Abstract
To date, both quality related high-molecular-weight (HMW) and low-molecular-weight (LMW) glutenin genes associated with dough extensibility and viscoelasticity traits were investigated separately in wheat. Therefore, the present study was designed to characterize at molecular level, nine spring wheat genotypes for desirable bread quality attributes, using gene/allele-specific DNA markers for both HMW and LMW glutenin and validating these results by conducting different bread quality analysis. The PCR results indicated that UAF-10,137 and Akbar-19 genotypes carried those HMW & LMW-Gs alleles that had previously been associated with good bread quality. These genotypes had Ax2*, Bx7 + By8 and Dx5 + Dy10 allelic combinations at Glu-1 loci, while gluA3b and gluB3b alleles were only present in UAF-10,137 at Glu-3 loci. However, Akbar-19 only had gluB3b allele at Glu-3 loci. Furthermore, the PCR investigation in UAF-10,123, Subhani-21, UAF-10,136 and Dilkash-20 genotypes confirmed the presence of some unknown alleles at both Glu-1 and Glu-3 loci thus indicating moderate bread making quality. In contrary to this, UAF-9515 and M.H-21 showed the presence of unknown alleles at Glu-A1, Glu-B1, Glu-A3, Glu-B3 loci and showed poor performance for bread quality parameters. Similar results were observed by using various bread quality-related tests such as farinograph, extensograph, sedimentation and bread volume. The results of these tests were in line with the findings of molecular investigations performed on the same wheat genotypes. In conclusion, genotypes UAF-10,137 and Akbar-19 were identified for having good bread making quality attributes and can be used as parents or as a good source of bread quality genes/alleles in future breeding programs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The rheological characteristics of the dough, which are used to further process it into a wide variety of food products, are determined by several seed storage proteins found in wheat. Based on their solubility in various solvents, the wheat storage proteins have been divided into four groups: albumins, globulins, glutenins, and gliadins (Osborne 1907). Of these, gluten, which makes up 80% of all the proteins in mature wheat grain, is formed when glutenin and gliadin interact. Essentially, the value addition of wheat into various products is carried out by the gluten protein. Together, glutenins and gliadins play a critical role in controlling the bread or dough quality of wheat flour. Contrarily, it is not believed that globulins and albumins have a significant role in determining the wheat flour’s ability to make dough or bread (Jones et al. 2006). Gliadin helps to increase the dough’s elasticity and strength, whereas glutenin is known to control the dough’s viscosity. As a result, the flour’s ability to make bread depends on the ideal ratio of both proteins (Khatkar and Schofield 1997; Wieser et al. 2006). HMW-GS (High-Molecular-weight Glutenin Subunits) and LMW-GS (Low-Molecular-weight Glutenin Subunits) are the two other broad types of glutenins. Molecular research has also shown that HMW glutenins are further classified into x-type (higher-molecular-weight) and y-type (low-molecular-weight) glutenins based on their isoelectric points and electrophoretic mobility. Genetic research has demonstrated that the Glu-1 locus is located on the homologous group-1 (A, B, and D) chromosomes. Glu-A1, B1, and D1 loci are responsible for encoding HMW-GS (Bietz et al. 1975). Additionally, each Glu-1 locus has genes that encode the x and y-types of glutenin protein, respectively. These genes are closely linked to one another (Harberd et al. 1986; Payne 1987; Shewry et al. 2003). But LMW-GS is encoded by the Glu-3 locus found on the Glu-A3, B3, and D3 loci of homologous group-1(A, B, and D) chromosomes (Gupta and Shepherd 1990).
The composition of HMW-GS has a considerable impact on the qualities associated with dough or bread making process (Payne 1987; Shewry et al. 1992). Similarly, the LMW-GS are crucial in determining the bread’s physical characteristics. In contrast to the LMW-GS, which accounts for 50% of the total gluten due to the expression of 30–40 genes in each individual genotype and contributes only 30% for gluten and dough-related parameters, the HMW-GS only accounts for 10% of the total gluten proteins due to the small number of protein subunits (3–5) present in each genotype. The roles of both HMW and LMW glutenin proteins have been revealed in earlier research. According to reports, the LMW-G alleles contribute significantly more to dough extensibility and resistance than the HMW-G alleles (Gupta et al. 1989; Cornish et al. 2001). Overall, HMW and LMW glutenin are connected to one another by disulfide bonds to produce macropolymers of gluten that provide qualities like extensibility and viscoelasticity to dough and eventually affect the dough’s capabilities for usage (Luo et al. 2001).
