intramolecular HB eliminates one of the MEP minima associated with carbonyl oxygen and it could be argued that this would place the compound outside the applicability domain of a model trained with data for carbonyl groups with pairs of very similar Vmin values. Nevertheless, the MEP calculations capture essential features of the intramolecular HB such as the reduced availability of the remaining oxygen ‘lone pair’. The intramolecular HBs for these three compounds are likely to persist in the aqueous phase and this would be expected to facilitate prediction of logPalk, especially for a method like ClogPalk that uses a single conformation to represent a structure.
Modeling logPoct for aza analogs of benzene and naphthalene
Although alkane/water partition coefficients represent the main focus of this study, hydrogen bonding also influences their octanol/water equivalents. Figure 8 illustrates the relationship between the effect of aza-substitution on logPoct and the pKBHX calculated for nitrogen. The analysis has been performed on a per-nitrogen basis and the data points fall into two groups according to whether or not a carbocyclic ring is present in the molecular structure of the aza-analog. The small residuals observed for phthalazine and cinnoline suggest that proximity of HB acceptors is much less of a problem for prediction of logPoct than for logPalk. Calculated values of logPoct for aza analogs of benzene and naphthalene are shown in Fig. 9. On a technical note, aza-analogs of benzene and naphthalene should be considered outside the applicability domains of these models if they are substituted (even with alkyl).
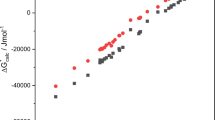
Relationship between effect on logPoct of aza substitution of benzene or naphthalene and pKBHX calculated for aromatic nitrogen using model M08 (Table 1). Differences in logPoct values were scaled by number of aza substitutions and points are grouped according to whether or not the aza-substituted species is fused with a carbocycle (1,5-naphthyridine is grouped with the aza-benzenes in green). The data was fitted using models M15 (green) and M16 (red) in Table 1
Prediction of logPoct for some six-membered heteroaromatic rings. Substructural polarity (qaromN) associated with nitrogen atoms values were calculated from Vmin using models M12 (Table 1). Each logPoct value is calculated by subtracting qaromN values from measured logPoct for either benzene (2.13) or naphthalene (3.30)
One aspect of lipophilicity control in molecular design is to achieve a balance between the polar and non-polar portions of molecular structures. The logPoct values for benzene (2.1) and 4-propylpyridine (2.1) suggest that aza-substitution of benzene will counter the effect of a propyl substituent. However, in the hexadecane/water partitioning system, 4-propylpyridine (logPhxd = 1.3) is 0.8 units less lipophilic than benzene (logPhxd = 2.1) suggesting that aza-substitution will more than compensate for the presence of a propyl group. Differences like these raise the question of which partitioning system is ‘right’ for lead optimization and even whether there is a single ‘right’ partitioning system for all applications. Despite its limitations, logPoct is likely to remain a useful design parameter for lead optimization and knowledge of HB acidity and basicity can help the medicinal chemist minimize the impact of the limitations. Lead optimization is usually carried out against structural series that are defined by scaffolds and HB acceptors/donors (and ionizable groups) tend to be relatively conserved within series. This means that the choice of partitioning system becomes less important when working within a series than when performing data analysis for structurally diverse sets of compounds [20]. If a plot of pIC50 against logPoct shows a compound to be deviating sharply from the trend line, it is advisable to assess the hydrogen bonding characteristics of the compound before jumping to the conclusion that the observed potency is especially unusual. The medicinal chemist should also be cautious when attempting to extrapolate trends (e.g. response of aqueous solubility to logPoct) observed for one series to another series and be especially wary of any analysis in which continuous data has been transformed to categorical data [12].
Conclusions
We show how logPalk measurements can be analyzed to define polarity for both compounds and substructures. Using a number of illustrative examples, we make a connection between polarity defined in terms of partitioning and hydrogen bonding defined in terms of 1:1 complex stability. Defining polarity in this way highlights the hydration imbalance between the HB donor and acceptor of the amide group. Two insights relevant to molecular design are that aromatic chloro substituents may be less hydrophobic that is commonly believed and that hydration of adjacent HB acceptors (or donors) is likely to be frustrated. We show how pKBHX values calculated for aromatic nitrogen and carbonyl oxygen can be used in prediction of partition coefficients.
