Abstract
Tuberculosis (TB) caused by Mycobacterium tuberculosis (MTB) is a global threat to human health hence better understanding of the MTB pathogenesis for improved therapeutics requires immediate attention. Emergence of drug-resistant strains has stimulated an urgent need for adopting new strategies that could be implemented to control TB. One of the contributing mechanisms by which MTB evades drug doses is overexpression of drug efflux pumps. Thus blocking or modulating the functionality of efflux pumps represents an attractive approach to combat drug resistance. Iron is a critical micronutrient required for MTB survival and not freely available inside the host. In this study, we demonstrated that iron deprivation impairs drug efflux pump activity and confers synergism for anti-TB drugs in presence of efflux pump inhibitors against MTB. Mechanistic insights revealed that iron deprivation inhibit resistance nodulation division superfamily transporter activity. This was evident from enhanced Nile red accumulation and reduced expression of MmpL3, a transmembrane promising target involved in mycolic acid transport across membrane. Furthermore, iron deprivation led to abrogated MA transport particularly of class methoxy-MA which was confirmed by TLC and mass spectrometry based lipidome analysis. Additionally, iron deprivation leads to enhanced membrane fluidity in MTB. Together, MmpL3 being a promiscuous anti-TB target, metal chelation strategy could be adopted to boost the effectiveness of current anti-TB drug regimes to combat drug resistance TB.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
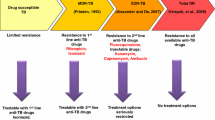
Tuberculosis (TB) caused by Mycobacterium tuberculosis (MTB) remains one of the important underlying causes of mortality worldwide. Widespread and prolonged deployment of anti-TB agents has led to emergence of drug resistance in MTB, which poses a serious threat to available therapy. Multidrug resistant tuberculosis (MDR-TB) results due to simultaneous resistance towards two frontline anti-TB drugs, isoniazid and rifampicin (Mezwa et al. 2018). Similarly, extensively drug resistant TB (XDR-TB) is caused by MTB strains which are resistant to isoniazid, rifampicin, a fluoroquinolone and one of the three second line injectables: amikacin, kanamycin, or capreomycin (WHO 2016). Therefore, it is imperative to understand effective strategies to cope with the challenges of drug resistance.
Among major mechanisms of MDR which includes alteration in membrane permeability and mutations in genes, overexpression of drug efflux pumps is a significant contributor that helps MTB to counter the adverse effects of antibiotics (Tanwar et al. 2014; Hegeto et al. 2018). Rapid efflux in resistant strains ensures that the drug does not get accumulated to lethal levels. Thus much attention has been focused on the search of efflux pumps inhibitors which may act as adjuvants of TB therapy therefore identification of strategies capable to block the function or overexpression of efflux pumps remains an area of immense significance. Efflux pumps are transmembrane proteins which expel the incoming drugs thereby prevent toxic intracellular accumulation of drugs resulting in development of MDR (Machado et al. 2012; Schmalstieg et al. 2012). Based on the utilization of energy from ATP or proton motive force (PMF) there are five main classes of transport proteins which are mainly responsible for the development of resistance; (i) ATP-binding cassette (ABC) superfamily, (ii) the major facilitator superfamily (MFS); (iii) the multidrug and toxic compound extrusion (MATE) family, (iv) the small multidrug resistance (SMR) family; and (v) the resistance nodulation division (RND) superfamily (Balganesh et al. 2012; Pal et al. 2014).
Host’s sequestration of essential micronutrients to arrest the growth and development of MTB during an active infection, is termed as nutritional immunity. Iron is one of the most significant micronutrient required for both host and pathogen. It is estimated that 25-30% of all enzymes require metals as cofactor; among which iron is critical for cellular events like DNA biosynthesis, TCA cycle and oxidative stress defense (Conor 2007). However, iron being a transition metal is not readily available inside the host hence there is constant struggle between the host and MTB for the limited amount of iron. Siderophore driven iron acquisition in MTB is required for virulence and establishment of infection (Sritharan 2016). Targeting iron-dependent processes thus represents a viable anti-TB strategy. Indeed, there are a growing number of evidences which suggests that targeting iron homeostasis is a new approach that enhances the efficiency of known anti-TB drugs (Hameed et al. 2015; Kurthkoti et al. 2017; Patel et al. 2018). The previously we have established the effect of iron deprivation on drug susceptibilities of known anti-TB drugs where disruption of membrane homeostasis (Pal et al. 2015) and virulence traits (Pal et al. 2016) were demonstrated. The present study was therefore designed for addition in the existing literature and to evaluate the impact of iron deprivation on drug efflux transporter activity in not only Mycobacterium smegmatis but pathogenic MTB. To the best of our knowledge, this is the first report elaborating the effect of iron deprivation on efflux pump activities. Understanding the underlying mechanisms could provide an important insight into TB pathogenesis and predict the future strategies to combat MDR-TB.
Materials and methods
Material
All Media chemicals Middlebrook 7H9 broth, Middlebrook 7H10 agar, albumin/dextrose/catalase (ADC), oleic acid/albumin/dextrose/catalase (OADC) supplements were purchased from BD Biosciences (USA). Tween-80, Ethambutol (EMB), Isonizid (INH), 1,6 diphenyl hexatriene (DPH) were purchased from Sigma-Aldrich (St. Louis, MO, USA). 2,2, Bipyridyl (2,2 BP), ethidium bromide (EtBr), dinitrophenol (2,4, DNP), rifampicin (RIF) and Nile red (NR) were purchased from Himedia (Mumbai, India). Dimethyl sulfoxide (DMSO), potassium chloride (KCl), sodium chloride (NaCl), di-sodium hydrogen orthophosphate (Na2HPO4), potassium di-hydrogen orthophosphate (KH2PO4), sodium dodecyl sulphate (SDS), glycerol and d-glucose were obtained from Fischer Scientific.
