Abstract
Little study was performed to know how microRNAs (miRNAs) are responsive to cadmium (Cd) stress in barley (Hordeum vulgare). In this study, 16 small RNA libraries of shoot and root tissues from a wild barley accession (WB-1) and cultivated barley (Golden Promise) with contrasting Cd tolerance were constructed and sequenced. Moreover, a degradome library was constructed and analyzed to identify target genes of the miRNAs. Based on high-throughput sequencing, 216 conserved miRNAs (in 59 miRNA families) and 87 novel miRNAs were identified. A total of 238 target genes for 149 miRNAs (113 conserved and 36 novel miRNAs) were detected by the degradome analysis. Among these miRNAs, 45 miRNAs (40 conserved and 5 novel miRNAs) and 43 miRNAs (40 conserved and 3 novel miRNAs) showed differential expression in roots and shoots of two genotypes under Cd conditions. Compared with cultivar Golden Promise, the wild genotype WB-1 had genotype-dependent responses of miR156, miR159, miR166, miR167, miR171 and miR393, which regulate target genes including SPL, MYB, HD-Zip, ARF, GRAS and TIR. Correspondingly, WB-1 had lower Cd concentration and stronger Cd tolerance than Golden Promise. It indicates that miRNAs may play critical roles underlying genotypic difference of Cd tolerance in barley.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cadmium (Cd) is a toxic element, recently becoming a global agricultural contaminant in the form of Cd2+ (ATSDR 2008). Soil Cd pollution is the consequences of metal mining, phosphate fertilizer overusing and other factors (Clemens et al. 2013; Khan et al. 2017). Unfortunately, Cd causes serious harm to human health including nephrotoxicity, osteoporosis and even cancer, mainly through the food chain (Jarup and Akesson 2009; Liu et al. 2009). Therefore, it is important to reduce Cd accumulation in crops. Cd can be readily taken up by plant roots, causing damages to plant growth, photosynthesis and nutrient uptakes (Khan et al. 2015; Uraguchi et al. 2009). In plants, many genes involved in Cd uptake, translocation and distribution have been identified, which are members of ABC (ATP-binding cassette), HMA (heavy metal ATPase), NRAMP (natural resistance-associated macrophage proteins) and ZIP (ZRT, IRT-like proteins) family genes (Kim et al. 2007; Siemianowski et al. 2014; Wu et al. 2016; Guerinot 2000). However, little study was performed to understand regulatory networks of microRNAs (miRNAs) involved in Cd tolerance in plants.
miRNAs are a special kind of endogenous non-coding small RNAs with the length of approximately 21 nucleotides (Voinnet 2009). Currently, there are 7,057 miRNAs from 73 plant species stored in the database miRBase (version 21; http://www.mirbase.org). It is well-known that miRNAs are involved in many biological processes such as plant growth, tissue development and various stress responses (Khraiwesh et al. 2012; Tang and Chu 2017). Furthermore, miRNAs were also reported in post-transcriptional regulatory networks under heavy metal stresses, such as arsenic (Ghosh et al. 2017; Yu et al. 2012), chromium (Bukhari et al. 2015), lead (He et al. 2014; Wang et al. 2015) and cadmium (Ding et al. 2011; He et al. 2016; Zhou et al. 2012). For instance, miRNA166 was confirmed to regulate Cd accumulation and tolerance in rice, and overexpression of miR166 could reduce Cd accumulation in grains and Cd translocation from roots to shoots (Ding et al. 2018). However, miRNAs involved in Cd tolerance have not been reported in barley (Hordeum vulgare).
Barley is the fourth important cereal crop in the world, which is widely used in processing malts and livestock feed due to its high nutrition value. Therefore, Cd accumulation in barley grains poses a great threat to human health. In our previous study, the Cd concentration in barley grains was analyzed in 100 barley core accessions planted in the natural fields for 2 years. The results showed a wide genotypic difference in Cd accumulation (Wu et al. 2015). However, the molecular mechanism of Cd accumulation and tolerance in barley is still unknown. In this study, we identified miRNAs in response to Cd exposure in wild and cultivated barley, and performed a degradome library to reveal target genes of these miRNAs. Furthermore, we compared the differential expression profiles of miRNAs between Golden Promise and WB-1 to identify their distinct miRNA regulation of Cd tolerance. These Cd-responsive miRNAs and their target genes could provide useful information for molecular mechanisms underlying Cd tolerance in barley.
