Abstract
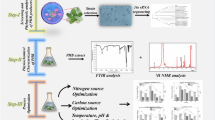
Polymers are currently used in the industry as raw material, yet they are rapidly eliminated and largely contaminate the environment. To address this issue, there is a special interest in biodegradable polymers, namely, polyhydroxyalkanoates (PHAs), produced by microorganisms. This study identifies PHA-producing bacteria from two industrial wastewaters of Manizales, Colombia. The samples were cultured in mineral salt medium with glucose as the carbon source in the presence of Nile red stain. The fluorescent colonies were independently transferred to another medium and assessed through fluorescence microscopy with Nile blue stain. The fluorescent strains under Nile blue staining were purified in Nutrient Agar, and their morphological and microbiological characteristics were determined. The bacteria positive for red-orange fluorescence were purified in Nutrient Agar medium, and molecular analyses were performed by PCR amplification of a 650-bp fragment of the 16S ribosomal DNA gene. The bacteria were also assessed in terms of PHA production. We confirmed the identity of 12 out of 14 PHA-positive strains, which belonged to the following genera: Bacillus, Lactococcus, Citrobacter, Enterobacter, and Acinetobacter. Five of the isolates (Enterobacter cloacae, Enterobacter sp., Enterobacter ludwigii, Bacillus thuringiensis, and Bacillus safensis) are promising strains for PHA production, with production values ranging from 0.360 to 0.9960 g/L. Bacteria that produce more than 0.3 g/L are considered useful for the industrial manufacture of bioplastic. We recommend performing large-scale studies on these strains to assess their use for the industrial production of biopolymers, allowing to generate high-impact bioconversion processes of industrial interest.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The expanding population, urbanization, and industrialization have led to the production of an excessive amount of wastewater that generates high contamination loads (Bengtsson et al. 2008). Industrial wastewater can produce waste material that is used by microorganisms (Bitton 2005). In particular, wastewater can be a culture medium that allows the growth of certain microbial populations. These microorganisms can interact and perform metabolic reactions that degrade the organic matter and remove nutrients from the wastewater (Bravo et al. 2005).
Industries rely on wastewater treatment plants to remove organic loads and deposit solid wastes through aerobic and anaerobic processes (Harding et al. 2007). These biological processes produce mixed microbial consortia (MMC), which are a potential raw material for biopolymer production (Reddy and Mohan 2012). These consortia contain microorganisms such as bacteria, yeast, and fungi that are capable of synthesizing biopolymers in the presence of an excess carbon source and limited nitrogen or phosphorous in the growth medium (Kumar et al. 2016).
Biopolymers, commonly known as polyhydroxyalkanoates (PHAs), were discovered in the early twentieth century in Bacillus megaterium (Keshavarz and Roy 2010). Over 300 species of microorganisms accumulate PHAs, among them halophiles can reach yields above 70% (Guzmán et al. 2017). PHAs are a group of biodegradable and biocompatible polyesters of natural origin that are synthesized by a wide range of Gram-negative and Gram-positive bacteria as carbon reserves (Neira and Pardo 2010). Some of these bacteria are industrially useful due to their efficient substrate transformation processes and the final concentration of biopolymer inside the cells (Barbosa et al. 2005). Gram-negative bacteria that efficiently produce PHAs are Cupriavidus necator (formally Alcaligenes eutrophus), Alcaligenes latus, Pseudomona putida, Pseudomona oleovorans, and Azotobacter vinelandii, as well as the recombinant strain of Escherichia coli that contains the PHA biosynthesis operon from C. necator (Barbosa et al. 2005). Similarly, some Gram-positive bacteria are also PHA producers, including several species of Bacillus (e.g., Bacillus megaterium and Bacillus cereus). In addition, fungi of the genus Streptomyces (Actinomycetes) are also known to produce PHAs (Yilmaz and Beyatli 2005; Franco et al. 2009; Reddy and Mohan 2012).
Due to their lipidic nature, the detection and isolation of PHA-producing bacteria are based on the use of lipophilic stains such as Sudan black (Schlegel et al. 1970), Nile blue (Ostle and Holt 1982), fluorescent oxazone, and Nile red (Spiekermann et al. 1999). However, the detection of PHA-producing bacteria is supported by confirmatory PCR-based molecular assays to determine the genus and species of the bacteria as well as the presence of one or several PHA biosynthesis genes (Solaiman et al. 2000; Ciesielski et al. 2006). In this regard, the 16S rDNA gene provides high-grade resolution for the taxonomical determination; therefore, if two strains share less than a 97% identity in their 16S rDNA sequence it can be inferred that these do not belong to the same species (Rosselló-Mora and Amann 2001).
