Abstract
miR393, which is encoded by MIR393a and MIR393b in Arabidopsis, post-transcriptionally regulates mRNAs for the F-box auxin receptors TIR1 (Transport Inhibitor Response Protein 1), AFB1 (Auxin Signaling F-box Protein 1), AFB2 and AFB3. However, biological functions of the miR393-TIR1/AFBs module in auxin response and plant development is not fully understood. In the study herein, we demonstrate that miR393 accumulated in response to exogenous IAA treatment, and its induction was due to enhanced MIR393b transcription but not MIR393a. Overexpression of a miR393-resistant form of TIR1 (mTIR1) enhanced auxin sensitivity and led to pleiotropic effects on plant development including inhibition of primary root growth, overproduction of lateral roots, altered leave phenotype and delayed flowering. Furthermore, miR393 level was increased in 35S:mTIR1 plant, suggesting that TIR1 promoted the expression of miR393 by a feedback loop. The interaction between miR393 and its target indicates a fine adjustment to the roles of the miR393-TIR1 module, which is required for auxin responses in plant development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The microRNAs (miRNAs) are endogenous ~21-nucleotide noncoding RNAs that target complementary mRNA transcripts for cleavage or transcriptional repression (Bartel and Bartel 2003; Bartel 2004; Carrington and Ambros 2003; Mallory and Vaucheret 2004). In plants, miRNAs have been demonstrated to play a crucial role in various biological processes including embryogenesis, flowering, leaf and root development, and plant responses to biotic and abiotic stresses (Llave 2004; Sunkar and Zhu 2004).
The identity of miRNA targets suggests that several miRNAs play a role in auxin signaling. miR393 targets auxin receptors TIR1 and three closely related F-box proteins (Jones-Rhoades and Bartel 2004). The transcripts of several Arabidopsis auxin response factors (ARFs) are either directly or indirectly regulated by miRNAs (Allen et al. 2005; Kasschau et al. 2003; Rhoades et al. 2002). These miRNAs involved in auxin signaling have been shown to modulate plant auxin responses and the expression of auxin-induced genes (Mallory et al. 2005; Wang et al. 2005).
miR393 is a conserved family that has been identified in many plants. In Arabidopsis, this family is encoded by two loci, MIR393a and MIR393b (Jones-Rhoades and Bartel 2004; Jones-Rhoades et al. 2006). Four F-box genes, TIR1 (At3g62980), AFB1 (At4g03190), AFB2 (At3g26810), and AFB3 (At1g12820), have been identified and validated as targets of miR393 (Jones-Rhoades and Bartel 2004; Navarro et al. 2006; Parry et al. 2009). The TIR1/AFBs constitute a small subset of F-box-containing auxin receptors, and function as a component of the Skp1-Cullin1-F-box protein (SCF) ubiquitin ligase complexes. TIR1/AFBs regulate auxin signaling by proteolysis of auxin/indole-3-acetic acids (Aux/IAA) repressors, and by releasing the activities of auxin response factors (ARFs) (Dharmasiri et al. 2005a; Kepinski and Leyser 2005; Ruegger et al. 1998). Previous studies have shown that mutations in TIR1/AFBs genes result in decreased auxin sensitivity in plants. However, only quadruple tir1/afb mutants exhibit a severe embryonic phenotype and a variety of growth defects, which indicates that these genes have overlapping functions, and collectively mediate auxin response during plant development (Dharmasiri et al. 2005b).
It has been shown that TIR1, AFB2, and AFB3 are negatively regulated by miR393 in response to pathogen attack, and overexpression of miR393 results in decreased levels of TIR1 mRNA and enhanced antibacterial resistance (Navarro et al. 2006). More recently, it was reported that the miR393/AFB3 regulatory module controls root system architecture in response to external and internal nitrate availability in Arabidopsis (Vidal et al. 2010).
However, it is not clear whether miR393 is regulated by auxin. To gain insights into miR393 regulation by auxin, we investigated the expression of two MIR393 genes for their responses to auxin treatments in Arabidopsis seedlings. Our results showed that exogenous IAA treatment induced miR393 accumulation resulted from enhanced MIR393b transcription. Overexpression of a miR393-resistant form of TIR1 (mTIR1) enhanced auxin sensitivity and led to pleiotropic effects. In addition, overexpression of mTIR1 could promote miR393 expression by a feedback loop. Our data suggested that the miR393-mediated regulation of TIR1 contributed to regulate auxin responses in plant normal development.
Materials and methods
Plant material, growth conditions and treatments
Arabidopsis thaliana mutant tir1-1 and the tir1-1 afb1-1 afb2-1 afb3-1 quadruple mutant were kindly provided by Mark Estelle (Section of Cell and Developmental Biology, The University of California, San Diego). dcl1-9 [CS3828] was obtained from the ABRC. Both wild-type (Columbia-0, Ws ecotype), mutants and transgenic plants seeds were surface sterilized with 70% ethanol and 10% bleach. Sterilized seeds were sown on B5-agar plates. Plates were vernalized in darkness for 2 days at 4°C and then transferred to a growth chamber at 22°C and 70% humidity under a 16-h-light/8-h-dark photoperiod. After 7–10 days, seedlings were potted in soil and placed under the same conditions.
For root elongation and lateral root assays, 5-day-old seedlings germinated on B5-agar plates were transferred onto new agar medium with or without hormone and grown vertically under constant light for designated times. Lateral root numbers were counted under a dissection microscope. Root elongation was measured using the ImageJ software.