Bacterial strains and culture conditions
MTB (H37Rv) and ∆PknG (mutant of Protein kinase G) along with M. smegmatis mc2155 (wild type) and siderophore mutant ∆011-14 were used in this study. Mycobacterium cells were grown in 7H9 Middlebrook (BD Biosciences) broth supplemented with 0.05% Tween-80, 0.2% glycerol and 10% ADC in 100 ml flasks (Schott Duran) and cultures were incubated at 37 °C at 150 rpm till the exponential phase reaches.
Establishment of iron deprived condition
Iron restricted condition was established by the use of 2,2, BP, a well known specific iron chelator, as described previously (Pal et al. 2015). M. smegmatis cells were exposed with different concentrations of 2,2, BP. Growth curve was performed for 72 h with 0.1 OD600 of cells in the presence of 2,2, BP (25, 30 and 35 µg/ml) and found that 35 µg/ml was subinhibitory enough for growth of M. smegmatis (Fig. S1A). Similarly, the sub-inhibitory concentration of 2,2, BP against MTB was standardized and subsequent growth curve experiment was performed for 21 days with 0.1 OD600 cells in the presence of 2,2, BP (40, 50 and 60 µg/ml) and found that 40 µg/ml was sub-inhibitory for MTB (Fig. S1B). Sub-inhibitory concentration is defined as concentration at which 2,2 BP was sufficient to chelate iron without causing noticeable growth defect. Concentration higher than 35 μg/ml in M. smegmatis and 40 μg/ml in MTB trigger growth inhibition and hence cannot be used for further experiments (Table 1). Each experiment was performed three times, and growth curves were generated using Microsoft Excel (Microsoft, USA). Additionally, the specificity of 2,2, BP was further confirmed by supplementation of growth inhibition by 2,2, BP with various metals that could only be reverted by iron (Fig. S1C).
Checkerboard assay
Two-dimensional broth microdilution checkerboard assay was performed as described earlier (Jin et al. 2010). Two hundred microliters of sterile deionized water was added to all outer-perimeter wells of the plates. The plates were prepared by dispensing the serially diluted antibiotics (INH, EMB, RIF) in the y-axis and 2,4, DNP in the x-axis in a iron sufficient or deprived media. Aliquots of 0.1 ml of inoculum adjusted to OD600 0.1 and diluted 1:100 were transferred to each well of the 96-well plate and incubated at 37 °C. The inoculated plates were sealed in plastic bags and incubated at 37 °C for 7 days and the results were interpreted by taking absorbance at 600 nm. The fractional inhibitory concentration index (FICI) was calculated and interpreted as follows: FICI ≤ 0.5, synergism; FICI > 0.5 < 4, indifference and FICI ≥ 4, antagonism (Odds 2003).
EtBr efflux
The efflux of EtBr was determined by using protocol described previously (Jin et al. 2010; Rodrigues et al. 2011). Approximately 1 × 106 cells were incubated until exponential phase under iron deprivation conditions. Cells were pelleted, washed twice with phosphate buffer saline (PBS) and resuspended in such a way so as to form 2% cell suspension (0.2 g of cell pellet weight dissolved in 10 ml PBS). The cells were then de-energized with an efflux pump inhibitor 2,4 DNP (20 µg/ml) in PBS (without glucose). The de-energized cells were pelleted, washed, and then resuspended as a 2% cell suspension (w/v) in PBS to which EtBr was added at a final concentration of 4 µg/ml and incubated for 45 min at 30 °C. The equilibrated cells with EtBr were then washed and resuspended in such a way so as to form 2% cell suspension (w/v) in PBS (with or without glucose 0.4%) for EtBr. Samples with a volume of 2 ml were withdrawn at the indicated time points as mentioned in the figure and centrifuged at 10,000 rpm for 1 min. The supernatant was collected, and absorption was measured at 285 nm. Glucose-free negative controls were included in all the experiments.
EtBr and NR influx
The EtBr accumulation was assessed by fluorescence assay with minor modifications (Rodrigues et al. 2011). Briefly, mid-log-phase cultures were washed with PBS containing 0.05% Tween-80 (PBST) and then stained with 4 µg/ml of EtBr. For NR uptake, mid-log-phase cultures were washed with PBS and then stained with 1 µg/ml NR (Chuang et al. 2015). For both assays, cells were incubated in 12-well plates, and analysis was performed after 30 min at excitation 544 nm and emission 590 nm with fluorescent microscope (CosLab). All experiments were repeated at least three times to confirm reproducibility.
RT-PCR
RNA was extracted from the control and iron deprived (2,2 BP) MTB cells by standard TRIzol protocol (Pal et al. 2018a). Reverse transcriptase (RT) PCR was performed as described in the RevertAid H Minus kit (Invitrogen) (Pal et al. 2018a). Briefly, 5 μg isolated RNA was DNase treated at 37 °C for 30 min and reaction was terminated by adding 1 μl of 25 mM EDTA and incubated at 65 °C for 60 min. RNA was subsequently primed with oligo (dT)18 for cDNA synthesis at 42 °C for 60 min. The synthesized cDNA product (2 μl) was directly used for PCR amplification reaction (50 μl) using gene specific forward and reverse primers. The synthesized cDNA product (2 μl) was directly used for PCR amplification reaction (50 μl) using gene specific forward and reverse primers (Table S1). The amplified products were gel electrophoresed and the densities of bands (for gene of interest) were measured and quantified using GelDoc by normalizing to that of the constitutively expressed 16 s gene using software.
