Abstract
Sucrose transporters (SUTs) play an important role in regulating carbohydrate homogeneity, and well understand of the regulatory control of sugars into demanding sink is important for plant growth and seed yield. Nevertheless, key sucrose transporters that are involved in this process are not identified or characterized in soybean. Here, a sucrose transporter gene, GmSUT4.2, belonging to the SUT4 subfamily was cloned and functionally characterized. It encodes a protein of 513 amino acids which is localized in the plasma membrane. Real-time quantitative PCR showed that the expression of this gene was induced by sucrose and the sucrose transport capability could be functionally recovered by the expression of GmSUT4.2 in a sucrose transport dysfunction mutant yeast SUSY7/ura3. In soybean (Glycine max L.), overexpression of GmSUT4.2 significantly promoted sucrose-stimulated hairy root growth and improved the capacity of sucrose uptake. However, the knockdown of GmSUT4.2 in soybean by CRISPR/Cas9 caused a small leaf phenotype, which could also be observed in the ethyl methane sulfonate (EMS)-induced GmSUT4.2 mutant. In addition, the significantly low level of sucrose and soluble sugars were recorded in these mutants compared with wild-type plants (WT). As a result, the soybean yield and biomass in mutants were decreased by more than ~ 30% compared with WT under greenhouse conditions. While overexpression of GmSUT4.2 (OE) in Arabidopsis showed a pleiotropic phenotype with more rosette leaves and branches, resulting in a higher yield (40.07%) than those of wild types. These results suggest that the GmSUT4.2 played key roles in the regulation of plant growth and development as well as yield formation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sucrose is the main form of assimilated carbon produced during photosynthesis and critical for the overall growth and performance of crop plants (Aluko et al. 2021; Bavnhoj et al. 2023), serving as the energy for cellular metabolism, with additional associated roles as a signaling molecule (Koch 2004; Yoon et al. 2021). Sucrose fluxes from source leaf towards demanding sink organs via the phloem was coordinated by sucrose transporters (SUTs) (Carpaneto et al. 2005; Ayre 2011; Nino-Gonzalez et al. 2019; Peng et al. 2020), which are key determinants of plant productivity and crop yield (Ishibashi et al. 2014; Saalbach et al. 2014; Wang et al. 2015; Mathan et al. 2021; Wen et al. 2022; Gong et al. 2023).
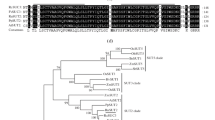
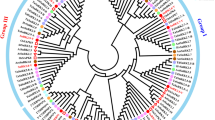
SUTs belong to the Major Facilitator Super Family (MFS), which is one of the largest known transporter families (Reddy 2013). Based on phylogenetic relationships analysis, SUTs are divided into five subgroups, namely SUT1-SUT5 (Kuhn and Grof 2010). Among them, the SUT1 subfamily members are indigenous to dicotyledons, which is so far the best characterized SUT members. SUT3 and SUT5 are native to monocotyledons, and SUT5 members have not been characterized. SUT2 and SUT4 clade members are found in both monocotyledons and dicotyledons (Reddy et al. 2012), and the SUT4 clade in most species was further divided into two subclades, SUT4.1 and SUT4.2 (Doidy et al. 2019).
Different members of each SUT clade show different functions. The SUT1 clade members are exclusively located in the plasma membrane and plays role in the apoplasmic phloem loading of sucrose (Slewinski et al. 2009; Oner-Sieben and Lohaus 2014; Nieberl et al. 2017; Dobbelstein et al. 2019). In addition, SUT1 genes also play essential roles in a variety of developmental and physiological processes in plants. For example, the SUT1 subfamily participates in regulation of vegetative growth (Gottwald et al. 2000), flowering time(Gu et al. 2020), pollen development (Lemoine et al. 1999; Stadler et al. 1999), anthocyanin accumulation(Sivitz et al. 2008), crop yield(Hackel et al. 2006; Sonnewald and Fernie 2018; Lu et al. 2020; Wang et al. 2021). The SUT2 and SUT3 subfamily were primally described as sugar sensor (Barker et al. 2000; Sauer 2007). However, the subsequent studies confirmed that members of SUT2 clade are involved in phloem loading(Hackel et al. 2006) or as a sucrose transporter (Ma et al. 2016; Ahmed et al. 2018), affecting plant yield, sugar accumulation (Ma et al. 2017) and resistance to abiotic stress (Ma et al. 2019).
In contrast, the SUT4 clade promotes sucrose uptake with low affinity and has high transport capacity within the SUT family (Weise et al. 2000). Members of the SUT4 clade are located at the plasma membrane or vacuolar membrane (Payyavula et al. 2011; Wang et al. 2020; Peng et al. 2020), and SUT4 transgenic plants do not show a consistent phenotype in different species. In Arabidopsis, AtSUC4 releases sucrose from the vacuole into the cytoplasm (Schulz et al. 2011), but the mutants have no observable phenotype. OsSUT2 mutant of rice exhibits a significantly reduced transport ability to export sucrose from the adult plant leaves compared with the wild type, resulting in severely delayed plant growth and reduced grain yield (Atkins et al. 2011). Similarly, the loss function of the ZmSUT2 gene in maize resulted in hyperaccumulation of foliar sucrose and greatly reduced growth (Leach et al. 2017). In addition, down-regulation of the StSUT4 gene resulted in early flowering, fewer leaves at flowering time, shorter internodes, and higher tuber production (Chincinska et al. 2008). In StSUT4-RNAi plants, the delayed export of sucrose and the accumulation of soluble sugars and starch in source leaves, suggest that StSUT4 is involved in circadian clock control (Garg et al. 2022). Thus, these results suggest that SUT4 clade members seem to have overwhelming importance for sucrose transport and plant development. In soybean, there are very few sucrose transporters functionally characterized to date. Doidy et al. (2019) indicated that two SUT4 accessions, GmSUT4.1 and GmSUT4.2, was identified in soybean. GmSUT4.1 is homologous to Ccjanus cajan CcSUT4.1 while GmSUT4.2 has the closest homology relationship to Ccjanus cajan CcSUT4.2, Vigna unguiculata VuSUT4.2 and Phaseolus vulgaris PvSUT4.2.
At present, the function of GmSUT4 in sucrose transport has not been demonstrated yet. In this context, a plasma membrane-localized sucrose transporter GmSUT4.2 gene was cloned from soybean, and its function was analyzed using yeast, Arabidopsis, and soybean. The results showed that GmSUT4.2 could transport sucrose and regulates leaf expanding and seed production, as well as sucrose homeostasis in Arabidopsis and soybean. Our findings improve the understanding of the SUTs mechanisms by which they mediate sucrose transport in plants to support the growth and development process.
Materials and methods
Plant materials and growth conditions
The soybean cultivar ‘Guixia1’, used for the analysis of the spatio-temporal expression levels of GmSUT4.2, was grown in potting soil, in a climate chamber. Tissues including those of root, steam, leaf, flower, source leaf, and seed tissues at the different developmental stages of trefoil, flowering, and seed development were sampled and kept in -80 °C.
For the analysis of the sucrose-dependent response, the soybean seedlings with the same growth rate were selected and treated with 1/4 Hoagland nutrient solution containing 1% sucrose. Subsequently, samples of root were collected at 0, 1, 2, 4, 8, 12, 24 and 48 h, for qRT-PCR analysis.
The EMS-induced GmSUT4.2 gene mutation lines NJAU0264 (GmSUT4-M1) and NJAU1191 (GmSUT4-M2) were provided by Nanjing Agricultural University, these two mutants were developed by using the cultivar Williams 82 (Zhang et al. 2022). All soybean plants used in this study were grown in pot cultures in a greenhouse (27 °C with a 14/10 h, light/dark).
