Abstract
Cancer-initiating cells (CIC) are the driving force in tumor progression. There is strong evidence that CIC fulfill this task via exosomes (TEX), which modulate and reprogram stroma, nontransformed cells, and non-CIC. Characterization of CIC, besides others, builds on expression of CIC markers, many of which are known as metastasis-associated molecules. We here discuss that the linkage between CIC/CIC-TEX and metastasis-associated molecules is not fortuitously, but relies on the contribution of these markers to TEX biogenesis including loading and TEX target interactions. In addition, CIC markers contribute to TEX binding- and uptake-promoted activation of signaling cascades, transcription initiation, and translational control. Our point of view will be outlined for pancreas and colon CIC highly expressing CD44v6, Tspan8, EPCAM, claudin7, and LGR5, which distinctly but coordinately contribute to tumor progression. Despite overwhelming progress in unraveling the metastatic cascade and the multiple tasks taken over by CIC-TEX, there remains a considerable gap in linking CIC biomarkers, TEX, and TEX-initiated target modulation with metastasis. We will try to outline possible bridges, which could allow depicting pathways for new and expectedly powerful therapeutic interference with tumor progression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
1.1 Historic overview on stem cells, metastasis markers, and exosomes in metastasis
In the late 1980, the first reports described metastasis-associated molecules [1,2,3]. To name a few, integrin α6β4 exits hemidesmosomes during carcinoma progression, facilitating tumor progression by cooperating with and amplifying signaling via the growth factor receptors EGFR, ERBB2, cMET, and others. It contributes to invasion by affecting promoter DNA demethylation of metastasis-associated S100A4 and autotaxin and upregulation/activation of NFATFootnote 1 and NFκB1, tumor-promoting transcription factors (TF) [4]. The metastasis-promoting activity of CD44v6 was first described in 1991 [5]. Overexpression of MTA11, noted in 1994 [6], is an important component of NURD1 acting as corepressor and coactivator of many genes including p53 and c-myc [7, 8]. Other metastasis-promoting surface markers were identified by proteome analysis [9, 10]. Importantly, metastasis-associated markers are not oncogenes, but molecules expressed in nontransformed cells, possibly upregulated during activation processes. This culminated in an early review outlining metastasis and tumorigenesis being two independent events [11].
About a decade formerly, the discussion on cancer stem cells started, pushed by the availability of hematopoietic stem cells (SC) [12] and the translation toward leukemia SC [13]. Cancer stem cells/cancer-initiating cells (CIC) were first defined by the capacity of human cancer cells to grow in xenogeneic, immunocompromised mice [14, 15]. Additional features like EMT (epithelial–mesenchymal transition), still disputed to be required [16], and the so-called CIC markers soon came into play, the latter largely overlapping with metastasis-associated markers [17,18,19].
Independently, the slowly starting story on exosomes (Exo) turned from a wastebasket [20, 21] into a magic bullet, with tumor exosomes (TEX) and CIC-TEX being suggested to affect non-CIC, endothelial, and hematopoietic cells as well as host and tumor stroma, supporting tumor growth and angiogenesis, deviating immune reactivity, and contributing in assembling a niche for migrating tumor cell settlement [22,23,24,25,26,27].
With excellent reviews on all these topics, we will introduce CIC markers, EMT-related TF, noncoding (nc)RNA, and TEX in pancreas and colorectal cancer to discuss the connection between these components suggested to be the essential cues.
1.2 Colorectal and pancreatic cancer
Colorectal cancer (CoCa), the third most common cancer, shows a slight tendency toward decrease [28, 29]. The mortality rate varies greatly depending on the disease stage. Over 90% of stage I patients have a survival rate of > 5 years, which drops to 10–20% in patients with metastases and to 25–50%, if metastases are resectable. The incidence of metastasis at diagnosis is in the range of 10%, but 50% of patients with a resectable CoCa at diagnosis develop metastasis after resection [29,30,31].
Pancreatic cancer (PaCa), the fourth leading cause of cancer-related death in Western countries, is the most lethal cancer with an overall 5-year survival of approximately 5% [28, 31, 32]. Surgery is the only curative treatment option, but 80% of patients are inoperable at diagnosis, and due to local recurrence and metastatic spread, the survival rate does not exceed 15–20% even after resection [33]. This is particularly alarming, and the incidence of pancreatic ductal adenocarcinoma, the most frequent subtype [33], steeply increasing, is expected to become the second leading cause of cancer-related death after lung cancer by 2030 [34]. The high mortality, due to early spread and radio- and chemotherapy resistance [35], is provoked by the small population of CIC [36].
1.3 Cancer-initiating cells
Development of a tumor is a stepwise process, where multiple genetic and epigenetic alterations, like aberrant DNA methylation, histone modification, and altered ncRNA expression [37], are culminating to reach unlimited growth and invasion. This accounts also for CoCa and PaCa [38, 39]. However, metastases require additional alterations, which mostly are restricted to CIC [40, 41]. CIC characterization and sharing with embryonic and adult stem cells (ESC, ASC) will be briefly introduced.
Stem cells are a rare population of cells with self-renewal and differentiation capacity. SC are defined as omnipotent, pluripotent, and multipotent. The zygote and early blastomeres are totipotent as they form all tissue from the embryo and the supportive extraembryonic tissue. ESC (inner mass of the blastocyst) are pluripotent generating the ectoderm, mesoderm, and endoderm of the developing embryo, but not the placenta. Multipotent, tissue-restricted ASC generate the cell types according to their tissue location [42, 43]. Notably, somatic cell nuclear transfer proved the nuclear equivalence throughout development, with a sheep being the first mammalian to demonstrate [44]. Soon thereafter, it was shown that TF play central roles in specifying cell fate [45, 46].
The pancreas develops from the junction of the foregut and midgut from dorsal and ventral pancreatic buds [47], the cells representing early multipotent progenitors, which form all lineages—acinar, ductal, and endocrine—of the pancreas [48]. They express and are critically dependent on PDX11. However, studies to establish PDX1 expressing pancreatic SC and particularly insulin-secreting cells from pluripotent ESC or human-induced (hi)PSC failed or were of very low efficiency and burdened by occasional development of teratoma [49], which led less than 4 years ago to the notion “pancreatic stem cell remain unresolved” [50]. The problem recently was sorted out based on increasing technological progress combining clonal tracing and whole-mount reconstruction [51]. The authors describe that self-renewing precursors at the termini of growing ducts drive ductal bifurcation, while multipotent precursors become fate restricted giving rise to acinar-committed precursors as well as ductal progenitors and endocrine cells providing an answer to large-scale patterning of pancreatic subcompartments [52]. This outstanding work offers a solid base for progress in the therapy of diabetes mellitus and for defining PaCIC.
The intestine contains two stem cell populations (ISC). ISC at the bottom of the crypt can divide rapidly giving rise to transit amplifying cells, which move to the inner surface of the crypt and differentiate into epithelial cells. These ISC are called active (A)-ISC. There is a second low cycling population (+4 cells), also called quiescent (Q)-ISC located above the Paneth cells. Q-ISC can replace A-ISC upon loss. More recently, it was suggested that the +4 cells are nondividing precursors of secretory cells that can dedifferentiate to replace damaged ISC [53]. The main markers of A-ISC are LGPCR 51, PHLDA11, ASCL21, OLFM41, and SOX91; the main markers of Q-ISC are BMI11, LRIG11, TERT1, DCLK11, MSI11, and HOPX1 (rev. in [54]). ISC reside in a niche composed of surrounding tissue cells including Paneth cells, which are essential for ISC expansion. They secrete large amounts of EGF1, WNT31, and DLL41 providing the essential components for LGR5+ ISC (rev. in [55]). Concerning about TF and signaling in ISC [56], the Wnt pathway is essential for crypt formation and renewal [57, 58]. RSPO1 binds LGR51 and homologs and enhances Wnt signaling [59, 60].
There is compelling evidence that hematopoietic and solid organ–derived malignancies also contain a small population of CIC. Originally discussed to derive from ASC, CIC are now considered to evolve independently [61,62,63].
CIC have the capacity of self-renewal and differentiation [64, 65]. They are characterized by rare division [66,67,68], longevity [69, 70], drug and radiation resistance [71,72,73,74], and migratory activity [75,76,77]. Upon serial transplantation of human CIC in xenogeneic, immunocompromised hosts, outgrowing tumors resemble the original heterogeneous population [78, 79]. CIC were first identified by Lapidot and Dick in human AML (acute myeloid leukemia) as a CD34+CD38− subpopulation [80]. Al-Hajj et al. identified a tumorigenicity of CD44+/CD24−/±/lineage− breast cancer cells as [81]. Meanwhile, CIC were identified in solid tumors of epithelial and mesenchymal origin of most organs (rev. in [15]). Besides, by serial transplantation [82], CIC are defined as side population by Hoechst 33342 exclusion [83], by the capacity of spheroid growth in serum-free medium [84], and by altered metabolism. CIC display high ALDH1, predominantly ALDH1A1 and ALDH3A1, that contributes to metabolic reprogramming by oxidizing absorbed vitamin A to RA1 acting as cotranscription factor for RARB1 in the nucleus, c-Myc1, CCND11, and others, which affects various CIC properties (rev. in [70, 85,86,87]). Finally, CIC are enriched by flow cytometry according to CIC marker expression (rev. in [88, 89]).
With great hope being given to CIC marker-based therapy [90, 91], we proceed with an overview on prominent PaCIC and CoCIC markers.
2 Function-relevant PaCIC and CoCIC components
CIC are suggested accounting for metastatic spread. The components that may be of major importance are CIC/metastasis markers, ncRNA, particularly mi (micro)RNA and long-nc (lnc)RNA, EMT-related TF, and Exo that help transfer the CIC message. CIC sharing many markers and features with ESC and ASC can pose a problem on their elimination but is helpful in unraveling their mode of action.
2.1 Stem cell markers
CIC markers, different to oncogenes, are expressed in nontransformed cells. The most relevant PaCIC and CoCIC markers are EPCAM1, mediating homotypic adhesion [92], CD44s/CD44v6 (CD44 standard/CD44 variant 6 isoform) [93], and CD1661, which belongs to the Ig superfamily and displays homotypic and heterotypic binding to CD6. Its function in CIC may rely on coexpression with LGR5, MSI1, and DCLK1, the cells being suggested to provide a kind of reservoir to shift from homeostasis to induced SC [94]. EPHB21 [95], cMET1 [96], and CXCR41 [97] also are considered as CIC markers in gastrointestinal cancer (rev. in [98, 99]). We briefly describe CD133 and LGR5 and, in some detail, CD44v6, Tspan8 (tetraspanin 8), EPCAM, and claudin7.
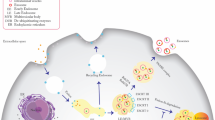
CD133 is a 5-transmembrane molecule [100] whose expression is enhanced by intracellular binding molecules HDAC61 and PTPRK1 [101, 102]. CD133 is confined to protruding membrane subdomains, where it interacts with cholesterol-based lipid rafts [103]. It is supposed to be engaged in cell polarity and to be integrated in cell–cell and cell–extracellular matrix (ECM) interactions [104]. By its integration in microdomains harboring signal transduction molecules, it becomes engaged in signaling cascades [105]. Finally, it is recovered in extracellular vesicles, originally called prominosomes that contribute to intercellular communication [106] (Fig. 1a).
CIC markers in pancreatic and colorectal cancer. a–h Schematic presentation of the most prominent CIC markers CD133, LGR5, CD44/CD44v6, Tspan8, a6b1/a6b4, claudin7, EPCAM, and CD24 in PaCa and CoCa including some of the ligands and signaling pathways. Arrows indicate associations between the distinct CIC markers. Full names of protein symbols are listed in Table S1
LGR51 [107] is expressed in ASC, best explored for ISC, where it promotes Wnt signaling through binding to its ligand RSPO [108]. In the absence of Wnt, a complex is formed between FZD1, LRP5/61, and RNF431, an E3 ubiquitin ligase, which promotes FZD ubiquitination and degradation. Upon RSPO binding to LGR4/5, RNF43 becomes phosphorylated and sequestered, and a more stable complex between RSPO, LRP5/6, and Wnt-FZD is established that promotes liberation of CTNNB11and β-catenin-LEF/TCF1 induction of Wnt genes (rev. in [109]). With LGR5 being recovered in CIC of many malignancies, it became tempting to speculate that elimination of LGR5 may suffice for tumor eradication. Though local tumor growth was only transiently retarded, metastatic growth could be inhibited. The only transient retardation of local growth is due to the plasticity of CIC, where differentiated cells can revert to LGR5+ CIC [110, 111]. Proving the essential role of LGR5 in tumor progression, the findings also stress SC plasticity (Fig. 1b).
2.1.1 CD44/CD44v6
The type I transmembrane glycoprotein CD44 varies in size due to N- and O-glycosylation and insertion of alternatively spliced exon products [112,113,114]. The standard isoform (CD44s) has seven extracellular domains, a transmembrane, and a cytoplasmic domain. By alternative splicing between exons 5 and 6, different combinations of 1–10 variant exon products are inserted (CD44v) [114]. CD44 belongs to the cartilage link protein family, conserved cysteines stabilizing the globular structure and two cysteines in the flanking region accounting for link domain folding [115]. The globular domain is followed by the heavily glycosylated stalk-like region, containing putative proteolytic cleavage sites and variable exon products [116,117,118]. The transmembrane region supports oligomerization and glycolipid-enriched membrane domain recruitment (GEM), important for interactions with extracellular ligands and other transmembrane and cytoplasmic molecule associations [119]. The cytoplasmic tail binds to cytoskeletal proteins [120, 121]. Wide CD44s expression differs from CD44v expression only in epithelial and hematopoietic cell subpopulations and frequent upregulation in CIC [122].
CD44 has multiple ligands. The link domain binds collagen, LN1, FN1, and E- and L-selectin [123, 124]. A basic motif outside the link domain binds to HA1, with CD44 being the major HA receptor [125] and having two additional glycosaminoglycans (GAG) binding sites [126, 127]. Post-translational CD44v modifications support growth factor (GF) binding: CD44v6 binding HGF1, VEGF1, and OPN1 [128,129,130], thereby CD44v promotes RTK (receptor tyrosine kinase) and GPCR activation [131]. The cytoplasmic tail binds ankyrin, ezrin, radixin, moesin (ERM) cytoskeletal linker proteins [121, 132]. HA-dependent adhesion and motility is aided by ankyrin contacting spectrin [121]. The activated ERM proteins N-terminus binds CD44 and the C-terminus F-actin, regulating migration, cell shape, and protein resorting [132,133,134]. Cytoskeletal linker protein binding contributes to expand CD44-mediated downstream signaling pathway activation [121, 133].
The lateral associations with distinct proteins are central to understanding the multitude of CD44/CD44v activities. cMET activation through CD44v6 HGF binding requires the interaction of the CD44 cytoplasmic tail with ERM proteins for Ras–MAPK1 pathway activation [135]. CD44v6–ECM binding also promotes cMET transcription [136]. Similar observations account for CD44v6 cross-linking–induced activation of IGF1R1 and PDGFR1 [137]. CD44 also associates with the GPCR CXCR4 [138], ABC1 transporters [139], additional antiapoptotic proteins [140, 141], and membrane-bound MMP141 and HYAL21 [142]. Upon GEM recruitment, CD44v6 associates with Tspan8. By the LRP6 association, CD44 contributes to EMT-related transcription factor Wnt signaling pathway activation [143]. The engagement in non-RTK pathways proceeds via activated RTK, GPCR, and cytoskeletal linker proteins or directly via GEM-recruited CD44v [144,145,146] (Fig. 1c).
We will discuss in detail how CD44 ligand binding and lateral associations contribute to CIC maintenance, apoptosis resistance, EMT, and the metastatic cascade.
2.1.2 Tetraspanins
Tetraspanins, 4-transmembrane proteins, have a small and a large extracellular loop [147]. The latter is engaged in dimerization and association with nontetraspanin partners, prominently integrins and proteases [148, 149]. Tetraspanins also bind cytoskeleton and cytosolic signal transduction molecules [150,151,152]. Tetraspanins form TEM-located (tetraspanin and glycolipid-enriched microdomains) webs, which are prone for internalization. Palmitoylation of intracellular, juxtamembrane cysteines supports tetraspanin web formation, protects from lysosomal degradation, and links tetraspanins to cholesterol and gangliosides. Importantly, after fission and scission, the tetraspanin webs are maintained during intracellular vesicle trafficking and are recovered after excretion in exosomes [153,154,155,156] (Fig. 1d,e).
Tspan8 is upregulated in ovarian, hepatocellular, and gastric cancer; malignant melanoma; and glioma [157,158,159,160,161] and is enriched in CoCIC and PaCIC [162,163,164]. Tspan8 promotes migration, invasion, and tumor progression [162,163,164,165], which relies in part on integrin recruitment, accompanied by integrin activation and initiation of downstream signaling [166, 167] and the cooperation with proteases [158, 167,168,169]. By associating with CD44v6, cMET and additional RTK become recruited [167, 170]. Finally, Tspan8 is engaged in the crosstalk with the tumor surrounding and the tissue in premetastatic niches [156] and promotes endothelial cell (EC) progenitor maturation and activation [171, 172]. These activities are a sequel of the Tspan8 engagement in Exo biogenesis and binding [156, 173].
In brief, the CIC marker Tspan8 contributes to tumor progression by arranging and clustering integrins and RTK, facilitating downstream signaling induction and the interaction with the surrounding. This accounts for CIC as well as CIC-TEX.
2.1.3 EPCAM
The epithelial cell adhesion molecule (EPCAM), mediating homophilic cell–cell adhesion [174], is overexpressed in many epithelial cancers, serving as a diagnostic and therapeutic target [175]. Besides interfering with E-cadherin-mediated adhesion, oncogenic and tumor progression supporting activity of EPCAM relies on engagement in Wnt/β-catenin signaling, on controlling motility by PRKC1 downregulation and MMP7 upregulation [176, 177]. After nuclear translocation, the cleaved intracellular domain (ICD) acts as a cotranscription factor for c-myc, cyclinA/E, Oct4, Nanog1, and others [178, 179]. EPCAM expression varies at different stages of tumor progression, with a transient downregulation during EMT being discussed [180,181,182]. Nonetheless, its CIC activity is supported by strong overexpression in metastasizing tumor cells [183] and is endorsed by its contribution to ESC pluripotency maintenance [184, 185].
Summarizing, a cell–cell adhesion molecule, expected to hamper metastasis, contributes to tumor progression in part by the cleaved ICD acting as a cotranscription factor. Notably, EPCAM can be recruited via claudin7 (cld7) into TEM, which adds a further dimension to its multiple activities (Fig. 1f,g).
2.1.4 Claudin7
Claudin7 is a member of four-pass proteins, central components of tight junctions (TJ) [186, 187]. A cld7-knockout (ko) being lethal within 10 days after birth due to intestine destruction [188, 189] might be due to a missing integrin association and a striking MMP3 upregulation [188] or as suggested by an intestine-specific conditional cld7ko due to enhancement of paracellular small organic solute flux across the TJ, which includes a major bacterial product initiating colonic inflammation [189]. The latter argues for loss of TJ-integrated cld7 accounting for lethality, and the former could also rely on non-TJ–integrated cld7. There are two modes for cld7 recovery outside of TJ [190, 191]. First, TJ are continuously remodeled. Claudins are PRKA and C and MLCK1 targets and cld phosphorylation prohibits TJ integration, with dysregulation of TJ being accompanied by loss of epithelial cell polarization and barrier function causing cell death and inflammation [192,193,194]. Internalized, TJ-excluded cld are recruited into Exo [195]. Cld7 also can become palmitoylated, a prerequisite for partitioning into TEM [196, 197], an important item in Exo biogenesis [198]. Palmitoylated, TEM-located cld7 associates with EPCAM and Tspan8 (Fig. 1f). Thus, cld7 is recovered in two distinct Exo populations, derived from TEM or from vesicles harboring “TJ-excluded” cld7 [199].
2.2 CIC markers and the contribution of epithelial–mesenchymal transition and transcription factors to SC and CIC maintenance
Transcription factors are of central importance in EMT during development and tumor progression. As there are strong links between EMT and SC/CIC including CIC markers, we will provide a condensed overview with emphasis on TF and PaCIC/CoCIC marker connections.
2.2.1 The network of stem cell transcription factors
During embryogenesis, a cell continuously requires changing the phenotype. It was sought that an endpoint was reached at the stage of a terminally differentiated somatic cell. This viewpoint changed, noting that a differentiated cell can be reprogrammed to pluripotency, the so-called iPSC reprogramming relying on an interplay between TF, chromatin modifiers, and regulatory motifs [200, 201]. Importantly, the phenotype of a cell is dictated by sets of TF that are regulated by other TF responding to extracellular signals, with the TF networks being central in regulating SC/CIC fate. There are two classes of TF: general TF, which bind to promoters and recruit polymerase II initiating transcription, and tissue-specific TF, which bind to promoter or enhancer regions and recruit general TF. Furthermore, several TF can function synergistically by binding to a superenhancer, a cluster of several enhancers that are highly cell-type specific and recruit additional cofactors like RNA polymerase II. Superenhancers may bridge some TF deficits lowering the binding threshold provoking a transcriptional burst. Last, but not least, extracellular factors play an important role in initiating TF activation, most prominently Wnt, LIF1, and TGFβ1 (rev. in [202]).
2.2.2 EMT and transcription factors contributing to PaCIC and CoCIC maintenance
PaCIC share with ESC a variety of TF, mostly OCT41, SOX2, and NANOG that, however, are distinctly regulated in CIC [203]. SOX2 expression is driven by HH1–EGFR1 signaling. SOX2 is critical for proliferation, dedifferentiation, and gain of stem cell features, with cyclin inhibitors being SOX2 targets. SOX2 also drives sphere formation and CIC marker expression and induces SNAIL1, Slug1, and Twist1, contributing to EMT [204,205,206]. EMT-TF adding to PaCIC progression, an elegant study of Roe et al. unraveled massive changes in enhancer activity driven by FOXA11, activating a transcriptional program of embryonic foregut endoderm [207].
Heterogeneity of CoCIC relies on context and surrounding-dependent patterns of gene expression and methylation, with the differentiation states of CIC and non-CIC changing in both directions [208, 209]. Signaling pathways regulating the plasticity of CoCIC are not fully unraveled. APC1 mutations contribute to Wnt/β-catenin pathway activation only in cells with a high level of Wnt. Wnt/β-catenin pathway activation also is achieved by KRAS1 mutation, NOTCH, HH, BMP1, PI3K1/AKT1 activation or metabolic changes, and high level of ROS (reactive oxygen species) activating the NFκB pathway [210,211,212,213,214,215]. A contribution via the interaction of Wnt with the HIPPO pathway is still discussed, with the HIPPO transducer YAP1 being a transcriptional coactivator. In ISC, YAP supports stemness by inducing β-catenin nuclear localization. Instead, in CoCa, YAP contributes to tumorigenesis and a β-catenin–YAP complex induces anti-apoptotic gene expression (rev. in [216]). The four NOTCH receptors, NOTCH signaling regulating self-renewal and repression toward secretory cell differentiation [217], distinctly regulate CIC-related TF (rev. in [218]). NOTCH 1 expression, correlating with cMET and CD44, increases migration and anchorage-independent growth via Slug, Smad3, and Jagged1 [219]. Further approaching the role of NOTCH in CoCIC homeostasis revealed engagement of the circadian clock gene PER31, overexpression decreasing Notch1, Jagged1, β-catenin, c-Myc, and LGR5 expression, accompanied by reduced drug resistance and self-renewal [211]. STRAP1 antagonizes NOTCH signal activation by competitively disrupting the association of the chromatin modifier PRC21 subunits. The authors suggest the STRAP–NOTCH1–HES11 axis as an important CoCIC regulator [220]. CoCIC are also regulated by BMP2 and BMP4, interfering with stemness by promoting differentiation through antagonizing Wnt/β-catenin signaling, where the zinc-finger TF GATA61 drives LGR5 expression in CoCIC and restricts BMP signaling to differentiated tumor cells by competing with β-catenin/TCF4 binding to a distal BMP4 regulatory region [213]. However, explaining the repeatedly described opposing effects of BMP on Wnt signaling, BMP inhibits Wnt signaling only when p53 and SMAD4 are unaffected [221]. Thus, depending on STAT31 activation, BMP2 supports CoCIC/EMT via activation of pSMAD1/5 and SNAIL [222]. BMP4, a direct target of intracellular TH1 that induces differentiation, modulates in a positive autoregulatory feedback loop TH signaling, mitigating Wnt activity [223].
Finally, the surrounding tissue impacts PaCIC and CoCIC (rev. in [224, 225]). The tumor stroma is characterized by an abundance of connective tissue and mesenchymal lineage-derived cells, most prominently fibroblasts. Cancer-associated fibroblasts (CAF) can have different origin and are characterized by amply secreting factors that affect CIC, including TGFβ, HFG, EGF, FGF1, OPN, and SDF11, which contribute to progenitor reprogramming toward CIC [93, 226]. CAF also secrete inflammatory IL6 and IL8, which activate the JAK1/STAT pathway and recruit inflammatory cells [227], and IL17A, which increases CIC self-renewal and invasion [228]. CAF can convert to αSMA1 expressing myofibroblasts and support anaerobic glucose metabolism [229, 230]. Vascular EC promote CIC survival and self-renewal through provision of DLL1, which assists Notch signaling [231]. Nonetheless, the view of the tumor stroma severely aggravating tumor progression recently changed. Stroma-derived HH controls epithelial SC and CIC via BMP, in PaCa and CoCa loss of stroma HH being accompanied by decreased BMP activity. Thus, stroma-specific HH acts as a metastasis suppressor via modulation of BMP signaling in CIC [232, 233].
2.3 Noncoding RNA
The recovery of function-relevant ncRNA has become a milestone in cell biology, being of central importance in development, homeostasis maintenance, and disease. The group of ncRNA still increasing, miRNA and lncRNA being so far the best explored in SC and CIC regulation [234], is shortly introduced.
2.3.1 MiRNA
MiRNA, the effectors of the endogenous RNA interference machinery, inhibit translation of protein-coding genes [235]. Long, capped and polyadenylated transcripts (pri-miRNA) form hairpins [236] that are processed by the ribonuclease III Drosha and the RNA-binding protein microprocessor complex subunit DGCR81, which generates 60–70 nt pre-miRNA [237]. Pre-miRNA is exported and is processed in the cytoplasm by Dicer1 to mature miRNA [238]. The mature miRNA is loaded into RISC (RNA-induced silencing complex) [239]. RISC-loaded miRNA bind to target mRNA in the 3′ UTR, which represses mRNA translation by degradation or blocking [240] (Fig. 2a). Due to their multiple targets, miRNA cover ~ 30% of all mRNA. In cancer, miRNA can function as oncogenes (oncomir), which inhibit tumor suppressor genes or as tumor suppressors, which inhibit oncogenes. MiRNA also promote tumor invasion and metastasis and some miRNA are engaged in CIC maintenance [241, 242].
MicroRNA processing and lncRNA contribution to and interference with miRNA. a Pathway of miRNA processing and mode of action. b Contribution of lncRNA to miRNA provision from intra-exonic or intronic regions being processed accordingly to miRNA genes. c LncRNA contains competive endogenous sequences that correspond to the target sequence for miRNA. By sponging the miRNA, the target mRNA is released from repression. Full names of gene symbols are listed in Table S1
Metastasis-related miRNA significantly up- or downregulated in PaCa and CoCa are summarized in Table S2. For more comprehensive lists, we recommend some recent reviews on PaCa [243] and CoCa [244, 245]. The miRNA list in Table S2 is by no means complete, which appears nearly impossible taking into account that miRNA mostly have multiple targets. Even sorting in oncomirs and tumor suppressor miR should be considered with some caution, opposing activities being reported for several miRNA, which we mention without providing detailed information. Finally, the view on miRNA may profoundly change with progress in deep sequencing (DS) and in silico network analyses, expected to furnish a complete list of miRNA, their targets and their regulation, with lncRNA being one miRNA controlling component.
