Abstract
Textile industries release major fraction of dyestuffs in effluents leading to a major environmental concern. These effluents often contain more than one dyestuff, which complicates dye degradation. In this study ten reactive dyes (Reactive Yellow 145, Reactive Yellow 160, Reactive Orange 16, Reactive Orange 107, Reactive Red 195, Reactive Blue 21, Reactive Blue 198, Reactive Blue 221, Reactive Blue 250, and Reactive Black 5) that are used in textile industries were subjected to biodegradation by a bacterial consortium VITPBC6, formulated in our previous study. Consortium VITPBC6 caused single dye degradation of all the mentioned dyes except for Reactive Yellow 160. Further, VITPBC6 efficiently degraded a five-dye mixture (Reactive Red 195, Reactive Orange 16, Reactive Black 5, Reactive Blue 221, and Reactive Blue 250). Kinetic studies revealed that the five-dye mixture was decolorized by VITPBC6 following zero order reaction kinetic; Vmax and Km values of the enzyme catalyzed five-dye decolorization were 128.88 mg L−1 day−1 and 1003.226 mg L−1 respectively. VITPBC6 degraded the dye mixture into delta-3,4,5,6-Tetrachlorocyclohexene, sulfuric acid, 1,2-dichloroethane, and hydroxyphenoxyethylaminohydroxypropanol. Phytotoxicity, cytogenotoxicity, microtoxicity, and biotoxicity assays conducted with the biodegraded metabolites revealed that VITPBC6 lowered the toxicity of five-dye mixture significantly after biodegradation.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Textile industries significantly contribute in the global economic development. Textile exportation is the highest in China trailed by the European Union, India, and the USA. The worldwide textile market was valued at USD 993.6 billion in 2021, and a compound annual growth rate of 4% is expected during 2022–2030 (Grand View Research 2022). The market of textile application is majorly led by fashion sector, followed by technical sector, and household sector (Ul-Islam 2017). Textile industries on one hand contributed to global economic progression, however on the other hand it caused major hazardous impacts on the environment. The industry generates huge quantity of pollutants during the wet processing stages of textile manufacturing (Uddin 2019). It majorly causes water pollution as well as soil and air pollution. Textile industries consume huge volumes of water during the manufacturing process, consequently, huge volumes of effluent are released by these industries. The effluents contain at least 72 toxic chemicals and 30 of these chemicals are difficult to treat with the conventional treatment processes. Textile industries are the largest consumer of dyes (∼ 70%). The dye market is predominated by reactive dyes and disperse dyes (44% of the market value) (Shukla et al. 2021). Reactive dyes are made up of azo or anthraquinone group as chromophores and various types of reactive groups (Asgher 2012). The discharge of wastewater containing dyestuffs occurs primarily due to the improper intake of dyes and the extent of dye fixation onto substrates. Dyes are noticeable in the effluents even in a minute quantity of 1 ppm due to its significant tinctorial property (Singh and Arora 2011). The release of untreated or partially treated wastewater is detrimental to both the biotic and abiotic factors of an ecosystem. A major concern is that the conventional effluent treatment processes are high energy consuming, time consuming, lacking a good infrastructure, and costing high. Moreover, in economically weaker countries wastewater treatment plants remain mostly non-functional due to reluctance and lack of profit gain. Diseases, such as contact dermatitis, allergic conjunctivitis, rhinitis, occupational asthma, and other allergic reactions have been documented, during work-related exposure to reactive dyes. A clinical and immunological investigation stated that 15% out of 400 workers subjected to reactive dye exposure suffered from respiratory issues (Docker et al. 1987). Frequent cases of colon and bladder cancer have been reported from reactive dye exposure (Senthil Kumar et al. 2020). Reactive dyes can bind to proteins found inside the human body, and thus pose a major health hazard (Al-Tohamy et al. 2022).
The biological treatment methods utilizing filamentous fungi, bacteria, actinomycetes, algae, and yeasts for dye degradation or mineralization is an excellent alternative for reducing dye pollution. Biological treatment methods have exponentially grown over the last two decades. The pros of using this treatment method are: (1) cheap, (2) production of low amount of sludge, (3) no secondary pollution, (4) low water requirement, (5) environment friendly (Saratale et al. 2011, 2013). Many studies have reported efficient dye degradation by various bacterial species. Dye biodegradation occurs with the help of oxidoreductive enzymatic system present in bacteria, such as azoreductase, nicotinamide adenine dinucleotide-2,6-dichlorophenolindophenol (NADH-DCIP) reductase, laccase, tyrosinase, lignin peroxidase, manganese peroxidase, and dye decolorizing peroxidase (Mishra et al. 2022). It has been documented in a significant number of studies that bacterial consortium is better able to degrade dyes than that of single bacterial species (pure culture). The metabolic diversity in a bacterial consortium allows them to degrade dyes more efficiently than that of pure cultures (Stolz 2001; Shade et al. 2012). Afrin et al. (2021) constructed a consortium of Pseudomonas aeruginosa and Enterococcus faecium that exhibited a higher rate of dye degradation more rapidly compared to that of the individual strains. Similarly, Mohanty and Kumar (2021) built up a consortium composed of Bacillus flexus, Proteus mirabilis, and P. aeruginosa. The consortium showed a higher average dye decolorization rate, reduction in TOC and COD removal, and induction of various oxidoreductase enzymes compared to that of the individual strains.
Over the last decade, bioremediation studies were carried out majorly with single dyes. Hence, the significance of the study relies on the use of dye mixtures, since real time effluents contains a mixture of different dyes. In the present study, consortium VITPBC6, a previously constituted bacterial consortium in our dye decolorization study, will be used to treat dye mixtures (Saha et al. 2022). The end products of biodegradation will be determined with the help of analytical techniques. The reaction kinetics involved during dye degradation will be studied. Lastly, different toxicity studies will be performed to determine any toxicity that might have arisen after dye biodegradation.
Materials and methods
Reactive dyes and their mixtures utilized
Different reactive dyes were obtained from a textile industry in Mysuru, Karnataka. Ten commercial dyes were used in this study including C.I. Reactive Yellow 145 (RY-145), C.I. Reactive Yellow 160 (RY-160), C.I. Reactive Orange 16 (RO-16), C.I. Reactive Orange 107 (RO-107), C.I. Reactive Red 195 (RR-195), C.I. Reactive Blue 21 (RB-21), C.I. Reactive Blue 198 (RB-198), C.I. Reactive Blue 221 (RB-221), C.I. Reactive Blue 250 (RB-250), and C.I. Reactive Black 5 (RB-5). Two dye mixtures were prepared from the above-mentioned dyes, including a ten-dye mixture (composed of RY-145, RY-160, RO-16, RO-107, RR-195, RB-21, RB-198, RB-221, RB-250, and RB-5) and a five-dye mixture (composed of RR-195, RO-16, RB-5, RB-221, and RB-250). Initially, dye standard solutions were prepared at 20 mg L−1. Necessary dilutions were made from the standard solution for all the future studies. Maximum absorption (λmax) of each of the dyes was determined using UV–Vis spectrophotometer (Shimadzu, UV-1280) (200–700 nm). The λmax of the dye mixtures were determined using the colorimetric method as illustrated by APHA (APHA 2005).
Building up of consortium VITPBC6
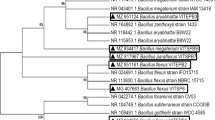
Consortium VITPBC6 was built up with industrial effluent-adapted bacterial isolates (textile industry and leather industry) in equal proportions (1:1:1:1:1:1), as discussed in our previous study (Saha et al. 2022). Bacterial isolates were obtained from the effluents using an enrichment technique method. Decolorization assay was then performed to identify the most potent isolates. Finally, compatibility of the isolates was determined in Mueller Hinton agar. The isolates thus selected to build up the consortium were identified as Bacillus firmus VITEPB1 (NCBI accession number: MZ934656), Bacillus flexus VITEPB2 (NCBI accession number: MZ951161), Bacillus aryabhattai VITEPB3 (NCBI accession number: MZ951124), B. flexus VITSP6 (NCBI accession number: MG407663), Bacillus paraflexus VITSPB7 (NCBI accession number: MZ817967), and Bacillus megaterium VITSPB9 (NCBI accession number: MZ934417).
