Abstract
Purpose
STK3 has a central role in maintaining cell homeostasis, proliferation, growth, and apoptosis. Previously, we investigated the functional link between STK3/MST2, and estrogen receptor in MCF-7 breast cancer cells. To expand the investigation, this study evaluated STK3’s higher expression and associated genes in breast cancer intrinsic subtypes using publicly available data.
Methods
The relationship between clinical pathologic features and STK3 high expression was analyzed using descriptive and multivariate analysis.
Results
Increased STK3 expression in breast cancer was significantly associated with higher pathological cancer stages, and a different expression level was observed in the intrinsic subtypes of breast cancer. Kaplan–Meier analysis showed that breast cancer with high STK3 had a lower survival rate in IDC patients than that with low STK3 expression (p < 0.05). The multivariate analysis unveiled a strong correlation between STK3 expression and the survival rate among IDC patients, demonstrating hazard ratios for lower expression. In the TCGA dataset, the hazard ratio was 0.56 (95% CI 0.34–0.94, p = 0.029) for patients deceased with tumor, and 0.62 (95% CI 0.42–0.92, p = 0.017) for all deceased patients. Additionally, in the METABRIC dataset, the hazard ratio was 0.76 (95% CI 0.64–0.91, p = 0.003) for those deceased with tumor. From GSEA outcomes 7 gene sets were selected based on statistical significance (FDR < 0.25 and p < 0.05). Weighted Sum model (WSM) derived top 5% genes also have higher expression in basal and lower in luminal A in association with STK3.
Conclusion
By introducing a novel bioinformatics approach that combines GSEA and WSM, the study successfully identified the top 5% of genes associated with higher expression of STK3.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer have many potential causes, often including a combination of genetic, hormonal, environmental, and lifestyle factors [1]. The diagnosis and treatment of breast cancer can be achieved by identification prognostic risk factors [2]. Sometimes mutations in certain genes, such as BRCA1 and BRCA2, and abnormality in signaling pathways are linked to an increased risk, which are complex and tightly controlled in normal development and regulations [1]. It has been shown that hippo pathways, which control cell proliferation, growth, and cell differentiation, are dysregulated in breast cancer compared to normal breast [3,4,5,6,7]. The Hippo pathway is an evolutionarily conserved regulator of tissue growth and cell fate during development, and regeneration and keeps tissues homeostasis. [8,9,10]. Mammalian Sterile 20-like kinases (STKs such as STK3 and STK4 which, respectively, known as MST2 and MST1), large tumor suppressor (LATS) kinases, Salvador homolog 1 (SAV1) scaffolding protein, monopolar spindle-one-binder kinase activator protein 1 (MOB1), and YAP (Yes-associated protein) are the main proteins that make up the canonical Hippo pathway in mammals. Mutation and deregulation for a subset of Hippo pathway genes have been reported in several malignancies, including breast cancer [8]. This relationship aids in the regulation of Lats1/2-Mob1 complexes by MST1/2, which phosphorylates and retains YAP/TAZ in the cytoplasm, inactivating downstream targets [11,12,13]. MST1/2 activation has been linked to tumor suppression and apoptosis, according to functional investigations [14, 15]. As a tumor suppressor, MST1 inhibits the spread of tumors and triggers apoptosis in breast cancer [14]. Patients who had diminished MST1 expression in breast cancer had a considerably reduced lifespan compared to individuals with high MST1 expression. There has been hypothesis suggesting MST1 expression is a predictive factor for people with breast cancer [16]. An essential interaction between ER and MST2 in breast cancer is implied by our prior analysis, which showed that the excellent predictive benefit of low MST2 was only detected within ER-positive breast cancer patients as opposed to ER-negative patients. Due to the fact that other cancer types did not exhibit this association.
The purpose of this study is to evaluate the STK3 higher expression patterns in different subtypes and their association with pathological stages of breast cancer patients. Patients were classified into higher and lower expression phenotypes of the STK3 gene using a median-based cut-off expression value.