Materials and methods
Plant materials and hydroponics
In this study, a barley cultivar Golden Promise and a wild barley genotype WB-1 were used. All materials were preserved by the Provincial Key Laboratory of Crop Gene Resources of Zhejiang University, China. Seeds of the two genotypes were disinfected with 2% H2O2 for 30 min, washed and soaked in deionized water for 2 h. Then, the seeds were germinated on moist filter papers in a growth chamber at 22 °C/18 °C (day/night) under dark conditions (Shen et al. 2017a, b). Light was supplied 5 days later. Seven-day-old seedlings were transferred to 6-L plastic pots filled with aerated one-fifth Hoagland (pH 6.0) solution, containing 1 mM KNO3, 1 mM Ca(NO3)2, 0.4 mM MgSO4, 0.2 mM NH4H2PO4, 20 µM Fe-EDTA, 3 µM H3BO3 and 1.0 µM CuSO4. The nutrient solution was renewed every 3 days.
Twelve-day-old seedlings were treated with 5 µM CdCl2. The solution without CdCl2 addition was used as the control. After 10 days treatments, root and shoot tissues of the two genotypes under treated and control conditions were sampled. Roots were washed 3 times with 5 mM CaCl2 solution, and then were separated from the shoots. Three biological replicates were sampled for physiological analysis. Meanwhile, two biological replicates were set for small RNA library construction, which were frozen immediately in liquid nitrogen and stored at − 80 °C for use.
Microelement and cadmium concentration determination
The root and shoot samples prepared as described above were dried in an oven at 70 °C for 2 days, and then digested completely in concentrated nitric acid solution using a microwave digestion instrument (Multiwave 3000, Anton Paar GmbH, Australia). There are interaction effects between Cd and other mineral elements. Therefore, the concentration of Cd and other mineral elements including Mn, Cu, Fe, and Zn was determined by ELAN® DRC-e ICP-MS (PerkinElmer SCIEX™, Concord, ON, Canada) following the manufacturer’s procedure. Cd uptake by roots was calculated by the formula: Cd uptake = Cd content per plant/root dry weight. While Cd translocation to shoots was calculated by the formula: Cd translocation (%) = shoot Cd content/Cd content per plant × 100%. Three biological replicates were performed for the analysis.
RNA extraction
Total RNA was extracted by Trizol reagents (Invitrogen, CA, USA) according to the manufacturer’s instructions. Sixteen samples from two genotypes Golden Promise and WB-1 were used for small RNA library construction. The concentration and quality of total RNA were tested by Bioanalyzer 2100 (Agilent, CA, USA).
Small RNA and degradome libraries construction and sequencing
Small RNA library construction was performed using TruSeq Small RNA Sample Prep Kit (Illumina, San Diego, USA) following the vender’s guidelines. In brief, 1 µg of total RNA was ligated to RNA 3′ and 5′ adapters. Reverse transcript reactions were carried out to create cDNA constructs, and the cDNAs were amplified with two primers that anneal to the ends of the adapters. Then, 140–160 bp (total length of RNA fragment and adapters) products were purified by 6% TBE PAGE Gel (Novex, USA). Finally, the constructed libraries were single-end sequenced (50 bp) by the Illumina Hiseq2500 platform (LC-BIO, Hangzhou, China).
To figure out the target genes for miRNAs, a degradome library was constructed by RNA pools used in the small RNA sequencing based on the methods as described by Ma et al. (2010). Briefly, poly (A)-enriched mRNA was isolated, and annealed with biotinylated random primers. RNAs containing 5′-monophosphates were ligated to a 5′ RNA adapter. Then, the ligated products were used to generate first-strand cDNA by the reverse transcription and the PCR reactions. Finally, we performed single-end sequencing (50 bp) by the Illumina Hiseq2500 as mentioned above.
The raw reads were deposited in the SRA database (http://www.ncbi.nlm.nih.gov/sra/) at NCBI with BioProject accession number PRJNA485436 and SRA accession number SRP157400 (Small RNA sequencing: SRR7687211-SRR7687226; Degradome sequencing: SRR7687210).
Bioinformatics analysis
Raw reads of high-throughput small RNA sequencing were processed by the software ACGT101-miR in the LC-Sciences (Hangzhou, China) to remove the adapter sequences and junk reads through mapping against barley mRNA database (ftp://ftp.ensemblgenomes.org/pub/plants/release-36/fasta/hordeum_vulgare/cds/Hordeum_vulgare.Hv_IBSC_PGSB_v2.cds.all.fa.gz), Rfam (http://rfam.janelia.org) and Repbase (http://www.girinst.org/repbase). The clean sequences with the length of 18–25 nt were blasted against miRBase (version: 21; http://www.mirbase.org). The matched reads mapped to the barley reference genome database(http://plants.ensembl.org/Hordeum_vulgare/Info/Index), which were considered as conserved miRNAs. Meanwhile, the unmatched reads and their flanking 120 nt sequences were used to predict the hairpin structures by RNAfold software (http://rna.tbi.univie.ac. at/cgi-bin/RNAfold.cgi) to identify novel miRNAs.