SDS-PAGE technology has been used in earlier studies to better understand the composition of HMW and LMW-GS and their function in bread quality (Bietz et al. 1975; Singh and Shepherd 1988). Based on their molecular weight, this approach classifies the proteins that affect the quality of the flour. The main issue with utilizing SDS-PAGE to characterize HMW and LMW protein subunits on a broad scale is that it is unable to identify allele-specific variations of HMW and LMW glutenin that are correlated with the rheological characteristics of the bread. Researchers have developed several DNA-based markers as an efficient substitute that can more quickly and accurately detect allele-specific variation for both LMW and HMW-GS (Ma et al. 2003; Andersen and Lübberstedt 2003; Zhang et al. 2004; Wang et al. 2010). These DNA markers are also well recognized to correlate with the rheological characteristics of bread. Furthermore, to find genotypes with good bread-making qualities, both HMW and LMW glutenin genes related with dough extensibility and viscoelasticity traits were investigated separately in earlier research work. Prior to the current work, however, the combined effect of the HMW & LMW glutenin alleles on bread or dough-making properties had not been explored.
The aim of the present study was to use a combination of both HMW and LMW glutenin allele-specific markers for identifying genotypes having good bread-making qualities. Furthermore, the testing of flour for different dough rheological and bread-making properties by using different quality tests to confirm the efficiency of these markers for selecting genotypes having good bread-making properties. This study also sought to determine the best glutenin allelic combination associated with desirable bread-related characteristics and to confirm the effectiveness of these molecular markers for genotype selection with favourable dough’s rheological and bread-making features.
Materials and methods
Planting material
Five indigenous elite lines of wheat (Table 1) were taken from the Department of Plant Breeding and Genetics, University of Agriculture, Faisalabad. Similarly, four Pakistani wheat cultivars (Dilkash-20, Subhani-21, Akbar-19 and M.H-21) were sourced from Wheat Research Institute, AARI, Faisalabad. The wheat genotypes were planted in Randomized Complete Block Design (RCBD) layout with three replications. Harvested seed was used for further quality related analysis.
DNA extraction and PCR amplification
The DNA was extracted by using CTAB method (Doyle and Doyle 1987). The PCR reactions were carried out to identify HMW & LMW-GS contributing to bread-making properties using twelve gene or allele-specific primers listed in Table 2.
Quality testing for dough rheological traits
The harvested seeds were cleaned, and flour was extracted at 14% moisture content from the collected samples of each genotype separately. The dough-rheological properties such as WA% (Water Absorption Percentage), DDT (Dough Development Time), DST (Dough Stability Time) and MTI (Mixing Tolerance Index) was determined by using Brabender Farinograph with a bowl capacity of 50 g according to AACC 54−21 method (AACC 2000). Extensograph was used to determine the RE (Resistance to Extension) and E (Extensibility) using AACC 54−10 method (AACC 2000). Moreover, Sodium dodecyl sulfonate (SDS)-Sedimentation volume was measured using 12% sodium dodecyl sulfate and 85% lactic acid in a standard cylinder tube by following AACC 56–61 method (AACC 2000). The bread volume (BV) was also measured by AACC 10−05 method (AACC 2000).
Statistical analysis
All tests were repeated in triplicate and the obtained data for each trait were subjected to Analysis of Variance (ANOVA) according to Steel et al. (1997). Moreover, the means comparison test was also applied to determine the differences among all the genotypes for the studied quality traits at the probability level of 5% (P < 0.05). The statistical analysis was performed using Statistica software v8.1.
Results
Glu-1 and Glu-3 loci gene/allele identification
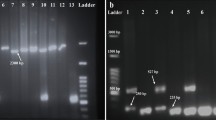
Twelve HMW and LMW glutenin genes/alleles were used in this work to describe the genotypes of Pakistani spring wheat. At Glu-1 loci, these alleles are known to encode the HMW glutenin subunits Ax1, Ax2, Ax2*, Bx7, Bx7*, Dx2, Dx5, and Dy12. However, in order to encode LMW glutenin subunits at Glu-3 loci, gluA3b, gluA3e, gluA3f, gluB3b, gluB3i, and gluB3c are required. PCR analysis showed that the PS1 primer pair did not amplify any PCR product for the Ax1 and Ax2 alleles in any wheat genotype and showed no allelic frequency (Tables 3 and 4). Similar to this, the PS2 primer pair amplified the PCR product for the Ax2* allele and confirmed its existence in the UAF-10,141, UAF-10,123, UAF-10,136, UAF-10,137, Subhani-21, and Akbar-19 genotypes by giving a single band of 1319 bp (Table 3; Fig. 1A). Contrary to this, the UAF-9515, Dilkash-20, and M.H-21 genotypes completely lacked the Ax2* allele. The Ax2* allele’s allelic frequency at the Glu-A1 locus was found to be 66.67% (Table 4).
