Abstract
Global warming significantly impacts aquatic ecosystems, with changes in the salt environment negatively affecting the physiological responses of fish. We investigated the impact of hyposalinity on the physiological responses and intestinal microbiota of Sebastes schlegelii under the context of increased freshwater influx due to climate change. We focused on the osmoregulatory capacity, oxidative stress responses, and alterations in the intestinal microbiome of S. schlegelii under low-salinity conditions. Our findings revealed compromised osmoregulatory capacity in S. schlegelii under low-salinity conditions, accompanied by the activation of oxidative stress responses, indicating physiological adaptations to cope with environmental stress. Specifically, changes in Na+/K+-ATPase (NKA) activity in gill tissues were associated with decreased osmoregulatory capacity. Furthermore, the analysis of the intestinal microbiome led to significant changes in microbial diversity. Exposure to low-salinity environments led to dysbiosis, with notable decreases in the relative abundance of Gammaproteobacteria at the class level and specific genera such as Enterovibrio, and Photobacterium. Conversely, Bacilli classes, along with genera like Mycoplasma, exhibited increased proportions in fish exposed to low-salinity conditions. These findings underscore the potential impact of environmental salinity changes on the adaptive capacity of fish species, particularly in the context of aquaculture. Moreover, they highlight the importance of considering both physiological and microbial responses in understanding the resilience of aquatic organisms to environmental stress. Additionally, they highlight the importance of intestinal microbiota analyses in understanding the immune system and disease management in fish.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Various environmental factors, such as water temperature, salinity, and pH, influence the ecology, metabolism, immune response, and physiological reactions of aquatic organisms (Becker et al. 2019; Li et al. 2022). Salinity is a crucial environmental factor that affects the ecological distribution, metabolism, and reproduction of aquatic organisms in both marine and freshwater environments (Kültz 2015). During summer monsoons, increased rainfall leads to an influx of freshwater into the oceans, producing freshwater dilution that results in decreased salinity in the affected marine environment (Kwak et al. 2023). For instance, while the typical salinity of seawater is 33–35 psu (practical salinity units), salinities ranging from 24 to 32 psu have been measured near the East China Sea because of the influx of low-salinity water. Specifically, salinity along the western coast of Jeju Island, located in the East China Sea off South Korea, has been shown to decrease to 26 psu before the passage of typhoons (Choi et al. 2021).
Freshwater dilution directly influences the ion concentration and osmolality of body fluids in marine fish via their interaction with aquatic habitats as fish regulate ion exchange within their body fluids to maintain homeostasis (Velotta et al. 2022). Gills play a crucial role in osmoregulation by allowing the transport of osmotically active solutes via specialized membrane transport proteins (Lee et al. 2020). Na+/K+ pumps are present in the plasma membrane of all animal cells. The cell utilizes energy derived from ATP hydrolysis to maintain membrane potential via the transport of Na+ and K+ ions (NKA) (Hiroi and McCormick 2012). Indeed, the modulation of NKA activity plays an important role in regulating ion exchange in the bodily fluids of fish exposed to various saline environments, with decreased ion excretion through the gills typically being observed in marine fish exposed to low-salinity environments to maintain body fluid osmolality (Li et al. 2022).
Shifts in the salinity of oceans (or changes in other environmental variables, such as temperature and pH) have negative impacts on the physiology of marine organisms, inducing antioxidant reactions within their bodies (Hennersdorf et al. 2016; Sin et al. 2020). In short, exposure to low-salinity environments causes a stress response in these organisms, which leads to the production of reactive oxygen species (ROS), such as superoxide anion radicals (O2−) and hydrogen peroxide (H2O2). If toxic ROS persist within the body without adequate removal, they can induce damage to DNA and cell membrane structures and impair protein function (Kiran et al. 2020). Consequently, the organisms alter their metabolism through the activation of antioxidant enzymes, such as superoxide dismutase (SOD) and catalase (CAT) (Huang et al. 2021). These enzymes convert ROS into water and oxygen to maintain internal equilibrium and ensure survival (Jomova et al. 2023), thereby playing a crucial role in enabling marine organisms to cope with stress from the external environment.
Moreover, unsuitable alterations in the external environment alter the relative abundance of microbial communities residing within host organisms, including that of intestinal microbiota in fish (Bissett et al. 2013). Such changes in intestinal microbial communities are pivotal in the health of the host, affecting physiological functions, hormonal changes, digestion, susceptibility to disease, and other factors (Sullam et al. 2012; Wang et al. 2018). These microbes normally promote intestine health by boosting the immune system of the host, generating beneficial metabolites, and performing various advantageous functions, such as strengthening the intestine barrier (Rastelli et al. 2019). However, despite the considerable contribution of intestinal microbiota to host health, research on the impact of changes in environmental salinity on fish intestinal microbiota remains limited (Tian et al. 2020; Zhao et al. 2020).
