Abstract
Stem cells are unique cells that can self-renew and differentiate into many cell types. Plasticity is a fundamental characteristic of stem cells and it is regulated by reversible epigenetic modifications. Although gene-restriction programs are established during embryonic development when cell lineages are formed, stem cells retain a degree of flexibility that is essential for tissue regeneration. For instance, quiescent adult stem cells can be induced to proliferate and trans-differentiate in response to injury. The same degree of plasticity is observed in cancer, where cancer cells with stem cell characteristics (or cancer stem cells) are formed by transformation of normal stem cells or de-differentiation of somatic cells. Reprogramming experiments with normal somatic cells and cancer cells show that epigenetic landscapes are more plastic than originally thought and that their manipulation can induce changes in cell fate. Our knowledge of stem cell function is still limited and only by understanding the mechanisms regulating developmental potential together with the definition of epigenetic maps of normal and diseased tissues we can reveal the true extent of their plasticity. In return, the control of plastic epigenetic programs in stem cells will allow us to develop effective treatments for degenerative diseases and cancer.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
How are cells in our body programmed to maintain their identity and function throughout life? The answer to this fundamental question is based on important processes that are initiated during embryo development and maintained in adulthood. This book chapter will describe and discuss the mechanisms that control cell and tissue homeostasis and how these are altered in cancer.
Cell identity is established during embryogenesis when the developmental potential of embryonic cells is restricted by differentiation programs that channel their fate to tissue-specific stem cells and specialised cell types. These dynamic events occur in cells with the same genetic information, thus cell fate depends on the epigenetic regulation of that genetic code. “Epigenetics” can be defined as regulation of gene expression that occurs by modifications imposed on the chromatin without change in the DNA sequence (Bird 2007). It is by changes in chromatin organisation that epigenetic modifications establish heritable transcriptional states responsible for the maintenance of cell function.
Epigenetic regulation includes DNA methylation, modification of histone tails and modulation by non coding RNAs (ncRNAs). Together with chromatin remodelling complexes, these modifications control chromatin organisation and regulate gene transcription (Jaenisch and Bird 2003).
DNA methylation is responsible for gene silencing and occurs at position 5 of cytosine (5mC) within CpG dinucleotides present in repetitive sequences and CpG islands in gene promoters and intragenic regions (Ball et al. 2009; Sharma et al. 2010). DNA methylation is maintained or established de novo by the DNA methyltransferases enzymes DNMT1 and DNMT3A/3B/3L, respectively (Bird 2002). Histone modifications comprise a vast range of post-translational modifications, such as acetylation, methylation, phosphorylation, ubiquitylation and ribosylation. These modifications can induce both activation and repression of transcription and their interactions function as a “code” defining cellular states (Turner 2007). Nucleosome remodelling and modulation by ncRNAs are the most important non covalent epigenetic modifications. Non coding RNAs, including microRNAs (miRNAs) and long non coding RNAs (lncRNAs), are single stranded transcripts involved in mRNA degradation and chromatin remodelling (Pauli et al. 2011). While heritable, epigenetic modifications are reversible and their dynamic interplay provides cells with ability to respond to environmental cues. Therefore it is easy to imagine how the epigenetic landscape created by these modifications can regulate phenotype plasticity in different cell types during normal development, but also cause disease if abnormally regulated.
Cancer is a disease characterised by abnormal cell proliferation and it is associated with both genetic lesions and epigenetic abnormalities. Because it can be portrayed as a process of aberrant cell proliferation and differentiation, cancer has been described as “a problem of developmental biology” where a marked resemblance between cancer cells and embryonic cells exists (Pierce and Johnson 1971). Indeed, cancer cells re-initiate epigenetic programs that favour cell growth and survival at the expense of differentiation, thus behaving like undifferentiated embryonic cells and stem cells. As cancer cells depend on those mechanisms that maintain stem cell plasticity (Garraway and Sellers 2006), it is not a coincidence that many tumour suppressor genes that are epigenetically silenced in cancer are developmental genes involved in the regulation of stem cells (Barrero et al. 2010). It is precisely how stem cell plasticity is programmed in development and cancer that will be the focus of our discussion.
2 Epigenetics and Development
Epigenetic modifications regulate the acquisition of totipotency and subsequent progressive restriction of totipotent potential during embryonic development. Acquisition of totipotency is associated with two epigenetic reprogramming events: the formation of the zygote and the germ line (Hemberger et al. 2009). Both developmental stages require resetting of a differentiated epigenetic landscape to establish a new state with augmented developmental potency. Differentiation of somatic cells then requires the establishment of specific epigenetic programs that restrict their potential and maintain lineage memory. This section will describe the epigenetic modifications occurring during embryo development and explain how embryonic developmental potential is programmed in embryonic cells, somatic cells and germ cells.
2.1 Epigenetic Reprogramming During Embryo Development
Embryo development initiates with the fusion of the male and female pronuclei after fertilisation. The formation of the zygote is followed by epigenetic reprogramming of the specialised gametic genomes to ensure that the embryonic genome acquires totipotency, defined as the ability of a cell to form an entire organism. Immediately after fertilisation, the paternal nucleus undergoes profound chromatin remodelling. This involves exchange of protamines for histones in the nucleosomes and active DNA demethylation (Oswald et al. 2000; Mayer et al. 2000; Santos et al. 2002). Although a specific DNA demethylase enzyme has not been identified, a process involving DNA repair through the intermediate 5-hydroxymethylcytosine (5hmC) has been proposed (Wossidlo et al. 2010, 2011; Hemberger et al. 2009). After fusion, the progressive decline in DNA methylation up to the morula stage is due to passive loss of methylated cytosine marks during DNA replication (Howell et al. 2001). Some genomic sequences escape this demetylation, including some repetitive sequences and most imprinted genes (Meissner 2010). Concurrent with DNA demethylation, reprogramming of histone modifications also takes place. The newly incorporated histones in the paternal pronucleus gradually increase active marks, such as acetylation of histone H3 at lysine 9 (H3K9ac), methylation of histone H3 at lysine 4 (H3K4me3), and repressive marks, e.g. methylated histone H3 at lysine 9 (H3K9me1, H3K9me2) and methylated histone H3 at lysine 27 (H3K27me3) (Meissner 2010). Subsequent to the first cleavage divisions, the embryo undergoes segregation of the first two lineages, the inner cell mass (ICM) and the trophectoderm. The cells of the ICM are pluripotent embryonic cells, able to differentiate to all somatic lineages and the germ line (Wray et al. 2010). Epigenetic programming of ICM cells includes de novo DNA methylation, acquisition of H3K9ac, H3K27me3, H3K4me3, H3K9me2 and H3K9me3 (Morgan et al. 2005). Re-establishment of DNA methylation is essential for normal embryonic development, as demonstrated by knockout experiments where deletion of DNMTs and other epigenetic modifiers participating in DNA methylation (LSH and G9a) causes embryonic lethality (Okano et al. 1999; Myant et al. 2011). During gastrulation and differentiation of embryonic cells to somatic lineages, a progressive decrease in plasticity is observed and this is accomplished by a program of epigenetic modifications that restricts cell fate, retains cell memory and confers cellular specialisation.
