Abstract
The thiophene bearing pyrazole derivatives (7a-j) were synthesized and examined for their in vitro cyclooxygenase, 5-lipoxygenase, and tumour inducing factor-α inhibitory activities followed by the in vivo analgesic, anti-inflammatory, and ulcerogenic evaluations. The synthesized series (7a-j) were characterized using 1H NMR, 13C NMR, FT-IR, and mass spectral analysis. Initially, the compounds (7a-j) were evaluated for their in vitro cyclooxygenase, 5-lipoxygenase, and tumour inducing factor-α inhibitory activities and the compound (7f) with two phenyl substituents in the pyrazole ring and chloro substituent in the thiophene ring and the compound (7g) with two phenyl substituents in the pyrazole ring and bromo substituent in the thiophene ring were observed as potent compounds among the series. The compounds (7f and 7g) with effective in vitro potentials were further analyzed for analgesic, anti-inflammatory, and ulcerogenic evaluations. Also, to ascertain the binding affinities of compounds (7a-j), docking assessments were carried out and the ligand (7f) with the highest binding affinity was docked to know the interactions of the ligand with amino acids of target proteins.
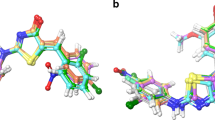
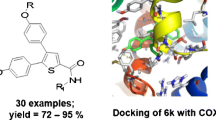
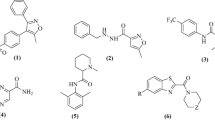
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Among the body's defense mechanisms, inflammation aids in defending the body from pathogens, injuries, and foreign objects. If a local acute inflammation is not monitored, it could create serious systemic or inflammatory problems (Serhan and Petasis 2011; Puttaswamy, Malojiao et al., 2018). The conventional, non-selective non-steroidal anti-inflammatory drugs (NSAIDs) target both cyclooxygenase (COX) enzymes, their potent anti-inflammatory capabilities and apparent adverse effects are indicated by their broad inhibitory characteristics (El-Miligy et al. 2017; Khadriet al. 2022). Various types of inflammatory agents such as prostaglandins, histamine, interleukins, tumour necrosis factor (TNF-α), nitrogen oxide, serotonin, etc. are released to activate the inflammatory action (Abd El-Karim et al. 2021). One of the key phospholipids that are comprised in the membrane of the cell is arachidonic acid, which is formed by the enzyme phospholipase A2 acting on phospholipids. The isoforms of COX include the constitutive COX-1, that are produced in various sites like the kidney and epithelium, the inducible COX-2 is liable for the release of prostaglandins, which contributes to inflammation (Qandeel et al. 2020). Besides that, arachidonic acid is a substrate that facilitates the 5-lipoxygenase (5-LOX) enzyme, responsible for creating pro-inflammatory leukotrienes (LT) (Youssif et al. 2019). A further consequence, of inhibiting COX-1/COX-2 regulated arachidonic acid metabolism, is an upsurge in LT generation through the 5-LOX pathway.
NSAIDs like aspirin and ibuprofen were found to inhibit COX-1, which contributed to the onset of gastrointestinal negative symptoms such as gastric bleeding and ulcers. In this context, the need for the advancement of drugs that block specific COX-2 enzymes, like celecoxib and its equivalents has drawn the attention of researchers globally (Alanazi et al. 2015). Yet persistent administration of selective COX-2 inhibitors triggered an alteration towards metabolism of the arachidonic acid route forming the 5-LOX enzyme, resulting in relatively high levels of LTs that aggravate bronchoconstriction (Omar et al. 2018). In this connection, a substantial attempt was undertaken to develop dual COX/5-LOX inhibitors (Abdelgawad et al. 2018; Chaaban et al. 2018; Youssif et al. 2019; Khadriet al. 2022). Also, TNF-α is a cytokine that serves a vital part in inflammation as well as the rapid increase in cell growth, metabolism, apoptosis, and also functions on immune cells (Rajput and Ware 2016).
The establishment of pharmacologically effective and sophisticated organic compounds has focused a great deal of interest on heterocyclic compounds, which have atoms preferably from two distinctive components as part of their ring (Küçükgüzel and ŞenkardeŞ 2015). In particular, pyrazole derivatives which are a five membered ring comprising two adjacent nitrogen heteroatoms exhibited potent biological applications that are very diverse and widespread including anti-inflammatory (Puttaswamyet al 2018; Shaker et al. 2022), anticancer (Ahmed et al. 2022), antiviral (Wu et al. 2021), antimicrobial (Yan et al. 2021), antioxidant (Alfi et al. 2022; Khadri et al. 2023), antitubercular (Shingare et al. 2022), analgesic (Bekhit et al. 2022), antiAlzheimer (Narayanan et al. 2021) activities. Also, another effective pharmacophore observed in many medications is the thiophene moiety, a five membered sulphur containing heterocyclic compound, that has been associated with several biological applications including anti-inflammatory (Qandeel et al. 2020), anticancer (Megally Abdo et al. 2020; Sumi et al. 2023), antimicrobial (Malani et al. 2016), antioxidant (Sumi et al. 2023), antiviral (Kang et al. 2020), antitubercular (Obu et al. 2021), and analgesic (Kuppusamy et al. 2022) activities. Thus, based on the literature survey and our group research on anti-inflammatory activity, it was intended to integrate these heterocyclic moieties and investigate them nti-inflammatory potential. Initially, the final compounds 5-{4-[3-(2-thiophene)-3-oxoprop-1-enyl] benzamide}-N-phenylpyrazoles (7a-j) were administered for COX-1/2, 5-LOX, and TNF-α inhibitory activities. The compounds (7f and 7g) with effective in vitro potentials were further analyzed for analgesic, anti-inflammatory, and ulcerogenic evaluations. Also, to ascertain the binding affinities of compounds (7a-j), docking assessments were carried out and the ligand (7f) with the highest binding affinity was docked to know the interactions of the ligand with amino acids of target proteins.
Results and discussion
Design based structure
From the literature review, it was found that the pyrazole and thiophene moieties exhibit various biological applications.
Specifically, it was found from the survey, pyrazole moiety displays potent anti-inflammatory activity (Arunkumar et al. 2009; Zabiulla et al. 2019; Priya et al. 2022). The commercially available drugs bearing the pyrazole group like celecoxib, ramifenazone, lonazolac, and rimonabant are reported to demonstrate good anti-inflammatory activity (Mantzanidou et al. 2021). Also, the thiophene molecule exhibited significant anti-inflammatory activity (Nayak et al. 2020; da Cruz et al. 2021). Furthermore, some of the thiophene containing commercially available drugs are zileuton, tenidap, tiaprofenic acid, and tinoridine (da Cruz et al. 2021). Thus, it was intended to integrate pyrazole and thiophene moieties as shown in Fig. 1.
Chemistry
The title compounds 5-{4-[3-(2-thiophene)-3-oxoprop-1-enyl] benzamide}-N-phenyl pyrazoles (7a-j) were generated as revealed in Scheme 1 and the synthesized compounds were characterized by infrared (IR), nuclear magnetic resonance (NMR), and mass spectra. Initially, phenylhydrazine (1) and 3-oxoalkyl/aryl nitriles (2a-c) were refluxed in ethanol and glacial acetic acid to yield substituted 5-aminopyrazoles (3a-c). Taking compound (3a) as a representative example in this series, the formation of IR bands at 1638 and 3229–3495 cm−1 suggested the presence of C=N and –NH2 groups, correspondingly. The 1H NMR data revealed the presence of three aromatic methyl protons with a singlet peak at δ 3.42, two amino protons with a singlet peak at δ 5.31, and six aromatic protons with a multiplet peaks in the range δ 7.27–7.59. Besides, the mass spectral data showed an M + 1 peak at m/z 174 which confirms the formation of the compound (3a). Further, 4-[3-(2-thiophene)-3-oxoprop-1-enyl] benzoic acids (6a-d) were synthesized using substituted 2-acetyl-thiophenes (4a-d) and 4-formyl benzoic acid (5). The IR data corresponded to the representative compound (6a) confirm the absorption bands at 1665, 1723, and 3359–3500 cm−1 for (C=O) of keto, (C=O) of carboxyl, and (OH) of hydroxyl groups of carboxylic acid, respectively. Further, 1H NMR data revealed the presence of a COCH proton with a doublet peak at δ 7.76, seven aromatic protons as multiplet peaks at δ 7.32–7.34, δ 7.97–8.01, and δ 8.37–8.38, C=CH proton as another doublet peak at δ 8.06, and COOH proton as another singlet peak at δ 13.12. In addition, the mass spectral data displayed a stable M + 1 peak at m/z 259. Furthermore, the final compounds 5-{4-[3-(2-thiophene)-3-oxoprop-1-enyl] benzamide}-N-phenylpyrazoles (7a-j) were generated by integrating the compounds (6a-d) and (3a-c). The IR data of compound (7a) as the representative example displayed characteristic bands at 1631, 1672, 1722, and 3130–3300 cm−1 corresponded to the groups like C=N, keto (C=O), amide (C=O), and –NH, respectively. The 1H NMR data revealed the presence of three aromatic methyl protons with a singlet peak at δ 2.54, COCH proton as a doublet peak at δ 7.78, thirteen aromatic protons with multiplet peaks in the range δ 7.33–7.73, δ 7.98–8.03, and δ 8.38–8.39, C=CH proton as another doublet peak at δ 8.08, and amide -NH proton as a broad singlet peak at δ 13.38. Further, the mass spectral data with an M + 1 peak at m/z 414 supported the elucidation of the final compound (7a).