Molecular cloning of GmSUT4.2 gene
The coding sequence of the GmSUT4.2 gene (Glyma04g09460) was downloaded from SoyBase (https://www.soybase.org) and primers (Supplementary in Table S1) used to clone the open reading frame (ORF) of GmSUT4.2 were designed using Premier 5.0. Total RNA was extracted from soybean leaves using TRNzol universal reagent (Vazyme Biotech Co., Ltd) following the manufacturer’s instructions. First-strand cDNA was synthesized from total RNA using a reverse transcription kit (Vazyme Biotech Co., Ltd). Polymerase chain reaction (PCR) products were recovered by FastPure Gel DNA Extraction Mini Kit (Vazyme Biotech Co., Ltd) and cloned using pEASY®-Blunt Cloning Kit (TransGen Biotech). After screening the positive clones, the strains were sequenced by BGI Genomics Co., Ltd.
Subcellular localization
The coding sequence of GmSUT4.2 without stop codon was cloned into the pBI121 vector with a green fluorescent protein (pBI121-GFP) between the Xba I and Kpn I to generate the GmSUT4.2-GFP plasmid. The vector of the fusion construct was transformed into the Agrobacterium strain EHA105, which was injected into five-week-old tobacco plants as described by Yue et al. (2022).The epidermis of leaves was infiltrated with the suspended bacterial solutions, and the resulting images were captured 48 h later using a fluorescence microscope (BX63, Olympus, Tokyo, Japan).
Functional analysis of GmSUT4.2 in yeast
To verify the sucrose transport activity of the SUT4.2 protein, the full length of GmSUT4.2 was sub-cloned into the pDR196 vector to produce the plasmid of pDR196-GmSUT4.2, and subsequently transformed it into yeast strain SUSY7/ura3 which is deficient in cell sucrose metabolism. Empty pDR196 vector was used as a negative control. Positive transformants were selected and grown in SD liquid medium (containing 6.7 g L−1 yeast nitrogen base without amino acids, 1.29 g L−1 SD/-ura base, 2% glucose, and a pH of 5.8) at 30 °C with continuous shaking at 220 rpm until an OD600 value of 1.2 was reached. The supernatant was removed by centrifugation at 4 000 rpm for 5 min, and the pellet was resuspended in sterile water to an optical density of 1.0 at OD600. The yeast cell suspension was then diluted 101, 102, 103, and 104 fold, and 2 μL of each dilution reaction was added to modified SD agar medium (SD/-ura liquid medium, 20 g L−1 agar) supplemented with 2% sucrose or 2% glucose. Yeast growth conditions were observed and recorded after 3 days of incubation at 30 °C in the dark.
Transformation of soybean hairy roots
Hairy roots were generated according to the previously described method with slight modifications (Fan et al. 2020). Briefly, soybean seeds were surface sterilized by treatment with chlorine gas, which was generated by adding 12 N HCl (4 mL) and 5.25% NaClO (50 mL) in an airtight dryer for 14 h. The seeds were cleaned three times with distilled water, and germinated in MS medium for 3 days. The imbibed seeds were used to produce explants in which the cotyledon nodes running parallel to the hypocotyl axis were gently scratched 3 ~ 4 times with a scalpel. The wounded explants were then transferred to Agrobacterium co-cultivation medium (CCM) for 30 min at 28 °C with constant shaking at 120 rpm. For the Agrobacterium CCM medium, expression of GmSUT4.2 was mediated by Agrobacterium rhizogenes K599 with the exogenous plasmid pCAMBIA1301-GmSUT4.2. After infection, 9 explants were placed evenly on each agar CCM coated with filter paper and co-cultured at 24 °C in the dark for 3 days. After co-culture, explants were washed six times with distilled water containing 300 mg L−1 cephalosporin. The explants were then transferred to a rooting medium and kept in an incubator at 24 °C on a 16/8 h light/dark cycle for 12 days.
CRISPR/Cas9 expression vector construction and soybean transformation
The online tool CRISPR-GE (http://skl.scau.edu.cn/) was used to predict and design the targets of sequences of the GmSUT4.2 gene. The single guide RNA (sgRNA) expression cassettes containing target sequences was conducted by overlapping PCR and subsequently built into the pYLCRISPR/Cas9Ppubi-H vector according to the protocol reported by (Ma et al. 2015). The positive plasmids were introduced into Agrobacterium tumefaciens strain EHA105 for soybean transformation. The soybean cultivar ‘Guixia1’ was transformed using the Agrobacterium-mediated cotyledon node system using established protocols (Zeng et al. 2004). The T0 generation transgenic seedlings were verified by PCR amplification using the specific primer Cas9-F/R and Hpt-F/R. In addition, gene editing status in the confirmed transgenic plants was examined by amplifying and sequencing fragments of the regions spanning the targets using primer Sp-F/R.
GmSUT4.2 overexpression vector construction and transgenic Arabidopsis plants acquisition
The One-step Cloning Kit (Vazyme Biotech Co., Ltd) and homologous recombination method was used for the amplification of the full-length GmSUT4.2 coding region and construction of the vector. The recombinant pCAMBIA1301-GmSUT4.2 plasmid was transformed into Agrobacterium tumefaciens GV3101, which was then used to transfect Arabidopsis plants by the floral dip method (Clough 2005). The T1 generation seedlings of the transgenic plants were tested on 1/2 MS media supplemented with 30 mg L–1 hygromycin and further verified by PCR and RT-qPCR.
Morphological and biomass parameters measurements
For leaf area measurements, fully expanded leaves at stage V1 (the single leaf is fully extended), R1 (the first flower appeared), and R3 (2 cm pods at the nodes) were photographed in vitro and the area was quantified using ImageJ software (Schroeder et al. 2021). The leaves and stems of the plants at R3 stage were collected and killed at 105 °C for 30 min before being dried to a constant weight at 85 °C and weighed to determine the dry biomass.
Sugar and starch content determination
Soluble sugar was extracted from 200 mg of fresh leaves using 1 mL of 80% ethanol. The determination of sucrose content was performed using a method modified from Lee et al. (2020). Starch content was determined using the method described by Huang et al. (2020) with some modifications. After sugar extraction, the pellet was dried, suspended with 1 mL of distilled water, and heated at 100 °C for 15 min. The sample was further incubated with 9.2 mol L−1 perchloric acid at 100 °C for 15 min. After cooling, the pellet was centrifuged further at 12 000 rpm for 10 min, and the supernatant was used to measure the starch content at 620 nm by the anthranone colorimetric method.
RT-qPCR analysis
The RNA extraction and cDNA transcription were same as mentioned above. Gene expression was quantified with specific primers q-F and q-R. Actin was used as the control. All primers used for assays are shown in Table S2. TransStart Top Green qPCR SuperMix (TransGen Biotech Co., Ltd.,) was used for the RT-qPCR reactions, and the system and operating procedures were performed according to the instructions. The 2−ΔΔCt method (Zhang et al. 2020b) was used to calculate relative transcript levels normalized by actin. The experiments were performed with three biological replicates, each with three plants.
Statistical analysis
Three biological replicates were used for each experiment. All experimental data were analyzed using SPSS version 22.0 and GraphPad 8.2. One-way ANOVA test was used for significance analysis (*P < 0.05, **P < 0.01).
