Abstract
The European wildcat (Felis silvestris silvestris) underwent a severe decline across Europe in the early twentieth century. Remaining populations are often very small and isolated, though there are indications that wildcat populations are currently expanding their range. However, linear landscape elements such as rivers and roads are thought to present barriers to dispersal, inhibiting gene flow and, thus, affecting the recolonization process. In this study, we investigated the fine-scale genetic structure of wildcats in the Upper Rhine Valley. We specifically analysed wildcats on both sides of the Rhine River by genotyping 55 individual wildcats, using 20 microsatellite loci. Genetic differentiation was weak and positive spatial autocorrelation was found up to a distance of 10 km (females: 5 km, males: 10 km) indicating substantial gene flow among sampling sites. High levels of gene flow, even across the Rhine River, indicated that the water body itself does not necessarily have a strong barrier effect, which is in contrast to other studies. Our findings could best be explained by the populations’ history, a local extinction east of the River Rhine and a current ongoing population expansion. Our study highlights that potential barriers, such as rivers, may have different effects in different local wildcat populations and that the history of the populations is important to interpret genetic results. As many wildcats still occur in isolated and patchy forest fragments, maintaining connectivity between populations is crucial to ensure their viability in the long term.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Today’s major threat to wildlife is often the loss and fragmentation of habitat, leading to small and isolated habitat patches (Wilcove et al. 1986; Brooks et al. 2002). Limited connectivity between such habitat patches additionally contributes to the negative effects of habitat fragmentation. Potential barriers for species are often linear landscape elements, such as rivers or roads (Balkenhol and Waits 2009; Holderegger and Di Giulio 2010). Although natural structures, like rivers, can impose a direct barrier to some species (Cozzi and Broekhuis 2013), their effect is often amplified through extensive use by humans. Additionally, anthropogenic structures, like roads or railways, often hinder landscape connectivity. Barriers have, for instance, shown to alter ranging behaviour and prevent migration, dispersal and gene flow of animals (Balkenhol and Waits 2009; Holderegger and Di Giulio 2010). Because of their relatively large spatial requirements, low densities and, partially, due to direct persecution from humans mammalian carnivores are regarded as particularly sensitive to landscape fragmentation (Noss et al. 1996; Woodroffe and Ginsberg 1998). For example, the range expansion of wolves (Canis lupus) was shown to be constrained by a river in Spain (Blanco et al. 2005). Negative effects of landscape barriers are even higher for habitat specialists (Devictor et al. 2008) which depend on specific habitat structures like forested habitats. One such specialist is the endangered European wildcat (Felis silvestris silvestris, Schreber 1777).
Initially, the distribution range of the European wildcat spanned the whole of Europe, except Turkey, Caucasus and Scandinavia (Stahl and Artois 1991). As a result of large-scale hunting in the early twentieth century and human-mediated habitat loss and fragmentation, the European wildcat has undergone a severe reduction in both population numbers and sizes, resulting in a fragmented distribution (Piechoki 1990; Stahl and Artois 1991; Nowell and Jackson 1996a). In Western Europe, the remaining populations are mostly small and fairly isolated (Eckert 2003), while the distribution in Eastern Europe appears to be less fragmented (Piechoki 1990; Stahl and Artois 1991; Nowell and Jackson 1996). Interestingly, the species is in the process of recovering in many areas, which can be attributed to the legal protection of the species under the Bern Convention (Appendix II) and the European Habitat Directive 92/43/EEC (Annex IV). Many of the former agents causing the reduction of wildcat populations have now been reduced or partially eliminated, e.g. changes in forest management and agriculture, the reduction of rodenticides in agriculture and changes in game law. As a result, wildcat populations could recover in some parts of Europe (Stahl and Artois 1991) and are currently expanding their range (Say et al. 2012; Steyer et al. 2012). However, habitat fragmentation is still persistent and the expansion and re-colonization processes might be hindered not only by unsuitable habitat, but also by dispersal barriers between habitat patches. For example, Hartmann et al. (2013) showed that both the Rhine River and a highway in the Taunus and Hunsrueck mountain ranges in Germany were hindering gene flow between wildcat populations.