Constructs and generation of transgenic plants
To generate the 35S:MIR393 constructs, a 778- and a 755-bp fragment surrounding the miRNA sequence including the fold-back structure were amplified from genomic DNA with the primers for 35S:MIR393a and 35S:MIR393b, respectively. The amplified fragments were sequenced and cloned downstream of the CaMV (Cauliflower mosaic virus) 35S promoter in pCAMBIA13011.
For the promoter:GUS constructs, 2.0- and 1.5-kb fragments upstream from the predicted fold-back of miR393 were amplified and sequenced. Fragments were double digested by PstI/NcoI and substituted the 35S promoter before GUS exon in pCAMBIA1301 to obtain pMIR393a:GUS and pMIR393b:GUS, respectively. To generate the 35S:TIR1 constructs, the TIR1 (At3g62980) coding sequence was amplified and cloned into the pCAMBIA13011 under a 35S promoter. For a miR393-resistant version of TIR1 (mTIR1), mutations were introduced into the miR393 binding sequence by an overlapping PCR. The resulting product was digested and cloned into pCAMBIA13011.
All of the constructs described were electroporated into Agrobacterium tumefaciens GV3101, and used to transform Arabidopsis by the floral dip method (Clough and Bent 1998). Homozygous T3 seeds were used for further study.
All primers used in this work are listed in Supplementary Table 1. The clones used for vector construction were verified by sequencing.
Gene expression analysis
Total RNA was isolated from plant tissue using TRIzol reagent (Invitrogen, Shanghai, CHN), and treated with RNase-free DNaseI (TakaRa, Dalian, CHN). Treated RNA (1 μg) was used for the first-strand cDNA synthesis using PrimeScript RT Reagent Kit (TaKaRa). Real-time PCR was performed by the LightCycler 480 (Roche) with the SYBR Premix Ex Taq (Perfect Real Time) Kit (Takara). Primers of TIR1 and AFB1/2/3 were designed from each side of the miR393 cleavage site, and relative transcript levels were normalized using UBQ5 as a standard. The primers used for qRT-PCR are also described in Supplemental Table 1.
RNA blot analysis
RNA was extracted using TRIzol reagent (Invitrogen). Total RNA was separated on 1.1% formaldehyde-MOPS agarose gels. TIR1 probe was randomly labeled with digoxigenin-dUTP (Roche) using a cDNA fragment spanning the miR393 cleavage site. For small RNAs, low molecular weight RNA was separated on 17% polyacrylamide gels under denaturing conditions (7 M urea). Blots were hybridized using either radioactively labeled or digoxigenin end-labeled locked nucleic acid (LNA) oligonucleotide probes with the sequence 5′-gAtcAatGcgAtcCctTtgGa-3′ (capital alphabets for LNA).
Histochemical detection of GUS activity
Histochemical localization of GUS staining was performed by incubating seedlings in a solution of 1 mg mL−1 5-bromo-4-chloro-3-indolyl-β-d-glucuronic acid, 1 mM potassium ferricyanide, 0.1% Triton X-100, 0.1 M sodium phosphate buffer, pH 7.0 and 10 mM EDTA overnight at 37°C, followed by clearing with 70% ethanol.
Results
Auxin enhanced MIR393b expression
The miR393 levels in wild-type seedlings were determined to evaluate the effects of exogenous IAA treatments. Northern blot analyses showed that miR393 was induced at IAA concentrations of 10 and 50 μM. The increased level of miR393 was apparent after 12 h of exposure to 10 and 50 μM IAA, and the level continued to increase by approximately threefold after 24 h exposure (Fig. 1a).
Auxin treatment enhanced MIR393b expression. a RNA gel blot analysis of miR393 in response to exogenous IAA treatment. 10-day-old wild-type seedlings were transferred to liquid B5 medium containing IAA, and seedlings were harvested after 6, 12, and 24 h of treatment. 10 μg low molecular RNA was loaded and rRNA was stained by EtBr as loading control. Numbers indicate fold change relative to control sample. b qRT-PCR analysis of precursor transcripts of MIR393 family members in response to exogenous IAA. Quantifications were normalized to the expression of UBQ5. The relative expression levels in control plants were set to 1.0. Error bars represent SD from three PCR results, and similar results were obtained in two independent experiments. Asterisks denote significantly different from the control plants (P < 0.05, Student’s t test). c Response of MIR393a and MIR393b promoter:GUS to exogenous IAA treatment. 10-day-old homozygous transgenic seedlings grown on B5-agar plates were transferred to new agar medium with or without 10 μM IAA. Seedlings were stained for GUS activity after 12 h of treatment. d Quantification of GUS expression in response to IAA treatments in pMIR393a:GUS and pMIR393b:GUS transgenic plants. Two representative transgenic lines were analyzed. Error bars represent SD. Asterisks denote significantly different from the control MIR393 promoter report lines (P < 0.05, Student’s t test)
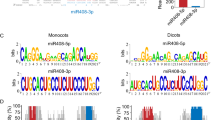
MIR393a and MIR393b produce two identical mature miR393. To determine whether the responsiveness of MIR393 to IAA is locus-specific, we measured precursor abundance of MIR393a and MIR393b. The precursor transcripts of MIR393b, but not those of MIR393a, were elevated under the IAA treatment (Fig. 1b). We also generated promoter:GUS transgenic lines with ~2-kb putative promoter sequences upstream of the predicted fold-back structure of MIR393a and MIR393b. In transgenic seedlings with no IAA treatment, GUS activity was observed in the shoot apical meristem, and vascular bundles of roots and leaves. After a 12 h treatment with 10 μM IAA, only pMIR393b:GUS transgenic seedlings showed an increase of GUS intensity, especially in leaves and hypocotyls (Fig. 1c). qRT-PCR analysis of GUS expression revealed a twofold increase in two pMIR393b:GUS transgenic lines after 12 h of 10 μM IAA treatment (Fig. 1d). These data indicated that the increased level of miR393 primary transcripts after the IAA treatment most probably due to the enhanced transcription of MIR393b.