Results
Molecular cloning and sequence analysis of GmSUT4.2
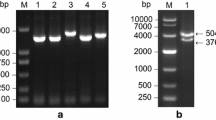
To investigate the function of sucrose transporters in the growth and development of soybean, we cloned the GmSUT4.2 gene by PCR amplification. The total nucleotide sequence length of the GmSUT4.2 gene was 1 765 bp, with an open reading frame (ORF) of 1 542 bp (Fig. S1). The encoded protein had a length of 513 amino acid residues and contained 12 transmembrane domains. The predicted molecular weight of the GmSUT4.2 protein was 57 869.35 Da, and the theoretical isoelectric point was 9.43 (Fig. S2a). Conserved domain analysis of the protein revealed the presence of GPH and MFS functional domains in the GmSUT4.2 protein (Fig. S2b).
GmSUT4.2 encodes a plasma membrane protein
The GmSUT4-GFP vector of the GmSUT4.2 gene was constructed and transformed into tobacco leaf subepidermal tissue using Agrobacterium-mediated transformation. After 48 h of co-cultivation, the tissues were observed at 480 nm using a fluorescence microscope. GFP signals were visible in the cell membranes of tobacco cells (Fig. S3), but not in other parts of the cells, indicating that GmSUT4.2 is localized in the cell membranes of tobacco cells.
Tissue-specific and exogenous sucrose sensitivity analysis of GmSUT4.2
Root, stem, leaf, flower, and seed samples (15, 25, 35, and 45 days after flowering, DAF) of soybean were harvested and used to analyze the tissue-specific expression of GmSUT4.2 by RT-qPCR. The highest transcript abundance of GmSUT4.2 was found in roots, followed by seed 35 DAF, and mature leaf, and the lowest expression abundance was found in the stem, seed 15 DAF, and flower (Fig. 1a). Additional experiments to analyze the sucrose-dependent response analysis of sucrose transporter GmSUT4.2 in roots revealed that the expression of GmSUT4.2 was upregulated by 1% exogenous sucrose and reached a maximum after 24 h (Fig. 1b).
Analysis of expression patterns of GmSUT4.2. a The transcript level of GmSUT4.2 in different soybean tissues (stem, root, young leaf, mature leaf, flower, seed 15DAF, seed 25DAF, seed 35DAF and seed 45DAF) was measured by RT-qPCR (n = 3). b Root samples under 1% exogenous sucrose treatment were collected at different time points (0, 1, 2, 4, 8, 12, 24 and 48h) to analyze GmSUT4.2 expression by RT-qPCR (n = 3). Different letters represent significant differences according to One-way ANOVA (P < 0.05)
Analysis of sucrose transport activity of GmSUT4.2
To investigate the function of GmSUT4.2 as a sucrose-uptake carrier, we constructed the pDR196-GmSUT4.2 vector and transformed it into SUSY7/ura3. At a dilution ratio of 101 or 102, yeast transformed with GmSUT4.2 gene grew better than yeasts transformed with an empty pDR196 vector on the SD medium containing 2% sucrose, even at a dilution ratio of 103 or 104, SUSY7/ura3 expressed GmSUT4.2 could grow normally. However, there was no difference in the growth of the yeast transformed with pDR196-GmSUT4.2 and pDR196 empty vector at all dilutions on the SD medium containing 2% glucose (Fig. 2). Expression of GmSUT4.2 in SUSY7/ura3 was able to recover a normal sucrose transportation capacity for yeast, suggesting that GmSUT4.2 encodes a functional transporter in sucrose transport.
Overexpression GmSUT4.2 promotes sucrose uptake in soybean hairy roots
We exposed soybean hairy roots overexpressing GmSUT4.2 (OE) and expressing empty vector (VC) to sucrose concentrations of 0.1% (MS0), 1% (MS1), and 3% (MS3) in MS medium following confirmation of GmSUT4.2 expression (Fig. S4). The transgenic roots overexpressing GmSUT4.2 grew well on MS1 and MS3 medium compared with control roots under normal conditions. However, no significant difference was observed in the growth rate between OE and VC grown on MS0 medium (Fig. 3). Furthermore, the metabolic conversion of exogenous sucrose to soluble sugar and endogenous sucrose was investigated. The results showed that the content of soluble sugar and sucrose in both roots overexpressing the GmSUT4.2 gene and control roots gradually increased with increasing sucrose concentration. Compared with the control roots, GmSUT4.2 overexpressing roots grown on MS1 and MS3 medium showed an increase in soluble sugar content of 78.43% and 155.35%, and sucrose content of 9.85% and 28.08% respectively. Similar content of sucrose (1.22 mg. g−1 and 1.25 mg. g−1) and soluble sugar (4.40 mg. g−1 and 4.49 mg. g−1) was observed in OE and VC grown on MS0 medium (Fig. 3). These results suggest that the expression of the sucrose transporter GmSUT4.2 leads to the rapid uptake of sucrose in overexpressing hairy roots.
Loss of function of GmSUT4.2 affects leaf size and seed production in soybean
GmSUT4.2 mutations in the EMS-induced mutants GmSUT4-M1 and GmSUT4-M2 were confirmed by sequencing (Fig.S5). GmSUT4-M1 carries an EMS-induced G to A mutation in GmSUT4.2 at 204 amino acid position in the coding region. GmSUT4-M2 carries an EMS-induced C to T mutation in GmSUT4.2 at 446 amino acid that introduces a premature stop codon in place of Asparagine. Phenotypic observation of mutant and wild-type plants in the greenhouse showed that non-synonymous mutations of GmSUT4.2 in soybean significantly inhibited plant growth (Fig. 4). Leaf area measurements showed that the small leaf phenotype of mutant plants appeared at the development stage V1 (Fig. 4c). The defective phenotype gradually becomes evident as the leaf develops and severely affects the growth of the plants in the R stages. Moreover, we characterized agronomic traits of GmSUT4-M1, GmSUT4-M2, and WT in the greenhouse (Fig. 4d-f). Growth of the mutants was poor compared with wild-type soybean, with fewer pods, seeds, and lower aerial biomass. Accordingly, seed weight per plant was significantly lower in the mutants than in the wild-type plants.
Organs phenotype and measurement of agronomic traits of EMS-induced soybean mutants. a The organs phenotype of GmSUT4-M1 and GmSUT4-M2 at different stages. b Phenotype of soybean plants at R3 stage. c, d Leaf area and plant height of WT and mutants at stage V1, R1 and R3 (n = 3). e Biomass of mutants and WT (n = 4). f Agronomic traits of mutant GmSUT4-M1 and GmSUT4-M2 (n = 6). *, P < 0.05, **, P < 0.001, according to One-way ANOVA
To confirm that the mutant phenotypes (GmSUT4-M1 and GmSUT4-M2) were directly caused due to the mutation of GmSUT4.2 and independent of other gene(s) mutations in NJAU0264 and NJAU1191, we constructed a CRISPR/Cas9-GmSUT4.2 vector containing two target adaptors, which were chosen for mutagenesis of this gene in soybean using the CRISPR/Cas9 technology (Fig. 5). The Arabidopsis U3b, and U6-1 promoters were used to drive the individual expression of the 2 sgRNA expression cassettes containing the designed target sites. We transformed this vector into Agrobacterium tumefaciens strain EHA105 to produce soybean transgenic plants. After screening the progeny by PCR and sanger sequencing, we obtained a Cas9-null homozygous mutants with frameshifts in the coding region of Cas-GmSUT4 (1-bp deletion) (Fig. 5a). Consistent with the GmSUT4-M1 and GmSUT4-M2, the plant growth and seed weight were decreased in mutant Cas-GmSUT4 (Fig. 5b-g). Compared to the wild type, seed weight per plant of T1 generation of Cas-GmSUT4 plants was decreased by 31.25%. Therefore, the combined evidence points to the importance of GmSUT4.2 in regulating plant growth, development, and agronomic yield of soybean plants.