Decolorization property of consortium VITPBC6 for different reactive dyes and their mixtures
Decolorization assay was performed in Luria Bertani (LB) broth under optimized conditions (40 °C, pH 7.38, 0.61% arabinose, 0.55% peptone, 2.5% Na2SO4, and 7.5% inoculum). The optimized conditions were determined by D-optimal design in response surface methodology, in our previous study (Saha et al. 2022). All the reactive dyes and two dye-mixtures (ten-dye mixture and five-dye mixture) were examined for decolorization. The individual dyes and dye mixtures were decolorized at an initial concentration of 50 and 30 mg L−1 respectively. Initial and final absorbance of test solutions were measured at λmax, and decolorization percentage was calculated using the following formula:
Maximum dye mixture tolerance assay by consortium VITPBC6
Initial concentration of the ten- and five-dye mixture was changed from 0 to 180 mg L−1 and 0 to 300 mg L−1 respectively. Experimentation was done in LB broth supplemented with different dye concentrations under the optimized physico–chemical parameters. The inoculated flasks were incubated (Thermo Scientific Heratherm Advanced Protocol Microbiological Incubator) for 6 days at 37 °C until decolorization occurred. After incubation, decolorization percentage was calculated according to Eq. 1.
Reaction and enzyme kinetics of five-dye mixture decolorization by consortium VITPBC6
Kinetics of the five-dye mixture decolorization process by consortium VITPBC6 was studied in LB broth under optimized physico-chemical parameters. LB broth was supplemented with different initial concentrations (C0 in mg L−1) of the five-dye mixture spanning from 0 to 300 mg L−1. Decolorization reaction was observed for 144 h. Absorbance changes were monitored at λmax, after every 6 h. Reaction orders are represented in Eqs. 2–4 (Atkins et al. 2006). Graphs were then plotted as residual dye concentration (Ct in mg L−1) vs. time (t) for the different reaction orders to determine the best fit. Zero, first, and second order graphs were plotted as Ct vs. t, ln Ct vs. t, and 1/Ct vs. t respectively.
Zero order:
First order:
Second order:
Michaelis constant (km) and maximum substrate consumption rate (Vmax) were determined using the Lineweaver–Burk double reciprocal plot. The plot has been represented in Eq. 5 (Berg et al. 2002). Reaction rate at different dye mixture initial concentration was calculated to plot the graph. The Km and Vmax values were then obtained from the plot.
Lineweaver–Burk equation:
where, V is the rate of substrate concentration (mg L−1 h−1), and S is substrate concentration (mg L−1).
Analysis of the five-dye mixture decolorization metabolites
UV–Vis spectrometric analysis
The five-dye mixture (30 mg L−1) was subjected to decolorization by consortium VITPBC6 in LB broth under the optimized physico-chemical parameters. Absorbance of the culture flask was recorded before and after decolorization. Cells were removed from the culture flasks by centrifuging (REMI C-24 Plus) the broth at 15,000×g for 15 min. The resulting supernatant was scanned in an UV–Vis spectrophotometer (200–700 nm). Dye degradation or transformation is indicated by a shift in the λmax.
Extraction of five-dye mixture decolorization metabolites
Decolorization of the five-dye mixture was conducted in large volumes of 1.5 L. The experiment was conducted in LB broth supplemented with the five-dye mixture (120 mg L−1) under the optimized physico–chemical parameters. The flasks were incubated till complete decolorization occurred (37 °C for 6 days). Then, completely decolorized broths were centrifuged at 12,000×g for 15 min to remove cellular debris. The supernatant was passed through 0.22 μm CA filter and collected. Decolorization metabolites present in the supernatant were collected using the liquid-liquid extraction method. Acetonitrile was used as the solvent of choice for metabolite extraction. The feed solution and acetonitrile were poured in equal volumes in a separating funnel. Separation was carried out for 30 min. The acetonitrile extracts were collected and dried over anhydrous Na2SO4. The organic extract was completely dehydrated in powder form using a rotary evaporator (BUCHI, Rotavapor R-300) and utilized for further analysis (Dawkar et al. 2008).
Fourier transform infrared spectroscopic (FTIR) analysis
The five-dye mixture and its corresponding degraded metabolite samples were analyzed through FTIR (Thermo Nicolet AVATAR 330) to determine any variations of molecular structure between the two samples. The analysis was conducted using the KBr disc method. Alterations in transmittance percentage at different wavenumbers (400–4000 cm−1) were recorded (scan speed 16).
Ultra-high performance liquid chromatography (UHPLC) analysis
The five-dye mixture and its corresponding degraded metabolite samples were separated using reverse phase UHPLC (Waters, ACQUITY H class) analysis. The mobile phase was a blend of HPLC grade carbinol and water (1:1). A C18 reverse column was used for achieving sample separation, and the corresponding separated compounds were detected by a photo diode array (PDA) detector. Sample flow was 0.8 mL min−1. Spectra were developed within the wavelength range of 200–700 nm. The separation was conducted for 20 min. The chromatograms of dyes and its corresponding degraded dye metabolite were compared.
Liquid chromatography high resolution mass spectrometry–quadrupole time-of-flight (LCHRMS–QTOF)
The five-dye mixture degraded metabolite sample was analyzed through LCHRMS–QTOF (Agilent technologies, 1290 Infinity UHPLC System, TOF 6500 series) to identify the compounds present in the sample. Samples were separated in a HYPERSIL Gold C18 column (100 × 2.1 mm, 3 μm). Samples were prepared by dissolving these in deionized water (1 µg µL−1). Water was used as the mobile phase, which was flown at 0.2 mL min−1 for 30 min. Electrospray ionization (ESI) was operated in both positive and negative mode to record the mass spectra under nitrogen (N2) gas flow (1 mL min−1). Fragment voltage was maintained at 175 V.
Toxicity analysis
The toxic nature of the five-dye mixture and its degraded metabolites were studied on different biological organisms, such as plants, microorganisms, and animals.
Phytotoxicity assay
The seeds of Vigna radiata and Cicer arietinum and the roots of the bulbs of Allium cepa were chosen for analysis.
The seeds were used for germination assay. These were grouped according to the treatment they received. The treatments included untreated five-dye mixture solution, treated dye degraded metabolites solution, and control (distilled water). The treatments were used in two concentrations (500 and 1500 ppm). The seeds were germinated in sterilized Petri plates covered with filter paper. Ten seeds were placed in each Petri plate. The treatment volume used for the seeds was 10 mL on the first day and 5 mL for the rest of the days. The seeds were treated for 7 days. V. radiata seeds were grown at 35 °C and C. arietinum seeds were grown at 15 °C (Rao and Prasad 2014; Mondal and Debnath 2022). Seed growth was documented every day. Relative seed germination (RSG) (Eq. 6), germination index (GI) (Eq. 7), and seedling characteristics were calculated at the end of the treatment period.
where, Ms represents the number of germinated seeds under treatment, Ds represents the number of germinated seeds under control treatment, Rs represents the average root length of the seeds under treatment, and Rc represents the average root length of the seeds under control treatment.
Allium cepa bulbs were used for root growth assay. The bulbs were grown under hydroponic conditions. Like the germination assay, the bulbs were distributed into groups based on the treatments (untreated five-dye mixture solution, treated dye degraded metabolites solution, and control (distilled water) that they received. Treatments were used in two concentrations (500 and 1500 ppm). The bulbs were incubated for 7 days at 23 °C (Haq et al. 2016; Alaguprathana et al. 2022). Root growth was documented regularly. At the end of the incubation period, the number of roots and mean root length of the five longest roots in each of the bulbs were recorded.