Biological studies
Structure activity relationship
Initially, the plan of synthesis was executed according to the literature's relevance to the anti-inflammatory capability of heterocyclic compounds thiophene and pyrazole. Thus, the final compounds 5-{4-[3-(2-thiophene)-3-oxoprop-1-enyl] benzamide}-N-phenyl pyrazoles (7a-j) were achieved with electron withdrawing and donating groups substituted in both thiophene and pyrazole scaffolds, and subjected for in vitro COX, 5-LOX, and TNF-α inhibitory activities. It was revealed that the compound (7f) with two phenyl substituents in the pyrazole ring and chloro substituent in the thiophene ring and the compound (7 g) with two phenyl substituents in the pyrazole ring and bromo substituent in the thiophene ring displayed potent COX inhibition. Whereas, the compound (7e) with two phenyl substituents in the pyrazole ring and no substituent in the thiophene ring revealed low COX-1 in comparison to COX-2 activity. Thus, the compounds (7f) and (7 g) with electron withdrawing halo groups in the thiophene ring and two phenyl substituents in the pyrazole ring inhibited both COX isoforms, whereas compound (7e) with two phenyl substituents in the pyrazole ring and no substituent in the thiophene ring particularly displayed good COX-2 activity in comparison to COX-1 inhibition. Also, the potent 5-LOX and TNF-α inhibitory activities, among the series, were shown by the compounds (7f) and (7 g). Additionally, the potent compounds (7f) and (7 g) were examined for in vivo analgesic, anti-inflammatory, and ulcerogenic effects.
In vitro COX-1 and COX-2 inhibitory activity
The final compounds (7a-j) were examined for in vitro COX inhibitory activity using the standards celecoxib and indomethacin. From Table 1, it was revealed that the compound (7 g) with two phenyl substituents in the pyrazole ring and bromo substituent in the thiophene ring (IC50 = 9.35 µM for COX-1; IC50 = 1.01 µM for COX-2), compound (7f) with two phenyl substituents in pyrazole ring and chloro substituent in thiophene ring (IC50 = 10.21 µM for COX-1; IC50 = 1.76 µM for COX-2), and compound (7i) with tert-butyl and phenyl substituents in pyrazole ring and bromo substituent in thiophene ring (IC50 = 7.98 µM for COX-1; IC50 = 1.99 µM for COX-2) showed substantial COX inhibiting activities. Besides, the compound (7d) with methyl and phenyl substituents in the pyrazole ring and chloro substituent in the thiophene ring showed similar COX-1 (14.87 µM) and COX-2 (15.06 µM) inhibitions. Surprisingly, the compound (7 h) with tert-butyl and phenyl substituents in the pyrazole ring and no substituent in the thiophene ring also revealed similar COX-1 and COX-2 inhibition with IC50 values of 39.99 and 42.78 µM for COX-1 and COX-2, correspondingly. Further, the compound (7c) with methyl and phenyl substituents in the pyrazole ring and bromo substituent in the thiophene ring exhibited higher COX-1 activity (IC50 = 7.99 µM), than COX-2 activity (IC50 = 26.76 µM). While the compounds (7b) with methyl and phenyl substituents in the pyrazole ring and another methyl substituent in the thiophene ring and (7j) with tert-butyl and phenyl substituents in the pyrazole ring and chloro substituent in the thiophene ring displayed low COX-2 activities with IC50 values > 170 µM.
Interestingly, the compounds (7a) and (7e) with no substituents in the thiophene ring revealed low COX-1 activities with IC50 values > 130 µM and good COX-2 inhibition with IC50 values of 26.45 and 28.78 µM, correspondingly. In comparison to standards celecoxib (IC50 values 14.98 and 0.039 µM for COX-1 and COX-2, respectively) and indomethacin (IC50 values 0.29 and 3.78 µM for COX-1 and COX-2, correspondingly). The compound (7e) displayed a selective index (SI) value of 14.63 with significant COX-2 inhibition and low COX-1 inhibitory activity. The compounds (7f) and (7 g) showed good SI values of 5.80 and 9.25, respectively.
In vitro 5-LOX inhibitory activity
The in vitro 5-LOX inhibitory activity of the final compounds (7a-j) using the standard nordihydroguaiaretic acid (NDGA) was studied.
Interestingly, the compounds (7f) and (7 g) were found to be potent in the 5-LOX inhibitory assay with IC50 values of 0.27 and 0.29 µM, respectively (Table 2). Whereas, the compounds (7b), (7c), and (7 h) displayed moderate activity with IC50 values ranging from 3.06 to 3.89 µM, in comparison to the standard NDGA (IC50 = 0.49 µM). The rest of the compounds showed declined 5-LOX inhibitory activity with IC50 values higher than 4 µM.
In vitro TNF-α inhibitory activity
The in vitro TNF-α inhibitory activity of the final compounds (7a-j) using the standard dexamethasone was studied as revealed in Table 3. The majority of the compounds showed very low activity with IC50 values > 500 µM, while compounds (7a), (7e), and (7 h) were inactive toward inhibition. But the compounds (7f) and (7g) displayed weaker activity with IC50 values of 444.2 and 401.2 µM, respectively, compared to the standard dexamethasone (IC50 = 10.45 µM). The compounds (7f) and (7g) which revealed potent in vitro abilities were further evaluated for analgesic, anti-inflammatory, and ulcerogenic effects.
Analgesic activity
The analgesic activities of the potent compounds (7f) and (7 g) with the standard piroxicam were evaluated utilizing the acetic acid writhing method and the results are revealed in Table 4. The compounds (7f) and (7 g) revealed % inhibitions of 63.18 and 62.56, respectively, which were comparable to the % inhibition of the piroxicam (65.10% inhibition).
Anti-inflammatory activity
The compounds (7f) and (7g), which displayed potential in vitro activities, were examined for anti-inflammatory activity utilizing the carrageenan induced paw edema method using the standards celecoxib and indomethacin (Table 5).
Data are expressed as mean ± SE. Statistical analysis of the data was done using one-way ANOVA. Probability levels of p < 0.05 were considered statistically significant. ANOVA followed by Duncan’s test for multiple group comparisons. Probability levels of p < 0.05 were considered statistically significant.
Ulcerogenicity evaluation
The development of gastrointestinal ulcers due to the administration of potent compounds (7f), (7g), and standards (celecoxib and indomethacin) were evaluated (Table 6). The compounds (7f) displayed mild ulceration, while the compound (7g) exhibited no signs of ulceration compared to the standards celecoxib and indomethacin with significant ulcerogenic effects.
Docking study
Molecular docking simulation
The binding affinities of the final compounds (7a-j) were calculated and the compound (7f) with the highest binding affinity was implicated to understand the docking interactions along with the standards celecoxib and indomethacin against the targets COX-1 and COX-2, standard NDGA against the target 5-LOX, and standard dexamethasone against the target TNF-α.
Molecular docking simulation of compounds (7a-j), celecoxib, and indomethacin with COX-1
The compounds (7a-j) along with celecoxib and indomethacin were docked against COX-1. Among them, compound (7f) displayed the highest binding affinity (− 9.4 kcal/mol) by interacting with the amino acids of the COX-1 binding site.
Compound (7f) established ten intermolecular interactions, three of them constituted the hydrogen bonds with amino acids Asn515 (2.55 Å), Asn515 (2.76 Å), and Pro514 (3.45 Å), the remaining included the pi-alkyl, pi-pi T shaped, and pi-cation interactions.
The binding affinities of the celecoxib and indomethacin were -7.4 and -7.5 kcal/mol, respectively. Celecoxib established seven intermolecular interactions with the amino acids of the COX-1 binding site, four of which were hydrogen bonds, while indomethacin established six intermolecular interactions, three being hydrogen bonds (Table 7).
The interactions of compound (7f), celecoxib, and indomethacin with the amino acid residues of COX-1 have been represented in Table 8. The 3D and 2D representations of compound (7f), celecoxib, and indomethacin interacting with the amino acids of the COX-1 active site have been depicted in Fig. 2.
3D and 2D representations of compound (7f) and the standards celecoxib and indomethacin interacting with amino acids of COX-1 active site, (A) Active site of COX-1 with compound (7f) and standards-surface and magnified, (B-D) 3D representations of compound (7f) (dark blue), celecoxib (orange) and indomethacin (light blue) interacting with the COX-1 active site amino acid, (E–G) 2D representations of compound (7f) (dark blue), celecoxib (orange) and indomethacin (light blue) interacting with the COX-1 active site amino acids
b) Molecular docking simulation of compounds (7a-j), celecoxib, and indomethacin with COX-2
The compounds (7a-j) along with celecoxib and indomethacin were docked against COX-2. Among them, compound (7f) revealed the highest binding affinity (-11.5 kcal/mol) and was allowed to interact with the amino acids of the COX-2 binding site.