However, epigenetic restrictions imposed during differentiation are reprogrammed in the germ line. Germ cells derive from embryonic precursors of gametes defined as primordial germ cells (PGC), which are responsible for the development of a new organism in the next generation. Epigenetic reprogramming in the germ line is essential for the generation of a cellular state that will allow totipotency in the newly formed embryo. In addition, reprogramming of PGC ensures an equivalent epigenetic state in both sexes prior to differentiation into mature gametes and erasure of acquired epimutations which could be inherited in the next generation (Allegrucci et al. 2005). PGC are specified in the proximal epiblast and then migrate through the hindgut to the developing gonads. It is during migration and after colonisation of the gonads that extensive epigenetic reprogramming occurs in these cells. This involves loss of H3K9me2 and DNA methylation, and an increase in H3K27me3. It is thought that this epigenetic configuration, enriched in H3K27me3, H3K4me2/me3 and H3K9ac confers PGC with the required plasticity to regain pluripotency (Hemberger et al. 2009). In addition, loss of DNA methylation at imprinted genes ensures erasure of epimutations and correct re-establishment of monoallelic expression for gene dosage in the next generation (Allegrucci et al. 2005; Sasaki and Matsui 2008).
The epigenetic reprogramming and programming of cell plasticity during development is orchestrated by a battery of epigenetic modifiers. Their coordinated action ensures a correct program of cell proliferation and differentiation (Table 24.1).
3 Epigenetic Regulation of Stem Cells
In the previous section we have reviewed the epigenetic events that regulate development and program cell differentiation in the embryo. Although pluripotent cells exist only for a limited period of time before gastrulation, they can be isolated from the embryo and maintained in vitro as embryonic stem cells (ESC). Therefore ESC can be studied as in vitro model of naive embryonic cells and differentiated into many different cell types. Differentiation is not limited to embryonic development, but it continues in the adult as continuous supply of specialised cells is needed for tissue turn-over and repair. This is accomplished by lineage restricted multipotent cells, or adult stem cells (ASC). Correct stem cell function is essential during an individual’s life starting at the time when tissues are formed and later on, when they need to be regenerated and repaired. By analysing the epigenetic control of ESC and ASC, this section will describe how developmental plasticity of stem cells is programmed for correct function. Knowledge of how stem cell programs are regulated is important not only to advance stem cell-based therapies but also to understand how we can overcome diseases characteristic of stem cell dysfunction.
3.1 Control of Embryonic Stem Cells
ESC can be derived from the blastocyst ICM and their pluripotency maintained in vitro for many cell generations. ESC can symmetrically self-renew, hence giving rise to two identical stem cells. ESC ability to self-renew and to respond to developmental cues is controlled by a unique gene expression program. The ground state of pluripotent ESC is defined by the expression of a core network of transcription factors that include OCT4, SOX2 and NANOG. These factors act both as transcription activators and repressors, by activating genes involved in cell proliferation and self-renewal while repressing the expression of lineage-specific genes promoting differentiation (Young 2011). This bivalent state of ESC is essential for pluripotency and it is regulated epigenetically by the interplay of core transcription factors and Trithorax (TrxG) and Polycomb (PcG) epigenetic modifiers. TrxG-related proteins (SET/MLL) catalyse H3K4me3 at promoters of active genes, whereas PcG proteins catalyse histone modifications that are associated with gene silencing. PcG proteins include two complexes, PRC1 and PRC2, responsible for H3K27me3 and ubiquitylation of histone H2A at lysine 119 (H2AK119u), respectively (Meissner 2010). While PRC2 is required for initial gene silencing, recruitment of PRC1 stabilises the established transcriptionally repressive state. It is the presence of both active H3K4me3 and repressive H3K27me3 marks (bivalent domain) at developmentally regulated genes that allows ESC to remain in a poised state, ready for activation upon differentiation. Therefore ESC show a global open chromatin structure, with about 75% of gene promoters enriched for H3K4me3. These promoters can be active or inactive, depending on H3K27me3 co-occupancy.
Among silencing mechanisms DNA methylation plays a fundamental role in ESC. ESC present about 60–80% of methylated CpG nucleotides, with a unique distribution (Meissner 2010). Comprehensive maps of DNA methylation in ESC have demonstrated that the majority of high CpG promoters (HCP) are lacking methylation and are enriched in H3K4me3. These represent housekeeping genes, pluripotency genes and key developmental genes. In contrast, tissue specific gene promoters with low CpG density (LCP) are mostly methylated (Mikkelsen et al. 2007; Meissner et al. 2008). Therefore an epigenetic landscape presenting either unmethylated promoters (HCP with H3K4me3 or bivalent domain with H3K4me3/H3K27me3), or methylated promoters (LCP) define ESC (Fig. 24.1). In addition to methylation of CpG dinucleotides, other modifications of the DNA have been discovered in ESC. These include cytosine methylation in a non CG context (Lister et al. 2009) and cytosine hydromethylation (5hmC). Hydroxylation of 5mC to 5hmC is catalysed by the TET family of enzymes (Koh et al. 2011) and it is believed to be involved in the demethylation of 5mC and prevention of DNMTs activity (Xu et al. 2011). A genome-wide study of 5hmC in ESC revealed that this mark is enriched in gene bodies, transcription start sites of HCP promoters and enhancers. Bivalent or PcG only marked promoters are also particularly enriched for 5hmC (Xu et al. 2011; Pastor et al. 2011). Although a clear function for 5hmC in transcription regulation is still elusive, its distribution suggests a role in preparing genomic loci for activation upon differentiation. Indeed, 5hmC has been shown to be present in ESC, but declines after differentiation (Tahiliani et al. 2009; Ficz et al. 2011). Finally, ncRNAs also participate in the epigenetic regulation of ESC plasticity. MicroRNAs regulate stability and translation of mRNAs involved in stem cell self-renewal and differentiation. ESC express a unique set of miRNA whose transcription is regulated by the core pluripotency factors, and these miRNA are involved both in sustaining self-renewal (e.g mir-290-295/302) and inducing rapid degradation of ESC transcription factors during differentiation (e.g mir-145) (Marson et al. 2008; Tay et al. 2008; Xu et al. 2009). In addition, many lineage-specific miRNA gene promoters are co-occupied by OCT4/NANOG/SOX2 and PcG and these are repressed in ESC, but become active upon differentiation (e.g let-7, mir-155, mir-124, mir-9) (Young 2011; Pauli et al. 2011). Balance between self-renewal and differentiation is a key characteristic of ESC. Altogether bivalent histone modifications, DNA methylation and miRNAs contribute to the establishment of an open chromatin state that allows undifferentiated cell function and the ability to respond to developmental signals in a timely fashion.