To elucidate the effect of miR393 accumulation, we also analyzed mRNA levels of TIR1, AFB1, AFB2 and AFB3 in Arabidopsis seedlings after exogenous IAA treatment. The results of real time RT-PCR did not show any remarkable changes in TIR1/AFBs transcripts after 12 h treatment with 10 μM IAA (Fig. 2a). Next, we examined the transcription of TIR1/AFBs using our previously described reporter lines TIR1/AFBs promoter:GUS (She et al. 2010). Quantitative GUS expression is shown in Fig. 2b. We found that the activities of the TIR1 and AFB1 promoters were elevated about 1.6-fold after 10 μM IAA treatment, indicating exogenous IAA increased transcription of TIR1 and AFB1. GUS staining also showed that exogenous IAA was able to transcriptionally induce expression of TIR1 in roots of pTIR1:GUS seedlings after 0.5 and 6 h treatments (Fig. 2c). According to these data, we proposed that the posttranscriptional repression of TIR1 by miR393 might contribute to the homeostasis of TIR1 mRNA in response to a high concentration of IAA treatment.
miR393 posttranscriptionally regulates TIR1/AFBs transcripts in response to exogenous IAA. a qRT-PCR analysis of TIR1/AFBs transcripts in response to auxin treatments. 10-day-old transgenic seedlings grown on B5-agar plates were treated with 10 μM IAA for 12 h. Quantifications were normalized to the expression of UBQ5. The relative expression levels in control plants were set to 1.0. Error bars represent SD from three PCR results, and similar results were obtained in two independent experiments. b Quantification of GUS expression in response to IAA treatments in pTIR1:GUS, pAFB1:GUS, pAFB2:GUS, and pAFB3:GUS transgenic plants. 10-day-old homozygous transgenic seedlings grown on B5-agar plates were transferred to new agar medium with or without 10 μM IAA. Error bars represent SD from three PCR results. c Response of TIR1 promoter:GUS to exogenous IAA treatment. Seedlings were stained for GUS activity after 0.5 or 6 h of 10 μM IAA treatment
Interaction of miR393 and TIR1 modulates plant auxin sensitivity
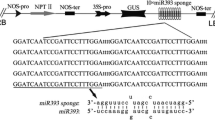
To investigate the role of miR393, we generated transgenic Arabidopsis plants harboring constructs of 35S:MIR393, 35S:TIR1 and 35S:mTIR1 (a miR393-resistant form of TIR1). Northern blot data showed more than threefold increases of miR393 accumulation in these transgenic 35S:MIR393a and 35S:MIR393b lines (Fig. 3a). Using qRT-PCR analysis, we detected moderate decrease of the TIR1 F-box family mRNA levels in all of these lines, which indicated the same effects of At-MIR393a and At-MIR393b overproduction (Fig. 3b). The magnitude of the decrease in their mRNA levels varied among different F-box genes, and AFB1 exhibited a partial resistance to miR393. For the 35S:mTIR1 construct, we introduced six mutations into the miR393 target site in the TIR1 coding sequence without alteration of the corresponding amino acid sequence ETMRSLW (Fig. 3c). Northern blot analysis revealed a significant increase of TIR1 mRNA abundance in 35S:TIR1 and a further doubling of the mRNA levels in 35S:mTIR1 (Fig. 3d). An introduced point mutation in mTIR1 also created an NsbI restriction site, which allowed us to distinguish RT-PCR products of mTIR1 transcripts from those of TIR1 (Supplemental Fig. 1). Our results suggested that release of mTIR1 transcripts from miR393 cleavage resulted in over-accumulation of the full-length transcripts, and confirmed that TIR1 mRNA is under negative regulation of miR393.
Altered TIR1 levels mediated by miR393 regulate plant auxin sensitivity. a Overexpression of miR393 in transgenic Arabidopsis plants. RNA gel blot analysis of miR393 levels in the wild-type and two representative transgenic lines. The blots also included RNA from the dcl1-9 mutant as a negative control. b Detection of TIR1/AFBs transcripts in 35S:MIR393 transgenic plants by real-time RT-PCR. Quantifications were normalized to the expression of UBQ5. The relative expression levels in wild-type plants were set to 1.0. Error bars represent SD from three PCR results, and similar results were obtained in two independent experiments. a, b, and c denote significantly different from wild-type Col-0 (P < 0.05, Student’s t test) for TIR1, AFB2 and AFB3, respectively. c Diagram of mTIR1 expression constructs. The introduced point mutations in the miR393 target site of TIR1 are shown. Introduced point mutation in the miR393 complementary sequence creates an NsbI site and is underlined. d RNA gel blot analysis of TIR1 mRNA. 40 μg total RNA extracted from 35S:TIR1, 35S:mTIR1 and wild-type plants were loaded and rRNAs on blot membrane were visualized by methylene blue trihydrate staining as a loading control. e Effects of decreased TIR1 levels on plant auxin sensitivity. 5-day-old seedlings on B5-agar plates were transferred to new agar medium containing IAA. Root length was measured after 5 days growth. Error bars represent SD. There are significant differences between wild-type and transgenic lines as well as tir1-1 under 0, 25, 50 and 100 nM IAA treatments, as determined by ANOVA (P < 0.05). f Effects of increased TIR1 levels on plant auxin sensitivity. Root length was measured after 5 days growth. Error bars represent SD. For 35S:mTIR1, there are significant differences from wild-type at all IAA concentration points (P < 0.01). For 35S:TIR1, only at 25 and 100 nM IAA (P < 0.05), as determined by ANOVA
To assess auxin responses, we determined the effect of exogenous IAA on root elongation. 5-day-old seedlings were transferred onto new B5-agar plates containing different concentrations of IAA, and primary root length was measured after 5 days. Both 35S:MIR393 and tir1-1 displayed reduced inhibition of primary root growth compared to the control (Fig. 3e). Opposite results have been obtained with 35S:TIR1 and 35S:mTIR1 seedlings. Root elongation of 35S:TIR1 plants were slightly shorter than wild-type under 25 and 50 nM IAA, but became significantly shorter compared with Col-0 under 100 nM IAA condition (P < 0.05, ANOVA) (Fig. 3f). Further, 35S:mTIR1 plants displayed an additive increase in auxin sensitivity compared to 35S:TIR1, so that root elongation was inhibited strongly even under low concentration of 25 nM IAA. These results demonstrated that overexpression of miR393 led to decreased TIR1 levels and reduced plant auxin sensitivity, while overexpression of mTIR1 enhanced auxin sensitivity. These data indicated that the balance between miR393 and TIR1 expression regulated auxin responses.