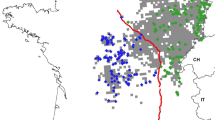
Here we investigate a potentially fragmented wildcat population in the border triangle of Germany, France and Switzerland, intersected by the Rhine River (see Fig. 1). In France, a continuous, recently expanding distribution is known in the woodland forest of the Vosges, with a sporadic presence in the lowland of the Alsace plain (Say et al. 2012). Wildcats also exist in neighboring regions in the North, in the Palatine Forest in Germany (Eckert et al. 2010) and in the South, in the Swiss Jura mountains (Nussberger et al. 2013). In all these regions, the presence of wildcats has been known for years. In contrast, east of the Rhine River, in Baden Wuerttemberg, Germany, the European wildcat was considered to be extinct (Piechoki 1990). However, in 2006, two carcasses confirmed the presence of the wildcat in Baden Wuerttemberg (Herdtfelder et al. 2007) and, since then, a small wildcat population has been confirmed in the Upper Rhine Valley (Streif et al. 2012).
a Location of the study area. b sample location of wildcats; all sampling locations are marked (empty circles), samples analysed (black filled circles). c selected part of the study area highlighting infrastructure: mosaic of forest patches, residential area, roads and the Rhine River within the study area
In our study, we focus on this newly discovered population of wildcats, investigating the fine scale genetic structure and gene flow in this and neighboring areas. Microsatellite markers are used to genotype individuals from those areas, estimating genetic diversity and gene flow across populations. We specifically test if the Rhine River acts as a potential barrier and try to identify from which area the newly discovered population originated. Two scenarios are possible: In scenario I the population east of the Rhine River has not been extinct in the twentieth century but could have survived in small refuges (e.g. Stromberg-Heuchelberg area) (Herrmann and Vogel 2005). In this region wildcats have probably a long time been undetected or confounded with free-roaming domestic cats. In recent years, the small population in the refuges could expand and has now established a growing population in the German Upper Rhine Valley. As a consequence, wildcats on the east and west side of the Rhine River would be differentiated and display signs of a genetic bottleneck. In Scenario II, we assume that the wildcat population east of the Rhine River got extinct in the twentieth century. In recent years, however, individuals could disperse to new areas and the growing population in the Vosges expanded their range across the Rhine River. In this case, we expect not to find a barrier effect of the Rhine River and no bottleneck of the small population east of the Rhine River.
Materials and methods
Study area
The study area is located in the border triangle of Switzerland, France and Germany, in the Upper Rhine Valley (Fig. 1). This valley (approximately 200 m a.s.l.) is 300 km in length and, on average, 40 km in width. It encompasses the lowlands from Basel-Landschaft and Swiss Jura (Switzerland) in the South, up to the Palatine Forest (Germany) in the North. The western border is the mountain range of the Vosges (France) and the eastern border is the low mountain range of the Black Forest (Germany). With a mean temperature of 18 °C in July and just 0 °C in January, the valley exhibits a relatively mild regional climate with a mean precipitation of 600–700 mm per year. The valley is divided by the Rhine River (Fig. 1). Within the study area, the river is between 83 and 240 m wide and has an almost constant discharge of 100 l/s. By the middle of the last century, the Upper Rhine was partly canalized between France and Germany to allow navigation between Basel and Strasbourg and to produce hydroelectricity. Most of the original wetland forest is now gone except for a forest line approximately 3 km wide along the river from Basel to Karlsruhe, on the German side, and scattered forest patches on both sides of the Rhine. The vegetation type has changed from floodplain vegetation to more desiccation tolerant tree species (e.g. robinia Robinia pseudoacacia, ash-leaf maple (Acer negundo), ash Fraxinus excelsior and common oak Quercus robur). Today, the region is mainly characterized by human-used agricultural fields, grasslands and vineyards, intersected by roads and settlements (Fig. 1c; for illustration purposes, a part of the study area with most samples was selected). Two main motorways and two major railways (two lanes, not fenced) cross the area, paralleling the Rhine River from north to south: on the German side, the federal motorway BAB5 (built in 1933, today more than 140,000 vehicles per day, at most parts six-lanes) and on the French side, the A35 (built in 1965, today more than 50,000 vehicles per day, at most parts four-lanes). The smaller motorway on the French site crosses also the wildcat distribution area while on the German side the BAB5 is at the border of the wildcat range.
Samples
A total of 101 wildcat samples were included in the present study (Fig. 1, Supplementary Tab. S1): 48 samples from west of the Rhine River and 53 samples from east of it. Wildcat samples were provided by François Léger (Office National de la Chasse et de la Faune Sauvage), Beatrice Nussberger (Institute of Evolutionary Biology and Environmental Studies, University of Zurich), Mathias Herrmann (ÖKO-LOG), Kathrin Steyer (Conservation Genetics, Senckenberg Research Institution and Natural History Museum Frankfurt), and the Zentrum für Fisch- und Wildtiermedizin (Vetsuisse-Faculty, University of Bern). It was checked genetically and, if possible, morphologically that all samples originated from pure wildcats. In total, 50 samples from wildcat carcasses (mostly road kills), 25 hair samples from captures (using lure sticks or carcasses), and 26 blood samples from captures were used. Genetic samples of captured cats were obtained as by-products of telemetry studies or routine analyses of veterinarians (Streif et al. 2012). Samples from living cats used in this study were collected in compliance with the respective local and national laws. No animal was harmed for the purposes of this study. Sampling was done between 1994 and 2013 by local experts (Supplementary Tab. S1) (annotation to the sampling period: only six of the samples used in final data analyses were sampled before the year 2000; running the analyses with only recent samples did not change results). Tissue samples were stored in 96 % ethanol and frozen at –20 °C, blood samples were stored in EDTA and frozen at −80 °C and hair samples were stored separately, in the dark, at +6 °C, prior to laboratory analysis.