Cytogenotoxicity assay
Onion root bulbs grown during the phytotoxicity assay were further examined for cytotoxic and genotoxic effects. The roots received four different treatments, including untreated five-dye mixture, treated dye degraded metabolites, distilled water (negative control), and H2O2 (positive control). The root-tips (2–2.5 cm) were cut off and stained by acetocarmine using the root-tip squash technique (Saha and Rao 2020). Meristematic root cells were then observed under light microscope (Magnus microscopes, model MLX). Mitotic index was calculated per 1000 cells scored (Eq. 8) and aberration index was also recorded (Eq. 9).
Where, MI is mitotic index, d is total dividing cells, c is total analyzed cells, AI is aberration index, and a is total chromosomal aberrations identified
Microtoxicity assay
The growth curves of Escherichia coli, Pseudomonas aeruginosa, and Bacillus cereus were used to reveal the toxicity of the untreated five-dye mixture and its corresponding degraded dye metabolite. The bacteria received three different treatments, including the five-dye mixture, degraded dye mixture metabolites, and without any treatment (control). The bacteria were inoculated (2%, v/v) in nutrient broth (NB). The treatments (200 ppm) were added in NB. The incubation of E. coli and P. aeruginosa was done at 37 ℃ and B. cereus at 30 ℃ for 8 h. The optical density (OD) of the bacterial culture broths (five-dye mixture treated broth at 560 nm, degraded dye metabolites treated broth, and control broth at 600 nm) were measured in a colorimeter (Systronics digital colorimeter 112) at an interval of 1 h (Assess et al. 2018).
Biotoxicity assay
Lethality assay was executed with Artemia salina (Rima et al. 2022). At first, Artemia cysts (4 g) were hatched in 1 L of sea water at 25 ℃ under continuous illumination and aeration conditions for 84 h. The nauplii were grown till they reached the instar II and III stages. Approximately 500 shrimps were transferred in 50 mL of sea water contained in a glass beaker (100 mL). The shrimps received four different treatments, including the untreated five-dye mixture, degraded dye mixture metabolites, K2Cr2O7 (positive control), and no treatment (negative control). The treatments were used at 0.125% and 0.5% concentration. All the treated brine shrimps were kept at 25 ℃ under continuous illumination and aeration for 24 h. Finally, the number of surviving shrimps under each treatment were counted to calculate survival percentage (Eq. 10). They were also observed under light microscope to spot any physiological alteration that might have occurred as a response to toxicity.
where, S indicates survival percentage, s indicates the number of survivors, and t indicates the total number of shrimps.
Statistical analysis
All the experiments were conducted in triplicates. Data in this research has been expressed as mean ± standard deviation, as per requirement. The statistical significance of the experimental data has been established by one way analysis of variance (ANOVA), two-way ANOVA, and coefficient of determination (R2) as per experimental requisite. The significances were determined based on probability (P-value). Analysis results with P-value less than or equal to 0.05 were considered significant.
Results and discussion
Details of reactive dyes and their mixtures
Table S1 contains the chemical structure and wavelength of maximum absorption (λmax) of the reactive dyes utilized for this study. The λmax of the ten-dye and five-dye mixtures were found to be 540 and 560 nm respectively.
Decolorization property of consortium VITPBC6 for different reactive dyes and their mixtures
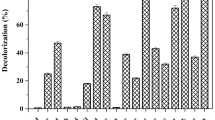
Consortium VITPBC6 efficiently decolorized RO-16 (89.29%), RB-5 (76.8%), RB-250 (76.32%), and RB-221 (76.01%). Moderate decolorization occurred with dyes RR-195 (63.01%) and RB-21 (44.9%). On the other hand, low decolorization occurred with dyes RY-145, RO-107, and RB-198; whereas no decolorization occurred with RY-160. Several factors play a role in determining dye degradation. High molecular weight and structural complexity hinder dye degradation. Other factors, such as substituent groups, azo bond number, electron withdrawing and releasing groups, and interaction of hydrogen between azo and hydroxyl group significantly affect dye degradation (Zhuang et al. 2020; Shi et al. 2021). Further, consortium VITPBC6 could decolorize dye-mixtures. Consortium VITPBC6 decolorized the five-dye mixture (72.98%) more effectively than the ten-dye mixture (48.85%) (Fig. 1). Five dye mixture was developed and used in the study since these five dyes (RR-195, RO-16, RB-5, RB-221, and RB-250) showed maximum decolorization potential among the other ten dyes that were decolorized individually. Several researchers have tried to use bacterial consortium for the decolorization of different dyes. Lade et al. (2015) reported a bacterial consortium composed of Providencia rettgeri and Pseudomonas sp. that efficiently decolorized RB 5, RO 16, Disperse Red 78, and Direct Red 81 (DR 81). Similarly, Guo et al. (2019) constructed a bacterial consortium composed of Pseudomonas, Lysinibacillus, Lactococcus, and Dysgonomonas that degraded multiple azo dyes, such as metanil yellow, direct fast black G, and acid brilliant scarlet GR. Another bacterial consortium composed of Proteus mirabilis, Morganella morganii, and Enterobacter cloacae decolorized four dyes Yellow EXF, Red EXF, Blue EXF, Black WNN, a mixture of these dyes, and a real textile industry effluent (Madhushika et al. 2019). Textile effluents contain a myriad of chemicals including a mixture of dyes. Hence, it is crucial to formulate a treatment capable of degrading dye mixtures. In the present study, VITPBC6 was able to decolorize as many as nine dyes and their mixtures as well. The decolorization potential of consortium VITPBC6 was thus higher than that of several previous reports.
Maximum dye-mixture tolerance assay by Consortium VITPBC6
The influence of dye mixture initial concentration on the decolorization potential of consortium VITPBC6 was investigated (Fig. 2). Decolorization of the ten-dye mixture was more hindered than the five-dye mixture by consortium VITPBC6. The highest decolorization (88.46%) of the five-dye mixture was observed at 120 mg L−1 and moderate decolorization was observed till 150 mg L−1. However, at the highest concentration of 300 mg L−1, decolorization dropped to 30.17%. On the other hand, the ten-dye mixture decolorization occurred maximally (61.44%) at 90 mg L−1. The decolorization was drastically reduced to 19.71% at the highest concentration of 180 mg L−1. The composition of textile industry effluent includes a concoction of dyes, metals, and other pollutants (Yaseen and Scholz 2019). However, the early dye degradation studies majorly concentrated on the study of single dyes (He et al. 2004; Asad et al. 2007; Ayed et al. 2010; Prasad and Rao 2013). Currently, several researchers have thus focused their studies on dye mixture remediation. The extent of dye decolorization by different consortia are as follows: 96.3% (100 mg L−1 of five dye mixture), 83% (50 mg L−1 of four dye mixture), 82% (1200 mg L−1 of three dye mixture), 75.2% (1000 mg L−1 of three dye mixture), 75% (100 mg L−1 of five dye mixture (Karim et al. 2018; Madhushika et al. 2019; Afrin et al. 2021; Khan et al. 2021; Thiruppathi et al. 2021; Biju et al. 2022). In this study, consortium VITPBC6 showed higher dye decolorization potential for the five-dye mixture at a high concentration compared to that of the previous studies. Dye toxicity is known to escalate on increasing azo bond number, substitutions of the aromatic rings, and dye concentration. This could be the probable reason for decreased decolorization of the ten-dye mixture.