In the present study, we selected Sebastes schlegelii, a fish species that is susceptible to hyposalinity changes to the marine environment, as the experimental subject (Oh et al. 2014). We aimed to investigate the effects of exposure to low-salinity environments on the physiological responses and intestinal microbial community in S. schlegelii, a coastal resident species in South Korea. We analyzed the expression and activity of NKA, which is related to osmoregulation in gill tissues, and examined changes in the plasma osmolality. Additionally, we confirmed the occurrence of ROS-mediated oxidative stress by analyzing hydrogen peroxide (H2O2) production in the liver and the expression levels of the antioxidant enzymes SOD and CAT. Lastly, we investigated changes in the intestinal microbial community using 16S rRNA gene sequencing to elucidate the impact of marine hyposalinity on the physiological responses of S. schlegelii.
Materials and methods
Study animals and experimental design
S. schlegelii individuals (body length 14.9 ± 0.9 cm, mass 57.5 ± 3.7 g) were purchased from the Tong-young fish market (South Korea) and transported to the laboratory, where they were acclimated for 1 week in six 350-L recirculating filtration tank systems filled with continuously oxygenated seawater. Seawater was maintained at a salinity of 34 ± 0.5 psu, temperature of 18.0 ± 1.0 ℃, and pH of 8.0 ± 0.1. To investigate the impact of relative hypoosmotic stress on S. schlegelii, the fish were divided into three treatment groups: a control group (100% seawater, 34 psu) and two low-salinity groups (80% seawater, 27 psu; and 60% seawater, 20 psu). In total, 6 experimental tanks were set up, with each tank containing 15 randomly selected fish. The experiment was conducted in duplicate. Low-salinity solutions were prepared by diluting standard seawater with freshwater. Water in the recirculating filtration tanks was continuously circulated. The fish were gradually acclimated to each salinity level and then transferred abruptly from 100% seawater to 60% and 80% seawater for 5 days (120 h) each. During the experimental period, no feed was provided and no mortalities were observed.
For each treatment group, individuals of S. schlegelii were exposed to salinity stress for 0, 12, 24, 72, and 120 h. After each exposure time, three individuals were randomly sampled from each tank and anesthetized using clove oil (Sigma-Aldrich, St. Louis, MO, USA). Blood was collected from the caudal vein using heparin-coated syringes. Plasma samples were separated from the blood by centrifugation (4 °C at 6000 × g for 10 min) and stored at − 80 °C until further analysis. At each sampling period, the liver, gill, and intestinal tissues were collected from each individual. The collected tissue samples were stored at − 80 °C until they were processed for analysis.
RNA extraction and real-time qPCR
Total RNA was extracted from the gill and liver tissues of S. schlegelii using TRI Reagent® (TR188, Molecular Research Center, Cincinnati, OH, USA) following the manufacturer’s instructions. The RNA purity (A260/A280) was determined to be approximately 1.9–2.0, and the contamination level (A260/A230) was around 2.0–2.2. Subsequently, 2 μg of the extracted RNA was reverse-transcribed into complementary DNA (cDNA) using an oligo-(dT)15 anchor primer (Cosmogenetech, Korea) and the Moloney Murine Leukemia Virus (M-MLV) cDNA Synthesis Kit (EZ006S, Ezynomics, Korea) according to the provided protocol. The synthesized cDNA was stored at − 20 °C and was diluted 1:49 prior to performing the real-time polymerase chain reaction (qPCR) analysis.
To assess the relative expression levels of NKA mRNA in the gill and SOD and CAT mRNA in the liver, qPCR was conducted. Primers for qPCR were designed using known mRNA sequences obtained from the National Center for Biotechnology Information (NCBI; Table 1). Amplification was carried out using a Bio-Rad CFX96™ real-time PCR detection system (Bio-Rad Laboratories, USA) along with iQ SYBR Green Supermix (Bio-Rad, USA). The internal control used for normalizing the relative mRNA expression levels was ef1-α. The thermal cycling protocol included an initial denaturation at 95 °C for 5 min, followed by 30 cycles of 95 °C for 10 s (denaturation), 51 °C for 15 s (annealing), and 72 °C for 20 s (extension). Results were expressed as changes relative to the corresponding 18S rRNA cycle threshold (ΔCt), with calibration performed using the 2−ΔΔCt method.