Laboratory techniques
To minimize contamination risks (after Taberlet et al. 1999), DNA isolation and amplification of invasive (blood and tissue) samples and post-PCR procedures were conducted spatially separated from DNA isolation of non-invasive (hair) samples, in two different lab rooms. For DNA isolation from hair samples, 10–15 single hairs were pooled per sample. A minimum of 10 hairs was found to be able to minimize genotyping errors (allelic dropout and false alleles) (Goossens et al. 1998) while not showing contamination with other individuals that potentially visited the same lure stick between two samplings (Steyer et al. 2012). Each hair was visually inspected under the microscope to ensure that a hair root was present. DNA isolation was done using the First-DNA all-tissue DNA-extraction kit (GEN-IAL, Troisdorf, Germany) according to the manufacturers’ protocol. Isolation from blood samples was done using the QIAamp DNA MicroKit (Qiagen, Hilden, Germany), and isolation from tissue samples using the DNeasy Blood and Tissue Kit (Qiagen, Hilden, Germany), following the manufacturers’ instructions. DNA isolates were stored at −20 °C until polymerase chain reaction (PCR).
For genotyping, 20 microsatellite loci (Table 1) and one sex-marker (Menotti-Raymond et al. 1999) were analyzed, summarized in six multiplex PCRs. Ten μl PCR volume contained 10–50 ng/μl template DNA, HotStart Mastermix (Genaxxon bioscience, Ulm, Germany) and 0.6 μM of each primer mix. The PCR protocol was as follows: initial denaturation at 95 °C for 15 min, 45 cycles of denaturation at 94 °C for 30 s, annealing at 52 °C for 90 s and extension at 72 °C for 60 s, final extension at 72 °C for 30 min. PCR of each sample was repeated three times to avoid genotyping errors by allelic dropout (Navidi et al. 1992). Negative controls containing no template DNA were included in each PCR to check for potential contamination. PCR products were sized on an ABI 3130 DNA Analyzer (Applied Biosystems, Darmstadt, Germany). Fragment length was scored using GeneMapper v.4.0 (Applied Biosystems, Darmstadt, Germany). Consensus genotypes for the three replicates per sample were generated manually.
Data analysis
The data set was checked for genotyping errors using the program DROPOUT ver. 2.3.1 (McKelvey and Schwartz 2005), which identifies both microsatellite loci and samples that likely contain genotyping errors. To test for possible hybrids and domestic cats in the data set, the program STRUCTURE ver. 2.3.4 (Pritchard et al. 2000) was run with the genotypes from this study and a reference set of 74 domestic cat genotypes and 19 wildcat genotypes (Wildlife Monitoring, Forest Research Institute Baden-Wuerttemberg (FVA), Germany, morphologically and genetically determined by FVA and Senckenberg Research Institute, Frankfurt, Germany). The following specifications were applied: K = 2, length of burn-in period: 10,000, number of MCMC repetitions after burn-in: 100,000, admixture model and correlated allele frequencies, iterations: 100. A probability of Q (individual proportion of membership) >0.8 was considered as correct assignment to a cluster (Pierpaoli et al. 2003; Oliveira et al. 2007). Samples that were assigned to the domestic cat cluster with a proportion >0.2 were excluded from further analyses.