2.3.2 LncRNA
LncRNA are > 200 bp long. They can be classified according to the genomic location as (1) located away from protein-coding genes (stand alone); (2) antisense transcripts (located on the opposite strand of transcript units); (3) pseudogenes, transcripts having lost protein coding potential by mutations; (4) long intronic ncRNA, the transcript deriving from annotated gene introns; and (5) divergent transcripts from sense and antisense directions of transcript start areas [246].
Mechanistically, lncRNA can interact with RNA, DNA, and proteins. LncRNA can hybridize by specific sequences with DNA or other RNA [247, 248] and can compete with the endogenous RNA network by sharing miRNA [249] due to tandem miRNA response element repeats allowing distinct miRNA or miRNA combination binding [250], targets being released from repression after miRNA decoy [251]. LncRNA can form secondary and tertiary structures that enable complex interactions with proteins. Functional annotation of lncRNA can be based on participating in biological processes at different levels such as chromatin remodeling, histone modification, DNA methylation, transcription, and translation [252]. A common classification sorts lncRNA by (1) recruiting and interacting with proteins, e.g., HOTAIR1 combines with the PRC21-regulating HOXD1 transcription [253]; (2) acting by decoy, e.g., PANDA1 decoys NF-YA1 [254]; (3) corepressor or coregulator activity such as SRA11, an androgen, estrogen, glucocorticoid, and retinoic acid receptor coactivator [255]; (4) miRNA sponging [256]; and (5) being a miRNA host gene [257] (Fig. 2b,c). So far, information is frequently limited to lncRNA acting as competing endogenous (ce)RNA.
The list of lncRNA is rapidly expanding with 565 publications on lncRNA in CoCa and 239 in PaCa. Only most prominent lncRNA and, where available, hints on the molecular mechanisms are presented in TableS3. For the overall, clinically relevant impact, we refer to recent reviews (CoCa: [258,259,260], PaCa: [261, 262], CIC: [263,264,265]).
LncRNA being supposed to regulate proliferation, apoptosis, differentiation, invasion, and metastasis, statistical evaluations of clinical samples accompanied by in silico predictions to obtain hints toward prognosis and therapeutic translation exceed studies searching for targets and the mode of lncRNA action, which are urgently required.
3 Exosomes
Discussing a contribution of CIC markers to tumor progression requires introducing exosomes. Exo are small 40–100 nm vesicles delivered by live cells [266]. Exo distribute throughout the body, being recovered in all body fluids [267]. Exo expressing donor cell-derived components provides an easy accessible diagnostic, prognostic, and therapy-controlling tool [268]. Importantly, Exo components are function-competent [269], message delivery [270], severely modulating target structures and reprogramming target cells in health and disease [271,272,273,274]. Thus, Exo are highly effective intercellular communicators expected to become a powerful therapeutics [275].
3.1 Exosome biogenesis
Exo biogenesis starts with early endosome (EE) formation, deriving from the trans-Golgi network or internalized membrane microdomains [276]. EE are guided toward multivesicular bodies (MVB) by distinct transport machineries [277]. Vesicles, called intraluminal vesicles (ILV), receive their cargo during inward budding into MVB [278,279,280]. Exo plasma loading with protein coding and ncRNA and DNA are nonrandom processes, SGPP11 and diaglycerol being engaged in cargo sorting, and LPAR11, Alix/PDCD6IP1, and HSP701 promote inward budding [281, 282]. Monoubiquitination, acylation, myristoylation, higher order oligomerization, or sphingolipids forming ceramide facilitate protein sorting [173, 283,284,285]. Annexin-II supports RNA sorting [286]. Alternatively, based on the affinity to the raft-like outer layer of the MVB membrane, continuous interaction of cellular RNA with the outer (cytoplasmic) MVB surface may account for ILV incorporation [287]. A zip code in the 3′ UTR and coupling of RISC to a specific EXOmotif (GGAG) of the sorting complex controls miRNA loading by binding to hnRNPA2B11 [279, 288]. The mechanism for selective recruitment of lncRNA is unknown [289]. After ILV inward budding, MVB are guided toward degradation in the proteasome or toward the plasma membrane, trafficking toward the plasma membrane involving microtubules and Rab proteins (rev. in [277, 290]). SNARE1 proteins and SYT1 are engaged in fusion with the plasma membrane (rev. in [276, 277, 290]). The released vesicles are called Exo. The cited excellent reviews provide detailed informations, including open questions and the diversity of Exo derived from individual cells.
3.2 Exosome composition
Though open questions remain on biogenesis pathways, aggravated by differences in biogenesis and the delivery of distinct exosomes by individual cells [291, 292], strong progress was achieved unraveling Exo composition. Exo are buildup by a transmembrane protein-containing lipid bilayer and proteins, mRNA, ncRNA, and DNA in the vesicle lumen. The latter being already introduced, Exo membrane components are briefly outlined.
The Exo lipid envelop contains phosphatidylcholine, phosphatidylethanolamine, phosphatidylinositol, prostaglandins, and lysobisphosphatidic acid and abundantly sphingomyelin, cholesterol, GM31/GRM61, and phosphatidylserine [293], the high phosphatidylserine content allowing differentiating Exo from microvesicles [294]. Though the lipid composition of TEX appears suited for diagnosis [295], only recently developed new methods will allow a precise judgment on exosome lipids, information so far confirming similarity with lipid rafts, a higher lipid order and stability against detergents [294].
Improved mass spectrometry (MS) [296] greatly facilitates Exo protein characterization, so far > 7000 being identified [297]. Structural proteins and proteins involved in vesicle biogenesis and traffic are constitutive Exo components, most abundantly 7–124-fold enriched tetraspanins [298, 299]. Adhesion molecules, proteases, MHC1 molecules, HSPs, TSG1011, Alix, annexins, cytoskeleton proteins, metabolic enzymes, cytosolic signal transduction molecules, and ribosomal proteins, some recruited via their association with biogenesis engaged proteins, also are copious (rev. in [300, 301]). Notably, the all-so-far described CIC markers are recovered in TEX, e.g., MART11 [302], EGFRVIII [303], MDR11 [304], EPCAM [199, 305], MET [306], mutant KRAS [307], CD44v6 [308], Tspan8 [307, 309], α6β4 [310, 311], cld7 [199, 312], LGR5, and CD133 [313, 314]. We interpret this finding that CIC markers may contribute to Exo activities [315, 316].
3.3 Exosome targeting and uptake
Exo targeting depends on their membrane structure and appropriate target ligands, which can be cell or ECM components. Exo binding promotes matrix and cell modulation [276, 297]. Bound Exo can be taken up, which requires different target structures and has distinct consequences for target cells [317].
Integrins, CD44, proteoglycans, and others are engaged in Exo binding to the ECM [318]. Exo proteases, mostly MMPs, IDE1, sialidase, and heparanase, contribute to matrix degradation and remodeling [319, 320]. Exo protease activity is accompanied by matrix-incorporated cytokine, chemokine, and protease liberation [321] and matrix-incorporated cell activation [322]. Exo binding also promotes cell movement through the ECM [323]. Finally, Exo lncRNA adds to ECM remodeling [324].
Only selected target cells bind and take up Exo. Binding frequently involves (tetraspanin-associated) integrins, partners among others for ICAM, FN, LN, proteoglycan-binding lectins, and phosphatidylserine binding TIM41, HAVCR1/TIM11, HAVCR2/TIM31, GAS61, MFGE81, STAB11, ADGRB11, and RAGE/AGER1 [317, 325, 326]. Other binding partners are galectins, selectins, and sialic acid–binding lectins [327,328,329]. We experienced Exo binding being greatly facilitated by clusters of adhesion molecules in both Exo and target cells [330].
Exo uptake proceeds by Exo fusion with the cell membrane [331, 332] or endocytosis, requiring actin cytoskeleton modulation [333]. Modes of uptake include phagocytosis, macropinocytosis, clathrin-dependent endocytosis, and lipid raft and caveolae uptake (rev. in [327]). Phagocytosis proceeds by formation of cup-like extensions, the tips fusing and becoming internalized, and phagocytic markers LAMP11 [334] and TIM4 binding phosphatidylserine facilitate phagocytosis [335]. Macropinocytosis relies on lamellipodia folding back and fusing with the plasma membrane [336]. Most frequent is endocytosis via clathrin-coated pits, rafts, cholesterol- and glycolipid-enriched membrane microdomains, like TEM [337] or caveolae [338]. Uptaken Exo may itinerate [339] but mostly are directed to MVB and are targeted to lysosomes for degradation their content modulating target cells directly or by stimulating target cells’ signaling cascades, transcription, and silencing processes [340,341,342].
3.4 Exosomes and target cell reprogramming
Target cell reprogramming by Exo can be initiated via binding or the uptaken Exo cargo. Target cell modulation by bound Exo relies on signal transduction and/or target cell membrane protein cleavage, e.g., Exo tissue factor binds to the GPCR PAR-21 in EC provoking heparin-binding EGF induction and ERK1/21 activation [343]; Exo HSP20 binding to VEGFR2 activates the VEGFR2 signaling cascade [344] and Exo binding to TRKA11 promotes activation and downstream signaling including FAK1 and Src1 [345]; Exo binding via α5β1 to target cell FN promotes IL1β, which does not require Exo uptake [346]. Due to technical difficulties differentiating binding- and uptake-induced target activation, only few studies explicitly explored binding-induced activation. Taking into account the ample presence of signaling receptors, integrins, CD44 and CAMs, and their ligands on Exo, respectively, target cells, we suggest Exo binding-induced target activation having not received adequate attention [337].
The uptaken Exo content could directly affect the target cell or provide an incentive hit. There are examples demonstrating the transferred Exo content directly changing the target cell. Transferred exo αvβ6 into αvβ6-negative prostate cancer cells localizes to the cell surface, with recipient cell de novo αvβ6 expression being excluded [347]. Also, tumor antigens are processed and loaded into newly synthesized MHC molecules in TEX-loaded dendritic cell (DC) [348]. Also, therapeutically tailored Exo loaded with large amounts of drugs or miRNA or signaling checkpoint inhibitors likely act via content transfer [349,350,351]. However, the naturally available amount of one type of Exo unlikely contains sufficient load for directly modulating targets. First, the small Exo plasma homes limited amounts of proteins and nucleotides [352]; second, TEX from a cloned tumor line distinctly affects tumor cells, fibroblasts, EC, and hematopoietic cells. Our hypothesis on activation of signal transduction and/or transcription/translation being the dominating mode of uptaken Exo activity is supported by DC-Exo uptake strongly affecting the immune synapse [353] and activating or inhibiting B cells, NK, and neutrophils, which also accounts for macrophage (Mϕ-Exo, SC-Exo, and TEX) [272, 354,355,356]. Also arguing for Exo-initiated activation of signal transduction are anterograde and retrograde information transfer by neurological synapses, which accounts for maintaining plasticity under physiological conditions and for pathological protein spread [357, 358]. In brief, the target cell–integrated Exo content can directly account for target modulation or provide an incentive hub.
Having outlined that signaling molecules/transcription factors, ncRNA, Exo, and host cells, are important contributors to CIC maintenance and activity, the question arose on a selective contribution of PaCIC and CoCIC biomarkers in the message exchange between CIC, non-CIC, and nontransformed host tissue [359, 360].
4 Coordinating CIC markers with molecules and exosomes engaged in tumor progression
We will provide some hints supporting coordinating activities of PaCIC and CoCIC biomarkers. In view of many open questions, this trial is doomed to be fragmentary, but worthwhile a shot.
4.1 The net of CIC markers
Asking for a possible connecting role of CIC markers, it should first be mentioned that many of them are associated.
Tspan8 most prominently contributes connecting PaCIC and CoCIC markers. Like all tetraspanins, it is located in TEM, where it associates with other tetraspanins, a multitude of adhesion molecules, proteases, and other molecules. In PaCIC and CoCIC, the Tspan8-associated molecules include the CIC markers CD44v6 [162, 170], α6β1 [361, 362], α6β4 [162, 361], EPCAM, and cld7 [162, 363, 364]. These associations mostly are not direct protein–protein interactions [361, 362], the molecules are not exclusively recovered in association with Tspan8, and some are only associated with Tspan8 in activated cells. This accounts particularly for the association with α6β4 [173, 361], a major contributor of hemidesmosomes in nonactivated cells [365]. It also applies to the TJ component cld7 [187, 196], which becomes recruited to the Tspan8 web only upon palmitoylation [366]. Finally, EPCAM is recruited in association with palmitoylated cld7 [199, 367,368,369], the association being based on a direct protein–protein interaction [366] (see arrows in Fig. 1c,g). The Tspan8 web becomes expanded by the recruitment of cMET and VEGFR2 via CD44v6-bound HGF and VEGF as well as by the association of HA-bound CD44v6 with GPCR (CXCR4) and the CD44v6 association with LRP5/6 [370]. Tspan8 also associates with α3β1, α4β1, and α5β1 [19, 172, 361]. Last but not least, CD44v6 provides a feedback on the Tspan8 net stability by promoting Tspan8 transcription [167].
The striking associations of several PaCIC and CoCIC markers may, in part, explain their contribution to CIC maintenance and activity, all these markers being engaged in transcription factor and signal transduction activation, facilitated by the TEM lipid composition that fosters the attachment of cytosolic molecules.
4.2 The engagement of CIC markers in exosome biogenesis
CIC marker contributions to CIC activities frequently rely on the location in membrane microdomains prone for internalization and EE formation and the loading process during inward budding of ILV.
4.2.1 CIC markers, early endosome formation, and endosome trafficking
Membrane microdomains that are doomed for curvation are in favor of invagination, scission, and fusion to form EE. Prominent membrane domains are caveolae, clathrin-coated pits, TEM, and proteolipid-enriched domains. EE formation by caveolae involves dynamin [371]. Clathrin-coated pit nucleation requires PIP2, AP21, and actin dynamics. Dynamin, actin, and myosins are involved in scission [372]. Highly hydrophobic proteolipids, recovered in detergent-resistant membrane microdomains characterized by solubility in organic solvents, are prone for internalization and EE formation [373, 374]. Invagination of TEM is facilitated by palmitoylation of tetraspanins or associated molecules and involves dynamin and, for some tetraspanins, intersectin 2 [148, 173, 375] (Fig. 3a).
CIC markers and exosome biogenesis. a Exosomes can derive from invaginated membrane microdomains that according to the lipid composition are prone for invagination. These microdomains include low-density lipid-enriched regions, caveolae, clathrin-coated pits, proteolipid-enriched domains, cholesterol-based lipid rafts, and TEM (tetraspanin- and glycolipid-enriched domains). Examples are presented for CIC markers located in internalization prone membrane domains, where TEM are of particular interest as tetraspanins associate with a multitude of proteins including the CIC markers α6β1, α6β4, CD44v6, cld7, and cld7-associated EPCAM). b By fission and scission of invaginated membrane domains, early endosomes (EE) are generated, which are transported toward multivesicular bodies (MVB) frequently involving ESCRT components and mostly guided by rab4 and rab5. However, cholesterol-based lipid raft and TEM-derived EE use ESCRT-independent pathways for trafficking, which for cholesterol-based lipid rafts include ceramide, neutral sphingomyelinase, and S1PR1. The transporters engaged in TEM-derived vesicles remain to be defined. It is important to note that invaginated membrane-derived EE maintain their organization, which includes membrane proteins, attached cytosolic proteins, and the selective lipid composition. So far, there is no evidence of different trafficking routes for MVB toward the plasma membrane or the armament required for exocytosis, after which the endosomes are called exosomes. c Invagination of endosomes, called ILV into MVB, is an energy-dependent process and includes a selection of cytoplasmic proteins, coding RNA, ncRNA, and DNA, which all require distinct supports that are not fully elucidated. Protein recruitment is facilitated by ANXAII and may be supported by cld-associated transporter molecules; miRNA recruitment requires TRPR, hnRNP, and RISC, with evidence for CD44v6 contributing by associated RNA processing proteins. The latter may also account for lncRNA recruitment where rules, however, are largely unknown. The mode of selective DNA recruitment also remains to be explored. d Pa-CIC-TEX is presented showing a selection of prominent protein markers. It should be noted that all CIC markers are recovered in TEX. Full names of protein and gene symbols are listed in Table S1
Early endosome traffic toward MVB mostly requires ESCRT (endosomal sorting complex required for transport). However, in ESCRT-depleted cells, MVB biogenesis is affected, but not absent [376], particularly TEM-derived endosome traffic frequently using ESCRT-independent pathways. Thus, PMEL1 is sorted in an ESCRT-dependent pathway toward MVB and becomes degraded. By sorting along a CD63-dependent pathway, PMEL evades degradation and amyloid fibers are generated [377]. Delivery of antigen-presenting Exo by DC also can follow different pathways of Exo biogenesis. Cognate interaction with antigen-specific CD4+ T cells triggers recruitment of MHCII into CD9 tetraspanin microdomains, where it does not become ubiquitinated and, after incorporation into MVB and Exo delivery, serves for peptide presentation to CD4+ T cells [378]. Uptake of antigen-loaded TEX by DC is strongly promoted by CD81, EE being driven into the MHCII-loading compartment [348]. Also, the intracellular domains of a pair of CD81 molecules form a pocket, which catches cholesterol contributing to distinct tetraspanin EE traffic [379]. Trafficking along the tetraspanin pathway interfering with ubiquination of associated molecules was also described for CD82 and ligand-induced EGFR ubiquitination, with CD82 controlling the activity of the E3 ubiquitin ligase CBL1 [380]. The TEM-guided pathways of EE trafficking also contribute to viral transport. The association of CD63 with syntenin and syntenin-1 interacting protein ALIX affects human papilloma viruses, virus disassembly, and post-uncoating processing being severely impaired in the absence of CD63 or syntenin [381]. The EBV-encoded oncoprotein LMP-11 is transferred via a CD63-dependent and a CD63-independent pathway into Exo, with only CD63-independent Exo biogenesis promoting pronounced MAPK/ERK and NFκB activation [382]. Other examples are the engagement of CD63+ EE in cation transporter cycling. hOCT21-associated CD63 colocalizes with Rab4, engaged in rapid endosome recycling to the plasma membrane, where transport of endosomal hOCT2 to basolateral membranes essentially requires CD63 [383]. In lymphoblastoid B cells, CD38 associates with CD81, HSP70, and LYN1, and the complex being recovered in Exo indicates maintenance during CD81-directed Exo biogenesis [384]. Trafficking of Tspan8–TEM awaits exploration, with preliminary evidence pointing toward CD81-like routing [385] (Fig. 3b).
The PaCIC and CoCIC markers CD133 [100, 386] and CD24 [387], recovered in Exo, are located in internalization prone rafts [387, 388]. Similar to TEM-guided EE trafficking, CD133 follows an ESCRT-independent pathway. EE traffic requiring ceramide and NSM1 relies on S1PR11 [389], confirmed by demonstrating that SNCA1, which causes expulsion of S1PR1 from lipid rafts, reduces MVB formation [390] (Fig. 3b).
Briefly, ESCRT-independent EE recruitment into MVB accounts for different types of lipid-enriched membrane microdomains, mainly demonstrated for TEM and classical rafts. These distinct pathways frequently circumvent ubiquitination and subsequent guidance to lysosomes. Notably, the microdomains are conserved during Exo biogenesis and all components including attached cytoplasmic proteins are recovered in Exo. The components required for ESCRT-independent EE traffic are not fully elucidated. Filling this gap will be important aiming to eliminate disease-promoting Exo. Finally, to our knowledge, ESCRT-independent membrane microdomain-guided Exo biogenesis does not influence the traffic of MVB toward the plasma membrane and the Exo release.
4.2.2 CIC markers and endosome loading
The selective processes of ILV loading are not fully explored. Nonetheless, it is worthwhile to remember that the interaction of RNA with the outer (cytoplasmic) surface of MVB may account for RNA recruitment [277], possibly also valid for protein recruitment. Furthermore, the special lipid composition of invagination-prone membrane domains supports the attachment of a large range of cytosolic signaling molecules, fostered by palmitoylation and myristoylation [391,392,393].
These modes of recruitment explain the high enrichment of tetraspanins and associated transmembrane and cytosolic molecules, in view of the recruitment of mRNA, miRNA, DNA, and the RNA splicing machinery (rev. in [277, 374, 394,395,396]), another observation that requires notion. Upon precipitating Exo lysates with anti-CD44v6, anti-CD44s, anti-EPCAM, anti-cld7, or anti-Tspan8, a wide range of RNA processing molecules including RNA splicing and alternative splicing and miRNA processing components only co-immunoprecipitated with CD44v6. As anti-Tspan8 precipitated mostly the panel of proteins recovered in TEM, the selective co-immunoprecipitation of CD44v6 with the mRNA processing machinery points toward an activity of cytoplasmic CD44v6 that—to our knowledge—did not receive attention, but could well contribute to Exo-promoted activities. Exo were described to process miRNA [394, 397]. They contain Dicer, AGO21, and TRBP1, which also are recovered in late endosomes [394, 397]. Furthermore, CD431 is guiding the RISC loading complex into late endosomes in breast cancer [394]. Thus, we speculate that CD44v6 performs this task in PaCa and CoCa, with preliminary evidence showing that CD44v6 contributes to miRNA and lncRNA loading. Palmitoylation-deficient cld7 precipitates rather exclusively transporter molecules or lipid-processing–engaged components, which would be in line with cld7-TEX contributing to lipid metabolism and ion transport (Fig. 3c).
In brief, protein incorporation into ILV may partly rely on integration of invagination-prone membrane domains, which includes in PaCIC and CoCIC particularly Tspan8-associated proteins. A potential contribution of TJ-derived cld7 in the recruitment of transporter proteins requires confirmation. Though less is known on coding and noncoding RNA recruitment, there is evidence for a decisive role of CD44v6. If confirmed, this would add a new dimension to the multiple metastasis-promoting activities of CD44v6. However, there is a need for further explorations of ILV loading, and new experimental tools for identifying RNA-associated motifs will accelerate progress in the future [398]. Last but not least, all PaCIC and CoCIC biomarkers are abundantly recovered on TEX (Fig. 3d).
4.3 CIC markers, exosome targeting, and exosome uptake
Exosomes are recovered in the ECM mesh [399]. The major components of the ECM are collagen, LN, and HA. CD44v6 binds HA [330], coll, and FN. Tspan8-associated α3β1 and α6β4 bind to collagens and LN [173, 310, 400], with the Tspan8–α6β4 complex particularly facilitating LN332-rich basement membrane attachment [401] (Fig. 4a).
CIC markers and exosome targeting. Exosomes distribute through the body and bind to selected targets. a Binding to the ECM is greatly facilitated by Tspan8 and associated integrins, which bind to collagens, FN, and LN; CD44v6 accounts predominantly to HA binding. b Target cell binding is facilitated by phosphatidylserine receptors which bind to phosphatidylserine in the Exo membrane. CD44v6 may bind to CD6, and TEM-derived Exo have a whole panel of potential target proteins. Exo binding can be followed by fusion with the plasma membrane or by uptake via macropinocytosis or GEM invagination. After uptake, the Exo membrane is digested and the content is released. Full names of protein symbols are listed in Table S1
With regard to cell binding, Exo tetraspanins play a decisive role, which relies on clustering associated molecules to increase the strength of binding either of individual or a range of tetraspanin partners that varies according to the Exo donor cell [375]. For PaCIC and CoCIC, Tspan8 clusters are dominating with likely some contribution of CD151 clusters [156]. The Tspan8 association with integrins is crucial for the contact with target cells. Thus, α6β4 binds cells in the premetastatic niche of the lung, whereas in the liver, integrin αvβ5 binding is dominant [310]. Binding to EC is promoted by α4β1 and α5β1 [171, 172]. Other Tspan8 partners like CD44v6 may contribute to selectin binding [402]. As only monomeric EPCAM associates with Tspan8, a possible contribution of EPCAM to targeting remains to be explored. Literature search did not provide hints toward an engagement of Tspan8–Exo in the crosstalk with leukocytes, which is in line with our experience [403]. Instead, CD151 and CP9 contribute particularly to platelet targeting [404, 405]. Finally, tailoring nontransformed cell Exo with Tspan8 greatly facilitates target cell binding [406, 407].
In PaCa and CoCa, target cell-bound Tspan8–Exo are readily taken up. Antibody blocking and proteome analysis after pulldown revealed preferential Tspan8–Exo uptake by molecules clustered in internalization-prone membrane domains. However, there is no evidence for a particular uptake by TEM [363], i.e., uptake by Tspan8-kd and Tspan8-ko cells is unaltered [405, 406]. TJ also were reported to exchange membrane particles [408], which was confirmed for the exchange of TJ components of DC and monocytes with lung epithelial cells [409]. Also, lymphatic vessels express cld7 at a high level [410] and TEX promote lymphatic vessel growth [411, 412]. Thus, we speculate that cld7 may contribute to CIC-TEX uptake. TJ are continuously remodeled [413], and proteins being either reshuttled or—evading degradation—are recovered in TEM-independent Exo [199]. A contribution of these Tspan8-devoid cld7–Exo to uptake by target cells remains to be experimentally confirmed (Fig. 4b). We are not aware of studies evaluating a possible contribution of the CIC markers CD133, CD24, and LGR5 to Exo targeting and uptake.
Taken together, the Pa- and CoCIC marker Tspan8 takes a prominent role in TEX targeting and uptake, guiding clustered integrins and CD44v6 toward their ligands, but target cell Tspan8 does not or not significantly add to uptake. For other TEX CIC markers, a contribution to targeting/uptake remains to be unraveled.
4.4 CIC markers and exosome message delivery
Exosome message delivery is hotly disputed as a possibly most powerful therapeutic tool in autoimmune disease, regenerative medicine, and cancer including cancer progression. There is an abundance of excellent reviews dealing with these diverse therapeutic options. Even with the restriction toward Pa- and CoCIC markers, we cannot cover the field and apologize for not citing outstanding publications. Instead, we aim to give a short overview on two topics: the contribution of CIC markers to modulation of the ECM and to changing expression profiles and signaling in TEX target cells.
4.4.1 CIC markers and modulation of the ECM
Being receptors for matrix proteins, integrins and CD44v6 are central components in TEX-promoted matrix protein binding. After binding, the process of matrix modulation will be initiated. So far, only few reports are explicitly concerned about the ECM, rather than the incorporated cells.
First to note, Exo can be coated with HA, which becomes deposited with the Exo in the ECM facilitating migrating tumor cell settlement by CD44 binding [319, 414]. Exo also carry FN, supporting docking to integrins and promoting tumor cell motility [323, 415], where Exo–FN binding depends on heparinsulfate binding annexins A2 and A6. These Exo also express the serine protease DPP41, with the activity in ECM modulation being not yet fully defined [416]. However, DPP4 is associated with Tspan8 and could contribute to ECM-incorporated protein digestion. Of particular interest with respect to ECM modulation is the Exo protease profile, which includes especially membrane-bound MMP14; ADAM10 and 171 and ADAMTS1, like ADAMTS51; and glucuronidases/sialidases like NEU11 [324, 417,418,419,420], elegantly reviewed by Bandari et al. [421]. To give a few examples, LOXL21 catalyzes the first step of collagen cross-linking [420]; MMP14 contributes to FN and VN1 degradation [422, 423]; IDE, routed via detergent-resistant membrane complexes into Exo, degrades matrix-deposited proteins, most well described for amyloid [424]; Exo heparanase degrades heparin sulfate in the ECM affecting the heparin-rich basement membrane [421].