Amplified PCR products obtained by using Nine gene/allele-specific markers of both HMW & LMW glutenin markers. A Ax2*, B Bx7 and Bx7*, C By8, D Dx5, E Dy10, F gluA3b, G gluB3b,
H gluB3c. 1–9 numbering indicates the name of different wheat genotypes i.e., UAF-9515, UAF-10,141, UAF-10,123, UAF-10,136, UAF-10,137, Subhani- 21, Dilkash-20, Akbar-19, M.H-21. M represents
the 100 bp gene ruler
The PS3 primer displayed a multi-banding pattern and amplified just one band (766 bp) for the Bx7* allele while displaying two bands of 670 and 770 bp for the Bx7 allele. Only the genotypes UAF-10,137, Subhani-21, Akbar-19, and M.H.-21 were discovered to carry the Bx7 allele. Moreover, Bx7* allele was found in three genotypes: UAF-10,141, UAF-10,136, and Dilkash-20 (Table 3; Fig. 1B). Additionally, (Table 4) showed that Bx7 and Bx7* had allelic frequencies of 44.45 and 33.33%, respectively, but Table 3 showed that PS4 primer failed to amplify the necessary band size. The PS5 primer set amplified a band with a size of 527 bp, confirming the existence of the By8 gene in all genotypes except for UAF-10,136, Subhani-21, and M.H-21, as shown in Table 3; Fig. 1C. Additionally, the Glu-B1 locus displayed an allelic frequency of 66.67%, as indicated in Table 4.
At Glu-D1 locus, Dx5 and Dy10 genes were detected in all genotypes as shown in Table 3; Fig. 1D and E. Bands of 450 and 576 bp were amplified by PS6 and PS7 primers. The Dy12 allele cannot be amplified by the PS7 primer, according to the PCR data. Due to their presence in all genotypes, the Dx5 and Dy10 genes displayed 100% allelic frequency at the Glu-D1 locus (Table 4).
While other genotypes did not demonstrate the presence of the gluA3b allele at the Glu-A3 locus, the PS8 primer pair amplified the PCR product of 894 bp for the gluA3b allele and validated its presence in UAF-9515, UAF-10,123, UAF-10,136, UAF-10,137, and Subhani-21 (Table 3; Fig. 1F). Only five genotypes had the gluA3b, and its allele frequency was 55.56%. (Table 4). But the gluA3e allele was not amplified by the PS9 primer, which confirmed its absence in all genotypes and showed no allelic frequency (Tables 3 and 4). Moreover, the PS10 primer pair did not amplify any PCR product for gluA3f allele and showed no allelic frequency (Tables 3 and 4).
Except for UAF-9515 and Subhani-21, the PS11 primer pair produced a PCR product of 1570 bp for the gluB3b allele and validated its presence in all genotypes examined, as shown in (Table 3) and (Fig. 1G). Additionally, it had a higher allelic frequency of 77.78% at the Glu-B3 locus since its frequency was at its highest in the genotypes under study, as seen in (Table 4). Table 3 shows that the PS12 primer pair failed to amplify the PCR result for the gluB3i allele, and that the allele frequency was 0%. (Table 4). While the remaining genotypes did not demonstrate the existence of the gluB3c allele, the PS13 primer produced a PCR product of 472 bp size for the gluB3c allele at the Glu-B3 locus in genotype UAF-9515 (Table 3). (Fig. 1H). Additionally, it had a low allelic frequency of 11.11% because of its rarity in the investigated germplasm (Table 4).
Variability in quality traits
To determine the variability for all the examined features, the results from the Farinograph, Extensograph, SDS-Sedimentation, and bread volume tests were subjected to analysis of variance (Supplementary Table 1). Water absorption, dough development and stability times, mixing tolerance index, resistance to extension, extensibility, SDS-sedimentation, and bread volume attributes all had significant variation (P < 0.05), according to the analysis of variance.
Farinograph studies
A farinograph is frequently used in the food industries to analyse the rheological characteristics of dough. It offers details on a variety of dough characteristics, including the quantity of water needed, dough development and stability times, and dough mixing tolerance. These characteristics have a key role in predicting the ultimate dough quality. The findings revealed that the UAF-10,137 genotype had the highest percentage of water absorption (61.3), whereas the UAF-9515 genotype had the lowest proportion (51%). Except for UAF-9515, all genotypes showed non-significant differences for the water absorption trait according to the Tukey HSD test. Significant changes between UAF-10,137, UAF-10,123, and UAF-9515 were found for DDT. The fastest dough formation times were 10.35 and 7.35 min for the genotypes UAF-10,137 and UAF-10,123, respectively. However, UAF-9515 had the shortest dough development time (3.35 min). (Table 3). DST was shown to significantly differ solely between the UAF-10,137 and UAF-9515 genotypes. UAF-10,137 had the longest dough stability time (18.8 min), whereas UAF-9515 had the shortest (5.85 min.). Additionally, there were notable differences between UAF-10,137 and M.H-21 in the mixing tolerance of the dough. According to Table 3, the genotype M.H-21 had the highest MTI value (51 min), whereas the genotype UAF-10,137 had the lowest MTI value (11.75 min.).