ARLEQUIN ver. 3.5 (Excoffier et al. 2005) was used to calculate Hardy–Weinberg-Equilibrium (HWE), observed (Ho) and expected heterozygosity (He), linkage disequilibrium, the number of alleles and allelic size range per locus. The most likely number of subpopulations for the set of samples was calculated using the program STRUCTURE. One to six clusters were tested, K = 1–6, all other specifications were applied as above. Both log-likelihood values calculated by STRUCTURE and structure harvester (Earl and VonHoldt 2011), implementing the Evanno method (Evanno et al. 2005), were used to infer the most likely number of clusters (K). The runs with the most likely K were pooled using the program CLUMPP ver. 1.1.2 (Jakobsson and Rosenberg 2007) and graphically displayed using the program DISTRUCT ver. 1.1 (Rosenberg 2003). TESS ver. 2.3.1 (Chen et al. 2007) was used to calculate the most likely pattern of structuring of the data set in a spatially explicit way. The following specifications were applied: multiple Ks ranging up to K = 6, runs per K = 100, sweeps: 50.000, burn-in sweeps: 10.000, admixture (CAR) model. The runs with the 10 % lowest deviance information criterion (DIC) values were averaged in CLUMPP (Jakobsson and Rosenberg 2007) and spatially displayed using the program R ver. 3.0.1 (R Development Core Team 2013) using the KridAdmixProportions script. Additionally, a discriminant analysis of principal components (DAPC) was performed using the adegenet package (Jombart2008; function dapc) for the software R (R Development Core Team 2013) to analyse the population structure and identify potential clusters of genetically related individuals. First, DAPC transforms the raw genetic data using principal components analysis (PCA), then the discriminant analysis attempts to maximize the differentiJombartation between groups while minimizing variation within groups (Jombart et al. 2010). The optimal number of PCA components to be retained was determined by applying the function optim.a.score (Jombart 2008). A Mantel test was performed in GenAlEx ver. 6.5 (Peakall and Smouse 2006; Peakall and Smouse 2012) to test isolation by distance (IBD; Wright 1943). To examine if the wildcat population east of the Rhine River could have survived in small refuges, a test of a recent expansion after a population reduction was executed with the program Bottleneck 1.2 (Piry et al. 1999). The program tests for an excess of heterozygosity, He, (after a population bottleneck) compared to the expected heterozygosity in a population at equilibrium, Heq. The program was run with 5000 iterations assuming a two-phase mutation model (TPM) with a proportion of mutations that follow the stepwise mutation model (SMM) set at 80 %. The significance was assessed with a Wilcoxon sign-rank test. GenAlEx was used to perform a spatial autocorrelation analysis, analysing the degree of dependency among observations in a geographic space. The spatial autocorrelation analysis was done for the whole data set, as well as for male and female individuals separately.
Results
DROPOUT identified neither microsatellite loci nor samples that were likely to contain genotyping errors. From the initial total set of 101 samples, 81 samples could be amplified successfully (with missing values in, at most, one quarter of the microsatellite loci; the total data set of 1620 values contained 36 missing values) and used in statistical analysis. The STRUCTURE analysis of the data set together with domestic cat and wildcat reference genotypes revealed 13 individuals with no explicit assignment (Q < 0.8) to the wildcat cluster. A Q-value less than 0.8 could be an indication for a recent hybridization or a former introgression with domestic cats. To be on the safe side, those 13 samples were excluded from the study, leaving a set of 68 presumably pure wildcat samples for further analyses. The number of alleles per locus varied from 4 (FCA23) to 11 (F37) with a mean of 7.8. We did not find any evidence for linkage disequilibrium and no significant deviations from Hardy–Weinberg-Equilibrium (HWE) could be detected after applying Bonferroni correction (Table 1). Observed heterozygosity (Ho) across all loci (mean Ho = 0.642) was not significantly (p = 0.57) lower than expected heterozygosity (mean He = 0.684).
An agglomeration of samples in the German Upper Rhine Valley led to an uneven distribution of samples. Therefore, we reduced half of the German samples, randomly, resulting in a total data set of 55 samples. We used the reduced data set in all subsequent analyses. Indicated by log-likelihood values, assignment analysis with STRUCTURE suggested a compilation of individuals into just one cluster (K = 1) (Fig. 2, Supplementary Tab S2). TESS did neither detect spatially discrete clusters of wildcat individuals (results not shown). In accordance, the DAPC did not reveal distinct clusters. Based on the Bayesian information criterion (BIC), the DAPC showed that all individuals belong to one cluster (Supplementary Fig. S1). Isolation by distance across the hole study site was found to be low but significant (R2 = 0.026, slope 0.019, P = 0.03). No significant signals of a recent bottleneck in the wildcat population east of the Rhine River were detected (one-tailed Wilcoxon sign-rank test for He excess, P = 0.61). The expected number of loci with heterozygosity excess was 11.85 under null hypothesis. The probability value was 0.57, meaning that the newly discovered wildcat population east of the Rhine River has not undergone a recent genetic bottleneck.
Spatial autocorrelation analysis for all individuals showed that, at a distance up to 10 km, relatedness among individuals was higher than expected under the assumption of randomness (Fig. 3). This effect was even stronger when analysing sex separately; males (N = 28) are more closely related to each other in distances below 10 km, whereas for females (N = 27) the distance was much shorter (5 km).
