Abstract
PCR was used to amplify low-molecular-weight (LMW) glutenin genes from the Glu-A3 loci of hexaploid wheat cultivars containing different Glu-A3 alleles. The complete coding sequence of one LMW glutenin gene was obtained for each of the seven alleles Glu-A3a to Glu-A3g. Chromosome assignment of PCR products using Chinese Spring nulli-tetrasomic lines confirmed the amplified products were from chromosome 1A. All sequences were classified as LMW-i-type genes based on the presence of an N-terminal isoleucine residue and eight cysteine residues located within the C-terminal domain of the predicted, mature amino acid sequence. All genes contained a single uninterrupted open reading frame, including the sequence from the Glu-A3e allele, for which no protein product has been identified. Comparison of LMW glutenin gene sequences obtained from different alleles showed a wide range of sequence identity between the genes, with between 1 and 37 single nucleotide polymorphisms and between one and five insertion/deletion events between genes from different alleles. Allele-specific PCR markers were designed based on the DNA polymorphisms identified between the LMW glutenin genes, and these markers were validated against a panel of cultivars containing different Glu-A3 alleles. This collection of markers represents a valuable resource for use in marker-assisted breeding to select for specific alleles of this important quality-determining locus in bread wheat.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Low-molecular-weight (LMW) glutenins represent approximately 40% of the total wheat gluten fraction, and their molecular weight is estimated to range from 30 kDa to 50 kDa (Ciaffi et al. 1999). The LMW glutenins have been found to have a pronounced effect in determining the physical properties of flour during bread-making (Payne et al. 1987; Gupta et al. 1989; Gupta and McRitchie 1994) and pasta-making (Pogna et al. 1990; Ruiz and Carrillo 1995). LMW glutenins are encoded at the Glu-A3, Glu-B3 and Glu-D3 loci on the short arms of group 1 homeologous chromosomes (Gupta and Shepherd 1990). The investigation of the influence of allelic variation of LMW glutenins on flour quality has received less attention than the effects of variation at the Glu-A1, Glu-B1 and Glu-D1 high-molecular-weight (HMW) glutenin loci located on the long arms of group 1 chromosomes (Payne et al. 1987). This may be attributed largely to the difficulty in scoring alleles of this complex multi-gene family by protein analysis (see below).
The effects of LMW-glutenin allelic composition on dough strength and extensibility have been demonstrated for both bread and durum wheat, although these effects are largely dependent on the association of Glu-3 and Glu-1 loci (Payne et al. 1987; Gupta et al. 1989, 1994; Nieto-Taladriz et al. 1994; Ruiz and Carrillo 1995; Sontag-Strohm et al. 1996; Redaelli et al. 1997). Characterisation of specific alleles at the Glu-3 loci has been conducted with respect to their effect on dough resistance and extensibility recently (Wesley et al. 1999; Brites and Carrillo 2001; Luo et al. 2001; Wesley et al. 2001). Several Glu-3 alleles have been ranked in terms of their dough quality effects (Gupta and Shepherd 1988; Gupta et al. 1989, 1990a, 1990b, 1991, 1994; Metakovsky et al. 1990). For R max (maximum dough resistance, an indicator of dough strength), the ranking of alleles at Glu-A3 was b>d>e>c; the alleles of Glu-B3, i>b=a>e=f=g=h>c; the alleles of Glu-D3, e>b>a>c>d. Cornish et al. (1993) found the Glu-3 composition bbb (at loci Glu-A3, Glu-B3 and Glu-D3, respectively) gave the best extensibility. Glu-3 composition bbc also had excellent extensibility. Glu-A3e, Glu-B3c, d and g alleles were associated with medium to weak dough properties.
Sodium dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE) is currently the most widely used method for determining LMW glutenin allelic composition of wheat cultivars and lines (Jackson et al. 1996). However, the difficulty of resolving the large number of LMW glutenin subunits (LMW-GS) expressed in hexaploid wheat has hindered the selection of specific LMW glutenin alleles in wheat breeding programs. Other fractionation techniques, with greater discriminating power, such as 2-D electrophoresis (Jackson et al. 1983; Skylas et al. 2000) and RP-HPLC (Burnouf and Bietz 1984; Marchylo et al. 1989; Sutton 1991; Margiotta et al. 1993) are available for LMW glutenin analysis. However, these are not routinely used in wheat breeding programs due to the complexity and the techniques and the problem of overlapping fractions (Weegels et al. 1996).