For degradome analysis, raw reads were obtained by the Illumina Pipeline software (version 1.5), and then filtered out adaptor sequences and low quality reads. The proprietary program ACGT101-DEG v3.1 (LC-BIO, Hangzhou, China) and the public software Cleaveland v3.0 (Addo-Quaye et al. 2009) were used to analyze sequencing data. The degradome reads were mapped to barley mRNA database (ftp://ftp.ensemblgenomes.org/pub/plants/release-36/fasta/hordeum_vulgare/cds/Hordeum_vulgare.Hv_IBSC_PGSB_v2.cds.all.fa.gz), and the annotations for target genes was obtained from the barley CDS database in IPK (http://webblast.ipk-gatersleben.de/barley_ibsc/downloads/160517_Hv_IBSC_PGSB_r1_CDS_HighConf_REPR_annotation.fasta.gz). All target genes were classified as category 0 to 4 in accordance with their abundance relative to the overall profiles of degradome reads (Addo-Quaye et al. 2008, 2009).
Identification of Cd-responsive miRNAs
Cd-responsive miRNAs were identified based on the criteria as decribed by Wu et al. (2018). The abundance of miRNAs was normalized according to Li et al. (2016). The fold change of miRNAs between the Cd-treated and the control samples was calculated as the following formula: fold change = log2N, N = Cd reads/control reads. The miRNAs were up-regulated with log2N ≥ 0.5, down-regulated with log2N ≤ − 0.5.
Results
The difference of Cd tolerance between Golden Promise and WB-1
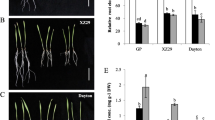
After 10 days of 5 µM Cd treatment, wild barley genotype WB-1 showed much stronger Cd tolerance than the cultivar Golden Promise (Fig. 1a). The dry weights of shoots and roots were reduced much more in Golden Promise than that in WB-1 after Cd treatment, especially 46.5% and 23.7% of the loss for shoot dry weights in Golden Promise and WB-1, respectively (Fig. 1b, c). Correspondingly, WB-1 accumulated much lower Cd concentration in the roots and shoots than Golden Promise, being 21.5% and 59.6% of Cd concentration in Golden Promise, respectively (Fig. 2a, b). In spite of much small biomass in Golden Promise, the Cd uptake by roots was significant larger in Golden Promise than that in WB-1, but it was opposite for Cd translocation from roots to shoots (Fig. 2c, d). It indicates that higher Cd concentration in Golden Promise is due to its stronger Cd uptake in comparison with WB-1. Furthermore, other micrometals also showed lower concentration in WB-1 than in Golden Promise including Mn, Cu, Fe and Zn (Supplemental Fig. S1). These results indicated that WB-1 was more Cd tolerant than Golden Promise, mainly due to its low Cd uptake.
Growth performace of two barley genotypes after Cd treatments. a Plant pictures of Golden Promise and WB-1 under 5 µM Cd and control (CK) conditions for 10 days. Dry weights of shoots (b) and roots (c) of Golden Promise and WB-1 after Cd treatments. Data are means ± SD of three biological replicates; * indicates significant difference at P < 0.05 by Tukey’s test and ** P < 0.01
The difference of Cd concentration, uptake and translocation between two barley genotypes. Cd concentration in roots (a) and shoots (b) of Golden Promise and WB-1 under control (CK) and 5 µM Cd conditions for 10 d was determined by the ICP-MS. (c) Cd uptake by roots and Cd translocation to shoots (d) in Golden Promise and WB-1 under Cd stress were calculated as described in the methods. Data are means ± SD of three biological replicates; * indicates significant difference at P < 0.05 by Tukey’s test and ** P < 0.01
Identification of conserved and novel miRNAs
To reveal molecular difference underlying contrasting Cd tolerance between Golden Promise and WB-1, 16 small RNA libraries from root and shoot tissues of two genotypes under Cd and control conditions were constructed. For small RNA libraries in shoots, 2,144,420, 2,178,580, 1,870,916 and 1,869,353 unique reads in the average were generated in Golden Promise (CK), Golden Promise (Cd), WB-1(CK) and WB-1(Cd), respectively (Supplemental Table 1). After raw reads filtering, 1,072,894, 1,091,675, 1,064,744 and 1,043,284 clean reads were obtained for the corresponding four groups, which were averagely accounted for 53.4% of the raw reads. Meanwhile, for small RNA libraries in roots, 2,325,952, 2,329,086, 3,452,555 and 2,864,046 unique reads in the average, corresponding to 1,353,683, 1,378,257, 2,349,012 and 1,730,848 clean reads were obtained in Golden Promise (CK), Golden Promise (Cd), WB-1(CK) and WB-1(Cd), respectively, averagely accounting for 61.5% of the raw reads (Supplemental Table 1). The length of the clean reads distributed from 18 to 25 nt and 24 nt was the most dominant in all libraries (Supplemental Fig. S2).