Cell surface lipid extraction
Cell surface lipids were extracted from Control and iron restricted (2,2, BP) MTB cells according to previously described methodology (Minnikin et al. 1975) with few modifications. Briefly, approximately 1 × 106 cells iron deprivation were incubated until exponential phase, pelleted, washed twice with phosphate buffer saline (PBS) and centrifuged at 10,000 rpm for 5 min. Pellet was then mixed with methanol (5 ml), Toluene (5 ml) and concentrated sulphuric acid (0.2 ml) in a 20-ml screw cap tube (Poly Tetra Fluoro Ethylene lined). The contents of the tube were mixed thoroughly and methanolysis was allowed to proceed for 12–16 h at 75 °C (stationary incubation). The reaction mixture was allowed to cool and mycolic acid (MA) was extracted by adding 1 ml of hexane. After vigorous shaking the mixture was allowed to settle and the upper hexane layer containing MA was collected. MA was spotted on TLC plates (silica gel G) in solvent system hexane/ethyl acetate (19:1, v/v) as previously described (Slayden and Barry 2001). For estimation carbol fuchsin dye was added in the ratio of (1:1) to the extracted hexane layer containing MA and spectrum was made between the wavelength 400 and 600 nm range (Khanuja et al. 2004).
Ultra Performance Liquid Chromatography–Electrospray Ionization and Mass Spectrometry (UPLC–ESI-MS)
The isolated cell surface lipids under iron deprivation were qualitatively analyzed on triple quadrupole tandem mass spectrometer ACQ-TQD#QBB1152. LC was done on a C18 column (100 mm × 3 mm, 2.6 µm) and elution was done for 30 min, using 5% isopropanol, 90% methanol and 5% ammonium acetate (5 mM, pH 6.5), at a flow rate, 0.1 ml/min. The source temperature was 120 °C, the desolvation temperature was 350 °C and the cone voltage was set at 40 volts (V). The sample was introduced using an autosampler with 1 µL of sample injection volume. Capillary voltage was set to 3.50 kV. The data was recorded in the mass range, m/z 200–2000 and processing was performed with the Masslynx software, where each chromatogram was smoothened and the background was subtracted. The analysis of the acquired data was performed by MS-LAMP software (Sabareesh and Singh 2013; Layre et al. 2011). This software is a graphical user interface (GUI) standalone programme built using Perl::Tk. It is a combination of MTB lipid database (www.mrl.colostate.edu) and Lipid Metabolites and Pathways Strategy Consortium (LIPID MAPS; www.lipidmaps.org). The most probable adduct was recognized to be [M–H]+ and 0.5 window range was used for analysis (Pal et al. 2017, 2018b). Further, it needs to be noted that the data herein are analyzed qualitatively only. All the experiments were performed in triplicates to ensure reproducibility and accuracy.
Membrane fluidity
MTB membrane fluidity was estimated using protocol described earlier with modifications (Bessa et al. 2018). Approximately 1 × 106 cells were grown for 12 days in 37 °C at 225 rpm in the absence (control) and presence of 2,2, BP. Cells were then harvested and washed twice with PBS at 13,000 rpm at 4 °C for 10 min, equalize the cells to absorbance 0.5 at wavelength 600 nm and fixed with 2% paraformaldehyde for 15 min. Cells were then washed with SOE buffer and resuspended in labeling buffer in such a way that each ml contains 4 × 108 cells. Later 4 µM DPH (in tetrahydrofuran) was added and incubated for 2 h with continuous shaking. Readings was recorded using spectrofluorimeter at both excitation wavelength 360 nm in 5 nm slit and emission wavelength 426 nm in 10 nm slit. Experiment was performed in replicates to get the mean value.
Statistical analysis
The results were represented by the mean expression values of three independent experiments run in triplicate ± SD values and analyzed using Student’s t test where in P < 0.05 was considered as statistically significant denoted by (*).
Results and discussion
Iron deprivation confers synergy to combination of drug efflux inhibitor and anti-TB drugs
Efflux pump inhibitors can influence drug susceptibility phenotypes in drug-susceptible, drug-tolerant and clinical isolates, although to varying degrees. Efflux pump inhibitors such as carbonyl cyanide m-chlorophenyl hydrazone (CCCP), verapamil and 2,4-dinitro phenol (2,4-DNP) have enhanced the accumulation of drugs possibly due to hindrance in active efflux transporter (Banerjee et al. 1996). Efflux transporter proteins are involved in resistance to anti-TB drugs, bedaquiline and clofazimine (Almeida et al. 2016; Kaniga et al. 2016). Likewise, there are numerous studies where combinatorial effect of anti-TB drugs has been studied with drug efflux pump inhibitors (Mullin et al. 2004; Gupta et al. 2014; Caleffi-Ferracioli et al. 2016; Chen et al. 2018). Hence, we sought to examine the effect of iron deprivation on FICI for the combination of anti-TB drugs (INH, RIF, EMB) with 2,4, DNP (efflux pump inhibitor). Using the two dimensional checkerboard method we studied the interaction profiles of various anti-TB drugs and drug efflux inhibitor firstly in surrogate of MTB i.e. M. smegmatis under iron deprivation. We observed that the FICI for anti-TB drugs and efflux pump inhibitor which was higher than 0.5 under iron sufficient condition became less than 0.5 under iron deprivation thus depicting synergy (Table 2). To confirm whether the observed synergy is the effect of iron deprivation only, we used M. smegmatis siderophore mutant (∆011-014). Siderophores are strong iron chelators synthesized by mycobacteria under iron deprivation to scavenge iron (Rodriguez and Smith 2003; Ojha and Hatfull 2007). Thus any defect of iron–exochelin siderophore uptake mutant results in inherent iron deprived condition without adding 2,2, BP. We observed that ∆011-014 corresponded with the results using iron chelator 2,2 BP (Table 2). Specifically, we could detect synergism with INH and RIF in comparison to EMB. To substantiate our findings in the pathogenic context, we performed checkerboard assay and to our expectation, we explored similar synergies with INH and RIF under iron deprivation in MTB (Fig. S2). This was further supported by use of MTB serine threonine protein kinase G mutant (∆pknG) which is known to govern nutrient utilization (Khan et al. 2017). We found that kinase mutant also displayed synergy albeit of low grade (Table 2). The degree of synergy was however variable for each drug and in the following order INH > RIF > EMB. Thus either pharmacological or genetic inhibition of iron availability renders synergistic response in MTB towards combination of anti-TB drugs and efflux pump inhibitor.