In this study, the publicly available data The Cancer Genome Atlas (TCGA) and Molecular Taxonomy of Breast Cancer International Consortium (METABRIC) are analyzed from cBioPortal which can be founded at (https://www.cbioportal.org/) and STK3 expression is associated with the pathological stages and intrinsic subtypes is displayed in descriptive statistical form. The patient’s survival analysis is performed using the Kaplan–Meier method. The gene sets from hallmark and breast cancer-related pathways in the context of STK3 high and low phenotypes using Gene Set Enrichment Analysis (GSEA) [17] were analyzed. The statistically significant gene sets (pathways) that were commonly enriched in both datasets were selected, and the leading-edge genes were examined using Weighted Sum Model (WSM) and Nominal Group Technique (NGT) [18] to identify the top 5% of genes associated with STK3. Moreover, a consistent expression pattern was observed across both datasets regarding the distribution of patients among intrinsic subtypes. We concluded that STK3’s higher expression has a vital role in overall invasive ductal carcinoma (IDC) patients by indicating that the survivability of breast cancer patients is significantly decreased after 5 years (p < 0.05) compared to its lower expression. Furthermore, based on the clinical-pathological stage association, it has been observed that patients with an advanced stage also exhibit higher expression of STK3. The basal subtype is most strongly associated with higher expression of STK3, while lower expression is correlated with luminal A-type, suggesting a connection between STK3 expression levels and the degree of cancer aggressiveness.
In general, the bioinformatic study of the genomic profile of breast cancer could offer a hint for discovering potential biomarkers and help with treating patients individually based on their gene expression. Based on our study results, it can be inferred that STK3’s higher expression and related highly ranked gene expressions have the potential to be candidate biomarkers for intrinsic subtypes, particularly in basal and luminal A breast cancer subtypes that exhibit considerable variation for differential analysis.
Materials and methods
Data collection
Large public databases containing cancer-related data are widely accessible for researchers. The study utilized two such databases, from cBioPortal, which provided access to genomic and clinical data from large studies such as TCGA and METABRIC.
TCGA data collection and preprocessing
The TCGA data underwent various processing steps to ensure consistency and accuracy in subsequent statistical analyses. Initially, data from 1084 patients (RNA seq count data and clinical data) were retrieved from the TCGA database via cBioPortal. Patients with more than 70% missing clinical data were excluded, and missing clinical features were obtained using the TNM (Tumor, node, and metastasis) staging system data. Unique identifiers-Case-Id for transcriptome data and bcr_patient_barcode for clinical data—were used to map clinical and transcriptomic data. To handle multiple transcriptomes for the same patient, the mean expression of transcriptomic records was used. To ensure an adequate sample size, the study exclusively examined transcriptomes from primary tumor tissues where as small number of patients samples (around 5 for metastatic and 113 for solid tissue normal) were excluded from study. The final analysis included transcriptome data from 780 IDC patients. All data processing and analysis were performed using R software (Version 4.2.2).
METABRIC data collection and preprocessing
The METABRIC database, comprising 2509 primary breast tumors and 548 matched normal samples (RNA seq count data and clinical data), was obtained from cBioportal. The samples were uniquely identified using the cancer study identifier brca_metabric. It should be noted that the normal samples only had clinical data and no transcriptomic records were excluded from the study. Furthermore, around 57 patients with primary tissue were also excluded due to missing transcriptomic data. To ensure the quality of the dataset and retrieve consistent outcomes, we removed a total of 605 samples without transcriptomic data and merged the remaining clinical and transcriptomic data, which were mapped based on the brca_metabric identifier. To conduct our final analysis, we focused on 1500 IDC patients out of the 1904 samples in the dataset.
STK3 higher expression association with clinical features
Descriptive statistical analysis
Patients in both datasets were categorized into low and high phenotypes based on the STK3 median gene expression value. The patients with STK3 expression less than the median value were considered low phenotype, while those with expression higher than the median value were considered high phenotype. The STK3 expression was visualized in box plots in clinical-pathological stages as well as molecular subtypes of breast cancer using R programming.
Survival analysis
The survival analysis of both datasets was conducted using the Kaplan–Meier method in R, based on the STK3 higher and lower phenotypes. The analysis utilized two parameters: the time from the first diagnosis to the last follow-up or death, and the status of patients (i.e., alive, or deceased). Furthermore, the Cox regression model was employed to examine the hazard rate of the low and high STK3 categories of patients’ data.