Extensensograph studies
Using this test, you may find out how strong the gluten is, how well the wheat makes bread, how long the fermentation process lasts, and how other ingredients affect the qualities of the dough. Using this method, we can calculate the dough’s RE (resistance to extension) and E (extensibility) values. The means comparison test for RE and E revealed that there were differences between genotypes that were statistically significant. Subhani-21 shown the least amount of resistance to extension, whereas genotype UAF-10,137 displayed the most (422 B.U. (336.5 B.U.). UAF-10,137 also displayed maximum extensibility values in terms of extensibility (277 mm). However, as shown in Table 3, genotype M.H-21 indicated minimum extensibility value (221 B.U).
Bread volume and SDS-sedimentation
The strength and quality of the gluten are also evaluated using the sedimentation test. It is a crude method that is used to have an understanding about baking quality related properties of the flour. Additionally, the main aspect of flour quality that determines whether a genotype is suitable for bread manufacturing or not is bread volume. Results showed that for sedimentation and bread volume, all the genotypes varied significantly. It was noted that UAF-10,137 had the greatest results for both bread volume (316 cm3) and sedimentation (55 ml), respectively. UAF-9515, on the other hand, had the lowest levels of sedimentation and bread volume, which were 29 ml and 249 cm3, respectively.
Discussion
Many studies were conducted to determine the composition of LMW & HMW glutenin subunits in bread wheat (Jin et al. 2011; Atanasova et al. 2009; Henkrar et al. 2017a, b). Their appropriate composition controls the quantity and quality of the gluten protein and is crucial in determining the rheological properties of the dough. The characterization of LMW & HMW glutenins subunits in bread wheat is crucial for developing genotypes with desired bread quality. SDS-PAGE was used in earlier investigations to analyze the composition of HMW and LMW-GS. However, because of its complexity and the similarity in electrophoretic mobility between gliadins and LMW-GS, this approach is not appropriate for high-throughput study (Liu et al. 2008; Gupta et al. 1994; Masci et al. 1998; Maruyama-Funatsuki et al. 2004). For the Glu-1 and Glu-3 loci, numerous thorough genomic studies have been described in recent studies. In recent studies, a lot of comprehensive genomic studies have been reported for characterizing the Glu-1 and Glu-3 loci. To study these, gene/allele-specific DNA markers were developed for HMW & LMW-GS for improving bread quality of wheat (Ahmad 2000; Radovanovic et al. 2002; Radovanovic and Cloutier 2003; Ma et al. 2003; Lei et al. 2006; Wang et al. 2010). Furthermore, Kuchel et al. (2007) also reported the use of MAS (Marker-Assisted Selection) approach to improve the bread or dough-related properties of wheat.
The PCR results indicated that all the genotypes except UAF-9515 and M.H-2 carried Ax2* allele and demonstrated an allelic frequency of 66.67% at Glu-A1 locus. Moreover, the absence of Ax1, Ax2 and Ax2* alleles in UAF-9515 and M.H-21 may indicate the presence of AxNull or any other type of allele at Glu-A1 locus. There is need to investigate that which alleles is present in UAF-9515 and M.H-21 genotypes. Similar findings were also reported by Nucia et al. (2019), who analysed 79 genotypes of spring wheat from all around the Europe and found that Ax2* had highest frequency (71%) at Glu-A1 locus. Jin et al. (2011), who evaluated 719 wheat genotypes from 20 countries to assess the composition of HMW-GS, complement the findings of the current study by demonstrating that Ax2* had a 43.3% allelic frequency among the studied genotypes. These studies also confirmed that the presence of Ax2* allele had strong influence on dough and bread related properties. Moreover, our PCR findings for Glu-B1 locus indicated the high variation and confirmed the presence of Bx7 and Bx7* alleles in the studied genotypes except from UAF-9515, Dilkash-20 and M.H-21. These genotypes might contain unidentified or other alleles like Bx17, Bx7OE, and Bx6. The presence of Bx7 and Bx7* showed an allelic frequency of 44.45 and 33.33%, respectively. Our findings conflict with prior research by Nucia et al. (2019), and Jin et al. (2011). They discovered that in the European wheat germplasm, Bx7* is the allele that is more prevalent than Bx7. Similarly, Esp’ et al. (2013Henkrar et al. (2017a, b) also noted a higher frequency of allele Bx7*. However, our PCR results for “y” type glutenin subunits indicated that By18 allele was absent and showed 0% allelic frequency, but By8 allele showed 66.67% allelic frequency due to its presence in all genotypes apart from UAF-10,136, Subhani-21, and M.H-21 at Glu-B1, as shown in Tables 3 and 4. These genotypes may carry the By9, Bynull, and By8* alleles. Therefore, it is necessary to identify the unknown alleles utilizing a variety of techniques, such as peptide mass sequencing and nucleotide sequencing. In contrast to the current work, Janni et al. (2017) identified 19 genotypes of bread and durum wheat and noted that the By8 allele had just 16% allelic frequency. Jin et al. (2011) also found that By8 had a higher allelic frequency (31.1%) than By9, which had only a 22.28% allelic frequency. In the present study, By8 also showed higher frequency but its absence in some genotypes concluded that higher variability is present at Glu-B1 locus in bread wheat.