Phenotypic identification and agronomic trait analysis of GmSUT4.2 mutant mediated by CRISPR/Cas9. a Targets location, diagram of the dual vector gRNA CRISPR/Cas9 and PCR-based identification of T0 transgenic plants; b Pods phenotype; c Plant height (n = 6); d Leaf size (n = 6); e Seeds phenotype; f, g Seeds number and seeds weight (n = 2) *, P < 0.05, **, P < 0.001, according to One-way ANOVA
GmSUT4.2 mutants have decreased sugar content in leaves
The function of sucrose transport of GmSUT4.2 makes us to hypothesize that differences in sugar distribution in mutants and wild types could account for the plant growth differences. To investigate this possibility, the sugar content of mature adult source leaves in both mutants and wild-type was examined (Fig. 6a). The contents of primary metabolites at the R3 stage showed a decrease of 57.63%, 43.91%, and 36.56% in sucrose content and 60.19%, 51.18%, and 29.97% in total soluble sugar content in the lines of GmSUT4-M1, GmSUT4-M2, and Cas-GmSUT4, respectively, which were significantly lower than those of wild-type plants (Williams 82, Guixia1). Similarly, the starch content of the mutant plants was also decreased compared with the wild type. Furthermore, RT-qPCR analysis revealed the differences in the expression levels of genes involved in sugar transport (GmSUT2, GmSWEET6, and GmSWEET11) and sucrose metabolism (GmSPP2, and GmCInv1), and sucrose signaling (GmSnRK1) in the mutants and wild type. The results showed that the expression of GmSPP2, GmCInv1, and GmSnRK1 was significantly down-regulated in the mutants compared with the wild type, but the expression of three sucrose transportation genes was up-regulated in GmSUT4-M1 and GmSUT4-M2 (Fig. 6b).
GmSUT4.2 regulates the development of rosette leaves, branches, and yield in Arabidopsis
To further elucidate the functional role of GmSUT4.2 in sucrose transport, the homozygous T3 generation of Arabidopsis lines overexpressing the GmSUT4.2 gene was generated. The transgenic plants with hygromycin resistance were further verified by PCR and RT-qPCR. Fragments of the required length could be amplified in the DNA genome of the transgenic plants, but not in the wild types (Fig. S7). Real-time fluorescence quantification results showed that transcription of GmSUT4.2 was significantly up-regulated in transgenic Arabidopsis line OE1, OE2 and OE4 (Fig. S8), and then we used these three lines for further experiments.
Phenotypic observation showed delayed leaf expansion in the transgenic plants compared with the wild type (Fig. 7a). Compared with the fully expanded rosette leaves of the wild type, the leaves of the transgenic plants were contracted in the middle and formed a central bundle, resulting in numerous rosette leaves (Fig. 7b). Transgenic line OE1, OE2 and OE4 had 25, 19, and 21 rosette leaves respectively, with an average of 22 leaves, which was significantly more than the number in WT (average of 13.5 rosette leaves). The number of rosette branches (average of 2.5 RI branches) was also higher in the transgenic plants than in the wild type (average of 1.5 RI branches) (Fig. 7c, d). Furthermore, we also evaluated the seed development of the transgenic lines. Generally, the number of pods per plant was significantly higher (38.44%) in the transgenic Arabidopsis than in the wild type. Consequently, the seed yield of the transgenic plants per plant was 40.07% higher than that of wild-type plants. However, we found no significant difference in 100-seed weight in the transgenic plants compared with the wild type (Fig. 7e, f).
Morphological analysis of the transgenic and wild-type plants. a Phenotype of the 12, 16 and 18 days seedings of WT and transgenic Arabidopsis plants. b, c Phenotype of the rosette leaves and branches of the transgenic and wild-type plants. d Wild-type plants and transgenic lines were grown under 16-h photoperiods. Measurement of morphological indicators including rosette leaves, rosette branches and cauline branches (n = 12). e Comparison of silique development between 6-week-old wild type and transgenic plants. f Agronomic traits (pods number per plant, seeds weight per plant and 100-seed weight) were quantified from transgenic (n = 36) and WT (n = 12) plants. *, P < 0.05, **, P < 0.001, according to One-way ANOVA
Sugar metabolism level in transgenic Arabidopsis plants
The measurement of the sucrose content of rosette leaves on day 30 (at the time of axillary bud formation) was conducted. In transgenic lines, the leaves had a higher accumulation of sucrose than the wild-type plants (Fig. 8a). Additionally, we measured photosynthetic activity and the transcript levels of genes related to sucrose metabolism and transport in line OE1 and OE2. The Photosynthetic rate and the expression of AtSPP2, AtCinv1, and AtSnRK1 were enhanced in transgenic lines compared with wild types (Fig. 8b, c). Further research found that transcript levels of sucrose efflux genes, such as AtSWEET11, AtSWEET12, and AtSWEET13, were also markedly up-regulated in transgenic plants compared with WT (Fig. 8d).
Physiological and molecular feedbacks in transgenic plants and WT. a, b Determination of sucrose content and photosynthetic activity in plants. c, d The transcriptional levels of genes related to sugar metabolism in wild-type and transgenic plants (n = 3). *, P < 0.05, **, P < 0.001, according to One-way ANOVA
Discussion
GmSUT4.2 is a functional protein localized to the plasma membrane
Subcellular localization is pivotal for the function of transport proteins (Schulz et al. 2011). Here, we observed that the fluorescence signal of GmSUT4-GFP was clearly present in the cell membranes of tobacco cells (Fig.S3). Previously reported that the results for subcellular localization of sucrose transporter genes belonging to the SUT4 subfamily have been mixed in different plants and even in the same plant species, with most of the members being reported to possess at least two different compartments (Schulz et al. 2011). For GeSUT4 from Gastrodia elata, it was identified as a plasma membrane protein in functional characterization of yeast, but GFP fusion showed that fluorescence was present in the plasma membrane and the tonoplast(Ho et al. 2021). Furthermore, a variety of SUT4 subfamily members, such as Arabidopsis AtSUT4 (Endler et al. 2006; Weise et al. 2000), Nicotiana tabacum NiSUT4 (Okubo-Kurihara et al. 2011), barley HvSUT2 (Endler et al. 2006), potato StSUT4 (Chincinska et al. 2013) and lotus LjSUT4 (Reinders et al. 2008), have been shown to reside in the vacuolar membrane and plasma membrane. The expression of GmSUT4.2 in SUSY7/ura3 yeast restored the sucrose-uptake ability which requires at least the localization of the protein to the plasma membrane (Fig. 2b). This result provided additional evidence that GmSUT4.2 functioned as a transporter in the plasma membrane.
Generally, SUTs transport sucrose; however, AtSUT5 also mediates the transport of biotin (Ludwig et al. 2000), AtSUT9 and LjSUT4 transport a variety of glucosides (Reinders et al. 2008), but not sucrose (AtSUT9) (Sivitz et al. 2007). In the current study, GmSUT4.2 was transferred into sucrose uptake-deficient yeast mutant SUSY7/ura3 and partially restored sucrose uptake in yeast implying that GmSUT4.2 to be a functional protein with sucrose transport activity (Fig. 2b).