With regard to PaCIC and CoCIC markers in matrix remodeling, the deposition of TEX-attached CD44-linked HA and CD44- or integrin-linked FN should be taken into account. For restructuring the matrix, the link between proteases and Tspan8/CD44v6 is important affecting collagen cross-linking as well as collagen and LN degradation and via CD44-linked hyaluronidase HA degradation [425]. The contribution of Tspan8-associated α6β4 binding to the LN332-rich basal membrane and of heparanase to the heparin-rich basement membrane may be prominent in promoting tumor cell migration. Tumor cell migration and settlement in distant organs receives additional support by liberation of growth factors, chemokines, and proteases deposited in the ECM [321] (Fig. 5a).
CIC markers and target modulation. CIC-Exo binding and uptake severely affects the targets a after ECM binding the Exo protease becomes active, which include (membrane-bound) MMP, ADAM, ADAMTS, uPAR, hyaluronidases, glucuronidases/sialidases, and IDE. Some of these proteases are associated with tetraspanins or integrins or CD44v6, which facilitates their concentration at the binding site. Besides collagens, FN, LN, HA, and heparansulfate, Exo also degrade deposited proteins and support activation of incorporated pre-proteases. Finally, deposited cytokines and chemokines become liberated and contribute to tumor cell activation, thereby a path for migrating tumor cells is generated and EC as well as tumor cells becomes activated. b Target cell activation by CIC markers expressed in bound or uptaken Exo is more difficult to decipher. A few examples are shown. CD24 binding to cortactin supports Notch stabilization and Nanog regulation. CD133 may contribute to NOTCH activation and promotes TIMP2 upregulation. Cld7 was reported to promote FASN, transporter, VEGFR3, and PDGFRB expression by not yet fully clarified pathways. Monomeric EPCAM promotes myosin and Mybbp1A upregulation. There is a large range of ligands for Tspan8-associated integrins and CD44v6, which could promote activation of multiple signaling pathways. CD44v6, additionally could bind to GPCR5B, which supports cMET activation; GLI1 becomes activated via EREG binding to the EGFR and promotes EMT gene, VEGFA, FGF, and IL8 expression. Transfer of OPN from CD44v6 to αvβ3 promotes MAPK, ERK1/2, and PI3K activation-supporting EGHR activation and CD44 expression. c For the crosstalk between miRNA and PaCIC and CoCIC markers, a few examples of an experimentally proven impact on metastasis are shown. d LncRNA also is engaged in CIC marker expression. So far, mostly release from repression by sponging miRNA was reported. However, there is increasing evidence that lncRNA predominantly act via chromatin modifications and regulation of transcription. Full names of protein symbols are listed in Table S1
4.4.2 CIC markers and exosome target cell activation and reprogramming
Four topics related to CIC marker engagement in TEX-promoted target cell modulation appear of particular interest: gene transcription and silencing, activation of signaling cascades, acquisition of a motile phenotype, and apoptosis resistance. Great effort is taken to explore these topics, which may in part be linked. Open questions surpassing available answers and few examples are given separated according to being protein- or ncRNA-mediated. We also will consider the ongoing dispute on the uptaken Exo content directly accounting for target cell modulation or acting as a hub [271, 426, 427].
CIC markers and exosome binding-induced signal transduction
The contribution of Exo-CIC markers on target modulation is best explored for tetraspanin-associated integrins and CD44/CD44v6. TEX tetraspanin-associated integrins account for premetastatic niche formation, with distinct integrins determining the organ specificity. This also holds for EC/EC progenitors, which are targeted by Tspan8–CD49d/CD49e Exo that induce CXCL51, MIF1, vWF1, and CCR11 mRNA upregulation. Induction of VEGFR2 required support by external VEGFA (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE18812). Upregulated mRNA were recovered after 1–5 days, which points toward induction of transcription and excludes upregulation directly relying on the transferred TEX content that expression is low in TEX [172]. Similarly, Tspan8–CD49f or Tspan8–CD104 TEX, but not Tspan8kd–CD49f or Tspan8kd–CD104 TEX, distinctly affect gene expression differing depending on the target cell. In fibroblasts, mostly proteases (ADAM17, MMP14, TIMP1,21) become upregulated. Instead, EC respond with upregulation of FGF, VEGFR1, and VEGFR2; BMC (bone marrow cells) with upregulation of TNF1 and STAT4 activation; LNC (lymph node cells) with upregulated TNF, TGFB1, and FoxP31; and tumor cells with vim1, Slug, and Snail expression. The target cell-dependent distinct responses argue for Exo providing an initiating hub [406]. Not taking into account tetraspanin expression, CD104–vinculin TEX were described to cope with resistance toward Taxol, a complex diterpene alkaloid [406]. This may rely on CD104-bound plectin recruitment into TEX, as blocking the transfer of plectin into Exo interfered with PaCa growth [428].
The PaCIC and CoCIC cell and TEX marker CD44/CD44v6 strongly force target cell reprogramming. Coculture of PaCa cells with CD44v6-competent, but not CD44v6kd TEX promotes, besides others, upregulation of tumor progression-engaged MMP3, ADAMTS-1, ADAMTS-5, ADAMTS-8, several chemokines, proteoglycan 4, COX21, SOD21, MDR1, PLA2G2A1, SSB11, FABP31, and MYH111 mRNA (ENA database accession no.: PRJEB25446), and for some of these mRNA, the direct transfer from TEX is excluded [429]. mRNA DS of human pancreatic non-CIC after coculture with CIC-TEX confirmed a compelling CD44v6-dependent impact. Two features should be particularly mentioned. First, the majority of upregulated mRNA were engaged in translation and splicing, followed by signaling components with a preponderance of RTK and EMT TF. Second, drug transporters were most strikingly upregulated. Most of these genes, being not affected in a Tspan8kd PaCa line, point toward the engagement of TEM-independent CD44v6 (ENA database accession no.: PRJEB25446). How can these strong effects be explained? Target cell modulation by CIC-CD44v6 was recently reviewed [430]. Thus, we will focus on the crosstalk between CD44v6-TEX and target cells. First, via cytokine binding, CD44v6 becomes linked to several RTK, which could account for initiation of signaling cascades. It was suggested that EGFR, ERBB21, and INSR1 are transferred from TEX to target cells, where they initiate MAPK signaling pathway activation [431]. TEX EGFR also is transferred into liver stroma cells, where HGF becomes activated, binds tumor cells, and facilitates tumor cell settling in the liver [432]. A similar pathway of activation was suggested accounting for bone metastasis [433]. Alternatively, TEX EREG1 binds to target cell EGFR inducing EMT by regulating GLI11 and increasing VEGFA, FGF, and IL8 expression [434]. It also was reported that cMET activation does not proceed by the transfer from TEX, but via orphan GPC5B1 that promotes MAPK and together with HGF cMET activation [435]. This suggestion appears very attractive and fits to the results of several studies, reporting on TEX-initiated target cell activation relying on the delivery of chemokines and cytokines. Thus, CXCR4 associates with CD44 upon HA cross-linking, which promotes signaling by CXCL12. Taking the HA coat of Exo, the delivery of Exo CXCL12 provides a convincing mode of CIC-Exo–initiated signaling [436], although activation of lymphatic EC by the direct transfer of Exo CXCR4 was also described [412]. A further mode of activation could rely on the release of Rantes/CCL51 from Exo that directly binds CD44 and promotes MAPK cascade activation [437]. Activation of CD44 signaling was also described for OPN and confirmed for Exo OPN, which proceeds via αvβ3 binding [438]. Finally, Exo tissue factor binding to its GPCR F2R1 promotes E-selectin upregulation and IL8 secretion [439].
Another PaCIC and CoCIC marker that is recovered in TEX is CD24. Its ligands are selectins, NCAM11, and CNTN11, a GPI-anchored member of the Ig-superfamily. TEX CD24 signals via contactin, promoting activation of the MAPK pathway. It is also engaged in EMT by NOTCH1 stabilization via p38MAPK and in STAT3-mediated NANOG regulation [440,441,442].
CD133, abundantly recovered in CIC-TEX, suppresses the RET1 tyrosine kinase via p38MAPK and PI3K signaling and regulates TIMP2 expression. A soluble form of EC-derived Jagged1 promotes colocalization of CD133 and Notch accompanied by NOTCH activation [231, 443, 444]. Whether these activities also account for TEX-CD133 remains to be explored.
Claudin7, recovered in two distinct Exo populations [199], targeting cell communication became of special interest. Proteome profiling of cells and TEX revealed that non-TEM–derived cld7 is dominating in TEX, and proteome analysis of immunoprecipitates uncovered that in cells and TEX-expressing TJ-derived and GEM-derived cld7, distinct components were prevalent with an abundance of proteins engaged in fatty acid biosynthesis/metabolism and TJ organization in cells expressing only TJ-derived cld7. In TJ-derived TEX, four independent network clusters related to TJ assembly, endocytosis, proteasome degradation, and a fourth, larger cluster of proteins engaged in DNA replication, RNA transport, AA synthesis, and metabolism were seen. However, after transfer of TJ-derived cld7-TEX, solely a pronounced upregulation of VEGFR3 and PDGFRB appeared to be linked to cld7, reinforcing the suggested contribution of TEX-cld7 to lymphangiogenesis [197].
Concomitantly with TEX-cld7, we searched for cld7-independent TEX-EPCAM-promoted activities. There were only 10 proteins not recovered in EPCAMkd-TEX, where the absence of the corepressor/coactivator MYBBP1A1 and of the actin-dependent motor proteins Myh10 and Myh14, engaged in cytokinesis, may be mentioned [445, 446]. Although there are a considerable number of proteins co-immunoprecipitated with TEX-EPCAM independent of the presence of cld7, signaling pathway analysis did not reveal hints toward EPCAM selectivity. Thus, whether and by which means TEX-EPCAM communicates with target cells is not yet answered.
There remains the CIC marker LGR5 that is engaged in Wnt signaling, where we did not succeed finding relevant notice on its activity in TEX (Fig. 5b).
Not related to prominent PaCIC- and CoCIC-TEX markers, we mention two additional aspects. Depending on TEX TGFβ1 or TGFβ2 and supported by TEX PDGF, FGF, and IL6, TEX promote the conversion of fibroblasts into CAF [447], which may involve activation of SMAD signaling [448]. This impact of TEX on the tumor surrounding will have a rebound on CIC. Finally, Exo display intrinsic metabolic activities. They carry lactate, PGE1, LDH1, pyruvate, and monocarboxylate transporters, implicated in fatty acid synthesis and cholesterol metabolism. They also can synthesize ATP by glycolysis [449,450,451]. The transfer of these lipids and glucose and particularly lipid metabolism-regulating components severely affects the metabolic state of target cells [452,453,454].
To summarize, PaCIC- and CoCIC-TEX marker-binding and transfer into target cells contributes to tumor progression, which includes tumor, stromal, endothelial, and hematopoietic lineage cells. The engagement of Tspan8 predominantly relies on its association with integrins, with contributions of α3β1, α4/α5β1, α6β4, and αvβ3 being well documented. Tspan8 also adds by associating with proteases. CD44v6 takes a leading role by engagement in RTK and GPCR associations as well as by the association with LRP6, strengthening Wnt signaling. The CD44/CD44v6 engagement is fostered by HA and FN as well as selectin binding. Cld7 mainly contributes to lymphangiogenesis and modulating lipid metabolism. Contributions of TEX CD24, CD133, and EPCAM remain to be substantiated.
Though pathways whereby CIC-TEX markers modulate targets are not fully elaborated, binding-initiated signal transduction plays a dominant role, and reports on activation of signaling cascades and in silico analyses depicting the connectivity of molecules in networks and between networks support this interpretation. This does not question exceptions, where the transfer from TEX into targets is unequivocally demonstrated. Nonetheless, TEX-transferred and TEX-induced proteins may act as initiators. This possibly also accounts for the transfer of coding and ncRNA.
CIC markers and exosome transfer of noncoding RNA
With rapidly increasing evidence on the importance of Exo ncRNA, particularly miRNA and lncRNA, a discussion on a possible connection to CIC-TEX markers should not be missed. However, three major unsolved or partially solved issues prohibit a round answer. Issue 1: There is no explanation on the abundance of lncRNA in Exo compared to cells, which differs from proteins and miRNA; only for few lncRNA the functional relevance was tackled, frequently restricted to lncRNA activity as miRNA sponge; it is also unknown whether Exo lncRNA is transported with/without their targets into acceptor cells [455]. Issue 2: There are hints that miRNA may be processed within Exo. Though the armament appears to be available, the functional importance of an intra-Exo processing and the consequences remain to be unraveled. Furthermore, many miRNA having several to > 100 targets, an assignment to target cell mRNA remaining sporadic and even knowing that a given miRNA targets one of the CIC markers, it is debatable whether the amount of transferred miRNA suffices affecting the activity of the respective CIC marker or associated/linked molecules. Third, as already mentioned, will the transferred Exo ncRNA be the actual effector or an initiator? Facing the limited state of knowledge, we only can give some examples on the link between transferred ncRNA and CIC markers.
Regarding CD44vv6, its engagement in miRNA recruitment into ILV was outlined [429]. Furthermore, expression of several miRNA and CD44 or CD44-associated molecules is linked. miR-146-5p targets ZNF831, resulting in pronounced migration and invasion and Frizzled6 and CD44v6-associated LRP6 upregulation [456]. Overexpression of the CD44 3′ UTR promotes motility, invasion, and metastasis. miR-328 targets the CD44 3′ UTR and COL1A11; miR-491, miR-512-3p, and miR-671 target CD44 and FN. The authors speculated that ECM protein synthesis could be corrected by provision of CD44 3′ UTR [457]. miR-34a targeting the 3′ UTR of CD44 prevents prostate Ca metastasis [458]; mir-218-4 targeting CD44-ROCK1 affects invasion. It is downregulated in squamous cell carcinoma [459]; miR-328-3p, upregulated in triple-negative breast cancer, targets AR1, which controls the expression of CD44 via miRNA-dependent and miRNA-independent pathways [460]; CD44-associated cMet, MMP2, and MMP9 become regulated by miR-340, which suppresses invasion and metastasis [461]. Exo miR-520c-3p targets CD44, which is accompanied by reduced extravillous trophoblast invasion [462].
CD133+ TEX from melanoma and CoCa contain many ESCRT and ESCRT-associated proteins, selectively harbor several miRNA, and are enriched in tumor progression-promoting proteins such as CD44, MAP2K41, GTP-binding proteins, ADAM10, and Annexin A2 as well as tetraspanins. The proteolipid assembly resembles that of TEM. The authors demonstrate that CD133+ TEX uptake, including the selectively enriched miRNA, strengthens metastatic potential [271]. On the other hand, miR-142-3p binds CD133, LGR5, and ABCG2 acting as a tumor suppressor and being downregulated in CoCa [463].
LGR5 also becomes regulated by miRNA. miR-363, downregulated in CoCa, targets GATA6, which enhances LGR5 expression [464]. LGR5 is also targeted by miR-100, downregulated in CoCa [465] (Fig. 5c).
EPCAM+ CoCIC-TEX also are enriched in selective miRNA, where miR-16-5p, miR-23a-3p, miR-23b-3p, miR-27a-3p, miR-27b-3p, miR-30b-5p, miR-30c-5p, and miR-222-3p recovery decreases after tumor excision, which supports these miRNA being tumor-derived and contributing to tumor progression [466]. Furthermore, a study separately collecting miRNA from extracellular vesicles (EV) and EPCAM+ TEX of a CoCa line revealed distinctly enriched miRNA clusters in EV and TEX with selectivity of some miRNA enrichment in EPCAM+ TEX [467].
There are also hints on the engagement of lncRNA in regulating PaCIC and CoCIC marker-promoted activities.
The lncRNA GAPLINC1 contributes to CD44-dependent invasiveness. Upregulation, associated with shorter survival in gastric cancer, correlates with CD44 expression, with CD44 targeting miR-211-3p being sponged by GAPLINC [468]. uc002kmd.1 is highly expressed in CoCa. It regulates CD44 by competing with miR-211-3p, which affects CoCa growth in xenogeneic mice [469]. DGCR5 is overexpressed in NSCLC (nonsmall cell lung carcinoma)-CIC. It targets miR-330-5p, which releases CD44 from repression [470].
LncRNA are also engaged in EPCAM expression. Upregulated LinC00152/CYTOR1 in HCC (hepatocellular carcinoma) promotes proliferation and tumor growth in vivo and in vitro. It binds to the EPCAM promoter promoting MTOR1 pathway activation [471]. TINCR1 is downregulated in CoCa and expression inversely correlates with metastasis. Pulldown assays revealed that TINCR binds EPCAM RNA, with TINCR downregulation being associated with the release of EPCAM–ICD and Wnt–β-catenin signaling [472]. TCF7 promotes glioma cell self-renewal, accompanied by EPCAM upregulation, which relies on TCF7 sponging miR-200c [473]. A clinical study reported upregulated BCYRN11 promoting gastric cancer progression and being accompanied by EPCAM upregulation. The underlying mechanism remains to be explored [474].
CASC151 promotes gastric cancer metastasis. It sponges miR-4310 accompanied by the release of LGR5 from repression [475].
KRTAP5-AS11 binds miR-596 and miR-3620-3p and TUBB2A1 binds miR-3620p in gastric cancer. This is accompanied by cld4 release from repression and promotes upregulation of EMT genes [476]. PlncRNA1/CBR3-AS11 and miR-34c are engaged in regulating TJ proteins in inflammatory bowel disease. PlncRNA1 targets miR-34c thereby releasing the miR-34c target MAZ1 from repression, which regulates ZO11 and occludin expression, and PlncRNA1 strongly mitigates inflammation-induced TJ dysfunction. As cld7 is highly expressed in the gastrointestinal tract [477], we suggest an additional involvement of cld7 that remains to be approved (Fig. 5d).
Finally, a comprehensive study in CoCa revealed 1028 lncRNA selectively enriched in TEX, with the co-existence of RNU1-1 and RNU1-21 in TEX suggesting a possible link to recipient cell splicing events [478].
DS of PaCIC-TEX and of the corresponding Tspan8kd and CD44v6kd line revealed that from 142 lncRNA, 37 displayed a higher and 28 a lower signal strength in Tspan8kd cells, whereas in CD44v6kd cells, 73 lncRNA were not and 12 lncRNA were detected at a lower level, with only 23 lncRNA being recovered at a higher level. The finding reinforces the contribution of CD44v6 to ncRNA loading during Exo biogenesis.
Expression of the three lncRNA with the highest score in TEX, LRRC75A-AS1, ZFAS11, and SNHG81, was strikingly reduced in CD44v6kd cells. LRRC75A-AS1 was described to be a prognostic factor of AML. Network analysis revealed that it mostly affects TP53 and ETV61 [479]. ZFAS1 represents a snoRNA host gene that produces a ncRNA. Increased ZFAS1 expression or locus amplification is associated with metastasis. The transcript regulates the expression of differentiation involved genes. It may act as a molecular sponge by directly interacting with miR-484. ZFAS1 also acts as ceRNA for miR-486, which promotes osteosarcoma progression. ZFAS1 enhances Wnt/β-catenin signaling with multiple effects on proliferation, EMT marker, and protease expression in gastric cancer [480,481,482]. SNHG8 affects several gastric cancer-specific pathways and targets EBV (Epstein–Barr virus) genes, which are largely enlaced. In line with this report, SNHG8 transcript levels are significantly higher in cultured EBV-associated gastric cancer cells than in normal gastric mucosal cells or EBV-negative gastric cancer cells. A SNHG8kd arrests the cell cycle in the G0/G1 phase, inhibits proliferation and colony formation, and suppresses tumor growth in vivo. In endometrial cancer, too, SNHG8 expression is significantly increased. It sponges miR-152, which targets cMET. Though not directly related to cancer, it is interesting to note that in muscle SC transcription of SNHG8 and the lncRNA GM26917 are regulated by FoxM1 that binds to their promoters [473, 483, 484].
The tumor suppressors lncRNA HOTAIRM1, LINC-PINT1, and SLC25A25-AS11 were recovered at a very low level in PaCIC-TEX and not in CD44v6kd cells. HOTAIRM is downregulated in CoCa and suggested to be a promising candidate for diagnosis. So far, most studies were concerned about its role in myelogenesis, where it acts as a ceRNA for miR-20a/106b and miR-125b, promoting ULK11, E2F11, and DRAM21 release from repression. It also regulates oncoprotein degradation [485, 486]. LINC-PINT is downregulated in multiple types of cancer and acts as a tumor suppressor by reducing the invasive phenotype of cancer cells. A highly conserved sequence element specifically interacts with PRC2, necessary for the LINC-PINT-dependent repression of a pro-invasion signature of genes regulated by EGR11. The authors suggest that LINC-PINT by proximity of co-regulated genomic loci affects the availability of free PRC2 [487]. SLC25A25-AS1 overexpression significantly inhibits proliferation and colony formation in CoCa lines and downregulation enhances chemoresistance and promotes EMT, accompanied by ERK and p38 signaling pathway activation [488]. In PaCa, TEX expression is low and reduced compared to cells.
These and additional lncRNA in PaCIC-TEX are listed in TableS4 including available information on functional relevance. Besides the most striking reduction of lncRNA in CD44v6kd cells and a relative abundance of intronic lncRNA, there was a dominance of lncRNA that cooperate with chromatin modifiers and affect transcription.
Last but not least, lncRNA regulation in PaCa and CoCa can also proceed independent of CIC markers. LINC-ROR1 sponges miR-145 and miR-205, regulating Oct4, Sox2, and Nanog [489]. IL22 induces H19 lncRNA1via STAT3 signaling. H19 promoting proliferation binds a cluster of proliferation inhibiting miRNA (let-7, mir-34a). It inhibits p53 but promotes MycN and FOXM1, which are targets of the H19-regulated miRNA [490].
Taken together, there is evidence for a crosstalk between metastasis-promoting CIC markers and cellular as well as TEX miRNA and lncRNA. We expect that further progress in lncRNA activities will strengthen information on the network between lncRNA, miRNA, and CIC-TEX markers. The still hypothetical engagement of CD44v6 in miRNA and lncRNA recruitment into TEX could add another knot to the engagement of CIC markers into tumor progression. Without striving for an irreplaceable position of CIC-TEX markers in tumor progression, their central and networking activity is beyond question.
5 Conclusions, open questions, and outlook
It is well appreciated that CIC markers are suited for CIC enrichment and can serve for non-/minimally invasive diagnosis, prognosis, and therapy response control due to recovery of TEX in body fluids. This review aimed to collect available information on the advantage for CIC and CIC-TEX in tumor progression by expressing these biomarkers.
-
I.
At the present state of knowledge, CIC mostly profit from CD44v6 expression as it recruits several ligands for RTK, which become activated via the association of signaling molecules/cytoskeletal linker proteins with the CD44ICD. LRP5 binding facilitates Wnt signaling pathway activation. The major CD44 ligand, HA, adds to the range of CIC-supporting CD44v6 activities at multiple levels facilitating binding and tumor cell migration, receptor cross-linking, and uptake. It contributes to inflammatory response induction and affects the metabolism after uptake. The CD44/CD44v6 linkage to ABC transporters, strongly supporting drug resistance, is of high clinical relevance. The engagement in hyaluronan synthase and hyaluronidase transcription and stabilization enforces matrix remodeling. CIC do not directly profit from Tspan8 expression. However, via the association with integrins, predominantly α6β4, CIC gain in motility and the association with proteases promote invasion. LGR5 is important in transferring Wnt signaling. Whether EPCAM promotes metastasis or rather oncogenesis remains an open question. A significant contribution of cellular cld7 and CD133 to tumor progression also has not been unequivocally demonstrated.
-
II.
Looking at exosomes, we are confronted with a different scenario. Tpsan8 plays a dominant role in TEX biogenesis as it contributes to EE formation, guiding all associated/loosely attached molecules into EE. In addition, tetraspanins use a distinct, ESCRT-independent trafficking route toward MVB, where due to monoubiquitination, endosome proteins are largely deviated from lysosomes and degradation. The associated CIC markers α6β4, CD44v6, cld7, and EPCAM profit from this Tspan8 activity. All other PaCIC and CoCIC markers are also located in membrane domains preponed for invagination and EE formation, but are not linked to one another.
The small cytoplasm of ILV is loaded during inward budding. There is evidence for TEM-independent CD44v6 being associated with several components of the RNA processing machinery, thereby contributing to ILV loading. Whether the recruited RNA processing machinery suffices for RNA processing is still disputed. TEM-independent cld7 is associated particularly with transporter molecules and lipid metabolism components and apparently has a share in recruitment into ILV. There is no evidence for CIC markers adding to MVB transport toward the cell membrane and Exo release.
-
III.
TEX-TEM contribute to target selection, which is involved in PaCIC and CoCIC Tspan8-associated integrins and CD44v6. The power of Tspan8 builds on offering densely packed receptors to target cell ligands, which decreases the threshold level and facilitates uptake. Target cell ligands mostly are located in synapses also prone for internalization, but target cell tetraspanins provide no special contribution to TEX uptake. A share of TEM-independent cld7, LGR5, and CD133 to TEX uptake remains to be explored.
-
IV.
Exo can be captured by the ECM. For CIC markers, this relies on the HA coat, on CD44 binding to FN and HA and on collagen- and LN-binding Tspan8-associated integrins. ECM binding frequently initiates ECM remodeling, which is provoked by Tspan8- and CD44v6-associated proteases. ECM remodeling is important in creating a path for migrating tumor cells. Liberation of deposited cytokines, chemokines, and matrix degradation products facilitates CIC settlement and activation, angiogenesis, and leukocyte recruitment.
-
V.
CIC-TEX binding and uptake by non-CIC and nontransformed cells, which can be of endoderm, mesoderm, or ectoderm origin, account for induction of a more aggressive phenotype including EMT in non-CIC, (lymph)angiogenesis, niche preparation, and deviation of hematopoietic cell maturation. Induction of signaling pathway activation includes TF, RTK, GPCR, membrane-integrated adhesion molecules, and proteases. Contributions were described for all PaCIC- and CoCIC-TEX markers. CD44v6, due to its multiple ligands and engagement in several receptor-initiated pathways, plays a dominant role. The situation is less clearcut for uptaken and digested TEX, with the TEX content still being disputed to act as effector or initiator. Different target cells showing distinct responses to the same TEX preparation argues in favor of the latter. Irrespective of the mode of transferred TEX activity, the strongest response was seen in CD44v6-deficient targets, followed by cld7-deficient targets, whereas EPCAM- and Tspan8-deficient targets displayed only few Tspan8- or EPCAM-specific responses. The rebound of CD44v6-deficient targets is dominated by activation of signal transduction, followed by transporter activation and transcription/translation modulation. This accounted for mRNA and miRNA, and lncRNA requires further evaluation. The weaker reaction of cld7kd cells is ruled at the mRNA/protein level by signaling, followed by adhesion and structural molecules. Few selective changes in miRNA were mostly related to trafficking molecules. Many of the cited publications approving these effects in vivo still appear like having touched the tip of an (nonmelting) iceberg, which leads to the point of open questions.
-
VI.
The TEX composition is quite well explored, and trafficking pathways of CIC markers located in lipid-enriched microdomains require further evaluation. Loading of ILV, too, needs additional scrutinized elaboration particularly with regard to the CIC marker contribution to miRNA and lncRNA loading. Required techniques being established will facilitate achieving the goal.
-
VII.
In PaCIC- and CoCIC-TEX and possibly additional gastrointestinal CIC-TEX, particularly, the tetraspanin Tspan8 plays a dominating role in target selection. A contribution of CD133, LGR5, and CD24 awaits an answer. More questions remain on the target cell ligands. There is evidence for molecules embedded in internalization-prone microdomains being preferably targeted, but only few ligands were unequivocally defined and the contribution of lipids is largely unanswered. Clarifying these issues is essential approaching tailored therapeutic Exo/Exo mimetics for message delivery into selective targets.
-
VIII.