GSEA analysis
To determine whether a set of previously defined genes exhibits statistically significant differences between two biological states, a computational technique known as GSEA is utilized [17]. In this study, GSEA was used to identify the associated upregulated pathways in the STK3 higher expression phenotype. The Hall Mark (h.all.v2022.1.Hs.symbols) and a query-driven gene set using the query “STK3, breast cancer, tumor microenvironment, nuclear translocation, genes regulations, hippo pathway, hypoxia, cell proliferation” from Gene card were used for GSEA analysis [20, 21]. Further analysis was performed by selecting pathways that were upregulated in the higher STK3 phenotype and were common to both data sets, with a p-value of less than 0.05 and an FDR of less than 25%. It is worth mentioning that no gene set with a significant p-value and matching threshold FDR value was observed in the STK3 low phenotype.
Leading edge genes analysis and top 5% genes derivation in association with STK3
After the selection of significant pathways enriched in higher expression of STK3 using a threshold of NES > 1.5, p < 0.05, and FDR < 25% were visualized in higher and lower expression context of STK3. To analyze the mean expression of leading edge genes for each subtype, heat map visualizations were used. The violin plots were used to display the expression pattern of significant pathways among breast cancer patients of different subtypes, categorized by SKT3 phenotype as either low or high. The line plots were used to depict the patient percentage for higher mean expression of leading edge genes across all significant pathways in TCGA and METABRIC datasets to observe trends of higher mean expression in the context of STK3’s higher phenotype across different subtypes of breast cancer. After all leading edge genes of significant pathways in SKT3 higher phenotypes were subjected to statistical analysis using WSM. The WSM used several features of genes, including the normalized enrichment score (NES) of the gene’s pathway, the running enrichment score (RES) of a gene in the gene sets/pathway, the total number of pathways that shared a given gene, and the scaled mean expression of the gene between the TCGA and METABRIC datasets (Table 1). We evaluated two sets of weights (Wt1, Wt2) from our team members closely working on STK3’s role in different diseases. The WSM was used to calculate the ranking based on the four criteria (Table 1). In WSM, the weights for each criterion were decided based on the NGT [18]. NGT is a group process that assists in selecting appropriate solutions to a problem based on the majority group member consensus. The problem was assigning the most moderate weights to criteria based on experts’ domain knowledge in the gene ranking process. This model enables the identification of the top 5% of genes that are associated with STK3.
where \(C_{i }\) represents gene ranking criteria shown in Table 2 i.e. \(C_i \left\{ {Gene_{NES} , Gene_{RES} , Gene_{cpw} ,Gene_{Exp} } \right\}and\; NGT\left( {C_i } \right)\) is a weight to \(C_{i }\) assign through NGT process.
Results
STK3 higher expression association with clinical characteristics
We analyzed STK3’s higher expression associated with the pathological stages and the patient’s distribution in intrinsic subtypes of breast cancer patients in TCGA and METABRIC datasets.
TCGA data (Fig. 1a) shows the patient proportion for STK3 higher expression increase precisely by moving from pathological lower stage I to higher stages III (I–II: 39% to 49%, II–III: 49% to 61%). In stage IV, the patient distribution is observed lower compared to stage III. So, the results remain inconclusive due to the relatively small proportion of patients (2.18%) in stage IV.
Based on the METABRIC data analysis (Fig. 1b), there is a clear increase in the proportion of patients with high-risk STK3 expression as breast cancer progresses from lower stage I to higher stages IV. Specifically, the proportion of patients with high-risk STK3 expression increases from 43 to 51% when moving from stage I to II, from 51 to 56% when moving from stage II to III, and from 56 to 67% when moving from stage III to IV. These findings suggest a positive association between higher STK3 expression and advanced stages of breast cancer.
STK3 is expressed differently amongst intrinsic molecular subtypes of breast cancer in TCGA and METABRIC datasets (Fig. 1c, d). STK3 higher expression pattern in each subtype for both dataset is given below.
-
Patients with Basal subtype showed higher STK3 expression levels in both TCGA and METABRIC datasets, with percentages of 62% and 68%, respectively. Additionally, the patient proportions of the Basal subtype in TCGA and METABRIC were 19.62% and 12.40%, respectively.