Gupta and Shepherd (1990) conducted an extensive survey of LMW glutenin proteins in hexaploid wheat cultivars and detected 20 different banding patterns: 6 controlled by Glu-A3, 9 by Glu-B3 and 5 by Glu-D3. The complexity of the banding patterns varied widely, with some cultivars showing no LMW glutenin protein detected from the Glu-A3 locus (Glu-A3e), and some Glu-B3 banding patterns consisting of at least eight distinct bands (Glu-B3 h and i). Thirty-nine different LMW subunits were identified in one bread wheat cultivar based on N-terminal amino acid sequences (Lew et al. 1992). Cassidy et al. (1998) estimated that the copy number of LMW glutenin-like DNA sequences is 30–40 in another bread wheat cultivar based on Southern blotting analysis. Approximately 60 genes or pseudogenes of the LMW-GS family have been cloned and sequenced from seven hexaploid wheat cultivars at this time (Bartels and Thompson 1983; Pitts et al. 1988; Colot et al. 1989; Cassidy et al. 1998; Benmoussa et al. 2000; Cloutier et al. 2001; Ikeda et al. 2002). These sequences were classified into 12 groups based on the alignment of the N- and C-terminal conserved domains of the deduced amino-acid sequences (Ikeda et al. 2002). Based on the gene sequence polymorphisms between LMW glutenin genes, an LMW glutenin-specific PCR marker (D’Ovidio 1993) was developed to distinguish durum wheat cultivars according to their quality characteristics and a microsatellite marker has been developed for use in mapping of hexaploid wheat (Devos et al. 1995). The Glu-3 LMW glutenin loci are tightly linked to the Gli-1 loci that encode γ and ω gliadins (Brown and Flavell 1981). Linkage of a γ-gliadin allele with gluten strength in durum wheat, possibly due to the linkage of the Gli-1 and Glu-3 loci, has been reported (Payne et al. 1984). The gene sequences in the complex γ-gliadin gene family of bread wheat have been used to identify single nucleotide polymorphisms (SNPs) and insertion/deletions (InDels) specific for individual genes (Zhang et al. 2003). Allele-specific PCR markers based on SNPs of γ-gliadin genes were subsequently used to detect specific alleles linked to the Glu-A3, Glu-B3 and Glu-D3 LMW glutenin loci, respectively (Zhang et al. 2003). InDels have also been reported in the LMW glutenin gene family recently (Benmoussa et al. 2000; Ikeda et al. 2002). Here, we describe the isolation and characterisation of LMW glutenin genes from different Glu-A3 alleles of bread wheat and the subsequent development of PCR markers to facilitate marker assisted selection of specific alleles of this important quality-determining locus.
Materials and methods
Plant materials
Wheat germplasm included in this study was maintained at CSIRO Plant Industry or obtained from the Australian Winter Cereals Collection in Tamworth, Australia. Chinese Spring euploid and its nulli-tetrasomic lines were kindly provided by Dr. E. Lagudah (CSIRO Plant Industry, Canberra) and derived from stocks described in Sears and Miller (1985).
Allelic composition at the Glu-A3 loci by SDS-PAGE
The Glu-A3 allelic compositions of all wheat cultivars used in this study were confirmed by SDS-PAGE as described by Zhang et al. (2003). The cultivars used as DNA template for amplification and sequencing of LMW-GS genes were Chinese Spring (Glu-A3a), Gabo (Glu-A3b), Cheyenne (Glu-A3c), Suneca (Glu-A3d), Halberd (Glu-A3e), Rescue (Glu-A3f) and Glenlea (Glu-A3g).
DNA isolation and PCR amplification
DNA was isolated from pools of five 3- to 5-day-old seedlings with a modified CTAB procedure (Gale et al. 2001). PCR was performed using Hotstar Taq polymerase (1 U, Qiagen) in 10 μL of reaction buffer (Qiagen, containing 1.5 mM MgCl2) containing 50 ng genomic DNA, 100 μM of each dNTP and 5 pmol of each PCR primer. PCR was 95°C for 3 min, followed by 40 cycles of 95°C for 30 s, 59°C for 30 s and 72°C for 1 min.
Chromosome assignment of specific PCR markers
Assignment of specific PCR markers to specific chromosomes was accomplished through PCR amplification using template DNA from each of the three Chinese Spring, group 1 nulli-tetrasomic lines: N1AT1B, N1BT1A and N1DTA.
Genome walking
The method described by Siebert et al. (1995) was applied with some modification. Genomic DNA of wheat cv. Halberd (Glu-A3e allele) was digested with restriction endonucleases DraI, EcoRV, PvuII or StuI (New England Biolabs). The digested genomic DNA was ligated with adaptor (Universal GenomeWalker Kit, Clontech PT3042–2) and nested PCR performed using the supplied adaptor-specific primers (AP1 and AP2) in combination with the Glu-A3e gene-specific primers GluA3eF1 (5′-ACTCGGGCAACAACCTCAACAACAACAG-3′) and GluA3eF2 (5′-CCTTTTTGCAGCCACACCAGATAGCCCAA-3′), respectively. PCR amplifications were performed as described in the manufacturer’s protocol using Hotstar Taq polymerase (Qiagen) and with the addition of incubation at 94°C for 3 min before the cycling. The PCR products were directly sequenced as described below.
Sequencing of amplification products
Individual amplification products (2 μl) were treated with 0.25 U shrimp alkaline phosphatase (USB) and 2.5 U exonuclease I (USB) at 37°C for 30 min and followed by 80°C for 15 min in PCR reaction buffer (Qiagen, 1.5 mM MgCl2). The PCR products were then sequenced using the appropriate PCR primers (the same primers as used for PCR and a primer GluA3R3 (5′-GGATTGCTCGGGGATTTGCC-3′ common to all public LMW-i-type sequences) using BigDye terminator mix (Applied Biosystems) as per the manufacturer’s instructions. The cycle sequencing reactions were performed on a FTS-1 Thermal Sequencer (Corbett Research) using 25 cycles of (95°C for 30 s, 50°C for 20 s and 60°C for 1 min). The extension products were precipitated by addition of 1/10 vol. of 3M sodium acetate pH 5.2 and 2 vol. of ethanol and sequenced on an ABI PRISM 377XL Genetic Analyzer (Applied Biosystems) according to the manufacturer’s instructions.
Allele-specific PCR primer design
Allele-specific PCR primers were designed manually from DNA sequence alignments as described previously (Zhang et al. 2003).