Altered TIR1 levels mediated by miR393 affect early auxin-responsive gene expression
It has been well known that primary auxin-response gene transcription can be activated by exogenous IAA (Hagen and Guilfoyle 2002). The changes of auxin sensitivity regulated by the miR393-TIR1 module prompted us to determine whether these transgenic lines altered the expression of auxin-responsive genes. We used qRT-PCR to monitor the transcripts of three primary auxin-response genes DFL1/GH3.6 (At5g54510), AXR5/IAA1 (At4g14560) and MSG2/IAA19 (At3g15440). The basal expression of all of these genes were slightly repressed in miR393 overexpression lines, and further repressed in tir1-1 and tir1/afb quadruple seedlings. In contrast, we observed slightly increased transcripts of these genes in 35S:TIR1 lines and a further increase in 35S:mTIR1 (Fig. 4a–c, black columns).
Transcript levels of three primary auxin-responsive genes in response to exogenous IAA. Relative mRNA levels of GH3.6 a, IAA1 b and IAA19 c were measured by qRT-PCR. 10-day-old seedlings grown on B5-agar plates were transferred to B5 liquid medium containing 10 μM IAA for 12 h. Quantifications were normalized to the expression of UBQ5 and the relative mRNA levels in control Col-0 plants were set to 1.0. Error bars represent SD from three PCR results, and similar results were obtained in two independent experiments
To reveal the effect of the miR393-TIR1 regulatory module on auxin response, we tested the expression levels of these genes under exogenous IAA treatment. After 12 h treatment with 10 μM IAA, these genes exhibited substantial changes in transcript abundance in all of these seedlings compared with their own control samples respectively. It showed that 35S:MIR393 inhibited the upregulation of the three auxin responsive genes compared with wild-type, while 35S:TIR1 and 35S:mTIR1 enhanced the expression of these genes when exogenous IAA was applied (white columns). These data suggested that miR393-mediated TIR1 regulation influenced the expression of some primary auxin-responsive genes, which contributed to the alteration of plant auxin sensitivity.
Ectopic expression of miR393-resistant TIR1 causes pleiotropic effects
To determine the in vivo consequences of disrupting miR393 regulation, phenotypic analysis was performed in 35S:mTIR1 transgenic plants. Ectopic overexpression of mTIR1 led to pleiotropic effects including shorter primary roots, more lateral roots, altered leaf phenotype and delayed flowering time, while overexpression of wild-type TIR1 had mild effects on morphology.
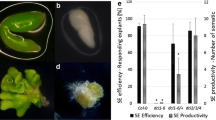
We measured the primary root length of 5-day-old transgenic 35S:MIR393, 35S:TIR1 and 35S:mTIR1 seedlings and the lateral root numbers of 10-day-old transgenic seedlings under long day conditions on vertically oriented plates. It showed that transgenic 35S:TIR1 and 35S:mTIR1 seedlings exhibited phenotypes opposite to those of 35S:MIR393 lines, as well as tir1-1 lines. Compared with wild-type plants, 35S:MIR393 seedlings displayed slightly longer primary root and fewer lateral roots, while 35S:TIR1 seedlings had shorter primary roots and more lateral roots (Table 1, Fig. 5). Furthermore, 35S:mTIR1 exhibited more dramatic changes in both average root length and lateral root number than 35S:TIR1. Along with the observation that 35S:mTIR1 accumulated more TIR1 transcripts than 35S:TIR1, our results provided evidence that regulation of TIR1 mRNA levels by miR393 is an important control point for primary root elongation and lateral root initiation.