Detection of NKA activity in gill tissues
Gill tissues were initially soaked in a solution of 200 mM sucrose, 5 mM Na2 EDTA, and 100 mM imidazole–HCl buffer (pH 7.6) and then homogenized with SEI solution and 0.1% sodium deoxycholate. After centrifuging at 4 °C and 2000 × g for 5 min, 10 μL of the supernatant was combined with reaction mixture A (125 mM NaCl, 75 mM KCl, 7.5 mM MgCl2, 5 mM Na2 ATP, and 100 mM imidazole–HCl buffer, pH 7.6) and reaction mixture B (which consists of mixture A and 10 mM ouabain). The mixture was incubated at 37 °C for 30 min, and the reaction was terminated by adding 50 μL of 30% trichloroacetic acid. The levels of inorganic phosphorus, with and without ouabain, were determined using a phosphate colorimetric assay kit (BioVision, US), and the protein concentration of the tissues was assessed using a Pierce BCA Protein Kit (Thermo Fisher Scientific, Waltham, MA, USA).
Osmolality and H2O2measurements
Plasma osmolality was measured using a VAPRO® Vapor Pressure Osmometer (ELITechGroup, USA). Hydrogen peroxide (H2O2) concentrations were determined using a commercial kit (PeroxiDetect™ Kit, PD1-1KT, Sigma, USA), and absorbance was read at 560 nm using a 2030 multilabel reader (Victor X3, Perkin Elmer, Waltham, MA, USA).
Sample collection from intestinal mucosa and DNA extraction
At the final exposure time point (120 h), DNA was extracted from the intestinal mucosa of three randomly selected fish. Then, 30 mg of visceral membrane tissue, with feces removed, was preserved in 300 μL of AccuStool Preservation Buffer (AccuGene, South Korea). DNA extraction was conducted using the AccuBlood DNA preparation kit (AccuGene). The preserved tissue was treated with 300 μL of lysis buffer and 30 μL of proteinase K, and then incubated at 60 °C with shaking at 2000 rpm for 1 h using a thermomixer (Eppendorf, Germany). Following centrifugation at 21,952 rpm for 1 min, the supernatant was collected into a new tube. An equal volume of 100% ethanol was added to the supernatant, which was then transferred to a silica column. All subsequent steps were performed according to the manufacturer’s instructions.
Intestinal microbiome analysis
The hypervariable V3-V4 region of the 16S-rRNA gene was amplified from DNA extracts using KAPA HiFi HotStart ReadyMix (Roche) and barcoded fusion primers (341F: 5′-CCTACGGGNGGCWGCAG-3′ and 805R: 5′-GACTACHVGGGTATCTAATCC-3′) containing Nextera adaptors (King et al. 2019) over 25 PCR cycles. PCR products were purified with HiAccuBeads (AccuGene). The amplicon libraries were pooled at equal molar ratios and sequenced on an Illumina MiSeq platform using the MiSeq Reagent Kit v3 (Illumina, San Diego, CA, USA). Post-sequencing, quality control was performed to remove adapters and low-quality sequences (score < 20) using FastQC v.0.11.8 and Cutadapt v.1.18. The preprocessed sequence data were clustered at a 0.01 cutoff, ensuring a sequence similarity of over 99% for the amplicon sequence variants (ASVs). The ASV abundance and ASV taxonomy are provided in the Supplementary Tables (Table S1 and S2). Further analyses involved assessing α-diversity and the Shannon and Chao1 indices to evaluate species dominance within each sample. Principal coordinate analysis (PCoA) plots for ASV-based groups were generated using the QIIME2 platform (Bolyen et al. 2019). Bacterial taxonomy classification from phylum to genus was performed using the SILVA reference database version 138.1 (Quast et al. 2012). The raw sequence data from the intestinal mucosa sequencing have been submitted to the public database NCBI Sequence Read Archive (SRA) (Bioproject number: PRJNA1122984).
Statistical analysis
Data analysis was performed using SPSS Statistics version 25.0 (IBM SPSS Inc., Armonk, NY, USA). Differences in salinity across the same exposure days were evaluated using one-way ANOVA. To examine the interaction effects of salinity and exposure duration, a two-way ANOVA was conducted, followed by Tukey’s post hoc test for further analysis. A significance level of 95% (p < 0.05) was applied to determine significant differences in the parameters assessed. Results are presented as the mean ± standard error (SE).
Results
Changes in NKA mRNA expression and activity in gills
The expression of NKA mRNA and activity of NKA were investigated in the gills of S. schlegelii fish following their exposure to low-salinity conditions (Fig. 1). In both the 60% and 80% groups, NKA mRNA expression decreased significantly after exposure to a low-salinity environment for 12 h (p < 0.05), and this state was maintained up to 120 h of exposure (day 5). However, there was no significant change in NKA mRNA expression or activity in the control group (100%) during the experiment (p > 0.05) (Fig. 1a). The activity of NKA in the gills showed a trend similar to that of NKA mRNA expression (Fig. 1b). No change was observed in the 100% group; in contrast, NKA activity decreased significantly at 12 h of exposure to low-salinity conditions in the 60% and 80% groups (p < 0.05) and remained constant thereafter (p > 0.05).