Dye mixture (Ten-dye mixture composition: C.I. Reactive Yellow, C.I. Reactive Yellow 160, C.I. Reactive Orange 16, C.I. Reactive Orange 107, C.I. Reactive Red 195, C.I. Reactive Blue 21, C.I. Reactive Blue 198, C.I. Reactive Blue 221, C.I. Reactive Blue 250, and C.I. Reactive Black 5; Five-dye mixture composition: C.I. Reactive Red 195, C.I. Reactive Orange 16, C.I. Reactive Black 5, C.I. Reactive Blue 221, C.I. Reactive Blue 250) tolerance assay with bacterial consortium VITPBC6 under optimized physico–chemical conditions (40 °C, pH 7.38, 0.61% arabinose, 0.55% peptone, 2.5% Na2SO4, and 7.5% inoculum)
Reaction and enzyme kinetics of five-dye mixture decolorization by consortium VITPBC6
The initial five-dye mixture concentration effect on the decolorization potential of consortium VITPBC6 has been tabulated (Table S2). The table consists of the Ct values recorded at different time gaps for the experimental C0 values. The five-dye mixture batch decolorization exhibited somewhat linear and gradual decline in the Ct values during 144 h for the given C0 values. The corresponding reaction order graphs derived from the batch decolorization test data has been presented in Fig. 3. The graphs of zero and first order model developed a more linear relationship than that of the second order kinetic model. Further, the reaction rate constants of all the three kinetic models at the specified C0 values and the corresponding R2 values were calculated (Table S3). The zero order model had R2 values in the range of 0.9314–9857 and was closer to unity than that of the first and second order models. Thus, it can be concluded that the consortium decolorized the five-dye mixture following zero order reaction kinetics. Zero order reaction was mostly followed during anaerobic dye decolorization or when substrate concentration was increased. Mordant Yellow 10 reduction by anaerobic granular sludge followed zero order kinetics (Field and Brady 2003; dos Santos et al. 2004). Sarioglu et al. (2007) investigated Basic Red 46 decolorization by an anaerobic mixed culture. The dye decolorization obeyed first order kinetics at the concentration range of 50–250 mg L−1, whereas zero order kinetics was followed at the concentration range of 500–1000 mg L−1. Zahran et al. (2019) conducted a study with azoreductase producing microbes from the gut microbiota to reduce food colorants. Azoreductase activity of Escherichia coli (brilliant black, sunset yellow, and tartrazine), Enterococcus faecalis (amaranth, brilliant black, and tartrazine), Enterococcus avium (amaranth, brilliant black, sunset yellow, and tartrazine), and Bacillus cereus (amaranth and brilliant black) followed zero order reaction kinetics for dye decolorization. However, interestingly azoreductase activity from B. cereus followed first order kinetics at low dye concentration (10 μm), but zero order kinetics at high dye concentration (20 and 30 μm) during the decolorization of sunset yellow and tartrazine. Similarly, in our previous study RO-16 decolorization by consortium VITPBC6 exhibited a zero order reaction (Saha et al. 2022). The results of this study are concomitant with the above-mentioned reports. The zero order reaction of the dye mixture decolorization might have stemmed from the high substrate (dye) concentration chosen for this study.
The decolorization rate with respect to different initial dye concentrations has been calculated and presented in Table S4. The corresponding data was then applied to the Lineweaver–Burk equation. A graph (1/V vs. 1/S) was plotted to verify the applicability of the equation. The double reciprocal plot for the five-dye mixture decolorization has been presented in Fig. 4. The Lineweaver–Burk model equation was a good fit (R2 value of 0.9765) for the obtained experimental data of the five-dye mixture decolorization. The interception (1/Vmax) and slope (Km/Vmax) of the graph was used to obtain Vmax (128.88 mg L−1 day−1) and Km values (1003.226 mg L−1) of the enzymatic reaction involved in the five-dye mixture decolorization. Enzyme kinetic studies have been majorly conducted with purified enzymes for single dye degradation (Wanyonyi et al. 2017; Iark et al. 2019; Edoamodu et al. 2021; Zhao et al. 2021). To the best of our understanding, the enzyme kinetic studies for dye-mixture decolorization by bacterial consortium have not been reported so far. Both the Vmax and Km values were high in this study. A high Km value points to a low enzyme affinity towards its substrate. The high Km value might be linked to the very high initial substrate concentrations coupled to lower volume of enzymes produced by VITPBC6 that were used in this study.
Reaction order kinetic models of five-dye mixture (C.I. Reactive Red 195, C.I. Reactive Orange 16, C.I. Reactive Black 5, C.I. Reactive Blue 221, C.I. Reactive Blue 250) decolorization at different initial dye concentrations a Zero order reaction was plotted as residual dye concentration (Ct) vs. t b First order reaction was plotted as ln Ct vs. t c Second order reaction was plotted as 1/Ct vs. t
Lineweaver–Burk double reciprocal plot of five-dye mixture (C.I. Reactive Red 195, C.I. Reactive Orange 16, C.I. Reactive Black 5, C.I. Reactive Blue 221, C.I. Reactive Blue 250) decolorization kinetics. The intercept (1/Vmax) and slope (Km/Vmax) of the straight-line equation (y = mx + c) was used to determine the Km and Vmax values.
Analysis of the five-dye mixture decolorization metabolites
UV–Vis spectrometric analysis
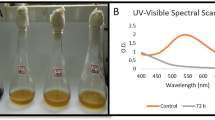
The five-dye mixture and its biodegraded metabolite spectra are illustrated in Fig. 5. The spectrum of the five-dye mixture exhibited a broad peak due to the presence of multiple chromophores with a maximum absorbance at 560 nm. However, the maximum absorbance peak in the spectrum of degraded dye mixture metabolite occurred at 285 nm. Additionally, the peak intensity in the degraded metabolite spectrum was also reduced significantly. Therefore, it can be presumed that the five-dye mixture decolorization by consortium VITPBC6 might have arisen from a cleavage or transformation of the chromophores i.e., biodegradation.
FTIR analysis
The FT-IR spectra of the five-dye mixture and its corresponding degraded metabolite have been illustrated in Fig. 6. A broad peak at 3417.86 cm−1 represents –OH group. The signature –N=N– group peaked at 1589.34 cm−1. Two peaks of primary –NH bend and tertiary –NH bend were noted at 1492.90 and 1176.58 cm−1 respectively. SO3 group appeared at 1128.36 cm−1. In the fingerprint region, aromatic –CH out of bend was noted at 896.90 cm−1 and 731.02 cm−1. On the other hand, the FTIR spectrum of the degraded five-dye mixture metabolite revealed fewer peaks compared to that of the dye mixture spectrum. The spectrum exhibited –OH group at 3305.99 cm−1, C–H deformation vibration in halogenated compound at 1398.39 cm−1, and –SO4 stretching vibration at 1093.64 cm−1. In the fingerprint region, skeletal vibration in alkene was observed at 609.51 cm−1. The spectrum represented the presence of functional groups corresponding to aliphatic compounds, and the absence of the functional group corresponding to the dye chromophore. Hence, it can be presumed that the dye mixture underwent biodegradation.
UHPLC analysis
The UHPLC elution profiles of the five-dye mixture and its corresponding degraded metabolite have been depicted in Fig. 7. The five-dye mixture chromatogram exhibited the presence of six sharp peaks at retention times 1.2 min, 1.3 min, 1.8 min, 4.9 min, 5.5 min, and 6.3 min respectively. The multiple peaks occurred owing to the presence of multiple dyes in the mixture. On the other hand, the degraded dye mixture metabolite chromatogram exhibited the appearance of three new peaks at 1.096 min, 1.225 min, and 2.254 min respectively. The occurrence of new peaks in the degraded dye metabolite chromatogram points towards the presence of compounds different from the untreated dye mixture compounds. Additionally, fewer number of peaks formed in the degraded dye mixture metabolite chromatogram than that of the dye chromatogram might suggest that the dye mixture was degraded into simple compounds.