We also showed that 5-day-old 35S:TIR1 seedlings displayed downward cotyledons and upward petioles, while cotyledons in 35S:mTIR1 were more severely curled (Fig. 6a–c). 10-day-old 35S:mTIR1 but not 35S:TIR1 plants produced downward curly true leaves with long and twisted petioles (Fig. 6d–f). 4-week-old 35S:mTIR1 plants displayed extremely curled leaves with margins bending to the abaxial surface. Furthermore, the rosette leaves from 35S:mTIR1 were much smaller in size and fewer in number than that of wild-type plants, while 35S:TIR1 plants had normal rosette leaves (Fig. 6g–j). After 8-weeks growth, 35S:mTIR1 plants displayed an obvious delay in bolting and flowering time, whereas the 35S:TIR1 seedlings displayed no developmental anomalies except for reduced silique numbers (Fig. 6k). Moreover, we found that 35S:mTIR1 plants exhibited strong apical dominance, with only one shorter shoot observed in all transgenic lines. Meanwhile, the plant vegetative phase was prolonged in 35S:mTIR1 lines (Fig. 6k).
Ectopic expression of miR393-resistant TIR1 causes pleiotropic effects. a–c 5-day-old 35S:mTIR1 transgenic plants displayed downward cotyledons and upward petioles. Seedlings of wild-type, 35S:TIR1 and 35S:mTIR1 plants grown on B5-agar plates were photographed. d–f 10-day-old 35S:mTIR1 plants displayed downward curly true leaves with long and twisted petioles. Wild-type, 35S:TIR1 and 35S:mTIR1 plants grown in pots were photographed. g–i The rosette leaves from 4-week-old 35S:mTIR1 were much smaller in size and fewer in number. Wild-type, 35S:TIR1 and 35S:mTIR1 transgenic plants grown under short-day conditions were photographed with side view (left) and top view (right). j Margins of 35S:mTIR1 rosette leaves were bending toward the abaxial surface. Leaves from 4-week-old wild-type, 35S:TIR1 and 35S:mTIR1 transgenic plants were cut and arranged in the order of appearance. Both sides of 35S:mTIR1 leaves are shown. Scale bar = 1 cm. k 8-week-old 35S:mTIR1 plants exhibited strong apical dominance
In 35S:MIR393 plants, no other changes of development and growth were observed, except that the time producing the first leaf with the serrate margins was slightly delayed (Supplemental Fig. 2).
Possible feedback regulation of miR393 expression
Feedback loops in which miRNA-regulated genes regulate the transcription of their miRNA have been described in a number of animals and plants (Kim et al. 2007; Varghese and Cohen 2007; Wu et al. 2009). In order to determine whether miR393 is regulated by its target TIR1, we analyzed the effect of TIR1 level on accumulation of mature miR393 using 35S:TIR1, 35S:mTIR1, tir1-1 and tir/afb quadruple mutants. The level of miR393 was elevated about 1.5-fold in 10-day-old 35S:mTIR1 but not in 35S:TIR1 plants. Conversely, it was reduced about 30% in tir1-1 and tir/afb quadruple mutants (Fig. 7). These data demonstrated that miR393 was positively regulated by the level of TIR1, and the expression of TIR1 was modulated by a negative feedback loop.
Discussion
Auxin regulation of miR393 expression
Several plant miRNAs have been predicted to regulate mRNAs involved in auxin signaling pathway. To date, however, only miR164 and miR390 have been reported to response to IAA (Guo et al. 2005; Marin et al. 2010; Yoon et al. 2010). In this work, we show that miR393 in Arabidopsis seedlings can be induced by exogenous IAA application. Northern blot analysis revealed a 2- to threefold increase of miR393 accumulation after 12 h IAA exposure. Analysis of precursor transcripts and promoter:GUS reporter lines confirmed that miR393 accumulation could be controlled at the transcriptional level. Unlike most auxin-responsive genes, which can rapidly regulate their transcription after a short period of treatment, the level of miR393 increased only at high IAA concentration and a long time course. The slower kinetics of miR393 induction may create a homeostatic mechanism to maintain the transcripts of TIR1/AFBs in response to high concentration of IAA treatment.
Another feature of miR393 accumulation in response to auxin is the locus-specific control of its transcription. There are two MIR393 loci: MIR393a (At2g39885) and MIR393b (At3g55734), which produce two identical mature miR393. Recent studies provided evidence that closely related miRNAs, which are predicted to target the same genes, have in fact different functions during development or stress responses (Li et al. 2008; Sieber et al. 2007). Previous studies showed that a flagellin-derived peptide induces miR393 accumulation, which results from increased transcription of MIR393a (Navarro et al. 2006). Our results indicated that MIR393b specifically induced their transcription in response to exogenous auxin treatment. Since the expression of the MIR393 family was under the control of their corresponding promoters, we proposed that the locus-specific control of MIR393 transcription can provide an additional layer of regulation for the auxin signaling network through repressing target gene expression.
The interaction between miR393 and TIR1
According to the results of 5′-RACE assay, four F-box genes, TIR1, AFB1, AFB2, as well as AFB3 have been validated to be the targets of miR393 (Jones-Rhoades and Bartel 2004; Navarro et al. 2006). In the present study herein, we showed that overexpression of miR393 decreased TIR1 mRNA level, while miR393 resistant form of TIR1 (mTIR1) resulted in over-accumulation of the mTIR1 transcripts. These data further confirmed that TIR1 mRNA is under negative regulation by miR393.
The regulation of TIR1 mediated by miR393 can also be observed during plant development. Using promoter:GUS analysis, it has been found that TIR1, AFB2, and AFB3 are broadly transcribed throughout the plant, while their corresponding GUS fusion proteins are restricted to the primary and lateral root tips, young leaves and young flower buds, indicating significant posttranscriptional regulation of these genes (Dharmasiri et al. 2005b; Parry et al. 2009). As a prime candidate for this regulation, it is reasonable to propose that a developmentally based regulation of miR393 production, which in turn controls the level of its target gene transcripts, is required for the auxin-related developmental processes.
