Abstract
Microalgal and cyanobacterial communities have a key role in sustaining the fertility of aquatic and terrestrial habitats. Thus, understanding the actual biodiversity of these communities is a task of utmost importance. However, this particular task suffers from several technical constraints and challenges. Sampling procedures and criteria for counting individuals of various microalgal and cyanobacterial species in such systems have not been standardized. Biodiversity indices are considered promising; however, ambiguity in respect of species concept and characterization criteria of microalgal and cyanobacterial forms makes the determination of biodiversity indices a complicated task. Recently, DNA barcoding was employed for the identification of microalgal and cyanobacterial species. However, it needs sufficient experimental validation. The functional diversity and zeta diversity, which are helpful in ecosystem process assessment, are largely unexplored for microalgal and cyanobacterial communities. Adequate knowledge of sampling designs, methods for detecting outliers and errors, and data transformation in biodiversity studies are crucial. Several analytical tools, such as analysis of variance (ANOVA), analysis of similarity (ANOSIM), multidimensional scaling (MDS) and cluster analysis are obligatory for understanding the compositional differences of different microbial communities. Regression and multiple correlations are important in realizing the relationships of different environmental factors. Principal component analysis (PCA) and canonical correspondence analysis (CCA) are effective in interpreting the influence of environmental factors on the distribution of microalgal and cyanobacterial species in a geographical region or a land patch. Nevertheless, statistical software packages are the backbone of research activities these days. So, development of new biodiversity software packages specific to microalgae and cyanobacteria is required.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microalgae and cyanobacteria are well known for possessing tremendous potential to colonize a range of habitats starting from the comfiest one to the extreme conditions like thermal springs, cold springs, and hypersaline systems (Badger et al. 2006; Ward et al. 2012; Singh et al. 2018; Malavasi et al. 2020). These organisms are extremely important for sustaining vital processes of aquatic and terrestrial systems, including agroecosystems (Sharma et al. 2012; Sharma 2015; Singh et al. 2014; Rai et al. 2019; Chittora et al. 2020). They may play important role in providing resilience to the rhizospheric microbial community (Ahmed et al. 2010) as they do in aquatic ecosystems (Akins et al. 2018). However, microalgae and their community composition also change under the influence of environmental disturbances (Chaurasia 2015; Lear et al. 2017; Ward et al. 2017). Hence, the biodiversity of microalgae and cyanobacteria and their responses to diverse physicochemical factors should be explored appropriately (Chaurasia 2015). Unfortunately, microalgae and cyanobacteria exist as a marginalized component of the biodiversity research (Nabout et al. 2013; Rejmánková et al. 2004; Chaurasia 2015; Suganya et al. 2016), and the role of their community composition has often been ignored in modelling of agriculture ecosystem processes (Allison and Martiny 2008).
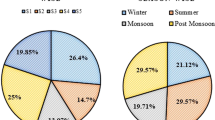
Owing to their small size and highly variable pattern of distribution, adequate assessment of the biodiversity of microorganisms is a more challenging task than that of higher plants. Disparities in pattern of distribution are indeed due to variations in prevailing environmental parameters which may be experienced by these organisms even on proximate sites due to vertical and horizontal gradients of the factors, such as pH, temperature, humidity and nutrients availability (Armitage et al. 2012; Shen et al. 2015; O’Brien et al. 2016; Van Der Putten 2017; Banerjee et al. 2020). Sometimes, temporal variations are crucial in shaping the structure of a microbial community. For example, in early spring, the diatom populations in freshwater systems increase due to the availability of nutrients but when light intensity increases during summer, species richness of green algae and cryptophytes rises. Subsequently, green algae are replaced by large-sized diatom species and cyanobacteria during early autumn, while the community is consecutively re-dominated by diatoms with the return of winter (Sommer et al. 1986, 2012). Similar kinds of changes in the microbial community of agriculture systems can also be expected during different crops seasons. Thus, the timing of field study becomes critically important in a biodiversity study and hence it needs adequate attention of the researchers (Fattorini 2003; Pagliarella et al. 2018).