The most urgent, but also the most demanding, question relates to the activity of uptaken Exo. The query may preferentially be answered with pluripotent SC to avoid overlaps between oncogenes and metastasis-associated genes. After a comprehensive analysis of the mode of uptaken Exo actions, clarifying the main level of activity requires evaluation. It is unknown whether the main activity centers on proteins, mRNA, miRNA, or lncRNA or whether the different Exo components are equally important. For TEX-derived lncRNA, a contribution by moving toward the nucleus getting involved in chromatin modulation and transcription also requires clarification. Minor, but still unanswered, points are the number of miRNA molecules needed for suppressing a given mRNA. Progress should also be strived for judging the efficacy of lncRNA as ceRNA and the contribution of lncRNA-incorporated miRNA. Clarifying metastasis-related CIC-TEX activities may greatly facilitate gaining knowledge on Exo activity in nontransformed cells. All PaCIC- and CoCIC-TEX biomarkers are being important in the crosstalk with non-CIC and nontransformed targets, but the mode of action of none of the TEX-CIC markers being fully explored strives for answers with high priority.
Is there too much noise about a few CIC biomarkers and small vesicles? Distinct to tumor induction relying on oncogenes, tumor progression builds on overexpression of nonmutated genes and is driven by the TEX-mediated crosstalk between CIC and the near surrounding as well as distant organs. There is convincing evidence that the PaCIC/CoCIC markers CD44v6, Tspan8, Tspan8-associated integrins, cld7, EPCAM, CD133, and LGR5 contribute to tumor progression, which includes the shift or a partial shift toward EMT, apoptosis resistance, motility/docking, matrix modulation, and (lymph)angiogenesis. Due to the location in specialized microdomains, these markers also add to TEX biogenesis and facilitate selective targeting. Amply demonstrated and approved in clinical studies, many details require further exploration, above all the suggested interplay between CIC markers, miRNA, and lncRNA. Answering the open questions will provide a unique chance for tailoring Exo/Exo mimetics that could prevent any step in the metastatic cascade. With cancer mortality largely being a sequel of metastasis, there is not too much noise about CIC markers and TEX that, based on profound answers to open demands, may allow turning metastasis in a curatively treatable disease.
Data availability
Proteome Analysis (files: ZW612, ZW2484, SH2726, SH2769), Functional proteome analysis, German Cancer Research Center, Heidelberg.
Notes
Full names of proteins and genes are listed in Table S1.
References
Steck, P. A., North, S. M., & Nicolson, G. L. (1987). Purification and partial characterization of a tumour-metastasis-associated high-Mr glycoprotein from rat 13762NF mammary adenocarcinoma cells. The Biochemical Journal, 242(3), 779–787.
Raz, A., Pazerini, G., & Carmi, P. (1989). Identification of the metastasis-associated, galactoside-binding lectin as a chimeric gene product with homology to an IgE-binding protein. Cancer Research, 49(13), 3489–3493.
Rao, C. N., Castronovo, V., Schmitt, M. C., Wewer, U. M., Claysmith, A. P., Liotta, L. A., et al. (1989). Evidence for a precursor of the high-affinity metastasis-associated murine laminin receptor. Biochemistry, 28(18), 7476–7486.
Stewart, R. L., & O'Connor, K. L. (2015). Clinical significance of the integrin α6β4 in human malignancies. Laboratory Investigation, 95(9), 976–986. https://doi.org/10.1038/labinvest.2015.82.
Günthert, U., Hofmann, M., Rudy, W., Reber, S., Zöller, M., Haussmann, I., et al. (1991). A new variant of glycoprotein CD44 confers metastatic potential to rat carcinoma cells. Cell, 65(1), 13–24.
Toh, Y., Pencil, S. D., & Nicolson, G. L. (1994). A novel candidate metastasis-associated gene, mta1, differentially expressed in highly metastatic mammary adenocarcinoma cell lines. cDNA cloning, expression, and protein analyses. Journal of Biological Chemistry, 269(37), 22958–22963.
Kaur, E., Gupta, S., & Dutt, S. (2014). Clinical implications of MTA proteins in human cancer. Cancer Metastasis Reviews, 33(4), 1017–1024. https://doi.org/10.1007/s10555-014-9527-z.
Malisetty, V. L., Penugurti, V., Panta, P., Chitta, S. K., & Manavathi, B. (2017). MTA1 expression in human cancers—clinical and pharmacological significance. Biomedicine & Pharmacotherapy, 95, 956–964. https://doi.org/10.1016/j.biopha.2017.09.025.
Karhemo, P. R., Hyvönen, M., & Laakkonen, P. (2012). Metastasis-associated cell surface oncoproteomics. Frontiers in Pharmacology, 3, 192. https://doi.org/10.3389/fphar.2012.00192.
Zhang, Y. Y., Chen, B., & Ding, Y. Q. (2012). Metastasis-associated factors facilitating the progression of colorectal cancer. Asian Pacific Journal of Cancer Prevention, 13(6), 2437–2244.
Gupta, P. B., Mani, S., Yang, J., Hartwell, K., & Weinberg, R. A. (2005). The evolving portrait of cancer metastasis. Cold Spring Harbor Symposia on Quantitative Biology, 70, 291–297.
Dexter, T. M. (1979). Haemopoiesis in long-term bone marrow cultures. A review. Acta Haematologica, 62(5–6), 299–305.
Leventhal, B. G., & Konior, G. S. (1976). Leukemia: a critical review. Seminars in Oncology, 3(3), 319–325.
Ailles, L. E., & Weissman, I. L. (2007). Cancer stem cells in solid tumors. Current Opinion in Biotechnology, 18(5), 460–466.
Tirino, V., Desiderio, V., Paino, F., De Rosa, A., Papaccio, F., La Noce, M., et al. (2013). Cancer stem cells in solid tumors: an overview and new approaches for their isolation and characterization. The FASEB Journal, 27(1), 13–24. https://doi.org/10.1096/fj.12-218222.
Brabletz, T., Kalluri, R., Nieto, M. A., & Weinberg, R. A. (2018). EMT in cancer. Nature Reviews. Cancer, 18(2), 128–134. https://doi.org/10.1038/nrc.2017.118.
Woodward, W. A., & Sulman, E. P. (2008). Cancer stem cells: markers or biomarkers? Cancer Metastasis Reviews, 27(3), 459–470. https://doi.org/10.1007/s10555-008-9130-2.
Keysar, S. B., & Jimeno, A. (2010). More than markers: biological significance of cancer stem cell-defining molecules. Molecular Cancer Therapeutics, 9(9), 2450–2457. https://doi.org/10.1158/1535-7163.MCT-10-0530.
Murar, M., & Vaidya, A. (2015). Cancer stem cell markers: premises and prospects. Biomarkers in Medicine, 9(12), 1331–1342. https://doi.org/10.2217/bmm.15.85.
Johnstone, R. M., Adam, M., Hammond, J. R., Orr, L., & Turbide, C. (1987). Vesicle formation during reticulocyte maturation. Association of plasma membrane activities with released vesicles (exosomes). The Journal of Biological Chemistry, 262(19), 9412–9420.
Rashed, M. H., Bayraktar, E., Helal, G. K., Abd-Ellah, M. F., Amero, P., Chavez-Reyes, A., et al. (2017). Exosomes: from garbage bins to promising therapeutic targets. International Journal of Molecular Sciences, 18(3), E538. https://doi.org/10.3390/ijms18030538.
Lobb, R. J., Lima, L. G., & Möller, A. (2017). Exosomes: key mediators of metastasis and pre-metastatic niche formation. Seminars in Cell & Developmental Biology, 67, 3–10. https://doi.org/10.1016/j.semcdb.2017.01.004.
Steinbichler, T. B., Dudás, J., Riechelmann, H., & Skvortsova, I. I. (2017). The role of exosomes in cancer metastasis. Seminars in Cancer Biology, 44, 170–181. https://doi.org/10.1016/j.semcancer.2017.02.006.
Wu, J., Qu, Z., Fei, Z. W., Wu, J. H., & Jiang, C. P. (2017). Role of stem cell-derived exosomes in cancer. Oncology Letters, 13(5), 2855–2866. https://doi.org/10.3892/ol.2017.5824.
Sharma, A. (2018). Role of stem cell derived exosomes in tumor biology. International Journal of Cancer, 142(6), 1086–1092. https://doi.org/10.1002/ijc.31089.
Sato, S., & Weaver, A. M. (2018). Extracellular vesicles: important collaborators in cancer progression. Essays in Biochemistry, 62(2), 149–163. https://doi.org/10.1042/EBC20170080.
Abak, A., Abhari, A., & Rahimzadeh, S. (2018). Exosomes in cancer: small vesicular transporters for cancer progression and metastasis, biomarkers in cancer therapeutics. PeerJ, 6, e4763. https://doi.org/10.7717/peerj.4763.
Siegel, R. L., Miller, K. D., & Jemal, A. (2016). Cancer statistics, 2016. CA: a Cancer Journal for Clinicians, 66(1), 7–30. https://doi.org/10.3322/caac.21332.
Engelhardt, E. G., Révész, D., Tamminga, H. J., Punt, C. J. A., Koopman, M., Onwuteaka-Philipsen, B. D., et al. (2018). Clinical usefulness of tools to support decision-making for palliative treatment of metastatic colorectal cancer: a systematic review. Clinical Colorectal Cancer, 17(1), e1–e12. https://doi.org/10.1016/j.clcc.2017.06.007.
Brenner, H., Kloor, M., & Pox, C. P. (2014). Colorectal cancer. Lancet, 383, 1490–1502.
Jemal, A., Siegel, R., Ward, E., Hao, Y., Xu, J., Murray, T., et al. (2008). Cancer statistics. CA: a Cancer Journal for Clinicians, 58(2), 71–96.
Ferlay, J., Steliarova-Foucher, E., Lortet-Tieulent, J., Rosso, S., Coebergh, J. W., Comber, H., et al. (2013). Cancer incidence and mortality patterns in Europe: estimates for 40 countries in 2012. European Journal of Cancer, 49(6), 1374–1403. https://doi.org/10.1016/j.ejca.2012.12.027.
Ahrendt, S. A., & Pitt, H. A. (2002). Surgical management of pancreatic cancer. Oncology (Williston Park), 16(6), 725–734 discussion 734, 736–728, 740, 743.
Rahib, L., Smith, B. D., Aizenberg, R., Rosenzweig, A. B., Fleshman, J. M., & Matrisian, L. M. (2014). Projecting cancer incidence and deaths to 2030: the unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Research, 74(11), 2913–2921. https://doi.org/10.1158/0008-5472.CAN-14-0155.
Del Chiaro, M., Segersvärd, R., Lohr, M., & Verbeke, C. (2014). Early detection and prevention of pancreatic cancer: is it really possible today? World Journal of Gastroenterology, 20, 12118–12131.
Ajani, J. A., Song, S., Hochster, H. S., & Steinberg, I. B. (2015). Cancer stem cells: the promise and the potential. Seminars in Oncology, 42(Suppl 1), S3–S17.
Weinstein, I. B. (1987). Growth factors, oncogenes, and multistage carcinogenesis. Journal of Cellular Biochemistry, 33(3), 213–224.
Hong, S. N. (2018). Genetic and epigenetic alterations of colorectal cancer. Intest Res, 16(3), 327–337. https://doi.org/10.5217/ir.2018.16.3.327.
Aguirre, A. J., & Collisson, E. A. (2017). Advances in the genetics and biology of pancreatic cancer. Cancer Journal, 23(6), 315–320. https://doi.org/10.1097/PPO.0000000000000286.
Shiozawa, Y., Nie, B., Pienta, K. J., Morgan, T. M., & Taichman, R. S. (2013). Cancer stem cells and their role in metastasis. Pharmacology & Therapeutics, 138(2), 285–293. https://doi.org/10.1016/j.pharmthera.2013.01.014.
Li, S., & Li, Q. (2014). Cancer stem cells and tumor metastasis (review). International Journal of Oncology, 44(6), 1806–1812. https://doi.org/10.3892/ijo.2014.2362.
Daley, G. Q. (2015). Stem cells and the evolving notion of cellular identity. Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences, 370(1680), 20140376. https://doi.org/10.1098/rstb.2014.0376.
Forsberg, E. C., Bhattacharya, D., & Weissman, I. L. (2006). Hematopoietic stem cells: expression profiling and beyond. Stem Cell Reviews, 2(1), 23–30.
Wilmut, I., Schnieke, A. E., McWhir, J., Kind, A. J., & Campbell, K. H. (1997). Viable offspring derived from fetal and adult mammalian cells. Nature, 385(6619), 810–813.
Takahashi, K., & Yamanaka, S. (2006). Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell, 126(4), 663–676.
Park, I. H., Zhao, R., West, J. A., Yabuuchi, A., Huo, H., Ince, T. A., et al. (2008). Reprogramming of human somatic cells to pluripotency with defined factors. Nature, 451(7175), 141–146.
Jonsson, J., Carlsson, L., Edlund, T., & Edlund, H. (1994). Insulin-promoter-factor 1 is required for pancreas development in mice. Nature, 371(6498), 606–609.
Gu, G., Dubauskaite, J., & Melton, D. A. (2002). Direct evidence for the pancreatic lineage: NGN3+ cells are islet progenitors and are distinct from duct progenitors. Development, 129(10), 2447–2457.
Rostovskaya, M., Bredenkamp, N., & Smith, A. (2015). Towards consistent generation of pancreatic lineage progenitors from human pluripotent stem cells. Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences, 370(1680), 20140365. https://doi.org/10.1098/rstb.2014.0365.
Jiang, F. X., & Morahan, G. (2014). Pancreatic stem cells remain unresolved. Stem Cells and Development, 23(23), 2803–2812. https://doi.org/10.1089/scd.2014.0214.
Larsen, H. L., & Grapin-Botton, A. (2017). The molecular and morphogenetic basis of pancreas organogenesis. Seminars in Cell & Developmental Biology, 66, 51–68. https://doi.org/10.1016/j.semcdb.2017.01.005.
Sznurkowska, M. K., Hannezo, E., Azzarelli, R., Rulands, S., Nestorowa, S., Hindley, C. J., et al. (2018). Defining lineage potential and fate behavior of precursors during pancreas development. Developmental Cell, 46(3), 360–375.e5. https://doi.org/10.1016/j.devcel.2018.06.028.
Buczacki, S. J., Zecchini, H. I., Nicholson, A. M., Russell, R., Vermeulen, L., Kemp, R., et al. (2013). Intestinal label-retaining cells are secretory precursors expressing Lgr5. Nature, 495(7439), 65–69. https://doi.org/10.1038/nature11965.
Zhang, Z., & Huang, J. (2013). Intestinal stem cells—types and markers. Cell Biology International, 37(5), 406–414. https://doi.org/10.1002/cbin.10049.
Clevers, H. C., & Bevins, C. L. (2013). Paneth cells: maestros of the small intestinal crypts. Annual Review of Physiology, 75, 289–311. https://doi.org/10.1146/annurev-physiol-030212-183744.
Kriz, V., & Korinek, V. (2018). Wnt, RSPO and Hippo signalling in the intestine and intestinal stem cells. Genes (Basel), 9(1), E20. https://doi.org/10.3390/genes9010020.
Krausova, M., & Korinek, V. (2014). Wnt signaling in adult intestinal stem cells and cancer. Cellular Signalling, 26(3), 570–579. https://doi.org/10.1016/j.cellsig.2013.11.032.
Clevers, H., Loh, K. M., & Nusse, R. (2014). Stem cell signaling. An integral program for tissue renewal and regeneration: Wnt signaling and stem cell control. Science, 346(6205), 1248012. https://doi.org/10.1126/science.1248012.
Park, S., Cui, J., Yu, W., Wu, L., Carmon, K. S., & Liu, Q. J. (2018). Differential activities and mechanisms of the four R-spondins in potentiating Wnt/β-catenin signaling. The Journal of Biological Chemistry, 293(25), 9759–9769. https://doi.org/10.1074/jbc.RA118.002743.
Yan, K. S., Janda, C. Y., Chang, J., Zheng, G. X. Y., Larkin, K. A., Luca, V. C., et al. (2017). Non-equivalence of Wnt and R-spondin ligands during Lgr5+ intestinal stem-cell self-renewal. Nature, 545(7653), 238–242. https://doi.org/10.1038/nature22313.
Passegué, E., & Weisman, I. L. (2005). Leukemic stem cells: where do they come from? Stem Cell Reviews, 1(3), 181–188.
Johnsen, H. E., Kjeldsen, M. K., Urup, T., Fogd, K., Pilgaard, L., Boegsted, M., et al. (2009). Cancer stem cells and the cellular hierarchy in haematological malignancies. European Journal of Cancer, 45(Suppl 1), 194–201. https://doi.org/10.1016/S0959-8049(09)70033-4.
Shah, M., & Allegrucci, C. (2013). Stem cell plasticity in development and cancer: epigenetic origin of cancer stem cells. Sub-Cellular Biochemistry, 61, 545–565. https://doi.org/10.1007/978-94-007-4525-4_24.
Verga Falzacappa, M. V., Ronchini, C., Reavie, L. B., & Pelicci, P. G. (2012). Regulation of self-renewal in normal and cancer stem cells. The FEBS Journal, 279(19), 3559–3572. https://doi.org/10.1111/j.1742-4658.2012.08727.x.
Liu, J. (2018). The dualistic origin of human tumors. Seminars in Cancer Biology, 2018. https://doi.org/10.1016/j.semcancer.2018.07.004.
Reya, T., Morrison, S. J., Clarke, M. F., & Weissman, I. L. (2001). Stem cells, cancer, and cancer stem cells. Nature, 414(6859), 105–111.
Mantamadiotis, T., & Taraviras, S. (2011). Self-renewal mechanisms in neural cancer stem cells. Front Biosci (Landmark Ed), 16, 598–607.
Hinge, A., & Filippi, M. D. (2016). Deconstructing the complexity of TGFβ signaling in hematopoietic stem cells: quiescence and beyond. Curr Stem Cell Rep, 2(4), 388–397. https://doi.org/10.1007/s40778-016-0069-x.
Soteriou, D., & Fuchs, Y. (2018). A matter of life and death: stem cell survival in tissue regeneration and tumour formation. Nature Reviews. Cancer, 18(3), 187–201. https://doi.org/10.1038/nrc.2017.122.
Alison, M. R., Guppy, N. J., Lim, S. M., & Nicholson, L. J. (2010). Finding cancer stem cells: are aldehyde dehydrogenases fit for purpose? The Journal of Pathology, 222(4), 335–344. https://doi.org/10.1002/path.2772.
Easwaran, H., Tsai, H. C., & Baylin, S. B. (2014). Cancer epigenetics: tumor heterogeneity, plasticity of stem-like states, and drug resistance. Molecular Cell, 54(5), 716–727. https://doi.org/10.1016/j.molcel.2014.05.015.
Colak, S., & Medema, J. P. (2014). Cancer stem cells—important players in tumor therapy resistance. The FEBS Journal, 281(21), 4779–4791. https://doi.org/10.1111/febs.13023.
Skvortsova, I., Debbage, P., Kumar, V., & Skvortsov, S. (2015). Radiation resistance: cancer stem cells (CSCs) and their enigmatic pro-survival signaling. Seminars in Cancer Biology, 35, 39–44. https://doi.org/10.1016/j.semcancer.2015.09.009.
Lipinska, N., Romaniuk, A., Paszel-Jaworska, A., Toton, E., Kopczynski, P., & Rubis, B. (2017). Telomerase and drug resistance in cancer. Cellular and Molecular Life Sciences, 74(22), 4121–4132. https://doi.org/10.1007/s00018-017-2573-2.
Yan, Y., Zuo, X., & Wie, D. (2015). Concise review: Emerging role of CD44 in cancer stem cells: a promising biomarker and therapeutic target. Stem Cells Translational Medicine, 4(9), 1033–1043. https://doi.org/10.5966/sctm.2015-0048.
de Lucas, B., Pérez, L. M., & Gálvez, B. G. (2018). Importance and regulation of adult stem cell migration. Journal of Cellular and Molecular Medicine, 22(2), 746–754. https://doi.org/10.1111/jcmm.13422.
Hamidi, H., & Ivaska, J. (2018). Every step of the way: integrins in cancer progression and metastasis. Nature Reviews. Cancer. https://doi.org/10.1038/s41568-018-0038-z.
Smith, G. H., & Boulanger, C. A. (2003). Mammary epithelial stem cells: transplantation and self-renewal analysis. Cell Proliferation, 36(Suppl 1), 3–15.
Whittle, J. R., Lewis, M. T., Lindeman, G. J., & Visvader, J. E. (2015). Patient-derived xenograft models of breast cancer and their predictive power. Breast Cancer Research, 17, 17. https://doi.org/10.1186/s13058-015-0523-1.
Lapidot, T., Sirard, C., Vormoor, J., Murdoch, B., Hoang, T., Caceres-Cortes, J., et al. (1994). A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature, 367(6464), 645–648.
Al-Hajj, M., Wicha, M. S., Benito-Hernandez, A., Morrison, S. J., & Clarke, M. F. (2003). Prospective identification of tumorigenic breast cancer cells. Proceedings of the National Academy of Sciences of the United States of America, 100(7), 3983–3988.
Moghbeli, M., Moghbeli, F., Forghanifard, M. M., & Abbaszadegan, M. R. (2014). Cancer stem cell detection and isolation. Medical Oncology, 31(9), 69. https://doi.org/10.1007/s12032-014-0069-6.
Telford, W. G. (2013). Stem cell identification by DyeCycle Violet side population analysis. Methods in Molecular Biology, 946, 163–179. https://doi.org/10.1007/978-1-62703-128-8_11.
Ishiguro, T., Ohata, H., Sato, A., Yamawaki, K., Enomoto, T., & Okamoto, K. (2017). Tumor-derived spheroids: relevance to cancer stem cells and clinical applications. Cancer Science, 108(3), 283–289. https://doi.org/10.1111/cas.13155.
Ma, I., & Allan, A. L. (2011). The role of human aldehyde dehydrogenase in normal and cancer stem cells. Stem Cell Reviews, 7(2), 292–306. https://doi.org/10.1007/s12015-010-9208-4.
Duan, J. J., Cai, J., Guo, Y. F., Bian, X. W., & Yu, S. C. (2016). ALDH1A3, a metabolic target for cancer diagnosis and therapy. International Journal of Cancer, 139(5), 965–975. https://doi.org/10.1002/ijc.30091.
Mele, L., Liccardo, D., & Tirino, V. (2018). Evaluation and isolation of cancer stem cells using ALDH activity assay. Methods in Molecular Biology, 1692, 43–48. https://doi.org/10.1007/978-1-4939-7401-6_4.
Gopalan, V., Islam, F., & Lam, A. K. (2018). Surface markers for the identification of cancer stem cells. Methods in Molecular Biology, 1692, 17–29. https://doi.org/10.1007/978-1-4939-7401-6_2.
Pelosi, E., Castelli, G., & Testa, U. (2015). Targeting LSCs through membrane antigens selectively or preferentially expressed on these cells. Blood Cells, Molecules & Diseases, 55(4), 336–346. https://doi.org/10.1016/j.bcmd.2015.07.015.
Bao, B., Ahmad, A., Azmi, A.S., Ali, S., & Sarkar, F.H. (2013). Overview of cancer stem cells (CSCs) and mechanisms of their regulation: implications for cancer therapy. Curr Protoc Pharmacol, Chapter 14:Unit 14.25. doi: 10.1002/0471141755.ph1425s61.
Guzman, M. L., & Allan, J. N. (2014). Concise review: Leukemia stem cells in personalized medicine. Stem Cells, 32(4), 844–851. https://doi.org/10.1002/stem.1597.
Munz, M., Baeuerle, P. A., & Gires, O. (2009). The emerging role of EpCAM in cancer and stem cell signaling. Cancer Research, 69(14), 5627–5629. https://doi.org/10.1158/0008-5472.CAN-09-0654.
Todaro, M., Gaggianesi, M., Catalano, V., Benfante, A., Iovino, F., Biffoni, M., et al. (2014). CD44v6 is a marker of constitutive and reprogrammed cancer stem cells driving colon cancer metastasis. Cell Stem Cell, 14(3), 342–356. https://doi.org/10.1016/j.stem.2014.01.009.
Smith, N. R., Davies, P. S., Levin, T. G., Gallagher, A. C., Keene, D. R., Sengupta, S. K., et al. (2017). Cell adhesion molecule CD166/ALCAM functions within the crypt to orchestrate murine intestinal stem cell homeostasis. Cellular and Molecular Gastroenterology and Hepatology, 3(3), 389–409. https://doi.org/10.1016/j.jcmgh.2016.12.010.
Jung, P., Sato, T., Merlos-Suárez, A., Barriga, F. M., Iglesias, M., Rossell, D., et al. (2011). Isolation and in vitro expansion of human colonic stem cells. Nature Medicine, 17(10), 1225–1227. https://doi.org/10.1038/nm.2470.
Li, C., Wu, J. J., Hynes, M., Dosch, J., Sarkar, B., Welling, T. H., et al. (2011). c-Met is a marker of pancreatic cancer stem cells and therapeutic target. Gastroenterology, 141(6), 2218–2227.e5. https://doi.org/10.1053/j.gastro.2011.08.009.
Hermann, P. C., Huber, S. L., Herrler, T., Aicher, A., Ellwart, J. W., Guba, M., et al. (2007). Distinct populations of cancer stem cells determine tumor growth and metastatic activity in human pancreatic cancer. Cell Stem Cell, 1(3), 313–323. https://doi.org/10.1016/j.stem.2007.06.002.
Dalerba, P., Dylla, S. J., Park, I. K., Liu, R., Wang, X., Cho, R. W., et al. (2007). Phenotypic characterization of human colorectal cancer stem cells. Proceedings of the National Academy of Sciences of the United States of America, 104(24), 10158–10163.
Ricci-Vitiani, L., Lombardi, D. G., Pilozzi, E., Biffoni, M., Todaro, M., Peschle, C., et al. (2007). Identification and expansion of human colon-cancer-initiating cells. Nature, 445(7123), 111–115.
Ren, F., Sheng, W. Q., & Du, X. (2013). CD133: a cancer stem cells marker, is used in colorectal cancers. World Journal of Gastroenterology, 19(17), 2603–2611. https://doi.org/10.3748/wjg.v19.i17.2603.
Mak, A. B., Nixon, A. M., Kittanakom, S., Stewart, J. M., Chen, G. I., Curak, J., et al. (2012). Regulation of CD133 by HDAC6 promotes β-catenin signaling to suppress cancer cell differentiation. Cell Reports, 2(4), 951–963. https://doi.org/10.1016/j.celrep.2012.09.016.
Shimozato, O., Waraya, M., Nakashima, K., Souda, H., Takiguchi, N., Yamamoto, H., et al. (2015). Receptor-type protein tyrosine phosphatase κ directly dephosphorylates CD133 and regulates downstream AKT activation. Oncogene, 34(15), 1949–1960. https://doi.org/10.1038/onc.2014.141.
Röper, K., Corbeil, D., & Huttner, W. B. (2000). Retention of prominin in microvilli reveals distinct cholesterol-based lipid micro-domains in the apical plasma membrane. Nature Cell Biology, 2(9), 582–592.
Giebel, B., Corbeil, D., Beckmann, J., Höhn, J., Freund, D., Giesen, K., et al. (2004). Segregation of lipid raft markers including CD133 in polarized human hematopoietic stem and progenitor cells. Blood, 104(8), 2332–2338.
Simons, K., & Toomre, D. (2000). Lipid rafts and signal transduction. Nature Reviews. Molecular Cell Biology, 1(1), 31–39.
Fonseca, A. V., Bauer, N., & Corbeil, D. (2008). The stem cell marker CD133 meets the endosomal compartment—new insights into the cell division of hematopoietic stem cells. Blood Cells, Molecules & Diseases, 41(2), 194–195. https://doi.org/10.1016/j.bcmd.2008.04.004.