Iron deprivation leads to enhanced EtBr accumulation
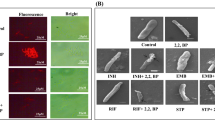
Taking clue from above observations, we examined the effect of iron deprivation on efflux pump activities by monitoring the intracellular accumulation of EtBr which is a widely used model substrate to measure the influx and efflux in MTB (Mullin et al. 2004). We estimated the EtBr accumulation through fluorescence imaging in M. smegmatis and MTB and found that most EtBr is accumulated in both de-energized controls in absence of glucose depicted by high fluorescence (Fig. 1). As expected, in energized cells with glucose, the efflux pumps are active hence negligible fluorescence is observed since most EtBr is effluxed out of the cell (Fig. 1). Contrary, under iron deprivation, despite the presence of glucose, the fluorescence is observed in both the mycobacterial strains (Fig. 1). These observations confirm that iron deprivation causes EtBr accumulation possibly due to some defect in efflux pump activities and/or expression.
Effect of iron deprivation on accumulation of EtBr. Fluorescence microscopy images depicting EtBr (4 μg/ml) accumulation under different iron deprived conditions in the absence and presence of glucose. Left panel depict EtBr accumulation in M. smegmatis whereas right panel depicts EtBr accumulation in MTB. Images were taken at × 100 magnification. Scale bar depicts 50 μm length. Bar graphs represent fluorescence integrated intensity of left and right panels. Error bar was calculated by taking the average of intensities measured using Image J software
Iron deprivation leads to abrogated EtBr efflux
Next, we studied the effect of iron deprivation on drug efflux activity more closely. Firstly, we monitored the extracellular EtBr efflux in time dependent manner in M. smegmatis and found that EtBr efflux was considerably inhibited under iron deprivation (Fig. 2a). Similarly, when EtBr efflux was monitored in MTB, we observed that extracellular concentration of EtBr was diminished in both iron deprived and ∆PknG mutant cells (Fig. 2b). These observations suggested that iron deprivation affects efflux pump activities.
EtBr efflux under iron deprivation. a EtBr efflux displayed by extracellular concentrations of EtBr for M. smegmatis cells grown in the absence (control), presence of 2,2, BP (35 μg/ml) and ∆011-14. Mean of OD285 ± SD of three independent sets of experiments are depicted on y-axis with respect to time (minutes) on x-axis (*depicts significant difference with P < 0.05). b EtBr efflux displayed by extracellular concentrations of EtBr for MTB cells grown in the absence (control), presence of 2,2, BP (40 μg/ml) and ∆PknG. Mean of OD285 ± SD of three independent sets of experiments are depicted on y-axis with respect to time (minutes) on x-axis (*depicts significant difference with P < 0.05)
Further we accessed the efflux activity of Mycobacterium cells under iron deprivation in the presence of anti-TB drugs (INH, RIF and EMB). In M. smegmatis, we observed that in comparison to the EtBr efflux in presence of anti-TB drugs alone, EtBr efflux in presence of same anti-TB drugs were considerably reduced under iron deprivation and in ∆011-14 cells (Fig. 3a). Similar results could be observed in MTB depicting that EtBr efflux in presence of same anti-TB drugs were considerably reduced under iron deprivation and in ∆PknG mutant (Fig. 3b). The decrease in EtBr efflux was however variable for each drug and were in the following order INH > RIF > EMB.
EtBr Efflux in presence of anti-TB drugs (INH, RIF, EMB) under iron deprivation. a EtBr efflux displayed by extracellular concentrations of EtBr for M. smegmatis cells grown with INH, RIF, EMB alone (control), presence of 2,2, BP (35 μg/ml) and ∆011-14. Mean of OD285 ± SD of three independent sets of experiments are depicted on y-axis with respect to time (minutes) on x-axis (*depicts significant difference with P < 0.05). b EtBr efflux displayed by extracellular concentrations of EtBr for MTB cells grown with INH, RIF and EMB alone (control), presence of 2,2, BP (40 μg/ml) and ∆PknG. Mean of OD285 ± SD of three independent sets of experiments are depicted on y-axis with respect to time (minutes) on x-axis (*depicts significant difference with P < 0.05)
Iron deprivation affects MmpL3 transporter of RND superfamily
Although mutations in KatG and InhA genes are the principal mechanism for the development of resistance against INH drug, there are reports (Machado et al. 2012; Jaiswal et al. 2017) which suggests that about 20–30% of the INH resistance in MTB isolates do not have mutation in any of the gene associated with INH resistance instead demonstrates efflux pumps as the causal reason of INH resistance (Rodrigues et al. 2012; Garima et al. 2015; Li et al. 2015; Narang et al. 2017; Unissa et al. 2017). The present study showed greater retardation of EtBr efflux under iron deprivation for INH in comparison to other anti-TB drugs. Moreover, resistance against INH is mainly attributed to RND superfamily being its substrate (Pasca et al. 2005; Silva and Palomino 2011; Sarathy et al. 2012). Hence, we studied the effect of iron deprivation on RND superfamily intricately. NR is a specific substrate of RND superfamily of transporters (Bohnert et al. 2010) and strong lipophilic fluorescent dye which is retained in phospholipid layer showing weak fluorescence in aqueous solutions and strong fluorescence in non-polar environment. We estimated the NR accumulation through fluorescence imaging in M. smegmatis and MTB and found that iron deprived cells showed more NR accumulation as compared to control cells (Fig. 4a). These observations indicated that iron deprivation causes some defect in RND superfamily efflux pump activity. Efflux-mediated drug resistance in MTB could be due to one or more efflux pumps working alone or in coordination. This is possible because of the redundancy of their functions, which may overlap extensively (Poole 2005). Hence, the fact that efflux pumps belonging to other superfamilies (MFS, SMR and MATE) are not affected cannot be ruled out and needs further validation.