Changes in the a mRNA expression of Na + /K + -ATPase (NKA) and b activity of NKA in gill tissues of S. schlegelii exposed to low salinity for 120 h (5 days). Different letters indicate significant differences between the same experimental groups for different exposure times (p < 0.05). Different numbers indicate significant differences between different experimental groups for the same exposure duration (p < 0.05). All values represent the mean ± standard error (n = 3). Control group, 34 psu (100% seawater); low-salinity groups, 20 psu (60% seawater) and 27 psu (80% seawater)
Changes in plasma osmolarity
Plasma osmolarity levels were investigated following the exposure of S. schlegelii to low salinity (Fig. 2). The plasma osmolarity changed with decreasing salinity (p < 0.05); however, no significant differences were observed in the 100% group (p > 0.05). After 12 h of exposure to low-salinity conditions, the plasma osmolarity initially decreased gradually and then significantly according to the saline environment. After 24 h, plasma collected from the 60% and 80% groups exhibited their lowest osmolarity. Osmolarity levels tended to recover in the 60% group after 120 h.
Changes in the plasma osmolarity of S. schlegelii exposed to low-salinity conditions for 120 h (5 days). Different letters indicate significant differences between the same experimental groups for different exposure times (p < 0.05). Different numbers indicate significant differences between different experimental groups for the same exposure duration (p < 0.05). All values represent the mean ± standard error (n = 3). Control group, 34 psu (100% seawater); low-salinity groups, 20 psu (60% seawater) and 27 psu (80% seawater)
Oxidative stress responses to low salinity
The expression of SOD and CAT mRNA in the livers of S. schlegelii fish exposed to different salinity conditions was analyzed using qPCR (Fig. 3a). In the 60% and 80% exposure groups, SOD mRNA expression increased with increasing exposure time. At 72 h, SOD expression was significantly higher in the 60% than in the 80% group (p < 0.05). Hepatic CAT expression similarly increased with exposure time (Fig. 3b). CAT mRNA expression was significantly higher in the 60% group after 24 h of exposure than in the other groups (p < 0.05). However, after 120 h of exposure, CAT mRNA expression reduced significantly from the levels recorded at 72 h (p < 0.05).
Changes in mRNA expression of a superoxide dismutase (SOD) and b catalase (CAT) in the liver and c levels of H2O2 in the plasma of S. schlegelii exposed to low salinity for 120 h (5 days). Different letters indicate significant differences between the same experimental groups for different exposure times (p < 0.05). Different numbers indicate significant differences between different experimental groups for the same exposure duration (p < 0.05). All values represent the mean ± standard error (n = 3). Control group, 34 psu (100% seawater); low-salinity groups, 20 psu (60% seawater) and 27 psu (80% seawater)
We also investigated the H2O2 levels in blood plasma following the exposure of S. schlegelii to low-salinity conditions (Fig. 3c). The H2O2 levels increased with an increased time of exposure to low salinity in the 60% and 80% groups. The highest H2O2 increase was observed in the 60% group after 120 h (p < 0.05).
Intestinal microbiome diversity
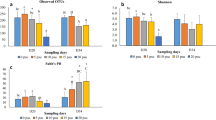
We evaluated the α-diversity to determine how the richness and diversity of the intestinal microbiome of S. schlegelii was altered by changes in water salinity (Fig. 4). The observed features and Shannon index showed significant differences between the intestinal microbiota of the 60% group and those of the other two groups (p < 0.05). Chao1 indices did not differ significantly among any of the groups (p > 0.05). To compare the entire structure of the intestinal microbial community, the β-diversity was evaluated using PCoA (Fig. 5). The control group exhibited a distinct microbiome cluster, whereas the 60% and 80% groups had overlapping microbiome clusters.
Effect of the exposure of S. schlegelii to low-salinity conditions on the β-diversity of intestinal microbiota using principal coordinate analysis (PCoA). Each point represents a single sample, with different colors representing different experimental groups. Control, 34 psu (100% seawater); low-salinity groups, 20 psu (60% seawater) and 27 psu (80% seawater)
Relative abundance of intestinal microbiota
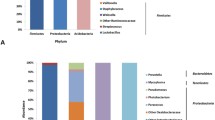
To determine whether exposure to a low-salinity environment causes dysbiosis in the intestinal of S. schlegelii, we determined the proportions of altered intestinal microbiota. Comparisons at the phylum level revealed a decrease in Proteobacteria in the low salinity groups with compared to the control group (100%, 66.4%; 80%, 21.8% decrease; 60%, 28.5% decrease). In contrast, Firmicutes showed a significant increase in the low-salinity exposure groups (100%, 11.2%; 80%, 16.5% increase; 60%, 18.7% increase compared 100%). At the class level, the relative abundance of Gammaproteobacteria decreased the most (100%, 57.1%; 80%, 16.0% decrease; 60%, 23.9% decrease) compared to that of the control group (Fig. 6b). Conversely, when compared to the control group, the two low-salinity groups exhibited increases in their proportions of Bacilli and Chlamydia classes in the intestines. At the genus level, the intestines of fish exposed to low-salinity conditions had decreased proportions of Enterovibrio (100%, 25.9%; 80%, 22.8% decrease; 60%, 23.7% decrease) and Photobacterium (p < 0.05) (Fig. 6c). Conversely, the abundance of Mycoplasma and Vibrio significantly increased in the low-salinity groups.