Kemper, K., Prasetyanti, P. R., De Lau, W., Rodermond, H., Clevers, H., & Medema, J. P. (2012). Monoclonal antibodies against Lgr5 identify human colorectal cancer stem cells. Stem Cells, 30(11), 2378–2386. https://doi.org/10.1002/stem.1233.
de Lau, W., Barker, N., Low, T. Y., Koo, B. K., Li, V. S., Teunissen, H., et al. (2011). Lgr5 homologues associate with Wnt receptors and mediate R-spondin signalling. Nature, 476(7360), 293–297. https://doi.org/10.1038/nature10337.
Koo, B. K., & Clevers, H. (2014). Stem cells marked by the R-spondin receptor LGR5. Gastroenterology, 147(2), 289–302. https://doi.org/10.1053/j.gastro.2014.05.007.
de Sousa e Melo, F., Kurtova, A. V., Harnoss, J. M., Kljavin, N., Hoeck, J. D., Hung, J., et al. (2017). A distinct role for Lgr5+ stem cells in primary and metastatic colon cancer. Nature, 543(7647), 676–680. https://doi.org/10.1038/nature21713.
Leung, C., Tan, S. H., & Barker, N. (2018). Recent advances in Lgr5+ stem cell research. Trends in Cell Biology, 28(5), 380–391. https://doi.org/10.1016/j.tcb.2018.01.010.
Idzerda, R. L., Carter, W. G., Nottenburg, C., Wayner, E. A., Gallatin, W. M., & John, T. (1989). Isolation and DNA sequence of a cDNA clone encoding a lymphocyte adhesion receptor for high endothelium. Proceedings of the National Academy of Sciences of the United States of America, 86, 4659–4663.
Goldstein, L. A., & Butcher, E. C. (1990). Identification of mRNA that encodes an alternative form of H-CAM (CD44) in lymphoid and nonlymphoid tissues. Immunogenetics, 32, 389–397.
Screaton, G. R., Bell, M. V., Jackson, D. G., Cornelis, F. B., Gerth, U., & Bell, J. I. (1992). Genomic structure of DNA encoding the lymphocyte homing receptor CD44 reveals at least 12 alternatively spliced exons. Proceedings of the National Academy of Sciences of the United States of America, 89, 12160–12164.
Ishii, S., Ford, R., Thomas, P., Nachman, A., Steele, G., Jr., & Jessup, J. M. (1993). CD44 participates in the adhesion of human colorectal carcinoma cells to laminin and type IV collagen. Surgical Oncology, 2, 255–264.
Bennett, K. L., Jackson, D. G., Simon, J. C., Tanczos, E., Peach, R., Modrell, B., et al. (1995). CD44 isoforms containing exon v3 are responsible for the presentation of heparin-binding growth factor. The Journal of Cell Biology, 128, 687–698.
Neame, S. J., & Isacke, C. M. (1993). The cytoplasmic tail of CD44 is required for basolateral localization in ephitelial MDCK cells but does not mediate association with the detergent-insoluble cytoskeleton of fibroblasts. The Journal of Cell Biology, 121, 1299–1310.
Liu, D., & Sy, M. S. (1997). Phorbol myristate acetate stimulates the dimerization of CD44 involving a cysteine in the transmembrane domain. Journal of Immunology, 159, 2702–2711.
Föger, N., Marhaba, R., & Zöller, M. (1999). Raft associated interaction of CD44 with the cytoskeleton. Journal of Cell Science, 114, 1169–1178.
Oliferenko, S., Paiha, K., Harder, T., Gerke, V., Schwärzler, C., Schwarz, H., et al. (1999). Analysis of CD44-containing lipid rafts: recruitment of annexin II and stabilization by the actin cytoskeleton. The Journal of Cell Biology, 146, 843–854.
Lokeshwar, V. B., Fregien, N., & Bourguignon, L. Y. (1994). Ankyrin-binding domain of CD44(Gp85) is required for the expression of hyaluronic acid-mediated adhesion function. The Journal of Cell Biology, 126, 1099–1109.
Ruiz, P., Schwärzler, C., & Günthert, U. (1995). CD44 isoforms during differentiation and development. Bioessays, 17, 17–24.
Jalkanen, S., & Jalkanen, M. (1992). Lymphocyte CD44 binds the COOH-terminal heparin-binding domain of fibronectin. The Journal of Cell Biology, 116, 817–825.
Toyama-Sorimachi, N., & Miyasaka, M. (1994). A novel ligand for CD44 is sulfated proteoglycan. International Immunology, 6, 655–660.
Aruffo, A., Stamenkovic, I., Melnick, M., Underhill, C. B., & Seed, B. (1990). CD44 is the principal cell surface receptor for hyaluronate. Cell, 61, 1303–1313.
Greenfield, B., Wang, W. C., Marquardt, H., Piepkorn, M., Wolff, E. A., Aruffo, A., et al. (1999). Characterization of the heparan sulfate and chondroitin sulfate assembly sites in CD44. The Journal of Biological Chemistry, 274, 2511–2517.
Higman, V. A., Briggs, D. C., Mahoney, D. J., Blundell, C. D., Sattelle, B. M., Dyer, D. P., et al. (2014). A refined model for the TSG-6 link module in complex with hyaluronan: use of defined oligosaccharides to probe structure and function. The Journal of Biological Chemistry, 289, 5619–5634. https://doi.org/10.1074/jbc.M113.542357.
Orian-Rousseau, V., & Ponta, H. (2008). Adhesion proteins meet receptors: a common theme? Advances in Cancer Research, 101, 63–92.
Tremmel, M., Matzke, A., Albrecht, I., Laib, A. M., Olaku, V., Ballmer-Hofer, K., et al. (2009). A CD44v6 peptide reveals a role of CD44 in VEGFR-2 signaling and angiogenesis. Blood, 114, 5236–5244. https://doi.org/10.1182/blood-2009-04-219204.
Kim, M. S., Park, M. J., Moon, E. J., Kim, S. J., Lee, C. H., Yoo, H., et al. (2005). Hyaluronic acid induces osteopontin via the phosphatidylinositol 3-kinase/Akt pathway to enhance the motility of human glioma cells. Cancer Research, 65, 686–691.
Orian-Rousseau, V. (2015). CD44 acts as a signaling platform controlling tumor progression and metastasis. Frontiers in Immunology, 6, 154. https://doi.org/10.3389/fimmu.2015.00154.
Mori, T., Kitano, K., Terawaki, S., Maesaki, R., Fukami, Y., & Hakoshima, T. (2008). Structural basis for CD44 recognition by ERM proteins. The Journal of Biological Chemistry, 283, 29602–29612.
Fehon, R. G., McClatchey, A. I., & Bretscher, A. (2010). Organizing the cell cortex: the role of ERM proteins. Nature Reviews. Molecular Cell Biology, 11, 276–287.
Stamenkovic, I., & Yu, Q. (2010). Merlin, a “magic” linker between extracellular cues and intracellular signaling pathways that regulate cell. Motility, proliferation, and survival. Current Protein & Peptide Science, 11, 471–484.
Orian-Rousseau, V., Morrison, H., Matzke, A., Kastilan, T., Pace, G., Herrlich, P., et al. (2007). Hepatocyte growth factor-induced Ras activation requires ERM proteins linked to both CD44v6 and F-actin. Molecular Biology of the Cell, 18, 76–83. https://doi.org/10.1091/mbc.E06-08-0674.
Adamia, S., Maxwell, C. A., & Pilarski, L. M. (2005). Hyaluronan and hyaluronan synthases: potential therapeutic targets in cancer. Current Drug Targets. Cardiovascular & Haematological Disorders, 5, 3–14.
Misra, S., Toole, B. P., & Ghatak, S. (2006). Hyaluronan constitutively regulates activation of multiple receptor tyrosine kinases in epithelial and carcinoma cells. The Journal of Biological Chemistry, 281, 34936–34941.
Kozovska, Z., Gabrisova, V., & Kucerova, L. (2014). Colon cancer: cancer stem cells markers, drug resistance and treatment. Biomedicine & Pharmacotherapy, 68, 911–916. https://doi.org/10.1016/j.biopha.2014.10.019.
Grass, G. D., Dai, L., Qin, Z., Parsons, C., & Toole, B. P. (2014). CD147: regulator of hyaluronan signaling in invasiveness and chemoresistance. Advances in Cancer Research, 123, 351–373. https://doi.org/10.1016/B978-0-12-800092-2.00013-7.
Bourguignon, L. Y. (2008). Hyaluronan-mediated CD44 activation of RhoGTPase signaling and cytoskeleton function promotes tumor progression. Seminars in Cancer Biology, 18, 251–259.
Ghatak, S., Misra, S., & Toole, B. P. (2005). Hyaluronan constitutively regulates ErbB2 phosphorylation and signaling complex formation in carcinoma cells. The Journal of Biological Chemistry, 280, 8875–8883. https://doi.org/10.1074/jbc.M410882200.
Heldin, P., Basu, K., Kozlova, I., & Porsch, H. (2014). HAS2 and CD44 in breast tumorigenesis. Advances in Cancer Research, 123, 211–229. https://doi.org/10.1016/B978-0-12-800092-2.00008-3.
Xu, H., Tian, Y., Yuan, X., Wu, H., Liu, Q., Pestell, R. G., et al. (2015). The role of CD44 in epithelial-mesenchymal transition and cancer development. Onco Targets Ther, 8, 3783–3792. https://doi.org/10.2147/OTT.S95470.
Nastase, M. V., Janicova, A., Wygrecka, M., & Schaefer, L. (2017). Signaling at the crossroads: matrix-derived proteoglycan and reactive oxygen species signaling. Antioxidants & Redox Signaling, 27(12), 855–873. https://doi.org/10.1089/ars.2017.7165.
Ekyalongo, R. C., Nakayama, H., Kina, K., Kaga, N., & Iwabuchi, K. (2015). Organization and functions of glycolipid-enriched microdomains in phagocytes. Biochimica et Biophysica Acta, 1851, 90–97. https://doi.org/10.1016/j.bbalip.2014.06.009.
Korcsmaros, T., & Schneider, M. V. (2017). Superti-Furga G. Next generation of network medicine: interdisciplinary signaling approaches. Integr Biol (Camb), 9, 97–108. https://doi.org/10.1039/c6ib00215c.
Stipp, C. S., Kolesnikova, T. V., & Hemler, M. E. (2003). Functional domains in tetraspanin proteins. Trends in Biochemical Sciences, 28, 106–112.
Hemler, M. E. (2005). Tetraspanin functions and associated microdomains. Nature Reviews. Molecular Cell Biology, 6, 801–811.
Levy, S., & Shoham, T. (2005). Protein-protein interactions in the tetraspanin web. Physiology (Bethesda), 20, 218–224.
Halova, I., & Draber, P. (2016). Tetraspanins and transmembrane adaptor proteins as plasma membrane organizers—mast cell case. Frontiers in Cell and Development Biology, 4, 43. https://doi.org/10.3389/fcell.2016.00043.
Berditchevski, F., & Odintsova, E. (2007). Tetraspanins as regulators of protein trafficking. Traffic, 8, 89–96.
Yáñez-Mó, M., Gutiérrez-López, M. D., & Cabañas, C. (2011). Functional interplay between tetraspanins and proteases. Cellular and Molecular Life Sciences, 68, 3323–3335. https://doi.org/10.1007/s00018-011-0746-y.
Stepanek, O., Draber, P., & Horejsi, V. (2014). Palmitoylated transmembrane adaptor proteins in leukocyte signaling. Cellular Signalling, 26, 895–902. https://doi.org/10.1016/j.cellsig.2014.01.007.
Termini, C. M., & Gillette, J. M. (2017). Tetraspanins function as regulators of cellular signaling. Frontiers in Cell and Development Biology, 5, 34. https://doi.org/10.3389/fcell.2017.00034.
Schmidt, T. H., Homsi, Y., & Lang, T. (2016). Oligomerization of the tetraspanin CD81 via the flexibility of its δ-loop. Biophysical Journal, 110, 2463–2474. https://doi.org/10.1016/j.bpj.2016.05.003.
Yue, S., Zhao, K., Erb, U., Rana, S., & Zöller, M. (2017). Joint features and complementarities of Tspan8 and CD151 revealed in knockdown and knockout models. Biochemical Society Transactions, 45, 437–447. https://doi.org/10.1042/BST20160298.
Park, C. S., Kim, T. K., Kim, H. G., Kim, Y. J., Jeoung, M. H., Lee, W. R., et al. (2016). Therapeutic targeting of tetraspanin8 in epithelial ovarian cancer invasion and metastasis. Oncogene, 35, 4540–4548. https://doi.org/10.1038/onc.2015.520.
Fang, T., Lin, J., Wang, Y., Chen, G., Huang, J., Chen, J., et al. (2016). Tetraspanin-8 promotes hepatocellular carcinoma metastasis by increasing ADAM12m expression. Oncotarget, 7, 40630–40643. doi: 10.18632/oncotarget.9769.
Wie, L., Li, Y., & Suo, Z. (2015). TSPAN8 promotes gastric cancer growth and metastasis via ERK MAPK pathway. International Journal of Clinical and Experimental Medicine, 8(6), 8599–8607.
El Kharbili, M., Robert, C., Witkowski, T., Danty-Berger, E., Barbollat-Boutrand, L., Masse, I., et al. (2017). Tetraspanin 8 is a novel regulator of ILK-driven β1 integrin adhesion and signaling in invasive melanoma cells. Oncotarget, 8(10), 17140–17155. https://doi.org/10.18632/oncotarget.15084.
Pan, S. J., Wu, Y. B., Cai, S., Pan, Y. X., Liu, W., Bian, L. G., et al. (2015). Over-expression of tetraspanin 8 in malignant glioma regulates tumor cell progression. Biochemical and Biophysical Research Communications, 458, 476–482. https://doi.org/10.1016/j.bbrc.2015.01.128.
Wang, H., Rana, S., Giese, N., Büchler, M. W., & Zöller, M. (2013). Tspan8, CD44v6 and alpha6beta4 are biomarkers of migrating pancreatic cancer-initiating cells. International Journal of Cancer, 133(2), 416–426. https://doi.org/10.1002/ijc.28044.
Madhavan, B., Yue, S., Galli, U., Rana, S., Groß, W., Müller, M., et al. (2015). Combined evaluation of a panel of protein and miRNA serum-exosome biomarkers for pancreatic cancer diagnosis increases sensitivity and specificity. International Journal of Cancer, 136(11), 2616–2627. https://doi.org/10.1002/ijc.29324.
Greco, C., Bralet, M. P., Ailane, N., Dubart-Kupperschmitt, A., Rubinstein, E., Le Naour, F., et al. (2010). E-cadherin/p120-catenin and tetraspanin Co-029 cooperate for cell motility control in human colon carcinoma. Cancer Research, 70(19), 7674–7683. https://doi.org/10.1158/0008-5472.CAN-09-4482.
Ailane, N., Greco, C., Zhu, Y., Sala-Valdés, M., Billard, M., Casal, I., et al. (2014). Effect of an anti-human Co-029/tspan8 mouse monoclonal antibody on tumor growth in a nude mouse model. Frontiers in Physiology, 5, 364. https://doi.org/10.3389/fphys.2014.00364.
Pan, S. J., Zhan, S. K., Pan, Y. X., Liu, W., Bian, L. G., Sun, B., et al. (2015). Tetraspanin 8-rictor-integrin α3 complex is required for glioma cell migration. International Journal of Molecular Sciences, 16, 5363–5374. https://doi.org/10.3390/ijms16035363.
Wang, Z., von Au, A., Schnölzer, M., Hackert, T., & Zöller, M. (2016). CD44v6-competent tumor exosomes promote motility, invasion and cancer-initiating cell marker expression in pancreatic and colorectal cancer cells. Oncotarget, 7(34), 55409–55436. https://doi.org/10.18632/oncotarget.10580.
Yue, S., Mu, W., & Zöller, M. (2013). Tspan8 and CD151 promote metastasis by distinct mechanisms. European Journal of Cancer, 49(13), 2934–2948. https://doi.org/10.1016/j.ejca.2013.03.032.
Schmidt, F., Müller, M., Prox, J., Arnold, P., Schönherr, C., Tredup, C., et al. (2016). Tetraspanin 8 is an interactor of the metalloprotease meprin β within tetraspanin-enriched microdomains. Biological Chemistry, 397(9), 857–869. https://doi.org/10.1515/hsz-2016-0126.
Zhu, Y., Ailane, N., Sala-Valdés, M., Haghighi-Rad, F., Billard, M., Nguyen, V., et al. (2017). Multi-factorial modulation of colorectal carcinoma cells motility—partial coordination by the tetraspanin Co-029/tspan8. Oncotarget, 8(16), 27454–27470. https://doi.org/10.18632/oncotarget.16247.
Gesierich, S., Berezovskiy, I., Ryschich, E., & Zöller, M. (2006). Systemic induction of the angiogenesis switch by the tetraspanin D6.1A/CO-029. Cancer Research, 66, 7083–7094.
Nazarenko, I., Rana, S., Baumann, A., McAlear, J., Hellwig, A., Trendelenburg, M., et al. (2010). Cell surface tetraspanin Tspan8 contributes to molecular pathways of exosome-induced endothelial cell activation. Cancer Research, 70(4), 1668–1678. https://doi.org/10.1158/0008-5472.CAN-09-2470.
Rana, S., Claas, C., Kretz, C. C., Nazarenko, I., & Zöller, M. (2011). Activation-induced internalization differs for the tetraspanins CD9 and Tspan8: impact on tumor cell motility. The International Journal of Biochemistry & Cell Biology, 43(1), 106–119. https://doi.org/10.1016/j.biocel.2010.10.002.
Litvinov, S. V., Velders, M. P., Bakker, H. A., Fleuren, G. J., & Warnaar, S. O. (1994). Ep-CAM: a human epithelial antigen is a homophilic cell-cell adhesion molecule. The Journal of Cell Biology, 125(2), 437–446.
Patriarca, C., Macchi, R. M., Marschner, A. K., & Mellstedt, H. (2012). Epithelial cell adhesion molecule expression (CD326) in cancer: a short review. Cancer Treatment Reviews, 38(1), 68–75. https://doi.org/10.1016/j.ctrv.2011.04.002.
Imrich, S., Hachmeister, M., & Gires, O. (2012). EpCAM and its potential role in tumor-initiating cells. Cell Adhesion & Migration, 6, 30–38.
Maghzal, N., Vogt, E., Reintsch, W., Fraser, J. S., & Fagotto, F. (2010). The tumor-associated EpCAM regulates morphogenetic movements through intracellular signaling. The Journal of Cell Biology, 191, 645–659.
Maetzel, D., Denzel, S., Mack, B., Eggert, C., Bärr, G., & Gires, O. (2009). Nuclear signalling by tumour-associated antigen EpCAM. Nature Cell Biology, 11, 162–171.
Lin, C. W., Liao, M. Y., Lin, W. W., Wang, Y. P., Lu, T. Y., & Wu, H. C. (2012). Epithelial cell adhesion molecule regulates tumor initiation and tumorigenesis via activating reprogramming factors and epithelial-mesenchymal transition genes expression in colon cancer. The Journal of Biological Chemistry, 287, 39449–39459.
Wang, H., Stoecklein, N. H., Lin, P. P., & Gires, O. (2017). Circulating and disseminated tumor cells: diagnostic tools and therapeutic targets in motion. Oncotarget, 8(1), 1884–1912. https://doi.org/10.18632/oncotarget.12242.
Herreros-Pomares, A., Aguilar-Gallardo, C., Calabuig-Fariñas, S., Sirera, R., Jantus-Lewintre, E., & Camps, C. (2018). EpCAM duality becomes this molecule in a new Dr. Jekyll and Mr. Hyde tale. Critical Reviews in Oncology/Hematology, 126, 52–63. https://doi.org/10.1016/j.critrevonc.2018.03.006.
Biddle, A., Liang, X., Gammon, L., Fazil, B., Harper, L. J., Emich, H., et al. (2011). Cancer stem cells in squamous cell carcinoma switch between two distinct phenotypes that are preferentially migratory or proliferative. Cancer Research, 71(15), 5317–5326. https://doi.org/10.1158/0008-5472.CAN-11-1059.
Gires, O., Klein, C. A., & Baeuerle, P. A. (2009). On the abundance of EpCAM on cancer stem cells. Nature Reviews. Cancer, 9(2), 143; author reply 143. https://doi.org/10.1038/nrc2499-c1.
González, B., Denzel, S., Mack, B., Conrad, M., & Gires, O. (2009). EpCAM is involved in maintenance of the murine embryonic stem cell phenotype. Stem Cells, 27(8), 1782–1791. https://doi.org/10.1002/stem.97.
Lu, T. Y., Lu, R. M., Liao, M. Y., Yu, J., Chung, C. H., Kao, C. F., et al. (2010). Epithelial cell adhesion molecule regulation is associated with the maintenance of the undifferentiated phenotype of human embryonic stem cells. The Journal of Biological Chemistry, 285(12), 8719–8732. https://doi.org/10.1074/jbc.M109.077081.
Tamura, A., & Tsukita, S. (2014). Paracellular barrier and channel functions of TJ claudins in organizing biological systems: advances in the field of barriology revealed in knockout mice. Seminars in Cell & Developmental Biology, 36, 177–185. https://doi.org/10.1016/j.semcdb.2014.09.019.
Van Itallie, C. M., & Anderson, J. M. (2014). Architecture of tight junctions and principles of molecular composition. Seminars in Cell & Developmental Biology, 36, 157–165. https://doi.org/10.1016/j.semcdb.2014.08.011.
Ding, L., Lu, Z., Foreman, O., Tatum, R., Lu, Q., Renegar, R., et al. (2012). Inflammation and disruption of the mucosal architecture in claudin-7-deficient mice. Gastroenterology, 142, 305–315. https://doi.org/10.1053/j.gastro.2011.
Tanaka, H., Takechi, M., Kiyonari, H., Shioi, G., Tamura, A., & Tsukita, S. (2015). Intestinal deletion of Claudin-7 enhances paracellular organic solute flux and initiates colonic inflammation in mice. Gut, 64, 1529–1538. https://doi.org/10.1136/gutjnl-2014-308419.
Lal-Nag, M., & Morin, P. J. (2009). The claudins. Genome Biology, 10, 235.
Van Itallie, C. M., & Anderson, J. M. (2013). Claudin interactions in and out of the tight junction. Tissue Barriers, 1, e25247.
Sjö, A., Magnusson, K. E., & Peterson, K. H. (2010). Protein kinase C activation has distinct effects on the localization, phosphorylation and detergent solubility of the claudin protein family in tight and leaky epithelial cells. The Journal of Membrane Biology, 236, 181–189.
Su, L., Nalle, S. C., Shen, L., Turner, E. S., Singh, G., Breskin, L. A., et al. (2013). TNFR2 activates MLCK-dependent tight junction dysregulation to cause apoptosis-mediated barrier loss and experimental colitis. Gastroenterology, 145(2), 407–415. https://doi.org/10.1053/j.gastro.2013.04.011.
Shen, L. (2012). Tight junctions on the move: molecular mechanisms for epithelial barrier regulation. Annals of the New York Academy of Sciences, 1258, 9–12518.
Stamatovic, S. M., Keep, R. F., & Andjelkovic, A. V. (2011). Tracing the endocytosis of claudin-5 in brain endothelial cells. Methods in Molecular Biology, 762, 303–320. https://doi.org/10.1007/978-1-61779-185-7_22.
Heiler, S., Mu, W., Zöller, M., & Thuma, F. (2015). The importance of claudin-7 palmitoylation on membrane subdomain localization and metastasis-promoting activities. Cell Communication and Signaling: CCS, 13, 29.
Thuma, F., Heiler, S., Schnölzer, M., & Zöller, M. (2016). Palmitoylated claudin7 captured in glycolipid-enriched membrane microdomains promotes metastasis via associated transmembrane and cytosolic molecules. Oncotarget, 7, 30659–30677. https://doi.org/10.18632/oncotarget.8928.
Rao, Y., Rückert, C., Saenger, W., & Haucke, V. (2012). The early steps of endocytosis: from cargo selection to membrane deformation. European Journal of Cell Biology, 91, 226–233. https://doi.org/10.1016/j.ejcb.2011.02.004.
Tauro, B. J., Greening, D. W., Mathias, R. A., Mathivanan, S., Ji, H., & Simpson, R. J. (2013). Two distinct populations of exosomes are released from LIM1863 colon carcinoma cell-derived organoids. Molecular & Cellular Proteomics, 12, 587–598.
Deshmukh, A., Binju, M., Arfuso, F., Newsholme, P., & Dharmarajan, A. (2017). Role of epigenetic modulation in cancer stem cell fate. The International Journal of Biochemistry & Cell Biology, 90, 9–16. https://doi.org/10.1016/j.biocel.2017.07.003.
Godoy, P., Schmidt-Heck, W., Hellwig, B., Nell, P., Feuerborn, D., Rahnenführer, J., et al. (2018). Assessment of stem cell differentiation based on genome-wide expression profiles. Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences, 373(1750), 20170221. https://doi.org/10.1098/rstb.2017.0221.
Niwa, H. (2018). The principles that govern transcription factor network functions in stem cells. Development, 145(6), 157420. https://doi.org/10.1242/dev.157420.
Herreros-Villanueva, M., Bujanda, L., Billadeau, D. D., & Zhang, J. S. (2014). Embryonic stem cell factors and pancreatic cancer. World Journal of Gastroenterology, 20(9), 2247–2254. https://doi.org/10.3748/wjg.v20.i9.2247.
Herreros-Villanueva, M., Zhang, J. S., Koenig, A., Abel, E. V., Smyrk, T. C., Bamlet, W. R., et al. (2013). SOX2 promotes dedifferentiation and imparts stem cell-like features to pancreatic cancer cells. Oncogenesis, 2, e61. https://doi.org/10.1038/oncsis.2013.23.
Rhim, A. D., Mirek, E. T., Aiello, N. M., Maitra, A., Bailey, J. M., McAllister, F., et al. (2012). EMT and dissemination precede pancreatic tumor formation. Cell, 148(1–2), 349–361. https://doi.org/10.1016/j.cell.2011.11.025.
Wang, S., Huang, S., & Sun, Y. L. (2017). Epithelial-mesenchymal transition in pancreatic cancer: a review. BioMed Research International, 2017, 2646148. https://doi.org/10.1155/2017/2646148.
Roe, J. S., Hwang, C. I., Somerville, T. D. D., Milazzo, J. P., Lee, E. J., Da Silva, B., et al. (2017). Enhancer reprogramming promotes pancreatic cancer metastasis. Cell, 170(5), 875–888.e20. https://doi.org/10.1016/j.cell.2017.07.007.
Kreso, A., & Dick, J. E. (2014). Evolution of the cancer stem cell model. Cell Stem Cell, 14(3), 275–291. https://doi.org/10.1016/j.stem.2014.02.006.
Katoh, M. (2017). Canonical and non-canonical WNT signaling in cancer stem cells and their niches: cellular heterogeneity, omics reprogramming, targeted therapy and tumor plasticity (review). International Journal of Oncology, 51(5), 1357–1369. https://doi.org/10.3892/ijo.2017.4129.
Fearon, E. R., & Wicha, M. S. (2014). KRAS and cancer stem cells in APC-mutant colorectal cancer. Journal of the National Cancer Institute, 106(2), djt444. https://doi.org/10.1093/jnci/djt444.
Zhang, F., Sun, H., Zhang, S., Yang, X., Zhang, G., & Su, T. (2017). Overexpression of PER3 inhibits self-renewal capability and chemoresistance of colorectal cancer stem-like cells via inhibition of notch and β-catenin signaling. Oncology Research, 25(5), 709–719. https://doi.org/10.3727/096504016X14772331883976.