References
van de Waterbeemd H, Smith DA, Jones BC (2001) Lipophilicity in PK design: methyl, ethyl, futile. J Comput Aided Mol Des 15:273–286
Giaginis C, Tsantili-Kakoulidou A (2008) Alternative measures of lipophilicity: from octanol–water partitioning to IAM retention. J Pharm Sci 97:2984–3004
Waring MJ (2010) Lipophilicity in drug discovery. Expert Opin Drug Discov 5:235–248
Sarkar A, Kellogg GE (2010) Hydrophobicity—shake flasks, protein folding and drug discovery. Curr Top Med Chem 10:67–83
Collander R (1937) Permeability. Ann Rev Biochem 6:1–18
Lindemann B, Solomon AK (1962) Permeability of luminal surface of intestinal mucosal cells. J Gen Physiol 45:801–810
Oldendorf WH (1974) Lipid solubility and drug penetration of the blood brain barrier. Exp Biol Med 147:813–816
Banks WA, Kastin A (1985) Peptides and the blood–brain barrier: lipophilicity as a predictor of permeability. Brain Res Bull 15:287–292
Yalkowsky SH, Valvan SC (1980) Solubility and partitioning I: solubility of nonelectrolytes in water. J Pharm Sci 69:912–922
Hansch C, Björkroth JP, Leo A (1987) Hydrophobicity and central nervous system agents: on the principle of minimal hydrophobicity in drug design. J Pharm Sci 76:663–687
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (1997) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 23:3–25
Kenny PW, Montanari CA (2013) Inflation of correlation in the pursuit of drug-likeness. J Comput Aided Mol Des 27:1–13
Nernst W (1891) Verteilung eines Stoffes zwischen zwei Lösungsmitteln und zwischen Lösungsmittel und Dampfraum. Z Phys Chem 8:110–139
Leo A, Hansch C, Elkins D (1971) Partition coefficients and their uses. Chem Rev 71:525–616
Dearden JC, Bresnen GM (1988) The measurement of partition coefficients. Quant Struct Act Relatsh 7:133–144
Mannhold R, Poda GI, Ostermann C, Tetko IV (2009) Calculation of molecular lipophilicity: state-of-the-art and comparison of log P methods on more than 96,000 compounds. J Pharm Sci 98:861–893
Kenny PW, Montanari CA, Prokopczyk IM (2013) ClogPalk: a method for predicting alkane/water partition coefficient. J Comput Aided Mol Des 27:389–402
Harris MJ, Higuchi T, Rytting JH (1973) Thermodynamic group contributions from ion pair extraction equilibriums for use in the prediction of partition coefficients. Correlation of surface area with group contributions. J Phys Chem 77:2694–2703
Scherrer RA, Donovan SF (2009) Automated Potentiometric Titrations in KCl/Water-saturated octanol: method for quantifying factors influencing ion-pair partitioning. Anal Chem 81:2768–2778
Kenny PW, Leitão A, Montanari CA (2014) Ligand efficiency metrics considered harmful. J Comput Aided Mol Des 28:699–710
Andrews PR, Craik DJ, Martin JL (1984) Functional group contributions to drug-receptor interactions. J Med Chem 27:1648–1657
Leach AR, Hann MM, Burrows JN, Griffen EJ (2006) Fragment screening: an introduction. Mol BioSyst 2:429–446
Albert JS, Blomberg N, Breeze AL, Brown AJH, Burrows JN, Edwards PD, Folmer RHA, Geschwindner S, Griffen EJ, Kenny PW, Nowak T, Olsson L, Sanganee H, Shapiro AB (2007) An integrated approach to fragment-based lead generation: philosophy, strategy and case studies from AstraZeneca’s drug discovery programmes. Curr Top Med Chem 7:1600–1629