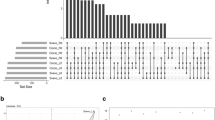
A total of 303 miRNAs including 216 conserved miRNAs belonging to 59 families and 87 novel miRNAs were identified (Fig. 3 and Supplemental Table 2). For the length of these miRNAs, 21 nt miRNAs was the most dominant, which accounted for 63.4% and 49.4% of conserved and novel miRNAs, respectively (Fig. 3c). There were 262 and 255 miRNAs identified in shoot and root libraries, respectively (Fig. 3a, b). Among them, 235 miRNAs were found in both genotypes (Supplemental Table 2). The miR169 family was the largest miRNA families with 21 members identified in this study. Moreover, 6 families (miR156, miR166, miR167, miR171, miR396 and miR1122) were identified with more than 10 miRNAs (Supplemental Table 2).
Genotypic difference of Cd responsive miRNAs in roots
In roots, 45 miRNAs were indentified with differentially expressed levels after Cd treatment including 40 conserved miRNAs mainly belonging to miR156, miR166, miR167, miR169, miR396 families and 5 novel miRNAs (Table 1). For two genotypes, 17 up-regulated, 17 down-regulated and 10 slightly changed miRNAs were identified in Golden Promise, while 17 up-regulated, 11 down-regulated, and 17 slightly changed miRNAs were found in WB-1, respectively (Table 1). Some miRNAs were identified in both genotypes including 8 up-regulated miRNAs (5 families: miR167, miR319, miR394, miR396 and miR397) and 6 down-regulated miRNAs (3 families: miR169, miR172, miR396 and a novel miRNA). Based on degradome analysis and bioinformatics prediction, the target genes of these miRNAs mainly included TCP family transcription factor, growth-regulating factor, elongation factor, cytochrome P450 superfamily protein, laccase, calnexin, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein and disease resistance (Table 1).
On the other hand, there was genotypic difference of Cd responsive miRNAs between two genotypes. There were 7 miRNAs (5 families: miR159, miR166, miR168, miR390 and miR396) up-regulated in WB-1, which were slightly changed in Golden Promise, and these miRNAs target genes mainly included MYB domain protein, homeobox-leucine zipper protein family, receptor-like kinase, etc. Three miRNAs (2 members from miR156 and hvu-miR169c-3p_2) were down-regulated in WB-1, but were not in Golden Promise. The target genes of these miRNAs encoded P-loop containing nucleoside triphosphate hydrolases superfamily protein, structure-specific endonuclease subunit and receptor-like protein kinase (Table 1).
Genotypic difference of Cd responsive miRNAs in shoots
In shoots, 43 miRNAs were differentially expressed in response to Cd stress, including 40 conserved miRNAs mainly belonging to miR156, miR164, miR167, miR169, miR396, miR397 families and 3 novel miRNAs (Table 2). For two genotypes, 15 up-regulated, 16 down-regulated and 11 slightly changed miRNAs were identified in Golden Promise, while 8 up-regulated, 18 down-regulated and 17 slightly changed miRNAs were identified in WB-1, respectively (Table 2). For both genotypes, 3 miRNAs (hvu-miR167c-5p, hvu-MIR397a-3p and hvu-MIR397a-3p) up-regulated and 6 miRNAs (hvu-miR156a-3p_1, hvu-miR164_1 and four miR169 members) down-regulated were found for both genotypes (Table 2). The target genes of these miRNAs mainly included ubiquitin carboxyl-terminal hydrolase, cytochrome P450 superfamily protein, laccase, P-loop containing nucleoside triphosphate hydrolases superfamily protein, NAC domain containing protein, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein, zinc finger protein and elongation factor.