This study aimed to characterize PHA-producing bacteria using microbiological and molecular techniques to search for native strains useful for the industrial production of biopolymers. The identification of these bacteria can promote the use of an economical substrate derived from wastewater treatment plants, which are a primary source of activated sludge containing polymers for bioplastic production. These microorganisms can be used as an alternative for wastewater treatment since industrial wastes can represent settings for biological degradation and the production of bioplastics, therefore, promoting biotechnological development in the region.
Materials and methods
Study area
The study area comprised wastewater treatment plants (WWTP) from two food-processing factories, namely, Super de Alimentos (5° 36′ 25″ N, 75° 45′ and 62″ N 75° 45′ 63″ W) and Industrias Lácteas Normandy (5° 34′ 97″ N and 75° 45′ 39″ W), located in the city of Manizales, Caldas, Colombia. We monitored three sites at each treatment plant: (a) entrance to the wastewater treatment plant (percolator filter), (b) sedimentation tank (activated sludge chamber), and (c) exit tank (decanting chamber) (Fig. 1).
Sample collection and selection of isolates with PHA-producing capacity
Sample collection was conducted on a normal production day at each wastewater treatment plant. The samples were collected at 1-h intervals during 4 continuous hours. The samples were taken at a depth of 1.2 m and stored in clean and clear wide-mouth 250-mL glass jars, carefully closed and labeled, according to the recommendation of the IDEAM (Guía para el Monitoreo de Vertimientos, Aguas Superficiales y Subterráneas 2003). The samples were taken to the biotechnology laboratory of Tecnoparque SENA—Manizales, Colombia, for processing and analyses.
The samples were cultured by spreading 1 mL of the sample on Nutrient Agar medium. After, the bacterial morphotypes were isolated by the streak plate method on a solid mineral salt medium (MSM), pH 7.0, supplemented with 2% glucose, 0.2% yeast extract, and 1 mL/L 0.1% Nile red stain in acetone solution to detect PHA-producing colonies under an ultraviolet transilluminator at a wavelength of 340 nm (Spiekermann et al. 1999).
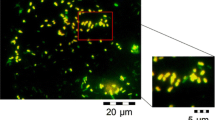
The cultures were incubated at 30 °C and monitored every 24 h under ultraviolet light for 72 h. After, the fluorescent colonies under Nile red staining were further isolated through streak-plating on Nutrient Agar medium. A confirmatory test was performed using Nile blue stain, according to Ostle and Holt (1982), using a Nikon CI-S INTENSILIGHT fluorescent microscope at 450 nm and × 100. The red-orange fluorescent bacteria indicated a possible production of PHAs, and these were purified in Nutrient Agar (Fernández et al. 2005).
Morphological and molecular identification of PHA-producing strains
Each isolate presumably capable of producing biopolymers was assessed by morphology based on the macroscopic and microscopic characteristics proposed by Bou et al. (2011). Finally, the isolates were Gram-stained and stored in cryovials with 15% glycerol at − 80 °C (Capuccino and Sherman 2007).
For the molecular analyses, DNA was extracted with the UltraCleanTM Microbial DNA Isolation kit (Vidal et al. 2007) and an ~ 650-bp fragment of the V1–V6 regions of the 16S rDNA gene was amplified by PCR using primers 27F 5′AGAGTTTGATCMTGGCTCAG 3′ and 104R 5′CGGTGTGTACAAGACCC 3′ (Kuske et al. 1997). The amplicons were visualized through horizontal electrophoresis on 1% agarose gels with 1X TBE pH 8.0 at 110 v/50 mA. SYBR Safe® dye was used to visualize the DNA bands using a GelDoc-It®2 310 Imager (UVP) photodocumentor device. The PCR products were purified using the QIAquick PCR purification (Qiagen®) kit, following the manufacturer’s recommendations, and sequenced at Macrogen (South Korea). The sequences were viewed and edited using Geneious Trial v8.14 (Drummond et al. 2009) and BioEdit 7.2.6. The sequences were searched against the public databases (i.e., GenBank) with MegaBLAST to retrieve similar sequences.