Effect of iron deprivation on NR accumulation. a Fluorescence microscopy images depicting NR (1 μg/ml) accumulation under different iron deprived conditions. Left panel depict NR accumulation in M. smegmatis whereas right panel depicts NR accumulation in MTB. Images were taken at × 40 magnification. Scale bar depicts 50 μm length. b RT-PCR of differentially regulated MmpL3 gene in response to iron deprivation. Bar graph depicts the quantitation (density expressed as Intensity/mm2) of transcript normalized with 16S constitutively expressed gene. Experiment was replicated to calculate the mean Intensity of gene
MmpL3 (Mycobacterium membrane protein Large 3) is a significant transporter belonging to RND family and is a common target for various small molecules (inhibitors) identified by high throughput screens against MTB. It is crucial for MA (Belardinelli et al. 2016) and flippase that transport trehalose monomycolate (TMM) to the periplasmic space and then to membrane (Xu et al. 2017; Zheng et al. 2018). MmpL3 also contributes to a number of significant mycobacterial phenotypes such as membrane potential, cell wall composition and antibiotic susceptibility (McNeil et al. 2017). In fact MmpL3 is also involved in iron acquisition through heme uptake pathway (Owens et al. 2013). These phenotypic properties of MmpL3 make it an attractive target for TB treatment (Kozikowski et al. 2017; Degiacomi et al. 2017). Hence, we examined the gene expression of MmpL3 in MTB and found that the transcript level of MmpL3 gene was diminished under iron deprivation (Fig. 4b). This confirms that enhanced NR accumulation observed above could be at least due to reduced expression of MmpL3, however, other members of RND transporters being not affected could not be ruled out in the present study. Since MmpL3 is a promising target for novel anti-TB agents, hence vulnerability of MmpL3 expression under iron deprivation needs to be exploited further.
Iron deprivation blocks MmpL3 driven mycolic acid transport
MmpL3 in MTB is crucial inner membrane transporter involved in the export of MA which is required for the maintenance and formation of membrane integrity of all mycobacteria and the possible reason underlying the excellent promiscuity of this target. The defining feature of the mycobacterial outer membrane is the presence of MAs, which in part provides hydrophobicity to the lipid bilayer and impermeability to external insults, including many anti-TB drugs. As a known fact of matter, out of 13 MmpL proteins encoded by MTB genome, only MmpL3 is needed for the transport of TMM to the outer membrane for the formation of trehalose dimycolate (TDM) which forms MA. MA is synthesized inside the cell in the form of TMM and transported by MmpL3 to outer membrane to form the precursor of TDM and mycolates bound to arabinogalactan (Degiacomi et al. 2017). Previously we have showed that iron deprivation alters the membrane permeability and cell surface properties (Pal et al. 2015, 2016). Hence we speculated that MA levels may also be affected under iron deprivation. To confirm this hypothesis, we isolated cell surface lipids and performed TLC which depicted that bands at respective positions corresponding to methoxy-MA were missing under iron deprivation (Fig. 5a). We also estimated the MA levels using carbol-fuchsin stain and found that MA content is decreased under iron deprivation (Fig. 5a). Therefore we adopted lipidomics approach for better assessment of the changes in the cell surface lipids from MTB under iron deprivation. The chromatogram showed m/z peaks for the MA spectra obtained (Figs. 5b, S3–S6). This was then analyzed using MS-LAMP software (Table S2) with window range 0.5 in [M–H]+ mode where we detected 7 m/z values in control and 3 m/z values under iron deprivation (Fig. 5b). These 7 m/z values encoded seven lipid molecules: alpha-MA, Keto-MA, methoxy-MA, GMM (alpha-MA), two GMM (methoxy-MA), and TMM (methoxy-MA) whereas 3 m/z values under iron deprivation encoded three lipid molecules: alpha-MA, Keto-MA and GMM (alpha-MA) (Fig. 5c). Thus, all these observations specify that iron deprivation affects the TMM translocation to the surface of MTB. Furthermore TLC analyses and lipidomic analyses confirm the probability of reduced expression of MmpL3 leading to abrogated transport of TMM and accumulation of TDM to cell surface. This is also commensurate with the fact that MmpL3 has been tipped as TMM flippase which got impaired in presence of MmpL3 inhibitor (Xu et al. 2017).
Effect of iron deprivation on MA transport. a TLC (left panel) showing alteration in MTB MA profile using solvent (hexane/ethyl acetate (19:1, v/v) and MA spectra (right panel) scanned between 400 and 600 nm in the presence and absence of 2,2, BP (40 μg/ml). b Representative mass spectra of control and 2,2, BP treated cell surface lipids (see Supplementary Figs. S1–S6). c Left panel showing bar graph depicting total number of lipid molecules identified from m/z values observed from chromatogram at WR 0.5 through MS-LAMP. Arrows depicts lipid moieties not detected under iron deprivation. Right panel depicts classes and molecular formula of identified lipids
Iron deprivation alters membrane fluidity
The cellular membrane serves as a sensor of its organism’s environment. MTB cell membrane is the major barrier due to its lipid complexity hence a foremost crucial drug target. Its lipid structure is a key to determine the physicochemical environment of the membrane, where the molecular packing of lipids acts as a direct determinant of membrane fluidity. Thus any alteration in fluidity signifies the disturbance in the lipid composition of MTB. Moreover, it is already been established that alteration of membrane fluidity is directly linked to the MA levels (Barkan et al. 2009). Notably, previously we have showed that iron deprivation led to enhanced membrane permeability (Pal et al. 2015). Therefore we estimated the membrane fluidity under iron deprivation by spectrofluoremetric analysis using fluorescent probe DPH. The binding of DPH is based on the environment where it exhibits no fluorescence in aqueous solutions however when adhering to the hydrophobic region of the mycobacterial membrane which gets exposed only during alterations in fluidity, the fluorescence intensity increases (do Canto et al. 2016). We found that membrane fluidity was enhanced under iron deprivation in both M. smegmatis (Fig. 6a) and MTB (Fig. 6b). This could be attributed to diminished MA levels under iron deprivation rendering the cell envelope more vulnerable and fluid. Moreover, disruption of MmpL3 leading to enhance cell envelope permeability (Xiong et al. 2017) corroborates with present observations.