Results of spatial autocorrelation analysis a for female individuals, b for male individuals, c for all individuals; black r, grey upper and lower bounds of the 95 % confidence interval about the null hypothesis of no spatial genetic structure as determined by permutation. If the calculated r value falls outside this confidence belt, significant (p < 0.05) spatial autocorrelation exists
Discussion
Here we studied a presumably expanding wildcat population in the Upper Rhine Valley, in a fragmented and highly anthropogenically cultivated landscape. In our study, we asked if the Rhine River acts as a potential barrier between the wildcats east and west of the river and tried to answer where the newly discovered wildcat population on the east side originated from. We only found weak barrier effects for the wildcat populations east and west of the river, supporting our scenario II, a recolonisation of the region. The weak population structure within the studied individuals confirms that all studied individuals belong to one wildcat population presumably to the western wildcat population distributed in France and Germany, which is genetically separated from a central German population (Eckert et al. 2010; Hertwig et al. 2009). Furthermore, genetic diversity by means of observed heterozygosity (Ho = 0.64) was comparable to values found in other recent studies of this population, e.g. in Germany (Ho = 0.65, Hartmann et al. 2013) and France (Ho = 0.69, Say et al. 2012). As such, our results indicate a better exchange and higher crossing capability of the species in this region than previously assumed.
No barrier effect of the Rhine River
Here we specifically investigated if the Rhine River does, indeed, represent a barrier to wild cat movement and thus gene flow. We investigated restrictions to gene flow between neighboring wildcat populations, on either side of the river. However, we did not find a substantial barrier to gene flow and no genetic differentiation between the wildcats east and west of the Rhine River. All analysed individuals could be assigned to one single wildcat population.
Those genetic results are further supported by observational data. In 2010, a radio-tracked individual proved that wildcats are able to cross the Rhine River, within the study area, by swimming across the Rhine to an island (Stéphanie Kraft, FVA, pers. comm.). Such islands may serve as stepping stones to cross the river. Besides swimming, wildcats can cross the Rhine River using bridges or dams. Earlier studies (Klar et al. 2009) already found that wildcats regularly use crossing structures (such as over- and underpasses). In the study area, several possibilities for crossing the Rhine are available, such as watergates and bridges at power plants with low night traffic, amongst others.
Our results, indicating recent gene flow across the Rhine River and its associated anthropogenic structures, differ from the results of a study on wildcats in the Taunus and Hunsrueck mountains (Rhineland Palatinate, Germany). The study found a strong genetic separation of wildcat populations on either side of the river, although they argued that some individuals actually did migrate from one area to the other (Hartmann et al. 2013). This area is located only about 200 km downstream from our study area. What can explain these somehow contradicting results? Firstly, the geographical situations at the two study sites are different. River width, water level and flow velocity are lower in our study area compared to Hartmann et al. (2013), possibly facilitating river crossing. In addition, at our study site, the Rhine River encompasses several islands where wildcats already have been detected (F. Légér, pers. comm.). In addition, both study sites have different population histories. The newly discovered wildcat population in Baden Wuerttemberg was possibly colonized by neighboring populations that dispersed to Germany across the Rhine River in recent years. Thus, gene flow can be detected across the Rhine. In contrast, in Rhineland Palatinate, wildcat populations are known to have survived on either side of the Rhine River since the last century (Raimer Raimer 1988). As such, isolation effects might be more pronounced, as populations on each side of the river could have differentiated through genetic drift over time. The role of rivers as potential barriers to gene flow was the subject of research in many studies, with differing outcomes. Strong barrier effects of rivers were, for example, found in hyenas, wild dogs, cheetahs (Cozzi and Broekhuis 2013) and tamarins (Vallinoto et al. 2006), while only moderate barrier effects were identified in wolves (Blanco et al. 2005) and roe deer (Hepenstrick et al. 2012). Thus, barrier effects of rivers may not only be dependent on the characteristics of animals, such as size and ability to swim, but even more on the characteristics of the river, such as river width, flow rate, crossing structures, as well as surrounding matrix.
Range expansion across the Rhine River
East of the Rhine River, the wildcat population was assumed to be extinct between 1912 and 2006. After rediscovering wildcats in the Upper Rhine Valley in 2006, it was discussed whether a small population had survived undetected and is now growing again or whether this population really had been extinct and has only recently recolonized the area. Here we provide evidence supporting scenario II, recolonisation. Firstly, the absence of population differentiation is best explained by the hypothesis that the population originates from the expanding French population crossing the Rhine River. Secondly, we could not detect a recent genetic bottleneck in the wildcat population east of the Rhine River, which should have been the case if the population had survived in small refuges and was expanding, recently. Thirdly, if a small population had survived, this population should be present, not only in the Upper Rhine Valley, but also in suitable habitats of the Black Forest, as the dispersal time would have been adequate. This indicates recent immigration and ongoing range expansion from neighboring populations across the Rhine River. A possible barrier to colonization of Black Forest habitats could be transportation infrastructure, such as the federal motorway BAB5. This motorway runs from north to south, with four lanes and heavy traffic, six road-killed wildcats have been found there since 2011. In several other studies roads were already identified as barriers to wildcat dispersal (Klar et al. 2009; Hartmann et al. 2013).