Results
Sequence alignment and Glu-A3 locus LMW-i-type sequence specific PCR
All publicly available LMW glutenin gene sequences were used for alignment using GCG software (WebANGIS). Based on this sequence alignment, five LMW glutenin gene sequences were found to cluster, and these genes were predicted to encode LMW glutenins with an isoleucine amino acid residue at the N-terminus of the mature peptide. The hexaploid wheat cultivars from which these genes were obtained were: cv. Yamhill (X07747, Pitts et al. 1988), cv. Cheyenne (U86030, Cassidy et al. 1998), cv. 1CW (AB008497, Maruyama et al. 1998), cv. Glenlea (pGH3.1, Cloutier et al. 2001) and cv. Norin 61 (AB062876 and AB062878, Ikeda et al. 2002). Two of these genes, pGH3.1 and AB062878, were previously shown to be located at the Glu-A3 locus on chromosome 1A (Cloutier et al. 2001; Ikeda et al. 2002).
Using this information, a primer set (GluA3F1/GluA3R1, Table 1) was designed to facilitate the PCR amplification of the entire open reading frame (ORF) from seven wheat varieties with different Glu-A3 alleles. Using this primer pair, a single band was amplified from cv. Chinese Spring (Glu-A3a), cv. Gabo (Glu-A3b), cv. Cheyenne (Glu-A3c), cv. Suneca (Glu-A3d), cv. Rescue (Glu-A3f) and cv. Glenlea (Glu-A3 g), but not from cv. Halberd that contains the Glu-A3e allele (Fig. 1). A second reverse primer, GluA3R2 (Table 1) located near the 3′ end of the ORF was therefore designed. The primer set GluA3F1/GluA3R2 was used for amplification from the same seven wheat varieties described above. A single band was amplified from the six varieties containing the Glu-A3 a, b, c, e, f and g alleles, respectively, although no amplification product was obtained from cv. Suneca (Glu-A3d allele, Fig. 1).
Locus-specific PCR for low-molecular-weight (LMW)-i-type genes from different Glu-A3 alleles using primer sets GluA3F1/GluA3R1 and GluA3F1/GluA3R2. Template DNAs were as follows: a Chinese Spring (Glu-A3a), b Gabo (Glu-A3b), c Cheyenne (Glu-A3c), d Suneca (Glu-A3d), e Halberd (Glu-A3e), f Rescue (Glu-A3f), g Glenlea (Glu-A3 g). The marker used was the 1-kb Plus DNA Ladder (Invitrogen)
Chromosome assignment of PCR products
Two primer sets (GluA3F1/GluA3R1 and GluA3F1/GluA3R2) were used to amplify products from Chinese Spring and its group 1 nulli-tetrasomic lines. A single band was amplified with Chinese Spring, N1BT1A and N1DT1A, but no PCR products were amplified with N1AT1B (Fig. 2). These results confirmed that the products were amplified from chromosome 1A, the location of the Glu-A3 locus.
Level of sequence polymorphism between Glu-A3 genes and relationships to previously characterised genes
The PCR products obtained using the GluA3F1/GluA3R1 and GluA3F1/GluA3R2 primer sets were directly sequenced. The PCR primers were used for sequencing with one additional primer, GluA3R3, required to obtain full DNA sequence from the PCR product obtained from cv. Suneca (Glu-A3d, see Materials and methods). DNA sequences of the PCR products were assembled using GCG software.
To obtain the whole ORF sequence from cv. Halberd (Glu-A3e), a genomic DNA walking technique was applied as described in Materials and methods. GenBank accession numbers for the genes characterised from Glu-A3a through to Glu-A3g are AY453155 to AY453160, respectively.
The seven DNA sequences derived from cultivars containing different Glu-A3 alleles in the current study were used to construct a 1,199-bp alignment (Fig. 3). Numerous SNPs and several InDels were found between the LMW glutenin gene sequences obtained in this study (Tables 2 and 3, respectively). The sequences obtained from the cultivars containing the Glu-A3 a, b and c alleles showed the highest level of sequence identity (Tables 2, 3). Two single-nucleotide differences were identified between the sequence from cv. Glenlea (Glu-A3 g) and pGH3.1 (also from Glenlea, Cloutier et al. 2001). The sequence from Glu-A3e showed a high level of sequence identity with GenBank sequence AB008497 (Maruyama et al. 1998), with three SNPs and one three-base pair InDel difference. The sequence obtained from cv. Rescue (Glu-A3f) showed a high level of sequence identity with GenBank sequence X07747 (Pitts et al. 1988) with one SNP and one three-base pair InDel difference. The sequence from cv. Suneca (Glu-A3d) was most similar to AB062878 (Ikeda et al. 2002), with 27 SNPs and four InDels difference.