Assessment of polyhydroxyalkanoate-producing strains
We assessed the capacity of the isolates to produce biopolymers using a modified synthetic culture medium proposed by Fernández et al. (2005), enriched with glucose to facilitate the measurement of biopolymer production. For the assessment, we inoculated 9 × 108 cells mL/L (McFarland nephelometric scale) of the isolates in the synthetic substrate in batch bioreactors. We recovered the precipitate at 24, 48, and 72 h of incubation by centrifugation and resuspension in 5% sodium hypochlorite and 10 mM EDTA. After, we incubated the suspension at 60 °C for 1.5 h, then, we centrifuged and washed the pellet with distilled water. The precipitate was resuspended in acetone, centrifuged, and the supernatant was removed. Then, we diluted the pellet in cold methanol, centrifuged once more, recovered the precipitate, and left it to dry at room temperature for 12 h. Finally, the resulting biomass was weighed, and this value was used to determine the total PHA content per dry mass based on the volume and dry weight of the sample (Fernández et al. 2005). As positive control we included Bacillus megaterium (cepa ATCC 14581), and as negative control we used the modified synthetic culture medium proposed by Fernández et al. (2005).
We determined which strains from the dairy (Industrias Lácteas Normandy) or candy (Super de Alimentos) factories showed the highest biopolymer production for 24, 48, and 72 h by performing a bootstrap test (1000 replicas) according to Crawley (2005). In addition, we also used bootstrapping (1000 replicas) to compare the production of biopolymers between the dairy and candy factories. These bootstrap analyses were conducted due to the small sample sizes (diary n = 8 and candy n = 6) and non-normal distribution of the data. Finally, we compared PHA median production of strains by Kruskal-Wallis test (H). The statistical analyses were performed in R version 3.3.3 (R Core Team 2017).
Results
Selection of the isolates with PHA-producing capacity
We identified 86 bacterial morphotypes growing on Nutrient Agar medium. These morphotypes grouped 106 strains; among them 31 were fluorescent under Nile red staining (Table 1). The confirmatory test (Nile blue staining) allowed identifying 14 strains capable of producing PHAs (Table 1).
Morphological and molecular identification of PHA-producing strains
The 14 strains screened for PHA-production displayed macroscopic features including spherical, circular, and irregular shapes; creamy textures; and smooth and rough surfaces, among others. Microscopically, we observed bacilli and cocci, including 4 Gram-positive and 10 Gram-negative strains.
The amplification of the 650-bp fragment of the 16S rDNA gene confirmed the identity of 12 strains (Table 1). The 16S partial sequences showed 97% and 99% identity to reported sequences for Bacillus, Lactococcus, Citrobacter, Enterobacter, and Acinetobacter (Table 2). The GenBank accession numbers for the 16 rDNA nucleotide sequences obtained in this study are [MN013897, MN013898, MN013899, MN013902, MN013903, MN013904, MN013905, MN013906, MN013907, MN013908, and MN013909].
Assessment of the PHA-producing strains
We determined the identity of 12 PHA-positive bacteria, including 8 strains isolated from Industrias Lácteas Normandy, namely Bacillus thuringiensis that showed the highest PHA production (960.67 mg/L), followed by Bacillus safensis (330.33 mg/L) (Fig. 2). These two strains showed a higher biopolymer production than the other six strains at 24, 48, and 72 h (P < 0.025). The PHA production differ among trains in Industrias Lácteas Normandy (H = 18.01; P = 0.012). Furthermore, 6 of the 12 PHA-positive strains were obtained from Super de Alimentos. Of these, we identified four strains, namely, Enterobacter cloacae and Enterobacter ludwigii, which showed a high PHA production capacity (730.33 mg/L and 800 mg/L, respectively) (Fig. 3). Enterobacter cloacae (Manizales) showed a higher biopolymer production at 24 h (P < 0.025), while E. ludwigii (Manizales) showed a higher production at 48 and 72 h (P < 0.025). The PHA production differ among trains in Super de Alimentos (H = 14.30, P = 0.014). Finally, no significant differences in PHA-production were found between the dairy and candy industries at 24 h (P = 0.516), 48 h (P = 0.515), or 72 h (P = 0.487). However, future research involving these PHA-positive bacteria must assess the purity of the PHA, as described in Amaro et al. (2019).