Batsaikhan, B. E., Yoshikawa, K., Kurita, N., Iwata, T., Takasu, C., Kashihara, H., et al. (2014). Cyclopamine decreased the expression of Sonic Hedgehog and its downstream genes in colon cancer stem cells. Anticancer Research, 34(11), 6339–6344.
Whissell, G., Montagni, E., Martinelli, P., Hernando-Momblona, X., Sevillano, M., Jung, P., et al. (2014). The transcription factor GATA6 enables self-renewal of colon adenoma stem cells by repressing BMP gene expression. Nature Cell Biology, 16(7), 695–707. https://doi.org/10.1038/ncb2992.
Chen, J., Shao, R., Li, F., Monteiro, M., Liu, J. P., Xu, Z. P., et al. (2015). PI3K/Akt/mTOR pathway dual inhibitor BEZ235 suppresses the stemness of colon cancer stem cells. Clinical and Experimental Pharmacology & Physiology, 42(12), 1317–1326. https://doi.org/10.1111/1440-1681.12493.
Pelicci, P. G., Dalton, P., & Giorgio, M. (2013). The other face of ROS: a driver of stem cell expansion in colorectal cancer. Cell Stem Cell, 12(6), 635–636. https://doi.org/10.1016/j.stem.2013.05.023.
Hong, A. W., Meng, Z., & Guan, K. L. (2016). The Hippo pathway in intestinal regeneration and disease. Nature Reviews. Gastroenterology & Hepatology, 13(6), 324–337. https://doi.org/10.1038/nrgastro.2016.59.
Sikandar, S. S., Pate, K. T., Anderson, S., Dizon, D., Edwards, R. A., Waterman, M. L., et al. (2010). NOTCH signaling is required for formation and self-renewal of tumor-initiating cells and for repression of secretory cell differentiation in colon cancer. Cancer Research, 70(4), 1469–1478. https://doi.org/10.1158/0008-5472.CAN-09-2557.
Apostolou, P., Toloudi, M., Ioannou, E., Kourtidou, E., Chatziioannou, M., Kopic, A., et al. (2013). Study of the interaction among Notch pathway receptors, correlation with stemness, as well as their interaction with CD44, dipeptidyl peptidase-IV, hepatocyte growth factor receptor and the SETMAR transferase, in colon cancer stem cells. Journal of Receptor and Signal Transduction Research, 33(6), 353–358. https://doi.org/10.3109/10799893.2013.828072.
Fender, A. W., Nutter, J. M., Fitzgerald, T. L., Bertrand, F. E., & Sigounas, G. (2015). Notch-1 promotes stemness and epithelial to mesenchymal transition in colorectal cancer. Journal of Cellular Biochemistry, 116(11), 2517–2527. https://doi.org/10.1002/jcb.25196.
Jin, L., Vu, T., Yuan, G., & Datta, P. K. (2017). STRAP promotes stemness of human colorectal cancer via epigenetic regulation of the NOTCH pathway. Cancer Research, 77(20), 5464–5478. https://doi.org/10.1158/0008-5472.CAN-17-0286.
Voorneveld, P. W., Kodach, L. L., Jacobs, R. J., van Noesel, C. J., Peppelenbosch, M. P., Korkmaz, K. S., et al. (2015). The BMP pathway either enhances or inhibits the Wnt pathway depending on the SMAD4 and p53 status in CRC. British Journal of Cancer, 112(1), 122–130. https://doi.org/10.1038/bjc.2014.560.
Kim, B. R., Oh, S. C., Lee, D. H., Kim, J. L., Lee, S. Y., Kang, M. H., et al. (2015). BMP-2 induces motility and invasiveness by promoting colon cancer stemness through STAT3 activation. Tumour Biology, 36(12), 9475–9486. https://doi.org/10.1007/s13277-015-3681-y.
Catalano, V., Dentice, M., Ambrosio, R., Luongo, C., Carollo, R., Benfante, A., et al. (2016). Activated thyroid hormone promotes differentiation and chemotherapeutic sensitization of colorectal cancer stem cells by regulating Wnt and BMP4 signaling. Cancer Research, 76(5), 1237–1244. https://doi.org/10.1158/0008-5472.CAN-15-1542.
Xue, R., Jia, K., Wang, J., Yang, L., Wang, Y., Gao, L., et al. (2018). A rising star in pancreatic diseases: pancreatic stellate cells. Frontiers in Physiology, 9, 754. https://doi.org/10.3389/fphys.2018.00754.
Zeuner, A., Todaro, M., Stassi, G., & De Maria, R. (2014). Colorectal cancer stem cells: from the crypt to the clinic. Cell Stem Cell, 15(6), 692–705. https://doi.org/10.1016/j.stem.2014.11.012.
Calon, A., Tauriello, D. V., & Batlle, E. (2014). TGF-beta in CAF-mediated tumor growth and metastasis. Seminars in Cancer Biology, 25, 15–22. https://doi.org/10.1016/j.semcancer.2013.12.008.136.
Wang, K., & Karin, M. (2015). Tumor-elicited inflammation and colorectal cancer. Advances in Cancer Research, 128, 173–196. https://doi.org/10.1016/bs.acr.2015.04.014.
Lotti, F., Jarrar, A. M., Pai, R. K., Hitomi, M., Lathia, J., Mace, A., et al. (2013). Chemotherapy activates cancer-associated fibroblasts to maintain colorectal cancer-initiating cells by IL-17A. The Journal of Experimental Medicine, 210(13), 2851–2872. https://doi.org/10.1084/jem.20131195.
Koukourakis, M. I., Giatromanolaki, A., Harris, A. L., & Sivridis, E. (2006). Comparison of metabolic pathways between cancer cells and stromal cells in colorectal carcinomas: a metabolic survival role for tumor-associated stroma. Cancer Research, 66(2), 632–637.
Koliaraki, V., Pallangyo, C. K., Greten, F. R., & Kollias, G. (2017). Mesenchymal cells in colon cancer. Gastroenterology, 152(5), 964–979. https://doi.org/10.1053/j.gastro.2016.11.049.
Lu, J., Ye, X., Fan, F., Xia, L., Bhattacharya, R., Bellister, S., et al. (2013). Endothelial cells promote the colorectal cancer stem cell phenotype through a soluble form of Jagged-1. Cancer Cell, 23(2), 171–185. https://doi.org/10.1016/j.ccr.2012.12.021.
Roberts, K. J., Kershner, A. M., & Beachy, P. A. (2017). The stromal niche for epithelial stem cells: a template for regeneration and a brake on malignancy. Cancer Cell, 32(4), 404–410. https://doi.org/10.1016/j.ccell.2017.08.007.
Gerling, M., Büller, N. V., Kirn, L. M., Joost, S., Frings, O., Englert, B., et al. (2016). Stromal Hedgehog signalling is downregulated in colon cancer and its restoration restrains tumour growth. Nature Communications, 7, 12321. https://doi.org/10.1038/ncomms12321.
Nicolas, F. E. (2017). Role of ncRNAs in development, diagnosis and treatment of human cancer. Recent Patents on Anti-Cancer Drug Discovery, 12(2), 128–135. https://doi.org/10.2174/1574892812666170105113415.
Bartel, D. P. (2004). MicroRNAs: genomics, biogenesis, mechanism, and function. Cell, 116(2), 281–297.
Basyuk, E., Suavet, F., Doglio, A., Bordonné, R., & Bertrand, E. (2003). Human let-7 stem-loop precursors harbor features of RNase III cleavage products. Nucleic Acids Research, 31(22), 6593–6597.
Lee, Y., Ahn, C., Han, J., Choi, H., Kim, J., Yim, J., et al. (2003). The nuclear RNase III Drosha initiates microRNA processing. Nature, 425(6956), 415–419.
Chendrimada, T. P., Gregory, R. I., Kumaraswamy, E., Norman, J., Cooch, N., Nishikura, K., et al. (2005). TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature, 436(7051), 740–744.
Denli, A. M., Tops, B. B., Plasterk, R. H., Ketting, R. F., & Hannon, G. J. (2004). Processing of primary microRNAs by the microprocessor complex. Nature, 432(7014), 231–235.
Seok, H., Ham, J., Jang, E. S., & Chi, S. W. (2016). MicroRNA target recognition: insights from transcriptome-wide non-canonical interactions. Molecules and Cells, 39(5), 375–381. https://doi.org/10.14348/molcells.2016.0013.
Nicoloso, M. S., Spizzo, R., Shimizu, M., Rossi, S., & Calin, G. A. (2009). MicroRNAs—the micro steering wheel of tumour metastases. Nature Reviews. Cancer, 9(4), 293–302. https://doi.org/10.1038/nrc2619.
Acunzo, M., Romano, G., Wernicke, D., & Croce, C. M. (2015). MicroRNA and cancer—a brief overview. Adv Biol Regul, 57, 1–9. https://doi.org/10.1016/j.jbior.2014.09.013.
Giovannetti, E., van der Borden, C. L., Frampton, A. E., Ali, A., Firuzi, O., & Peters, G. J. (2017). Never let it go: stopping key mechanisms underlying metastasis to fight pancreatic cancer. Seminars in Cancer Biology, 44, 43–59. https://doi.org/10.1016/j.semcancer.2017.04.006.
Liu, X., Fu, Q., Du, Y., Yang, Y., & Cho, W. C. (2016). MicroRNA as regulators of cancer stem cells and chemoresistance in colorectal cancer. Current Cancer Drug Targets, 16(9), 738–754.
Mamoori, A., Gopalan, V., Smith, R. A., & Lam, A. K. (2016). Modulatory roles of microRNAs in the regulation of different signalling pathways in large bowel cancer stem cells. Biology of the Cell, 108(3), 51–64. https://doi.org/10.1111/boc.201500062.
Kung, J. T., Colognori, D., & Lee, J. T. (2013). Long noncoding RNAs: past, present, and future. Genetics, 193(3), 651–669. https://doi.org/10.1534/genetics.112.146704.
Poliseno, L., Salmena, L., Zhang, J., Carver, B., Haveman, W. J., & Pandolfi, P. P. (2010). A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature, 465(7301), 1033–1038. https://doi.org/10.1038/nature09144.
Johnsson, P., Ackley, A., Vidarsdottir, L., Lui, W. O., Corcoran, M., Grandér, D., et al. (2013). A pseudogene long-noncoding-RNA network regulates PTEN transcription and translation in human cells. Nature Structural & Molecular Biology, 20(4), 440–446. https://doi.org/10.1038/nsmb.2516.
Li, T., Mo, X., Fu, L., Xiao, B., & Guo, J. (2016). Molecular mechanisms of long noncoding RNAs on gastric cancer. Oncotarget, 7(8), 8601–8612. https://doi.org/10.18632/oncotarget.6926.
Sanchez-Mejias, A., & Tay, Y. (2015). Competing endogenous RNA networks: tying the essential knots for cancer biology and therapeutics. Journal of Hematology & Oncology, 8, 30. https://doi.org/10.1186/s13045-015-0129-1.
Ebert, M. S., Neilson, J. R., & Sharp, P. A. (2007). MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells. Nature Methods, 4(9), 721–726.
Kopp, F., & Mendell, J. T. (2018). Functional classification and experimental dissection of long noncoding RNAs. Cell, 172(3), 393–407. https://doi.org/10.1016/j.cell.2018.01.011.
Rinn, J. L., Kertesz, M., Wang, J. K., Squazzo, S. L., Xu, X., Brugmann, S. A., et al. (2007). Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell, 129(7), 1311–1323.
Hung, T., Wang, Y., Lin, M. F., Koegel, A. K., Kotake, Y., Grant, G. D., et al. (2011). Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nature Genetics, 43(7), 621–629. https://doi.org/10.1038/ng.848.
Liu, C., Wu, H. T., Zhu, N., Shi, Y. N., Liu, Z., Ao, B. X., et al. (2016). Steroid receptor RNA activator: biologic function and role in disease. Clinica Chimica Acta, 459, 137–146. https://doi.org/10.1016/j.cca.2016.06.004.
Thomson, D. W., & Dinger, M. E. (2016). Endogenous microRNA sponges: evidence and controversy. Nature Reviews. Genetics, 17(5), 272–283. https://doi.org/10.1038/nrg.2016.20.
Dykes, I. M., & Emanueli, C. (2017). Transcriptional and post-transcriptional gene regulation by long non-coding RNA. Genomics, Proteomics & Bioinformatics, 15(3), 177–186. https://doi.org/10.1016/j.gpb.2016.12.005.
Deng, H., Wang, J. M., Li, M., Tang, R., Tang, K., Su, Y., et al. (2017). Long non-coding RNAs: new biomarkers for prognosis and diagnosis of colon cancer. Tumour Biology, 39(6), 1010428317706332. https://doi.org/10.1177/1010428317706332.
Yang, Y., Junjie, P., Sanjun, C., & Ma, Y. (2017). Long non-coding RNAs in colorectal cancer: progression and future directions. Journal of Cancer, 8(16), 3212–3225. https://doi.org/10.7150/jca.19794.
Yang, S., Sun, Z., Zhou, Q., Wang, W., Wang, G., Song, J., et al. (2018). MicroRNAs, long noncoding RNAs, and circular RNAs: potential tumor biomarkers and targets for colorectal cancer. Cancer Management and Research, 10, 2249–2257. https://doi.org/10.2147/CMAR.S166308.
Han, T., Hu, H., Zhuo, M., Wang, L., Cui, J. J., Jiao, F., et al. (2016). Long non-coding RNA: an emerging paradigm of pancreatic cancer. Current Molecular Medicine, 16(8), 702–709.
Duguang, L., Jin, H., Xiaowei, Q., Peng, X., Xiaodong, W., Zhennan, L., et al. (2017). The involvement of lncRNAs in the development and progression of pancreatic cancer. Cancer Biology & Therapy, 18(12), 927–936. https://doi.org/10.1080/15384047.2017.1385682.
Huang, X., Xiao, R., Pan, S., Yang, X., Yuan, W., Tu, Z., et al. (2017). Uncovering the roles of long non-coding RNAs in cancer stem cells. Journal of Hematology & Oncology, 10(1), 62. https://doi.org/10.1186/s13045-017-0428-9.
Heery, R., Finn, S. P., Cuffe, S., & Gray, S. G. (2017). Long non-coding RNAs: key regulators of epithelial-mesenchymal transition, tumour drug resistance and cancer stem cells. Cancers (Basel), 9(4), E38. https://doi.org/10.3390/cancers9040038.
Chi, H. C., Tsai, C. Y., Tsai, M. M., Yeh, C. T., & Lin, K. H. (2017). Roles of long noncoding RNAs in recurrence and metastasis of radiotherapy-resistant cancer stem cells. International Journal of Molecular Sciences, 18(9), E1903. https://doi.org/10.3390/ijms18091903.
Théry, C., Zitvogel, L., & Amigorena, S. (2002). Exosomes: composition, biogenesis and function. Nature Reviews. Immunology, 2(8), 569–579. https://doi.org/10.1038/nri855.
Boukouris, S., & Mathivanan, S. (2015). Exosomes in bodily fluids are a highly stable resource of disease biomarkers. Proteomics. Clinical Applications, 9, 358–367. https://doi.org/10.1002/prca.201400114.
Jia, S., Zocco, D., Samuels, M. L., Chou, M. F., Chammas, R., Skog, J., et al. (2014). Emerging technologies in extracellular vesicle-based molecular diagnostics. Expert Review of Molecular Diagnostics, 14, 307–321. https://doi.org/10.1586/14737159.2014.893828.
Valadi, H., Ekström, K., Bossios, A., Sjöstrand, M., Lee, J. J., & Lötvall, J. O. (2007). Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nature Cell Biology, 9, 654–659.
Lo Cicero, A., Stahl, P. D., & Raposo, G. (2015). Extracellular vesicles shuffling intercellular messages: for good or for bad. Current Opinion in Cell Biology, 35, 69–77. https://doi.org/10.1016/j.ceb.2015.04.013.
Javeed, N., & Mukhopadhyay, D. (2017). Exosomes and their role in the micro-/macro-environment: a comprehensive review. Journal of Biomedical Research, 31(5), 386–394. https://doi.org/10.7555/JBR.30.20150162.
Burrello, J., Monticone, S., Gai, C., Gomez, Y., Kholia, S., & Camussi, G. (2016). Stem cell-derived extracellular vesicles and immune-modulation. Frontiers in Cell and Development Biology, 4, 83. https://doi.org/10.3389/fcell.2016.00083.
Todorova, D., Simoncini, S., Lacroix, R., Sabatier, F., & Dignat-George, F. (2017). Extracellular vesicles in angiogenesis. Circulation Research, 120(10), 1658–1673. https://doi.org/10.1161/CIRCRESAHA.117.309681.
Rajagopal, C., & Harikumar, K. B. (2018). The origin and functions of exosomes in cancer. Frontiers in Oncology, 8, 66. https://doi.org/10.3389/fonc.2018.00066.
Yang, B., Chen, Y., & Shi, J. (2018). Exosome biochemistry and advanced nanotechnology for next-generation theranostic platforms. Advanced Materials, e1802896. https://doi.org/10.1002/adma.201802896.
Colombo, M., Raposo, G., & Théry, C. (2014). Biogenesis, secretion, and intercellular interactions of exosomes and other extracellular vesicles. Annual Review of Cell and Developmental Biology, 30, 255–289. https://doi.org/10.1146/annurev-cellbio-101512-122326.
van Niel, G., D’Angelo, G., & Raposo, G. (2018). Shedding light on the cell biology of extracellular vesicles. Nature Reviews. Molecular Cell Biology, 19(4), 213–228. https://doi.org/10.1038/nrm.2017.125.
Abels, E. R., & Breakefield, X. O. (2016). Introduction to extracellular vesicles: biogenesis, RNA cargo selection, content, release, and uptake. Cellular and Molecular Neurobiology, 36, 301–312. https://doi.org/10.1007/s10571-016-0366-z.
Villarroya-Beltri, C., Baixauli, F., Gutiérrez-Vázquez, C., Sánchez-Madrid, F., & Mittelbrunn, M. (2014). Sorting it out: regulation of exosome loading. Seminars in Cancer Biology, 28, 3–13. https://doi.org/10.1016/j.semcancer.2014.04.009.
Choi, D. S., Kim, D. K., Kim, Y. K., & Gho, Y. S. (2015). Proteomics of extracellular vesicles: exosomes and ectosomes. Mass Spectrometry Reviews, 34, 474–490. https://doi.org/10.1002/pmic.201200329.
Nabhan, J. F., Hu, R., Oh, R. S., Cohen, S. N., & Lu, Q. (2012). Formation and release of arrestin domain-containing protein 1-mediated microvesicles (ARMMs) at plasma membrane by recruitment of TSG101 protein. Proceedings of the National Academy of Sciences of the United States of America, 109, 4146–4151. https://doi.org/10.1073/pnas.1200448109.
Egea-Jimenez, A. L., & Zimmermann, P. (2018). Phospholipase D and phosphatidic acid in the biogenesis and cargo loading of extracellular vesicles. Journal of Lipid Research, jlr.R083964. https://doi.org/10.1194/jlr.R083964.
Kajimoto, T., Okada, T., Miya, S., Zhang, L., & Nakamura, S. (2013). Ongoing activation of sphingosine 1-phosphate receptors mediates maturation of exosomal multi-vesicular endosomes. Nature Communications, 4, 2712. https://doi.org/10.1038/ncomms3712.
Shen, B., Fang, Y., Wu, N., & Gould, S. J. (2011). Biogenesis of the posterior pole is mediated by the exosome/microvesicle protein-sorting pathway. The Journal of Biological Chemistry, 286, 44162–44176. https://doi.org/10.1074/jbc.M111.274803.
Guo, B. B., Bellingham, S. A., & Hill, A. F. (2015). The neutral sphingomyelinase pathway regulates packaging of the prion protein into exosomes. The Journal of Biological Chemistry, 290, 3455–3467. https://doi.org/10.1074/jbc.M115.684258.
Vedeler, A., Hollås, H., Grindheim, A. K., & Raddum, A. M. (2012). Multiple roles of annexin A2 in post-transcriptional regulation of gene expression. Current Protein & Peptide Science, 13, 401–412.
Janas, T., Janas, M. M., Sapoń, K., & Janas, T. (2015). Mechanisms of RNA loading into exosomes. FEBS Letters, 589(13), 1391–1398. https://doi.org/10.1016/j.febslet.2015.04.036.
Villarroya-Beltri, C., Gutierrez-Vazquez, C., Sanchez-Cabo, F., Pérez-Hernández, D., Vázquez, J., Martin-Cofreces, N., et al. (2013). Sumoylated hnRNPA2B1 controls the sorting of miRNAs into exosomes through binding to specific motifs. Nature Communications, 4, 2980. https://doi.org/10.1038/ncomms3980.
Gezer, U., Özgür, E., Cetinkaya, M., Isin, M., & Dalay, N. (2014). Long non-coding RNAs with low expression levels in cells are enriched in secreted exosomes. Cell Biology International, 38(9), 1076–1079. https://doi.org/10.1002/cbin.10301.
Hessvik, N. P., & Llorente, A. (2018). Current knowledge on exosome biogenesis and release. Cellular and Molecular Life Sciences, 75(2), 193–208. https://doi.org/10.1007/s00018-017-2595-9.
Ji, H., Greening, D. W., Barnes, T. W., Lim, J. W., Tauro, B. J., Rai, A., et al. (2013). Proteome profiling of exosomes derived from human primary and metastatic colorectal cancer cells reveal differential expression of key metastatic factors and signal transduction components. Proteomics, 13, 1672–1686. https://doi.org/10.1002/pmic.201200562.
Kowal, J., Arras, G., Colombo, M., Jouve, M., Morath, J. P., Primdal-Bengtson, B., et al. (2016). Proteomic comparison defines novel markers to characterize heterogeneous populations of extracellular vesicle subtypes. Proceedings of the National Academy of Sciences of the United States of America, 113, E968–E977. https://doi.org/10.1073/pnas.1521230113.
Subra, C., Grand, D., Laulagnier, K., Stella, A., Lambeau, G., Paillasse, M., et al. (2010). Exosomes account for vesicle-mediated transcellular transport of activatable phospholipases and prostaglandins. Journal of Lipid Research, 51, 2105–2120.
Skotland, T., Hessvik, N. P., Sandvig, K., & Llorente, A. (2018). Exosomal lipid composition and the role of ether lipids and phosphoinositides in exosome biology. Journal of Lipid Research, jlr.R084343. https://doi.org/10.1194/jlr.R084343.
Sharma, R., Huang, X., Brekken, R. A., & Schroit, A. J. (2017). Detection of phosphatidylserine-positive exosomes for the diagnosis of early-stage malignancies. British Journal of Cancer, 117, 545–552. https://doi.org/10.1038/bjc.2017.183.
Lesur, A., & Domon, B. (2015). Advances in high-resolution accurate mass spectrometry application to targeted proteomics. Proteomics, 15, 880–890. https://doi.org/10.1002/pmic.201400450.
Schey, K. L., Luther, J. M., & Rose, K. L. (2015). Proteomics characterization of exosome cargo. Methods, 87, 75–82. https://doi.org/10.1016/j.ymeth.2015.03.018.
Zöller, M. (2009). Tetraspanins: push and pull in suppressing and promoting metastasis. Nature Reviews. Cancer, 9, 40–55. https://doi.org/10.1038/nrc2543.
Mathivanan, S., Ji, H., & Simpson, R. J. (2010). Exosomes: extracellular organelles important in intercellular communication. Journal of Proteomics, 73, 1907–1920. https://doi.org/10.1016/j.jprot.2010.06.006.
Greening, D. W., Xu, R., Gopal, S. K., Rai, A., & Simpson, R. J. (2017). Proteomic insights into extracellular vesicle biology—defining exosomes and shed microvesicles. Expert Review of Proteomics, 14(1), 69–95. https://doi.org/10.1080/14789450.2017.1260450.
Rosa-Fernandes, L., Rocha, V. B., Carregari, V. C., Urbani, A., & Palmisano, G. (2017). A perspective on extracellular vesicles proteomics. Frontiers in Chemistry, 5, 102. https://doi.org/10.3389/fchem.2017.00102.
Mears, R., Craven, R. A., Hanrahan, S., Totty, N., Upton, C., Young, S. L., et al. (2004). Proteomic analysis of melanoma-derived exosomes by two-dimensional polyacrylamide gel electrophoresis and mass spectrometry. Proteomics, 4(12), 4019–4031.
Al-Nedawi, K., Meehan, B., Micallef, J., Lhotak, V., May, L., Guha, A., et al. (2008). Intercellular transfer of the oncogenic receptor EGFRvIII by microvesicles derived from tumour cells. Nature Cell Biology, 10, 619–624. https://doi.org/10.1038/ncb1725.
Osteikoetxea, X., Benke, M., Rodriguez, M., Pálóczi, K., Sódar, B. W., Szvicsek, Z., et al. (2018). Detection and proteomic characterization of extracellular vesicles in human pancreatic juice. Biochemical and Biophysical Research Communications, 499(1), 37–43. https://doi.org/10.1016/j.bbrc.2018.03.107.
Marhaba, R., Klingbeil, P., Nuebel, T., Nazarenko, I., Buechler, M. W., & Zöller, M. (2008). CD44 and EpCAM: cancer-initiating cell markers. Current Molecular Medicine, 8(8), 784–804.
Matsumoto, K., Umitsu, M., De Silva, D. M., Roy, A., & Bottaro, D. P. (2017). Hepatocyte growth factor/MET in cancer progression and biomarker discovery. Cancer Science, 108(3), 296–307. https://doi.org/10.1111/cas.13156.
Demory Beckler, M., Higginbotham, J. N., Franklin, J. L., Ham, A. J., Halvey, P. J., Imasuen, I. E., et al. (2013). Proteomic analysis of exosomes from mutant KRAS colon cancer cells identifies intercellular transfer of mutant KRAS. Molecular & Cellular Proteomics, 12, 343–355. https://doi.org/10.1074/mcp.M112.022806.
Jung, T., Castellana, D., Klingbeil, P., Cuesta Hernández, I., Vitacolonna, M., Orlicky, D. J., et al. (2009). CD44v6 dependence of premetastatic niche preparation by exosomes. Neoplasia, 11(10), 1093–1105.
Anami, K., Oue, N., Noguchi, T., Sakamoto, N., Sentani, K., Hayashi, T., et al. (2016). TSPAN8, identified by Escherichia coli ampicillin secretion trap, is associated with cell growth and invasion in gastric cancer. Gastric Cancer, 19(2), 370–380. https://doi.org/10.1007/s10120-015-0478-z.
Hoshino, A., Costa-Silva, B., Shen, T. L., Rodrigues, G., Hashimoto, A., Tesic Mark, M., et al. (2015). Tumour exosome integrins determine organotropic metastasis. Nature, 527(7578), 329–335. https://doi.org/10.1038/nature15756.
Paolillo, M., & Schinelli, S. (2017). Integrins and exosomes, a dangerous liaison in cancer progression. Cancers (Basel), 9(8), E95. https://doi.org/10.3390/cancers9080095.
Philip, R., Heiler, S., Mu, W., Büchler, M. W., Zöller, M., & Thuma, F. (2015). Claudin-7 promotes the epithelial-mesenchymal transition in human colorectal cancer. Oncotarget, 6(4), 2046–2063.
Marimpietri, D., Petretto, A., Raffaghello, L., Pezzolo, A., Gagliani, C., Tacchetti, C., et al. (2013). Proteome profiling of neuroblastoma-derived exosomes reveal the expression of proteins potentially involved in tumor progression. PLoS One, 8(9), e75054. https://doi.org/10.1371/journal.pone.0075054.