For genotypic difference, hvu-miR390-5p, hvu-miR394-5p, hvu-miR396a-5p_2 and hvu-miR444b_1 were up-regulated in WB-1, but slightly changed in Golden Promise, and the target genes of these miRNAs mainly included receptor-like protein kinase, calnexin, growth-regulating factor and MADS-box transcription factor. Moreover, 7 miRNAs (hvu-miR156a-5p, hvu-miR168-3p, hvu-miR1122b-3p, hvu-miR5168-3p and 3 miRNAs from miR169) were down-regulated in WB-1, but were not in Golden Promise. These miRNAs target genes mainly encoded squamosa promoter-binding-like protein, receptor-like protein kinase and homeobox-leucine zipper protein family. Moreover, hvu-miR167a-5p_1, hvu-miR167c-3p, hvu-miR530-5p, hvu-miR5048a, and PC-miR62 showed oppositely changed pattern between Golden Promise and WB-1, and the target genes of these miRNAs included auxin response factor, receptor kinase, UDP-glucose 4-epimerase and disease resistance protein (Table 2). In addition, hvu-MIR169g-3p targeted to non-specific phospholipase C2 was detected only in WB-1, which was down-regulated after Cd treatment.
Discussion
In this study, we attempted to identify miRNAs involved in Cd tolerance, to reveal their post-transcriptional mechanisms underlying Cd tolerance in barley. Two barley genotypes (cv. Golden Promise and a wild accession WB-1) were grown in hydroponic culture and treated with 5 µM Cd for 10 days. The growth performance of the wild accession WB-1 was much better than that of the cultivar Golden Promise after Cd treatment. Moreover, it was found that Cd concentration was much lower in shoots and roots of WB-1 than that in Golden Promise. These physiological parameters suggest that WB-1 had stronger Cd tolerance than Golden Promise due to its low Cd uptake (Fig. 2). On the other hand, 303 miRNAs were identified by sequencing small RNA libraries from root and shoot tissues of WB-1 and Golden Promise. Then, the miRNA expression profiles were compared between the two genotypes to figure out Cd-responsive miRNAs in barley. Furthermore, to explore the molecular mechanism of these Cd-responsive miRNAs, target genes for these miRNAs were validated by the degradome sequencing.
Previous studies on miRNAs in response to Cd stress were studied in many crops including soybean (Glycine max), tobacco (Nicotiana tabacum L.), Chinese flowering cabbage (Brassica parachinensis L.) and so on (Fang et al. 2013; He et al. 2016; Zhou et al. 2017). In the present study, 40 miRNAs and 5 novel miRNAs were identified in response to Cd stress in barley roots, and 43 Cd responsive miRNAs (40 conserved and 3 novel miRNAs) were detected in barley shoots (Table 2). In addition, several miRNAs were responded differently in Golden Promise or WB-1 after Cd treatment. For instance, there were 11 miRNAs with differentially expressed levels in roots of WB-1, but they were slightly changed in roots of Golden Promise. It indicates that different genotypes may have distinct regulatory mechanisms in response to Cd stress. Similar studies were reported in water spinach (Ipomoea aquatic Forsk.) and pakchoi (Brassica chinensis L.) (Shen et al. 2017a, b; Xue et al. 2014). Therefore, we hypothesized networks of Cd-responsive miRNAs in response to Cd stress in roots and shoots of barley (Supplemental Fig. S3 and S4).
Cd-responsive miRNAs could regulate transcriptional factors (TFs) and gene networks involved in ion homeostasis, plant development and metabolic processes, to enhance Cd tolerance in barley (Supplemental Fig. S3 and S4). GRAS (GAI, RGA and SCR) TFs are important for root development, radial patterning and vascular tissue differentiation (DiLaurenzio et al. 1996; Lim et al. 2000; Pysh et al. 1999). Previously, miR171/GRAS module regulating Cd tolerance in rice and B. napus was reported (Ding et al. 2011; Zhou et al. 2012); while miR393/TIR 1 (transport inhibitor response 1-like protein) module was identified in auxin signaling pathway for mediating root growth and stress responses (Bai et al. 2017; Chen et al. 2011). In this study, hvu-miR171-3p and hvu-miR393-5p were up-regulated in roots of Golden Promise after Cd treatment, but were not in WB-1, which probably caused the worse root growth in Golden Promise under Cd condition. However, hvu-miR159a targeting HvMYB33 was up-regulated in roots of WB-1, but not for Golden Promise. MYB TFs have been reported to be associated with metal tolerance (Shen et al. 2008; Hu et al. 2017). Some miRNAs targeting important genes were involved in metal uptake and accumulation in roots. Three members of the miR166 family targeting HD-Zip (homeodomain-leucine zipper) TFs were up-regulated only in roots of WB-1. In rice, overexpression of miR166 could enhance Cd tolerance and regulate the expression of metal transporters OsHMA2 and OsHMA3 genes (Ding et al. 2018). Thus, we hypothesized that miR159a and miR166 might partially explain the mechanism of Cd tolerance in the roots of WB-1.