Local spatial genetic structure
Spatial autocorrelation could be detected when analysing wildcats individually, indicating gene flow up to 10 km within the area, with higher distances for males than females. Those differences in structure between male and female individuals can be explained by differences in spatial movement, such as different dispersal distances and home range sizes (Stahl et al. 1988; Thiel 2004; Monterroso et al. 2009). In mammals, dispersal distances are generally higher (Greenwood 1980) in males than in females, and male home ranges are generally larger (Harestad and Bunnell 1979). Dispersal distances of wildcats are largely unknown, but several studies found that the home ranges of males are 2–3 times larger than the home ranges of females (Stahl et al. 1988; Thiel 2004; Monterroso et al. 2009) and can overlap with several female home ranges (Stahl et al. 1988; Biró et al. 2004). This fits well with observations in other felid species, where higher dispersal distances were found in males than in females (e.g. Iberian lynx (Lynx pardinus), Revilla et al. 2004; cougar (Puma concolor), Thompson and Jenks 2010). Low genetic spatial structure in our study area could, thus, be partially explained by higher dispersal distances and larger home ranges of males.
Implications for conservation
The results of this study indicate that the Upper Rhine Valley east of the Rhine River was recolonized by wildcats crossing the Rhine River from neighboring populations, presumably at the end of the twentieth century. The current study identifies the genetic status quo of this wildcat population and shows that, at least in the recent past, the crossing of the Rhine River was possible for individual wildcats. Expanding wildcat populations that were able to recolonize areas in spite of potential barriers have been reported previously (Say et al. 2012; Steyer et al. 2012), raising hope for the conservation of this, as well as other endangered species.
Nevertheless, existing crossing structures are essential for the survivability of the wildcat population east of the Rhine River, if not others. This population is small and hardly connected with other wildcat populations further east. At present, no further wildcat populations are known (e.g. in the Black Forest). Furthermore, within this study area, the habitat for wildcats surrounding the Rhine River (i.e. the amount of forest patches bordering on the river banks) is sparse and highly fragmented. Therefore, conservation strategies should focus on the maintenance of dispersal structures and on the realization of corridor networks, which have already been implemented in national strategies such as those in Baden Wuerttemberg (Müller et al. 2003) and throughout Germany (Birlenbach and Klar 2009). Of special significance for the wildcat population in our study area are, in this regard, structures to cross the BAB5 to allow further expansion towards the Black Forest. However, to ensure gene flow and, therefore, the long-term viability of the European wildcat, transnational networks on a larger scale are required.
References
Balkenhol N, Waits LP (2009) Molecular road ecology: exploring the potential of genetics for investigating transportation impacts on wildlife. Mol Ecol 18:4151–4164. doi:10.1111/j.1365-294X.2009.04322.x
Birlenbach K, Klar N (2009) Aktionsplan zum Schutz der Europäischen Wildkatze in Deutschland. Naturschutz Landsch 41:325–332
Biró Z, Szemethy L, Heltai M (2004) Home range sizes of wildcats (Felis silvestris) and feral domestic cats (Felis silvestris f. catus) in a hilly region of Hungary. Mamm Biol—Z Säugetierkd 69:302–310. doi:10.1078/1616-5047-00149
Blanco JC, Cortés Y, Virgós E (2005) Wolf response to two kinds of barriers in an agricultural habitat in Spain. Can J Zool 83:312–323
Brooks TM, Mittermeier RA, Mittermeier CG et al (2002) Habitat loss and extinction in the hotspots of biodiversity. Conserv Biol 16:909–923
Chen C, Durand E, Forbes F, François O (2007) Bayesian clustering algorithms ascertaining spatial population structure: A new computer program and a comparison study. Mol Ecol Notes 7:747–756
Cozzi G, Broekhuis F (2013) Comparison of the effects of artificial and natural barriers on large African carnivores: implications for interspecific relationships and connectivity. J Anim Ecol 82:707–715
Devictor V, Julliard R, Jiguet F (2008) Distribution of specialist and generalist species along spatial gradients of habitat disturbance and fragmentation. Oikos. doi:10.1111/j.2008.0030-1299.16215.x