At Glu-D1, a single allelic combination Dx5 + Dy10 was observed in all the genotypes and showed 100% allele frequency. This indicated that both Dx5 and Dy10 alleles are more common in Pakistani spring wheat genotypes. A similar study was conducted by Ali et al. (2013), who also confirmed the high allelic frequency (95%) of Dx5 + Dy10 combination in Pakistani spring wheat genotypes. MoreoverDias et al. (2017); Henkrar et al. (2017a, b) also confirmed the high allelic frequency of Dx5 + Dy10 in their studies that was 73% and 85%, respectively. Nucia et al. (2019) also observed that more than 80% European’s wheat genotypes had Dx5 + Dy10 allele at Glu-D1 locus. This allele showed a strong association with good bread-making properties as compared to Dx2 + Dy12 allelic combination (Costa et al. 2013; Barakat et al. 2018).
At Glu-A3 locus, only a few genotypes showed the presence of gluA3b allele and an allelic frequency of 55.56%. This lower frequency % age may point out the presence of other alleles such as the a and g allele at the Glu-A3 locus. Similar results were reported by Zhang et al. (2004), who discovered six distinct allelic variants of a single gene encoding LMW glutenin and developed six allele-specific markers to differentiate each allele from the others. Similarly, the presence of gluB3b, gluB3c, and other unknown alleles at the Glu-B3 locus also showed the existence of several other LMW glutenin alleles. Additionally, the allelic frequencies for the gluB3b and gluB3c alleles were 77.78 and 11%, respectively as shown in Tables 3 and 4. The observed differences in allelic frequency also evident the presence of several LMW glutenin alleles at the Glu-B3 locus. Different alleles of the Glu-B3 locus, including alleles a, b, c, d, e, f, g, h, and i were also found by Gupta and Shepherd (1990). Additionally, Costa et al. (2013) discovered many alleles and revealed that the spring wheat genotypes under study had the highest frequency of gluB3b (33.33%). Our findings support earlier research, which reported the significant level of polymorphism at the Glu-B3 locus.
Molecular findings were further validated with various bread quality-related tests such as farinograph, extensograph, sedimentation, and bread volume. These analytical tests proved helpful in assessing the effectiveness of Marker-Assisted Selection (MAS) technique that was utilized in this study for genotyping and selection of wheat genotypes that contributed to good bread-making properties. The farinographic analysis indicated that, genotype UAF-10,137 absorbed more water than UAF-9515 by a wide margin. Similar findings were reported by Simon (1987), who concluded that greater water absorption is a sign of high-quality flour and is necessary for baking bread of excellent quality. Other genotypes, on the other hand, displayed non-significant differences for WA%; this could be because various alleles have a similar effect or because other flour quality parameters play an important role. Future research on this figure will require a thorough comprehension.
Additionally, the dough development time data demonstrated that all genotypes, except for Subhani-21, UAF-9515, and M.H-21 genotypes, showed largest and non-significant differences for DDT. These three genotypes might exhibit less DDT because of the different HMW and LMW glutenin subunits. Stronger flour is typically indicated by a longer dough development time, whereas a lower value denotes weaker flour. Safdar et al. (2009) also reported that genotypes with higher DDT indicated good quality flour as compared to genotypes with lower DDT. In addition to this, UAF-10,137 genotype indicated highest DST (Dough Stability Time) and lowest MTI (Mixing Tolerance Index) or dough softening whereas UAF-9515 had lowest value for DST and highest MTI value. The higher DST and lower MTI values also indicate strong flour quality, and these rheological properties are required to make good bread (Anjum and Walker 2000). Moreover, all the genotypes showed a minor difference for both DST and MTI values in the present study, it may be due to the minor differences in the protein quality and quantity in the studied genotypes. Similar findings were also reported in many studies (Rehman et al. 2001).