In shoots, hvu-miR156a-5p targeting SPL (squamosa promoter-binding-like proteins) was down-regulated in WB-1, but slightly changed in Golden Promise after Cd treatment. SPL TFs are involved in plant development (Cardon et al. 1999). Meanwhile, hvu-miR167a-5p_1 was up-regulated in Golden Promise, but was down-regulated in WB-1 under Cd stress. The target gene of hvu-miR167a-5p_1 encoded ARF6 (auxin response factor 6) protein, which played crucial roles in plant growth and development (Guilfoyle and Hagen 2007). Hence, these two miRNAs may be associated with the difference of shoot performance between two genotypes under Cd stress condition.
Additionally, some Cd-responsive miRNAs were identified with the same changed pattern in Golden Promise and WB-1 after Cd treatment, indicating that these miRNAs might be responsible for common adaptive responses to Cd stress in barley. For instance, miR397 was dramatically increased in roots and shoots of both genotypes after Cd treatment, as well as similar studies in (A) thaliana and (B) parachinensis (Gielen et al. 2016; Zhou et al. 2017). LAC17 (laccase 17), as the target gene of hvu-MIR397a-5p, encodes a kind of copper-containing polyphenol oxidase, which regulates internal Cu homeostasis in plants (Bao et al. 1993; Solomon et al. 1996). Thus, decreasing gene expression of LAC17 may enhance synthesis of essential copper-containing proteins such as SOD (Cu–Zn superoxide dismutase) and AO (ascorbate oxidase), in order to improve plant tolerance to oxidative stresses caused by Cd stress (Dixit et al. 2001; Hegedus et al. 2001; Abdel-Ghany and Pilon 2008). Hvu-miR319a-3p. 2 targeting TCP4 (Teosinte branched1/Cycloidea/proliferating cell factor 4) was up-regulated in roots of both genotypes. Previous studies revealed that TCP was related to cell proliferation and hormone pathways in plants (Danisman et al. 2012; Sarvepalli and Nath 2011a, b; Schommer et al. 2014). Overexpression of osa-miR319a in creeping bentgrass (Agrostis stolonifera) showed more tolerance to abiotic stresses than the wild-type plants (Zhou and Luo 2014). It was suggested that hvu-MIR397a-5p and hvu-miR319a-3p.2 probably played important roles in Cd tolerance for barley.
In conclusion, we compared the miRNAs profiles between wild and cultivated barley with contrasting Cd tolerance based on small RNA sequencing to identify 45 and 43 Cd-responsive miRNAs in roots and shoots. In comparison with cultivar Golden Promise, the wild genotype WB-1 had different responses of Cd-responsive miRNAs including miR156, miR159, miR166, miR167, miR171 and miR393, which might be related to Cd tolerance. The results could provide useful information for revealing molecular regulation mechanism of Cd tolerance in barley.
References
Abdel-Ghany SE, Pilon M (2008) MicroRNA-mediated systemic down-regulation of copper protein expression in response to low copper availability in arabidopsis. J Biol Chem 283:15932–15945
Addo-Quaye C, Eshoo TW, Bartel DP, Axtell MJ (2008) Endogenous siRNA and miRNA targets identified by sequencing of the Arabidopsis degradome. Curr Biol 18:758–762
Addo-Quaye C, Miller W, Axtell MJ (2009) CleaveLand: a pipeline for using degradome data to find cleaved small RNA targets. Bioinformatics 25:130–131
ATSDR (2008) Draft toxicological profile for cadmium. U.S. Department of Health and Human Services, Atlanta
Bai B et al (2017) miR393-mediated auxin signaling regulation is involved in root elongation inhibition in response to toxic aluminum stress in barley. Plant Cell Physiol 58:426–439
Bao W, Omalley DM, Whetten R, Sederoff RR (1993) A laccase associated with lignification in loblolly-pine xylem. Science 260:672–674
Bukhari SAH et al (2015) Genome-wide identification of chromium stress-responsive microRNAs and their target genes in tobacco (Nicotiana tabacum) roots. Environ Toxicol Chem 34:2573–2582
Cardon G, Hohmann S, Klein J, Nettesheim K, Saedler H, Huijser P (1999) Molecular characterisation of the Arabidopsis SBP-box genes. Gene 237:91–104
Chen ZH et al (2011) Regulation of auxin response by miR393-targeted transport inhibitor response protein 1 is involved in normal development in Arabidopsis. Plant Mol Biol 77:619–629