Epigenetic landscapes regulating stem cell plasticity in development and cancer. HCP and LCP gene promoters are enriched for genes with different epigenetic regulation in different cell types. The figure shows how housekeeping genes, pluripotency genes, developmental genes and tissue-specific genes are epigenetically regulated in ESC, ASC, differentiated (somatic) cells and cancer cells. During differentiation there is a decrease in cell plasticity due to loss of bivalent domains and acquisition of repressive chromatin marks that restrict cell fate. Cancer cells reactivate an epigenetic landscape that is more plastic and shifted towards self-renewal and proliferation at the expense of differentiation (⌂: H3K4me3; ○: 5hmC; ∆: H3K27me3; ●: 5mC; ∆: H3K9me2/3)
3.2 Control of Adult Stem Cells and Somatic Cells
Differentiation of pluripotent cells is associated with a loss of developmental potency that ensures cellular specialisation and committed identity. During lineage specification, ASC are formed and it is from these committed multipotent stem cells that specialised cells are derived. The role of stem cells in the adult is to maintain tissue homeostasis by regenerating aged or damaged cells. ASC reside in tissue-specific niches that control their asymmetrical self-renewal, defined as the ability to form a stem cell and a differentiated progenitor at cell division. ASC are more restricted in their differentiation potential compared to ESC as they can only give rise to multiple cell types within a tissue under physiological conditions. Genome-wide maps of epigenetic modifications in ASC and differentiated cells show that restriction in developmental potential is associated with a resolution of ESC open chromatin into a more restricted configuration (Fig. 24.1). Silencing of pluripotency genes is readily observed during differentiation, by loss of H3K4me3, gain of H3K27me3, H3K9me3, DNA methylation (Barrero et al. 2010) and expression of specific miRNAs (e.g. mir-134, mir-296, mir-470) (Tay et al. 2008). Differentiation into a specific lineage involves expression of genes specific to that cell type and silencing of genes expressed in other tissues. In this way, differentiation into the neural stem/progenitor cells (NSC) is accompanied by a decrease of H3K27me3 at neural genes silenced by bivalent marks, which correspond to increased gene expression. Genes poised or weakly induced retain bivalent marks, while H3K27me3 silencing is increased in non-neural lineage genes, together with H3K9me3 (Hawkins et al. 2010; Bernstein et al. 2006; Mikkelsen et al. 2007; Bracken et al. 2006). The same pattern is also observed in muscle and germ cell differentiation (Caretti et al. 2004; Chen et al. 2005; Asp et al. 2011), suggesting a PcG-mediated regulation of cell fate decisions.
Both H3K4me3 and bivalent HPC remain mostly unmethylated during differentiation. In contrast, resolution to univalent H3K27me3 mark results in an increase in DNA methylation and a complete loss of the bivalent marks results in DNA hypermethylation. A different epigenetic regulation is observed at LCP associated with tissue specific genes, as methylated LCP associated with neural genes gain H3K4me3 and non lineage specific genes retain DNA methylation (Meissner et al. 2008). Although overall DNA methylation levels are similar in pluripotent and differentiated cells, a small subset of genes displays tissue specific methylation. A recent study demonstrated 491 differentially methylated regions (DMR) being more methylated in fibroblasts compared to ESC (Lister et al. 2009), with DMR representing only 6–8% of CpG islands in different tissues (Berdasco and Esteller 2010). Important DNA methylation differences can be observed in ASC compared to differentiated cells of the same lineage. For instance, breast self-renewal and proliferation genes are hypomethylated in CD44+/CD24− stem cells compared to differentiated luminal CD24+ cells (Bloushtain-Qimron et al. 2008).
Many studies indicate DNA methylation as a mechanism providing long term gene silencing and epigenetic memory in differentiated cells, however experiments of conditional deletion of DNMT1 suggest that DNA methylation plays also an important role in maintaining ASC self-renewal and suppressing differentiation (Sen et al. 2010). Indeed, loss of methylation causes differentiation alterations in epithelial progenitor cells (EPC) (Sen et al. 2010) and hematopoietic stem cells (HSC) (Trowbridge et al. 2009; Broske et al. 2009). Other epigenetic mechanisms are involved in regulation ASC self-renewal, but their relation to DNA methylation is still unknown. For instance, the PCR1 PcG protein BMI-1 is required for NSC, HSC, mammary and intestinal stem cell proliferation (Molofsky et al. 2003; Lessard and Sauvageau 2003; Pietersen et al. 2008; Sangiorgi and Capecchi 2008), while overexpression of PCR2 PcG protein EZH2 blocks differentiation of myoblasts and EPC (Caretti et al. 2004; Sen 2011) and prevents HSC exhaustion (Kamminga et al. 2006). Because cell memory is set during development and inherent to each tissue-type, it was long assumed that differentiation of ASC is strictly specific to their lineage. However, recent studies demonstrate that under certain conditions, particularly after injury, ASC can trans-differentiate into cells of different tissues (Lotem and Sachs 2006). Therefore ASC, like ESC, show a differentiation plasticity that is conferred by epigenetic programs that can reversibly regulate transcription of genes expressed in different tissue according to physiological and pathological signals. The contribution of ASC plasticity to cancer will be described in the next section.
4 Cancer Stem Cells
The idea that cancer is caused by transformed cells with stem cell properties is not novel, but it has received renewed interest among scientists in recent years. The observation that tumours are formed by cells with functional heterogeneity has led to the postulation of two mutually exclusive models for the cellular origin of cancer: the stochastic model and the cancer stem cell hierarchy. The stochastic (or clonal evolution) model predicts that every cell can become tumorigenic under the influence of endogenous (transcription factors) and exogenous (microenvironment) factors that can generate their own heterogeneous sub clones (Nowell 1976). In the stochastic model, cancer cells fluctuate between several states, owing to their plasticity (Wang and Dick 2005). In contrast, the cancer stem cell (CSC) model considers that tumours originate from transformed stem cells and they are organised in a hierarchical manner, whereby CSC lies at the apex and the proliferating progenitors and terminally differentiated cancer cells reside at the bottom of the hierarchy (Bonnet and Dick 1997). Recent studies show that both models can act together depending on microenvironmental signals and that tumour initiating cells can originate both from transformation of normal ASC or epigenetic reprogramming of more differentiated cells (Campbell and Polyak 2007) (Fig. 24.2). Both theories converge on the idea that cancer arises from transformed cells that acquire growth and survival advantage, which is a landmark of stem cells. The theory that cancer could arise from embryo-like cells was proposed about 150 years ago (Virchow 1855) and was later developed by Cohnheim and Durante with the concept of “maturation arrest”, according to which cancer could develop from embryonic rudiments remaining in adult organs (Cohnheim 1867; Durante 1974). These theories were proven years later by studies on germ cell tumours demonstrating that teratocarcinomas contain CSC with very similar characteristics to ESC, with self-renewal and differentiation potential (Sell and Pierce 1994; Sperger et al. 2003). For a long time it was known that only a small population of cancer cells is tumorigenic and can propagate the tumour. Single-cell analysis of leukaemia revealed two different populations of cancer cells in terms of proliferative kinetics: the frequent large, fast-cycling cells and the rare, smaller slow-cycling cells with the same properties to that of normal HSC (Clarkson 1974). Through elegant studies, Dick and colleagues proved the existence of CSC by showing that in acute myeloid leukaemia (AML), a rare population of CSC with CD34+/CD38− cell surface expression were able to recapitulate the original disease over repeated transplantation into NOD/SCID (non-obese diabetic/severe combined immunodeficiency) mice (Lapidot et al. 1994). Since then, CSC have been identified and isolated in solid tumours