GmSUT4.2 function is essential for the normal growth of soybean
Mutants play an important role in identifying gene function (Greene et al. 2003; Kuhn and Grof 2010). In our study, we observed that loss function of GmSUT4.2 display severe growth defects such as reduced leaf area, plant height, and seed yield compared with the wild type (Fig. 4), similar to previous observations for the mutants of SUT4 clade members (ZmSUT2, StSUT4 and OsSUT2) in other species (Chincinska et al. 2008; Eom et al. 2011; Leach et al. 2017). Thus, these results suggested that GmSUT4.2 function is necessary for normal plant growth in soybean. In parallel, mutation of GmSUT4.2 resulted in decreased sugar capacity in leaf and led to a lower biomass compared with wild-type plants (Fig. 6a). In previous studies, sucrose loading function of SUT4 members in leaves has been thoroughly studied in maize and rice (Eom et al. 2011; Leach et al. 2017). Limitations in sucrose export promote accumulation of sugars in leaves, thereby inhibiting leaf photosynthesis. In recent years, increasing evidence shows that SUT4 members function as sugar transporter and contribute to sugar accumulation (Zheng et al. 2014; Zhang et al. 2018; Peng et al. 2020). In addition, several SUT4 clade members, including pear PbSUT2 (Wang et al. 2016), rice OsSUT2 (Zhang et al. 2020a) and Populus PtaSUT4(Frost et al. 2012), were found to be potentially associated with the photosynthetic rate. Our results support the hypothesis that GmSUT4.2 functions as a source leaf sucrose phloem loader and is involved in leaf photosynthesis, and additional research is necessary to test this hypothesis.
Manipulation of the source-sink interaction determines the seed formation. In this study, the expression of the sucrose transporter gene GmSUC2 and GmSWEET6 is upregulated in the mutants (Fig. 6b). These results suggested that sucrose export is not blocked from the source leaf. There is little evidence that SUT4 orthologs somehow mediate sucrose export from source leaves to sink organs. The growth is severely reduced in null mutants of SUT4 members in rice and maize, while the sucrose export and long-distance translocation are normal (Julius et al. 2017). An explanation is that the loss of function of sucrose transporters and the imbalance of sugar accumulation in plants may induce compensatory expression of other sugar supply or transformation genes.
GmSUT4.2 regulates Arabidopsis growth in a sucrose-dependent manner
Overexpression of GmSUT4.2 strongly affects leaf development in the transgenic Arabidopsis plants, resulting in cluster growth in the middle of the leaves and eventually more rosette leaves (Fig. 7a, b). Leaf growth and development are primarily driven by cell proliferation and expansion (Lee et al. 2022). Recent studies have reported sugar availability is closely to apical meristem cell proliferation, affecting overall growth responses and developmental transitions in short-lived plants such as Arabidopsis (Zhang et al. 2019). Determination of sugar content showed that the overexpressing lines accumulated more sucrose in leaves compared with the wild type (Fig. 8a). This result is consistent with those observed in the transgenic tobacco overexpression of RuSUT2 (Yan et al. 2021). This also confirms recent work in peas and other crops demonstrating that the expression of sucrose transporters in plants had higher photosynthesis capacity than control plants, increased sugar accumulation, and significantly improved the storage of sinks (Ding et al. 2019; Lu et al. 2020; Wang et al. 2016). The intrinsic mechanism by which sucrose transporters affect photosynthetic activity in plants is unclear. Recent studies suggest that sucrose transport depends on transporter activity in leaves, which in turn regulates the transport load of the phloem and ultimately controls photosynthesis (Bush 2020).
Moreover, ectopic expression of GmSUT4.2 promoted the buds to be released from a dormant state, resulting in increased rosette branches, and finally has a strong additive effect on the yield of Arabidopsis (Fig. 7c, f; Fig. 8a, b). Given the established function of sucrose as a signal for the availability of plant bud branches at the top advantage (Barbier et al. 2015, 2019; Fichtner et al. 2017; Martin-Fontecha et al. 2018). At our consideration, whether sucrose sensing and signaling functions or transport ability was altered by the expression of GmSUT4.2 in Arabidopsis thaliana. T6P is a sucrose signaling metabolite and acts as a negative feedback regulator of sucrose levels (Yadav et al. 2014; Figueroa and Lunn 2016). SnRK1 is a sugar signaling node and appears to be repressed by high-energy signals such as trehalose-6-P (T6P) (Crepin and Rolland 2019), and CINV1 activity is an important checkpoint for coordinating sucrose catabolism plant growth and development mediated by sugar signaling and metabolism (Meng et al. 2021). As a result, RT-qPCR analysis revealed that the expression of SnRK1 and CINV1 was higher in transgenic plants than in wild-type plants (Fig. 8c). We assumed that the development of the plant growth and development was influenced by the interaction of SnRK1 and GmSUT4.2-mediatied sucrose signaling and metabolism because the final development of organs is primarily under the control of genetic programs. These findings call for further research to verify this possibility.
Conclusion
In this study, a soybean GmSUT4.2 gene was isolated and functionally characterized. Subcellular localization revealed that GmSUT4.2 localized to the plasma membrane, and the heterologous expression of GmSUT4.2 enabled SUSY7/ura3 yeast cells to grow on a sucrose medium. Moreover, loss function of GmSUT4.2 in soybean inhibits plant growth, resulting in a decrease in biomass and seed yield under greenhouse conditions. While GmSUT4.2 overexpression in Arabidopsis showed the opposite trend, with more rosette leaves, branches, and higher seed yield compared to the wild-type plants. Thus, our findings provide evidence that the GmSUT4.2 gene is involved in the regulation of plant growth and development as well as seed yield. GmSUT4.2 could be used as a candidate gene for future soybean seed yield and quality genetic improvements.
Data availability
All data generated or analyzed during this study were included in this published article and its supplementary information files.
References
Ahmed SA, Zhang JJ, Ma WJ, Dell B (2018) Contributions of TaSUTs to grain weight in wheat under drought. Plant Mol Biol 98(4–5):333–347. https://doi.org/10.1007/s11103-018-0782-1
Aluko OO, Li CZ, Wang Q, Liu HB (2021) Sucrose utilization for improved crop yields: A review article. Int J Mol Sci 22(9):4704–4733. https://doi.org/10.3390/ijms22094704