Hopkins AL, Keserü GM, Leeson PD, Rees DC, Reynolds CH (2014) The role of ligand efficiency metrics in drug discovery. Nat Rev Drug Discov 13:105–121
Collander R (1951) Partition of organic compounds between higher alcohols and water. Acta Chem Scand 5:774–780
Dallas AJ, Carr PW (1992) A thermodynamic and solvatochromic investigation of the effect of water on the phase-transfer properties of octan-1-ol. J Chem Soc Perkin Trans 2 1992:2155–2161
Goldman S (1974) The determination and statistical mechanical interpretation of the solubility of water in benzene, carbon tetrachloride, and cyclohexane. Can J Chem 52:1668–1680
Abraham MH, Whiting GS, Fuchs R, Chambers EJ (1990) Thermodynamics of solute transfer from water to hexadecane. J Chem Soc Perkin Trans 2 1990:291–300
Finkelstein A (1976) Water and nonelectrolyte permeability of lipid bilayer membranes. J Gen Physiol 68:127–135
Mayer PT, Anderson BD (2002) Transport across 1,9-decadiene precisely mimics the chemical selectivity of the barrier domain in egg lecithin bilayers. J Pharm Sci 91:640–646
Radzicka A, Wolfenden R (1988) Comparing the polarities of the amino acids: side-chain distribution coefficients between the vapor phase, cyclohexane, 1-octanol, and neutral aqueous solution. Biochem 27:1664–1670
Shih P, Pedersen LG, Gibbs PR, Wolfenden R (1998) Hydrophobicities of the nucleic acid bases: distribution coefficients from water to cyclohexane. J Mol Biol 280:421–430
Bannan CC, Burley KH, Chiu M, Shirts MR, Gilson MK, Mobley DL (2016) Blind prediction of cyclohexane–water distribution coefficients from the SAMPL5 challenge. J Comput Aided Mol Des. doi:10.1007/s10822-016-9954-8
Rustenburg AS, Dancer J, Lin B, Feng JA, Ortwine DF, Mobley DL, Chodera JD (2016) J Comput Aided Mol Des. doi:10.1007/s10822-016-9971-7
Cabani S, Gianni P, Mollica V, Lepori L (1981) Group contributions to the thermodynamic properties of nonionic organic solutes in dilute aqueous solution. J Solut Chem 10:563–595
Dearden JC, Bresnen GM (2005) Thermodynamics of water–octanol and water–cyclohexane partitioning of some aromatic compounds. Int J Mol Sci 6:119–129
Golumbic C, Orchin M, Weller S (1949) Partition studies on phenols. I. Relation between partition coefficient and ionization constant. J Am Chem Soc 71:2624–2627
Delaney AD, Currie DJ, Holmes HL (1969) Partition coefficients of some N-alkyl and N, N-dialkyl derivatives of some cinnamamides and benzalcyanoacetamides in the system cyclohexane–water. Can J Chem 47:3273–3277
Seiler P (1974) Interconversion of lipophilicities from hydrocarbon/water systems into the octanol/water system. Eur J Med Chem 9:473–479
Riebesehl W, Tomlinson E (1984) Enthalpies of solute transfer between alkanes and water determined directly by flow microcalorimetry. J Phys Chem 88:4770–4775
Young RC, Mitchell RC, Brown TH, Ganellin CR, Griffiths R, Jones M, Rana KK, Saunders D, Smith IR, Sore NE, Wilks TJ (1988) Development of a new physicochemical model for brain penetration and its application to design of centrally acting H2 receptor histamine antagonists. J Med Chem 31:656–671