Effect of iron deprivation membrane fluidity. a Line graph of DPH fluorescence intensity scanned between the wavelengths 420–498 nm under iron deprived conditions in M. smegmatis. b Line graph of DPH fluorescence intensity scanned between the wavelengths 420–498 nm under iron deprived conditions in MTB
Conclusion
Drug resistance arising out of efflux pump overexpression has been an issue in clinical settings posing serious problems in the treatment of TB. This study establishes that iron deprivation affects efflux pump activity particularly of RND superfamily. Additionally, iron is indispensable for growth and maintenance of its membrane integrity due to inhibition of MmpL3 expression and subsequent MA transport blockage leading to altered membrane fluidity (Fig. 7). Considering the promiscuity of MmpL3 as target, this study could be helpful to fulfill the unmet demand for development of novel therapeutic strategy. Together, targeting the iron homeostasis in MTB could be a potential approach to combat MDR-TB infections.
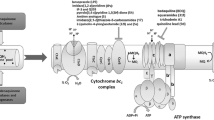
Model depicting effect of MmpL3 blockage under iron deprivation. Under iron sufficient (normal) condition, MmpL3 is functional leading to normal transport of drugs and TMM from cytoplasm to cell surface for mycolation. Right panel depicts iron deprived condition where blockage in MmpL3 leads to abrogated transport of drugs and TMM leading to defective mycolation. Note that the model is not to the scale and does not depict comprehensive structures of all constituents of the MTB cell surface
References
Almeida D, Ioerger T, Tyagi S, Li SY, Mdluli K, Andries K et al (2016) Mutations in pepQ confer low-level resistance to bedaquiline and clofazimine in Mycobacterium tuberculosis. Antimicrob Agents Chemother 60:4590–4599
Balganesh M, Dinesh N, Sharma S, Kuruppath S, Nair AV, Sharma U (2012) Efflux pumps of Mycobacterium tuberculosis play a significant role in antituberculosis activity of potential drug candidates. Antimicrob Agents Chemother 56(5):2643–2651
Banerjee SK, Bhatt K, Rana S, Misra P, Chakraborti PK (1996) Involvement of an efflux system in mediating high level of fluoroquinolone resistance in Mycobacterium smegmatis. Biochem Biophys Res Commun 226:362–368
Barkan D, Liu Z, Sacchettini JC, Glickman MS (2009) Mycolic acid cyclopropanation is essential for viability, drug resistance, and cell wall integrity of Mycobacterium tuberculosis. Chem Biol 16:499–509
Belardinelli JM, Yazidi A, Yang L, Fabre L, Li W, Jacques B et al (2016) Structure-function profile of MmpL3, the essential mycolic acid transporter from Mycobacterium tuberculosis. ACS Infect Dis 2(10):702–713
Bessa LJ, Ferreira M, Gameiro P (2018) Evaluation of membrane fluidity of multidrug-resistant isolates of Escherichia coli and Staphylococcus aureus in presence and absence of antibiotics. J Photochem Photobiol 181:150–156
Bohnert JA, Karamian B, Nikaido H (2010) Optimized Nile Red efflux assay of AcrAB-TolC multidrug efflux system shows competition between substrates. Antimicrob Agents Chemother 54:3770–3775
Caleffi-Ferracioli KR, Amaral RC, Demitto FO, Maltempe FG, Canezin PH, Scodro RB, Nakamura CV, Leite CQ, Siqueira VL, Cardoso RF (2016) Morphological changes and differentially expressed efflux pump genes in Mycobacterium tuberculosis exposed to a rifampicin and verapamil combination. Tuberculosis (Edinb) 97:65–72
Chen C, Gardete S, Jansen RS, Shetty A, Dick T, Rhee KY, Dartois V (2018) Verapamil targets membrane energetics in Mycobacterium tuberculosis. Antimicrob Agents Chemother 62(5):1–15
Chuang YM, Bandyopadhyay N, Rifat D, Rubin H, Bader JS, Karakousis PC (2015) Deficiency of the novel exopolyphosphatase Rv1026/PPX2 leads to metabolic downshift and altered cell wall permeability in Mycobacterium tuberculosis. MBio 6:e02428
Conor PD (2007) Host–pathogen interactions: the role of iron. J Nutr 137:341–1344
Degiacomi G, Benjak A, Madacki J, Boldrin F, Provvedi R, Palù G (2017) Essentiality of mmpL3 and impact of its silencing on Mycobacterium tuberculosis gene expression. Sci Rep 7:43495
do Canto AMTM, Robalo JR, Santos PD, Carvalho AJP, Ramalho JPP, Loura LMS (2016) Diphenylhexatriene membrane probes DPH and TMA-DPH: a comparative molecular dynamics simulation study. Biochim Biophys Acta 1858(11):2647–2661
Garima K, Pathak R, Tandon R, Rathor N, Sinha R, Bose M, Varma-Basil M (2015) Differential expression of efflux pump genes of Mycobacterium tuberculosis in response to varied subinhibitory concentrations of antituberculosis agents. Tuberculosis (Edinb). 95(2):155–161
Gupta S, Cohen KA, Winglee K, Maiga M, Diarra B, Bishai WR (2014) Efflux inhibition with verapamil potentiates bedaquiline in Mycobacterium tuberculosis. Antimicrob Agents Chemother 58:574–576