Atkins CA, Smith PM, Rodriguez-Medina C (2011) Macromolecules in phloem exudates–A review. Protoplasma 248(1):165–172. https://doi.org/10.1007/s00709-010-0236-3
Ayre BG (2011) Membrane-transport systems for sucrose in relation to whole-plant carbon partitioning. Mol Plant 4(3):377–394. https://doi.org/10.1093/mp/ssr014
Barbier F, Peron T, Lecerf M, Perez-Garcia MD, Barriere Q, Rolcik J, Boutet-Mercey S, Citerne S, Lemoine R, Porcheron B, Roman H, Leduc N, Le Gourrierec J, Bertheloot J, Sakr S (2015) Sucrose is an early modulator of the key hormonal mechanisms controlling bud outgrowth in Rosa hybrida. J Exp Bot 66(9):2569–2582. https://doi.org/10.1093/jxb/erv047
Barbier FF, Dun EA, Kerr SC, Chabikwa TG, Beveridge CA (2019) An update on the signals controlling shoot branching. Trends Plant Sci 24(3):220–236. https://doi.org/10.1016/j.tplants.20-18.12.001
Barker L, Kuhn C, Weise A, Schulz A, Gebhardt C, Hirner B, Hellmann H, Schulze W, Ward JM, Frommer WB (2000) SUT2, a putative sucrose sensor in sieve elements. Plant Cell 12(7):1153–1164. https://doi.org/10.1105/tpc.12.7.1153
Bavnhoj L, Driller JH, Zuzic L, Stange AD, Schiott B, Pedersen BP (2023) Structure and sucrose binding mechanism of the plant SUC1 sucrose transporter. Nat Plants. https://doi.org/10.1038/s41-477-023-01421-0
Bush DR (2020) Identifying the pathways that control resource allocation in higher plants. Proc Natl Acad Sci USA 117(16):8669–8671. https://doi.org/10.1073/pnas.2002581117
Carpaneto A, Geiger D, Bamberg E, Sauer N, Fromm J, Hedrich R (2005) Phloem-localized, proton-coupled sucrose carrier ZmSUT1 mediates sucrose efflux under the control of the sucrose gradient and the proton motive force. J Biol Chem 280(22):21437–21443. https://doi.org/10.1074/jbc.M50-1785200
Chincinska IA, Liesche J, Krugel U, Michalska J, Geigenberger P, Grimm B, Kuhn C (2008) Sucrose transporter StSUT4 from potato affects flowering, tuberization, and shade avoidance response. Plant Physiol 146(2):515–528. https://doi.org/10.1104/pp.107.112334
Chincinska I, Gier K, Krugel U, Liesche J, He HX, Grimm B, Harren FJM, Cristescu SM, Kuhn C (2013) Photoperiodic regulation of the sucrose transporter StSUT4 affects the expression of circadian-regulated genes and ethylene production. Front Plant Sci. https://doi.org/10.3389/fp01-3.00026
Clough SJ (2005) Floral dip: agrobacterium-mediated germ line transformation. Methods Mol Biol 286:91–102. https://doi.org/10.1385/1-59259-827-7:091
Crepin N, Rolland F (2019) SnRK1 activation, signaling, and networking for energy homeostasis. Curr Opin Plant Biol 51:29–36. https://doi.org/10.1016/j.pbi.2019.03.006
Ding XY, Zeng JY, Huang L, Li XB, Song SQ, Pei Y (2019) Senescence-induced expression of ZmSUT1 in cotton delays leaf senescence while the seed coat-specific expression increases yield. Plant Cell Rep 38(8):991–1000. https://doi.org/10.1007/s00299-019-02421-1
Dobbelstein E, Fink D, Oner-Sieben S, Czempik L, Lohaus G (2019) Seasonal changes of sucrose transporter expression and sugar partitioning in common European tree species. Tree Physiol 39(2):284–299. https://doi.org/10.1093/treephys/tpy120
Doidy J, Vidal U, Lemoine R (2019) Sugar transporters in Fabaceae, featuring SUT MST and SWEET families of the model plant Medicago truncatula and the agricultural crop Pisum sativum. PLoS ONE. https://doi.org/10.1371/journal.pone.0223173
Endler A, Meyer S, Schelbert S, Schneider T, Weschke W, Peters SW, Keller F, Baginsky S, Martinoia E, Schmidt UG (2006) Identification of a vacuolar sucrose transporter in barley and Arabidopsis mesophyll cells by a tonoplast proteomic approach. Plant Physiol 141(1):196–207. https://doi.org/10.1104/pp.106.079533
Eom JS, Cho JI, Reinders A, Lee SW, Yoo Y, Tuan PQ, Choi SB, Bang G, Park YI, Cho MH, Bhoo SH, An G, Hahn TR, Ward JM, Jeon JS (2011) Impaired function of the tonoplast-localized sucrose transporter in Rice, OsSUT2, limits the transport of vacuolar reserve sucrose and affects plant growth. Plant Physiol 157(1):109–119. https://doi.org/10.1104/pp.111.176982
Fan YL, Zhang XH, Zhong LJ, Wang XY, Jin LS, Lyu SH (2020) One-step generation of composite soybean plants with transgenic roots by Agrobacterium rhizogenes-mediated transformation. BMC Plant Biol 20(1):208–219. https://doi.org/10.1186/s12870-020-02421-4
Fichtner F, Barbier FF, Feil R, Watanabe M, Annunziata MG, Chabikwa TG, Hofgen R, Stitt M, Beveridge CA, Lunn JE (2017) Trehalose 6-phosphate is involved in triggering axillary bud outgrowth in garden pea (Pisum sativum L.). Plant J 92(4):611–623. https://doi.org/10.1111/tpj.13-705
Figueroa CM, Lunn JE (2016) A tale of two sugars: trehalose 6-phosphate and sucrose. Plant Physiol 172(1):7–27. https://doi.org/10.1104/pp.16.00417
Frost CJ, Nyamdari B, Tsai CJ, Harding SA (2012) The tonoplast-localized sucrose transporter in populus (PtaSUT4) regulates whole-plant water relations, responses to water stress, and photosynthesis. PLoS ONE. https://doi.org/10.1371/journal.pone.0044467
Garg V, Reins J, Hackel A, Kuhn C (2022) Elucidation of the interactome of the sucrose transporter StSUT4: sucrose transport is connected to ethylene and calcium signalling. J Exp Bot. https://doi.org/10.1093/jxb/erac378
Gong HL, Liu JB, Igiraneza C, Dusengemungu L (2023) Sucrose transporter StSUT2 affects potato plants growth, flowering time, and tuber yield. Curr Issues Mol Biol 45(3):2629–2643. https://doi.org/10.3390/cimb45030172
Gottwald JR, Krysan PJ, Young JC, Evert RF, Sussman MR (2000) Genetic evidence for the in planta role of phloem-specific plasma membrane sucrose transporters. Proc Natl Acad Sci USA 97(25):13979–13984. https://doi.org/10.1073/pnas.250473797
Greene EA, Codomo CA, Taylor NE, Henikoff JG, Till BJ, Reynolds SH, Enns LC, Burtner C, Johnson JE, Odden AR, Comai L, Henikoff S (2003) Spectrum of chemically induced mutations from a large-scale reverse-genetic screen in arabidopsis. Genetics 164(2):731–740. https://doi.org/10.1093/genetics/164.2.731
Gu JH, Zeng Z, Wang YR, Lyu YM (2020) Transcriptome analysis of carbohydrate metabolism genes and molecular regulation of sucrose transport gene LoSUT on the flowering process of developing oriental hybrid lily “Sorbonne” bulb. Int J Mol Sci. https://doi.org/10.3390/ijms21093092
Hackel A, Schauer N, Carrari F, Fernie AR, Grimm B, Kuhn C (2006) Sucrose transporter LeSUT1 and LeSUT2 inhibition affects tomato fruit development in different ways. Plant J 45(2):180–192. https://doi.org/10.1111/j.1365-313X.2005.02572
Ho LH, Lee YI, Hsieh SY, Lin IS, Wu YC, Ko HY, Klemens PA, Neuhaus HE, Chen YM, Huang TP, Yeh CH, Guo WJ (2021) GeSUT4 mediates sucrose import at the symbiotic interface for carbon allocation of heterotrophic Gastrodia elata (Orchidaceae). Plant Cell Environ 44(1):20–33. https://doi.org/10.1111/pce.13833