Rappa, G., Mercapide, J., Anzanello, F., Pope, R. M., & Lorico, A. (2013). Biochemical and biological characterization of exosomes containing prominin-1/CD133. Molecular Cancer, 12, 62. https://doi.org/10.1186/1476-4598-12-62.
Kumar, D., Gupta, D., Shankar, S., & Srivastava, R. K. (2015). Biomolecular characterization of exosomes released from cancer stem cells: possible implications for biomarker and treatment of cancer. Oncotarget, 10.18632/oncotarget.2462, 6, 3280, 3291.
Zöller, M. (2016). Exosomes in cancer disease. Methods in Molecular Biology, 1381, 111–149. https://doi.org/10.1007/978-1-4939-3204-7_7.
Mulcahy, L. A., Pink, R. C., & Carter, D. R. (2014). Routes and mechanisms of extracellular vesicle uptake. J Extracell Vesicles, 3. https://doi.org/10.3402/jev.v3.24641.
Buzás, E. I., Tóth, E. Á., Sódar, B. W., & Szabó-Taylor, K. É. (2018). Molecular interactions at the surface of extracellular vesicles. Seminars in Immunopathology. https://doi.org/10.1007/s00281-018-0682-0.
Rackov, G., Garcia-Romero, N., Esteban-Rubio, S., Carrión-Navarro, J., Belda-Iniesta, C., & Ayuso-Sacido, A. (2018). Vesicle-mediated control of cell function: the role of extracellular matrix and microenvironment. Frontiers in Physiology, 9, 651. https://doi.org/10.3389/fphys.2018.00651.
Sanderson, R.D., Bandari, S.K., & Vlodavsky, I. (2017). Proteases and glycosidases on the surface of exosomes: newly discovered mechanisms for extracellular remodeling. Matrix Biol, S0945-053X(17)30311-30316. doi: https://doi.org/10.1016/j.matbio.2017.10.007.
Mu, W., Rana, S., & Zöller, M. (2013). Host matrix modulation by tumor exosomes promotes motility and invasiveness. Neoplasia, 15, 875–887.
Wang, L., Hu, L., Zhou, X., Xiong, Z., Zhang, C., Shehada, H. M. A., et al. (2017). Exosomes secreted by human adipose mesenchymal stem cells promote scarless cutaneous repair by regulating extracellular matrix remodelling. Scientific Reports, 7(1), 13321. https://doi.org/10.1038/s41598-017-12919-x.
Sung, B. H., Ketova, T., Hoshino, D., Zijlstra, A., & Weaver, A. M. (2015). Directional cell movement through tissues is controlled by exosome secretion. Nature Communications, 6, 7164. https://doi.org/10.1038/ncomms8164.
Del Vecchio, F., Lee, G. H., Hawezi, J., Bhome, R., Pugh, S., Sayan, E., et al. (2018). Long non-coding RNAs within the tumour microenvironment and their role in tumour-stroma cross-talk. Cancer Letters, 421, 94–102. https://doi.org/10.1016/j.canlet.2018.02.022.
French, K. C., Antonyak, M. A., & Cerione, R. A. (2017). Extracellular vesicle docking at the cellular port: extracellular vesicle binding and uptake. Seminars in Cell & Developmental Biology, 67, 48–55. https://doi.org/10.1016/j.semcdb.2017.01.002.
Moller-Tank, S., & Maury, W. (2014). Phosphatidylserine receptors: enhancers of enveloped virus entry and infection. Virology, 468-470, 565–580. https://doi.org/10.1016/j.virol.2014.09.009.
Christianson, H. C., Svensson, K. J., van Kuppevelt, T. H., Li, J. P., & Belting, M. (2013). Cancer cell exosomes depend on cell-surface heparan sulfate proteoglycans for their internalization and functional activity. Proceedings of the National Academy of Sciences of the United States of America, 110, 17380–17385. https://doi.org/10.1073/pnas.1304266110.
Fei, F., Joo, E. J., Tarighat, S. S., Schiffer, I., Paz, H., Fabbri, M., et al. (2015). B-cell precursor acute lymphoblastic leukemia and stromal cells communicate through Galectin-3. Oncotarget, 6, 11378–11394. https://doi.org/10.18632/oncotarget.3409.
Gomes, J., Gomes-Alves, P., Carvalho, S. B., Peixoto, C., Alves, P. M., Altevogt, P., et al. (2015). Extracellular vesicles from ovarian carcinoma cells display specific glycosignatures. Biomolecules, 5, 1741–1761. https://doi.org/10.3390/biom5031741.
Rana, S., Yue, S., Stadel, D., & Zöller, M. (2012). Toward tailored exosomes: the exosomal tetraspanin web contributes to target cell selection. The International Journal of Biochemistry & Cell Biology, 44(9), 1574–1584. https://doi.org/10.1016/j.biocel.2012.06.018.
Montecalvo, A., Larregina, A. T., Shufesky, W. J., Stolz, D. B., Sullivan, M. L., Karlsson, J. M., et al. (2012). Mechanism of transfer of functional microRNAs between mouse dendritic cells via exosomes. Blood, 119, 756–766. https://doi.org/10.1182/blood-2011-02-338004.
Del Conde, I., Shrimpton, C. N., Thiagarajan, P., & López, J. A. (2005). Tissue-factor-bearing microvesicles arise from lipid rafts and fuse with activated platelets to initiate coagulation. Blood, 106, 1604–1611. https://doi.org/10.1182/blood-2004-03-1095.
Tian, T., Zhu, Y. L., Hu, F. H., Wang, Y. Y., Huang, N. P., & Xiao, Z. D. (2013). Dynamics of exosome internalization and trafficking. Journal of Cellular Physiology, 228, 1487–1495. https://doi.org/10.1002/jcp.24304.
Feng, D., Zhao, W. L., Ye, Y. Y., Bai, X. C., Liu, R. Q., Chang, L. F., et al. (2010). Cellular internalization of exosomes occurs through phagocytosis. Traffic, 11, 675–687. https://doi.org/10.1111/j.1600-0854.2010.01041.x.
Freeman, S. A., & Grinstein, S. (2014). Phagocytosis: receptors, signal integration, and the cytoskeleton. Immunological Reviews, 262, 193–215. https://doi.org/10.1111/imr.12212.
Nakase, I., Kobayashi, N. B., Takatani-Nakase, T., & Yoshida, T. (2015). Active macropinocytosis induction by stimulation of epidermal growth factor receptor and oncogenic Ras expression potentiates cellular uptake efficacy of exosomes. Scientific Reports, 5, 10300. https://doi.org/10.1038/srep10300.
Thuma, F., & Zöller, M. (2014). Outsmart tumor exosomes to steal the cancer initiating cell its niche. Seminars in Cancer Biology, 28, 39–50. https://doi.org/10.1016/j.semcancer.2014.02.011.
Nanbo, A., Kawanishi, E., Yoshida, R., & Yoshiyama, H. (2013). Exosomes derived from Epstein-Barr virus-infected cells are internalized via caveola-dependent endocytosis and promote phenotypic modulation in target cells. Journal of Virology, 87, 10334–10347. https://doi.org/10.1128/JVI.01310-13.
Lakkaraju, A., & Rodriguez-Boulan, E. (2008). Itinerant exosomes: emerging roles in cell and tissue polarity. Trends in Cell Biology, 18, 199–209. https://doi.org/10.1016/j.tcb.2008.03.002.
Leone, D. A., Peschel, A., Brown, M., Schachner, H., Ball, M. J., Gyuraszova, M., et al. (2017). Surface LAMP-2 is an endocytic receptor that diverts antigen internalized by human dendritic cells into highly immunogenic exosomes. Journal of Immunology, 199, 531–546. https://doi.org/10.4049/jimmunol.1601263.
Holder, B., Jones, T., Sancho Shimizu, V., Rice, T. F., Donaldson, B., Bouqueau, M., et al. (2016). Macrophage exosomes induce placental inflammatory cytokines: a novel mode of maternal-placental messaging. Traffic, 17, 168–178. https://doi.org/10.1111/tra.12352.
Heusermann, W., Hean, J., Trojer, D., Steib, E., von Bueren, S., Graff-Meyer, A., et al. (2016). Exosomes surf on filopodia to enter cells at endocytic hot spots, traffic within endosomes, and are targeted to the ER. The Journal of Cell Biology, 213, 173–184. https://doi.org/10.1083/jcb.201506084.
Svensson, K. J., Kucharzewska, P., Christianson, H. C., Sköld, S., Löfstedt, T., Johansson, M. C., et al. (2011). Hypoxia triggers a proangiogenic pathway involving cancer cell microvesicles and PAR-2-mediated heparin-binding EGF signaling in endothelial cells. Proceedings of the National Academy of Sciences of the United States of America, 108(32), 13147–13152. https://doi.org/10.1073/pnas.1104261108.
Zhang, X., Wang, X., Zhu, H., Kranias, E. G., Tang, Y., Peng, T., et al. (2012). Hsp20 functions as a novel cardiokine in promoting angiogenesis via activation of VEGFR2. PLoS One, 7(3), e32765. https://doi.org/10.1371/journal.pone.0032765.
Arscott, W. T., Tandle, A. T., Zhao, S., Shabason, J. E., Gordon, I. K., Schlaff, C. D., et al. (2013). Ionizing radiation and glioblastoma exosomes: implications in tumor biology and cell migration. Translational Oncology, 6(6), 638–648.
Atay, S., Gercel-Taylor, C., & Taylor, D. D. (2011). Human trophoblast-derived exosomal fibronectin induces pro-inflammatory IL-1β production by macrophages. American Journal of Reproductive Immunology, 66(4), 259–269. https://doi.org/10.1111/j.1600-0897.2011.00995.x.
Fedele, C., Singh, A., Zerlanko, B. J., Iozzo, R. V., & Languino, L. R. (2015). The αvβ6 integrin is transferred intercellularly via exosomes. The Journal of Biological Chemistry, 290, 4545–4551. https://doi.org/10.1074/jbc.C114.617662.
Gu, X., Erb, U., Büchler, M. W., & Zöller, M. (2015). Improved vaccine efficacy of tumor exosome compared to tumor lysate loaded dendritic cells in mice. International Journal of Cancer, 136, E74–E84. https://doi.org/10.1002/ijc.29100.
Lamichhane, T. N., Jeyaram, A., Patel, D. B., Parajuli, B., Livingston, N. K., Arumugasaamy, N., et al. (2016). Oncogene knockdown via active loading of small RNAs into extracellular vesicles by sonication. Cellular and Molecular Bioengineering, 9, 315–324. https://doi.org/10.1007/s12195-016-0457-4.
Saari, H., Lázaro-Ibáñez, E., Viitala, T., Vuorimaa-Laukkanen, E., Siljander, P., & Yliperttula, M. (2015). Microvesicle- and exosome-mediated drug delivery enhances the cytotoxicity of paclitaxel in autologous prostate cancer cells. Journal of Controlled Release, 220(Pt B), 727–737. https://doi.org/10.1016/j.jconrel.2015.09.031.
Kapustin, A. N., Schoppet, M., Schurgers, L. J., Reynolds, J. L., McNair, R., Heiss, A., et al. (2017). Prothrombin loading of vascular smooth muscle cell-derived exosomes regulates coagulation and calcification. Arteriosclerosis, Thrombosis, and Vascular Biology, 37, e22–e32. https://doi.org/10.1161/ATVBAHA.116.308886.
Zarovni, N., Corrado, A., Guazzi, P., Zocco, D., Lari, E., Radano, G., et al. (2015). Integrated isolation and quantitative analysis of exosome shuttled proteins and nucleic acids using immunocapture approaches. Methods, 87, 46–58. https://doi.org/10.1016/j.ymeth.2015.05.028.
Théry, C., Duban, L., Segura, E., Véron, P., Lantz, O., & Amigorena, S. (2002). Indirect activation of naïve CD4+ T cells by dendritic cell-derived exosomes. Nature Immunology, 3, 1156–1162. https://doi.org/10.1038/ni854.
Simhadri, V. R., Reiners, K. S., Hansen, H. P., Topolar, D., Simhadri, V. L., Nohroudi, K., et al. (2008). Dendritic cells release HLA-B-associated transcript-3 positive exosomes to regulate natural killer function. PLoS One, 3, e3377. https://doi.org/10.1371/journal.pone.0003377.
Viaud, S., Terme, M., Flament, C., Taieb, J., André, F., Novault, S., et al. (2009). Dendritic cell-derived exosomes promote natural killer cell activation and proliferation: a role for NKG2D ligands and IL-15Ralpha. PLoS, 4(3), e4942. https://doi.org/10.1371/journal.pone.0004942.
Vulpis, E., Cecere, F., Molfetta, R., Soriani, A., Fionda, C., Peruzzi, G., et al. (2017). Genotoxic stress modulates the release of exosomes from multiple myeloma cells capable of activating NK cell cytokine production: role of HSP70/TLR2/NF-kB axis. Oncoimmunology, 6, e1279372. https://doi.org/10.1080/2162402X.2017.1279372.
Budnik, V., Ruiz-Cañada, C., & Wendler, F. (2016). Extracellular vesicles round off communication in the nervous system. Nature Reviews. Neuroscience, 17, 160–172. https://doi.org/10.1038/nrn.2015.29.
Gong, J., Körner, R., Gaitanos, L., & Klein, R. (2016). Exosomes mediate cell contact-independent ephrin-Eph signaling during axon guidance. The Journal of Cell Biology, 214, 35–44. https://doi.org/10.1083/jcb.201601085.
Fitzgerald, T. L., & McCubrey, J. A. (2014). Pancreatic cancer stem cells: association with cell surface markers, prognosis, resistance, metastasis and treatment. Adv Biol Regul, 56, 45–50. https://doi.org/10.1016/j.jbior.2014.05.001.
Cherciu, I., Bărbălan, A., Pirici, D., Mărgăritescu, C., & Săftoiu, A. (2014). Stem cells, colorectal cancer and cancer stem cell markers correlations. Current Health Sciences Journal, 40(3), 153–161. https://doi.org/10.12865/CHSJ.40.03.01.
Gesierich, S., Paret, C., Hildebrand, D., Weitz, J., Zgraggen, K., Schmitz-Winnenthal, F. H., et al. (2005). Colocalization of the tetraspanins, CO-029 and CD151, with integrins in human pancreatic adenocarcinoma: impact on cell motility. Clinical Cancer Research, 11(8), 2840–2852.
Claas, C., Wahl, J., Orlicky, D. J., Karaduman, H., Schnölzer, M., Kempf, T., et al. (2005). The tetraspanin D6.1A and its molecular partners on rat carcinoma cells. The Biochemical Journal, 389(Pt 1), 99–110.
Kanatsu-Shinohara, M., Takashima, S., Ishii, K., & Shinohara, T. (2011). Dynamic changes in EPCAM expression during spermatogonial stem cell differentiation in the mouse testis. PLoS One, 6(8), e23663. https://doi.org/10.1371/journal.pone.0023663.
Le Naour, F., André, M., Greco, C., Billard, M., Sordat, B., Emile, J. F., et al. (2006). Profiling of the tetraspanin web of human colon cancer cells. Molecular & Cellular Proteomics, 5(5), 845–857.
Margadant, C., Frijns, E., Wilhelmsen, K., & Sonnenberg, A. (2008). Regulation of hemidesmosome disassembly by growth factor receptors. Current Opinion in Cell Biology, 20(5), 589–596. https://doi.org/10.1016/j.ceb.2008.05.001.
Ladwein, M., Pape, U. F., Schmidt, D. S., Schnölzer, M., Fiedler, S., Langbein, L., et al. (2005). The cell-cell adhesion molecule EpCAM interacts directly with the tight junction protein claudin-7. Experimental Cell Research, 309(2), 345–357.
Kuhn, S., Koch, M., Nübel, T., Ladwein, M., Antolovic, D., Klingbeil, P., et al. (2007). A complex of EpCAM, claudin-7, CD44 variant isoforms, and tetraspanins promotes colorectal cancer progression. Molecular Cancer Research, 5(6), 553–567.
Okada, T., Nakamura, T., Watanabe, T., Onoda, N., Ashida, A., Okuyama, R., et al. (2014). Coexpression of EpCAM, CD44 variant isoforms and claudin-7 in anaplastic thyroid carcinoma. PLoS One, 9(4), e94487. https://doi.org/10.1371/journal.pone.0094487.
Wu, C. J., Mannan, P., Lu, M., & Udey, M. C. (2013). Epithelial cell adhesion molecule (EpCAM) regulates claudin dynamics and tight junctions. The Journal of Biological Chemistry, 288(17), 12253–12268. https://doi.org/10.1074/jbc.M113.457499.
Matzke-Ogi, A., Jannasch, K., Shatirishvili, M., Fuchs, B., Chiblak, S., Morton, J., et al. (2016). Inhibition of tumor growth and metastasis in pancreatic cancer models by interference with CD44v6 signaling., 150(2), Gastroenterology, 513–Gastroent525.e10. https://doi.org/10.1053/j.gastro.2015.10.020.
Parton, R. G., & del Pozo, M. A. (2013). Caveolae as plasma membrane sensors, protectors and organizers. Nature Reviews. Molecular Cell Biology, 14(2), 98–112. https://doi.org/10.1038/nrm3512.
Lampe, M., Vassilopoulos, S., & Merrifield, C. (2016). Clathrin coated pits, plaques and adhesion. Journal of Structural Biology, 196(1), 48–56. https://doi.org/10.1016/j.jsb.2016.07.009.
Marsh, D., Horváth, L. I., Swamy, M. J., Mantripragada, S., & Kleinschmidt, J. H. (2002). Interaction of membrane-spanning proteins with peripheral and lipid-anchored membrane proteins: perspectives from protein-lipid interactions (review). Molecular Membrane Biology, 19(4), 247–255.
Bottini, M., Mebarek, S., Anderson, K. L., Strzelecka-Kiliszek, A., Bozycki, L., Simão, A. M. S., et al. (2018). Matrix vesicles from chondrocytes and osteoblasts: their biogenesis, properties, functions and biomimetic models. Biochimica et Biophysica Acta, 1862(3), 532–546. https://doi.org/10.1016/j.bbagen.2017.11.005.
Andreu, Z., & Yáñez-Mó, M. (2014). Tetraspanins in extracellular vesicle formation and function. Frontiers in Immunology, 5, 442. https://doi.org/10.3389/fimmu.2014.00442.
Stuffers, S., Sem Wegner, C., Stenmark, H., & Brech, A. (2009). Multivesicular endosome biogenesis in the absence of ESCRTs. Traffic, 925–937. https://doi.org/10.1111/j.1600-0854.2009.00920.x.
van Niel, G., Bergam, P., Di Cicco, A., Hurbain, I., Lo Cicero, A., Dingli, F., et al. (2015). Apolipoprotein E regulates amyloid formation within endosomes of pigment cells. Cell Reports, 13(1), 43–51. https://doi.org/10.1016/j.celrep.2015.08.057.
Buschow, S. I., Nolte-'t Hoen, E. N., van Niel, G., Pols, M. S., ten Broeke, T., Lauwen, M., et al. (2009). MHC II in dendritic cells is targeted to lysosomes or T cell-induced exosomes via distinct multivesicular body pathways. Traffic, 10(10), 1528–1542. https://doi.org/10.1111/j.1600-0854.2009.00963.x.
Zimmerman, B., Kelly, B., McMillan, B. J., Seegar, T. C. M., Dror, R. O., Kruse, A. C., et al. (2016). Crystal structure of a full-length human tetraspanin reveals a cholesterol-binding pocket. Cell, 167(4), 1041–1051.e11. https://doi.org/10.1016/j.cell.2016.09.056.
Odintsova, E., van Niel, G., Conjeaud, H., Raposo, G., Iwamoto, R., Mekada, E., et al. (2013). Metastasis suppressor tetraspanin CD82/KAI1 regulates ubiquitylation of epidermal growth factor receptor. The Journal of Biological Chemistry, 288(36), 26323–26334. https://doi.org/10.1074/jbc.M112.439380.
Gräßel, L., Fast, L. A., Scheffer, K. D., Boukhallouk, F., Spoden, G. A., Tenzer, S., et al. (2016). The CD63-syntenin-1 complex controls post-endocytic trafficking of oncogenic human papillomaviruses. Scientific Reports, 6, 32337. https://doi.org/10.1038/srep32337.
Li, X., Zhao, H., Gu, J., & Zheng, L. (2015). Prognostic value of cancer stem cell marker CD133 expression in pancreatic ductal adenocarcinoma (PDAC): a systematic review and meta-analysis. International Journal of Clinical and Experimental Pathology, 8(10), 12084–12092 eCollection 2015.
Schulze, U., Brast, S., Grabner, A., Albiker, C., Snieder, B., Holle, S., et al. (2017). Tetraspanin CD63 controls basolateral sorting of organic cation transporter 2 in renal proximal tubules. The FASEB Journal, 31(4), 1421–1433. https://doi.org/10.1096/fj.201600901R.
Zumaquero, E., Muñoz, P., Cobo, M., Lucena, G., Pavón, E. J., Martín, A., et al. (2010). Exosomes from human lymphoblastoid B cells express enzymatically active CD38 that is associated with signaling complexes containing CD81, Hsc-70 and Lyn. Experimental Cell Research, 316(16), 2692–2706. https://doi.org/10.1016/j.yexcr.2010.05.032.
Erb, U., Zhao, K., Wang, Z., Xiao, L., & Zöller, M. (2017). Murine and human pancreatic tumor exosome recovery in mouse serum: diagnostic and prognostic potential and target cell delivery. Cancer Letters, 403, 1–12. https://doi.org/10.1016/j.canlet.2017.06.005.
Hurwitz, S. N., Nkosi, D., Conlon, M. M., York, S. B., Liu, X., Tremblay, D. C., et al. (2017). CD63 regulates Epstein-Barr virus LMP1 exosomal packaging, enhancement of vesicle production, and noncanonical NF-κB signaling. Journal of Virology, 91(5), e02251–e02216. https://doi.org/10.1128/JVI.02251-16.
Kristiansen, G., Sammar, M., & Altevogt, P. (2004). Tumour biological aspects of CD24, a mucin-like adhesion molecule. Journal of Molecular Histology, 35(3), 255–262.
Corbeil, D., Joester, A., Fargeas, C. A., Jászai, J., Garwood, J., Hellwig, A., et al. (2009). Expression of distinct splice variants of the stem cell marker prominin-1 (CD133) in glial cells. Glia, 57(8), 860–874. https://doi.org/10.1002/glia.20812.
Trajkovic, K., Hsu, C., Chiantia, S., Rajendran, L., Wenzel, D., Wieland, F., et al. (2008). Ceramide triggers budding of exosome vesicles into multivesicular endosomes. Science, 319(5867), 1244–1247. https://doi.org/10.1126/science.1153124.
Badawy, S. M. M., Okada, T., Kajimoto, T., Hirase, M., Matovelo, S. A., Nakamura, S., et al. (2018). Extracellular α-synuclein drives sphingosine 1-phosphate receptor subtype 1 out of lipid rafts, leading to impaired inhibitory G-protein signaling. The Journal of Biological Chemistry, 293(21), 8208–8216. https://doi.org/10.1074/jbc.RA118.001986.
Hartman, N. C., & Groves, J. T. (2011). Signaling clusters in the cell membrane. Current Opinion in Cell Biology, 23(4), 370–376. https://doi.org/10.1016/j.ceb.2011.05.003.
Gonnord, P., Blouin, C. M., & Lamaze, C. (2012). Membrane trafficking and signaling: two sides of the same coin. Seminars in Cell & Developmental Biology, 23(2), 154–164. https://doi.org/10.1016/j.semcdb.2011.11.002.
Head, B. P., Patel, H. H., & Insel, P. A. (2014). Interaction of membrane/lipid rafts with the cytoskeleton: impact on signaling and function: membrane/lipid rafts, mediators of cytoskeletal arrangement and cell signaling. Biochimica et Biophysica Acta, 1838(2), 532–545. https://doi.org/10.1016/j.bbamem.2013.07.018.
Melo, S. A., Sugimoto, H., O'Connell, J. T., Kato, N., Villanueva, A., Vidal, A., et al. (2014). Cancer exosomes perform cell-independent microRNA biogenesis and promote tumorigenesis. Cancer Cell, 26(5), 707–721. https://doi.org/10.1016/j.ccell.2014.09.005.
Nag, A., & Steitz, J. A. (2012). Tri-snRNP-associated proteins interact with subunits of the TRAMP and nuclear exosome complexes, linking RNA decay and pre-mRNA splicing. RNA Biology, 9(3), 334–342. https://doi.org/10.4161/rna.19431.
Chang, A. Y., Castel, S. E., Ernst, E., Kim, H. S., & Martienssen, R. A. (2017). The conserved RNA binding cyclophilin, Rct1, regulates small RNA biogenesis and splicing independent of heterochromatin assembly. Cell Reports, 19(12), 2477–2489. https://doi.org/10.1016/j.celrep.2017.05.086.
Tran, N. (2016). Cancer exosomes as miRNA factories. Trends Cancer, 2(7), 329–331. https://doi.org/10.1016/j.trecan.2016.05.008.
Gao, T., Shu, J., & Cui, J. (2018). A systematic approach to RNA-associated motif discovery. BMC Genomics, 19(1), 146. https://doi.org/10.1186/s12864-018-4528-x.
Shapiro, I. M., Landis, W. J., & Risbud, M. V. (2015). Matrix vesicles: are they anchored exosomes? Bone, 79, 29–36. https://doi.org/10.1016/j.bone.2015.05.013.
Belov, L., Matic, K. J., Hallal, S., Best, O. G., Mulligan, S. P., & Christopherson, R. I. (2016). Extensive surface protein profiles of extracellular vesicles from cancer cells may provide diagnostic signatures from blood samples. J Extracell Vesicles, 5, 25355. https://doi.org/10.3402/jev.v5.25355.
Chase, S. D., Magnani, J. L., & Simon, S. I. (2012). E-selectin ligands as mechanosensitive receptors on neutrophils in health and disease. Annals of Biomedical Engineering, 40(4), 849–859. https://doi.org/10.1007/s10439-011-0507-y.
Levy, S., Todd, S. C., & Maecker, H. T. (1998). CD81 (TAPA-1): a molecule involved in signal transduction and cell adhesion in the immune system. Annual Review of Immunology, 16, 89–109.
Zech, D., Rana, S., Büchler, M. W., & Zöller, M. (2012). Tumor-exosomes and leukocyte activation: an ambivalent crosstalk. Cell Communication and Signaling: CCS, 10(1), 37. https://doi.org/10.1186/1478-811X-10-37.
Lau, L. M., Wee, J. L., Wright, M. D., Moseley, G. W., Hogarth, P. M., Ashman, L. K., et al. (2004). The tetraspanin superfamily member CD151 regulates outside-in integrin alphaIIbbeta3 signaling and platelet function. Blood, 104(8), 2368–2375.
Zhao, K., Erb, U., Hackert, T., Zöller, M., & Yue, S. (2018). Distorted leukocyte migration, angiogenesis, wound repair and metastasis in Tspan8 and Tspan8/CD151 double knockout mice indicate complementary activities of Tspan8 and CD51. Biochimica et Biophysica Acta, 1865(2), 379–391. https://doi.org/10.1016/j.bbamcr.2017.11.007.
Yue, S., Mu, W., Erb, U., & Zöller, M. (2015). The tetraspanins CD151 and Tspan8 are essential exosome components for the crosstalk between cancer initiating cells and their surrounding. Oncotarget, 6(4), 2366–2384.
Kyuno, D., Zhao, K., Bauer, N., Ryschich, E., & Zöller, M. (2018). Therapeutic targeting cancer-initiating cell markers by exosome miRNA: efficacy and functional consequences exemplified for claudin7 and EpCAM. Translational Oncology, 12(2), 191–199. https://doi.org/10.1016/j.tranon.2018.08.021.