Hameed S, Pal R, Fatima Z (2015) Iron acquisition mechanisms: promising target against Mycobacterium tuberculosis. Open Microbiol J 9:91–97
Hegeto LA, Caleffi-Ferracioli KR, Nakamura-Vasconcelos SS, Almeida AL, Baldin VP, Nakamura CV, Siqueira VLD, Scodro RBL, Cardoso RF (2018) In vitro combinatory activity of piperine and anti-tuberculosis drugs in Mycobacterium tuberculosis. Tuberculosis (Edinb) 111:35–40
Jaiswal I, Jain A, Verma SK, Singh P, Kant S, Singh M (2017) Effect of efflux pump inhibitors on the susceptibility of Mycobacterium tuberculosis to isoniazid. Lung India 34(6):499–505
Jin J, Zhang JY, Guo N, Sheng H, Li L et al (2010) Farnesol, a potential efflux pump inhibitor in Mycobacterium smegmatis. Molecules 15:7750–7762
Kaniga K, Cirillo DM, Hoffner S, Ismai NA, Kaur D, Lounis N et al (2016) A multilaboratory, multicountry study to determine bedaquiline MIC quality control ranges for phenotypic drug susceptibility testing. J Clin Microbiol 54:2956–2962
Khan MZ, Bhaskar A, Upadhyay S, Kumari P, Rajmani RS et al (2017) Protein kinase G confers survival advantage to Mycobacterium tuberculosis during latency-like conditions. J Biol Chem 292:16093–16108
Khanuja SPS, Srivastava S, Kumar TRS, Shasany AK (2004) A quick and sensitive method of quantifying mycolic acid. US Patent USOO6833249B2
Kozikowski AP, Onajole OK, Stec J, Dupont C, Viljoen A, Richard M et al (2017) Targeting mycolic acid transport by indole-2-carboxamides for the treatment of Mycobacterium abscessus Infections. J Med Chem 60(13):5876–5888
Kurthkoti K, Amin H, Marakalala MJ, Ghanny S, Subbian S, Sakatos A et al (2017) The capacity of Mycobacterium tuberculosis To survive iron starvation might enable it to persist in iron-deprived microenvironments of human granulomas. MBio 8(4):1–17
Layre E, Sweet L, Hong S, Madigan CA, Desjardins D, Young D et al (2011) A comparative lipidomics platform for chemotaxonomic analysis of Mycobacterium tuberculosis. Chem Biol 18:1537–1549
Li G, Zhang J, Guo Q, Jiang Y, Wei J, Zhao LL, Zhao X, Lu J, Wan K (2015) Efflux pump gene expression in multidrug-resistant Mycobacterium tuberculosis clinical isolates. PLoS ONE 10(2):e0119013
Machado D, Couto I, Perdigão J, Rodrigues L, Portugal I (2012) Contribution of efflux to the emergence of isoniazid and multidrug resistance in Mycobacterium tuberculosis. PLoS ONE 7:e34538
McNeil MB, Dennison D, Parish T (2017) Mutations in MmpL3 alter membrane potential, hydrophobicity and antibiotic susceptibility in Mycobacterium smegmatis. Microbiology 163:1065–1670
Mezwa K, McAnaw S, Chiang CY, Trébucq A, Horsburgh CR (2018) Insights into drug-resistant tuberculosis treatment: results of The Union DR-TB Working Group Survey, 2017. Public Health Action 8(3):141–144
Minnikin DE, Alshamaony L, Goodfellow M (1975) Differentiation of Mycobacterium, Nocardia, and related taxa by thin-layer chromatographic analysis of whole-organism methanolysates. J Gen Microbiol 88:194–200
Mullin S, Mani N, Grossman TH (2004) Inhibition of antibiotic efflux in bacteria by the novel multidrug resistance inhibitors biricodar (VX-710) and timcodar (VX-853). Antimicrob Agents Chemother 48:4171–4176
Narang A, Giri A, Gupta S, Garima K, Bose M, Varma-Basil M (2017) Contribution of putative efflux pump genes to isoniazid resistance in clinical isolates of Mycobacterium tuberculosis. Int J Mycobacteriol 6(2):177–183
Odds FC (2003) Synergy, antagonism, and what the chequerboard puts between them. J Antimicrob Chemother 52:1
Ojha A, Hatfull GF (2007) The role of iron in Mycobacterium smegmatis biofilm formation: the exochelin siderophore is essential in limiting iron conditions for biofilm formation but not for planktonic growth. Mol Microbiol 66(2):468–483
Owens CP, Chim N, Graves AB, Harmston CA, Iniguez A, Contreras H et al (2013) The Mycobacterium tuberculosis secreted protein Rv0203 transfers heme to membrane proteins MmpL3 and MmpL11. J Biol Chem 288:21714–21728
Pal R, Fatima Z, Hameed S (2014) Efflux pumps in drug resistance of Mycobacterium tuberculosis: a panoramic view. Int J Curr Microbiol Appl Sci 3:528–546
Pal R, Hameed S, Fatima Z (2015) Iron deprivation affects drug susceptibilities of Mycobacteria targeting membrane integrity. J Pathog 2015:938523
Pal R, Hameed S, Sharma S, Fatima Z (2016) Influence of iron deprivation on virulence traits of mycobacteria. Braz J Infect Dis 20:585–591
Pal R, Hameed S, Kumar P, Singh S, Fatima Z (2017) Comparative lipidomics of drug sensitive and resistant Mycobacterium tuberculosis reveals altered lipid imprints. 3 Biotech 7:325
Pal R, Ansari MA, Saibabu V, Das S, Fatima Z, Hameed S (2018a) Nonphotodynamic roles of methylene blue: display of distinct antimycobacterial and anticandidal mode of actions. J Pathog 2018:3759704