Clemens S, Aarts MGM, Thomine S, Verbruggen N (2013) Plant science: the key to preventing slow cadmium poisoning. Trends Plant Sci 18:92–99
Danisman S et al (2012) Arabidopsis class I and class II TCP transcription factors regulate jasmonic acid metabolism and leaf development antagonistically. Plant Physiol 159:1511–1523
DiLaurenzio L et al (1996) The SCARECROW gene regulates an asymmetric cell division that is essential for generating the radial organization of the Arabidopsis root. Cell 86:423–433
Ding Y, Chen Z, Zhu C (2011) Microarray-based analysis of cadmium-responsive microRNAs in rice (Oryza sativa). J Exp Bot 62:3563–3573
Ding Y et al (2018) MicroRNA166 modulates cadmium tolerance and accumulation in rice. Plant Physiol 177:1691–1703
Dixit V, Pandey V, Shyam R (2001) Differential antioxidative responses to cadmium in roots and leaves of pea (Pisum sativum L. cv. Azad). J Exp Bot 52:1101–1109
Fang X et al (2013) Identification and comparative analysis of cadmium tolerance-associated miRNAs and their targets in two soybean genotypes. PLoS ONE 8:e81471
Ghosh S et al (2017) Insights into the miRNA-mediated response of maize leaf to arsenate stress. Environ Exp Bot 137:96–109
Gielen H, Remans T, Vangronsveld J, Cuypers A (2016) Toxicity responses of Cu and Cd: the involvement of miRNAs and the transcription factor SPL7. BMC Plant Biol 16:145
Guerinot ML (2000) The ZIP family of metal transporters. Biochim Biophys Acta 1465:190–198
Guilfoyle TJ, Hagen G (2007) Auxin response factors. Curr Opin Plant Biol 10:453–460
He Q, Zhu S, Zhang B (2014) MicroRNA-target gene responses to lead-induced stress in cotton (Gossypium hirsutum L.). Funct Integr Genomic 14:507–515
He XY, Zheng WT, Cao FB, Wu FB (2016) Identification and comparative analysis of the microRNA transcriptome in roots of two contrasting tobacco genotypes in response to cadmium stress. Sci Rep 6:32805
Hegedus A, Erdei S, Horvath G (2001) Comparative studies of H2O2 detoxifying enzymes in green and greening barley seedlings under cadmium stress. Plant Sci 160:1085–1093
Hu S, Yu Y, Chen Q, Mu G, Shen Z, Zheng L (2017) OsMYB45 plays an important role in rice resistance to cadmium stress. Plant Sci 264:1–8
Jarup L, Akesson A (2009) Current status of cadmium as an environmental health problem. Toxicol Appl Pharm 238:201–208
Khan A, Khan S, Khan MA, Qamar Z, Waqas M (2015) The uptake and bioaccumulation of heavy metals by food plants, their effects on plants nutrients, and associated health risk: a review. Environ Sci Pollut Res 22:13772–13799
Khan MA, Khan S, Khan A, Alam M (2017) Soil contamination with cadmium, consequences and remediation using organic amendments. Sci Total Environ 601:1591–1605
Khraiwesh B, Zhu J-K, Zhu J (2012) Role of miRNAs and siRNAs in biotic and abiotic stress responses of plants. BBA-Gene Regul Mech 1819:137–148
Kim DY, Bovet L, Maeshima M, Martinoia E, Lee Y (2007) The ABC transporter AtPDR8 is a cadmium extrusion pump conferring heavy metal resistance. Plant J 50:207–218
Li X, Shahid MQ, Wu J, Wang L, Liu X, Lu Y (2016) Comparative small RNA analysis of pollen development in autotetraploid and diploid rice. Int J Mol Sci 17:499
Lim J et al (2000) Molecular analysis of the SCARECROW gene in maize reveals a common basis for radial patterning in diverse meristems. Plant Cell 12:1307–1318
Liu J, Qu W, Kadiiska MB (2009) Role of oxidative stress in cadmium toxicity and carcinogenesis. Toxicol Appl Pharmacol 238:209–214
Ma Z, Coruh C, Axtell MJ (2010) Arabidopsis lyrata small RNAs: transient MIRNA and small interfering RNA Loci within the Arabidopsis genus. Plant Cell 22:1090–1103
Pysh LD, Wysocka-Diller JW, Camilleri C, Bouchez D, Benfey PN (1999) The GRAS gene family in Arabidopsis: sequence characterization and basic expression analysis of the SCARECROW-LIKE genes. Plant J 18:111–119
Sarvepalli K, Nath U (2011a) Hyper-activation of the TCP4 transcription factor in Arabidopsis thaliana accelerates multiple aspects of plant maturation. Plant J 67:595–607