including breast (Al-Hajj et al. 2003), brain (Singh et al. 2003), melanoma (Fang et al. 2005), pancreatic (Hermann et al. 2007), prostate (Tang et al. 2007) and ovarian cancers (Bapat et al. 2005) (Table 24.2). CSC are a rare population of cells that resemble normal stem cells. They can self-renew, are long-lasting, remain relatively quiescent, and can generate all heterogeneous cell types comprising the tumour. CSC can lay dormant within their niche and therefore escape chemotherapy, which only targets highly proliferating cells. Their resistance to current cancer therapies (chemotherapy and radiotherapy) is also due to expression of ATP-binding cassette (ABC) transporters (pumping out harmful drugs), increased free radical scavenging and high expression of anti-apoptotic proteins (Visvader 2011). Since CSC retain many features of normal stem cells, their identification often relies on the expression of tissue specific stem cell markers. For example, leukemic stem cells can be identified by CD34 and CD38 which are expressed on the cells in the HSC hierarchy (Bonnet and Dick 1997). Other universal CSC markers are instead based on their ability to pump out toxicants, which defines them as “side population” cells able to extrude a Hoechst dye, expressing ABC transporters and the detoxifying enzyme ALDH1 (Visvader and Lindeman 2008). Functional assays for CSC identification include xenografts into immunocompromised mice and formation of spheroids in culture. The xenograft assay involves transplanting cancer cells into NOD/SCID mice for tumour formation. Isolated CSC are generally more tumorigenic than differentiated tumour cells and their serial transplantation shows that they can reproduce the original disease through every passage. Sphere forming assays, which involve culturing CSC under stem cell conditions, preserve survival of CSC while inducing cell death by apoptosis in non-CSC (Visvader and Lindeman 2008). However, regardless of the assay, a major challenge for studying CSC is their inherent developmental plasticity, which involves the co-existence of different epigenetic states during cancer progression. For instance, CD44+/CD24− breast CSC exist in a metastable state oscillating between differentiation and de-differentiation, with CSC giving rise to luminal CD24+ cells and luminal cells de-differentiating back into CSC (Meyer et al. 2009). The same has been observed in melanoma, where JARID1B+ CSC generate JARID1B− cells and vice versa (Roesch et al. 2010). In addition, CSC plasticity can often extend beyond their lineage and they can express genes normally expressed in different tissues. Consistent with the trans-differentiation potential of ASC after injury, CSC show the same plasticity resulting in abnormal tissue regeneration (Lotem and Sachs 2006). CSC plasticity is influenced by embryonic developmental programmes and a similar gene expression signature between highly malignant, poorly differentiated solid tumours and ESC has been reported (Ben-Porath et al. 2008). This is due to the ability of cancer to take control of normal developmental programs for selective advantage, albeit in part related to an upregulated Myc-regulatory network (Kim et al. 2010). For example, the epithelial-to-mesenchymal (EMT) transition, a reversible embryonic programme that allows transition between cellular phenotypes during gastrulation, contributes to CSC plasticity. EMT is recapitulated during tumour progression and metastasis by a transition from an epithelial to a mesenchymal phenotype with acquired cell motility. This is induced by activation of key signalling pathways (TGF-β, Notch, FGF) that drive epigenetic silencing of the adhesion molecule E-cadherin (Thiery et al. 2009). EMT is also important for maintenance of stem cell properties and CSC can hijack this program to regulate their plasticity. In addition to this, CSC establish their own niche by recruiting cells to recreate a similar microenvironment to that of a normal stem cell niche. The niche can induce and expand CSC by enhancing “stemness” features in non tumourigenic cells by overexpressing signals that are important for stem cell renewal and promote EMT through epigenetic alterations (Mani et al. 2008).
Cancer stem cell-of-origin model. Different cell types from the lineage hierarchy can undergo oncogenic events to transform into tumour cells. CSC can either originate from a transformed normal stem cell, transit-amplifying cell, progenitor cell, or a terminally differentiated cell to give rise to a heterogeneous population of cancer cells
4.1 Epigenetic Origin of Cancer Stem Cells
Epigenetic alterations are generally observed at early stages of tumorigenesis and are likely candidates for a mechanism of tumour initiation. ASC are long-lived and during their aging process they may undergo epigenetic insult which can induce survival programs and predispose to the onset of cancer after further genetic and epigenetic alterations (Feinberg et al. 2006; Baylin and Ohm 2006). Numerous evidences indicate a role for epigenetic defects in the development of CSC. Normal stem cells are vulnerable to epigenetic alteration when induced to sustained self-renewal. Extensive DNA methylation alterations have been reported in ESC after long term in culture, with changes which are inherited after differentiation and associated with cancer (Allegrucci et al. 2007). Similar alterations have been observed in NSC (Shen et al. 2006), with a recent study demonstrating hypermethylation of HPC after many generations and inherited after differentiation to astrocytes (Meissner et al. 2008). Hypermethylation of bivalent domain genes in stem cells is particularly important for tumorigenesis as tumour suppressor genes have bivalent promoters in ESC and ASC (Barrero et al. 2010) and hypermethylation of tumour suppressor genes is a hallmark of cancer (Jones and Baylin 2007). As bivalent genes are developmental genes and transcription factors that regulate stem cell fate, it seems apparent how their epigenetic silencing in ASC could generate stem cells locked in a self-renewal state with impaired or limited differentiation potential (Fig. 24.1). Several studies have demonstrated that PcG target genes are much more likely to become hypermethylated in cancer (Widschwendter et al. 2007; Ohm et al. 2007; Schlesinger et al. 2007) and a mechanism by which the PcG-H3K27me3 mark could direct DNA methylation has been proposed (Keshet et al. 2006; Vire et al. 2006). In addition, overexpression of PcG proteins BMI-1 and EZH2 are also often found in cancer and they both play a fundamental role in regulating stem cell function (Bracken and Helin 2009). DNA methylation alterations at other genomic regions can also participate in the development of CSC. Hypomethylation of the genome can induce chromosome instability together with aberrant activation of proto-oncogenes associated with stem cell self-renewal and proliferation (Sharma et al. 2010). Chromosome translocations producing MLL fusion proteins are involved in AML, with more than 50 different fusion partners being identified. Importantly, MLL-ENF fusion is able to transform HSC and committed progenitors, thus creating CSC with acquired self-renewal and de-differentiated phenotype (Milne et al. 2005).
Loss of imprinting (LOI) via DNA demethylation can also be associated with growth advantage in stem cells and biallelic expression of IGF2 accounts for half of Wilms tumours and predisposition to colon cancer. Other LOI involved in cancer include PEG1/MEST involved in lung cancer, CDK1C in pancreatic cancer, TP73 in gastric cancer and DIRAS3 in breast cancer (Feinberg et al. 2006). Finally, DNA methylation at promoters of miRNA genes involved in stem cell differentiation can lead to CSC. For instance, silencing of the miRNA let-7 contributes to breast, colon and lung cancer (Zimmerman and Wu 2011) and silencing of the mir-200 gene family induces EMT and CSC phenotype in breast, lung and ovarian cancer (Brabletz and Brabletz 2010). As epigenetic alterations can result in CSC and plasticity is a fundamental property of stem cells, we should not be surprised that CSC share this characteristic. What becomes apparent is that the effect of the environment is of primary importance for cancer initiation and progression while the behaviour of stem cells is completely dependent on physiological and pathological signals. As controlling environmental conditions is not an easy endeavour, we need to develop treatments that are able to completely eradicate CSC as their plasticity is a likely prospect of tumour recurrence.