Design of LMW-GS, allele-specific PCR markers
Based on the SNPs between sequences obtained from different Glu-A3 alleles, sets of primer pairs (Table 1) were designed to specifically amplify each Glu-A3 gene where possible. It was not possible to design allele-specific primers from the sequences from the cultivars containing the Glu-A3 b and c alleles, which required the use of a combination of two primer sets for scoring (see below). Primer set GluA3F1/GluA3aR was used to specifically amplify the 586-bp expected product from cv. Chinese Spring containing the Glu-A3a allele (Fig. 4a). Primer set GluA3bF/GluA3bR was shown to amplify the expected product (823 bp) from cvs. Chinese Spring (Glu-A3a), Gabo (Glu-A3b) and Cheyenne (Glu-A3c, Fig. 4b). Primer set GluA3bF/GluA3cR was shown to amplify the expected product (196 bp) from cvs. Chinese Spring (Glu-A3a) and Cheyenne (Glu-A3c, Fig. 4c). Primer set GluA3dF/GluA3dR was shown to amplify the expected product (488 bp) from cv. Suneca (Glu-A3d, Fig. 4d). Primer set GluA3eF/GluA3R2 was shown to amplify the expected product (1,151 bp) from cv. Halberd (Glu-A3e, Fig. 4e), Primer set GluA3F1/GluA3fR was shown to specifically amplify a 1,101-bp product from cv. Rescue (Glu-A3f, Fig. 4f) and GluA3gF/GluA3R2 produced an 861 bp product from cv. Glenlea (Glu-A3 g, Fig. 4g). The seven cultivars above mentioned for gene isolation and ten Australian wheat cultivars containing the Glu-A3b allele, ten cultivars with Glu-A3c, seven cultivars with Glu-A3d and two cultivars with Glu-A3e were selected to check the seven allele-specific markers. All the PCR results matched the corresponding Glu-A3 allele data (Table 4). These results further confirmed that the markers developed here are Glu-A3 allele specific.
Allele-specific PCR markers based on SNPs between different Glu-A3 alleles. Allele specificity is labelled above each gel. Glu-A3abc refers to specificity for the three alleles, a, b and c. Wheat varieties used as PCR templates were Chinese Spring (Glu-A3a), Gabo (Glu-A3b), Cheyenne (Glu-A3c), Suneca (Glu-A3d), Halberd (Glu-A3e), Rescue (Glu-A3f) and Glenlea (Glu-A3g). PCR product sizes are as listed in Table 1. The marker used was the 1-kb Plus DNA Ladder (Invitrogen)
Deduced amino acid sequences of genes from cultivars containing different Glu-A3 alleles
An alignment of the deduced protein sequences from the seven different genes isolated from cultivars containing Glu-A3 alleles a–g is shown in Fig. 5. All genes contain a single, uninterrupted ORF, including the gene isolated from the cultivar containing the Glu-A3e allele, from which no gene product has been identified (Singh and Shepherd 1988). No introns were observed, consistent with the general absence of introns from glutenin genes. Six of the deduced proteins contain an identical, hydrophobic 20 amino acid predicted signal peptide, typical of LMW-GS genes. The predicted protein from the cultivar containing the Glu-A3e allele, however, contains a cysteine residue at position 7 in the predicted signal sequence. All predicted mature proteins have an isoleucine at the truncated N-terminus, eight cysteine residues located within the C-terminal domain and two poly-glutamine repeats of up to 15 residues, as observed for LMW-i-type proteins reported previously (Cloutier et al. 2001; Ikeda et al. 2002).
Alignment of deduced protein sequences from seven genes amplified from seven cultivars containing Glu-A3 alleles a–g. The predicted 20-amino acid signal peptide is also included in the alignment. The start of the deduced mature peptide (matP) is indicated. Conserved cysteine residues are indicated by arrows. Dashes indicate gaps introduced into the alignment
Comparison of predicted and observed i-type LMW-GS from cultivars containing different Glu-A3 alleles by SDS-PAGE
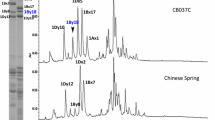
Reduced and alkylated glutenin subunits from the cultivars containing Glu-A3 alleles a–g were extracted and fractionated by one-step SDS-PAGE as described previously (Zhang et al. 2003). Molecular mass standards were used to compare mobility of the Glu-A3 B-type LMW-GS expressed by the different cultivars (Fig. 6). Values obtained are shown in Table 5.
SDS-PAGE analysis of proteins expressed from cultivars containing different Glu-A3 alleles. M Molecular weight standard (Benchmark Protein Ladder, Invitrogen), d cv. Suneca (Glu-A3d allele), g Glenlea (Glu-A3 g), b Gabo (Glu-A3b), c Cheyenne (Glu-A3c), a Chinese Spring (Glu-A3a), f Rescue (Glu-A3f), e Halberd (Glu-A3e). Bands corresponding to the B-type low-molecular-weight-glutenin subunits linked to each allele are highlighted on the left of each gel lane
Discussion
Approximately 60 LMW-GS genes or pseudogenes have been isolated from genomic or cDNA libraries (Bartels and Thompson 1983; Pitts et al. 1988; Colot et al. 1989; Cassidy et al. 1998; Ikeda et al. 2002) and via gene-specific PCR (Masci et al. 1998; Benmoussa et al. 2000; Cloutier et al. 2001; Ikeda et al. 2002). LMW-GS genes have been divided into three broad classes based on the first amino acid residue of the predicted mature protein: s-type (serine), m-type (methionine) and i-type (isoleucine, Cloutier et al. 2001). LMW-GS genes have been further classified into 12 groups based on alignment of the conserved N- and C-terminal domains of the deduced amino acid sequences (Ikeda et al. 2002). Whereas most previous studies have isolated LMW-GS genes from one specific hexaploid wheat cultivar, here we report the isolation and characterisation of one LMW-i-type gene from seven hexaploid wheat cultivars, each with a different Glu-A3 allele.