LCHRMS–QTOF
The chromatograms and mass spectra of the five-dye mixture degradation by consortium VITPBC6 have been illustrated in Fig. 8a–f. The chromatogram obtained, when ESI was operated in negative mode has been presented in Fig. 8a, and the chromatogram obtained when ESI was operated in positive mode has been presented in Fig. 8b . The mass spectra revealed that the dye mixture was degraded into simple compounds, viz., delta-3,4,5,6-Tetrachlorocyclohexene (mass 217.919), sulfuric acid (mass 97.9682), 1,2-dichloroethane (mass 97.9682), and hydroxyphenoxyethylaminohydroxypropanol (mass 227.1153). End-products delta-3,4,5,6-Tetrachlorocyclohexene, sulfuric acid, and 1,2-dichloroethane were obtained when ESI was operated in negative mode. delta-3,4,5,6-Tetrachlorocyclohexene appeared at a retention time of 1.163 min with ion peak at m/z 2116.9118 (Fig. 8c). Sulfuric acid appeared at a retention time of 1.167 min with ion peak at m/z 96.9609 (Fig. 8d). 1,2-dichloroethane appeared at a retention time of 1.514 min with ion peak at m/z 96.9609 (Fig. 8e). End-product hydroxyphenoxyethylaminohydroxypropanol was obtained when ESI was operated in positive mode. It had a retention time of 4.176 min and ion peak at m/z 210.1121 (Fig. 8f). It is notable that the complex dye mixture was biodegraded by consortium VITPBC6, which makes it a highly potential agent for reactive dye biodegradation.
Liquid chromatography-mass spectrophotometry analysis of biodegradation products of five-dye mixture (C.I. Reactive Red 195, C.I. Reactive Orange 16, C.I. Reactive Black 5, C.I. Reactive Blue 221, C.I. Reactive Blue 250) by consortium VITPBC6 a chromatogram (Electrospray ionization operated in negative mode) b chromatogram (Electrospray ionization operated in positive mode) c mass spectrum of delta-3,4,5,6-Tetrachlorocyclohexene, d mass spectrum of sulfuric acid, e Mass spectrum of 1,2- dichloroethane, f mass spectrum of hydroxyphenoxyethylaminohydroxypropanol
Toxicity assays
Phytotoxicity assay
Seed germination is a period when seeds are sensitive and highly susceptible to toxicity. Hence, after treating the seeds with distilled water, dyes, and degraded dye metabolites RSG, GI, root length, and shoot length were noted down (Table 1a, b). In case of V. mungo, the five-dye mixture treated seeds had lower RSG, GI, root and shoot length compared to that of the corresponding degraded dye mixture metabolite treated seeds. The root and shoot length of the degraded dye mixture metabolite treated seeds and control seeds were comparable. Statistical analysis indicated that the effect of treatment variation on the seeds were significantly different from each other (P-value 0.001 < 0.05, Fstatistics > Fcritical). However, concentration variation had insignificant effect on the seeds (P-value 0.322 > 0.05, Fstatistics < Fcritical) (Table 1a). In case of C. arietinum, RSG, GI, radicle and plumule length were lower in the seeds treated with the five-dye mixture than the seeds treated with the corresponding degraded dye metabolite. Therefore, it might be presumed that the degraded dye metabolite had significantly lower toxicity than that of the dye mixture. The data obtained with treatment variation on the seeds were statistically significant with P-value 0.01 < 0.05 and Fstatistical > Fcritical. However, like that observed in the P. mungo seeds, the concentration variation had insignificant effect on the C. arietinum seeds (P-value 0.196 > 0.05, Fstatistical < Fcritical) (Table 1b). Plant bioassays have been predominantly used by researchers to evaluate different environmental contaminants. Various plant seeds utilized for analyzing the toxic effect of dye degradation, include Vigna radiata, Vigna aconitifolia, Vigna mungo, Oryza sativa, Zea mays, Sorghum vulgare, Solanum leucopersicum, and Triticum aestivum (Pathak et al. 2014; Kumar et al. 2019; Thanavel et al. 2019; Singh and Dwivedi 2020).
Root growth is likely to be hampered under the influence of toxic compounds because of inhibited cell division. Hence, root growth analysis is a simple, rapid, and cost-inexpensive assay to identify toxicity (Saha end Rao 2020). Root numbers and lengths from the A cepa bulbs that underwent the various treatments have been tabulated in Table 2a. The highest number of roots developed from the bulbs under the control treatment followed by the bulbs under the degraded dye mixture metabolite treatment. The lowest root number and growth was observed from the bulbs treated with the untreated dye mixture. Statistical analysis revealed that the various treatments caused significant changes on the root number and length (P-value 0.04 < 0.05, Fstatistical > Fcritical). However, the concentration variations did not have any significant effect on the root growth of the bulbs (P-value 0.37 > 0.05, Fstatistical < Fcritical). Majority of phytotoxicity studies utilizing root growth as an indicator of toxicity have analyzed A. cepa bulbs (Kalyani et al. 2012; Ghosh et al. 2020; Yadav et al. 2021). Few studies have utilized other plants, such as Lactuca sativa, V. radiata, and Sinapis alba (Malachova et al. 2013; Almeida and Corso 2014; Pandiyan et al. 2022).
Cytogenotoxicity assay
An increase or decrease in MI denotes cytotoxicity and is used during environmental biomonitoring (Carita and Marin-Morales 2008). Aberrant cells consist of altered chromosomal number or morphology that occurred spontaneously in response to a toxic agent (Obe et al. 2002). The cytogenotoxic effects of the various treatments upon root tips have been tabulated (Table 2b). Roots treated with the degraded dye mixture metabolite solution documented the highest MI followed by the roots treated with the negative control solution. However, roots treated with the positive control solution documented the lowest MI followed by roots treated with the five-dye mixture solution. On the other hand, AI was highest in the positive control root cells followed by the five-dye mixture treated root cells. No aberrant cells were observed in the negative control cells, however the degraded five-dye mixture metabolite treated root-tip cells exhibited a low aberrant index. Therefore, it can be pointed out that toxicity substantially decreased after the dye mixture degradation resulting in less aberrant cells in the degraded dye metabolite treated roots. Statistical analysis revealed that the various treatments significantly affected the mitotic onion root-tip cells (P-values of MI (0.016266) and AI (0.000121) were < 0.05; Fstatistical > Fcritical in both cases). However, the various concentrations did not produce any significant changes in the root-tip cells (P-values of MI (0.56953379) and AI (0.0873) > 0.05, Fstatistical < Fcritical in both cases). Many previous studies have conducted cytogenotoxicity assay with A. cepa for identifying toxicity associated with dye degradation (Yadav et al. 2021; Patil and Jadhav 2013; Gurav et al. 2021).
Microtoxicity assay
The bacterial growth curves of E. coli, P. aeruginosa, and B. subtilis cultured with the five-dye mixture and degraded five-dye mixture supplementation as well as the control cultures without any supplementation were plotted (Fig. 9). After completion of the incubation period, the bacterial cell density was very low in cultures with the dye mixture supplementation and resulted in a flat curve. Thus, the dye treatment evidently inhibited bacterial cell growth. On the other hand, the bacterial cell density gradually increased in cultures with the degraded dye mixture metabolite supplement. The cellular growth density of E. coli, P. aeruginosa, and B. subtilis after the incubation period were 0.345, 0.325, and 0.36 respectively in the control cultures; and 0.385, 0.21, and 0.355 respectively in the degraded dye mixture metabolite treated cultures. The growth curves of the cultures with the different treatment supplementation significantly differed from each other (P-values were 1.75568E-07 < 0.05 (E. coli), 1.03E-07 < 0.05 (P. aeruginosa), and 1.97639E-07 < 0.05 (B. subtilis); Fstatistical > Fcritical). The results of this analysis implied that the degraded dye mixture metabolites did not stimulate any toxic effect on the test organisms. Other microorganisms used as an indicator of toxicity include soil microbial flora (Rhizobium radiobacter, Acinetobacter sp. Pseudomonas desmolyticum, Cellulomonas biazotea, isolated Escherichia coli DH5a, Micrococcus glutamicus and Proteus vulgaris), Azotobacter vinelandii, Rhizobium leguminosoma, Sinorhizobium meliloti, and Staphylococcus aureus (Kalyani et al. 2009; Saratale et al. 2010; Al-Tohamy et al. 2020; Yuan et al. 2020).