-
Both TCGA and METABRIC datasets showed a higher level of STK3 expression (58%) in patients with Luminal B subtype. The patient proportions for Luminal B subtype were 19.62% and 12.40% in TCGA and METABRIC, respectively.
-
Both TCGA and METABRIC datasets showed a higher expression of STK3 in 54% of patients (with a ratio of 9.23%) and 52% of patients (with a ratio of 13.00%), respectively in subtype Her2.
-
The proportion of patients with Luminal A subtype exhibiting higher STK3 expression was 37% and 40% in TCGA and METABRIC datasets, respectively. Notably, Luminal A patients represented the majority of samples in both datasets, accounting for 49.48% and 50.00% of TCGA and METABRIC samples, respectively.
The results suggest that the expression of STK3 varies among the intrinsic subtypes of breast cancer. Specifically, STK3’s higher expression is observed in the Basal subtype, while it is lower in the Luminal A subtype.
IDC patients’ survival in context of STK3 lower and higher expression
The Kaplan survival analysis for 5 to 10 years shows that breast cancer IDC patients with STK3 higher expression have low survivability compared to STK3 low expression both in TCGA and METABRIC data sets. The survival rates exhibit notable distinctions, with p-values of 0.027 and 0.016 in TCGA for patients classified as “dead with tumor” and all deceased IDC patients, respectively. In the METABRIC dataset, the survival rate for patients labeled as “dead with tumor” also demonstrates a significant difference with a p-value of 0.003. Figure 2 depicts the details of survival curves in both TCGA (a, b) and METABRIC (c).
The survival rate of IDC patients significantly correlates with STK3 expression, as indicated by multivariate analysis. In the TCGA dataset, lower expression shows a hazard ratio of 0.56 (95% CI 0.34–0.94, p = 0.029) for patients who died with tumors and 0.62 (95% CI 0.42–0.92, p = 0.017) for all deceased patients. In the METABRIC dataset, the hazard ratio for those deceased with tumors in lower expression is 0.76 (95% CI 0.64–0.91, p = 0.003).
Candidate Gene sets regulations in context of STK3 low and high expression
We get the most enriched pathways in both datasets with a p-value less than 0.05 and an FDR value < 25% using GSE analysis. A total of 7 pathways are commonly enriched in both datasets. The enrichements plots of 7 Pathways (Supplementary Figure S1) shows only pathways or genesets from which the top 5% of genes are associated with STK3 expression.
To identify signaling pathways that are differentially activated in breast cancer in both TCGA and METABRIC, GSEA was conducted between low and high STK3 expression data sets. GSEA reveals significant differences (FDR 0.25%, NOM p-value % 0.05) in the enrichment of MSigDB Collection (h.all. v2022. Symbols [22]). We selected the most significantly enriched signaling pathways based on their NES (Supplementary Figure S1 and Table S1). In Supplementary Figure S1 shows that GM2_check point, E2F targets, mitotic spindle, MTORC1 signaling, MYC targets V1, and unfolded_protein_response is differentially enriched in STK3 high-expression phenotype. Hallmarks can effectively associate with their corresponding protein activation phenotypes thus confirming their biological relevance.
Leading edge genes visualization in STK3 context
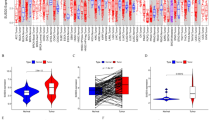
The seven significant gene sets and one selected top 5% genes in the context of STK3 low and higher phenotype and in different subtypes of breast cancer were visualized using heat maps and violin plots to display the leading edge genes. The results showed that for all significant gene sets, patients had a higher percentage of higher expression for the leading edge genes in the basal subtype and a lower percentage in Luminal A. This trend was observed in both data sets, TCGA and METABRIC (Fig. 3). [Higher resolution is shown in supplementary Fig. 3a].