In biodiversity researches, plant and animal ecologists generally employ advanced statistical concepts and tools, such as sampling designs, categorization, normalization, data extrapolation, regression analysis, ANOVA, factor analysis, cluster analysis, logistic regression, generalized linear and generalized additive modelling in order to enhance the quality of their studies (Guisan et al. 2002; Fattorini 2003; Chiarucci et al. 2011; Pagliarella et al. 2018). However, very few microbiologists have given suitable consideration to the above tools for assessing microbial diversity of diverse habitats (Ampe and Miambi 2000; Oliveira et al. 2020; Banerjee et al. 2020). Counting individuals present in the sampling area of a specified quadrat size has been recognized as a standard procedure all through studying the diversity of higher plants (Misra 1968; Kershaw 1973; Cox 1990; Elzinga et al. 2001). However, this approach cannot be straightforwardly employed for exploring microbial biodiversity. Here, we become either dependent on the haemocytometer-based microscopic counting or circumscribed to use the advanced but expensive molecular and omics approaches (Hill et al. 2003; Gupta et al. 2013; Emerson et al. 2017; Kushwaha et al. 2020). The development of molecular tools has changed the scenario of studying microbial diversity. This has greatly helped us in deciphering the existence of non-culturable microbial forms, which are considered as the major components of the soil ecosystem (Schleifer 2004; Bodor et al. 2020). However, to extract the comprehensible information, both the haemocytometer- and the molecular tool-based biodiversity assessment methodology ultimately require mathematical, statistical and software-based data analysis (Ampe and Miambi 2000; Hill et al. 2003; Oliveira et al. 2020; Banerjee et al. 2020).
The present study provides an overview of the biodiversity of microalgae and cyanobacteria in crop fields. It includes a specific discussion on biodiversity concept, types, components and different measures. The modern statistical tools, such as principal component analysis (PCA) and canonical correspondence analysis (CCA) have been covered as these tools support in identifying the major factors influencing the distribution of microorganisms in a specific habitat or community. ANOSIM and SIMPER have also been discussed subsequently since they are useful in the comparative analysis of the biodiversity between communities and at the landscape level. Efforts have also been made to describe the merits and shortcomings of some available software packages.
Diversity of microalgae and cyanobacteria in the agricultural fields
The importance of cyanobacteria in enhancing the fertility of agricultural fields or reclamation of usar lands has long been very well established (Singh 1950, 1961). The potential of microalgae and cyanobacteria in agriculture and other applied fields was recently reviewed by several researchers (Abdel-Raouf et al. 2012; Singh et al. 2014, 2016; Abinandan et al. 2019; Rai et al. 2019; Chittora et al. 2020). A list of microalgae and cyanobacteria often reported from agriculture fields can be seen in Table 1, with a brief mention of major mechanisms through which they may contribute to enhancing the quality of agricultural lands.
Based on laboratory and pilot-scale experiments, the beneficial effects of some selected microalgal and cyanobacterial species in improving the phosphorus, nitrogen, and carbon content of the soil has been regularly documented (Karthikeyan et al. 2009; Prasanna et al. 2012; Natarajan et al. 2012; Swarnalakshmi et al. 2013). But, the actual biodiversity of microalgae and cyanobacteria in agriculture fields has been sporadically investigated (Ahmed et al. 2010; Alvarez et al. 2021). Irissari et al. (2001) reported the diversity of unicellular, heterocystous and non-heterocystous cyanobacteria in the paddy fields of Uruguay. The diversity of N2-fixing cyanobacteria in agricultural fields of Thailand increased with the crop rotation process and was affected by environmental factors and season (Chunleuchanon et al. 2003). In the rice fields of Fujian (China), 11 genera of cyanobacteria were identified using 16S rRNA gene sequencing (Song et al. 2005). The occurrence of cyanobacteria and microalgae from the cornfields of north-eastern Italy showed a decrease in cyanobacterial diversity due to prolonged use of chemical fertilizers (Zancan et al. 2006). Hendrayanti et al. (2018) chronicled various cyanobacterial representatives from the paddy fields of Serang Mekar Village, Ciparay-South Bandung, West Java, Indonesia. The cyanobacterial richness in the agricultural lands of Al Diwaniyah city (Iraq) was represented by 96 species mostly belonging to N2-fixing unicellular and filamentous forms (Alghanmi and Jawad 2019).
Several unicellular, heterocystous and non-heterocystous cyanobacteria were documented from the rice fields of different states of India like Kerala, Meghalaya, Tamil Nadu, Assam, Bihar, Orissa, Uttar Pradesh, Telangana and Maharashtra (Prasanna and Nayak 2007; Srivastava et al. 2009; Dey et al. 2010; Bharadwaj and Baruah 2013; Singh et al. 2014; Khare et al. 2014; Vijayan and Ray 2015; Srinivas and Aruna 2016). Anabaena circinalis showed the maximum relative abundance among the diverse cyanobacterial species reported from the rice fields of Assam (Bharadwaj and Baruah 2013). The rice-based cropping systems of north and eastern India exhibited the presence of Nostoc, Anabaena and Phormidium with the predominance of heterocystous forms (Prasanna et al. 2013b). Singh et al. (2014) recorded 29 cyanobacterial strains from the paddy fields of Chhattisgarh. Of which, 15 were non-heterocystous. Likewise, 19 species of cyanobacteria (11 heterocystous and 7 non-heterocystous) were identified from paddy fields of Bihar, India (Khare et al. 2014). In the Kuttanadu Paddy Wetlands (Kerala, India), 45 species of cyanobacteria were documented. Here, Chroococcus turgidus showed the maximum relative abundance, while the highest species richness was observed during monsoon season when paddy crop attained the panicle growth stage (Vijayan and Ray 2015). Srinivas and Aruna (2016) reported the members of Nostocaceae, Chroococaceae, Scytonemataceae, Oscillatoriaceae in the rice fields of Telangana, India. Anabaena and Oscillatoria were abundant in the paddy fields of Patan and Karad (Maharashtra, India; Ghadage and Karande 2019).