Lambert WJ, Wright LA (1989) Development of a preformulation lipophilicity screen utilizing a C-18-derivatized polystyrene–divinylbenzene High-performance liquid chromatographic (HPLC) column. Pharm Res 7:577–586
El Tayar N, Tsai R-S, Testa B, Carrupt P-A, Leo A (1991) Partitioning of solutes in different solvent systems: the contribution of hydrogen-bonding capacity and polarity. J Pharm Sci 80:590–598
Leahy DE, Morris JJ, Taylor PJ, Wait AR (1992) Model solvent systems for QSAR. Part 2. Fragment values (f-values) for the critical quartet. J Chem Soc Perkin Trans 2 1992:723–731
El Tayar N, Testa B, Carrupt P-A (1992) Polar intermolecular interactions encoded in partition coefficients: an indirect estimation of hydrogen-bond parameters of polyfunctional solutes. J Phys Chem 96:1455–1459
Abraham MH, Chadha HS, Whiting GS, Mitchell RC (1994) Hydrogen bonding. 32. An analysis of water–octanol and water–alkane partitioning and the ∆logP parameter of Seiler. J Pharm Sci 83:1085–1100
Habgood MD, Liu ZD, Dehkordi LS, Khodr HH, Abbott J, Hider RC (1999) Investigation into the correlation between structure of hydroxypyridones and blood–brain barrier permeability. Biochem Pharmacol 57:1305–1310
Wohnsland F, Faller B (2001) High-throughput permeability pH profile and high-throughput alkane/water log P with artificial membranes. J Med Chem 44:923–930
Zissimos AM, Abraham MH, Barker MC, Box KJ, Tam KY (2002) Calculation of Abraham descriptors from solvent–water partition coefficients in four different systems; evaluation of different methods of calculation. J Chem Soc Perkin Trans 2 2002:470–477
Caron G, Ermondi G (2005) Calculating virtual log P in the alkane/water System (log PN alk) and its derived parameters ∆log PN oct–alk and log DpH alk. J Med Chem 48:3269–3279
Toulmin A, Wood JM, Kenny PW (2008) Toward prediction of alkane/water partition coefficients. J Med Chem 51:3720–3730
Wittekindt C, Klamt A (2009) COSMO-RS as a predictive tool for lipophilicity. QSAR Comb Sci 28:874–877
Shalaeva M, Giulia Caron G, Abramov YA, O’Connell TN, Plummer MS, Yalamanchi G, Farley KA, Goetz GH, Philippe L, Shapiro MJ (2013) Integrating intramolecular hydrogen bonding (IMHB) considerations in drug discovery using ∆logP as a tool. J Med Chem 56:4870–4879
Ermondi G, Visconti A, Esposito R, Caron G (2014) The Block Relevance (BR) analysis supports the dominating effect of solutes hydrogen bond acidity on ∆log Poct–tol. Eur J Pharm Sci 53:50–54
Chen D, Oezguen N, Urvil P, Ferguson C, Dann SM, Savidge TC (2016) Regulation of protein–ligand binding affinity by hydrogen bond pairing. Sci Adv 2:e1501240
Tsai R-S, Fan W, El Tayar N, Carrupt P-A, Testa B, Kier LB (1993) Solute-water interactions in the organic phase of a biphasic system. 1. Structural influence of organic solutes on the “water-dragging” effect. J Am Chem Soc 115:9632–9639
Bard B, Carrupt P-A, Martel S (2012) Determination of alkane/water partition coefficients of polar compounds using hydrophilic interaction chromatography. J Chromatogr A 1260:164–168
Lin B, Pease JH (2013) A novel method for high throughput lipophilicity determination by microscale shake flask and liquid chromatography tandem mass spectrometry. Comb Chem High Throughput Screen 16: 817–825.