a Heatmap Visualization of leading-edge genes (LEG) for statistically significant gene sets and the top 5% genes, in the context of STK3 in breast cancer subtypes [(BL: Basal, HR: HER2, LA: Luminal A, LB: Luminal B);(L: lowSTK3Expression, H: HighSTK3Expression)] of TCGA and METABRIC data sets. b LEG mean expression distribution across subtypes in the context of STK3 using data from TCGA and METABRIC. c Patterns of Patients percentage of LEG with high mean expression of statistically significant gene sets, in breast cancer subtypes Of TCGA and METABRIC datasets
Top 5% genes derivation in association with STK3
The WSM model derived the top 5% genes associated with STK3 relied on GSEA-derived features such as NES, RES, pathways (gene sets) number, and gene expression from patients TCGA and METABRIC data sets. The genes that were identified using this approach could provide valuable insights into the underlying mechanisms of breast cancer and may ultimately aid in the development of more effective treatments for this disease. These top 5% genes are HSPA8, HSP90AB1, NOP56, MCM6, TRA2B, and PRDX4, and have literature evidence to have a role in breast cancer proliferation [22,23,24,25,27]. Table 3 highlights the detailed features of each of the top 5% genes with overall score and ranks. Although, Wt1 score criteria were used, Table 2 also provided the gene score and overall score and ranks using Wt2 criteria.
Discussion
The Hippo pathway component has a substantial role in regulating the cell cycle, growth, proliferation, and maintaining tissue homeostasis. Furthermore, it inhibits the development and occurrences of malignancy tightly controlled under normal conditions depending on the types of signaling. In a study of human sarcoma tumorigenesis, the epigenetic alteration effect was observed for STK3(MST2) in signaling pathway of Sav-RASSF1-Hpo.[28]. Furthermore, STK3’s lower expression is correlated with poor prognosis in ovarian cancer, and higher expression inhibits the cell proliferation, and migration of ovarian cancer cells and promotes apoptosis [29].
In a study of gastric carcinogenesis, STK3 was discovered to be an independent prognostic biomarker that mediates cell cycle progression by activating Ras-MAPK pathways [19].
Our previous work has proved that the expression of two components STK3 (MST2) and SAV of the hippo pathway was associated with ERα phosphorylation and transactivation and represses ERα gene expression. Silencing of STK3 can inhibit breast cancer in vitro experiments using MCF-7 cells and showed that its higher expression leads to ERα activation in the absence of ligand [30].
We aimed to analyze STK3 as a potential prognostic molecular marker of poor survival. Bioinformatic analysis in this study showed that STK3 has higher expression levels in basal types and lower expression in luminal A type of breast cancer patients. Furthermore, its higher expression is closely related to the poor prognosis of IDC breast cancer patients.
We observed that the STK3 higher expression is associated with higher stages in both datasets, but for only TCGA data, the stage IV patients had comparatively low STK3 expression. One possible reason could be a smaller number of patients compared to other stages, or different biological and molecular mechanisms are involved.
By using the statistical model WSM and NGT based ranking on GSEA-derived features such as NES, enrichment score, and pathways number the top 5% genes were derived in STK3 higher phenotype-based GSEA leading edge genes. The genes that were identified using this approach could provide valuable insights into the underlying mechanisms of breast cancer and may ultimately aid in the development of more effective treatments for this disease. For example, HSPA8, HSP90AB1, NOP56, MCM6, TRA2B, and PRDX4, were identified as genes that were analyzed in STK3 higher expression phenotype study. These genes have already been observed through some preliminary studies which are evidently having a role in breast cancer proliferation [22,23,24,26, 31].
One study indicates the potential molecular mechanism that promotes the evolution of TNBC (triple negative breast cancer) related with the poor clinical outcome of TNBC is associated with high expression of HSPA8 [32]. High-level expression of HSP90AB1, one of cytoplasmic HSP90 isoforms was correlated with poor prognosis in different subtypes of breast cancer and was driven by chromosome coding region amplifications and were independent factors that led to death from breast cancer among patients with triple-negative (TNBC) and HER2-/ER + subtypes [23]. MCM6 is known as a specific biomarker of cancer in many cancer types including breast cancer. its expression level, and biological function in various types of cancer is complicated and have remain uncleared up to date [33]. TRA2B is in association with several other genes and its product involve in breast cancer metastasis and was identified as cancer hall mark [26]. PRDX4 antioxidant protein has been shown to causally facilitate tumor initiation and propagation, therapeutic resistance, and subsequent recurrence of many types of tumors. The mechanisms of how PRDX4 works in different cancers requires more in depth research [31]. NOP56 is located at the key crossroads of many signaling pathways and plays an important role related to the occurrence and development of various tumors. Although the role and mechanism of NOP56 are still unclear. However, it is one of many methylated genes, and examining the methylation status of genes can help identify tumor-specific markers and therapeutic targets for cancer patients [34].