Earl DA, VonHoldt BM (2011) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the evanno method. Conserv Genet Resour 4:359–361. doi:10.1007/s12686-011-9548-7
Eckert I (2003) DNA-Analysen zum genetischen Status der Wildkatze (Felis silvestris) in Deutschland. 1–101
Eckert I, Suchentrunk F, Markov G, Hartl GB (2010) Genetic diversity and integrity of German wildcat (Felis silvestris) populations as revealed by microsatellites, allozymes, and mitochondrial DNA sequences. Mamm Biol—Z Säugetierkd 75:160–174. doi:10.1016/j.mambio.2009.07.005
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. doi:10.1111/j.1365-294X.2005.02553.x
Excoffier L, Laval G, Schneider S (2005) Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Goossens B, Waits LP, Taberlet P (1998) Plucked hair samples as a source of DNA: reliability of dinucleotide microsatellite genotyping. Mol Ecol 7:1237–1241
Greenwood PJ (1980) Mating systems, philopatry and dispersal in birds and mammals. Anim Behav 28:1140–1162
Harestad AS, Bunnell FL (1979) Home range and body weight—a reevaluation. Ecology 60:389–402
Hartmann SA, Steyer K, Kraus RHS et al (2013) Potential barriers to gene flow in the endangered European wildcat (Felis silvestris). Conserv Genet 14:413–426. doi:10.1007/s10592-013-0468-9
Hepenstrick D, Thiel D, Holderegger R, Gugerli F (2012) Genetic discontinuities in roe deer (Capreolus capreolus) coincide with fenced transportation infrastructure. Basic Appl Ecol 13:631–638. doi:10.1016/j.baae.2012.08.009
Herdtfelder M, Strein M, Suchant R (2007) Wildkatzen am Kaiserstuhl. Naturschutz Landsch 10:320–328
Herrmann M, Vogel C (2005) Wildkatze Felis silvestris sivestris Schreber, 1777. In: Braun M, Dieterlen F (eds) Die Säugetiere Baden-Württembergs, vol 2. Eugen Ulmer, Stuttgart, pp 363–376
Hertwig ST, Schweizer M, Stepanow S et al (2009) Regionally high rates of hybridization and introgression in German wildcat populations (Felis silvestris, Carnivora, Felidae). J Zool Syst Evol Res 47:283–297
Holderegger R, Di Giulio M (2010) The genetic effects of roads: a review of empirical evidence. Basic Appl Ecol 11:522–531. doi:10.1016/j.baae.2010.06.006
Jakobsson M, Rosenberg N (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806. doi:10.1093/bioinformatics/btm233
Jombart T (2008) Adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24(11):1403–1405
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genetics 11(1):1
Klar N, Herrmann M, Kramer-Schadt S (2009) Effects and mitigation of road impacts on individual movement behavior of wildcats. J Wildl Manag 73:631–638. doi:10.2193/2007-574
McKelvey KS, Schwartz MK (2005) Dropout: a program to identify problem loci and samples for noninvasive genetic samples in a capture-mark-recapture framework. Mol Ecol Notes 5:716–718. doi:10.1111/j.1471-8286.2005.01038.x
Menotti-Raymond M, David VA, Lyons LA et al (1999) A genetic linkage map of microsatellites in the domestic cat (Felis catus). Genomics 57:9–23. doi:10.1006/geno.1999.5743
Monterroso P, Brito JC, Ferreras P, Alves PC (2009) Spatial ecology of the European wildcat in a Mediterranean ecosystem: dealing with small radio-tracking datasets in species conservation. J Zool 279:27–35. doi:10.1111/j.1469-7998.2009.00585.x
Müller U, Strein M, Suchant R (2003) Wildtierkorridore in Baden-Württemberg. Berichte Freiburger forstliche Forschung 48:46
Navidi W, Arnheim N, Waterman MS (1992) A multiple-tubes approach for accurate genotyping of very small DNA samples by using PCR: statistical considerations. Am J Hum Genet 50:347–359
Noss RF, Quigley HB, Hornocker MG et al (1996) Conservation biology and carnivore conservation in the Rocky mountains. Conserv Biol 10:949–963
Nowell K, Jackson P (1996) Wild cats. status survey and conservation action plan. IUCN/SSC Cat Specialist Group, Switzerland, pp 1–383
Nussberger B, Greminger MP, Grossen C et al (2013) Development of SNP markers identifying European wildcats, domestic cats, and their admixed progeny. Mol Ecol Resour 13:447–460. doi:10.1111/1755-0998.12075
Oliveira R, Godinho R, Randi E et al (2007) Molecular analysis of hybridisation between wild and domestic cats (Felis silvestris) in Portugal: implications for conservation. Conserv Genet 9:1–11. doi:10.1007/s10592-007-9297-z
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in excel. Population genetics software for teaching and research. Mol Ecol Notes 6:288–295