Negative and positive feedback loops of miRNA/target regulons have been described for the miRNA homeostasis regulation in plants (Marin et al. 2010; Rajagopalan et al. 2006; Wu et al. 2009). In Arabidopsis, miR172b and miR156a are also found to be positively regulated by the their targets (Wu et al. 2009). Interestingly, we also observed that the accumulation of miR393 was increased in 35S:mTIR1 plants, and was reduced in tir1-1 and tir/afb quadruple mutants (Fig. 7), though the related mechanism was unclear. We tested the TIR1 transcript levels in 35S:TIR1, 35S:mTIR1 and tir1-1 plants and found that the TIR1 level in tir1-1 plants was not obviously different from the level in wild-type (Supplementary Fig. 3). According to the previous reports, the tir1-1 mutant, which is caused by a glycine to aspartate substitution at position 147, is affected in auxin response (Ruegger et al. 1998). We proposed that the change of miR393 level might be due to deficient function of TIR1 protein.
Function of the miR393-TIR1 regulatory module
Auxin is a vital hormone that regulates many aspects of plant development. Individual cells interpret auxin largely by a signaling pathway that involves the F-box protein TIR1 as an auxin receptor. Auxin-dependent TIR1 activity leads to ubiquitination-based degradation of Aux/IAAs, and triggers predefined changes in developmental programs (Vanneste and Friml 2009).
It is well known that auxin can inhibit root growth and promote lateral root production. We analyzed the effect of overexpressing miR393 and miR393-resistant mTIR1 on root development. 35S:MIR393 seedlings displayed slightly longer primary root and fewer lateral roots, while 35S:mTIR1 and 35S:TIR1 seedlings had shorter primary roots and more lateral roots. These data suggested miR393-mediated TIR1 regulation was critical for plant root development.
In addition to the root phenotypes, we did not observe apparent developmental defects in our 35S:MIR393 plants, which were exhibited by tir1/afb mutants. Similar results have been reported by other researchers (Jones-Rhoades and Bartel 2004; Navarro et al. 2006). Our further analysis revealed that overexpression of miR393 only decreased TIR1, AFB2 and AFB3 transcripts, but not AFB1. A plausible explanation is that AFB1, one member of the auxin receptor F-box family, is partially resistant to miR393-guided negative regulation (Navarro et al. 2006). Since the TIR1/AFBs function in a redundant fashion to mediate auxin response, it is not surprising to find that 35S:MIR393 plants displayed no dramatic defects in development.
On the other hand, plants expressing a miR393-resistant version of TIR1 had dramatically increased TIR1 mRNA levels and altered accumulation of many auxin responsive genes when compared with 35S:TIR1. These expression changes were correlated with developmental abnormalities such as downward curly leaves, dwarfed stature, as well as growth retardation. Our results differ from a previous report describing the expression of a mTIR1 with four mutations, which has no effect on the pattern of GUS staining and plant phenotypes. However, no convincing data had been shown to explain the same effect of pTIR1:mTIR1-GUS as pTIR1:TIR1-GUS (Parry et al. 2009). In order to avoid the effect of constitutively expressed promoter, we have also generated the pTIR1:mTIR1 (in tir1-1 background) transgenic lines and have got 10 independent T1 transgenic plants. We found that 6 of them displayed the pleiotropic developmental defects, including downward curled leaves and upward petioles (supplemental Fig 4).
Furthermore, our 35S:mTIR1 plants phenocopy the artificial target mimics plants (MIM 393) reported by Weigel Lab (Todesco et al. 2010). In plant, miRNA target mimicry is an endogenous mechanism used to negatively regulate the activity of a specific miRNA family, through the production of a false target transcript that can not be cleaved (Todesco et al. 2010). Up to now, this technique has been successfully exploited to the study of miRNA functions (Gou et al. 2011; Wang et al. 2008, 2009; Wu et al. 2009). These data further indicates the important role of miR393 in plant development.
In contrast, no obvious phenotypic changes were observed in the 35S:TIR1 transgenic lines, though the mRNA level of TIR1 was 7–30-fold higher than that of wild-type (Supplementary Table 2). This is probably because transgene-derived TIR1 mRNAs are still repressed by relative abundant endogenous miR393. Our results implied that disrupting miR393-directed TIR1 regulation rather than an extra copy of TIR1 gave rise to severe developmental consequences in 35S:mTIR1 lines. In order to explain the pleiotropic effects in 35S:mTIR1, we examined the genome-wide response in 10-day-old seedlings by Affymetrix ATH1 microarrays. Using a per-gene variance of log2 (fold change) >1 and FDR (Q value) <0.01, we found 291 genes were upregulated and 72 genes were downregulated (Supplemental Table 3). These data demonstrated that ectopic expression of a miR393-resistant TIR1 resulted in the collective misregulation of many downstream genes, suggesting the function of the miR393-TIR1 module is critical for interpreting local auxin signals which are required for proper plant development.
In summary, we propose a model for the regulation of auxin response by miR393/TIR1 (Fig. 8). Auxin triggers a transcriptional induction of AtMIR393b, leading to increased miR393 level. The posttranscriptional repression of TIR1/AFBs genes by miR393 alters plant auxin responses. Meanwhile, high level of TIR1 promotes miR393 expression by a feedback loop. The specific regulation of MIR393 expression and the interaction between miR393 and its target indicates a fine adjustment to the roles of the miR393-TIR1 module, which is required for auxin responses in plant development. Elucidation of the mechanism of the miR393-TIR1 regulatory module and their eventual functions is still an important task for future research.