Compound (7f) formed twenty three intermolecular interactions, including two hydrogen bonds with Arg120 (2.60 Å), and Tyr355 (2.57 and 3.85 Å). Other interactions such as alkyl, pi-alkyl, pi-pi T shaped, pi-sulphur, pi-sigma, and amide-pi stacked were also observed during the docking simulation. The binding affinities of celecoxib and indomethacin were − 7.7 and − 7.5 kcal/mol, respectively. Celecoxib interacted with the amino acids at the binding site of COX-2 via ten intermolecular interactions, one among them was hydrogen bonding and the thirteen intermolecular interactions from indomethacin included three hydrogen bonds (Table 9). The interactions of compound (7f), celecoxib, and indomethacin with COX-2 have been represented in Table 10.
The 3D and 2D representations of compound (7f), celecoxib, and indomethacin interacting with amino acids of the active site of COX-2 have been depicted in Fig. 3.
3D and 2D representations of compound (7f) and the standards celecoxib and indomethacin interacting with the amino acids of COX-2 active site, (A) Active site of COX-2 with compounds (7f) and standards-surface and magnified, (B-D) 3D representations of compound (7f) (dark blue), celecoxib (orange) and indomethacin (green) interacting with amino acids of COX-2 active site, (E–G) 2D representations of compound (7f) (dark blue), celecoxib (orange) and indomethacin (green) interacting with amino acids of COX-2 active site
c) Molecular docking simulation of compounds (7a-j) and NDGA with 5-LOX
Compounds (7a-j) along with NDGA were docked against 5-LOX. Among them, compound (7f) exhibited the highest binding affinity (-8.8 kcal/mol) interacting with the amino acids of the COX-2 binding site. Further, compound (7f) revealed ten intermolecular interactions, including one hydrogen bond with Gln363 (2.56 Å). Other interactions such as pi-cation, pi-alkyl, pi-pi T shaped, and pi-sigma were also observed during docking simulation. The binding affinity of the NDGA was -7.3 kcal/mol and interacted with the amino acids of the binding site of 5-LOX by establishing five intermolecular interactions which included one hydrogen bond (Table 11). Interactions of the compound (7f) and NDGA with the amino acid residues of 5-LOX are represented in Table 12. The 3D and 2D representations of compound (7f) and NDGA interacting with the active site of amino acids of 5-LOX have been depicted in Fig. 4.
3D and 2D representations of compound (7f) and standard NDGA interacting with amino acids of 5-LOX active site, (A) Active site of 5-LOX with compounds (7f) and standard-surface and magnified, (B-C) 3D representations of compound (7f) (dark blue) and NDGA (orange) interacting with the active site amino acids of 5-LOX, (D-E) 2D representations of compound (7f) (dark blue) and NDGA (orange) interacting with the 5-LOX active site amino acids
d) Molecular docking simulation of compounds (7a-j) and dexamethasone with TNF-α
The compounds (7a-j) along with dexamethasone were docked against TNF-α. Among them, compound (7f) was recognized to display the highest binding affinity (-8.2 kcal/mol) during the interaction with the amino acids of the binding site of TNF-α. Compound (7f) developed eight intermolecular interactions, two among them constituted the hydrogen bonds with Tyr151 (2.23 Å) and Tyr59 (1.96 Å). Also, pi-sigma, pi-alkyl, and pi-pi stacked interactions were observed during the docking simulation. The binding affinity of the dexamethasone was -6.1 kcal/mol and interacted with the binding site of amino acids of TNF-α by forming two intermolecular interactions, both being hydrogen bonds (Table 13). The interactions of compound (7f) and dexamethasone with TNF-α have been represented in Table 14.
The 3D and 2D representations of compound (7f) and dexamethasone interacting with active site amino acids of TNF-α have been depicted in Fig. 5.
3D and 2D representations of compound (7f) and standard dexamethasone interacting with the amino acids of TNF-α active site, (A) Active site of TNF-α with compound (7f) and standard-surface and magnified, (B-C) 3D representations of compound (7f) (dark blue) and dexamethasone (orange) interacting with the TNF-α active site amino acids, (D-E) 2D representations of compound (7f) (dark blue) and dexamethasone (orange) interacting with the TNF-α active site amino acids
Experimental section
Materials and methods
The reagents and solvents were purchased from Sigma Aldrich Chemicals Private Limited. For monitoring the reactions, thin layer chromatography (TLC) was employed using 0.25 mm Merck 60 F 254 silica plates and various solvent systems, further, the TLC plate was visualized using UV light. The compounds were purified using Biotage Isolera flash chromatography. The Chemi line melting point apparatus was used to record the melting points of the compounds. The Infrared (KBr) spectra were obtained using Agilent Cary 630 FT-IR spectrophotometer and elemental analysis results were found to be within 0.6% of the calculated value. The NMR analysis was performed using Agilent VNMRS-400 MHz-NMR spectrophotometer in solvents deuterated dimethyl sulphoxide (DMSO). Further, mass spectral analysis was carried out using a VG70-70H spectrometer.
Chemistry: Plan of the synthesis
The final compounds (7a-j) were synthesized via a synthetic route as illustrated in Scheme 1. The phenylhydrazine (1) and substituted 3-oxoalkyl/aryl nitriles (2a-c) were refluxed with ethanol and glacial acetic acid to yield substituted 5-aminopyrazoles (3a-c). Later, 2-acetyl-thiophenes (4a-d) and 4-formyl benzoic acid (5) were stirred with potassium hydroxide and ethanol to yield 4-[3-(2-thiophene)-3-oxoprop-1-enyl] benzoic acids (6a-d). Furthermore, the compounds (6a-d) and (3a-c) were treated with N,N,N',N'-tetramethyl-O-(benzotriazol-1-yl) uranium tetrafluoroborate (TBTU) in DMSO along with 2,6-lutidine to furnish the final compounds 5-{4-[3-(2-thiophene)-3-oxoprop-1-enyl] benzamide}-N-phenylpyrazoles (7a-j).
General procedure for the synthesis of substituted 5-aminopyrazoles (3a-c):
The mixture of phenylhydrazine (1, 0.001 mol) and substituted 3-oxoalkyl/aryl nitriles (2a-c, 0.001 mol) was refluxed for 6 h in the presence of ethanol and glacial acetic acid. The reaction was monitored by TLC, utilizing ethyl acetate and hexane (1:9). Then, the contents were neutralized with 10% sodium carbonate solution and poured on crushed ice to get a solid mass. Later, the solid mass was filtered, washed with distilled water (3 × 20 ml), and followed by recrystallization with ethanol, to achieve the compounds (3a-c).
3-Methyl-N-phenyl-5-aminopyrazole (3a): Yield: 91%. Mp. 110–112 ℃. IR (KBr, cm−1): 1638 (C=N), 3229–3495 (NH2). 1H NMR (CDCl3): δ 2.31 (s, 3H, CH3), 5.31 (s, 2H, NH2), 7.27–7.59 (m, 6H, Ar–H). LC–MS m/z 174 (M + 1). Anal. Calcd. for C10H11N3 (173): C, 69.34; H, 6.40; N, 24.26. Found: C, 69.28; H, 6.32; N, 24.18%.
3-Phenyl-N-phenyl-5-aminopyrazole (3b): Yield: 89%. Mp. 115–117 ℃. IR (KBr, cm−1): 1641 (C=N), 3290–3485 (NH2). 1H NMR (CDCl3): δ 6.32 (s, 2H, NH2), 7.19–8.11, (m, 11H, Ar–H). LC–MS m/z 236 (M + 1). Anal. Calcd. for C15H13N3 (235): C, 76.57; H, 5.57; N, 17.86. Found: C, 76.51; H, 5.50; N, 17.79%.
3-(Tert-butyl)-N-phenyl-5-aminopyrazole (3c): Yield: 91%. Mp. 122–124 ℃. IR (KBr, cm−1): 1645 (C=N), 3300–3496 (NH2). 1H NMR (CDCl3): δ 1.27 (s, 9H, 3-CH3), 5.78 (s, 2H, NH2), 6.98–7.51 (m, 6H, Ar–H). LC–MS m/z 216 (M + 1). Anal. Calcd. for C13H17N3 (215): C, 72.52; H, 7.96; N, 19.52. Found: C, 72.45; H, 7.89; N, 19.48%.