5 Resetting Cancer by Epigenetic Re-programming
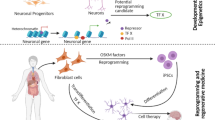
The developmental plasticity demonstrated by CSC and the reversible nature of epigenetic alterations has led to the development of epigenetic therapies as a new treatment option for cancer patients. Several epigenetic drugs that aim at halting tumour progression and restoring normal cell function are being tested in human clinical trials. Because epigenetic drugs can induce differentiation of ESC and ASC, their use for resumption of normal tissue differentiation is also been tested (Berdasco and Esteller 2010). The concept of differentiation therapy as an alternative or adjuvant treatment to chemotherapy has been inspired by many years of research. Landmark experiments have shown that embryonic microenvironments that program cell fate during development are able to reverse malignancy by resetting normal pathways of cell differentiation. For instance, teratocarcinoma cells can be induced to differentiate when transplanted into chimeric embryos, and malignancy can be reverted by injecting cancer cells into zebrafish, chicken and mouse embryos (Telerman and Amson 2009). Nuclear transfer experiments have shown that the epigenotype of cancer cells can be reprogrammed by oocyte molecules (Blelloch et al. 2004; Hochedlinger et al. 2004) and recent experiments with oocyte extracts have shown that this effect is mediated by chromatin remodelling and reactivation of silenced tumour suppressor genes resulting in reduction of tumour growth in mouse xenografts (Allegrucci et al. 2011). New insights have also come from studies of reprogramming to pluripotency. Yamanaka’s work demonstrated that the ectopic expression of core pluripotency factors (OCT4, SOX2, MYC, KLF4) in somatic cells can reprogram the epigenetic state of somatic cells to that of pluripotent cells giving rise to induced pluripotent stem cells (iPSC) (Takahashi and Yamanaka 2006). With a similar approach, cancer cells are reprogrammed to induced pluripotent cancer cells (iPC) with re-acquired differentiation potential (Sun and Liu 2011). Reprogramming studies suggest that even if finely regulated, epigenetic landscapes are plastic and reversible and that cellular transformation and de-differentiation share similar epigenetic mechanisms. This view is sustained by the evidence that many factors able to reprogram somatic/cancer cells to induced pluripotent cells act as oncogenes. This is the case for OCT4 and SOX2 (aberrant expression is tumorigenic in epithelial tissues), NANOG (overexpressed in germ cell tumours), KLF4 (associated with colorectal cancer), LIN28 (associated with hepatic cancer) and MYC (potent oncogene involved in many cancer types) (Daley 2008). Therefore, differentiation therapy could be the way to reset CSC to normal function and epigenetic programs that regulate stem cells are of particular interest as novel treatment targets.
6 Conclusions
More than 50 years have passed since Waddington proposed the idea that signals from embryonic environments can influence cell fate and behaviour according to an “epigenetic landscape” (Waddington 1940). He proposed that embryonic development can be visualised as a landscape delimited by hills and valley, where cells can take different directions depending on the signals received along the path. However, possible paths in this landscape are restricted by barriers dictated by hills and only defined valleys are available when cells are restricted to a defined initial trajectory. With a modern view, we identify these landscapes as epigenetic modifications regulating expression of developmental genes during differentiation in a coordinated fashion. Although cell fate is developmentally established, a degree of plasticity is retained for tissue turnover or acquired in pathological conditions. Waddington recognised that cancer cells escape the effect of developmental forces (Waddington 1935), introducing the concept of cancer as a defect in the mechanisms that control cell differentiation. We now know that epigenetic alterations are hallmark of cancer and they can transform normal cells to tumour cells with altered proliferation and differentiation. Our knowledge of cell plasticity has greatly increased over the last few years as stem cell technologies have developed and genome-wide mapping of epigenetic modifications of stem cells and somatic cells are being established. Defining the epigenome of cellular states in normal and cancer tissues is therefore a new challenge, but a most beneficial one as it holds the promise to eradicate or control cancer, an expected disease of an aging population.
References
Al-Hajj M, Wicha MS, Benito-Hernandez A, Morrison SJ, Clarke MF (2003) Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci USA 100:3983–3988
Allegrucci C, Thurston A, Lucas E, Young L (2005) Epigenetics and the germline. Reproduction 129:137–149
Allegrucci C, Wu YZ, Thurston A, Denning CN, Priddle H, Mummery CL, Ward-van Oostwaard D, Andrews PW, Stojkovic M, Smith N, Parkin T, Jones ME, Warren G, Yu L, Brena RM, Plass C, Young LE (2007) Restriction landmark genome scanning identifies culture-induced DNA methylation instability in the human embryonic stem cell epigenome. Hum Mol Genet 16:1253–1268
Allegrucci C, Rushton MD, Dixon JE, Sottile V, Shah M, Kumari R, Watson S, Alberio R, Johnson AD (2011) Epigenetic reprogramming of breast cancer cells with oocyte extracts. Mol Cancer 10:7
Asp P, Blum R, Vethantham V, Parisi F, Micsinai M, Cheng J, Bowman C, Kluger Y, Dynlacht BD (2011) PNAS Plus: genome-wide remodeling of the epigenetic landscape during myogenic differentiation. Proc Natl Acad Sci USA 108:E149–E158
Ball MP, Li JB, Gao Y, Lee J-H, LeProust EM, Park I-H, Xie B, Daley GQ, Church GM (2009) Targeted and genome-scale strategies reveal gene-body methylation signatures in human cells. Nat Biotechnol 27:361–368
Bapat SA, Mali AM, Koppikar CB, Kurrey NK (2005) Stem and progenitor-like cells contribute to the aggressive behavior of human epithelial ovarian cancer. Cancer Res 65:3025–3029
Barrero MJ, Boue S, Izpisua Belmonte JC (2010) Epigenetic mechanisms that regulate cell identity. Cell Stem Cell 7:565–570
Baylin SB, Ohm JE (2006) Epigenetic gene silencing in cancer – a mechanism for early oncogenic pathway addiction? Nat Rev Cancer 6:107–116
Ben-Porath I, Thomson MW, Carey VJ, Ge R, Bell GW, Regev A, Weinberg RA (2008) An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nat Genet 40:499–507
Berdasco M, Esteller M (2010) Aberrant epigenetic landscape in cancer: how cellular identity goes awry. Dev Cell 19:698–711