Sequences of the LMW-i-type gene from Glu-A3g (this study) and pGH3.1 (Cloutier et al. 2001) are both derived from cv. Glenlea. Two single SNPs were identified over a 1,199 bp alignment of these sequences. These SNPs may indicate these are two very similar but non-identical genes, or may be attributable to sequencing errors or errors generated during propagation in Escherichia coli (Cooper and Krawczak 1993). Sequences from Glu-A3c (this study) and U86030 (Cassidy et al. 1998) are both derived from cv. Cheyenne. However, 18 SNPs and three InDels were identified between these two sequences, indicating that these are probably paralogous genes. In addition, no gene with a high level of sequence identity with AB062878 (Ikeda et al. 2002) was observed for any of the Glu-A3-derived sequences in the current study. These results are consistent with the observation of Ikeda et al. (2002) that cultivars containing some Glu-A3 alleles (as classified according to the observed protein profile) may contain more than one LMW-i-type gene or pseudogene.
The polymorphism levels between LMW-i-type gene sequences obtained in the current study for cultivars containing different Glu-A3 alleles are much higher than expected for orthologous genes. The SNP rate in wheat low copy sequences (Bryan et al. 1999) and multi-copy-gene family γ-gliadin sequences (Zhang et al. 2003) was approximately one SNP per kilobase. A similar SNP level was also observed in the human genome (Cooper et al. 1985). In the current study, only comparison of the LMW-i-type genes derived from Glu-A3a and Glu-A3c shows a level of polymorphism consistent with this SNP level. Five to 35 times this SNP rate was observed between the other LMW-i-type gene sequences from different Glu-A3 alleles. Therefore, it appears unlikely that all the LMW-i-type genes sequenced from different Glu-A3 alleles in this study are from the same origin, as confirmed for two other low-copy wheat genes encoding plastid acetyl-CoA carboxylase and 3-phosphoglycerate kinase (Huang et al. 2002). The LMW-i-type genes from Glu-A3a, Glu-A3b and Glu-A3c probably represent orthologues, the LMW-i-type genes from Glu-A3e and Glu-A3 g may have the same origin, whereas LMW-i-type genes from Glu-A3d and Glu-A3f appear to be from separate origins. Therefore, there appear to be at least four different origins for the Glu-A3 loci LMW-i-type genes of hexaploid wheat. Another possible explanation is that a monomorphic progenitor Glu-A3 locus underwent sequence rearrangement during allopolyploidy formation as described by Ozkan et al. (2001) and Shaked et al. (2001).
It is estimated that several dozen genes or pseudogenes comprise the Glu-3 loci in hexaploid wheat (Cassidy et al. 1998). Alleles Glu-A3 a, b, c and f appear to encode a single, B-type LMW-GS, whereas the d allele encodes two B-type subunits and no product has been detected for the e allele (Gupta and Shepherd 1990). Just one gene was isolated from each cultivar containing a different Glu-A3 allele in this study. All seven LMW-GS genes characterised in this study contain a single, predicted ORF that encodes an LMW-GS i-type protein (Cloutier et al. 2001), also described as ‘type VI’ LMW-GS (Ikeda et al. 2002) and reported previously by Pitts et al. (1988), Cassidy et al. (1998) and Maruyama et al. (1998). Where known, the location of genes encoding i-type LMW-GS is chromosome 1A [the gene of Cloutier et al. (2001), two genes of Ikeda et al. (2002), seven genes in this study].
The i-type LMW-GS characterised by Cloutier et al. (2001) from cv. Glenlea (‘LMW-50’, encoded by clone pGH3.1) is almost identical to the sequence we obtained from the same cultivar (Glu-A3g, one amino acid difference) and these authors demonstrated that this gene encodes a protein of approximately 50 kDa observed by SDS-PAGE. This contrasted with a molecular mass of 42.6 kDa for the predicted mature protein. We observed differences of between 11 kDa and 13 kDa between the predicted proteins encoded by the genes from cultivars containing Glu-A3 alleles a–g and the SDS-PAGE-estimated molecular masses of the proteins encoded by these alleles (Fig. 6; Table 5), with an estimated molecular mass of 56 kDa for the Glu-A3g-encoded LMW-GS of cv. Glenlea. It is interesting to note that the ranking of the mobilities of the SDS-PAGE-observed proteins and the predicted proteins from the cultivars containing the corresponding Glu-A3 alleles are similar (predicted: g>b>c~a>d~f, observed: g>b>c>a>d~f, Table 5). It is known that HMW-GS do not migrate according to their predicted molecular masses, with the molecular mass estimated by SDS-PAGE being significantly higher than that predicted by amino acid composition (Bunce et al. 1985; Anderson et al. 1989; Shewry et al. 1989). Bunce et al. (1985) have attributed the over-estimation of the molecular weight of HMW-GS in SDS-PAGE to their particular structure and specifically to the central repetitive domain that may not be completely denatured under the conditions of SDS-PAGE (Goldsbrough et al. 1989). It is likely that the structurally similar repetitive domain of the LMW-GS is involved in the anomalous mobility of these proteins.
The deduced amino acid sequence of the gene isolated from the Glu-A3e-containing cultivar contains a single ORF despite no protein being observed for this allele. It is interesting to note that this predicted protein contains a cysteine in the signal sequence not found in any other LMW-GS signal sequence.