Biotoxicity assay
A. salina is a potential bioindicator for toxic compounds. The assay is simple, cost-inexpensive, gives rapid results, and able to generate results with less amount of test compounds (Sreedharan et al. 2021). The survival percentage of shrimps under the dye mixture and degraded dye mixture treatment has been presented in Fig. 10. The highest survival percentage was observed in the negative control treatment (100%) trailed by the degraded dye mixture metabolite treatment (87.15%), while the lowest survival percentage was observed in the positive control treatment (0.13%) followed by the dye mixture treatment (22.52%). The various treatments significantly differed from each other (P-value 0.001277 < 0.05, Fstatistical > Fcritical), however the concentration variation did not significantly affect the shrimp survival (P-value 0.297027 > 0.05, Fstatistical < Fcritical). Further, microscopic observation revealed visible dye build-up in the midgut of the dye mixture treated shrimps, while there was no dye buildup in the midgut of the positive control and degraded dye mixture metabolite treated shrimps (Fig. S1). Therefore, the results imply that significantly lower toxicity was induced on the shrimps by the degraded dye mixture metabolite than the untreated dye mixture. A. salina has been proven to be an excellent bioindicator for dye toxicity by several researchers (Priyaragini et al. 2014; Bilal et al. 2017; Ayed et al. 2019). Other animals and invertebrates that have been used for assessing dye toxicity include Daphnia magma, Rattus norvegicus (Swiss albino rats), Danio rerio (zebra fish), Xenopus laevis (frog) embryo, and Pheretima phosthuma (earthworm) (Birhanli and Ozmen 2005; Sharma et al. 2007; Kolekar et al. 2012; Barathi et al. 2020).
Biotoxicity assay of Artemia salina treated with five-dye mixture (C.I. Reactive Red 195, C.I. Reactive Orange 16, C.I. Reactive Black 5, C.I. Reactive Blue 221, C.I. Reactive Blue 250) (Dye mix) and degraded five-dye mixture metabolite (Degraded dye mix), K2Cr2O7 (positive control), and without any treatment (negative control)
Conclusion
Bacterial consortium VITPBC6 composed of Bacillus firmus VITEPB1, Bacillus flexus VITEPB2, Bacillus aryabhattai VITEPB3, B. flexus VITSP6, Bacillus paraflexus VITSPB7, and Bacillus megaterium VITSPB9 decolorized nine reactive dyes and two dye mixtures. Consortium VITPBC6 was most efficient in decolorizing the five-dye mixture. The dye mixture was maximally decolorized till a concentration of 150 mg L−1. Reaction and enzyme kinetic studies helped to shed an insight on the five-dye degradation process. The five-dye mixture was degraded following zero reaction order. Both Vmax and Km values were high for the degradation. Further, the five-dye mixture was successfully biodegraded by VITPBC6 into simple organic and inorganic compounds. The toxicity of the five-dye mixture was drastically reduced after biodegradation, and hence it can be considered safe for practical application. Therefore, consortium VITPBC6 is an excellent alternative for remediating dye pollution persistent in textile industries. Future scope of this study includes chalking out the pathway involved in the five-dye mixture biodegradation and studying the interaction of other pollutants, such as salt and heavy metals, with dyestuff present in the textile effluents.
References
Afrin S, Shuvo HR, Sultana B, Islam F, Rus’d AA, Begum S, Hossain MN (2021) The degradation of textile industry dyes using the effective bacterial consortium. Heliyon 7:e08102. https://doi.org/10.1016/j.heliyon.2021.e08102
Al-Tohamy R, Sun J, Fareed MF, Kenawy ER, Ali SS (2020) Ecofriendly biodegradation of reactive black 5 by newly isolated Sterigmatomyces halophilus SSA1575, valued for textile azo dye wastewater processing and detoxification. Sci Rep 10:12370. https://doi.org/10.1038/s41598-020-69304-4
Al-Tohamy R, Ali SS, Li F et al (2022) A critical review on the treatment of dye-containing wastewater: ecotoxicological and health concerns of textile dyes and possible remediation approaches for environmental safety. Ecotoxicol Environ Saf 231:113160. https://doi.org/10.1016/j.ecoenv.2021.113160
Alaguprathana M, Poonkothai M, Al-Ansari MM, Al-Humaid L, Kim W (2022) Cytogenotoxicity assessment in Allium cepa roots exposed to methyl orange treated with Oedogonium subplagiostomum AP1. Environ Res 213:113612. https://doi.org/10.1016/j.envres.2022.113612
Almeida EJR, Corso CR (2014) Comparative study of toxicity of azo dye procion red MX-5B following biosorption and biodegradation treatments with the fungi Aspergillus niger and Aspergillus terreus. Chemosphere 112:317–322. https://doi.org/10.1016/j.chemosphere.2014.04.060
APHA (2005) Standard methods for the examination of water and wastewater. American Water Works Association (AWWA), the Water Environment Federation (WEF) and the American Public Health Association (APHA), Washington D.C.
Asad S, Amoozegar MA, Pourbabaee A, Sarbolouki MN, Dastgheib SMM (2007) Decolorization of textile azo dyes by newly isolated halophilic and halotolerant bacteria. Bioresour Technol 98:2082–2088. https://doi.org/10.1016/j.biortech.2006.08.020
Asgher M (2012) Biosorption of reactive dyes: a review. Water Air Soil Pollut 223:2417–2435. https://doi.org/10.1007/s11270-011-1034-z
Asses N, Ayed L, Hkiri N, Hamdi M (2018) Congo red decolorization and detoxification by Aspergillus niger: removal mechanisms and dye degradation pathway. BioMed Res Int 2018:3049686. https://doi.org/10.1155/2018/3049686
Atkins P, de Paula J, Keeler J (2006) Atkins’ physical chemistry. Oxford University Press, Oxford
Ayed L, Khelifi E, Jannet HB, Miladi H, Cheref A, Achour S, Bakhrouf A (2010) Response surface methodology for decolorization of azo dye methyl orange by bacterial consortium: produced enzymes and metabolites characterization. Chem Eng J 165:200–208. https://doi.org/10.1016/j.cej.2010.09.018
Ayed L, Zmantar T, Bayar S, Cheref A, Achour S, Mansour HB, Mzoughi R (2019) Potential use of probiotic consortium isolated from kefir for textile azo dye decolorization. J Microbiol Biotechnol 29:1629–1635. https://doi.org/10.4014/jmb.1906.06019
Barathi S, Karthik C, Nadanasabapathi S, Padikasan IA (2020) Biodegradation of textile dye reactive blue 160 by Bacillus firmus (Bacillaceae: Bacillales) and non-target toxicity screening of their degraded products. Toxicol Rep 7:16–22. https://doi.org/10.1016/j.toxrep.2019.11.017
Berg JM, Tymoczko JL, Stryer L (2002) Biochemistry. W. H. Freeman and Company, New York
Biju LM, Pooshana V, Kumar PS, Gayathri KV, Ansar S, Govindaraju S (2022) Treatment of textile wastewater containing mixed toxic azo dye and chromium (VI) by haloalkaliphilic bacterial consortium. Chemosphere 287:132280. https://doi.org/10.1016/j.chemosphere.2021.132280
Bilal M, Iqbal HM, Hu H, Wang W, Zhang X (2017) Enhanced bio-catalytic performance and dye degradation potential of chitosan-encapsulated horseradish peroxidase in a packed bed reactor system. Sci Total Environ 575:1352–1360. https://doi.org/10.1016/j.scitotenv.2016.09.215
Birhanli A, Ozmen M (2005) Evaluation of the toxicity and teratogenity of six commercial textile dyes using the frog embryo teratogenesis assay–Xenopus. Drug Chem Toxicol 28:51–65. https://doi.org/10.1081/dct-39689