Bernstein BE, Mikkelsen TS, Xie X, Kamal M, Huebert DJ, Cuff J, Fry B, Meissner A, Wernig M, Plath K, Jaenisch R, Wagschal A, Feil R, Schreiber SL, Lander ES (2006) A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125:315–326
Bird A (2002) DNA methylation patterns and epigenetic memory. Genes Dev 16:6–21
Bird A (2007) Perceptions of epigenetics. Nature 447:396–398
Blelloch RH, Hochedlinger K, Yamada Y, Brennan C, Kim M, Mintz B, Chin L, Jaenisch R (2004) Nuclear cloning of embryonal carcinoma cells. Proc Natl Acad Sci USA 101:13985–13990
Bloushtain-Qimron N, Yao J, Snyder EL, Shipitsin M, Campbell LL, Mani SA, Hu M, Chen H, Ustyansky V, Antosiewicz JE, Argani P, Halushka MK, Thomson JA, Pharoah P, Porgador A, Sukumar S, Parsons R, Richardson AL, Stampfer MR, Gelman RS, Nikolskaya T, Nikolsky Y, Polyak K (2008) Cell type-specific DNA methylation patterns in the human breast. Proc Natl Acad Sci USA 105:14076–14081
Bonnet D, Dick JE (1997) Human acute myeloid leukemia is organized as a hierarchy that originates from a primitive hematopoietic cell. Nat Med 3:730–737
Brabletz S, Brabletz T (2010) The ZEB/miR-200 feedback loop–a motor of cellular plasticity in development and cancer? EMBO Rep 11:670–677
Bracken AP, Helin K (2009) Polycomb group proteins: navigators of lineage pathways led astray in cancer. Nat Rev Cancer 9:773–784
Bracken AP, Dietrich N, Pasini D, Hansen KH, Helin K (2006) Genome-wide mapping of Polycomb target genes unravels their roles in cell fate transitions. Genes Dev 20:1123–1136
Broske AM, Vockentanz L, Kharazi S, Huska MR, Mancini E, Scheller M, Kuhl C, Enns A, Prinz M, Jaenisch R, Nerlov C, Leutz A, Andrade-Navarro MA, Jacobsen SE, Rosenbauer F (2009) DNA methylation protects hematopoietic stem cell multipotency from myeloerythroid restriction. Nat Genet 41:1207–1215
Campbell LL, Polyak K (2007) Breast tumor heterogeneity: cancer stem cells or clonal evolution? Cell Cycle 6:2332–2338
Caretti G, Di Padova M, Micales B, Lyons GE, Sartorelli V (2004) The Polycomb Ezh2 methyltransferase regulates muscle gene expression and skeletal muscle differentiation. Genes Dev 18:2627–2638
Chen X, Hiller M, Sancak Y, Fuller MT (2005) Tissue-specific TAFs counteract Polycomb to turn on terminal differentiation. Science 310:869–872
Clarkson BD (1974) The survival value of the dormant state in neoplastic and normal cell populations. In: Clarkson B, Baserga R (eds) Control of proliferation in animal cells, vol 1, Cold spring harbor conferences on cell proliferation. Cold Spring Harbor Laboratory, New York, pp 945–972
Cohnheim J (1867) Ueber entzundung und eiterung. Path Anat Physiol Klin Med 40:1–79
Cox CV, Evely RS, Oakhill A, Pamphilon DH, Goulden NJ, Blair A (2004) Characterization of acute lymphoblastic leukemia progenitor cells. Blood 104:2919–2925
Dalerba P, Dylla SJ, Park IK, Liu R, Wang X, Cho RW, Hoey T, Gurney A, Huang EH, Simeone DM, Shelton AA, Parmiani G, Castelli C, Clarke MF (2007) Phenotypic characterization of human colorectal cancer stem cells. Proc Natl Acad Sci USA 104:10158–10163
Daley GQ (2008) Common themes of dedifferentiation in somatic cell reprogramming and cancer. Cold Spring Harb Symp Quant Biol 73:171–174
Durante F (1974) Nesso fisio-patologico tra la struttura dei nei materni e la genesi di alcuni tumori maligni. Arch Memori ed Osservazioni di Chirurgia Practica 11:217–2226
Eramo A, Lotti F, Sette G, Pilozzi E, Biffoni M, Di Virgilio A, Conticello C, Ruco L, Peschle C, De Maria R (2008) Identification and expansion of the tumorigenic lung cancer stem cell population. Cell Death Differ 15:504–514
Fang D, Nguyen TK, Leishear K, Finko R, Kulp AN, Hotz S, Van Belle PA, Xu X, Elder DE, Herlyn M (2005) A tumorigenic subpopulation with stem cell properties in melanomas. Cancer Res 65:9328–9337
Feinberg AP, Ohlsson R, Henikoff S (2006) The epigenetic progenitor origin of human cancer. Nat Rev Genet 7(1):21–33
Ficz G, Branco MR, Seisenberger S, Santos F, Krueger F, Hore TA, Marques CJ, Andrews S, Reik W (2011) Dynamic regulation of 5-hydroxymethylcytosine in mouse ES cells and during differentiation. Nature 473:398–402
Garraway LA, Sellers WR (2006) Lineage dependency and lineage-survival oncogenes in human cancer. Nat Rev Cancer 6:593–602
Ginestier C, Hur MH, Charafe-Jauffret E, Monville F, Dutcher J, Brown M, Jacquemier J, Viens P, Kleer CG, Liu S, Schott A, Hayes D, Birnbaum D, Wicha MS, Dontu G (2007) ALDH1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell 1:555–567
Goldstein AS, Stoyanova T, Witte ON (2010) Primitive origins of prostate cancer: in vivo evidence for prostate-regenerating cells and prostate cancer-initiating cells. Mol Oncol 4:385–396
Hawkins RD, Hon GC, Lee LK, Ngo Q, Lister R, Pelizzola M, Edsall LE, Kuan S, Luu Y, Klugman S, Antosiewicz-Bourget J, Ye Z, Espinoza C, Agarwahl S, Shen L, Ruotti V, Wang W, Stewart R, Thomson JA, Ecker JR, Ren B (2010) Distinct epigenomic landscapes of pluripotent and lineage-committed human cells. Cell Stem Cell 6:479–491
Hemberger M, Dean W, Reik W (2009) Epigenetic dynamics of stem cells and cell lineage commitment: digging Waddington’s canal. Nat Rev Mol Cell Biol 10:526–537
Hermann PC, Huber SL, Herrler T, Aicher A, Ellwart JW, Guba M, Bruns CJ, Heeschen C (2007) Distinct populations of cancer stem cells determine tumor growth and metastatic activity in human pancreatic cancer. Cell Stem Cell 1:313–323
Hochedlinger K, Blelloch R, Brennan C, Yamada Y, Kim M, Chin L, Jaenisch R (2004) Reprogramming of a melanoma genome by nuclear transplantation. Genes Dev 18:1875–1885
Howell CY, Bestor TH, Ding F, Latham KE, Mertineit C, Trasler JM, Chaillet JR (2001) Genomic imprinting disrupted by a maternal effect mutation in the Dnmt1 gene. Cell 104:829–838
Jaenisch R, Bird A (2003) Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat Genet 33(Suppl):245–254
Jiang F, Qiu Q, Khanna A, Todd NW, Deepak J, Xing L, Wang H, Liu Z, Su Y, Stass SA, Katz RL (2009) Aldehyde dehydrogenase 1 is a tumor stem cell-associated marker in lung cancer. Mol Cancer Res 7:330–338
Jones PA, Baylin SB (2007) The epigenomics of cancer. Cell 128:683–692
Kamminga LM, Bystrykh LV, de Boer A, Houwer S, Douma J, Weersing E, Dontje B, de Haan G (2006) The Polycomb group gene Ezh2 prevents hematopoietic stem cell exhaustion. Blood 107:2170–2179
Keshet I, Schlesinger Y, Farkash S, Rand E, Hecht M, Segal E, Pikarski E, Young RA, Niveleau A, Cedar H, Simon I (2006) Evidence for an instructive mechanism of de novo methylation in cancer cells. Nat Genet 38:149–153