Relative abundance of intestinal microbiota in S. schlegelii exposed to low salinity according to the a phylum, b class, and c genus level. Each bar represents the average relative abundance of a bacterial taxon for a treatment group. Control group, 34 psu (100% seawater); low-salinity groups, 20 psu (60% seawater) and 27 psu (80% seawater)
Discussion
In this study, we analyzed the changes in the osmoregulatory capacity, oxidative stress response, and intestinal microbiota of S. schlegelii following exposure to a low-salinity environment, thereby assessing the impact and extent of environmental salinity changes on the physiological responses and intestinal microbial dysbiosis of this fish species. Fish exhibit varying degrees of tolerance to changes in environmental salinity (euryhaline or stenohaline), which affects their sensitivity to saline fluctuations. Therefore, it is necessary to investigate the physiological responses and adaptive mechanisms of fish that are increasingly exposed to hyposalinity induced by climate change. Specifically, changes in osmolality in low-salinity environments affect ion exchange and transportation within bodily fluids and NKA membrane transport proteins, and an antioxidant reaction represents the primary response to such environmental changes. Furthermore, changes in the intestinal microbial communities of fish influence physiological functions, including digestion and nutrient absorption. Therefore, analyzing the intestinal microbiota in fish exposed to low-salinity environments is crucially associated with research concerning fish health.
Exposure of S. schlegelii to low-salinity environments resulted in decreased expression and activity of NKA mRNA (which is involved in osmoregulation in the gills), along with a tendency toward decreased plasma osmolality; at 120 h post-exposure, plasma osmolality had decreased slightly but not significantly. In a study on euryhaline Epinephelus fuscoguttatus groups exposed to a low-salinity environment, plasma Na+ and K+ ions were shown to decrease, and osmolality significantly decreased with decreasing salinity (Liu et al. 2022). NKA activity also decreased in euryhaline teleosts such as Siganus rivulatus (rivulated rabbitfish) and Larimichthys crocea (large yellow croaker) when exposed to low-salinity environments but tended to recover with increased exposure time (Saoud et al. 2007; Han et al. 2022). However, in our study, exposure of S. schlegelii to salinity levels of 20 and 27 psu for 5 days did not result in the recovery of NKA activity, suggesting a relatively lower adaptability to salinity changes compared to euryhaline species such as S. rivulatus and L. crocea. The degree of tolerance to low salinity varies among species, and a low-salinity environment (20 and 27 psu) appears to have extreme effects on ion transport enzymes and changes in plasma osmolality in S. schlegelii.
When investigating two representative antioxidant enzymes, SOD and CAT, we found that their mRNA expression was significantly higher in S. schlegelii exposed to a 20-psu low-salinity environment than those exposed to 27 psu, starting from the 24th hour of exposure across a 5-day period. While the mRNA expression of SOD as well as H2O2 levels continued to increase with the passage of exposure time, CAT mRNA expression decreased slightly at 120 h post-exposure. The exposure of Notopterus chitala to salinity stress environments has similarly resulted in a significant increase in the SOD and CAT activity of this fish (Moniruzzaman et al. 2022); in groups exposed to the highest salinity stress environment (0–12 ppt), antioxidant enzyme activity decreased. Organisms, including fish, activate oxidative stress responses under salinity stress to remove harmful substances, such as ROS, from the body in response to environmental changes (Mozanzadeh et al. 2021). However, when the antioxidant levels exceed the regulatory capacity of the body, ROS may not be completely eliminated. In the present study, an increase in H2O2 levels suggests that S. schlegelii in the treatment groups experienced salinity stress, leading to the increased mRNA expression of antioxidant enzymes (SOD and CAT) to reduce oxidative stress in the body. Nonetheless, the continued increase in H2O2 levels also indicates that salinity conditions of 20 and 27 psu pose a high stress environment that challenges the effective removal of ROS generated within S. schlegelii.