This finding suggests that these genes may play a role in the development of breast cancer in patients with high levels of STK3 expression. This study is novel to exploit STK3 expression in different intrinsic subtypes of breast cancer at the more granular level using two data sets. Consideration of underlying molecular mechanisms in association with hippo pathways genes especially STK3 could lead to develop the targeted therapy for a more aggressive type like basal or triple-negative cancer type of breast cancer and improve the patient’s life quality.
Several studies have implied the NGT to involve stakeholders and gather their views and opinions to develop consensus-based healthcare decisions [18]. Some common examples include establishing end-of-life care preferences, prioritizing treatment decisions, highlighting chronic disease issues, and developing research-based guidelines. This work employed NGT techniques to rank genes and reach a consensus on weighting gene attributes retrieved from GSEA. Bioinformaticians, and medical data scientists determined WSM model weights for Wt1 and Wt2, consequently finalized through NGT process. Wt1 was selected using consensus-based NGT integration.
Conclusion
STK3 has been studied in breast cancer as a potential prognostic molecular marker of poor survival. This study is novel to exploit STK3 expression in different intrinsic subtypes of breast cancer at the more granular level using two data sets. The weighted sum statistical model based on GSEA-derived leading edge genes in STK3 higher phenotypes are the genes having evidence in breast cancer proliferation. Consideration of underlying molecular mechanisms in association with hippo pathways genes especially STK3 could lead to develop the targeted therapy for a more aggressive type like basal or triple-negative cancer type of breast cancer and improve the patient’s life quality.
Data availability
The data used in this study is openly accessible and can be found at https://www.cbioportal.org/datasets. Specifically, the following datasets were employed: (1) Breast Invasive Carcinoma (TCGA, PanCancer Atlas 2018). (2) Breast Cancer (METABRIC, Nature 2012 & Nat Commun 2016).
References
Feng Y, Spezia M, Huang S, Yuan C, Zeng Z, Zhang L, Ji X, Liu W, Huang B, Luo W, Liu B, Lei Y, Du S, Vuppalapati A, Luu HH, Haydon RC, He TC, Ren G (2018) Breast cancer development and progression: risk factors, cancer stem cells, signaling pathways, genomics, and molecular pathogenesis. Genes Dis 5:77–106. https://doi.org/10.1016/j.gendis.2018.05.001
Yu F, Quan F, Xu J, Zhang Y, Xie Y, Zhang J, Lan Y, Yuan H, Zhang H, Cheng S, Xiao Y, Li X (2019) Breast cancer prognosis signature: linking risk stratification to disease subtypes. Brief Bioinform 20:2130–2140. https://doi.org/10.1093/bib/bby073
Pan D (2010) The hippo signaling pathway in development and cancer. Dev Cell 19:491–505. https://doi.org/10.1016/j.devcel.2010.09.011
Park Y, Park J, Lee Y, Lim W, Oh BC, Shin C, Kim W, Lee Y (2011) Mammalian MST2 kinase and human salvador activate and reduce estrogen receptor alpha in the absence of ligand. J Mol Med 89:181–191. https://doi.org/10.1007/s00109-010-0698-y
Sun YS, Zhao Z, Yang ZN, Xu F, Lu HJ, Zhu ZY, Shi W, Jiang J, Yao PP, Zhu HP (2017) Risk factors and preventions of breast cancer. Int J Biol Sci 13:1387–1397. https://doi.org/10.7150/ijbs.21635
Wu L, Yang X (2018) Targeting the hippo pathway for breast cancer therapy. Cancers (Basel). https://doi.org/10.3390/cancers10110422
Zheng Y, Pan D (2019) The Hippo signaling pathway in development and disease. Dev Cell 50:264–282. https://doi.org/10.1016/j.devcel.2019.06.003