Pal R, Hameed S, Sabareesh V, Kumar P, Singh S, Fatima Z (2018b) Investigations into isoniazid treated Mycobacterium tuberculosis by electrospray mass spectrometry reveals new insights into its lipid composition. J Pathog 2018:1454316
Pasca MR, Guglierame P, De Rossi E, Zara F, Riccardi G (2005) mmpL7 gene of Mycobacterium tuberculosis is responsible for isoniazid efflux in Mycobacterium smegmatis. Antimicrob Agents Chemother 49(11):4775–4777
Patel K, Butala S, Khan T, Suvarna V, Sherje A, Dravyakar B (2018) Mycobacterial siderophore: a review on chemistry and biology of siderophore and its potential as a target for tuberculosis. Eur J Med Chem 157:783–790
Poole K (2005) Efflux-mediated antimicrobial resistance. J Antimicrob Chemother 56:20–51
Rodrigues L, Ramos J, Couto I, Amaral L, Viveiros M (2011) Ethidium bromide transport across Mycobacterium smegmatis cell-wall: correlation with antibiotic resistance. BMC Microbiol 11:35
Rodrigues L, Machado D, Couto I, Amaral L, Viveiros M (2012) Contribution of efflux activity to isoniazid resistance in the Mycobacterium tuberculosis complex. Infect Genet Evol 12(4):695–700
Rodriguez GM, Smith I (2003) Mechanisms of iron regulation in mycobacteria: role in physiology and virulence. Mol Microbiol 47:1485–1494
Sabareesh V, Singh G (2013) Mass spectrometry based lipid(ome) analyzer and molecular platform: a new software to interpret and analyze electrospray and/or matrix-assisted laser desorption/ionization mass spectrometric data of lipids: a case study from Mycobacterium tuberculosis. J Mass Spectrom 48:465–477
Sarathy JP, Dartois V, Lee EJD (2012) The role of transport mechanisms in Mycobacterium tuberculosis drug resistance and tolerance. Pharmaceuticals 5(11):1210–1235
Schmalstieg AM, Srivastava S, Belkaya S, Deshpande D, Meek C, Leff R et al (2012) The antibiotic resistance arrow of time: efflux pump induction is a general first step in the evolution of mycobacterial drug resistance. Antimicrob Agents Chemother 56:4806–4815
Silva PEA, Palomino JC (2011) Molecular basis and mechanisms of drug resistance in Mycobacterium tuberculosis: classical and new drugs. J Antimicrob Chemother 66:1417–1430
Slayden RA, Barry CE III (2001) Analysis of the lipids of Mycobacterium tuberculosis. Methods Mol Med 54:229–245
Sritharan M (2016) Iron homeostasis in Mycobacterium tuberculosis: mechanistic insights into siderophore-mediated iron uptake. J Bacteriol 198:2399–2409
Tanwar J, Das S, Fatima Z, Hameed S (2014) Multidrug resistance: an emerging crisis. Interdiscip Perspect Infect Dis 2014:541340
Unissa AN, Dusthackeer VNA, Kumar MP, Nagarajan P, Sukumar S, Kumari VI, Lakshmi AR, Hanna LE (2017) Variants of katG, inhA and nat genes are not associated with mutations in efflux pump genes (mmpL3 and mmpL7) in isoniazid-resistant clinical isolates of Mycobacterium tuberculosis from India. Tuberculosis (Edinb) 107:144–148
WHO (2016) World Health Organization Report on Global Tuberculosis 2016. 322 Reference Number: WHO/HTM/TB/201613
Xiong LB, Liu HH, Xu LQ, Sun WJ, Wang FO, Wei DZ (2017) Improving the production of 22-hydroxy-23,24-bisnorchol-4-ene-3-one from sterols in Mycobacterium neoaurum by increasing cell permeability and modifying multiple genes. Microb Cell Fact 16:89
Xu Z, Meshcheryakov VA, Poce G, Chng SS (2017) MmpL3 is the flippase for mycolic acids in mycobacteria. Proc Natl Acad Sci USA 114:7993–7998
Zheng H, Williams JT, Coulson GB, Haiderer ER, Abramovitch RB (2018) HC2091 Kills Mycobacterium tuberculosis by targeting the MmpL3 mycolic acid transporter. Antimicrob Agents Chemother 62(7):1–34
Acknowledgements
Z.F. thanks from Board of Research in Nuclear Sciences (BRNS), Mumbai (2013/37B/45/BRNS/1903) for the financial assistance. We are grateful to Mandira Varma-Basil, Graham F. Hatfull, Sarman Singh and Yossef Av-Gay for providing MTB (H37Rv), siderophore mutant ∆011-14, M. smegmatis mc2155 and ∆PknG mutant as generous gifts respectively. We thank Sanjeev Kanojiya, Central Drug Research Institute (CDRI), Lucknow for assisting us in mass spectrometry experiments. We thank Varatharajan Sabareesh for his intellectual support in mass spectrometry data analysis.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
10534_2018_157_MOESM2_ESM.xls
Supplementary material 2 Spread sheets obtained from MS-LAMP software for Control (untreated) and 2,2, BP treated MTB cell surface lipids (XLS 31 kb)
Rights and permissions
About this article
Cite this article
Pal, R., Hameed, S. & Fatima, Z. Altered drug efflux under iron deprivation unveils abrogated MmpL3 driven mycolic acid transport and fluidity in mycobacteria. Biometals 32, 49–63 (2019). https://doi.org/10.1007/s10534-018-0157-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10534-018-0157-8