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in excel. Population genetic software for teaching and research–an update. Bioinformatics 28:2537–2539. doi:10.1093/bioinformatics/bts460
Piechoki R (1990) Die Wildkatze - Felis silvestris., Neue Brehm BüchereiA Ziemsen, Wittenberg Lutherstadt
Pierpaoli M, Biro ZS, Herrmann M et al (2003) Genetic distinction of wildcat (Felis silvestris) populations in Europe, and hybridization with domestic cats in Hungary. Mol Ecol 12:2585–2598. doi:10.1046/j.1365-294X.2003.01939.x
Piry S, Luikart G, Cornuet JM (1999) BOTTLENECK: a computer program for detecting recent reductions in the effective population size using allele frequency data. J Hered 90:502–503
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Raimer F (1988) Die Wildkatze in Hessen und Niedersachsen - Biotop, Umwelt, Verbreitung, Bestandsentwicklung, Gefährdung, Schutz
R Development Core Team (2013) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.r-project.org/
Revilla E, Wiegand T, Palomares F et al (2004) Effects of matrix heterogeneity on animal dispersal: from individual behavior to metapopulation-level parameters. Am Nat 164:E130–E153. doi:10.1086/424767
Rosenberg NA (2003) Distruct: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138. doi:10.1046/j.1471-8286.2003.00566.x
Say L, Devillard S, Léger F et al (2012) Distribution and spatial genetic structure of European wildcat in France. Anim Conserv 15:18–27. doi:10.1111/j.1469-1795.2011.00478.x
Stahl P, Artois M (1991) Status and conservation of the wildcat in Europe and around the Mediterranean rim. Council of Europe, Strasbourg
Stahl P, Artois M, Aubert MFA (1988) Organisation spatiale et déplacement des chats forestiers adultes (Felis silvestris Schreber 1777) en Lorraine. Rev d’Ecologie, Terre Vie 43:113–132
Steyer K, Simon O, Kraus RHS et al (2012) Hair trapping with valerian-treated lure sticks as a tool for genetic wildcat monitoring in low-density habitats. Eur J Wildl Res 59:39–46. doi:10.1007/s10344-012-0644-0
Streif S, Kraft S, Veith S et al (2012) Monitoring and research of the European wildcat (Felis silvestris) in Baden-Württemberg. Säugetierkundliche Informationen 8:411–416
Taberlet P, Waits L, Luikart G (1999) Noninvasive genetic sampling: look before you leap. Trends Ecol Evol 14:323–327
Thiel C (2004) Streifgebiete und Schwerpunkte der Raumnutzung von Felis silvestris silvestris (Schreber 1777) in der Nordeifel. 1–195
Thompson DJ, Jenks JA (2010) Dispersal movements of subadult cougars from the Black Hills: the notions of range expansion and recolonization. Ecosphere. doi:10.1890/ES10-00028.1
Vallinoto M, Araripe J, Rego PS et al (2006) Tocantins river as an effective barrier to gene flow in Saguinus niger populations. Genet Mol Biol 29:215–219
Wilcove DS, McLellan CH, Dobson AP (1986) Habitat fragmentation in the temperate zone. In: Soule ME (ed) Conservation biology: the Science of Scarcity and Diversity. Sinauer Associates, Sunderland, pp 237–256
Woodroffe R, Ginsberg J (1998) Edge effects and the extinction of populations inside protected areas. Science 280:2126–2128
Wright S (1943) Isolation by distance. Genetics 28(2):114
Acknowledgments
We are grateful to the numerous persons who helped to conduct this research, in particular, our collegues at the Forest Research Institute Baden Wuerttemberg, Stéphanie Kraft, Sarah Veith, Mara Sandrini and Rudi Suchant for their support and constructive discussions. Sampled wildcat material and expertise was provided by François Léger (Office National de la Chasse et de la Faune Sauvage), Beatrice Nussberger (Institute of Evolutionary Biology and Environmental Studies, University of Zurich), Mathias Herrmann (ÖKO-LOG), Kathrin Steyer (Conservation Genetics, Senckenberg Research Institution and Natural History Museum Frankfurt), the Zentrum für Fisch- und Wildtiermedizin (Vetsuisse-Faculty, University of Bern) and the numerous people who regularly controlled our lure sticks. We acknowledge the funding for this study from the Landesjagdabgabe Baden Wuerttemberg. Stephanie Witczak helped revising the English of the present paper. We thank the two anonymous referees for providing constructive comments and help in improving the contents of this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Würstlin, S., Segelbacher, G., Streif, S. et al. Crossing the Rhine: a potential barrier to wildcat (Felis silvestris silvestris) movement?. Conserv Genet 17, 1435–1444 (2016). https://doi.org/10.1007/s10592-016-0874-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-016-0874-x