Discussion
We identified PHA-producing bacterial strains isolated from wastewater treatment plants of dairy (Industrias Lácteas Normandy) and candy (Industria Super de Alimentos) factories in Manizales, Colombia. The data provide new evidence that supports the PHA-producing bacterial isolates in wastewater treatment plants and the use of these beneficial microorganisms for the industrial production of bioplastic.
The morphological and molecular identifications of the 12 PHA-producing isolates showed that these belonged to the genera Bacillus, Lactococcus, Citrobacter, Enterobacter, and Acinetobacter, which have been reported to produce PHAs (Cardona-Echavarria et al. 2013; González-García et al. 2013; Malagón-micán et al. 2017). Cardona-Echavarria et al. (2013) reported 38 PHA-producing bacterial strains isolated from the wastewaters of the dairy industry. González-García et al. (2013) found five Bacillus strains and one Enterobacter strain in wastes derived from the industrial production of saccharose and molasses, which showed a high capacity for PHA production.
Among the bacterial strains reported here, we found that B. thuringiensis and B. safensis (from Industrias Lácteas Normandy) showed the highest levels of production for this factory at a laboratory scale (960.67 mg/L and 330.33 mg/L, respectively). Otero-Ramírez and Fernández (2013) reported production values of 0.3 to 2 g/L for Bacillus strains. Similarly, Porwal et al. (2008) reported a production of 0.19 g/L in Bacillus grown on a medium supplemented with glucose as a carbon source, while Tam-Dogan and Sidal (2011) found a production value of 0.01 g/L for Bacillus using manitol as a carbon source.
The strains of Enterobacter cloacae and E. ludwigii from wastewaters of Super de Alimentos showed the highest production levels for this factory (730.33 and 800 mg/L, respectively). For E. cloacae, González-García et al. (2013) report a PHA production equal to over 94% of its dry weight and Malagón-Micán et al. (2017) found PHA production values close to 0.6 g/L, similar to those found in this study. Regarding E. ludwigii, we found that this bacterium had the highest polymer production capacity (800 mg/L). Mora et al. (2017) found that E. ludwigii, isolated from sub-products of sugarcane, is capable of producing PHA from various carbon sources. Finally, the genus Enterobacter, found here in wastewater from the candy factory, has been widely studied (Koller et al. 2012) and isolated from diverse agro-industrial wastes (Naheed et al. 2012).
Fernández et al. (2005) report that bacteria capable of producing more than 0.3 g/L of PHAs are considered promising for the industrial manufacture of bioplastic. Of the 14 strains found here, 5 are promising for PHA production (Enterobacter cloacae, Enterobacter sp., Enterobacter ludwigii, Bacillus thuringiensis, and Bacillus safensis), showing values between 0.360 and 0.9960 g/L. Based on our findings, we recommend conducting large-scale assays and evaluating the industrial production of biopolymers using these strains. This will allow generating high-impact bioconversion processes to allow recovering industrially useful metabolites, such as PHAs, from contaminating sub-products. Although solvent-based methods are economical and show high PHA yields, we must assess and implement more environmentally friendly extraction methods, such as the use of enzymes and detergents that remove cellular components while leaving PHAs intact. These strategies are currently more suitable for the environment but normally result in lower PHA yields (Suriyamongkol et al. 2007; Amaro et al. 2019).
References
Amaro, T. M., Rosa, D., Comi, G., & Iacumin, L. (2019). Prospects for the use of whey for polyhydroxyalkanoate (PHA) production. Frontiers in Microbiology, 10, 1–12.
Barbosa, M., Moreno, N., Espinosa, A., & Malagón, D. (2005). Producción de poli-B-hidroxibutirato (PHB) por Ralstonia eutropha ATCC 17697. Universitas Scientiarum, 10, 45–54.
Bengtsson, O., Jeppsson, M., Sonderegger, M., Parachin, N. S., Sauer, U., Hahn-Hägerdal, B., & Gorwa-Grauslund, M. F. (2008). Identificación de rasgos comunes en Saccharomyces cerevisiae con crecimiento mejorado de xilosa para ingeniería metabólica inversa. Levadura, 25(11), 835–847.
Bitton, G. (2005). Wastewater microbiology. 3rd Edition. Willey. WILEY Series in Ecological and Applied Microbiology.
Bou, G., Fernández-Olmos, A., García, C., Sáez-Nieto, J., & Valdezate, S. (2011). Métodos de Identificación Bacteriana En El Laboratorio de Microbiología. Enfermedades Infecciosas y Microbiología Clínica., 29, 601–608.