Carita R, Marin-Morales MA (2008) Induction of chromosome aberrations in the Allium cepa test system caused by the exposure of seeds to industrial effluents contaminated with azo dyes. Chemosphere 72:722–725. https://doi.org/10.1016/j.chemosphere.2008.03.056
Dawkar VV, Jadhav UU, Jadhav SU, Govindwar SP (2008) Biodegradation of disperse textile dye Brown 3REL by newly isolated Bacillus sp. VUS. J Appl Microbiol 105:14–24. https://doi.org/10.1111/j.1365-2672.2008.03738.x
Docker A, Wattie JM, Topping MD et al (1987) Clinical and immunological investigations of respiratory disease in workers using reactive dyes. Br J Ind Med 44:534–541. https://doi.org/10.1136/oem.44.8.534
dos Santos AB, Bisschops IA, Cervantes FJ, van Lier JB (2004) Effect of different redox mediators during thermophilic azo dye reduction by anaerobic granular sludge and comparative study between mesophilic (30 C) and thermophilic (55 C) treatments for decolourisation of textile wastewaters. Chemosphere 55:1149–1157. https://doi.org/10.1016/j.chemosphere.2004.01.031
Edoamodu CE, Nwodo UU (2021) Enterobacter sp. AI1 produced a thermo-acidic-tolerant laccase with a high potential for textile dyes degradation. Biocatal Agric Biotechnol 38:102206. https://doi.org/10.1016/j.bcab.2021.102206
Field JA, Brady J (2003) Riboflavin as a redox mediator accelerating the reduction of the azo dye mordant yellow 10 by anaerobic granular sludge. Water Sci Technol 48:187–193. https://doi.org/10.2166/wst.2003.0393
Ghosh SK, Saha PD, Di MF (2020) Recent trends in waste water treatment and water resource management. Springer, Berlin
Grand View Research (2022) Textile market size, share & trends analysis report by raw material (cotton, wool, silk, chemical), by product (natural fibers, nylon), by application (technical, fashion), by region, and segment forecasts, 2022–2030. https://www.grandviewresearch.com/industry-analysis/textile-market. Accessed 29 Nov 2022
Guo G, Tian F, Zhang C et al (2019) Performance of a newly enriched bacterial consortium for degrading and detoxifying azo dyes. Water Sci Technol 79:2036–2045. https://doi.org/10.2166/wst.2019.210
Gurav R, Bhatia SK, Choi TR et al (2021) Application of macroalgal biomass derived biochar and bioelectrochemical system with Shewanella for the adsorptive removal and biodegradation of toxic azo dye. Chemosphere 264:128539. https://doi.org/10.1016/j.chemosphere.2020.128539
Haq I, Kumari V, Kumar S, Raj A, Lohani M, Bhargava R (2016) Evaluation of the phytotoxic and genotoxic potential of pulp and paper mill effluent using Vigna radiata and Allium cepa. Adv Biol 2016:8065736. https://doi.org/10.1155/2016/8065736
He F, Hu W, Li Y (2004) Biodegradation mechanisms and kinetics of azo dye 4BS by a microbial consortium. Chemosphere 57:293–301. https://doi.org/10.1016/j.chemosphere.2004.06.036
Iark D, dos Reis Buzzo AJ, Garcia JAA et al (2019) Enzymatic degradation and detoxification of azo dye congo red by a new laccase from Oudemansiella canarii. Bioresour Technol 289:121655. https://doi.org/10.1016/j.biortech.2019.121655
Kalyani DC, Telke AA, Govindwar SP, Jadhav JP (2009) Biodegradation and detoxification of reactive textile dye by isolated Pseudomonas sp. SUK1. Water Environ Res 81:298–307. https://doi.org/10.2175/106143008x357147
Kalyani DC, Telke AA, Surwase SN, Jadhav SB, Lee JK, Jadhav JP (2012) Effectual decolorization and detoxification of triphenylmethane dye malachite green (MG) by Pseudomonas aeruginosa NCIM 2074 and its enzyme system. Clean Technol Environ Policy 14:989–1001. https://doi.org/10.1007/s10098-012-0473-6
Karim ME, Dhar K, Hossain MT (2018) Decolorization of textile reactive dyes by bacterial monoculture and consortium screened from textile dyeing effluent. J Genet Eng Biotechnol 16:375–380. https://doi.org/10.1016/j.jgeb.2018.02.005
Khan AU, Rehman MU, Zahoor M, Shah AB, Zekker I (2021) Biodegradation of brown 706 dye by bacterial strain Pseudomonas aeruginosa. Water 13:2959. https://doi.org/10.3390/w13212959
Kolekar YM, Nemade HN, Markad VL, Adav SS, Patole MS, Kodam KM (2012) Decolorization and biodegradation of azo dye, reactive blue 59 by aerobic granules. Bioresour Technol 104:818–822. https://doi.org/10.1016/j.biortech.2011.11.046
Kumar N, Sinha S, Mehrotra T, Singh R, Tandon S, Thakur IS (2019) Biodecolorization of azo dye acid black 24 by Bacillus pseudomycoides: process optimization using box Behnken design model and toxicity assessment. Bioresour Technol Rep 8:100311. https://doi.org/10.1016/j.biteb.2019.100311
Lade H, Kadam A, Paul D, Govindwar S (2015) Biodegradation and detoxification of textile azo dyes by bacterial consortium under sequential microaerophilic/aerobic processes. EXCLI J 14:158. https://doi.org/10.17179%2Fexcli2014-642
Madhushika HG, Ariyadasa TU, Gunawardena SH (2019) Biological decolourization of textile industry wastewater by a developed bacterial consortium. Water Sci Technol 80:1910–1918. https://doi.org/10.2166/wst.2020.010
Malachova K, Rybkova Z, Sezimova H, Cerven J, Novotny C (2013) Biodegradation and detoxification potential of rotating biological contactor (RBC) with Irpex lacteus for remediation of dye-containing wastewater. Water Res 47:7143–7148. https://doi.org/10.1016/j.watres.2013.07.050
Mishra A, Takkar S, Joshi NC, Shukla S, Shukla K, Singh A, Manikonda A, Varma A (2022) An integrative approach to study bacterial enzymatic degradation of toxic dyes. Front Microbiol 12:802544. https://doi.org/10.3389/fmicb.2021.802544
Mohanty SS, Kumar A (2021) Enhanced degradation of anthraquinone dyes by microbial monoculture and developed consortium through the production of specific enzymes. Sci Rep 11:7678. https://doi.org/10.1038/s41598-021-87227-6
Mondal NK, Debnath P (2022) Impact of two commercially available hair dyes on germination, morpho-physiology, and biochemistry of Cicer arietinum L. and cytotoxicity study on Allium cepa L. root tip. Environ Res 208:112681. https://doi.org/10.1016/j.envres.2022.112681
Obe G, Pfeiffer P, Savage JRK et al (2002) Chromosomal aberrations: formation, identification and distribution. Mutat Res 504:17–36. https://doi.org/10.1016/s0027-5107(02)00076-3
Pandiyan R, Dharmaraj S, Ayyaru S et al (2022) Ameliorative photocatalytic dye degradation of hydrothermally synthesized bimetallic Ag-Sn hybrid nanocomposite treated upon domestic wastewater under visible light irradiation. J Hazard Mater 421:126734. https://doi.org/10.1016/j.jhazmat.2021.126734
Pathak H, Soni D, Chauhan K (2014) Evaluation of in vitro efficacy for decolorization and degradation of commercial azo dye RB-B by Morganella sp. HK-1 isolated from dye contaminated industrial landfill. Chemosphere 105:126–132. https://doi.org/10.1016/j.chemosphere.2014.01.004
Patil AV, Jadhav JP (2013) Evaluation of phytoremediation potential of Tagetes patula L. for the degradation of textile dye reactive blue 160 and assessment of the toxicity of degraded metabolites by cytogenotoxicity. Chemosphere 92:225–232. https://doi.org/10.1016/j.chemosphere.2013.01.089
Prasad ASA, Rao KVB (2013) Aerobic biodegradation of azo dye by Bacillus cohnii MTCC 3616; an obligately alkaliphilic bacterium and toxicity evaluation of metabolites by different bioassay systems. Appl Microbiol Biotechnol 97:7469–7481. https://doi.org/10.1007/s00253-012-4492-3