Kim J, Woo AJ, Chu J, Snow JW, Fujiwara Y, Kim CG, Cantor AB, Orkin SH (2010) A Myc network accounts for similarities between embryonic stem and cancer cell transcription programs. Cell 143:313–324
Koh KP, Yabuuchi A, Rao S, Huang Y, Cunniff K, Nardone J, Laiho A, Tahiliani M, Sommer CA, Mostoslavsky G, Lahesmaa R, Orkin SH, Rodig SJ, Daley GQ, Rao A (2011) Tet1 and Tet2 regulate 5-hydroxymethylcytosine production and cell lineage specification in mouse embryonic stem cells. Cell Stem Cell 8:200–213
Lapidot T, Sirard C, Vormoor J, Murdoch B, Hoang T, Caceres-Cortes J, Minden M, Paterson B, Caligiuri MA, Dick JE (1994) A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature 367:645–648
Lessard J, Sauvageau G (2003) Bmi-1 determines the proliferative capacity of normal and leukaemic stem cells. Nature 423:255–260
Li C, Heidt DG, Dalerba P, Burant CF, Zhang L, Adsay V, Wicha M, Clarke MF, Simeone DM (2007) Identification of pancreatic cancer stem cells. Cancer Res 67:1030–1037
Lister R, Pelizzola M, Dowen RH, Hawkins RD, Hon G, Tonti-Filippini J, Nery JR, Lee L, Ye Z, Ngo QM, Edsall L, Antosiewicz-Bourget J, Stewart R, Ruotti V, Millar AH, Thomson JA, Ren B, Ecker JR (2009) Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462:315–322
Lotem J, Sachs L (2006) Epigenetics and the plasticity of differentiation in normal and cancer stem cells. Oncogene 25:7663–7672
Mani SA, Guo W, Liao MJ, Eaton EN, Ayyanan A, Zhou AY, Brooks M, Reinhard F, Zhang CC, Shipitsin M, Campbell LL, Polyak K, Brisken C, Yang J, Weinberg RA (2008) The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133:704–715
Marson A, Levine SS, Cole MF, Frampton GM, Brambrink T, Johnstone S, Guenther MG, Johnston WK, Wernig M, Newman J, Calabrese JM, Dennis LM, Volkert TL, Gupta S, Love J, Hannett N, Sharp PA, Bartel DP, Jaenisch R, Young RA (2008) Connecting microRNA genes to the core transcriptional regulatory circuitry of embryonic stem cells. Cell 134:521–533
Mayer W, Niveleau A, Walter J, Fundele R, Haaf T (2000) Demethylation of the zygotic paternal genome. Nature 403:501–502
Meissner A (2010) Epigenetic modifications in pluripotent and differentiated cells. Nat Biotechnol 28:1079–1088
Meissner A, Mikkelsen TS, Gu H, Wernig M, Hanna J, Sivachenko A, Zhang X, Bernstein BE, Nusbaum C, Jaffe DB, Gnirke A, Jaenisch R, Lander ES (2008) Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature 454:766–770
Meyer MJ, Fleming JM, Ali MA, Pesesky MW, Ginsburg E, Vonderhaar BK (2009) Dynamic regulation of CD24 and the invasive, CD44posCD24neg phenotype in breast cancer cell lines. Breast Cancer Res 11:R82
Mikkelsen TS, Ku M, Jaffe DB, Issac B, Lieberman E, Giannoukos G, Alvarez P, Brockman W, Kim TK, Koche RP, Lee W, Mendenhall E, O’Donovan A, Presser A, Russ C, Xie X, Meissner A, Wernig M, Jaenisch R, Nusbaum C, Lander ES, Bernstein BE (2007) Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448:553–560
Milne TA, Martin ME, Brock HW, Slany RK, Hess JL (2005) Leukemogenic MLL fusion proteins bind across a broad region of the Hox a9 locus, promoting transcription and multiple histone modifications. Cancer Res 65:11367–11374
Molofsky AV, Pardal R, Iwashita T, Park IK, Clarke MF, Morrison SJ (2003) Bmi-1 dependence distinguishes neural stem cell self-renewal from progenitor proliferation. Nature 425:962–967
Morgan HD, Santos F, Green K, Dean W, Reik W (2005) Epigenetic reprogramming in mammals. Hum Mol Genet 14(Spec No 1):R47–R58
Myant K, Termanis A, Sundaram AY, Boe T, Li C, Merusi C, Burrage J, de Las Heras JI, Stancheva I (2011) LSH and G9a/GLP complex are required for developmentally programmed DNA methylation. Genome Res 21:83–94
Nowell PC (1976) The clonal evolution of tumor cell populations. Science 194:23–28
O’Brien CA, Pollett A, Gallinger S, Dick JE (2007) A human colon cancer cell capable of initiating tumour growth in immunodeficient mice. Nature 445:106–110
Ohm JE, McGarvey KM, Yu X, Cheng L, Schuebel KE, Cope L, Mohammad HP, Chen W, Daniel VC, Yu W, Berman DM, Jenuwein T, Pruitt K, Sharkis SJ, Watkins DN, Herman JG, Baylin SB (2007) A stem cell-like chromatin pattern may predispose tumor suppressor genes to DNA hypermethylation and heritable silencing. Nat Genet 39:237–242
Okano M, Bell DW, Haber DA, Li E (1999) DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99:247–257
Oswald J, Engemann S, Lane N, Mayer W, Olek A, Fundele R, Dean W, Reik W, Walter J (2000) Active demethylation of the paternal genome in the mouse zygote. Curr Biol 10:475–478
Pastor WA, Pape UJ, Huang Y, Henderson HR, Lister R, Ko M, McLoughlin EM, Brudno Y, Mahapatra S, Kapranov P, Tahiliani M, Daley GQ, Liu XS, Ecker JR, Milos PM, Agarwal S, Rao A (2011) Genome-wide mapping of 5-hydroxymethylcytosine in embryonic stem cells. Nature 473:394–397
Pauli A, Rinn JL, Schier AF (2011) Non-coding RNAs as regulators of embryogenesis. Nat Rev Genet 12:136–149
Pierce GB, Johnson LD (1971) Differentiation and cancer. In Vitro 7:140–145
Pietersen AM, Evers B, Prasad AA, Tanger E, Cornelissen-Steijger P, Jonkers J, van Lohuizen M (2008) Bmi1 regulates stem cells and proliferation and differentiation of committed cells in mammary epithelium. Curr Biol 18:1094–1099
Prince ME, Sivanandan R, Kaczorowski A, Wolf GT, Kaplan MJ, Dalerba P, Weissman IL, Clarke MF, Ailles LE (2007) Identification of a subpopulation of cells with cancer stem cell properties in head and neck squamous cell carcinoma. Proc Natl Acad Sci USA 104:973–978
Rajasekhar VK, Studer L, Gerald W, Socci ND, Scher HI (2011) Tumour-initiating stem-like cells in human prostate cancer exhibit increased NF-kappaB signalling. Nat Commun 2:162
Ran D, Schubert M, Pietsch L, Taubert I, Wuchter P, Eckstein V, Bruckner T, Zoeller M, Ho AD (2009) Aldehyde dehydrogenase activity among primary leukemia cells is associated with stem cell features and correlates with adverse clinical outcomes. Exp Hematol 37:1423–1434
Roesch A, Fukunaga-Kalabis M, Schmidt EC, Zabierowski SE, Brafford PA, Vultur A, Basu D, Gimotty P, Vogt T, Herlyn M (2010) A temporarily distinct subpopulation of slow-cycling melanoma cells is required for continuous tumor growth. Cell 141:583–594
Sangiorgi E, Capecchi MR (2008) Bmi1 is expressed in vivo in intestinal stem cells. Nat Genet 40:915–920
Santos F, Hendrich B, Reik W, Dean W (2002) Dynamic reprogramming of DNA methylation in the early mouse embryo. Dev Biol 241:172–182
Sasaki H, Matsui Y (2008) Epigenetic events in mammalian germ-cell development: reprogramming and beyond. Nat Rev Genet 9:129–140
Schatton T, Murphy GF, Frank NY, Yamaura K, Waaga-Gasser AM, Gasser M, Zhan Q, Jordan S, Duncan LM, Weishaupt C, Fuhlbrigge RC, Kupper TS, Sayegh MH, Frank MH (2008) Identification of cells initiating human melanomas. Nature 451:345–349