Huang Y, Wang LL, Hu SL, Luo XG, Cao Y (2020) Overexpression of the bamboo sucrose synthase gene (BeSUS5) improves cellulose production, cell wall thickness and fiber quality in transgenic poplar. Tree Genet Genomes 16(5):75–89. https://doi.org/10.1007/s11295-020-01464-w
Ishibashi Y, Okamura K, Miyazaki M, Phan T, Yuasa T, Iwaya-Inoue M (2014) Expression of rice sucrose transporter gene OsSUT1 in sink and source organs shaded during grain filling may affect grain yield and quality. Environ Exp Bot 97:49–54. https://doi.org/10.1016/j.envexpbot.2013.08.0-05
Julius BT, Leach KA, Tran TM, Mertz RA, Braun DM (2017) Sugar transporters in plants: new insights and discoveries. Plant Cell Physiol 58(9):1442–1460. https://doi.org/10.1093/pcp/pcx090
Koch K (2004) Sucrose metabolism: regulatory mechanisms and pivotal roles in sugar sensing and plant development. Curr Opin Plant Biol 7(3):235–246. https://doi.org/10.1016/j.pbi.-2004.03.014
Kuhn C, Grof CP (2010) Sucrose transporters of higher plants. Curr Opin Plant Biol 13(3):288–298. https://doi.org/10.1016/j.pbi.2010.02.001
Leach KA, Tran TM, Slewinski TL, Meeley RB, Braun DM (2017) Sucrose transporter2 contributes to maize growth, development, and crop yield. J Integr Plant Biol 59(6):390–408. https://doi.org/10.1111/jipb.12527
Lee H, Lee BR, Islam MT, La VH, Park SH, Bae DW, Kim TH (2020) Cultivar variation in hormone- and sugar-response reveals abscisic acid-responsive sucrose phloem loading at the early regenerative stage is a significant determinant of seed yield in Brassica napus. Environ Exp Bot 169:103917–103926. https://doi.org/10.1016/j.envexpbot.2019.103917
Lee GH, Lee BH, Jung JH, Lee SJ, Mai TT, Kim JH (2022) Systematic assessment of the positive role of Arabidopsis thaliana growth-regulation factors in regulation of cell proliferation during leaf growth. J Plant Biol 65:413–422. https://doi.org/10.1007/s12374-022-09366-1
Lemoine R, Burkle L, Barker L, Sakr S, Kuhn C, Regnacq M, Gaillard C, Delrot S, Frommer WB (1999) Identification of a pollen-specific sucrose transporter-like protein NtSUT3 from tobacco. Febs Lett 454(3):325–330. https://doi.org/10.1016/S0014-5793(99)00843-1
Lu MZ, Snyder R, Grant J, Tegeder M (2020) Manipulation of sucrose phloem and embryo loading affects pea leaf metabolism, carbon and nitrogen partitioning to sinks as well as seed storage pools. Plant J 101(1):217–236. https://doi.org/10.1111/tpj.14533
Ludwig A, Stolz J, Sauer N (2000) Plant sucrose-H+ symporters mediate the transport of vitamin H. Plant J 24(4):503–509. https://doi.org/10.1046/j.1365-313x.2000.00900.x
Ma XL, Zhang QY, Zhu QL, Liu W, Chen Y, Qiu R, Wang B, Yang ZF, Li HY, Lin YR, Xie YY, Shen RX, Chen SF, Wang Z, Chen YL, Guo JX, Chen LT, Zhao XC, Dong ZC, Liu YG (2015) A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant 8(8):1274–1284. https://doi.org/10.1016/j.molp.2015.04.007
Ma QJ, Sun MH, Liu YJ, Lu J, Hu DG, Hao YJ (2016) Molecular cloning and functional characterization of the apple sucrose transporter gene MdSUT2. Plant Physiol Bioch 109:442–451. https://doi.org/10.1016/j.plaphy.2016.10.026
Ma QJ, Sun MH, Lu J, Liu YJ, Hu DG, Hao YJ (2017) Transcription factor AREB2 is involved in soluble sugar accumulation by activating sugar transporter and amylase genes. Plant Physiol 174(4):2348–2362. https://doi.org/10.1104/pp.17.00502
Ma QJ, Sun MH, Lu J, Kang H, You CX, Hao YJ (2019) An apple sucrose transporter MdSUT2.2 is a phosphorylation target for protein kinase MdCIPK22 in response to drought. Plant Biotechnol J 17(3):625–637. https://doi.org/10.1111/pbi.13003
Martin-Fontecha ES, Tarancon C, Cubas P (2018) To grow or not to grow, a power-saving program induced in dormant buds. Curr Opin Plant Biol 41:102–109. https://doi.org/10.1016/j.pbi.2017.10-001
Mathan J, Singh A, Ranjan A (2021) Sucrose transport and metabolism control carbon partitioning between stem and grain in rice. J Exp Bot 72(12):4355–4372. https://doi.org/10.1093/jxb/erab066
Meng LS, Bao QX, Mu XR, Tong C, Cao XY, Huang JJ, Xue LN, Liu CY, Fei Y, Loake GJ (2021) Glucose- and sucrose-signaling modules regulate the Arabidopsis juvenile-to-adult phase transition. Cell Rep. https://doi.org/10.1016/j.celrep.2021.109348
Nieberl P, Ehrl C, Pommerrenig B, Graus D, Marten I, Jung B, Ludewig F, Koch W, Harms K, Flugge UI, Neuhaus HE, Hedrich R, Sauer N (2017) Functional characterisation and cell specificity of BvSUT1, the transporter that loads sucrose into the phloem of sugar beet (Beta vulgaris L) source leaves. Plant Biol 19(3):315–326. https://doi.org/10.1111/plb.12546
Nino-Gonzalez M, Novo-Uzal E, Richardson DN, Barros PM, Duque P (2019) More transporters, more substrates: the Arabidopsis major facilitator superfamily revisited. Mol Plant 12(9):1182–1202. https://doi.org/10.1016/j.molp.2019.07.003
Okubo-Kurihara E, Higaki T, Kurihara Y, Kutsuna N, Yamaguchi J, Hasezawa S (2011) Sucrose transporter NtSUT4 from tobacco BY-2 involved in plant cell shape during miniprotoplast culture. J Plant Res 124(3):395–403. https://doi.org/10.1007/s10265-010-0377-7
Oner-Sieben S, Lohaus G (2014) Apoplastic and symplastic phloem loading in quercus robur and fraxinus excelsior. J Exp Bot 65(7):1905–1916. https://doi.org/10.1093/jxb/eru066
Payyavula RS, Tay KHC, Tsai CJ, Harding SA (2011) The sucrose transporter family in Populus: the importance of a tonoplast PtaSUT4 to biomass and carbon partitioning. Plant J 65(5):757–770. https://doi.org/10.1111/j.1365-313X.2010.04463.x
Peng Q, Cai YM, Lai EH, Nakamura M, Liao L, Zheng BB, Ogutu C, Cherono S, Han YP (2020) The sucrose transporter MdSUT4.1 participates in the regulation of fruit sugar accumulation in apple. BMC Plant Biol. https://doi.org/10.1186/s12870-020-02406-3
Reddy (2013) The major facilitator superfamily (MFS) revisited. Febs J 280(16):3975–3975. https://doi.org/10.1111/febs.12405
Reddy VS, Shlykov MA, Castillo R, Sun EI, Saier MH Jr (2012) The major facilitator superfamily (MFS) revisited. FEBS J 279(11):2022–2035. https://doi.org/10.1111/j.1742-4658.2012.08588.x
Reinders A, Sivitz AB, Starker CG, Gantt JS, Ward JM (2008) Functional analysis of LjSUT4, a vacuolar sucrose transporter from Lotus japonicus. Plant Mol Biol 68(3):289–299. https://doi.org/10.1007/s11103-008-9370-0
Saalbach I, Mora-Ramirez I, Weichert N, Andersch F, Guild G, Wieser H, Koehler P, Stangoulis J, Kumlehn J, Weschke W, Weber H (2014) Increased grain yield and micronutrient concentration in transgenic winter wheat by ectopic expression of a barley sucrose transporter. J Cereal Sci 60(1):75–81. https://doi.org/10.1016/j.jcs.2014.01.017
Sauer N (2007) Molecular physiology of higher plant sucrose transporters. Febs Lett 581(12):2309–2317. https://doi.org/10.1016/j.febslet.2007.03.048