Dong J, Feldmann G, Huang J, Wu S, Zhang N, Comerford SA, Gayyed MF, Anders RA, Maitra A, Pan D (2007) Elucidation of a universal size-control mechanism in drosophila and mammals. Cell 130:1120–1133. https://doi.org/10.1016/j.cell.2007.07.019
Fallahi E, O’Driscoll NA, Matallanas D (2016) The MST/hippo pathway and cell death: a non-canonical affair. Genes 7:28
Oka T, Mazack V, Sudol M (2008) Mst2 and lats kinases regulate apoptotic function of yes kinase-associated protein (YAP)*. J Biol Chem 283:27534–27546. https://doi.org/10.1074/jbc.M804380200
Ni L, Li S, Yu J, Min J, Brautigam Chad A, Tomchick Diana R, Pan D, Luo X (2013) Structural basis for autoactivation of human Mst2 kinase and its regulation by RASSF5. Structure 21:1757–1768. https://doi.org/10.1016/j.str.2013.07.008
Thompson BJ, Sahai E (2015) MST kinases in development and disease. J Cell Biol 210:871–882. https://doi.org/10.1083/jcb.201507005
Zhao B, Lei QY, Guan KL (2008) The Hippo–YAP pathway: new connections between regulation of organ size and cancer. Curr Opin Cell Biol 20:638–646. https://doi.org/10.1016/j.ceb.2008.10.001
Turunen SP, von Nandelstadh P, Öhman T, Gucciardo E, Seashore-Ludlow B, Martins B, Rantanen V, Li H, Höpfner K, Östling P, Varjosalo M, Lehti K (2019) FGFR4 phosphorylates MST1 to confer breast cancer cells resistance to MST1/2-dependent apoptosis. Cell Death Differ 26:2577–2593. https://doi.org/10.1038/s41418-019-0321-x
Wang Y, Li J, Gao Y, Luo Y, Luo H, Wang L, Yi Y, Yuan Z, Jim Xiao Z-X (2019) Hippo kinases regulate cell junctions to inhibit tumor metastasis in response to oxidative stress. Redox Biol 26:101233. https://doi.org/10.1016/j.redox.2019.101233
Lin XY, Cai FF, Wang MH, Pan X, Wang F, Cai L, Cui RR, Chen S, Biskup E (2017) Mammalian sterile 20-like kinase 1 expression and its prognostic significance in patients with breast cancer. Oncol Lett 14:5457–5463. https://doi.org/10.3892/ol.2017.6852
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102:15545–15550. https://doi.org/10.1073/pnas.0506580102
McMillan SS, Kelly F, Sav A, Kendall E, King MA, Whitty JA, Wheeler AJ (2014) Using the nominal group technique: how to analyse across multiple groups. Health Serv Outcomes Res Method 14:92–108. https://doi.org/10.1007/s10742-014-0121-1
Chen B, Chan WN, Mui CW, Liu X, Zhang J, Wang Y, Cheung AHK, Chan AKY, Chan RCK, Leung KT, Dong Y, Pan Y, Ke H, Liang L, Zhou Z, Wong CC, Wu WKK, Cheng ASL, Yu J, Lo KW, To KF, Kang W (2021) STK3 promotes gastric carcinogenesis by activating Ras-MAPK mediated cell cycle progression and serves as an independent prognostic biomarker. Mol Cancer 20:147. https://doi.org/10.1186/s12943-021-01451-2
Liberzon A, Birger C, Ghandi M, Jill P, Tamayo P, Jolla L, Jolla L (2015) The molecular signatures database (MSigDB) hallmark gene set collection. Cell Syst 1:417–425
Stelzer G, Rosen N, Plaschkes I, Zimmerman S, Twik M, Fishilevich S, Iny Stein T, Nudel R, Lieder I, Mazor Y, Kaplan S, Dahary D, Warshawsky D, Guan-Golan Y, Kohn A, Rappaport N, Safran MD (2016) Lancet, the genecards suite: from gene data mining to disease genome sequence analyses. Curr Protoc Bioinform 1.30.1-1.30.33
Liberzon A, Birger C, Thorvaldsdóttir H, Ghandi M, Mesirov Jill P, Tamayo P (2015) The molecular signatures database hallmark gene set collection. Cell Syst 1:417–425. https://doi.org/10.1016/j.cels.2015.12.004
Cheng Q, Chang JT, Geradts J, Neckers LM, Haystead T, Spector NL, Lyerly HK (2012) Amplification and high-level expression of heat shock protein 90 marks aggressive phenotypes of human epidermal growth factor receptor 2 negative breast cancer. Breast Cancer Res 14:R62. https://doi.org/10.1186/bcr3168