Bravo, M., Moreno, A., Henández, C., Yeomans, J., & Okumoto, S. (2005). Implementación y monitoreo de la etapa inicial del sistema de tratamiento de aguas residuales del laboratorio de procesamiento de alimentos de la Universidad EARTH. Tierra Tropical, 1, 89–97.
Capuccino, J., & Sherman, N. (2007). Microbiology: a laboratory manual. New York: Benjamin Cummings 544 p.
Cardona-Echavarria, A., Mora, A., & Marín, N. (2013). Identificación Molecular de Bacterias Productoras de Polihidroxialcanoatos en Subproductos de Lácteos y Caña de Azúcar. Revista Facultad Nacional de Agronomía, 66(2).
Ciesielski, S., Cydzik-Kwiatkowska, A., Pokoj, T., & Klimiuk, E. (2006). Molecular detection and diversity of medium chain length polyhydroxyalkanoates producing bacteria enriched from activated sludge. Journal of Applied Microbiology, 101(1), 190–199.
Crawley, M. J. (2005). Statistics: an introduction using R. Chichester: Wiley 327 pp.
Drummond, A.J., Ashton, B., Cheung, M., Heled, J., Kearse, M., Moir, R., Stones, H.S., Thierer, T., & Wilson, A. (2009). Geneious v.8.14. http://www.geneious.com. Accessed 13 Feb 2017.
Fernández, D., Rodríguez E., Bassas, M., Viñas-Solanas, A.M., Liorens, J., Marquéz, A.M., & Manresa, A. (2005). Agro-industrial oily wastes as substrates for PHA production by the new strain Pseudomonas aeruginosa NCIB 40045: Effect of culture conditions. Biochem. Eng. J., 26, 159–167.
Franco, M., Gómez, D., Medina, N., & Rendón, M. (2009). Polihidroxialcanoatos en actinomicetos nativos de suelos colombianos. Revista Peruana de Biología, 16, 115–118.
González-García, Y., Meza-contreras, J., González-Reynoso, O., & Córdova-López, J. (2013). Síntesis y biodegradación de polihidroxialcanoatos: plásticos de origen microbiano. Rev. Int. Contam. Ambie., 29(1), 77–115.
Guzmán, C., Hurtado, A., Carreño, C., & Casos, I. (2017). Production of polyhydroxyalkanoates by native halophilic bacteriausing Solanum tuberosum L. shell starch. Scientia Agropecuaria, 8(2), 109–118.
Harding, K., Dennis, J., Von Blottnitz, H., & Harrison, S. (2007). Environmental analysis of plastic production processes: Comparing petroleum—based polypropylene and polyethylene with biologically—based poly-β-hydroxybutyric acid using life cycle analysis. Journal of Biotechnology, 130, 57–66.
Keshavarz, T., & Roy, P. (2010). Polyhydroxyalkanoates: bioplastics with a green agenda. Current Opinion in Microbiology, 13(3), 321–326.
Koller, M., Salerno, A., Muhr, A., Reiterer, A., Chiellini, E., Casella, S., Horvat, P., & Brauneg, G. (2012). Chapter 2: Whey lactose as a raw material for microbial production of biodegradable polyester. In H. E.-D. M. Saleh (Ed.), Polyesters (pp. 19–60). Rijeka: InTech.
Kumar, M., Gupta, A., & Thakur, I. S. (2016). Carbon dioxide sequestration by chemo- lithotrophic oleaginous bacteria for production and optimization of poly- hydroxyalkanoate. Bioresource Technology, 213, 249–256.
Kuske, C. R., Bams, S. M., & Busch, J. D. (1997). Diverse uncultivated bacterial groups from soils of the arid Southwestern United States that are present in many geographic regions. Applied and Environmental Microbiology, 63(9), 3614–3621.
Malagón-Micán, M., López-López, S., & Martínez-Hernández, A. (2017). Síntesis de bioplásticos a partir de microorganismos. Fundación Universidad de América, Semilleros Formación Investigativa, 3(1), 127–135.
Mora, A., Gómez, J., Salazar, A., Sánchez, S., Cardona, A., Correa, G., Yepes, M., & Marín, M. (2017). Producción y caracterización de polihidroxialcanos a partir de residuos agroindustriales. Revista Colombiana de Materiales, 1, 5–7.