Henne, W. M., Liou, J., & Emr, S. D. (2015). Molecular mechanisms of inter-organelle ER-PM contact sites. Current Opinion in Cell Biology, 35, 123–130. https://doi.org/10.1016/j.ceb.2015.05.001.
Blank, F., Wehrli, M., Lehmann, A., Baum, O., Gehr, P., von Garnier, C., et al. (2011). Macrophages and dendritic cells express tight junction proteins and exchange particles in an in vitro model of the human airway wall. Immunobiology, 216(1–2), 86–95. https://doi.org/10.1016/j.imbio.2010.02.006.
Nelson, G. M., Padera, T. P., Garkavtsev, I., Shioda, T., & Jain, R. K. (2007). Differential gene expression of primary cultured lymphatic and blood vascular endothelial cells. Neoplasia, 9(12), 1038–1045.
Nogués, L., Benito-Martin, A., Hergueta-Redondo, M., & Peinado, H. (2018). The influence of tumour-derived extracellular vesicles on local and distal metastatic dissemination. Molecular Aspects of Medicine, 60, 15–26. https://doi.org/10.1016/j.mam.2017.11.012.
Li, M., Lu, Y., Xu, Y., Wang, J., Zhang, C., Du, Y., et al. (2018). Horizontal transfer of exosomal CXCR4 promotes murine hepatocarcinoma cell migration, invasion and lymphangiogenesis. Gene, S0378-1119(18), 30787-X. https://doi.org/10.1016/j.gene.2018.07.018.
Shen, L., Weber, C. R., & Turner, J. R. (2008). The tight junction protein complex undergoes rapid and continuous molecular remodeling at steady state. The Journal of Cell Biology, 181(4), 683–695. https://doi.org/10.1083/jcb.200711165.
Rilla, K., Siiskonen, H., Tammi, M., & Tammi, R. (2014). Hyaluronan-coated extracellular vesicles—a novel link between hyaluronan and cancer. Advances in Cancer Research, 123, 121–148. https://doi.org/10.1016/B978-0-12-800092-2.00005-8.
Purushothaman, A., Bandari, S. K., Liu, J., Mobley, J. A., Brown, E. E., & Sanderson, R. D. (2016). Fibronectin on the surface of myeloma cell-derived exosomes mediates exosome-cell interactions. The Journal of Biological Chemistry, 291(4), 1652–1663. https://doi.org/10.1074/jbc.M115.686295.
Dismuke, W. M., Klingeborn, M., & Stamer, W. D. (2016). Mechanism of fibronectin binding to human trabecular meshwork exosomes and its modulation by dexamethasone. PLoS One, 11(10), e0165326. https://doi.org/10.1371/journal.pone.0165326 eCollection 2016.
Shimoda, M., & Khokha, R. (2013). Proteolytic factors in exosomes. Proteomics, 13(10–11), 1624–1636. https://doi.org/10.1002/pmic.201200458.
Sevenich, L., & Joyce, J. A. (2014). Pericellular proteolysis in cancer. Genes & Development, 28(21), 2331–2347. https://doi.org/10.1101/gad.250647.114.
Silva, A. M., Teixeira, J. H., Almeida, M. I., Gonçalves, R. M., Barbosa, M. A., & Santos, S. G. (2017). Extracellular vesicles: immunomodulatory messengers in the context of tissue repair/regeneration. European Journal of Pharmaceutical Sciences, 98, 86–95. https://doi.org/10.1016/j.ejps.2016.09.017.
Than, U. T. T., Guanzon, D., Leavesley, D., & Parker, T. (2017). association of extracellular membrane vesicles with cutaneous wound healing. International Journal of Molecular Sciences, 18(5), E956. https://doi.org/10.3390/ijms18050956.
Bandari, S. K., Purushothaman, A., Ramani, V. C., Brinkley, G. J., Chandrashekar, D. S., Varambally, S., et al. (2018). Chemotherapy induces secretion of exosomes loaded with heparanase that degrades extracellular matrix and impacts tumor and host cell behavior. Matrix Biology, 65, 104–118. https://doi.org/10.1016/j.matbio.2017.09.001.
Shi, F., & Sottile, J. (2011). MT1-MMP regulates the turnover and endocytosis of extracellular matrix fibronectin. Journal of Cell Science, 124(Pt 23), 4039–4050. https://doi.org/10.1242/jcs.087858.
Jessen, T. N., & Jessen, J. R. (2017). VANGL2 interacts with integrin αv to regulate matrix metalloproteinase activity and cell adhesion to the extracellular matrix. Experimental Cell Research, 361(2), 265–276. https://doi.org/10.1016/j.yexcr.2017.10.026.
Nalivaeva, N. N., Belyaev, N. D., Kerridge, C., & Turner, A. J. (2014). Amyloid-clearing proteins and their epigenetic regulation as a therapeutic target in Alzheimer’s disease. Frontiers in Aging Neuroscience, 6, 235. https://doi.org/10.3389/fnagi.2014.00235.
Jung, T., Gross, W., & Zöller, M. (2011). CD44v6 coordinates tumor matrix-triggered motility and apoptosis resistance. The Journal of Biological Chemistry, 286(18), 15862–15874. https://doi.org/10.1074/jbc.M110.208421.
Quesenberry, P. J., Aliotta, J., Deregibus, M. C., & Camussi, G. (2015). Role of extracellular RNA-carrying vesicles in cell differentiation and reprogramming. Stem Cell Research & Therapy, 6, 153. https://doi.org/10.1186/s13287-015-0150-x.
Kanada, M., Bachmann, M. H., Hardy, J. W., Frimannson, D. O., Bronsart, L., Wang, A., et al. (2015). Differential fates of biomolecules delivered to target cells via extracellular vesicles. Proceedings of the National Academy of Sciences of the United States of America, 112(12), E1433–E1442. https://doi.org/10.1073/pnas.1418401112.
Shin, S. J., Smith, J. A., Rezniczek, G. A., Pan, S., Chen, R., Brentnall, T. A., et al. (2013). Unexpected gain of function for the scaffolding protein plectin due to mislocalization in pancreatic cancer. Proceedings of the National Academy of Sciences of the United States of America, 110(48), 19414–19419. https://doi.org/10.1073/pnas.1309720110.
Rana, S., Malinowska, K., & Zöller, M. (2013). Exosomal tumor microRNA modulates premetastatic organ cells. Neoplasia, 15(3), 281–295.
Wang, Z., Zhao, K., Hackert, T., & Zöller, M. (2018). CD44/CD44v6 a reliable companion in cancer-initiating cell maintenance and tumor progression. Frontiers in Cell and Development Biology, 6, 97. https://doi.org/10.3389/fcell.2018.00097.
Song, X., Ding, Y., Liu, G., Yang, X., Zhao, R., Zhang, Y., et al. (2016). Cancer cell-derived exosomes induce mitogen-activated protein kinase-dependent monocyte survival by transport of functional receptor tyrosine kinases. The Journal of Biological Chemistry, 291(16), 8453–8464. https://doi.org/10.1074/jbc.M116.716316.
Zhang, H., Deng, T., Liu, R., Bai, M., Zhou, L., Wang, X., et al. (2017). Exosome-delivered EGFR regulates liver microenvironment to promote gastric cancer liver metastasis. Nature Communications, 8, 15016. https://doi.org/10.1038/ncomms15016.
Bendinelli, P., Maroni, P., Matteucci, E., & Desiderio, M. A. (2017). Epigenetic regulation of HGF/Met receptor axis is critical for the outgrowth of bone metastasis from breast carcinoma. Cell Death & Disease, 8(2), e2578. https://doi.org/10.1038/cddis.2016.403.
Yang, W. W., Yang, L. Q., Zhao, F., Chen, C. W., Xu, L. H., Fu, J., et al. (2017). Epiregulin promotes lung metastasis of salivary adenoid cystic carcinoma. Theranostics, 7(15), 3700–3714. https://doi.org/10.7150/thno.19712.
Kwon, S. H., Liu, K. D., & Mostov, K. E. (2014). Intercellular transfer of GPRC5B via exosomes drives HGF-mediated outward growth. Current Biology, 24(2), 199–204. https://doi.org/10.1016/j.cub.2013.12.010.
Fuchs, K., Hippe, A., Schmaus, A., Homey, B., Sleeman, J. P., & Orian-Rousseau, V. (2013). Opposing effects of high- and low-molecular weight hyaluronan on CXCL12-induced CXCR4 signaling depend on CD44. Cell Death & Disease, 4, e819. https://doi.org/10.1038/cddis.2013.364.
Roscic-Mrkic, B., Fischer, M., Leemann, C., Manrique, A., Gordon, C. J., Moore, J. P., et al. (2003). RANTES (CCL5) uses the proteoglycan CD44 as an auxiliary receptor to mediate cellular activation signals and HIV-1 enhancement. Blood, 102(4), 1169–1177.
Zhu, B., Suzuki, K., Goldberg, H. A., Rittling, S. R., Denhardt, D. T., McCulloch, C. A., et al. (2004). Osteopontin modulates CD44-dependent chemotaxis of peritoneal macrophages through G-protein-coupled receptors: evidence of a role for an intracellular form of osteopontin. Journal of Cellular Physiology, 198(1), 155–167.
Che, S. P. Y., Park, J. Y., & Stokol, T. (2017). Tissue factor-expressing tumor-derived extracellular vesicles activate quiescent endothelial cells via protease-activated receptor-1. Frontiers in Oncology, 7, 261. https://doi.org/10.3389/fonc.2017.00261.
Gilliam, D. T., Menon, V., Bretz, N. P., & Pruszak, J. (2017). The CD24 surface antigen in neural development and disease. Neurobiology of Disease, 99, 133–144. https://doi.org/10.1016/j.nbd.2016.12.011.
Lim, J., Lee, K. M., Shim, J., & Shin, I. (2014). CD24 regulates stemness and the epithelial to mesenchymal transition through modulation of Notch1 mRNA stability by p38MAPK. Archives of Biochemistry and Biophysics, 558, 120–126. https://doi.org/10.1016/j.abb.2014.06.022.
Lee, T. K., Castilho, A., Cheung, V. C., Tang, K. H., Ma, S., & Ng, I. O. (2011). CD24(+) liver tumor-initiating cells drive self-renewal and tumor initiation through STAT3-mediated NANOG regulation. Cell Stem Cell, 9(1), 50–63. https://doi.org/10.1016/j.stem.2011.06.005.
Takenobu, H., Shimozato, O., Nakamura, T., Ochiai, H., Yamaguchi, Y., Ohira, M., et al. (2011). CD133 suppresses neuroblastoma cell differentiation via signal pathway modification. Oncogene, 30(1), 97–105. https://doi.org/10.1038/onc.2010.383.
Zhang, M., Liu, Y., Feng, H., Bian, X., Zhao, W., Yang, Z., et al. (2013). CD133 affects the invasive ability of HCT116 cells by regulating TIMP-2. The American Journal of Pathology, 182(2), 565–576. https://doi.org/10.1016/j.ajpath.2012.10.015.
Lee, J. W., Lee, Y. C., Na, S. Y., Jung, D. J., & Lee, S. K. (2001). Transcriptional coregulators of the nuclear receptor superfamily: coactivators and corepressors. Cellular and Molecular Life Sciences, 58(2), 289–297.
Mazzi, S., Lordier, L., Debili, N., Raslova, H., & Vainchenker, W. (2018). Megakaryocyte and polyploidization. Experimental Hematology, 57, 1–13. https://doi.org/10.1016/j.exphem.2017.10.001.
Heneberg, P. (2016). Paracrine tumor signaling induces transdifferentiation of surrounding fibroblasts. Critical Reviews in Oncology/Hematology, 97, 303–311. https://doi.org/10.1016/j.critrevonc.2015.09.008.
Cho, J. A., Park, H., Lim, E. H., & Lee, K. W. (2012). Exosomes from breast cancer cells can convert adipose tissue-derived mesenchymal stem cells into myofibroblast-like cells. International Journal of Oncology, 40(1), 130–138. https://doi.org/10.3892/ijo.2011.1193.
Record, M., Poirot, M., & Silvente-Poirot, S. (2014). Emerging concepts on the role of exosomes in lipid metabolic diseases. Biochimie, 96, 67–74. https://doi.org/10.1016/j.biochi.2013.06.016.
Fonseca, P., Vardaki, I., Occhionero, A., & Panaretakis, T. (2016). Metabolic and signaling functions of cancer cell-derived extracellular vesicles. International Review of Cell and Molecular Biology, 326, 175–199. https://doi.org/10.1016/bs.ircmb.2016.04.004.
Skotland, T., Sandvig, K., & Llorente, A. (2017). Lipids in exosomes: current knowledge and the way forward. Progress in Lipid Research, 66, 30–41. https://doi.org/10.1016/j.plipres.2017.03.001.
García-González, V., Díaz-Villanueva, J. F., Galindo-Hernández, O., Martínez-Navarro, I., Hurtado-Ureta, G., & Pérez-Arias, A. A. (2018). Ceramide metabolism balance, a multifaceted factor in critical steps of breast cancer development. International Journal of Molecular Sciences, 19(9), E2527. https://doi.org/10.3390/ijms19092527.
Hsu, M. C., & Hung, W. C. (2018). Pyruvate kinase M2 fuels multiple aspects of cancer cells: from cellular metabolism, transcriptional regulation to extracellular signaling. Molecular Cancer, 17(1), 35. https://doi.org/10.1186/s12943-018-0791-3.
Alcayaga-Miranda, F., González, P. L., Lopez-Verrilli, A., Varas-Godoy, M., Aguila-Díaz, C., Contreras, L., et al. (2016). Prostate tumor-induced angiogenesis is blocked by exosomes derived from menstrual stem cells through the inhibition of reactive oxygen species. Oncotarget, 7(28), 44462–44477. https://doi.org/10.18632/oncotarget.9852.
van Balkom, B. W., Eisele, A. S., Pegtel, D. M., Bervoets, S., & Verhaar, M. C. (2015). Quantitative and qualitative analysis of small RNAs in human endothelial cells and exosomes provides insights into localized RNA processing, degradation and sorting. J Extracell Vesicles, 4, 26760. https://doi.org/10.3402/jev.v4.26760.
Deng, X., Wu, B., Xiao, K., Kang, J., Xie, J., Zhang, X., et al. (2015). MiR-146b-5p promotes metastasis and induces epithelial-mesenchymal transition in thyroid cancer by targeting ZNRF3. Cellular Physiology and Biochemistry, 35(1), 71–82. https://doi.org/10.1159/000369676.
Rutnam, Z. J., & Yang, B. B. (2012). The non-coding 3’ UTR of CD44 induces metastasis by regulating extracellular matrix functions. Journal of Cell Science, 125(Pt 8), 2075–2085. https://doi.org/10.1242/jcs100818.
Li, J., & Lam, M. (2015). Reproducibility project: cancer biology. Registered report: the microRNA miR-34a inhibits prostate cancer stem cells and metastasis by directly repressing CD44. Elife, 4, e06434. https://doi.org/10.7554/eLife.06434.
Li, X., He, J., Shao, M., Cui, B., Peng, F., Li, J., et al. (2018). Downregulation of miR-218-5p promotes invasion of oral squamous cell carcinoma cells via activation of CD44-ROCK signaling. Biomedicine & Pharmacotherapy, 106, 646–654. https://doi.org/10.1016/j.biopha.2018.06.151.
Al-Othman, N., Hammad, H., & Ahram, M. (2018). Dihydrotestosterone regulates expression of CD44 via miR-328-3p in triple-negative breast cancer cells. Gene, S0378-1119(18), 30755–30758. https://doi.org/10.1016/j.gene.2018.06.094.
Wu, Z. S., Wu, Q., Wang, C. Q., Wang, X. N., Huang, J., Zhao, J. J., et al. (2011). miR-340 inhibition of breast cancer cell migration and invasion through targeting of oncoprotein c-Met. Cancer, 117, 2842–2852. https://doi.org/10.1002/cncr.25860.
Takahashi, H., Ohkuchi, A., Kuwata, T., Usui, R., Baba, Y., Suzuki, H., et al. (2017). Endogenous and exogenous miR-520c-3p modulates CD44-mediated extravillous trophoblast invasion. Placenta, 50, 25–31. https://doi.org/10.1016/j.placenta.2016.12.016.
Shen, W. W., Zeng, Z., Zhu, W. X., & Fu, G. H. (2013). MiR-142-3p functions as a tumor suppressor by targeting CD133, ABCG2, and Lgr5 in colon cancer cells. J Mol Med (Berl), 91(8), 989–1000. https://doi.org/10.1007/s00109-013-1037-x.
Tsuji, S., Kawasaki, Y., Furukawa, S., Taniue, K., Hayashi, T., Okuno, M., et al. (2014). The miR-363-GATA6-Lgr5 pathway is critical for colorectal tumourigenesis. Nature Communications, 5, 3150. https://doi.org/10.1038/ncomms4150.
Zhou, M. K., Liu, X. J., Zhao, Z. G., & Cheng, Y. M. (2015). MicroRNA-100 functions as a tumor suppressor by inhibiting Lgr5 expression in colon cancer cells. Molecular Medicine Reports, 11(4), 2947–2952. https://doi.org/10.3892/mmr.2014.3052.
Ostenfeld, M. S., Jensen, S. G., Jeppesen, D. K., Christensen, L. L., Thorsen, S. B., Stenvang, J., et al. (2016). miRNA profiling of circulating EpCAM(+) extracellular vesicles: promising biomarkers of colorectal cancer. J Extracell Vesicles, 5, 31488. https://doi.org/10.3402/jev.v5.31488.
Ji, H., Chen, M., Greening, D. W., He, W., Rai, A., Zhang, W., et al. (2014). Deep sequencing of RNA from three different extracellular vesicle (EV) subtypes released from the human LIM1863 colon cancer cell line uncovers distinct miRNA-enrichment signatures. PLoS One, 9(10), e110314. https://doi.org/10.1371/journal.pone.0110314.
Hu, Y., Wang, J., Qian, J., Kong, X., Tang, J., Wang, Y., et al. (2014). Long noncoding RNA GAPLINC regulates CD44-dependent cell invasiveness and associates with poor prognosis of gastric cancer. Cancer Research, 74(23), 6890–6902. https://doi.org/10.1158/0008-5472.CAN-14-0686.
Wu, X., He, X., Li, S., Xu, X., Chen, X., & Zhu, H. (2016). Long non-coding RNA ucoo2kmd.1 regulates CD44-dependent cell growth by competing for miR-211-3p in colorectal cancer. PLoS One, 11(3), e0151287. https://doi.org/10.1371/journal.pone.0151287.
Wang, R., Dong, H. X., Zeng, J., Pan, J., & Jin, X. Y. (2018). LncRNA DGCR5 contributes to CSC-like properties via modulating miR-330-5p/CD44 in NSCLC. Journal of Cellular Physiology, 233(9), 7447–7456. https://doi.org/10.1002/jcp.26590.
Ji, J., Tang, J., Deng, L., Xie, Y., Jiang, R., Li, G., et al. (2015). LINC00152 promotes proliferation in hepatocellular carcinoma by targeting EpCAM via the mTOR signaling pathway. Oncotarget, 6(40), 42813–42824. https://doi.org/10.18632/oncotarget.5970.
Zhang, Z.Y., Lu, Y.X., Zhang, Z.Y., Chang, Y.Y., Zheng, L., Yuan, L., et al. (2016). Loss of TINCR expression promotes proliferation, metastasis through activating EpCAM cleavage in colorectal cancer. Oncotarget, 7(16), 22639–22649. doi: 10.18632/oncotarget.8141.
Liu, J., Yang, C., Gu, Y., Li, C., Zhang, H., Zhang, W., et al. (2018). Knockdown of the lncRNA SNHG8 inhibits cell growth in Epstein-Barr virus-associated gastric carcinoma. Cellular & Molecular Biology Letters, 23, 17. https://doi.org/10.1186/s11658-018-0070-8.
Ren, H., Yang, X., Yang, Y., Zhang, X., Zhao, R., Wie, R., et al. (2017). Upregulation of LncRNA BCYRN1 promotes tumor progression and enhances EpCAM expression in gastric carcinoma. Oncotarget, 9(4), 4851–4861. https://doi.org/10.18632/oncotarget.23585.
Jing, N., Huang, T., Guo, H., Yang, J., Li, M., Chen, Z., et al. (2018). LncRNA CASC15 promotes colon cancer cell proliferation and metastasis by regulating the miR-4310/LGR5/Wnt/β-catenin signaling pathway. Molecular Medicine Reports, 18(2), 2269–2276. https://doi.org/10.3892/mmr.2018.9191.
Song, Y. X., Sun, J. X., Zhao, J. H., Yang, Y. C., Shi, J. X., Wu, Z. H., et al. (2017). Non-coding RNAs participate in the regulatory network of CLDN4 via ceRNA mediated miRNA evasion. Nature Communications, 8(1), 289. https://doi.org/10.1038/s41467-017-00304-1.
Chen, T., Xue, H., Lin, R., & Huang, Z. (2017). MiR-34c and PlncRNA1 mediated the function of intestinal epithelial barrier by regulating tight junction proteins in inflammatory bowel disease. Biochemical and Biophysical Research Communications, 486(1), 6–13. https://doi.org/10.1016/j.bbrc.2017.01.115.
Chen, M., Xu, R., Ji, H., Greening, D. W., Rai, A., Izumikawa, K., et al. (2016). Transcriptome and long noncoding RNA sequencing of three extracellular vesicle subtypes released from the human colon cancer LIM1863 cell line. Scientific Reports, 6, 38397. https://doi.org/10.1038/srep38397.
Wang, F., Tian, X., Zhou, J., Wang, G., Yu, W., Li, Z., et al. (2018). A three-lncRNA signature for prognosis prediction of acute myeloid leukemia in patients. Molecular Medicine Reports, 18(2), 1473–1484. https://doi.org/10.3892/mmr.2018.9139.
Xie, S., Ge, Q., Wang, X., Sun, X., & Kang, Y. (2018). Long non-coding RNA ZFAS1 sponges miR-484 to promote cell proliferation and invasion in colorectal cancer. Cell Cycle, 17(2), 154–161. https://doi.org/10.1080/15384101.2017.1407895.
Li, N., Sun, Z. H., Fang, M., Xin, J. Y., & Wan, C. Y. (2017). Long non-coding RNA ZFAS1 sponges miR-486 to promote osteosarcoma cells progression and metastasis in vitro and vivo. Oncotarget, 8(61), 104160–104170. https://doi.org/10.18632/oncotarget.22032.
Xu, W., He, L., Li, Y., Tan, Y., Zhang, F., & Xu, H. (2018). Silencing of lncRNA ZFAS1 inhibits malignancies by blocking Wnt/β-catenin signaling in gastric cancer cells. Bioscience, Biotechnology, and Biochemistry, 82(3), 456–465. https://doi.org/10.1080/09168451.2018.1431518.
Yang, C. H., Zhang, X. Y., Zhou, L. N., Wan, Y., Song, L. L., Gu, W. L., et al. (2018). LncRNA SNHG8 participates in the development of endometrial carcinoma through regulating c-MET expression by miR-152. European Review for Medical and Pharmacological Sciences, 22(6), 1629–1637. https://doi.org/10.26355/eurrev_201803_14698.
Chen, Z., Bu, N., Qiao, X., Zuo, Z., Shu, Y., Liu, Z., et al. (2018). Forkhead box M1 transcriptionally regulates the expression of long noncoding RNAs Snhg8 and Gm26917 to promote proliferation and survival of muscle satellite cells. Stem Cells. https://doi.org/10.1002/stem.2824.
Wan, L., Kong, J., Tang, J., Wu, Y., Xu, E., Lai, M., et al. (2016). HOTAIRM1 as a potential biomarker for diagnosis of colorectal cancer functions the role in the tumour suppressor. Journal of Cellular and Molecular Medicine, 20(11), 2036–2044. https://doi.org/10.1111/jcmm.12892.
Chen, Z. H., Wang, W. T., Huang, W., Fang, K., Sun, Y. M., Liu, S. R., et al. (2017). The lncRNA HOTAIRM1 regulates the degradation of PML-RARA oncoprotein and myeloid cell differentiation by enhancing the autophagy pathway. Cell Death and Differentiation, 24(2), 212–224. https://doi.org/10.1038/cdd.2016.111.
Marín-Béjar, O., Mas, A. M., González, J., Martinez, D., Athie, A., Morales, X., et al. (2017). The human lncRNA LINC-PINT inhibits tumor cell invasion through a highly conserved sequence element. Genome Biology, 18(1), 202. https://doi.org/10.1186/s13059-017-1331-y.
Li, Y., Huang, S., Li, Y., Zhang, W., He, K., Zhao, M., et al. (2016). Decreased expression of LncRNA SLC25A25-AS1 promotes proliferation, chemoresistance, and EMT in colorectal cancer cells. Tumour Biology, 37(10), 14205–14215. https://doi.org/10.1007/s13277-016-5254-0.
Jiao, F., Hu, H., Han, T., Yuan, C., Wang, L., Jin, Z., et al. (2015). Long noncoding RNA MALAT-1 enhances stem cell-like phenotypes in pancreatic cancer cells. International Journal of Molecular Sciences, 16(4), 6677–6693. https://doi.org/10.3390/ijms16046677.
Geng, H., Bu, H. F., Liu, F., Wu, L., Pfeifer, K., Chou, P. M., et al. (2018). In inflamed intestinal tissues and epithelial cells, interleukin 22 signaling increases expression of h19 long noncoding RNA, which promotes mucosal regeneration. Gastroenterology, 155(1), 144–155. https://doi.org/10.1053/j.gastro.2018.03.058.
Abbreviations
A-ISC active ISC, AML acute myeloid leukemia, ASC adult stem cells, BMC bone marrow cells, CAF cancer-associated fibroblasts, ceRNA competing endogenous RNA, CIC cancer-initiating cells/cancer stem cells, CoCa colorectal cancer, DC dendritic cells, DS deep sequencing, EBV Epstein–Barr virus, EC endothelial cells, ECM extracellular matrix, EE early endosome, EMT epithelial–mesenchymal transition, ERM ezrin, radixin, moesin, ESC embryonic stem cells, ESCRT endosomal sorting complex required for transport, EV extracellular vesicles, Exo exosome, GAG glycosaminoglycan, GEM glycolipid-enriched membrane domains, GPCR G protein-coupled receptor, HCC hepatocellular carcinoma, hiPSC human-induced pluripotent SC, ICD intracellular domain, ILV intraluminal vesicle, ISC intestinal SC, kd knockdown, ko knockout, LDL low-density lipoprotein, LN laminin, lnc long nc, LNC lymph node cells, Mϕ macrophage, MHC major histocompatibility complex, miRNA microRNA, MS mass spectrometry, MVB multivesicular body, nc noncoding, NK natural killer cells, NSCLC nonsmall cell lung carcinoma, PaCa pancreatic cancer, PSC pluripotent SC, R receptor, RA retinoic acid, RISC RNA-induced silencing complex, ROS reactive oxygen species, RTK receptor tyrosine kinase, Q-ISC quiescent ISC, SC stem cells, TEM tetraspanin- and glycolipid-enriched membrane microdomain, TEX tumor exosomes, TF transcription factor, TJ tight junction
Funding
This work was supported by the National Natural Science Foundation of China (ZW, NSFC. 81702877) and the German Cancer Research Aid (MZ, 110836). The funding had no impact on the design of the study and on collection, analysis and interpretation of data, and on writing the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Microarray
http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi? acc=GSE119031, GSE119032, -GSE 11903, -GSE50632, http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi? acc=GSE120185).
Deep sequencing
ENA database accession No: PRJEB25446
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 97 kb)
Rights and permissions
About this article
Cite this article
Wang, Z., Zöller, M. Exosomes, metastases, and the miracle of cancer stem cell markers. Cancer Metastasis Rev 38, 259–295 (2019). https://doi.org/10.1007/s10555-019-09793-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10555-019-09793-6