The microbial communities in fish intestines promote intestinal health by enhancing the host immune system and generating beneficial metabolites through interactions with the host (Rastelli et al. 2019). In our study, the exposure of S. schlegelii to a low-salinity environment of 20 psu altered the species diversity and relative abundance of intestinal microbial communities. Beta diversity analysis further revealed that the low-salinity exposure group formed distinct clusters compared to the control group (p < 0.05). Thus, salt stress was confirmed to alter the diversity of intestinal microbial communities in this species. Schmidt et al. (2015) reported that in Poecilia sphenops exposed to salinity changes, Gammaproteobacteria exhibited the highest relative abundance. Conversely, our study found that Gammaproteobacteria (Proteobacteria phylum) were predominant in the control group (34 psu) but decreased in relative abundance as salinity exposure was decreased to 27 and 20 psu. In addition, even at the Genus level, Vibrio of the Gammaproteobacteria family increased the most when compared to the intestinal microbial community in the low-salt environment. Disease-causing Vibrio species such as Vibrio harveyi are known to penetrate the skin or intestines of fish and cause infection, which can weaken the immune system of fish and cause disorders (Chen et al. 2024). Photobacterium, Alivibrio, and Enterovibrio are mainly found in marine life and are microorganisms that reproduce and survive well in high-salt environments (Gregory and Boyd 2021). Enterovibrio calviensis is a halophilic bacterium that cannot survive without NaCl (Pascual et al. 2009).
The microorganism with increased relative ratios in the low-salt environment was Mycoplasma belonging to class Bacilli (Firmicutes phylum), which was predominant when compared to that in the control group. Mycoplasma causes respiratory diseases and infections in animals (García-Galán et al. 2021). The relative ratios of Chlamydia at the class level were also increased in the low-salt environmental laboratory, which is a pathogen that causes infections in humans and animals and is known to cause gill epithelial cystitis and respiratory infections in fish (Pawlikowska-Warych and Deptu ła 2016). In addition, the interaction between the intestinal microorganisms of fish and the immune system of the host organism affects the digestion and nutritional absorption of fish, thereby playing an important role in fish health and disease management (Pickard et al. 2017). Therefore, this study also confirmed that changes in the salt environment change the intestinal microbiota of S. schlegelii at the class and genus levels, and this change in microbial composition (intestinal imbalance) seems to affect the health and physiological response of fish. In other words, changes in the intestinal microbiota of S. schlegelii increase the relative proportion of pathogenic microorganisms, which affects the immune system of the host, ultimately affecting its survival.
In conclusion, our study confirmed the association between environmental changes in salinity and the physiological responses and intestinal microbiota of S. schlegelii, a target species for aquaculture. Low-salinity environments affected the osmoregulatory capacity of the gills and plasma of S. schlegelii and activated oxidative stress responses to saline stress. Low salinity levels increased ROS-mediated oxidative stress. Salinity changes in the environment also influenced their intestinal microbial community, leading to dysbiosis, which may increase the incidence of diseases. Saline fluctuations could therefore cause extensive economic damage to this species in the fishing and aquaculture sectors. Further microbial investigations aimed at elucidating the interaction between S. schlegelii and its intestinal microbiota could contribute to preventing disease and strengthening the immune system of this commercial species. Moreover, this study provides insights into coping mechanisms and strategies that fish can employ upon exposure to various unfavorable environmental conditions resulting from climate change.
Data availability
No datasets were generated or analysed during the current study.
References
Becker EA, Forney KA, Redfern JV, Barlow J, Jacox MG, Roberts JJ, Palacios DM (2019) Predicting cetacean abundance and distribution in a changing climate. Divers Distrib 25(4):626–643. https://doi.org/10.1111/ddi.12867
Bissett A, Brown MV, Siciliano SD, Thrall PH (2013) Microbial community responses to anthropogenically induced environmental change: towards a systems approach. Ecol Lett 16:128–139. https://doi.org/10.1111/ele.12109