General procedure for the synthesis of substituted 4-[3-(2-thiophene)-3-oxoprop-1-enyl] benzoic acids (6a-d):
The mixture of substituted 2-acetyl-thiophenes (4a-d, 0.001 mol) and 4-formyl benzoic acid (5, 0.001 mol) in potassium hydroxide (0.0005 mol) and ethanol (15 ml) was stirred for 4 h at room temperature. After monitoring the reaction by TLC (ethyl acetate and hexane, in the ratio 1:9), dilute hydrochloric acid was added in cold condition for neutralization, consequently, a solid mass was formed. It was filtered, washed with distilled water (3 × 20 ml), and followed by the recrystallization utilizing isopropyl alcohol to attain the compounds (6a-d) in pure form.
4-[3-(2-Thiophene)-3-oxoprop-1-enyl] benzoic acid (6a): Yield: 89%. Mp. 118–120 ℃. IR (KBr, cm−1): 1665 (C=O), 1723 (acid, C=O), 3359–3500 (OH). 1H NMR (DMSO): δ 7.32–7.34 (m, 1H, Ar-H), 7.76 (d, 1H, COCH), 7.97–8.01 (m, 5H, Ar-H), 8.06 (d, 1H, C=CH), 8.37–8.38 (m, 1H, Ar-H), 13.12 (s, 1H, COOH). LC–MS m/z 259 (M + 1). Anal. Calcd. for C14H10O3S (258): C, 65.10; H, 3.90; O, 18.58. Found: C, 65.02; H, 3.84; O, 18.51%.
4-[3-(5-Bromo-2-thiophene)-3-oxoprop-1-enyl] benzoic acid (6b): Yield: 90%. Mp. 137–139 ℃. IR (KBr, cm−1): 1674 (C=O), 1725 (acid, C=O), 3300–3500 (OH). 1H NMR (DMSO): δ 7.25–7.27 (m, 1H, Ar-H), 7.32 (d, 1H, COCH), 7.54–7.67 (m, 4H, Ar–H), 7.66 (d, 1H, C=CH), 8.32–8.35 (m, 1H, Ar-H), 13.10 (s, 1H, COOH). LC–MS m/z 335 (M +), 337 (M + 2). Anal. Calcd. for C14H9BrO3S (335): C, 49.87; H, 2.69; O, 14.23. Found: C, 49.81; H, 2.63; O, 14.19%.
4-[3-(5-Chloro-2-thiophene)-3-oxoprop-1-enyl] benzoic acid (6c): Yield: 92%. Mp. 123–125 ℃. IR (KBr, cm−1): 1672 (C=O), 1725 (acid, C=O), 3361–3500 (OH). 1H NMR (DMSO): δ 7.28–7.30 (m, 1H, Ar-H), 7.38 (d, 1H, COCH), 7.51–7.64 (m, 4H, Ar–H), 7.70 (d, 1H, C=CH), 8.37–8.39 (m, 1H, Ar-H), 12.81 (s, 1H, COOH). LC–MS m/z 292 (M +), 294 (M + 2). Anal. Calcd. for C14H9ClO3S (292): C, 57.44; H, 3.10; O, 16.40. Found: C, 57.37; H, 3.03; O, 16.33%.
4-[3-(5-Methyl-2-thiophene)-3-oxoprop-1-enyl] benzoic acid (6d): Yield: 91%. Mp. 117–119 ℃. IR (KBr, cm−1): 1670 (C=O), 1723 (acid, C=O), 3355–3500 (OH). 1H NMR (DMSO): δ 2.32 (s, 3H, Ar-CH3), 7.25–7.27 (m, 1H, Ar-H), 7.42 (d, 1H, COCH), 7.61–7.71 (m, 4H, Ar–H), 7.74 (d, 1H, C=CH), 8.17–8.19 (m, 1H, Ar-H), 12.85 (s, 1H, COOH). LC–MS m/z 273 (M + 1). Anal. Calcd. for C15H12O3S (272): C, 66.16; H, 4.44; O, 17.63. Found: C, 66.09; H, 4.38; O, 17.58%.
General procedure for the synthesis of substituted 5-{4-[3-(2-thiophene)-3-oxoprop-1-enyl] benzamide}-N-phenylpyrazoles (7a-j)
The reaction mixture of 4-[3-(2-thiophene)-3-oxoprop-1-enyl] benzoic acids (6a-d, 0.001 mol) and substituted 5-aminopyrazoles (3a-c, 0.001 mol) in DMSO was stirred for 30 min in a round bottom flask by adding 2–3 drops of 2,6-lutidine. Then, the reaction temperature was lowered to 0–5 ℃ followed by the addition of TBTU (0.002 mol) and stirred overnight. Later, the reaction contents were drained on crushed ice after the reaction was completed. The consequent solid was filtered, washed with distilled water (3 × 20 ml), followed by diethyl ether (3 × 20 ml), and recrystallized with ethanol to achieve the final compounds (7a-j).
5-{4-[3-(2-Thiophene)-3-oxoprop-1-enyl] benzamide}-3-methyl-N-phenylpyrazole (7a): Yield: 85%. Mp. 134–136 ℃. IR (KBr, cm−1): 1631 (C=N), 1672 (C=O), 1722 (amide, C=O), 3130–3300 (NH). 1H NMR (DMSO): δ 2.54 (s, 3H, Ar-CH3), 7.33–7.73 (m, 5H, Ar-H), 7.78 (d, 1H, COCH), 7.98–8.03 (m, 7H, Ar–H), 8.08 (d, 1H, C=CH), 8.38–8.39 (m, 1H, Ar-H), 13.38 (bs, 1H, NH). 13C NMR (DMSO): δ 18.09 (1C, CH3-Ar), 111.76 (1C-Ar), 112.48 (1C-Ar), 115.21 (2C-Ar), 115.61 (1C-Ar), 120.90 (1C, CHCO), 124.56 (2C-Ar), 125.58 (1C-Ar), 126.45 (2C-Ar), 128.72 (2C-Ar), 129.64 (1C-Ar), 131.21 (1C-Ar), 141.17 (1C-Ar), 143.01 (1C-Ar), 147.78 (1C, CH-Ar), 150.86 (1C-Ar), 151.19 (1C-Ar), 151.31 (1C-Ar), 164.54 (1C, CONH), 182.74 (1C, CO). LC–MS m/z 414 (M + 1). Anal. Calcd. for C24H19N3O2S (413): C, 69.71; H, 4.63; N, 10.16. Found: C, 69.70; H, 4.65; N, 10.15%.
5-{4-[3-(5-Methyl-2-thiophene)-3-oxoprop-1-enyl] benzamide}-3-methyl-N-phenyl
Pyrazole (7b): Yield: 89%. Mp. 111–113 ℃. IR (KBr, cm−1): 1632 (C=N), 1675 (C=O), 1725 (amide, C=O), 3125–3295 (NH). 1H NMR (DMSO): δ 2.38 (s, 3H, Ar-CH3), 2.54 (s, 3H, Ar-CH3), 6.97–7.08 (m, 2H, Ar-H), 7.35 (d, 1H, COCH), 7.41–7.66 (m, 8H, Ar–H), 7.68 (d, 1H, C=CH), 7.99–8.01 (m, 2H, Ar-H),12.88 (bs, 1H, NH). 13C NMR (DMSO): δ 14.94 (1C, CH3-Ar), 16.44 (1C, CH3-Ar), 110.08 (C-Ar), 119.62 (1C, CHCO), 124.53 (2C-Ar), 124.98 (2C-Ar), 127.81 (1C-Ar), 129.42 (2C-Ar), 129.49 (2C-Ar), 137.32 (1C-Ar), 132.25 (1C-Ar), 134.62 (1C-Ar), 136.47 (1C-Ar), 139.10 (1C-Ar), 140.15 (1C-Ar), 142.20 (1C-Ar), 143.28 (1C-Ar), 145.78 (1C, CH-Ar), 152.82 (1C-Ar), 167.11 (1C, CONH), 182.28 (1C, CO). LC–MS m/z 428 (M + 1). Anal. Calcd. for C25H21N3O2S (427): C, 70.24; H, 4.95; N, 9.83. Found: C, 70.28; H, 4.91; N, 9.85%.
5-{4-[3-(5-Bromo-2-thiophene)-3-oxoprop-1-enyl] benzamide}-3-methyl-N-phenyl
Pyrazole (7c): Yield: 88%. Mp. 109–111 ℃. IR (KBr, cm−1): 1635 (C=N), 1676 (C=O), 1721 (amide, C=O), 3135–3319 (NH). 1H NMR (DMSO): δ 2.48 (s, 3H, Ar-CH3), 7.02–7.09 (m, 2H, Ar-H), 7.23 (d, 1H, COCH), 7.32–7.59 (m, 8H, Ar–H), 7.61 (d, 1H, C=CH), 7.81–7.84 (m, 2H, Ar-H), 13.21 (bs, 1H, NH). 13C NMR (DMSO): δ 14.94 (1C, CH3-Ar), 110.08 (1C-Ar), 120.62 (1C, CHCO), 124.12 (2C-Ar), 124.53 (2C-Ar), 125.98 (1C-Ar), 127.81 (1C-Ar), 129.42 (2C-Ar), 129.49 (2C-Ar), 132.82 (1C-Ar), 134.62 (1C-Ar), 136.47 (1C-Ar), 138.52 (1C-Ar), 139.10 (1C-Ar), 142.20 (1C-Ar), 143.28 (1C-Ar), 145.78 (1C, CH-Ar), 147.15 (1C-Ar), 167.71 (1C, CONH), 180.21 (1C, CO). LC–MS m/z 491 (M +), 493 (M + 2). Anal. Calcd. for C24H18BrN3O2S (491): C, 58.54; H, 3.68; N, 8.53. Found: C, 58.57; H, 3.69; N, 8.55%.