LMW glutenins have been shown to significantly effect wheat dough properties (Payne et al. 1987; Gupta et al. 1989, 1994; Pogna et al. 1990). The effects of individual Glu-3 alleles are largely additive with those of Glu-1 alleles (Gupta et al. 1989; Pogna et al. 1990; Sontag-Strohm et al. 1996; Redaelli et al. 1997; Wesley et al. 1999). Interactions between these loci also have significant effects (Gupta et al. 1994; Luo et al. 2001). The absence of a cysteine residue in the N-terminal or repetitive domains of the mature protein in i-type LMW-GS may have significant effects on gluten polymer formation. This could be due to the exclusion of the repetitive domain of i-type LMW-GS from the disulphide-linked network frame or by this LMW-GS type acting as a chain terminator (Ikeda et al. 2002). For the cv. Gleanlea (GluA3g), Cloutier et al. (2001) have reported positive effects for the Glu-A3 locus on dough strength. In Australia, specific LMW-GS alleles have been actively selected against is certain breeding programs, for example Glu-B3e (Lawrence 1986) and Glu-A3e (Eagles et al. 2001).
In order to obtain DNA markers for all of the Glu-A3 alleles, we designed a series of allele-specific primers according to the principles that we have successfully used for wheat γ-gliadin gene detection (Zhang et al. 2003). Five allele-specific primer sets were successfully designed for Glu-A3a, Glu-A3d, Glu-A3e, Glu-A3f and Glu-A3g detection. No Glu-A3b allele-specific primer could be designed because the only Glu-A3b-specific SNP is located in the repetitive sequence region of the gene. Detection of this allele therefore requires the use of the GluA3abc marker in combination with either the GluA3a or GluA3ac marker (depending on the cross). No Glu-A3c-allele specific primer could be designed as no allele-specific SNP could be identified relative to the sequences obtained from the other six Glu-A3 alleles. Detection of this allele therefore requires the use of the GluA3ac marker in combination with the GluA3a marker. Although the allele-specific markers described in this study are dominant, the use of pairs of markers together would provide the capacity to detect heterozygous plants, with the exception of the crosses Glu-A3 a × b, b × c and a × c.
A similar approach to that used in the current study may now be used to identify genes and markers linked to alleles of the Glu-B3 and Glu-D3 loci to provide a comprehensive set of LMW glutenin markers for use in marker-assisted breeding.
References
Anderson OD, Green FC (1989) The characterization and comparative analysis of high-molecular-weight glutenin genes from genomes A and B of a hexaploid bread wheat. Theor Appl Genet 77:689–700
Bartels D, Thompson RD (1983) The characterization of cDNA clones coding for wheat storage proteins. Nucleic Acids Res 11:2961–2977
Benmoussa M, Vézina L-P, Pagé M, Yelle S, Laberge S (2000) Genetic polymorphism in low-molecular-weight glutenin genes from Triticum aestivum, variety Chinese Spring. Theor Appl Genet 100:789–793
Brites C, Carrillo JM (2001) Influence of high molecular weight (HMW) and low molecular weight (LMW) glutenin subunits controlled by Glu-1 and Glu-3 loci on durum wheat quality. Cereal Chem 78:59–63
Brown JWS, Flavell RB (1981) Fractionation of wheat gliadin and glutenin subunits by two-dimensional electrophoresis and the role of group 6 and group 2 chromosomes in gliadins synthesis. Theor Appl Genet 59:349–359
Bryan GJ, Stephenson P, Collins A, Kirby J, Smith JB, Gale MD (1999) Low levels of DNA sequence variation among adapted genotypes of hexaploid wheat. Theor Appl Genet 99:192–198
Bunce NAC, White RP, Shewry PR (1985) Variation in estimates of molecular weight of cereal prolamins by SDS-PAGE. J Cereal Sci 3:131–142
Burnouf T, Bietz JA (1984) Reversed-phase high-performance liquid chromatography of reduced glutenin, a disulphide bonded protein of wheat endosperm. J Chromatogr 299:185–199
Cassidy BG, Dvorak J, Anderson OD (1998) The wheat low-molecular-weight glutenin genes: characterization of six new genes and progress in understanding gene family structure. Theor Appl Genet 96:743–750
Ciaffi M, Lee YK, Tamas L, Gupta R, Skerritt J, Appels R (1999) The low-molecular-weight glutenin subunit proteins of primitive wheats. III. The genes from D-genome species. Theor Appl Genet 98:135–148
Cloutier S, Rampitsch C, Penner GA, Lukow, OM (2001) Cloning and expression of a LMW-i glutenin gene. J Cereal Sci 33:143–154
Colot V, Bartels D, Thompson R, Flavell R (1989) Molecular characterization of a active wheat LMW glutenin gene and its relation to other wheat and barley prolamin genes. Mol Gen Genet 216:81–90
Cooper DN, Krawczak M (eds) (1993) Human gene mutation. Coronet Books, Bios Scientific, Oxford, UK
Cooper DN, Smith BA, Cooke H, Niemann S, Schmidtke, J (1985) An estimate of unique sequence heterozygosity in the human genome. Hum Genet 69:201–205