Jensen DA, Gary RK (2015) Estimation of alkane–water log P for neutral, acidic, and basic compounds using an alkylated polystyrene–divinylbenzene high-performance liquid chromatography column. J Chromatogr A 1417:21–29
Chung K, Park H (2016) Extended solvent-contact model approach to blind SAMPL5 prediction challenge for the distribution coefficients of drug-like molecules. J Comput Aided Mol Des. doi:10.1007/s10822-016-9928-x
Klamt A, Eckert F, Reinisch J, Wichmann K (2016) Prediction of cyclohexane–water distribution coefficients with COSMO-RS on the SAMPL5 data set. J Comput Aided Mol Des. doi:10.1007/s10822-016-9927-y
Bannan CC, Calabro G, Kyu DY, Mobley DL (2016) Calculating partition coefficients of small molecules in octanol/water and cyclohexane/water. J Chem Theor Comput 12:4015–4024
Kenny PW, Montanari CA, Prokopczyk IM, Ribeiro JFR, Sartori GR (2016) Hydrogen bond basicity prediction for medicinal chemistry design. J Med Chem 59:4278–4288
Abraham MH (1993) Scales of solute hydrogen-bonding: their construction and application to physicochemical and biochemical processes. Chem Soc Rev 22:73–83
Taft RW, Gurka D, Joris L, Schleyer PVR, Rakshys JW (1969) Studies of hydrogen-bonded complex formation with p-fluorophenol. V. Linear free energy relationships with OH reference acids. J Am Chem Soc 91:4801–4808
Abraham MH, Duce PP, Prior DV, Barratt DG, Morris JJ, Taylor PJ (1989) Hydrogen bonding. Part 9. Solute proton-donor and proton-acceptor scales for use in drug design. J Chem Soc Perkin Trans 2 1989:1355–1375
Laurence C, Berthelot M (2000) Observations on the strength of hydrogen bonding. Perspect Drug Discov Des 18:39–60
Laurence C, Brameld KA, Graton J, Le Questel J-Y, Renault E (2009) The pKBHX database: toward a better understanding of hydrogen-bond basicity for medicinal chemists. J Med Chem 52:4073–4086
Kenny PW (2009) Hydrogen bonding, electrostatic potential and molecular design. J Chem Inf Model 49:1234–1244
Murray JS, Ranganathan S, Politzer P (1991) Correlations between the solvent hydrogen bond acceptor parameter β and the calculated molecular electrostatic potential. J Org Chem 56:3734–3739
Kenny PW (1994) Prediction of hydrogen bond basicity from computed molecular electrostatic properties: implications for comparative molecular field analysis. J Chem Soc Perkin Trans 2 1994:199–202
Graton J, Le Questel J-Y, Maxwell P, Popelier PLA (2016) Hydrogen-bond accepting properties of new heteroaromatic rings chemical motifs: a theoretical study. J Chem Inf Model 56:322–334
Graton J, Berthelot M, Gal J-F, Laurence C, Lebreton J, Le Questel J-Y, Maria P-C, Robins R (2003) The nicotinic pharmacophore: thermodynamics of the hydrogen-bonding complexation of nicotine, nornicotine, and models. J Org Chem 68:8208–8221
Bissantz C, Kuhn B, Stahl M (2010) A medicinal chemist’s guide to molecular interactions. J Med Chem 53:5061–5084
Persch E, Dumele O, Diederich F (2015) Molecular recognition in chemical and biological systems. Angew Chem Int Ed 54:3290–3327
OEChem Toolkit. OpenEye Scientific Software. http://www.eyesopen.com/oechem-tk. Accessed 19 Aug 2016
Spicoli Toolkit. OpenEye Scientific Software. http://www.eyesopen.com/spicoli-tk. Accessed 19 Aug 2016
OpenEye Scientific Software, 9 Bisbee Court, Suite D, Santa Fe, NM 87508. http://www.eyesopen.com. Accessed 28 Feb 2013
SMARTS Theory Manual. Daylight Chemical Information Systems. http://www.daylight.com/dayhtml/doc/theory/theory.smarts.html
SMARTS at Wikipedia http://en.wikipedia.org/wiki/Smiles_arbitrary_target_specification