Schroeder AB, Dobson ETA, Rueden CT, Tomancak P, Jug F, Eliceiri KW (2021) The ImageJ ecosystem: Open-source software for image visualization, processing, and analysis. Protein Sci 30(1):234–249. https://doi.org/10.1002/pro.3993
Schulz A, Beyhl D, Marten I, Wormit A, Neuhaus E, Poschet G, Buttner M, Schneider S, Sauer N, Hedrich R (2011) Proton-driven sucrose symport and antiport are provided by the vacuolar transporters SUC4 and TMT1/2. Plant J 68(1):129–136. https://doi.org/10.1111/j.1365-313X.2011.04672.x
Sivitz AB, Reinders A, Johnson ME, Krentz AD, Grof CPL, Perroux JM, Ward JM (2007) Arabidopsis sucrose transporter AtSUC9. High-affinity transport activity, intragenic control of expression, and early flowering mutant phenotype. Plant Physiol 143(1):188–198. https://doi.org/10.1104/pp.106.089003
Sivitz AB, Reinders A, Ward JM (2008) Arabidopsis sucrose transporter AtSUC1 is important for pollen germination and sucrose-induced anthocyanin accumulation. Plant Physiol 147(1):92–100. https://doi.org/10.1104/pp.108.118992
Slewinski TL, Meeley R, Braun DM (2009) Sucrose transporter1 functions in phloem loading in maize leaves. J Exp Bot 60(3):881–892. https://doi.org/10.1093/jxb/ern335
Sonnewald U, Fernie AR (2018) Next-generation strategies for understanding and influencing source-sink relations in crop plants. Curr Opin Plant Biol 43:63–70. https://doi.org/10.1016/j.pbi.2018.01.004
Stadler R, Truernit E, Gahrtz M, Sauer N (1999) The AtSUC1 sucrose carrier may represent the osmotic driving force for anther dehiscence and pollen tube growth in Arabidopsis. Plant J 19(3):269–278. https://doi.org/10.1046/j.1365-313X.1999.00527.x
Wang L, Lu QT, Wen XG, Lu CM (2015) Enhanced sucrose loading improves rice yield by increasing grain size. Plant Physiol 169(4):2848–2862. https://doi.org/10.1104/pp.15.01170
Wang LF, Qi XX, Huang XS, Xu LL, Jin C, Wu J, Zhang SL (2016) Overexpression of sucrose transporter gene PbSUT2 from Pyrus bretschneideri, enhances sucrose content in Solanum lycopersicum fruit. Plant Physiol Biochem 105:150–161. https://doi.org/10.1016/j.plaphy.2016.04.019
Wang DD, Liu HJ, Wang HX, Zhang P, Shi CY (2020) A novel sucrose transporter gene IbSUT4 involves in plant growth and response to abiotic stress through the ABF-dependent ABA signaling pathway in Sweetpotato. BMC Plant Biol. https://doi.org/10.1186/s12870-020-02382-8
Wang GP, Wu Y, Ma L, Lin Y, Hu YX, Li MZ, Li WW, Ding YF, Chen L (2021) Phloem loading in rice leaves depends strongly on the apoplastic pathway. J Exp Bot 72(10):3723–3738. https://doi.org/10.1093/jxb/erab085
Weise A, Barker L, Kuhn C, Lalonde S, Buschmann H, Frommer WB, Ward JM (2000) A new subfamily of sucrose transporters, SUT4, with low affinity/high capacity localized in enucleate sieve elements of plants. Plant Cell 12(8):1345–1355. https://doi.org/10.1105/tpc.12.8.1345
Wen SY, Neuhaus HE, Cheng JT, Bie ZL (2022) Contributions of sugar transporters to crop yield and fruit quality. J Exp Bot 73(8):2275–2289. https://doi.org/10.1093/jxb/erac043
Yadav UP, Ivakov A, Feil R, Duan GY, Walther D, Giavalisco P, Piques M, Carillo P, Hubberten HM, Stitt M, Lunn JE (2014) The sucrose-trehalose 6-phosphate (Tre6P) nexus: specificity and mechanisms of sucrose signalling by Tre6P. J Exp Bot 65(4):1051–1068. https://doi.org/10.1093/jxb/ert457
Yan ZX, Yang HY, Zhang CH, Wu WL, Li WL (2021) Functional analysis of the blackberry sucrose transporter gene RuSUT2. Russ J Plant Phys 68(2):246–253. https://doi.org/10.1134/S102144372-1020217
Yoon J, Cho LH, Tun W, Jeon JS, An G (2021) Sucrose signaling in higher plants. Plant Sci. https://doi.org/10.1016/j.plantsci.2020.110703
Yue J, Tang MQ, Zhang H, Luo DJ, Cao S, Hu YL, Huang Z, Wu QJ, Wu X, Pan J, Chen CN, Wang CJ, Chen P (2022) The transcription factor HcERF4 confers salt and drought tolerance in kenaf (Hibiscus cannabinus L.). Plant Cell Tiss Org 150(1):207–221. https://doi.org/10.1007/s11240-022-02260-1
Zeng P, Vadnais DA, Zhang Z, Polacco JC (2004) Refined glufosinate selection in Agrobacterium-mediated transformation of soybean [Glycine max (L.) Merrill]. Plant Cell Rep 22(7):478–482. https://doi.org/10.1007/s00299-003-0712-8
Zhang CM, Bian Y, Hou SH, Li XG (2018) Sugar transport played a more important role than sugar biosynthesis in fruit sugar accumulation during Chinese jujube domestication. Planta 248(5):1187–1199. https://doi.org/10.1007/s00425-018-2971-1
Zhang N, Meng YY, Li X, Zhou Y, Ma LY, Fu LW, Schwarzlander M, Liu HT, Xiong Y (2019) Metabolite-mediated TOR signaling regulates the circadian clock in Arabidopsis. Proc Natl Acad Sci USA 116(51):25395–25397. https://doi.org/10.1073/pnas.1913095116
Zhang JS, Li DF, Xu X, Ziska LH, Zhu JG, Liu G, Zhu CW (2020a) The potential role of sucrose transport gene expression in the photosynthetic and yield response of rice cultivars to future CO2 concentration. Physiol Plantarum 168(1):218–226. https://doi.org/10.1111/ppl.12973
Zhang M, Liu YH, Cai HY, Guo ML, Chai MN, She ZY, Ye L, Cheng Y, Wang BR, Qin Y (2020b) The bZIP transcription factor GmbZIP15 negatively regulates salt- and drought-stress responses in soybean. Int J Mol Sci 21(20):7778–7797. https://doi.org/10.3390/ijms21207778
Zhang MZ, Zhang XY, Jiang XY, Qiu L, Jia GH, Wang LF, Ye WX, Song QX (2022) iSoybean: a database for the mutational fingerprints of soybean. Plant Biotechnol J 20(8):1435–1437. https://doi.org/10.1111/pbi.13844
Zheng QM, Tang Z, Xu Q, Deng XX (2014) Isolation, phylogenetic relationship and expression profiling of sugar transporter genes in sweet orange (Citrus sinensis). Plant Cell Tiss Org 119(3):609–624. https://doi.org/10.1007/s11240-014-0560-y
Acknowledgements
We are thankful to Professor Qiusheng Yang for providing mutant yeast SUSY7/ura3 and vector pDR196. We also thank Professor Qingxin Song for providing seeds of soybean cultivar Williams 82 and mutant line NJAU1191 and NJAU0264.
Funding
This research work was supported by the National Natural Science Foundation of China (Grant No. 31960368), and the China Agriculture Research System of MOF and MARA (CARS-16-E14).
Author information
Authors and Affiliations
Contributions
PC, conceived and designed the research. XW performed the experiments and analyzed the data. XW wrote the manuscript and JY, SM revised the manuscript. DJL, SC, CCW, QQW, HZ, CNC, MW, JZN participated in experiments and data collection. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no known competing financial interests.
Additional information
Communicated by Hong-Xia Zhang.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, X., Mubeen, S., Luo, D. et al. Function characterization of a soybean sucrose transporter GmSUT4.2 involved in plant growth, development, and crop yield. Plant Growth Regul 102, 529–543 (2024). https://doi.org/10.1007/s10725-023-01078-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-023-01078-x