The Extensograph results indicated that UAF-10,137 had highest RE (Resistance to Extension) and E (Extensibility). However, Subhani-21 and M.H-21 had lower RE and E, respectively. Our results suggested that genotypes had varying performance for both RE and E parameters of Extensograph. It implies that these two factors alone are insufficient to ascertain the rheological or viscoelastic characteristics of the dough. Moreover, the differences in the extensographic properties may be exhibited due to the presence of different allelic combinations at both Glu-1 and Glu-3 loci. Torbica et al. (2007) concluded that the differences in RE and E may occur due to imperfect balance between gliadin and glutenin content. The imperfect balance between these two proteins leads to increased extensibility whereas lower resistance to extension. Torbica et al. (2011) also observed that RE and E parameters of Extensograph are not enough to determine the visco-elastics properties of the bread. Now there is a need to study other parameters of extensgraph while selecting genotypes for good visco-elastics properties.
The SDS-Sedimentation volume is widely used to measure the gluten quality and its strength. Additionally, it also gives an information about bread-making properties. The findings of the sedimentation test revealed that UAF-10,137 had the maximum volume of sedimentation while UAF-9515 had the lowest amount. A significant positive correlation between loaf volume and sedimentation volume was found by Guzmán et al. (2022). Moreover, they also found that Glu-D1 locus had less 1% contribution to this character. However, at Glu-3 locus, gluA3b and gluB3b alleles were also showing a strong association with sedimentation volume. Our results are also in agreement with Guzmán et al. (2022). Our findings showed a substantial difference between the genotypes for bread volume. The genotype UAF-10,137 displayed the highest value for bread volume, whereas the genotype UAF-9515 displayed the lowest value. These significant variations could be the result of distinct HMW and LMW glutenin subunits. Similar results were reported by Guzmán et al. (2022). They concluded that environmental conditions and protein content, in addition to the makeup of different alleles, also affect bread volume. They also confirmed that gluA3b and gluB3b alleles had higher contributions to bread volume than Glu-D1, which has a minimal effect on bread volume. These findings are consistent with the current investigation.
Conclusion
Marker-Assisted Selection (MAS) and various quality-related test indicated that Ax2*, Bx7, By8, Dx5, Dy10, gluA3b, and gluB3b genes/alleles can be used for early selection of bread wheat genotypes with good dough or bread quality. Wheat genotypes (UAF-10,137 and Akbar-19) that are selected in this study, can be utilized as parents or as a source of high-quality contributing genes/alleles in breeding efforts to evolve genotypes that produce large quantity of grains while maintaining higher bread quality standards.
Future prospect
Several unknown alleles at the Glu-1 and Glu-3 loci that were discovered in the current study still need to be identified. Future mutant studies will also be required for a thorough investigation of the function of each allele at the Glu-1 and Glu-3 loci. These investigations will aid in clarifying the precise nature of the contribution of each allele to the final qualities of dough or bread. Additionally, it is important to research how each allele expresses itself in various environmental contexts. This will give specific information about how the environment may affect the quality of the bread. Additionally, it is essential to develop new testing methods that may aid breeders in validating the outcomes of marker-assisted selection (MAS).
Change history
01 July 2023
A Correction to this paper has been published: https://doi.org/10.1007/s10722-023-01651-3
References
Ahmad M (2000) Molecular marker-assisted selection of HMW glutenin alleles related to wheat bread quality by PCR-generated DNA markers. Theor Appl Genet 101:892–896
Ali N, Iqbal M, Yasmin S, Ahmed I, Asif M (2013) Genetic variation in loci controlling quality traits in spring wheat. Pakistan J Agri Sci 50:637–647
American Association of Cereal Chemists (AACC) (2000) Approved Methods of the AACC. 10th edition. American Assoc Cereal Chem, St. Paul, MN
Andersen JR, Lübberstedt T (2003) Functional markers in plants. Trends in Plant Sci 8:554–560
Anjum FM, Walker CE (2000) Grain, flour and bread-making properties of eight pakistani hard white spring wheat cultivars grown at three different locations for 2 years. Int J Food Sci Technol 35:407–416