Hou L, Chen M, Wang M, Cui X, Gao Y, Xing T, Li J, Deng S, Hu J, Yang H, Jiang J (2016) Systematic analyses of key genes and pathways in the development of invasive breast cancer. Gene 593:1–12. https://doi.org/10.1016/j.gene.2016.08.007
Issac MSM, Yousef E, Tahir MR, Gaboury LA (2019) MCM2, MCM4, and MCM6 in breast cancer: clinical utility in diagnosis and prognosis. Neoplasia 21:1015–1035. https://doi.org/10.1016/j.neo.2019.07.011
Park S, Brugiolo M, Akerman M, Das S, Urbanski L, Geier A, Kesarwani AK, Fan M, Leclair N, Lin K-T, Hu L, Hua I, George J, Muthuswamy SK, Krainer AR, Anczuków O (2019) Differential functions of splicing factors in mammary transformation and breast cancer metastasis. Cell Rep 29:2672-2688.e2677. https://doi.org/10.1016/j.celrep.2019.10.110
Schirmer AU, Driver LM, Zhao MT, Wells CI, Pickett JE, O’Bryne SN, Eduful BJ, Yang X, Howard L, You S, Devi GR, DiGiovanni J, Freedland SJ, Chi JT, Drewry DH, Macias E (2022) Non-canonical role of hippo tumor suppressor serine/threonine kinase 3 STK3 in prostate cancer. Mol Ther 30:485–500. https://doi.org/10.1016/j.ymthe.2021.08.029
Seidel C, Schagdarsurengin U, Blümke K, Würl P, Pfeifer GP, Hauptmann S, Taubert H, Dammann R (2007) Frequent hypermethylation of MST1 and MST2 in soft tissue sarcoma. Mol Carcinog 46:865–871. https://doi.org/10.1002/mc.20317
Wang X, Wang F, Zhang ZG, Yang XM, Zhang R (2020) STK3 Suppresses ovarian cancer progression by activating NF-κB signaling to recruit CD8(+) T-cells. J Immunol Res 2020:7263602. https://doi.org/10.1155/2020/7263602
Park J, Kim GH, Lee J, Phuong BTC, Kong B, Won JE, Won GW, Lee YH, Han HD, Lee Y (2020) MST2 silencing induces apoptosis and inhibits tumor growth for estrogen receptor alpha-positive MCF-7 breast cancer. Toxicol Appl Pharmacol 408:115257. https://doi.org/10.1016/j.taap.2020.115257
Jia W, Chen P, Cheng Y (2019) PRDX4 and its roles in various cancers. Technol Cancer Res Treat 18:1533033819864313. https://doi.org/10.1177/1533033819864313
Ying B, Xu W, Nie Y, Li Y (2022) HSPA8 is a new biomarker of triple negative breast cancer related to prognosis and immune infiltration. Dis Markers 2022:8446857. https://doi.org/10.1155/2022/8446857
Zeng T, Guan Y, Li YK, Wu Q, Tang XJ, Zeng X, Ling H, Zou J (2021) The DNA replication regulator MCM6: an emerging cancer biomarker and target. Clin Chim Acta 517:92–98. https://doi.org/10.1016/j.cca.2021.02.005
Zhao S, Zhang D, Liu S, Huang J (2023) The roles of NOP56 in cancer and SCA36. Pathol Oncol Res. https://doi.org/10.3389/pore.2023.1610884
Funding
This research was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education, Science, and Technology (2022R1F1A1069631) to YJL and by Korea Ministry of Environment (MOE) as Graduate School specialized in Climate Change.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Data curation, Methodology, Formal analysis, and Writing—original draft was performed by Rukhsana. Data curation, Writing—review & editing were performed by ATS. Conceptualization, Methodology, Writing—review & editing were performed by MH and Conceptualization, mentoring, Writing—review & editing were performed by YL. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rukhsana, Supty, A.T., Hussain, M. et al. STK3 higher expression association with clinical characteristics in intrinsic subtypes of breast cancer invasive ductal carcinoma patients. Breast Cancer Res Treat 206, 119–129 (2024). https://doi.org/10.1007/s10549-024-07248-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-024-07248-3