The model for the regulation of auxin response by miR393/TIR1. The schematic diagram illustrates that auxin triggers a transcriptional induction of AtMIR393b, leading to increased miR393 levels. The posttranscriptional repression of TIR1/AFBs genes by miR393 alters plant auxin responses. The possible feedback regulation between TIR1 and miR393 expression provides a homeostatic model in which miR393 and TIR1 abundance are tightly regulated
References
Allen E, Xie Z, Gustafson AM, Carrington JC (2005) microRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell 121(2):207–221
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116(2):281–297
Bartel B, Bartel DP (2003) MicroRNAs: at the root of plant development? Plant Physiol 132(2):709–717
Carrington JC, Ambros V (2003) Role of microRNAs in plant and animal development. Science 301(5631):336–338
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16(6):735–743
Dharmasiri N, Dharmasiri S, Estelle M (2005a) The F-box protein TIR1 is an auxin receptor. Nature 435(7041):441–445
Dharmasiri N, Dharmasiri S, Weijers D, Lechner E, Yamada M, Hobbie L, Ehrismann JS, Jurgens G, Estelle M (2005b) Plant development is regulated by a family of auxin receptor F box proteins. Dev Cell 9(1):109–119
Gou JY, Felippes FF, Liu CJ, Weigel D, Wang JW (2011) Negative regulation of anthocyanin biosynthesis in Arabidopsis by a miR156-targeted SPL transcription factor. Plant Cell 23(4):1512–1522
Guo HS, Xie Q, Fei JF, Chua NH (2005) MicroRNA directs mRNA cleavage of the transcription factor NAC1 to downregulate auxin signals for Arabidopsis lateral root development. Plant Cell 17(5):1376–1386
Hagen G, Guilfoyle T (2002) Auxin-responsive gene expression: genes, promoters and regulatory factors. Plant Mol Biol 49(3–4):373–385
Jones-Rhoades MW, Bartel DP (2004) Computational identification of plant microRNAs and their targets, including a stress-induced miRNA. Mol Cell 14(6):787–799
Jones-Rhoades MW, Bartel DP, Bartel B (2006) MicroRNAS and their regulatory roles in plants. Annu Rev Plant Biol 57:19–53
Kasschau KD, Xie Z, Allen E, Llave C, Chapman EJ, Krizan KA, Carrington JC (2003) P1/HC-Pro, a viral suppressor of RNA silencing, interferes with Arabidopsis development and miRNA function. Dev Cell 4(2):205–217
Kepinski S, Leyser O (2005) The Arabidopsis F-box protein TIR1 is an auxin receptor. Nature 435(7041):446–451
Kim J, Inoue K, Ishii J, Vanti WB, Voronov SV, Murchison E, Hannon G, Abeliovich A (2007) A microRNA feedback circuit in midbrain dopamine neurons. Science 317(5842):1220–1224
Li WX, Oono Y, Zhu J, He XJ, Wu JM, Iida K, Lu XY, Cui X, Jin H, Zhu JK (2008) The Arabidopsis NFYA5 transcription factor is regulated transcriptionally and posttranscriptionally to promote drought resistance. Plant Cell 20(8):2238–2251
Llave C (2004) MicroRNAs: more than a role in plant development? Mol Plant Pathol 5(4):361–366
Mallory AC, Vaucheret H (2004) MicroRNAs: something important between the genes. Curr Opin Plant Biol 7(2):120–125
Mallory AC, Bartel DP, Bartel B (2005) MicroRNA-directed regulation of Arabidopsis AUXIN RESPONSE FACTOR17 is essential for proper development and modulates expression of early auxin response genes. Plant Cell 17(5):1360–1375
Marin E, Jouannet V, Herz A, Lokerse AS, Weijers D, Vaucheret H, Nussaume L, Crespi MD, Maizel A (2010) miR390, Arabidopsis TAS3 tasiRNAs, and their AUXIN RESPONSE FACTOR targets define an autoregulatory network quantitatively regulating lateral root growth. Plant Cell 22(4):1104–1117
Navarro L, Dunoyer P, Jay F, Arnold B, Dharmasiri N, Estelle M, Voinnet O, Jones JD (2006) A plant miRNA contributes to antibacterial resistance by repressing auxin signaling. Science 312(5772):436–439
Parry G, Calderon-Villalobos LI, Prigge M, Peret B, Dharmasiri S, Itoh H, Lechner E, Gray WM, Bennett M, Estelle M (2009) Complex regulation of the TIR1/AFB family of auxin receptors. Proc Natl Acad Sci USA 106(52):22540–22545
Rajagopalan R, Vaucheret H, Trejo J, Bartel DP (2006) A diverse and evolutionarily fluid set of microRNAs in Arabidopsis thaliana. Genes Dev 20(24):3407–3425
Rhoades MW, Reinhart BJ, Lim L, Burge CB, Bartel B, Bartel DP (2002) Prediction of plant microRNA targets. Cell 110(4):513–520
Ruegger M, Dewey E, Gray WM, Hobbie L, Turner J, Estelle M (1998) The TIR1 protein of Arabidopsis functions in auxin response and is related to human SKP2 and yeast Grr1p. Genes Dev 12(2):198–207
She W, Lin W, Zhu Y, Chen Y, Jin W, Yang Y, Han N, Bian H, Zhu M, Wang J (2010) The gypsy insulator of Drosophila melanogaster together with its binding protein Su(Hw) (Suppressor of Hairy-wing) facilitate high and precise expression of transgenes in Arabidopsis thaliana. Genetics 185(4):1141–1150
Sieber P, Wellmer F, Gheyselinck J, Riechmann JL, Meyerowitz EM (2007) Redundancy and specialization among plant microRNAs: role of the MIR164 family in developmental robustness. Development 134(6):1051–1060