Atanasova D, Tsenov N, Todorov I, Ivanova I (2009) Glutenin composition of winter wheat varieties bred in dobrudzha agricultural institute. Bulgarian J Agri Sci 15:9–19
Barakat MN, Al-Doss AA, Ghazy AI, Moustafa KA, Elshafei AA, Ahmed EI (2018) Doubled haploid wheat lines with high molecular weight glutenin alleles derived from microspore cultures. New Z J Crop Hortic Sci 46:198–211
Bietz JA, Shephard KW, Wall JS (1975) Single-kernel analysis of glutenin: use in wheat genetics and breeding. Cereal Chem 52:513–532
Cornish GB, Bekes F, Allen HM, Martin DJ (2001) Flour proteins linked to quality traits in an australian doubled haploid wheat population. Aust J Agric Res 52(12):1339–1348
Costa MS, dos Scholz S, Franco MB CML (2013) Effect of high and low molecular weight glutenin subunits and subunits of gliadin on physicochemical parameters of different wheat genotypes. Food Sci Technol 33:163–170
Dias RO, Souza MA, Pirozi MR, Oliveira LC, Pimantel AJB (2017) Protein profile and molecular markers related to the baking quality of brazilian wheat cultivars. Cereal Chem 94:568–575
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Feldman M (2001) Origin of cultivated wheat. In: Bonjean AP, Angus WJ (eds) The World Wheat Book. A history of wheat breeding. Lavoisier Publishing, Paris, pp 3–53
Gupta RB, Shepherd KW (1990) Two-step one-dimensional SDS-PAGE analysis of LMW subunits of glutenin. I. Variation and genetic control of the subunits in hexaploid wheats. Theor Appl Genet 80:65–74
Gupta RB, Singh NK, Shepherd KW (1989) The cumulative effect of allelic variation in LMW and HMW glutenin subunits on dough properties in the progeny of two bread wheats. Theor Appl Genet 77(1):57–64
Gupta RB, Paul JG, Cornish GB, Palmer GA, Bekes F, Rathjen AJ (1994) Allelic variation at glutenin subunit and gliadin loci, Glu-1, Glu-3 and Gli-1 of common wheats. Its additive and interaction effects on dough properties. J Cereal Sci 19:917
Guzmán C, Crossa J, Mondal S, Govindan V, Huerta J, Crespo-Herrera L, Ibba MI (2022) Effects of glutenins (Glu-1 and Glu-3) allelic variation on dough properties and bread-making quality of CIMMYT bread wheat breeding lines. Field Crops Res 284:108585
Harberd NP, Bartels D, Thompson RD (1986) DNA restriction fragment variation in the gene family encoding high-molecular-weight (HMW) glutenin subunits of wheat. Biochem Genet 24:579–592
Henkrar F, El-Haddoury J, Iraqi D, Bendaou N, Udupa SM (2017a) Allelic variation at high molecular weight and low-molecular weight glutenin subunit genes in moroccan bread wheat and durum wheat cultivars. 3 Biotech 7:287
Henkrar F, El-Haddoury J, Iraqi D, Bendaou N, Udupa SM (2017b) Allelic variation at high-molecular weight and low-molecular weight glutenin subunit genes in moroccan bread wheat and durum wheat cultivars. 3 Biotech 7:1–10
Janni M, Cadonici S, Pignone D, Marmiroli N (2017) Survey and new insights in the application of PCR-based molecular markers for identification of HMW-GS at the Glu-B1 locus in durum and bread wheat. Plant Breed 136:467–473
Jin H, Yan J, Pena RJ, Xia XC, Morgounov A, Han LM, Zhang Y, He ZH (2011) Molecular detection of high- and low molecular-weight glutenin subunit genes in common wheat cultivars from 20 countries using allele-specific markers. Crop Pasture Sci 62:746–754
Jones BL, Morris CF, Bekes F, Wrigley CW (2006) Proteins that complement the roles of gliadin and glutenin. In: Wrigley CW, Bekes F, Bushuk W (eds) Gliadin and glutenin. The Unique Balance of Wheat Quality. AACC International, St. Paul, MN, pp 413–446
Khatkar BS, Schofield JD (1997) Molecular and physico-chemical basis of breadmaking properties of wheat gluten proteins: a critical appraisal. J Food Sci Technol 34:85–102
Kuchel H, Fox R, Reinheimer J, Mosionek L, Willey N, Bariana H, Jefferies S (2007) The successful application of a marker-assisted wheat breeding strategy. Mol Breed 20:295–308
Lei ZS, Gale KR, He ZH, Gianibelli C, Larroque O, Xia XC, Butow BJ, Ma W (2006) Y-type gene specific markers for enhanced discrimination of high-molecular weight glutenin alleles at the Glu-B1 locus in hexaploid wheat. J Cereal Sci 43:94–101
Liang X, Zhen S, Han C, Wang C, Li X, Ma W, Yan Y (2015) Molecular characterization and marker development for hexaploid wheat-specific HMW glutenin subunit 1By18 gene. Mol Breed 35:1–16
Liu S, Chao S, Anderson JA (2008) New DNA markers for high molecular weight glutenin subunits in wheat. Theor Appl Genet 118:177–183