5-{4-[3-(5-Chloro-2-thiophene)-3-oxoprop-1-enyl] benzamide}-3-methyl-N-phenyl
Pyrazole (7d): Yield: 91%. Mp. 127–129 ℃. IR (KBr, cm−1): 1641 (C=N), 1675 (C=O), 1720 (amide, C=O), 3133–3314 (NH). 1H NMR (DMSO): δ 2.35 (s, 3H, Ar-CH3), 7.14–7.18 (m, 2H, Ar-H), 7.34 (d, 1H, COCH), 7.37–7.55 (m, 8H, Ar–H), 7.59 (d, 1H, C=CH), 7.79–7.81 (m, 2H, Ar-H), 13.21 (bs, 1H, NH). 13C NMR (DMSO): δ 14.94 (1C, CH3-Ar), 115.08 (1C-Ar), 121.59 (1C, CHCO), 124.53 (2C-Ar), 124.98 (2C-Ar), 127.81 (2C-Ar), 129.42 (2C-Ar), 129.82 (1C-Ar), 134.62 (1C-Ar), 134.92 (1C-Ar), 136.47 (1C-Ar), 139.10 (1C-Ar), 140.74 (1C-Ar), 141.24 (1C-Ar), 142.20 (1C-Ar), 143.28 (1C-Ar), 145.78 (1C, CH-Ar), 145.91 (1C-Ar), 167.71 (1C, CONH), 180.21 (1C, CO). LC–MS m/z 447 (M +), 449 (M + 2). Anal. Calcd. for C24H18ClN3O2S (447): C, 64.35; H, 4.05; N, 9.38. Found: C, 64.32; H, 4.08; N, 9.40%.
5-{4-[3-(2-Thiophene)-3-oxoprop-1-enyl] benzamide}-3-phenyl-N-phenylpyrazole (7e): Yield: 86%. Mp. 125–127 ℃. IR (KBr, cm−1): 1640 (C=N), 1672 (C=O), 1720 (amide, C=O), 3128–3311 (NH). 1H NMR (DMSO): δ 7.09–7.12 (m, 3H, Ar-H), 7.17 (d, 1H, COCH), 7.37–7.48 (m, 8H, Ar–H), 7.66 (d, 1H, C=CH), 7.64–7.87 (m, 7H, Ar-H), 12.58 (bs, 1H, NH). 13C NMR (DMSO): δ 110.08 (1C-Ar), 119.62 (1C-Ar), 124.53 (1C, CHCO), 124.98 (1C-Ar), 127.48 (2C-Ar), 127.81 (2C-Ar), 128.65 (1C-Ar), 128.98 (2C-Ar), 129.32 (2C-Ar), 129.42 (1C-Ar), 129.49 (1C-Ar), 130.19 (1C-Ar), 132.85 (1C-Ar), 133.10 (2C-Ar), 134.62 (1C-Ar), 136.47 (1C-Ar), 139.12 (2C-Ar), 139.75 (1C-Ar), 142.15 (1C-Ar), 143.28 (1C-Ar), 145.78 (1C, CH-Ar), 167.31 (1C, CONH), 181.98 (1C, CO). LC–MS m/z 476 (M + 1). Anal. Calcd. for C29H21N3O2S (475): C, 73.24; H, 4.45; N, 8.84. Found: C, 73.27; H, 4.46; N, 8.87%.
5-{4-[3-(5-Chloro-2-thiophene)-3-oxoprop-1-enyl] benzamide}-3-phenyl-N-phenyl
Pyrazole (7f): Yield: 90%. Mp. 111–113 ℃. IR (KBr, cm−1): 1641 (C=N), 1671 (C=O), 1726 (amide, C=O), 3114–3311 (NH). 1H NMR (DMSO): δ 7.01–7.05 (m, 3H, Ar-H), 7.30 (d, 1H, COCH), 7.32–7.68 (m, 7H, Ar–H), 7.68 (d, 1H, C=CH),7.71–7.98 (m, 7H, Ar-H), 12.58 (bs, 1H, NH). 13C NMR (DMSO): δ 110.08 (1C-Ar), 119.62 (1C-Ar), 124.53 (1C-Ar), 124.98 (1C, CHCO), 127.51 (2C-Ar), 127.81 (2C-Ar), 128.75 (1C-Ar), 129.29 (2C-Ar), 129.42 (2C-Ar), 129.49 (1C-Ar), 128.82 (1C-Ar), 133.25 (2C-Ar), 134.62 (1C-Ar), 135.88 (1C-Ar), 136.47 (2C-Ar), 138.12 (1C-Ar), 139.75 (1C-Ar), 142.20 (1C-Ar), 143.28 (1C-Ar), 145.78 (1C-Ar), 145.95 (1C, CH-Ar), 167.31 (1C, CONH), 181.98 (1C, CO). LC–MS m/z 509 (M +), 511 (M + 2). Anal. Calcd. for C29H20ClN3O2S (509): C, 68.30; H, 3.95; N, 8.24. Found: C, 68.32; H, 3.97; N, 8.25%.
5-{4-[3-(5-Bromo-2-thiophene)-3-oxoprop-1-enyl] benzamide}-3-phenyl-N-phenyl
Pyrazole (7 g): Yield: 89%. Mp. 121–123 ℃. IR (KBr, cm−1): 1639 (C=N), 1674 (C=O), 1722 (amide, C=O), 3126–33,240 (NH). 1H NMR (DMSO): δ 6.97–7.04 (m, 3H, Ar-H), 7.29 (d, 1H, COCH), 7.35–7.63 (m, 7H, Ar–H), 7.66 (d, 1H, C=CH), 7.71–7.94 (m, 7H, Ar-H), 11.98 (bs, 1H, NH). 13C NMR (DMSO): δ 110.08 (1C-Ar), 119.62 (1C-Ar), 124.42 (1C-Ar), 124.53 (1C, CHCO), 124.98 (2C-Ar), 127.55 (1C-Ar), 127.81 (2C-Ar), 128.80 (2C-Ar), 129.21 (1C-Ar), 129.42 (2C-Ar), 129.49 (1C-Ar), 132.18 (1C-Ar), 133.11 (2C-Ar), 134.62 (C-Ar), 137.47 (2C-Ar), 139.12 (1C-Ar), 138.85 (1C-Ar), 142.20 (1C-Ar), 143.28 (1C-Ar), 145.78 (1C, CH-Ar), 147.73 (1C-Ar), 167.31 (1C, CONH), 181.98 (1C, CO). LC–MS m/z 553 (M +), 555 (M + 2). Anal. Calcd. for C29H20BrN3O2S (553): C, 62.82; H, 3.64; N, 7.58. Found: C, 62.84; H, 3.65; N, 7.56%.
5-{4-[3-(2-Thiophene)-3-oxoprop-1-enyl] benzamide}-3-(tert-butyl)-N-phenylpyrazole (7 h): Yield: 87%. Mp. 114–116 ℃. IR (KBr, cm−1): 1631 (C=N), 1672 (C=O), 1722 (amide, C=O), 3129–3302 (NH). 1H NMR (DMSO): δ 1.32 (s, 9H, -CH3), 7.10–7.14 (m, 2H, Ar-H), 7.21 (d, 1H, COCH), 7.25–7.68 (m, 7H, Ar–H), 7.68 (d, 1H, C=CH), 7.71–7.82 (m, 4H, Ar-H), 12.58 (bs, 1H, NH). 13C NMR (DMSO): δ 29.65 (3C, CH3), 31.51 (1C, CH), 110.08 (1C-Ar), 119.62 (1C-Ar), 124.53 (1C, CHCO), 124.98 (2C-Ar), 127.81 (2C-Ar), 128.98 (2C-Ar), 129.42 (1C-Ar), 129.49 (2C-Ar), 130.17 (1C-Ar), 132.81 (1C-Ar), 134.62 (1C-Ar), 136.47 (1C-Ar), 139.12 (1C-Ar), 139.48 (1C-Ar), 142.20 (1C-Ar), 145.78 (1C, CH-Ar), 161.38 (1C-Ar), 167.31 (1C, CONH), 181.98 (1C, CO). LC–MS m/z 456 (M + 1). Anal. Calcd. for C27H25N3O2S (455): C, 71.18; H, 5.53; N, 9.22. Found: C, 71.15; H, 5.54; N, 9.21%.