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, ... Caporaso JG (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37(8), 852–857. https://doi.org/10.1038/s41587-019-0209-9
Chen M, Wan Q, Xu M, Chen Z, & Guo S (2024) Transcriptome analysis of host anti-Vibrio harveyi infection revealed the pathogenicity of V. harveyi to American eel (Anguilla rostrata). Mar Biotechnol 1–18. https://doi.org/10.1007/s10126-024-10298-9
Choi JK, Son YB, Park MS, Hwang DJ, Ahn JH, Park YG (2021) The applicability of the geostationary ocean color imager to the mapping of sea surface salinity in the East China Sea. Remote Sens 13(14):2676. https://doi.org/10.3390/rs13142676
García-Galán A, Seva J, Gómez-Martín Á, Ortega J, Rodríguez F, García-Muñoz Á, De la Fe C (2021) Importance and antimicrobial resistance of mycoplasma bovis in clinical respiratory disease in feedlot calves. Animals 11(5):1470. https://doi.org/10.3390/ani11051470
Gregory GJ, Boyd EF (2021) Stressed out: Bacterial response to high salinity using compatible solute biosynthesis and uptake systems, lessons from Vibrionaceae. Comput Struct Biotechnol J 19:1014–1027. https://doi.org/10.1016/j.csbj.2021.01.030
Han K, Zhou L, Zeng X, Zhang Z, Zou P, Huang W, Wang Y (2022) Effects of low-salinity acclimation on the Na+/K+ ATPase activity and expression of osmoregulatory-related genes in large yellow croaker (Larimichthys crocea). Aquac Rep 26:101326. https://doi.org/10.1016/j.aqrep.2022.101326
Hennersdorf P, Kleinertz S, Theisen S, Abdul-Aziz MA, Mrotzek G, Palm HW, Saluz HP (2016) Microbial diversity and parasitic load in tropical fish of different environmental conditions. PLoS ONE 11(3):e0151594. https://doi.org/10.1371/journal.pone.0151594
Hiroi J, McCormick SD (2012) New insights into gill ionocyte and ion transporter function in euryhaline and diadromous fish. Respir Physiol Neurobiol 184(3):257–268. https://doi.org/10.1016/j.resp.2012.07.019
Huang M, Yang X, Zhou Y, Ge J, Davis DA, Dong Y, Gao Q, Dong S (2021) Growth, serum biochemical parameters, salinity tolerance and antioxidant enzyme activity of rainbow trout (Oncorhynchus mykiss) in response to dietary taurine levels. Mar Life Sci Technol 1–14. https://doi.org/10.1007/s42995-020-00088-2
Jomova K, Raptova R, Alomar SY, Alwasel SH, Nepovimova E, Kuca K, Valko M (2023) Reactive oxygen species, toxicity, oxidative stress, and antioxidants: Chronic diseases and aging. Arch Toxicol 97(10):2499–2574. https://doi.org/10.1007/s00204-023-03562-9
King WL, Siboni N, Williams NL, Kahlke T, Nguyen KV, Jenkins C, Dove M, O’Connor WR, Seymour J, Labbate M (2019) Variability in the composition of Pacific oyster microbiomes across oyster families exhibiting different levels of susceptibility to OsHV-1 μvar disease. Front Microbiol 10:473. https://doi.org/10.3389/fmicb.2019.00473
Kiran KR, Deepika VB, Swathy PS, Prasad K, Kabekkodu SP, Murali TS, Satyamoorthy K, Muthusamy A (2020) ROS-dependent DNA damage and repair during germination of NaCl primed seeds. J Photochem Photobiol, B 213:112050. https://doi.org/10.1016/j.jphotobiol.2020.112050
Kültz D (2015) Physiological mechanisms used by fish to cope with salinity stress. J Exp Biol 218(12):1907–1914. https://doi.org/10.1242/jeb.118695
Kwak DH, Song YS, Choi YH, Kim KM, Jeong YH (2023) Influence of sluice gate operation on salinity stratification and hypoxia development in a brackish estuary dam. Regional Studies in Marine Science 57:102731. https://doi.org/10.1016/j.rsma.2022.102731
Lee SY, Lee HJ, Kim YK (2020) Comparative transcriptome profiling of selected osmotic regulatory proteins in the gill during seawater acclimation of chum salmon (Oncorhynchus keta) fry. Sci Rep 10(1):1987. https://doi.org/10.1038/s41598-020-58915-6
Li X, Shen Y, Bao Y, Wu Z, Yang B, Jiao L, Zhang C, Tocher DR, Zhou Q, Jin M (2022) Physiological responses and adaptive strategies to acute low-salinity environmental stress of the euryhaline marine fish black seabream (Acanthopagrus schlegelii). Aquaculture 554:738117. https://doi.org/10.1016/j.aquaculture.2022.738117
Liu LL, Luo M, Chen FX, Liu JY (2022) Effects of low salinity on osmoregulation, Na+/K+-ATPase activity and related gene expression of juvenile Epinephelus fuscoguttatus. Mar Fish 44(3):315–327
Moniruzzaman M, Mukherjee M, Kumar S, Chakraborty SB (2022) Effects of salinity stress on antioxidant status and inflammatory responses in females of a “Near Threatened” economically important fish species Notopterus chitala: a mechanistic approach. Environ Sci Pollut Res 29(49):75031–75042. https://doi.org/10.1007/s11356-022-21142-9