Sunkar R, Zhu JK (2004) Novel and stress-regulated microRNAs and other small RNAs from Arabidopsis. Plant Cell 16(8):2001–2019
Todesco M, Rubio-Somoza I, Paz-Ares J, Weigel D (2010) A collection of target mimics for comprehensive analysis of microRNA function in Arabidopsis thaliana. PLoS Genet 6(7):e1001031
Vanneste S, Friml J (2009) Auxin: a trigger for change in plant development. Cell 136(6):1005–1016
Varghese J, Cohen SM (2007) microRNA miR-14 acts to modulate a positive autoregulatory loop controlling steroid hormone signaling in Drosophila. Genes Dev 21(18):2277–2282
Vidal EA, Araus V, Lu C, Parry G, Green PJ, Coruzzi GM, Gutierrez RA (2010) Nitrate-responsive miR393/AFB3 regulatory module controls root system architecture in Arabidopsis thaliana. Proc Natl Acad Sci USA 107(9):4477–4482
Wang JW, Wang LJ, Mao YB, Cai WJ, Xue HW, Chen XY (2005) Control of root cap formation by microRNA-targeted auxin response factors in Arabidopsis. Plant Cell 17(8):2204–2216
Wang JW, Schwab R, Czech B, Mica E, Weigel D (2008) Dual Effects of miR156-Targeted SPL genes and CYP78A5/KLUH on plastochron length and organ size in Arabidopsis thaliana. Plant Cell 20(5):1231–1243
Wang J, Czech B, Weigel D (2009) miR156-regulated SPL transcription factors define an endogenous flowering pathway in Arabidopsis thaliana. Cell 138(4):738–749
Wu G, Park MY, Conway SR, Wang JW, Weigel D, Poethig RS (2009) The sequential action of miR156 and miR172 regulates developmental timing in Arabidopsis. Cell 138(4):750–759
Yoon EK, Yang JH, Lim J, Kim SH, Kim SK, Lee WS (2010) Auxin regulation of the microRNA390-dependent transacting small interfering RNA pathway in Arabidopsis lateral root development. Nucleic Acids Res 38(4):1382–1391
Acknowledgments
We sincerely thank Dr. Mark Estelle (Section of Cell and Developmental Biology, UCSD) for kindly providing the tir1-1 and tir1-1 afb1-1 afb2-1 afb3-1 quadruple mutants and Dr. C-Y Huang (The University of Adelaide, Australia) for helpful discussions and critical reading the manuscript. This work was supported by the National Science Foundation of China (Grant No. 30571197, No. 30972016), National High Technology Research and Development Program of China (863 Program) (No. 2007AA10Z141) and Zhejiang Provincial Natural Science Foundation of China (Grant No. Y3080323).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
11103_2011_9838_MOESM1_ESM.tif
Supplementary material 1 (TIFF 1991 kb). Supplemental Figure 1. Determination of endogenous TIR1 in wild-type, 35S:mTIR1 and 35S:TIR1 transgenic lines. Endogenous and miR393-resistant TIR1 (mTIR1) transcripts were amplified by RT-PCR and distinguished by digestion with the restriction enzyme NsbI, which cuts only mTIR1
11103_2011_9838_MOESM2_ESM.tif
Supplementary material 2 (TIFF 3279 kb). Supplemental Figure 2. Rosette leaves of 4-week-old 35S:MIR393 plants. a The first leaf with serrate margins was slightly delayed in 35S:MIR393 plants. Leaves from 4-week-old wild-type, 35S:MIR393a and 35S:MIR393b transgenic plants were cut and arranged in the order of appearance. Scale bar = 1 cm. b Rosette leaves of 4-week-old 35S:MIR393 plants displayed no remarkable differences in size or numbers. Wild-type, 35S:MIR393a and 35S:MIR393b transgenic plants grown under short-day conditions were photographed with side view (left) and top view (right)
11103_2011_9838_MOESM3_ESM.tif
Supplementary material 3 (TIFF 2053 kb). Supplemental Figure 3. Relative expression levels of TIR1 in Col-0 and tir1-1 plants. TIR1 level in tir1-1 plants was not significantly different from the level in wild-type. Relative transcript levels were normalized using UBQ5 as a standard. The relative expression levels in wild-type plants were set to 1.0. Error bars represent SD from three PCR results, and similar results were obtained in three independent experiments
11103_2011_9838_MOESM4_ESM.tif
Supplementary material 4 (TIFF 3373 kb). Supplemental Figure 4. pTIR1:mTIR1-tir1-1 plants display pleiotropic effects similar to 35S:mTIR1. a 5-day-old pTIR1:mTIR1-tir1-1 transgenic plants displayed downward cotyledons and upward petioles. b 10-day-old pTIR1:mTIR1-tir1-1 plants displayed downward curly true leaves with long and twisted petioles. c–d The narrow and bended rosette leaves from 4-week-old pTIR1:mTIR1-tir1-1 were much smaller in size and fewer in numbers. Transgenic plants grown under short-day conditions were photographed with side view (c) and top view (d)
Rights and permissions
About this article
Cite this article
Chen, ZH., Bao, ML., Sun, YZ. et al. Regulation of auxin response by miR393-targeted transport inhibitor response protein 1 is involved in normal development in Arabidopsis . Plant Mol Biol 77, 619–629 (2011). https://doi.org/10.1007/s11103-011-9838-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-011-9838-1