5-{4-[3-(5-Bromo-2-thiophene)-3-oxoprop-1-enyl] benzamide}-3-(tert-butyl)-N-phenyl
Pyrazole (7i): Yield: 89%. Mp. 132–133 ℃. IR (KBr, cm−1): 1632 (C=N), 1675 (C=O), 1722 (amide, C=O), 3126–3305 (NH). 1H NMR (DMSO): δ 1.32 (s, 9H, 3-CH3), 7.15–7.18 (m, 2H, Ar-H), 7.21 (d, 1H, COCH), 7.27–7.59 (m, 6H, Ar–H), 7.64 (d, 1H, C=CH), 7.65–7.85 (m, 4H, Ar-H), 12.35 (bs, 1H, NH). 13C NMR (DMSO): δ 30.15 (3C, CH3), 31.51 (1C, CH), 110.08 (1C-Ar), 119.62 (1C-Ar), 124.28 (1C-Ar),124.53 (1C, CHCO), 124.98 (2C-Ar), 127.81 (2C-Ar), 129.42 (2C-Ar), 129.49 (2C-Ar), 132.21 (1C-Ar), 134.62 (1C-Ar), 136.47 (C-Ar), 138.94 (1C-Ar), 139.12 (1C-Ar), 142.20 (1C-Ar), 145.78 (1C, CH-Ar), 147.84 (1C-Ar), 161.28 (1C-Ar), 167.31 (1C, CONH), 181.98 (1C, CO). LC–MS m/z 533 (M +), 535 (M + 2). Anal. Calcd. for C27H25BrN3O2S (533): C, 60.68; H, 4.53; N, 7.86. Found: C, 60.67; H, 4.55; N, 7.87%.
5-{4-[3-(5-Chloro-2-thiophene)-3-oxoprop-1-enyl] benzamide}-3-(tert-butyl)-N-phenyl
Pyrazole (7j): Yield: 91%. Mp. 111–113 ℃. IR (KBr, cm−1): 1630 (C=N), 1674 (C=O), 1722 (amide, C=O), 3121–3298 (NH). 1H NMR (DMSO): δ 1.32 (s, 9H, CH3), 7.16–7.20 (m, 2H, Ar-H), 7.23 (d, 1H, COCH), 7.31–7.61 (m, 6H, Ar–H), 7.64 (d, 1H, C=CH), 7.75–8.00 (m, 4H, Ar-H), 12.27 (bs, 1H, NH). 13C NMR (DMSO): δ 30.32 (3C, CH3), 31.51 (1C, CH), 110.08 (1C-Ar), 119.62 (1C-Ar), 124.16 (1C, CHCO), 124.98 (2C-Ar), 127.81 (2C-Ar), 128.42 (2C-Ar), 129.49 (2C-Ar), 129.78 (1C-Ar), 134.62 (1C-Ar), 135.68 (1C-Ar), 136.47 (1C-Ar), 139.12 (1C-Ar), 140.54 (1C-Ar), 142.20 (1C-Ar), 145.78 (1C, CH-Ar), 145.98 (1C, C-Ar), 164.28 (1C, C-Ar), 167.31 (1C, CONH), 181.98 (1C, CO). LC–MS m/z 489 (M +), 491 (M + 2). Anal. Calcd. for C27H24ClN3O2S (489): C, 66.18; H, 4.94; N, 8.58. Found: C, 66.15; H, 4.93; N, 8.59%.
Biological studies
In vitro COX-1 and COX-2 inhibitory activity
The potential of the synthesized compounds (7a-j) to inhibit COX-1 and COX-2 enzymes was studied. From the reported method, the assays were performed using the Cayman colorimetric COX (ovine) inhibitor screening assay kit (Catalogue No. 760111) provided by Cayman Chemicals, Ann Arbour, MI, USA (Xie et al. 1991; Blobaum and Marnett 2007).
In vitro 5-LOX inhibitory activity
The capability of compounds (7a-j) to block the human recombinant 5-LOX enzyme was tested. Measuring the value of absorbance at 234 nm using assay buffer composed of 50 mM tris-buffer (pH = 7.5) containing EDTA, calcium chloride, and adenosine triphosphate at a volume of 2 mM each, newly produced conjugated dienes like hydroperoxyeicosatetraenoic acid (HPETE) and hydroxyeicosatetraenoic acid (HETE) catalyzed by lipoxygenase was identified. The volume of each quartz cuvette was 2 ml, and the absorbance was constantly recorded for 300 s at 234 nm for each enzyme process. Different concentrations of test compounds (7a-j) were utilized to measure the inhibitor activity and 20 mM linoleic acid was used to initiate the process.
In vitro TNF-α inhibitory activity
The capacity of the compounds (7a-j) for the TNF-α inhibition (IC50 value, μM) was evaluated. The six-well culture dishes with RAW cells were placed in Dulbecco's Modified Eagle medium consisting of 10% fetal bovine serum at a cell density ranging from 1.5 × 105 to 2 × 105 cells/ml. The cells were exposed to test compound concentrations with 1 μg/ml of lipopolysaccharide, after 24 h; they were incubated (37 ℃) for 4 h with 5% of carbon dioxide. Following the incubation, the cell supernatant was gathered, and separated by centrifugation. Using a standard ELISA kit (Mouse TNF-alpha ELISA, Cat No. ELMTNF-1, Ray Biotech, USA), the TNF-α levels in the cell supernatants were calculated as per the standard kit methodology.
Analgesic activity
The compounds (7f) and (7 g) that displayed potential in vitro activity were evaluated for analgesic activity along with the standard piroxicam via acetic acid induced writhing approach (Collier et al. 1968). Adult male albino mice weighing about 20–25 g were procured from the animal house of Farooquia Pharmacy College, Mysore, Karnataka, India, and utilized for this study. For one week prior to the trials, the rats were housed in a typical laboratory environment with standard lighting and temperature. A day before the drug regimen all of the rats were given an intraperitoneal injection with a dosage of 0.20–0.25 ml of 0.01% acetic acid solution to test their sensitivity. Animals for analgesic experiments were chosen based on their positive writhing reaction within 30 min of administration of the injection. Suspension of the test compounds (7f), (7 g), and piroxicam in 2% tween 80 was prepared. The animals were bifurcated into four groups consisting of five in each group. Later, the 2% tween 80 suspensions of test compounds (7f), (7 g), and piroxicam were tested orally on the standard groups at the dosage of 0.028 mM per kg body weight, whereas, the control group was tested orally with 2% tween 80 alone. The intraperitoneal injection of 0.01% acetic acid solution was given after an hour. After 5 min of injection, the injected animals were observed for 10 min and the frequency of writhing was counted. The percentage protection of analgesic activity was estimated using the equation:
Anti-inflammatory activity
The compounds (7f) and (7 g) that displayed potential in vitro activity were examined for anti-inflammatory activity by carrageenan induced rat paw edema method using the standards celecoxib and indomethacin. The rats were labeled and split into five groups, each with six rats. The initial group (control) was given 1 ml of saline. The preceding two groups received 10 mg/kg of the compounds (7f) and (7 g), respectively. The final two groups (positive control groups) each were given 10 mg/kg of the standards celecoxib and indomethacin, correspondingly. After one hour of introducing the compounds and the standards, 1% carrageenan was given to the treated rats. Further, the initial rat paw volume was measured and paw volumes over 1–4 h at a time interval of 1 h were recorded (Table 5). The edema percentage was computed with the equation:
Ulcerogenicity evaluation
Furthermore, the compounds (7f) and (7 g) were studied for ulcerogenic effect with the standards celecoxib and indomethacin. The rats were grouped into five consisting of five rats in each group. The compounds (7f), (7 g), standards, and saline (control) were given orally (10 mg/kg body weight to rats that had been kept starving for 18 h previously). Four hours later, the animals were sacrificed and their stomachs were examined for the prevalence of ulceration and lesions.