Priyaragini S, Veena S, Swetha D, Karthik L, Kumar G, Rao KB (2014) Evaluating the effectiveness of marine actinobacterial extract and its mediated titanium dioxide nanoparticles in the degradation of azo dyes. J Environ Sci 26:775–782. https://doi.org/10.1016/S1001-0742(13)60470-2
Rao KVB, Prasad ASA (2014) Biodecolourisation of azo dye reactive red 22 by Bacillus infantis strain AAA isolated from seawater and toxicity assessment of degraded metabolites. Nat Environ Pollut Technol 13:369–374
Rima SAJ, Paul GK, Islam S, Akhtar-E-Ekram M, Zaman S, Saleh MA, Uddin MS (2022) Efficacy of Pseudomonas sp. and Bacillus sp. in textile dye degradation: a combined study on molecular identification, growth optimization, and comparative degradation. J Hazard Mater Lett 3:100068. https://doi.org/10.1016/j.hazl.2022.100068
Saha P, Rao KVB (2020) Biotransformation of reactive Orange 16 by alkaliphilic bacterium Bacillus flexus VITSP6 and toxicity assessment of biotransformed metabolites. Int J Environ Sci Technol 17:99–114. https://doi.org/10.1007/s13762-019-02256-z
Saha P, Sivaramakrishna A, Rao KVB (2022) Bioremediation of reactive orange 16 by industrial effluent-adapted bacterial consortium VITPBC6: process optimization using response surface methodology (RSM), enzyme kinetics, pathway elucidation, and detoxification. Environ Sci Pollut Res. https://doi.org/10.1007/s11356-022-24501-8
Saratale RG, Saratale GD, Chang JS, Govindwar SP (2010) Decolorization and biodegradation of reactive dyes and dye wastewater by a developed bacterial consortium. Biodegradation 21:999–1015. https://doi.org/10.1007/s10532-010-9360-1
Saratale RG, Saratale GD, Chang JS, Govindwar SP (2011) Bacterial decolorization and degradation of azo dyes: a review. J Taiwan Inst Chem Eng 42:138–157. https://doi.org/10.1016/j.jtice.2010.06.006
Saratale RG, Gandhi SS, Purankar MV, Govindwar SP, Oh SE, Saratale GD (2013) Decolorization and detoxification of sulfonated azo dye C.I Remazol red and textile effluent by isolated Lysinibacillus sp. RGS. J Biosci Bioeng 115:658–667. https://doi.org/10.1016/j.jbiosc.2012.12.009
Sarioglu M, Bali U, Bisgin T (2007) The removal of CI Basic Red 46 in a mixed methanogenic anaerobic culture. Dyes Pigm 74:223–229. https://doi.org/10.1016/j.dyepig.2006.02.001
Senthil Kumar J, Ponmurugan P, Vinothkanna A (2020) A primer on earth pollution: pollution types and disposal. Bentham Science Publishers, USA
Shade A, Peter H, Allison SD, Baho D, Berga M, Bürgmann H, Huber DH, Langenheder S, Lennon JT, Martiny JB, Matulich KL (2012) Fundamentals of microbial community resistance and resilience. Front Microbiol 3:417. https://doi.org/10.3389%2Ffmicb.2012.00417
Sharma S, Kalpana AS, Suryavathi V, Singh PRS, Sharma K (2007) Toxicity assessment of textile dye wastewater using swiss albino rats. Aust J Ecotoxicol 13:81–85. https://doi.org/10.3316/informit.377001889081181
Shi Y, Yang Z, Xing L et al (2021) Recent advances in the biodegradation of azo dyes. World J Microbiol Biotechnol 37:137–155. https://doi.org/10.1007/s11274-021-03110-6
Shukla CA, Kute MS, Kulkarni AA (2021) Towards sustainable continuous production of azo dyes: possibilities and techno-economic analysis. Green Chem 23:6614–6624. https://doi.org/10.1039/D1GC01133B
Singh K, Arora S (2011) Removal of synthetic textile dyes from wastewaters: a critical review on present treatment technologies. Crit Rev Environ Sci Technol 41:807–878. https://doi.org/10.1080/10643380903218376Uddin
Singh G, Dwivedi SK (2020) Decolorization and degradation of Direct Blue-1 (azo dye) by newly isolated fungus Aspergillus terreus GS28, from sludge of carpet industry. Environ Technol Innov 18:100751. https://doi.org/10.1016/j.eti.2020.100751
Sreedharan V, Saha P, Rao KVB (2021) Dye degradation potential of Acinetobacter baumannii strain VITVB against commercial azo dyes. Bioremediat J 25:347–368. https://doi.org/10.1080/10889868.2020.1871317
Stolz A (2001) Basic and applied aspects in the microbial degradation of azo dyes. Appl Microbiol Biotechnol 56:69–80. https://doi.org/10.1007/s002530100686
Thanavel M, Bankole PO, Kadam S, Govindwar SP, Sadasivam SK (2019) Desulfonation of the textile azo dye acid fast yellow MR by newly isolated Aeromonas hydrophila SK16. Water Resour Ind 22:100116. https://doi.org/10.1016/j.wri.2019.100116
Thiruppathi K, Rangasamy K, Ramasamy M, Muthu D (2021) Evaluation of textile dye degrading potential of ligninolytic bacterial consortia. Environ Chall 4:100078. https://doi.org/10.1016/j.envc.2021.100078
Uddin F (2019) Textile manufacturing processes. IntechOpen, UK
Ul-Islam S (2017) Advanced materials for wastewater treatment. Scrivener Publishing, USA
Wanyonyi WC, Onyari JM, Shiundu PM, Mulaa FJ (2017) Biodegradation and detoxification of malachite green dye using novel enzymes from Bacillus cereus strain KM201428: kinetic and metabolite analysis. Energy Procedia 119:38–51. https://doi.org/10.1016/j.egypro.2017.07.044
Yadav A, Yadav P, Singh AK et al (2021) Decolourisation of textile dye by laccase: process evaluation and assessment of its degradation bioproducts. Bioresour Technol 340:125591. https://doi.org/10.1016/j.biortech.2021.12559
Yaseen DA, Scholz M (2019) Textile dye wastewater characteristics and constituents of synthetic effluents: a critical review. Int J Environ Sci Technol 16:1193–1226. https://doi.org/10.1007/s13762-018-2130-z
Yuan H, Chen L, Cao Z, Hong FF (2020) Enhanced decolourization efficiency of textile dye reactive blue 19 in a horizontal rotating reactor using strips of BNC-immobilized laccase: optimization of conditions and comparison of decolourization efficiency. Biochem Eng J 156:107501. https://doi.org/10.1016/j.bej.2020.107501
Zahran SA, Ali-Tammam M, Hashem AM, Aziz RK, Ali AE (2019) Azoreductase activity of dye-decolorizing bacteria isolated from the human gut microbiota. Sci Rep 9:5508. https://doi.org/10.1038/s41598-019-41894-8
Zhao J, Wu QX, Cheng XD et al (2021) Biodegradation and detoxification of the triphenylmethane dye coomassie brilliant blue by the extracellular enzymes from mycelia of Lactarius deliciosus. Front Chem Sci Eng 15:421–436. https://doi.org/10.1007/s11705-020-1952-7
Zhuang M, Sanganyado E, Zhang X et al (2020) Azo dye degrading bacteria tolerant to extreme conditions inhabit nearshore ecosystems: optimization and degradation pathways. J Environ Manage 261:110222. https://doi.org/10.1016/j.jenvman.2020.110222
Acknowledgements
The authors are thankful to the management for providing with the necessary infrastructure to carry out this research work.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
PS was responsible for conceptualization of the work, carrying out the entire work and formal analysis of research data, and writing the manuscript and KVBR provided the necessary resources, validated the obtained data, supervised the entire research work. Both the authors reviewed the manuscript after completion.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Saha, P., Rao, K.V.B. Biodegradation of commercial textile reactive dye mixtures by industrial effluent adapted bacterial consortium VITPBC6: a potential technique for treating textile effluents. Biodegradation 35, 173–193 (2024). https://doi.org/10.1007/s10532-023-10047-0
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-023-10047-0