Sarvepalli K, Nath U (2011b) Interaction of TCP4-mediated growth module with phytohormones. Plant Signal Behav 6:1440–1443
Schommer C, Debernardi JM, Bresso EG, Rodriguez RE, Palatnik JF (2014) Repression of cell proliferation by miR319-regulated TCP4. Mol Plant 7:1533–1544
Shen J, Xu X, Li T, Cao D, Han Z (2008) An MYB transcription factor from Malus xiaojinensis has a potential role in iron nutrition. J Integr Plant Biol 50:1300–1306
Shen C et al (2017a) Comparative analysis of cadmium responsive microRNAs in roots of two Ipomoea aquatica Forsk. cultivars with different cadmium accumulation capacities. Plant Physiol Biochem 111:329–339
Shen QF, Fu LB, Qiu L, Feng X, Zhang GP, Wu DZ (2017b) Time-course of ionic responses and proteomic analysis of a Tibetan wild barley at early stage under salt stress. Plant Growth Regul 81:11–21
Siemianowski O, Barabasz A, Kendziorek M, Ruszczynska A, Bulska E, Williams LE, Antosiewicz DM (2014) HMA4 expression in tobacco reduces Cd accumulation due to the induction of the apoplastic barrier. J Exp Bot 65:1125–1139
Solomon EI, Sundaram UM, Machonkin TE (1996) Multicopper oxidases and oxygenases. Chem Rev 96:2563–2605
Tang J, Chu C (2017) MicroRNAs in crop improvement: fine-tuners for complex traits. Nat Plants 3:17077
Uraguchi S, Mori S, Kuramata M, Kawasaki A, Arao T, Ishikawa S (2009) Root-to-shoot Cd translocation via the xylem is the major process determining shoot and grain cadmium accumulation in rice. J Exp Bot 60:2677–2688
Voinnet O (2009) Origin, biogenesis, and activity of plant microRNAs. Cell 136:669–687
Wang Y et al (2015) Identification of radish (Raphanus sativus L.) miRNAs and their target genes to explore miRNA-mediated regulatory networks in lead (Pb) stress responses by high-throughput sequencing and degradome analysis. Plant Mol Biol Rep 33:358–376
Wu DZ, Sato K, Ma JF (2015) Genome-wide association mapping of cadmium accumulation in different organs of barley. New Phytol 208:817–829
Wu DZ, Yamaji N, Yamane M, Kashino-Fujii M, Sato K, Ma JF (2016) The HvNramp5 transporter mediates uptake of cadmium and manganese, but not iron. Plant Physiol 172:1899–1910
Wu LY, Yu JH, Shen QF, Huang L, Wu DZ, Zhang GP (2018) Identification of microRNAs in response to aluminum stress in the roots of Tibetan wild barley and cultivated barley. BMC Genom 19:560
Xue M, Zhou YH, Yang ZY, Lin BY, Yuan JG, Wu SS (2014) Comparisons in subcellular and biochemical behaviors of cadmium between low-Cd and high-Cd accumulation cultivars of pakchoi (Brassica chinensis L.). Front Env Sci 8:226–238
Yu LJ et al (2012) Comparative transcriptome analysis of transporters, phytohormone and lipid metabolism pathways in response to arsenic stress in rice (Oryza sativa). New Phytol 195:97–112
Zhou M, Luo H (2014) Role of microRNA319 in creeping bentgrass salinity and drought stress response. Plant Signal Behav 9:e28700
Zhou ZS, Song JB, Yang ZM (2012) Genome-wide identification of Brassica napus microRNAs and their targets in response to cadmium. J Exp Bot 63:4597–4613
Zhou Q, Yang YC, Shen C, He CT, Yuan JG, Yang ZY (2017) Comparative analysis between low- and high-cadmium-accumulating cultivars of Brassica parachinensis to identify difference of cadmium-induced microRNA and their targets. Plant Soil 420:223–237
Acknowledgements
This study was supported by the National Natural Science Foundation of China (31771685, 31620103912), China Agriculture Research System (CARS-05), Jiangsu Collaborative Innovation Center for Modern Crop Production (JCIC-MCP) and the Fundamental Research Funds for the Central Universities.
Author information
Authors and Affiliations
Contributions
JH Yu and DZ Wu designed the research. JH Yu, LY Wu, LB Fu, QF Shen and LH Kuang performed the research. JH Yu, GP Zhang and DZ Wu analyzed the data. JH Yu and DZ Wu wrote the article.
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yu, J., Wu, L., Fu, L. et al. Genotypic difference of cadmium tolerance and the associated microRNAs in wild and cultivated barley. Plant Growth Regul 87, 389–401 (2019). https://doi.org/10.1007/s10725-019-00479-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-019-00479-1