Naheed, N., Jamil, N., Hasnain, S., & Abbas, G. (2012). Biosynthesis of polyhydroxybutyrate in Enterobacter sp. SEL2 and Enterobacteriaceae Bacterium sp. PFW1 using sugar cane molasses as media. African Journal of Biotechnology, 11(16), 3321–3332.
Neira, A., & Pardo, L. (2010). Trabajo revisión bibliográfica biopolímeros: Poliésteres. Universidad pedagógica y tecnológica de Colombia. Recuperado de http://poliesteres.files.wordpress.com/2010/04/ojotrabajo-final-poliesteres.pdf . Consultado (20-05-2017).
Ostle, A., & Holt, J. (1982). Nile Blue A as a fluorescent stain for poly beta hydroxybutyric acid. Applied and Environmental Microbiology, 44(1), 238–241.
Otero-Ramírez, I., & Fernández, P. (2013). Bioprospección de bacterias productoras de polihidroxialcanoatos (PHAs) en el departamento de Nariño. Biotecnología en el Sector Agropecuario y Agroindustrial, 2(12), 60–71.
Porwal, S., Kumar, T., Lal, S., Rani, A., Kumar, S., Cheema, S., Purohit, H., Sharma, R., Kumar, S., Patel, S., & Kalia, V. (2008). Hydrogen and polyhydroxybutyrate producing abilities of microbes from diverse habitats by dark fermentative process. Bioresource Technology, 99, 5444–5451.
R Core Team (2017). R: A language and environment for statistical computing. R Foundation for statistical computing, Vienna, Austria. URL https://www.R-project.org/.
Reddy, M. V., & Mohan, S. V. (2012). Effect of substrate load and nutrients concentration on the polyhydroxyalkanoates (PHA) production using mixed consortia through waste-water treatment. Bioresource Technology, 114, 573–582.
Rossello-Mora, R., & Aman, R. (2001). The species concept for prokaryotes. FEMS Microbiology Reviews, 25, 39–67.
Schlegel, H., Lafferty, R., & Krauss, I. (1970). The isolation of mutants not accumulating poly-β-hydroxybutyric acid. Archiv für Mikrobiologie, 70, 283–294.
Solaiman, D., Ashby, R., & Foglia, A. (2000). Rapid and specific identification of medium-chain-length polyhydroxyalkanoate synthase gene by polymerase chain reaction. Applied Microbiology and Biotechnology, 53, 690–694.
Spiekermann, P., Rehm, B., Kalscheuer, R., Baumeister, D., & Steinbuchel, A. (1999). A sensitive, viable colony staining method using Nile red for direct screening of bacteria that accumulate poly lyhydroxyalkanoic acids and other lipid storage compounds. Archives of Microbiology, 171(2), 73–80.
Suriyamongkol, P., Weselake, R., Narine, S., Moloney, M., & Shah, S. (2007). Biotechnological approaches for the production of polyhydroxyalkanoates in microorganisms and plants—a review. Biotechnology Advances, 2, 148–175.
Tam-Dogan, N., & Sidal, U. (2011). Investigation of poly-hydroxybutyrate (PHB) production by Bacillus subtilisATCC 6633 under different con- ditions. KafkasUniv Vet FakDerg, 17, 173–176.
Vidal, N., Delmas, A. S., Patrick, D., Cruaud, C., Couloux, A., & Hedges, B. (2007). The phylogeny and classification of Caenophidian snakes inferred from seven nuclear protein-coding genes. CR Biologies, 330, 182–187.
Yilmaz, M., & Beyatli, Y. (2005). Poly-β-hydroxybutyrate (PHB) production by a Bacillus cereus M5 strain in sugar beet molasses. Zuck- erindustrie, 130, 109–112.
Funding
This work was financed by Servicio Nacional de Aprendizaje - SENA, Colombia, through grant number SENNOVA 016-2015 and Vicerrectoría de Investigaciones y Postgrados from University of Caldas through grant number 0923517—Joint Call for Promotion of Applied Research and Technological Development, between the Universidad de Caldas, Universidad Nacional de Colombia, Manizales, Colombia.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Giraldo-Montoya, J.M., Castaño-Villa, G.J. & Rivera-Páez, F.A. Bacteria from industrial waste: potential producers of polyhydroxyalkanoates (PHAs) in Manizales, Colombia. Environ Monit Assess 192, 480 (2020). https://doi.org/10.1007/s10661-020-08461-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10661-020-08461-5