Cornish GB, Burridge PM, Palmer GA, Wrigley CW (1993) Mapping the origins of some HMW and LMW glutenin subunit alleles in Australian germplasm. Proceedings of the 42nd Australian Cereal Chemistry Conference, Sydney, pp 255–260
Devos KM, Bryan GJ, Collins AJ, Stephenson P, Gale MD (1995) Application of two microsatellite sequences in wheat storage proteins as molecular markers. Theor Appl Genet 90:247–252
D’Ovidio R (1993) Single-seed PCR of LMW glutenin genes to distinguish between durum wheat cultivars with good and poor technological properties. Plant Mol Biol 22:1173–1176
Eagles HA, Bariana HS, Ogbonnaya FC, Rebetzke GJ, Hollamby GJ, Henry RJ, Henschke PH, Carter M (2001) Implementation of markers in Australian wheat breeding. Aust J Agric Res 52:1349–1356
Gale KR, Ma W, Zhang W, Rampling L, Hill AS, Appels, R, Morris P, Morell M (2001) Simple high-throughput DNA markers for genotyping in wheat. In: Eastwood R, et al (eds) 10th Australian Wheat Breeding Assembly Proceedings, pp 26–31
Goldsbrough AP, Bulleid NJ, Freedman RB, Flavell RB (1989) Conformational differences between two wheat (Triticum aestivum) high molecular weight glutenin subunits are due to a short region containing six amino acids differences. Biochem J 263:837–842
Gupta RB, MacRitchie F (1994) Allelic variation at glutenin subunit and gliadin loci, Glu-1, Glu-3, and Gli-1, of common wheats. II. Biochemical basis of the allelic effects on dough properties. J Cereal Sci 19:19–29
Gupta RB, Shepherd KW (1988) LMW-GS in wheat: their variation, inheritance, and association with bread-making quality. In: Proceedings of the 7th International Wheat Genetics Symposium, Cambridge, UK, pp 943–949
Gupta RB, Shepherd KW (1990) Two-step one-dimensional SDS-PAGE analysis of LMW subunits of glutenin. I. Variation and genetic control of the subunits in hexaploid wheats. Theor Appl Genet 80:65–74
Gupta RB, Singh NK, Sheperd KW (1989) The cumulative effects of allelic variation in LMW and HMW glutenin subunits on physical dough properties in the progeny of two bread wheats. Theor Appl Genet 77:57–62
Gupta RB, Bekes F, Wrigley CW (1990a) Predicting values of LMW glutenin alleles for dough quality of bread wheat. In: Bushuk W, Tkachuk R (eds) Gluten proteins 1990. American Association of Cereal Chemists, St. Paul, Minnesota, USA, pp 615–621
Gupta RB, Bekes F, Wrigley CW, Moss HJ (1990b) Prediction of wheat dough quality in breeding on the basis of LMW and HMW glutenin subunit composition. In: Proceedings of the 6th Australian Wheat Breeders Society Assembly, Tamworth, Australia, pp 217–225
Gupta RB, Bekes F, Wrigley CW (1991) Prediction of physical dough properties from glutenin subunit composition in bread wheats: correlation studies. Cereal Chem 68:328–333
Gupta RB, Paul JG, Cornish GB, Palmer GA, Bekes F, Rathjen AJ (1994) Allelic variation at glutenin subunit and gliadin loci, Glu-1, Glu-3, and Gli-1, of common wheats. I. Its additive and interaction effects on dough properties. J Cereal Sci 19:9–17
Huang S, Sirikhachornkit A, Su X, Faris J, Gill B, Haselkorn R, Gornicki P (2002) Genes encoding plastid acetyl-CoA carboxylase and 3-phosphoglycerate kinase of the Triticum/Aegilops complex and the evolutionary history of polyploid wheat. Proc Natl Acad Sci USA 99:8133–8138
Ikeda TM, Nagamine T, Fukuoka H, Yano H (2002) Identification of new low-molecular-weight glutenin subunit genes in wheat. Theor Appl Genet 104:680–687
Jackson EA, Holt LM, Payne PI (1983) Characterization of high-molecular-weight gliadin and low-molecular-weight glutenin subunits of wheat endosperm by two-dimensional electrophoresis and the chromosomal location of their controlling genes. Theor Appl Genet 66:29–37
Jackson EA, Morel M-H, Sontag-Strohm T, Branlard G, Metakovsky, Radaelli R (1996) Proposal for combining the classification systems of alleles of Gli-1 and Glu-3 loci in bread wheat (Triticum aestivum L.) J Genet Breed 50:321–336
Lawrence GJ (1986) The high-molecular-weight glutenin subunit composition of Australian wheat cultivars. Aust J Agric Res 37:125–133
Lew EJL, Kuzmicky DD, Kasarda DD (1992) Characterization of low molecular weight glutenin subunits by reversed-phase high-performance liquid chromatography, sodium dodecyl sulfate-polyacrylamide gel electrophoresis, and N-terminal amino acid sequencing. Cereal Chem 69:508–515
Luo C, Griffin WB, Branlard G, McNeil DL (2001) Comparison of low- and high molecular-weight wheat glutenin allele effects on flour quality. Theor Appl Genet 102:1088–1098
Marchylo BA, Kruger JE, Hatcher DW (1989) Quantitative reverse-phase high-performance liquid chromatographic analysis of wheat storage proteins as a potential quality prediction tool. J Cereal Sci 9:113–130
Margiotta B, Colaprico G, D’Ovidio R, Lafiandra D (1993) Characterization of high Mr subunits of glutenin by combined chromatographic (RP-HPLC) and electrophoretic separation and restriction fragment length polymorphism (RFLP) analyses of their encoding genes. J Cereal Sci 17:221–236
Maruyama N, Ichise K, Katsube T, Kishimoto T, Kawase S, Matsumura Y, Takeuchi Y, Sawada T, Utsumi S (1998) Identification of major wheat allergens by means of the Escherichia coli expression system. Eur J Biochem 255:739–745