Docking study
Molecular dynamics simulation
The X-ray crystallographic structures of COX-1 (PDB ID: 3KK6), COX-2 (PDB ID: 6BL4), 5-LOX (PDB ID: 6N2W), and TNF-α (PDB ID: 2AZ5) were retrieved via Research Collaboratory for Structural Bioinformatics (RCSB) PDB database (https://www.rcsb.org) (Accessed on March 2023), the molecules of the protein were developed using the Auto Dock Tools 1.5.7. In the first instance, the removal of the water and heteroatoms was done, followed by the inclusion of polar hydrogens were accomplished to stabilize the protein structure (Patil et al. 2021; Reshma M Martiz et al. 2022a, b). Using Kollmann-united and Gasteiger charges, the reduction of the protein structure energy was achieved. Post energy minimization, an Auto Dock 4 (AD4) atom type was allocated for all atoms, prior to the final protein structure being obtained in PDBQT format allowing molecular docking simulation and prediction of the binding site. The grid box measuring 40 × 40 × 40 Å3 comprising the binding pocket and active site aspects was established at x = − 30.957491 Å, y = 47.607252 Å, and z = − 6.611637 Å for COX-1, x = − 37.400227 Å, y = − 27.861640 Å, and z = 21.802171 Å for COX-2, x = 35.820614 Å, y = 65.513114 and z = 38.411455 for 5-LOX, and x = -19.409600 Å, y = 74.650750 Å, and z = 33.849550 Å for TNF-α. Phytochemical structures were generated for the docking simulation employing the Auto Dock Tools 1.5.7 for ligand (7f) preparation (Patil et al. 2021), whereas the 3D SDF structures were retrieved from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/) and transformed to PDBQT format, and Kollmann-united and Gasteiger charges were incorporated to decrease the energy. To execute the docking simulation, the ligand (7f) was saved in PDBQT format in the identical directory as the molecules of protein after energy minimization (Patil et al. 2022; Reshma et al. 2022). A command-line program, Auto Dock Vina 1.1.2 was implemented for finalizing the virtual screening of the compounds. It applies the Brayden-Fletcher-Goldfarb-Shanno (BGFS) algorithm to disrupt, allocate ligand (7f) to the target site, and assess the score function associated with every ligand conformation (Patil et al. 2022; Reshma et al. 2022). In contrast to proteins, which were believed to be rigid throughout the docking simulation, ligand (7f) was flexible owing to the huge number of torsions enabled throughout its formation. For ligand molecule (7f), however, 10 degrees were permitted, and the initial binding pose with zero root-mean-square deviation (RMSD) of atomic locations was by far the most realistic. Moreover, it has the most significant binding affinity at all the positions, which suggests that the binding is more effectively achieved. The open-source visualizing GUI program Biovia Discovery Studio Visualizer 2021 was implemented to carry out the visualization of the molecular docking simulation. Considering binding affinity, the total number of intermolecular bonds, and the total number of hydrogen bonds, the level of ligand interaction has been estimated (Gurupadaswamy et al. 2022; Patil et al. 2022; Shivanna et al. 2022).
Binding free energy calculations
The molecular dynamics simulation results for complexes and standards were subjected to binding free energy estimation employing the MM-PBSA method. Further, the MmPbStat.py script and GROMACS 2018.1 input trajectories were used with the g mmpbsa tool for ligand–protein combination to know the binding free energy (Jyothi et al. 2022; Prabhakaran et al., 2022). The g mmpbsa programme employs three factors to estimate the binding free energy: molecular mechanical, polar, and apolar solvation energies. For the computation, trajectories of molecular dynamics during the previous 50 ns were utilized to calculate ΔG having frames of dt 1000.
Conclusion
Nitrogen and sulphur containing heterocyclic compounds have been recognized to be important molecules in medicinal chemistry research. The current study reveals the in vitro COX, 5-LOX, and TNF-α inhibitory activities of the final compounds 5-{4-[3-(2-thiophene)-3-oxoprop-1-enyl] benzamide}-N-phenyl pyrazoles (7a-j) and the potent compounds (7f) and (7 g) from the series were additionally evaluated for in vivo analgesic, anti-inflammatory, and ulcerogenic evaluations. The compound (7f) with two phenyl substituents in the pyrazole ring and chloro substituent in the thiophene ring and the compound (7 g) with two phenyl substituents in the pyrazole ring and bromo substituent in the thiophene ring were revealed to be potent among the series. Also, the binding energies and molecular docking interactions were carried out.
References
Abd El-Karim SS et al (2021) Design, synthesis and molecular docking of new pyrazole-thiazolidinones as potent anti-inflammatory and analgesic agents with TNF-α inhibitory activity. Bioorg Chem 111(January):104827. https://doi.org/10.1016/j.bioorg.2021.104827
Abdelgawad MA et al (2018) Design, synthesis, analgesic, anti-inflammatory activity of novel pyrazolones possessing aminosulfonyl pharmacophore as inhibitors of COX-2/5-LOX enzymes: Histopathological and docking studies. Bioorgan Chem 78:103–114. https://doi.org/10.1016/j.bioorg.2018.03.011
Ahmed AHH et al (2022) Design, synthesis, and molecular docking of novel pyrazole-chalcone analogs of lonazolac as 5-LOX, iNOS and tubulin polymerization inhibitors with potential anticancer and anti-inflammatory activities. Bioorganic Chemistry 129:106171. https://doi.org/10.1016/j.bioorg.2022.106171
Alanazi AM et al (2015) Structure-based design of phthalimide derivatives as potential cyclooxygenase-2 (COX-2) inhibitors: anti-inflammatory and analgesic activities. Eur J Med Chem 92:115–123. https://doi.org/10.1016/j.ejmech.2014.12.039
Alfi AA et al (2022) Molecular modeling and docking studies of new antioxidant pyrazole-thiazole hybrids. J Mol Struct 1267:133582. https://doi.org/10.1016/j.molstruc.2022.133582
Arunkumar S et al (2009) Synthesis and Anti-inflammatory Activity of Some Novel Pyrazole Derivatives of Gallic Acid. E J Chem 6:128586. https://doi.org/10.1155/2009/128586
Bekhit AA et al (2022) Investigation of the anti-inflammatory and analgesic activities of promising pyrazole derivative. Eur J Pharm Sci 168:106080. https://doi.org/10.1016/J.EJPS.2021.106080
Blobaum AL, Marnett LJ (2007) Structural and Functional basis of cyclooxygenase inhibition. J Med Chem 50(7):1425–1441. https://doi.org/10.1021/jm0613166
Chaaban I et al (2018) Synthesis, anti-inflammatory screening, molecular docking, and COX-1,2/-5-LOX inhibition profile of some novel quinoline derivatives. Bioorg Chem 78:220–235. https://doi.org/10.1016/j.bioorg.2018.03.023
Collier HO et al (1968) The abdominal constriction response and its suppression by analgesic drugs in the mouse. Br J Pharmacol Chemother 32(2):295–310. https://doi.org/10.1111/j.1476-5381.1968.tb00973.x
da Cruz RMD et al (2021) Thiophene-based compounds with potential anti-inflammatory activity. Pharmaceuticals 14(7):692. https://doi.org/10.3390/ph14070692
El-Miligy MMM et al (2017) New hybrid molecules combining benzothiophene or benzofuran with rhodanine as dual COX-1/2 and 5-LOX inhibitors: Synthesis, biological evaluation and docking study. Bioorg Chem 72:102–115. https://doi.org/10.1016/j.bioorg.2017.03.012
Gurupadaswamy HD et al (2022) Competent synthesis of biaryl analogs via asymmetric Suzuki-Miyaura cross-coupling for the development of anti-inflammatory and analgesic agents. J Iran Chem Soc 19(6):2421–2436. https://doi.org/10.1007/s13738-021-02460-0
Jyothi M et al (2022) Microwave-assisted synthesis, characterization, docking studies and molecular dynamic of some novel phenyl thiazole analogs as xanthine oxidase inhibitor. J Iran Chem Soc 19(9):3919–3933. https://doi.org/10.1007/s13738-022-02574-z
Kang D et al (2020) Exploring the hydrophobic channel of NNIBP leads to the discovery of novel piperidine-substituted thiophene[3,2-d]pyrimidine derivatives as potent HIV-1 NNRTIs. Acta Pharm Sin B 10(5):878–894. https://doi.org/10.1016/j.apsb.2019.08.013
Khadri MJN, Prashanth T et al (2022) Synthesis, analgesic, anti-inflammatory, COX/5-LOX inhibition, ulcerogenic evaluation, and docking study of benzimidazole bearing indole and benzophenone analogs. J Mol Struct 1259:132741. https://doi.org/10.1016/j.molstruc.2022.132741
Khadri MJN, Khamees HA et al (2022) Synthesis, analgesic, anti-inflammatory, ulcerogenic evaluation, and docking study of (benzoylphenoxy)-N-{5-[2-methylphenyl-6-chlorobenzoxazole]} acetamides as COX/5-LOX inhibitor. J Mol Struct 1272:134240. https://doi.org/10.1016/j.molstruc.2022.134240