Weininger D (1988) SMILES, a chemical language and information system. 1. Introduction to methodology and encoding rules. J Chem Inf Comp Sci 28:31–36
Weininger D, Weininger A, Weininger JL (1989) SMILES. 2. Algorithm for generation of unique SMILES notation. J Chem Inf Comp Sci 29:97–101
OMEGA. OpenEye Scientific Software http://www.eyesopen.com/omega
Hawkins PCD, Skillman AG, Warren GL, Ellingson BA, Stahl MT (2010) Conformer generation with OMEGA: algorithm and validation using high quality structures from the protein databank and Cambridge structural database. J Chem Inf Model 50:572–584
Halgren TA (1999) MMFF VI. MMFF94S option for energy minimization studies. J Comp Chem 20:720–729
SZYBKI. OpenEye Scientific Software http://www.eyesopen.com/szybki
Bondi A (1964) van der Waals volumes and radii. J Phys Chem 68:441–451
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA, Nakatsuji H, Caricato M, Li X, Hratchian HP, Izmaylov AF, Bloino J, Zheng G, Sonnenberg JL, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Montgomery JA, Peralta JE, Ogliaro F, Bearpark M, Heyd JJ, Brothers E, Kudin KN, Staroverov VN, Kobayashi R, Normand J, Raghavachari K, Rendell A, Burant JC, Iyengar SS, Tomasi J, Cossi M, Rega N, Millam JM, Klene M, Knox JE, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Martin RL, Morokuma K, Zakrzewski VG, Voth GA, Salvador P, Dannenberg JJ, Dapprich S, Daniels AD, Farkas Ö, Foresman JB, Ortiz JV, Cioslowski J, Fox DJ (2009) Gaussian 09, Revision A.1, Gaussian, Inc., Wallingford
Szabo A, Ostlund NS (1996) Modern quantum chemistry. Introduction to advanced electronic structure theory. Dover, Mineola
Becke AD (1993) Density-functional thermochemistry. III. The role of exact exchange. J Chem Phys 98:5648–5652
Lee C, Yang W, Parr RG (1988) Development of the Colle-Salvetti correlation-energy formula into a functional of the electron density. Phys Rev B 3:785–789
Møller C, Plesset MS (1934) Note on the approximation treatment for many-electron systems. Phys Rev 46:618–622.
Frisch MJ, Head-Gordon M, Pople JA (1990) A direct MP2 gradient method. Chem Phys Lett 166:275–280
Hehre WJ, Pople JA (1971) Self-consistent molecular-orbital methods. IX. Extended Gaussian-type basis for molecular-orbital studies of organic molecules. J Chem Phys 54:724–728
Frisch MJ, Pople JA, Binkley JS (1984) Self-consistent molecular orbital methods. 25. Supplementary functions for Gaussian basis sets. J Chem Phys 80:3265–3269
Spitznagel GW, Clark T, Chandrasekhar J, Schleyer PVR (1982) Stabilization of methyl anions by first-row substituents. The superiority of diffuse function-augmented basis sets for anion calculations. J Comput Chem 3:363–371
Hansch C, Leo A, Hoekman D (1995) Exploring QSAR. American Chemical Society, Washington DC
Kenny PW, Montanari CA, Prokopczyk IM, Sala FA, Sartori GR (2013) Automated molecule editing in molecular design. J Comput Aided Mol Des 27:655–664
Kenny PW, Sadowski J (2005) Structure modification in chemical databases. Methods and principles in medicinal chemistry. In: Oprea T (ed) Chemoinformatics in drug discovery 23:271–285
Leach AG, Jones HD, Cosgrove DA, Kenny PW, Ruston L, MacFaul P, Wood JM, Colclough N, Law B (2006) Matched molecular pairs as a guide in the optimization of pharmaceutical properties; a study of aqueous solubility, plasma protein binding and oral exposure. J Med Chem 49:6672–6682
Hussain J, Rea C (2010) Computationally efficient algorithm to identify matched molecular pairs (MMPs) in large data sets. J Chem Inf Model 50:339–348
Hu X, Hu Y, Vogt M, Stumpfe D, Bajorath J (2012) MMP-Cliffs: systematic identification of activity cliffs on the basis of matched molecular pairs. J Chem Inf Model 52:1138–1145