Masci S, D’Ovidio R, Lafiandra, D, Kasarda DD (1998) Characterization of a low-molecular weight glutenin subunit gene from bread wheat and the corresponding protein that represents a major subunit of the glutenin polymer. Plant Physiol 118:1147–1158
Metakovsky EV, Wrigley CW, Bekes F Gupta RB (1990) Gluten polypeptides as useful genetic markers of dough quality in Australian wheats. Aust J Agric Res 41:289–306
Nieto-Taladriz MT, Perretant MR, Rousset M (1994) Effect of gliadins and HMW and LMW subunits of glutenin on dough properties in the F1 recombinant inbred lines from a bread wheat cross. Theor Appl Genet 88:81–88
Ozkan H, Levy AA, Feldman M (2001) Allopolyploidy-induced rapid genome evolution in the wheat (Aegilops-Triticum) group. Plant Cell 13:1735–1747
Payne PI, Jackson EA, Holt LM (1984) The association between γ-gliadin 45 and gluten strength in durum varieties: a direct causal effect or the result of genetic linkage? J Cereal Sci 2:73–81
Payne PI, Seekings JA, Worland AJ, Jarvis MG, Holt LM (1987) Allelic variation of glutenin subunits and gliadins and its effect on bread making quality in wheat: analysis of F5 progeny from Chinese Spring × Chinese Spring (Hope 1A). J Cereal Sci 6:103–118
Pitts EG, Rafalski JA, Hedgcoth C (1988) Nucleotide sequence and encoded amino acid sequence of a genomic gene region for a low molecular weight glutenin. Nucleic Acids Res 16:11376
Pogna NE, Autran JC, Mellini F, Lafiandra D, Feillet P (1990) Chromosome 1B-encoded gliadins and glutenin subunits in durum wheat: genetics and relationship to glutenin strength. J Cereal Sci 11:15–34
Redaelli R, Pogna NE, Ng PKW (1997) Effects of prolamins encoded by chromosomes 1B and 1D on the rheological properties of dough in near-isogenic lines of bread wheat. Cereal Chem 74:102–107
Ruiz M, Carrillo JM (1995) Separate effects on gluten strength of Gli-1 and Glu-3 prolamine genes on chromosomes 1A and 1B in durum wheat. J Cereal Sci 21:137–144
Sears ER, Miller TE (1985) The history of Chinese Spring wheat. Cereal Res Commun 13:261–263
Shaked H, Kashkush K, Ozkan H, Feldman M, Levy AA (2001). Sequence elimination and cytosine methylation are rapid and reproducible responses of the genome to wide hybridization and allopolyploidy in wheat. Plant Cell 13:1749–1759
Shewry PR, Halford NG, Tatham AS (1989) The high molecular weight subunits of wheat, barley and rye: genetics, molecular biology, chemistry and role in wheat gluten structure and functionality. In: Miflin BJ (ed) Oxford surveys of plant and molecular cell biology, vol 6. Oxford University Press, London, pp 163–219
Siebert PD, Chenchik A, Kellogg DE, Lukyanov KA, Lukyanov SA (1995) An improved PCR method for walking in uncloned genomic DNA. Nucleic Acids Res 23:1087–1088
Singh NK, Shepherd KW (1988) Linkage mapping of genes controlling endosperm storage proteins in wheat. 1. Genes on the short arms of group 1 chromosomes. Theor Appl Genet 75:628–641
Skylas DJ, Mackintosh JA, Cordwell SJ,Walsh BJ, Harry J, Blumenthal C, Copeland L,Wrigley CW, Rathmell WG (2000) Proteome approach to the characterisation of protein composition in the developing and mature wheat-grain endosperm. J Cereal Sci 52:169–188
Sontag-Strohm T, Payne PI, Saovaara H (1996) Effect of allelic variation of glutenin subunits and gliadins on baking quality in the progeny of two biotypes of bread wheat cv. Ulla. J Cereal Sci 24:115–124
Sutton KH (1991) Qualitative and quantitative variation among high molecular weight subunits of glutenin detected by reverse-phase high-performance liquid chromatography. J Cereal Sci 14:25–34
Weegels PL, Hamer RJ, Schofield JD (1996) Functional properties of wheat glutenin. J Cereal Sci 23:1–18
Wesley AS, Lukow OM, Ames N, Kovaks MIP, McKenzie RIH, Brown D (1999) Effect of single substitution of glutenin or gliadin proteins on flour quality of Alpha 16, a Canada Prairie Spring wheat breeders line. Cereal Chem 76:743–747
Wesley AS, Lukow OM, McKenzie RIH, Ames N, Brown D (2001) Effect of multiple substitution of glutenin or gliadin proteins on flour quality of Canada Prairie Spring wheat. Cereal Chem 78:69–73
Zhang W, Gianibelli MC, Ma W, Rampling L, Gale KR (2003) Identification of SNPs and development of AS-PCR markers for γ-gliadin alleles in Triticum aestivum. Theor Appl Genet 107:130–138
Acknowledgements
This work was supported by Graingene, a research consortium of CSIRO Plant Industry, AWB Ltd and the Grains Research and Development Corporation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by P. Langridge
Rights and permissions
About this article
Cite this article
Zhang, W., Gianibelli, M.C., Rampling, L.R. et al. Characterisation and marker development for low molecular weight glutenin genes from Glu-A3 alleles of bread wheat (Triticum aestivum. L). Theor Appl Genet 108, 1409–1419 (2004). https://doi.org/10.1007/s00122-003-1558-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-003-1558-8