Mozanzadeh MT, Safari O, Oosooli R, Mehrjooyan S, Najafabadi MZ, Hoseini SJ, Saghavi H, Monem J (2021) The effect of salinity on growth performance, digestive and antioxidant enzymes, humoral immunity and stress indices in two euryhaline fish species: Yellowfin seabream (Acanthopagrus latus) and Asian seabass (Lates calcarifer). Aquaculture 534:736329. https://doi.org/10.1016/j.aquaculture.2020.736329
Oh SY, Kim CK, Jang YS, Choi HJ, Myoung JG (2014) Effect of salinity on survival, oxygen consumption and blood physiology of Korean rockfish Sebastes schlegelii. Ocean Polar Res 36(2):135–143. https://doi.org/10.4217/OPR.2014.36.2.135
Pascual J, Macian MC, Arahal DR, Garay E, Pujalte MJ (2009) Description of Enterovibrio nigricans sp. nov., reclassification of Vibrio calviensis as Enterovibrio calviensis comb. nov. and emended description of the genus Enterovibrio Thompson et al. 2002. Int J Syst Evol Microbiol 59(4):698–704. https://doi.org/10.1099/ijs.0.001990-0
Pawlikowska-Warych M, Deptuła W (2016) Characteristics of chlamydia-like organisms pathogenic to fish. J Appl Genet 57:135–141. https://doi.org/10.1007/s13353-015-0303-8
Pickard JM, Zeng MY, Caruso R, Núñez G (2017) Gut microbiota: role in pathogen colonization, immune responses, and inflammatory disease. Immunol Rev 279(1):70–89. https://doi.org/10.1111/imr.12567
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2012) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41(D1):D590–D596. https://doi.org/10.1093/nar/gks1219
Rastelli M, Cani PD, Knauf C (2019) The gut microbiome influences host endocrine functions. Endocr Rev 40(5): 1271–1284. https://doi.org/10.1210/er.2018-00280
Saoud IP, Kreydiyyeh S, Chalfoun A, Fakih M (2007) Influence of salinity on survival, growth, plasma osmolality and gill Na+–K+–ATPase activity in the rabbitfish Siganus rivulatus. J Exp Mar Biol Ecol 348(1–2):183–190. https://doi.org/10.1016/j.jembe.2007.05.005
Schmidt VT, Smith KF, Melvin DW, Amaral-Zettler LA (2015) Community assembly of a euryhaline fish microbiome during salinity acclimation. Mol Ecol 24(10):2537–2550. https://doi.org/10.1111/mec.13177
Sin E, Ahn IY, Park S, Kim T (2020) Effects of low pH and low salinity induced by meltwater inflow on the behavior and physical condition of the Antarctic limpet, Nacella concinna. J Mar Sci Eng 8(10):822. https://doi.org/10.3390/jmse8100822
Sullam KE, Essinger SD, Lozupone CA, O’Connor MP, Rosen GL, Knight ROB, Kil SS, Russell JA (2012) Environmental and ecological factors that shape the gut bacterial communities of fish: a meta-analysis. Mol Ecol 21(13):3363–3378. https://doi.org/10.1111/j.1365-294X.2012.05552.x
Tian L, Tan P, Yang L, Zhu W, Xu D (2020) Effects of salinity on the growth, plasma ion concentrations, osmoregulation, non-specific immunity, and intestinal microbiota of the yellow drum (Nibea albiflora). Aquaculture 528:735470. https://doi.org/10.1016/j.aquaculture.2020.735470
Velotta JP, McCormick SD, Whitehead A, Durso CS, Schultz ET (2022) Repeated genetic targets of natural selection underlying adaptation of fishes to changing salinity. Integr Comp Biol 62(2):357–375. https://doi.org/10.1093/icb/icac072
Wang AR, Ran C, Ringø E, Zhou ZG (2018) Progress in fish gastrointestinal microbiota research. Rev Aquac 10(3):626–640. https://doi.org/10.1111/raq.12191
Zhao R, Symonds JE, Walker SP, Steiner K, Carter CG, Bowman JP, Nowak BF (2020) Salinity and fish age affect the gut microbiota of farmed Chinook salmon (Oncorhynchus tshawytscha). Aquaculture 528:735539. https://doi.org/10.1016/j.aquaculture.2020.735539
Funding
This work was supported by Korea Institute of Marine Science and Technology Promotion (KIMST) funded by the Ministry of Oceans and Fisheries (20220559).
Author information
Authors and Affiliations
Contributions
Jin A Kim: Conceptualization, Data curation, Writing—Original Draft. Young-Su Park: Methodology, Formal analysis. Jun-Hwan Kim: Validation, Writing – Review & Editing. Cheol Young Choi: Writing – Review & Editing, Supervision. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Declarations
Approval for animal research was obtained from the Institutional Animal Care and Use Committee of Korea Maritime and Ocean University (approved no # KMOU IACUC 2023–05).
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kim, J.A., Park, YS., Kim, JH. et al. Hyposalinity elicits physiological responses and alters intestinal microbiota in Korean rockfish Sebastes schlegelii. Fish Physiol Biochem (2024). https://doi.org/10.1007/s10695-024-01387-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10695-024-01387-6