Schlesinger Y, Straussman R, Keshet I, Farkash S, Hecht M, Zimmerman J, Eden E, Yakhini Z, Ben-Shushan E, Reubinoff BE, Bergman Y, Simon I, Cedar H (2007) Polycomb-mediated methylation on Lys27 of histone H3 pre-marks genes for de novo methylation in cancer. Nat Genet 39:232–236
Sell S, Pierce GB (1994) Maturation arrest of stem cell differentiation is a common pathway for the cellular origin of teratocarcinomas and epithelial cancers. Lab Invest 70:6–22
Sen GL (2011) Remembering one’s identity: the epigenetic basis of stem cell fate decisions. FASEB J 25(7):2123–2128
Sen GL, Reuter JA, Webster DE, Zhu L, Khavari PA (2010) DNMT1 maintains progenitor function in self-renewing somatic tissue. Nature 463:563–567
Sharma S, Kelly TK, Jones PA (2010) Epigenetics in cancer. Carcinogenesis 31:27–36
Shen Y, Chow J, Wang Z, Fan G (2006) Abnormal CpG island methylation occurs during in vitro differentiation of human embryonic stem cells. Hum Mol Genet 15:2623–2635
Singh SK, Clarke ID, Terasaki M, Bonn VE, Hawkins C, Squire J, Dirks PB (2003) Identification of a cancer stem cell in human brain tumors. Cancer Res 63:5821–5828
Sperger JM, Chen X, Draper JS, Antosiewicz JE, Chon CH, Jones SB, Brooks JD, Andrews PW, Brown PO, Thomson JA (2003) Gene expression patterns in human embryonic stem cells and human pluripotent germ cell tumors. Proc Natl Acad Sci USA 100:13350–13355
Sun C, Liu YK (2011) Induced pluripotent cancer cells: progress and application. J Cancer Res Clin Oncol 137:1–8
Tahiliani M, Koh KP, Shen Y, Pastor WA, Bandukwala H, Brudno Y, Agarwal S, Iyer LM, Liu DR, Aravind L, Rao A (2009) Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324:930–935
Takahashi K, Yamanaka S (2006) Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126:663–676
Takaishi S, Okumura T, Tu S, Wang SS, Shibata W, Vigneshwaran R, Gordon SA, Shimada Y, Wang TC (2009) Identification of gastric cancer stem cells using the cell surface marker CD44. Stem Cells 27:1006–1020
Tang DG, Patrawala L, Calhoun T, Bhatia B, Choy G, Schneider-Broussard R, Jeter C (2007) Prostate cancer stem/progenitor cells: identification, characterization, and implications. Mol Carcinog 46:1–14
Tay Y, Zhang J, Thomson AM, Lim B, Rigoutsos I (2008) MicroRNAs to Nanog, Oct4 and Sox2 coding regions modulate embryonic stem cell differentiation. Nature 455:1124–1128
Telerman A, Amson R (2009) The molecular programme of tumour reversion: the steps beyond malignant transformation. Nat Rev Cancer 9:206–216
Thiery JP, Acloque H, Huang RY, Nieto MA (2009) Epithelial-mesenchymal transitions in development and disease. Cell 139(5):871–890
Tirino V, Desiderio V, d’Aquino R, De Francesco F, Pirozzi G, Graziano A, Galderisi U, Cavaliere C, De Rosa A, Papaccio G, Giordano A (2008) Detection and characterization of CD133+ cancer stem cells in human solid tumours. PLoS One 3:e3469
Trowbridge JJ, Snow JW, Kim J, Orkin SH (2009) DNA methyltransferase 1 is essential for and uniquely regulates hematopoietic stem and progenitor cells. Cell Stem Cell 5:442–449
Turner BM (2007) Defining an epigenetic code. Nat Cell Biol 9:2–6
Virchow R (1855) Editorial Archiv fuer pathologische Anatomie und Physiologie und fuer klinisque Madizin 8:23–54
Vire E, Brenner C, Deplus R, Blanchon L, Fraga M, Didelot C, Morey L, Van Eynde A, Bernard D, Vanderwinden JM, Bollen M, Esteller M, Di Croce L, de Launoit Y, Fuks F (2006) The Polycomb group protein EZH2 directly controls DNA methylation. Nature 439:871–874
Visvader JE (2011) Cells of origin in cancer. Nature 469:314–322
Visvader JE, Lindeman GJ (2008) Cancer stem cells in solid tumours: accumulating evidence and unresolved questions. Nat Rev Cancer 8:755–768
Waddington CH (1935) Cancer and the theory of organisers. Nature 135:606–608
Waddington CH (1940) Organisers & genes. Cambridge University Press, Cambridge, UK
Wang JCY, Dick JE (2005) Cancer stem cells: lessons from leukemia. Trends Cell Biol 15:494–501
Widschwendter M, Fiegl H, Egle D, Mueller-Holzner E, Spizzo G, Marth C, Weisenberger DJ, Campan M, Young J, Jacobs I, Laird PW (2007) Epigenetic stem cell signature in cancer. Nat Genet 39:157–158
Wossidlo M, Arand J, Sebastiano V, Lepikhov K, Boiani M, Reinhardt R, Scholer H, Walter J (2010) Dynamic link of DNA demethylation, DNA strand breaks and repair in mouse zygotes. EMBO J 29:1877–1888
Wossidlo M, Nakamura T, Lepikhov K, Marques CJ, Zakhartchenko V, Boiani M, Arand J, Nakano T, Reik W, Walter J (2011) 5-Hydroxymethylcytosine in the mammalian zygote is linked with epigenetic reprogramming. Nat Commun 2:241
Wray J, Kalkan T, Smith AG (2010) The ground state of pluripotency. Biochem Soc Trans 38:1027–1032
Xu N, Papagiannakopoulos T, Pan G, Thomson JA, Kosik KS (2009) MicroRNA-145 regulates OCT4, SOX2, and KLF4 and represses pluripotency in human embryonic stem cells. Cell 137:647–658
Xu Y, Wu F, Tan L, Kong L, Xiong L, Deng J, Barbera AJ, Zheng L, Zhang H, Huang S, Min J, Nicholson T, Chen T, Xu G, Shi Y, Zhang K, Shi YG (2011) Genome-wide regulation of 5hmC, 5mC, and gene expression by Tet1 hydroxylase in mouse embryonic stem cells. Mol Cell 42:451–464
Yang ZF, Ho DW, Ng MN, Lau CK, Yu WC, Ngai P, Chu PW, Lam CT, Poon RT, Fan ST (2008) Significance of CD90+ cancer stem cells in human liver cancer. Cancer Cell 13:153–166
Young RA (2011) Control of the embryonic stem cell state. Cell 144:940–954
Zhang S, Balch C, Chan MW, Lai HC, Matei D, Schilder JM, Yan PS, Huang TH, Nephew KP (2008) Identification and characterization of ovarian cancer-initiating cells from primary human tumors. Cancer Res 68:4311–4320
Zimmerman AL, Wu S (2011) MicroRNAs, cancer and cancer stem cells. Cancer Lett 300:10–19
Acknowledgements
The authors would like to acknowledge funding from the University of Nottingham, the Royal Society of London and EvoCell Ltd. We are obliged to Dr Ramiro Alberio for valuable discussion and critical evaluation of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media Dordrecht
About this chapter
Cite this chapter
Shah, M., Allegrucci, C. (2013). Stem Cell Plasticity in Development and Cancer: Epigenetic Origin of Cancer Stem Cells. In: Kundu, T. (eds) Epigenetics: Development and Disease. Subcellular Biochemistry, vol 61. Springer, Dordrecht. https://doi.org/10.1007/978-94-007-4525-4_24
Download citation
DOI: https://doi.org/10.1007/978-94-007-4525-4_24
Published:
Publisher Name: Springer, Dordrecht
Print ISBN: 978-94-007-4524-7
Online ISBN: 978-94-007-4525-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)