Dossetter AG, Griffen EJ, Leach AG (2013) Matched molecular pair analysis in drug discovery. Drug Discov Today 18:724–731
Kramer C, Fuchs JE, Whitebread S, Gedeck P, Liedl KR (2014) Matched molecular pair analysis: significance and the impact of experimental uncertainty. J Med Chem 57:3786–3802
JMP version 12.0, SAS Institute, Cary, NC 27513. http://www.jmp.com
Ritchie TJ, Macdonald SJF, Pickett SD (2015) Insights into the impact of N- and O-methylation on aqueous solubility and lipophilicity using matched molecular pair analysis. MedChemComm 6:1787–1797
Mobley DL, Baker JR, Barber AE, Fennell CJ, Dill KA (2008) Charge asymmetries in hydration of polar solutes. J Phys Chem B 112:2405–2414
Mukhopadhyay A, Fenley AT, Tolokh IS, Onufriev AV (2012) Charge hydration asymmetry: the basic principle and how to use it to test and improve water models. J Phys Chem B 116:9776–9783
Reif MM, Hünenberger PH (2016) Origin of asymmetric solvation effects for ions in water and organic solvents investigated using molecular dynamics simulations: the Swain acity-basity scale revisited. J Phys Chem B 120:8485–8517
Ritchie TJ, Macdonald SJF (2014) Physicochemical descriptors of aromatic character and their use in drug discovery. J Med Chem 57:5206–5215
Adler TK, Albert A (1960) Diazaindenes (“azaindoles”). Part I. Ionization constants and spectra. J Chem Soc 1960:1794–1797
Topliss JG (1972) Utilization of operational schemes for analog synthesis in drug design. J Med Chem 15:1006–1011
Brown DG, Gagnon MM, Boström J (2015) Understanding our love affair with p-chlorophenyl: present day implications from historical biases of reagent selection. J Med Chem 58:2390–2405
Leahy DE, Morris JJ, Taylor PJ, Wait AR (1994) Model solvent systems for QSAR. Part IV. The hydrogen bond acceptor behaviour of heterocycles. J Phys Org Chem 7:743–750
Edwards JO, Pearson RG (1962) The factors determining nucleophilic reactivities. J Am Chem Soc 84:16–24
Jorgensen WL, Pranata J (1990) Importance of secondary interactions in triply hydrogen bonded complexes: guanine-cytosine vs uracil-2,6-diaminopyridine. J Am Chem Soc 112:2008–2010
Hann MM, Leach AR, Harper G (2001) Molecular complexity and its impact on the probability of finding leads for drug discovery. J Chem Inf Comp Sci 41:856–864
Johnson ME, Malardier-Jugroot C, Murarka RK, Head-Gordon T (2009) Hydration water dynamics near biological interfaces. J Phys Chem B 113:4082–4092
Bethel PA, Gerhardt S, Jones EV, Kenny PW, Karoutchi GI, Morley AD, Oldham K, Rankine N, Augustin M, Krapp S, Simader H, Steinbacher S (2009) Design of selective cathepsin inhibitors. Bioorg Med Chem Lett 19:4622–4625
Murray CW, Rees DC (2016) Opportunity knocks: organic chemistry for fragment-based drug discovery (FBDD). Angew Chem Int Ed 55:488–492
Acknowledgements
We thank FAPESP (Fundação de Amparo à Pesquisa do Estado de São Paulo; Grant No. 2013/18009-4) and CNPq (Conselho Nacional de Pesquisa; Grant No. 303991/2014-3) for financial support. NMB and IMP thank Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) and JFRR and GRS thank CNPq for scholarships. We are grateful to OpenEye Scientific Software for an academic software license. We also thank the two anonymous reviewers of the manuscript for their constructive and insightful comments.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Borges, N.M., Kenny, P.W., Montanari, C.A. et al. The influence of hydrogen bonding on partition coefficients. J Comput Aided Mol Des 31, 163–181 (2017). https://doi.org/10.1007/s10822-016-0002-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-016-0002-5