Luo C, Griffin WB, Branlard G, McNeil DL (2001) Comparison of low- and high- molecular-weight wheat glutenin allele effects on flour quality. Theor Appl Genet 102:1088–1098
Ma W, Zhang W, Gale KR (2003) Multiplex-PCR typing of high molecular weight glutenin alleles in wheat. Euphytica 134:51–60
Maruyama-Funatsuki W, Takata K, Nishio Z, Tabiki T, Yahata E, Kato A, Saito K, Funatsuki H, Saruyama H, Yamauchi H (2004) Identification of low-molecular weight glutenin subunits of wheat associated with bread-making quality. Plant Breed123:355–360
Masci S, D’Ovidio R, Lafiandra D, Kasarda DD (1998) Characterization of a low-molecular-weight glutenin subunit gene from bread wheat and the corresponding protein that represents a major subunit of the glutenin polymer. Plant Physiol 118:1147–1158
Nucia A, Okoń S, Tomczyńska-Mleko M (2019) Characterization of HMW glutenin subunits in european spring common wheat (Triticum aestivum L). Genet Resour Crop Evol 66:579–588
Osborne TB (1907) The proteins of the Wheat Kernel. Carnegie Institution of Washington, Washington, DC, USA, Publ., p 84
Payne PI (1987) Genetics of wheat storage proteins and the effect of allelic variation on bread-making quality. Annu Rev Plant Physiol 38:141–153
Radovanovic N, Cloutier S (2003) Gene-assisted selection for high molecular weight glutenin subunits in wheat doubled haploid breeding programs 12:51–60
Radovanovic N, Cloutier S, Brown D, Humphreys DG, Lukow OM (2002) Genetic variance for gluten strength contributed by high molecular weight glutenin proteins. Cereal Chem 79:843–849
Rehman S, Nazir Y, Hussain S, Huma N (2001) Study on the evaluation of wheat varieties of Sindh Province for the production of ring doughnuts. J Anim Plant Sci 3:135–138
Safdar MN, Naseem K, Siddiqui N, Amjad M, Hameed T, Khalil S (2009) Quality evaluation of different wheat varieties for the production of Unleavened flat Bread (Chapatti). Pakistan J Nutr 8:1773–1778
Shewry PR, Halford NG, Tatham AS (1992) High molecular weight subunits of wheat glutenin. J Cereal Sci 15:105–120
Shewry PR, Halford NG, Lafiandra D (2003) Genetics of wheat gluten proteins. Adv Genet 49:111–184
Simon SJ (1987) More wheat with superior baking quality is reached. Cereals Foods World 32:323–332
Singh NK, Shepherd KW (1988) Linkage mapping of genes controlling endosperm storage proteins in wheat. Theor Appl Genet 75:628–641
Steel RGD, Torrie JH, Dickey DA (1997) Principles and procedures of statistics: a biometrical approach, 3rd edn. Mc Graw-Hill, New York
Torbica A, Antov M, Mastilović J, Knežević D (2007) The influence of changes in Gluten Complex structure on Technological Quality of Wheat (Triticum aestivum L). Food Res Int 40(8):1038–1045
Torbica A, Živančev D, Mastilović J, Knežević D, Bodroža-Solarov M (2011) Impact of Changes in Climate Conditions on the Technological Quality of Wheat, Proceedings of 46th Croatian & 6th International Symposium on Agriculture, pp. 617–621, ISBN 978-953-6135-71-4, Opatija, Hrvatska, February 14–18, 2011
Wang LH, Zhao XL, He ZH, Ma W, Appels R, Peña RJ, Xia XC (2009) Characterization of low-molecular-weight glutenin subunit Glu-B3 genes and development of STS markers in common wheat (Triticum aestivum L). Theor Appl Genet 118:525–539
Wang L, Li G, Peña RJ, Xia X, He Z (2010) Development of STS markers and establishment of multiplex PCR for Glu-A3 alleles in common wheat (Triticum aestivum L). J Cereal Sci 51:305–312
Wieser H, Bushuk W, MacRitchie F (2006) The polymeric glutenins. In: Wrigley CW, Bekes F, Bushuk W (eds) Gliadin and glutenin. The Unique Balance of Wheat Quality. AACC International, St. Paul, MN, pp 213–240
Zhang W, Gianibelli MC, Rampling L, Gale KR (2004) Characterisation and marker development for low molecular weight glutenin genes from Glu-A3 alleles of bread wheat (Triticum aestivum L). Theor Appl Genet 108:1409–1419
Funding
This post graduate research work was carried out under the joint collaboration of Agricultural Biotechnology Research Institute (ABRI) and Department of Plant Breeding and Genetics (PBG), University of Agriculture (UAF), Faisalabad, Pakistan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There is no potential conflict of interest among authors of this manuscript.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bashir, M.A., Kashif, M., Habib, I. et al. Molecular characterization of bread quality contributing high and low-molecular-weight glutenin subunits in Pakistani spring wheat genotypes. Genet Resour Crop Evol 71, 285–295 (2024). https://doi.org/10.1007/s10722-023-01622-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-023-01622-8