Khadri MJN et al (2023) Synthesis and docking studies of pyrazole-benzamide-benzothiazole conjugates as xanthine oxidase inhibitor candidates. Journal of Molecular Structure 1290:135937. https://doi.org/10.1016/j.molstruc.2023.135937
Küçükgüzel G, ŞenkardeŞ S (2015) Recent advances in bioactive pyrazoles. Eur J Med Chem 97(1):786–815. https://doi.org/10.1016/j.ejmech.2014.11.059
Kuppusamy R et al (2022) ‘Benzo[b]thiophene-2-carboxamides as novel opioid receptor agonists with potent analgesic effect and reduced constipation. Eur J Med Chem 243:114728. https://doi.org/10.1016/j.ejmech.2022.114728
Malani K et al (2016) Synthesis, characterization and in silico designing of diethyl-3-methyl-5-(6-methyl-2-thioxo-4-phenyl-1,2,3,4-tetrahydropyrimidine-5-carboxamido) thiophene-2,4-dicarboxylate derivative as anti-proliferative and anti-microbial agents. Bioorg Chem 68:265–274. https://doi.org/10.1016/j.bioorg.2016.09.001
Mantzanidou M, Pontiki E, Hadjipavlou-Litina D (2021) ‘Pyrazoles and pyrazolines as anti-inflammatory agents. Molecules (Basel, Switzerland) 26(11):3439. https://doi.org/10.3390/molecules26113439
Martiz RM et al (2022) Defining the role of isoeugenol from ocimum tenuiflorum against diabetes mellitus-linked alzheimer & disease through network pharmacology and computational methods. Molecules. https://doi.org/10.3390/molecules27082398
Martiz Reshma Mary, Patil SM, Ramu R et al (2022) Discovery of novel benzophenone integrated derivatives as anti-Alzheimer’s agents targeting presenilin-1 and presenilin-2 inhibition: a computational approach. PLoS ONE 17(4):e0265022. https://doi.org/10.1371/journal.pone.0265022
Martiz RM, Patil SM, Thirumalapura Hombegowda D et al (2022) Phyto-Computational intervention of diabetes mellitus at multiple stages using isoeugenol from ocimum tenuiflorum: a combination of pharmacokinetics and molecular modelling approaches. Molecules 27(19):6222. https://doi.org/10.3390/molecules27196222
Megally Abdo NY, Samir EM, Mohareb RM (2020) Synthesis and evaluation of novel 4H-pyrazole and thiophene derivatives derived from chalcone as potential anti-proliferative agents, Pim-1 kinase inhibitors, and pains. J Heterocycl Chem 57(4):1993–2009. https://doi.org/10.1002/jhet.3928
Narayanan SE et al (2021) Design, synthesis and biological evaluation of substituted pyrazoles endowed with brominated 4-methyl 7-hydroxy coumarin as new scaffolds against Alzheimer’s disease. Future J Pharm Sci 7(1):161. https://doi.org/10.1186/s43094-021-00278-4
Nayak SG, Poojary B, Kamat V (2020) Novel pyrazole-clubbed thiophene derivatives via Gewald synthesis as antibacterial and anti-inflammatory agents. Archiv der Pharmazie 353(12):2000103. https://doi.org/10.1002/ardp.202000103
Obu QS et al (2021) Synthesis, spectra (FT-IR, NMR) investigations, DFT study, in silico ADMET and molecular docking analysis of 2-amino-4-(4-aminophenyl)thiophene-3-carbonitrile as a potential anti-tubercular agent. J Mol Struct 1244:130880. https://doi.org/10.1016/j.molstruc.2021.130880
Omar YM, Abdu-Allah HHM, Abdel-Moty SG (2018) Synthesis, biological evaluation and docking study of 1,3,4-thiadiazole-thiazolidinone hybrids as anti-inflammatory agents with dual inhibition of COX-2 and 15-LOX. Bioorg Chem 80:461–471. https://doi.org/10.1016/j.bioorg.2018.06.036
Patil SM et al (2021) Comparative molecular docking and simulation analysis of molnupiravir and remdesivir with SARS-CoV-2 RNA dependent RNA polymerase (RdRp). Bioinformation 17(11):932–939. https://doi.org/10.6026/97320630017932
Patil SM, Martiz Reshma M et al (2022) Discovery of novel coumarin derivatives as potential dual inhibitors against α-glucosidase and α-amylase for the management of post-prandial hyperglycemia via molecular modelling approaches. Molecules. https://doi.org/10.3390/molecules27123888
Patil SM, Martiz Reshma Mary et al (2022) Evaluation of flavonoids from banana pseudostem and flower (quercetin and catechin) as potent inhibitors of α-glucosidase: an in silico perspective. J Biomol Struct Dyn 40(23):12491–12505. https://doi.org/10.1080/07391102.2021.1971561
Patil SM, Al-Mutairi KA et al (2022) Pharmacoinformatics based screening discovers swertianolin from Lavandula angustifolia as a novel neuromodulator targeting epilepsy, depression, and anxiety. South Afr J Bot 149:712–730. https://doi.org/10.1016/j.sajb.2022.06.054
Prabhakaran S et al (2022) One-pot three-component synthesis of novel phenyl-pyrano-thiazol-2-one derivatives and their anti-diabetic activity studies. Results in Chemistry 4:100439
Priya D et al (2022) Structural insights into pyrazoles as agents against anti-inflammatory and related disorders. ChemistrySelect 7(5):e202104429. https://doi.org/10.1002/slct.202104429
Puttaswamy N, Rekha ND et al (2018) Anti-inflammatory and antioxidant activity of salicylic acid conjugated dihydropyrazoline analogues. J Appl Pharm Sci 8(2):060–064. https://doi.org/10.7324/JAPS.2018.8209
Puttaswamy N, Malojiao VH et al (2018) Synthesis and amelioration of inflammatory paw edema by novel benzophenone appended oxadiazole derivatives by exhibiting cyclooxygenase-2 antagonist activity. Biomed Pharmacother 103:1446–1455. https://doi.org/10.1016/j.biopha.2018.04.167
Qandeel NA et al (2020) Synthesis, in vivo anti-inflammatory, COX-1/COX-2 and 5-LOX inhibitory activities of new 2,3,4-trisubstituted thiophene derivatives. Bioorgan Chem 102:103890. https://doi.org/10.1016/j.bioorg.2020.103890
Rajput A, Ware CF (2016) ‘Tumor necrosis factor signaling pathways’, Encyclopedia of Cell Biology. Edited by R.A. Bradshaw and P.D.B.T.-E. of C.B. Stahl 3:354–363. https://doi.org/10.1016/B978-0-12-394447-4.30048-7
Serhan CN, Petasis NA (2011) Resolvins and protectins in inflammation resolution. Chem Rev 111(10):5922–5943. https://doi.org/10.1021/cr100396c
Shaker AMM et al (2022) Novel 1,3-diaryl pyrazole derivatives bearing methylsulfonyl moiety: design, synthesis, molecular docking and dynamics, with dual activities as anti-inflammatory and anticancer agents through selectively targeting COX-2. Bioorganic Chemistry 129:106143. https://doi.org/10.1016/j.bioorg.2022.106143
Shingare R et al (2022) Docking stimulations and primary assessment of newly synthesized benzene sulfonamide pyrazole oxadiazole derivatives as potential antimicrobial and antitubercular agents. Polycycl Arom Compd 43:1799–1811. https://doi.org/10.1080/10406638.2022.2036771
Shivanna C et al (2022) Synthesis, Characterization, Hirshfeld Surface Analysis, Crystal Structure and Molecular Modeling Studies of 1-(4-(Methoxy(phenyl)methyl)-2-methylphenoxy)butan-2-one Derivative as a Novel α-Glucosidase Inhibitor. Crystals. https://doi.org/10.3390/cryst12070960
Sumi M, Nevaditha NT, Sindhu Kumari B (2023) Synthesis, structural evaluation, antioxidant, DNA cleavage, anticancer activities and molecular docking study of metal complexes of 2-amino thiophene derivative. J Mol Struct 1272:134091. https://doi.org/10.1016/j.molstruc.2022.134091
Wu Z et al (2021) In vivo antiviral activity and disassembly mechanism of novel 1-phenyl-5-amine-4-pyrazole thioether derivatives against Tobacco mosaic virus. Pestic Biochem Physiol 173:104771. https://doi.org/10.1016/j.pestbp.2021.104771
Xie WL et al (1991) ‘Expression of a mitogen-responsive gene encoding prostaglandin synthase is regulated by mRNA splicing. Proc Natl Acad Sci USA 88(7):2692–2696. https://doi.org/10.1073/pnas.88.7.2692
Yan R et al (2021) Synthesis and activity evaluation of some pyrazole-pyrazoline derivatives as dual anti-inflammatory and antimicrobial agents. Polycyclic Aromatic Compounds 42:1–14. https://doi.org/10.1080/10406638.2021.1919156
Youssif BGM et al (2019) Novel aryl carboximidamide and 3-aryl-1,2,4-oxadiazole analogues of naproxen as dual selective COX-2/15-LOX inhibitors: Design, synthesis and docking studies. Bioorganic Chemistry 85:577–584. https://doi.org/10.1016/j.bioorg.2019.02.043
Zabiulla et al (2019) Design, synthesis and molecular docking of benzophenone conjugated with oxadiazole sulphur bridge pyrazole pharmacophores as anti inflammatory and analgesic agents. Bioorganic Chemistry 92(August):103220. https://doi.org/10.1016/j.bioorg.2019.103220
Acknowledgements
Nagesh Khadri M J is thankful to KSTePS, DST, Govt. of Karnataka for providing fellowship. Shaukath Ara Khanum thankfully acknowledges the VGST, Bangalore, under CISEE Programme [Project sanction order: No. VGST/ CISEE/282].
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Khadri, M.J.N., Ramu, R., Simha, N.A. et al. Synthesis, molecular docking, analgesic, anti-inflammatory, and ulcerogenic evaluation of thiophene-pyrazole candidates as COX, 5-LOX, and TNF-α inhibitors. Inflammopharmacol 32, 693–713 (2024). https://doi.org/10.1007/s10787-023-01364-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10787-023-01364-0