Some researchers have documented cyanobacterial species from soils other than paddy fields. Zancan et al. (2006) reported the cyanobacterial diversity of cornfields of north-eastern, Italy. Ahlesaadat et al. (2017) characterized the cyanobacterial diversity of wheat fields of Yazd province, Iran. Recently, Alghanmi and Jawad (2019) have explored cyanobacterial diversity from soils of a variety of crops of Al Diwaniyah city, Iraq. Rai et al. (2018) investigated the diversity of cyanobacterial forms along the rural-urban gradient. These latter authors concluded that urbanization adversely affected the diversity and microbial community composition but favoured heterocystous forms.
It is concerning to note that a majority of the above-mentioned studies are restricted to cyanobacteria totally ignoring the microalgal component of the ecosystem. Further, most of these studies usually lack adequate quantitative estimates of the microalgal and cyanobacterial diversity and also miserably fail to furnish sufficient sampling details. The fact that most of the field studies underestimate the cyanobacterial diversity is attributable inter alia to (i) low sampling efforts, (ii) sensitivity of molecular markers used, and (iii) definition of species as per the researcher (Dvořák et al. 2015).
Biodiversity and its types
Biodiversity (Wilson 1988) refers to all kinds of variations in organisms starting from gene to biosphere levels. One may come across a variety of terms, such as genetic diversity, phylogenetic diversity, species diversity, ecological diversity, ecosystem diversity, functional diversity, etc., in the existing literature. All these terms are used either to express the different levels of understanding of biodiversity or to reflect its diverse ecological and functional perspectives. Whittaker (1972) introduced the concept of alpha, beta and gamma diversity, which is an illustration of biodiversity within the community, between-community and at the landscape level. Gamma diversity represents the total diversity of the landscape, while alpha diversity is the diversity of the sub-communities residing at a local scale. These two diversities are straightforward to comprehend and measure. Beta diversity, however, is comparative and represents the differences between the two sub-communities. Ecologists first estimate alpha and gamma diversity and then derive beta diversity from these two. Initially, beta diversity was proposed to involve multiplicative portioning (i.e., DαDβ = Dγ), however, the latter additive formulation was proposed (i.e., Dα + Dβ = Dγ) taking into account that alpha and beta diversity are not necessarily independent (Daly et al. 2018). It seems worth mentioning here that though additive and multiplicative partitioning of biodiversity are appreciated and widely used due to offering a single set of values of alpha and beta diversity, both methods suffer from the disadvantage of significant loss of information (Daly et al. 2018).
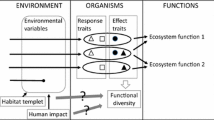
Biodiversity which focuses on the functional roles of species in communities and ecosystems is termed functional diversity (Laureto et al. 2015). The functional diversity of habitat, niche space, community or ecosystems is of immense importance as it is directly related to the diverse aspects of ecosystem processes, such as productivity, nutrient cycling, ecosystem stability and sustainability (Petchey and Gaston 2006; Costanza et al. 2007; Laureto et al. 2015). The idea of plant functional traits has emerged from here and now has attracted a great deal of attention of modern-day ecologists, working in the field of higher plants diversity (Petchey and Gaston 2006; Laureto et al. 2015). Different kinds of models, such as the sampling effect model and niche differentiation model, have been proposed by ecologists to assess the effects of functional diversity on the productivity of the ecosystem. Species redundancy hypothesis and niche complementarity model help understand the relationship between functional diversity and ecosystem processes (Goswami et al. 2017). The two widely used models, rivets and idiosyncratic are useful in comprehending the interdependency of species richness and functional diversity for the stability of an ecosystem (Ehrlich and Ehrlich 1981; Lawton 1994). Nevertheless, the concept of functional diversity has not been adequately explored in the case of microalgae and cyanobacteria (Goswami et al. 2017). Most of the microbial studies conducted so far are devoted largely to discovering the species and phylogenetic diversities of the microbial communities in question.
Basic components of biodiversity
Species richness and species concept in cyanobacteria and microalgae
Species richness and evenness are the two primary components of biodiversity. Almost all kinds of indices incorporate these two components for providing a quantitative assessment of biodiversity. Another component, which has gained less attention from researchers, is disparity (Daly et al. 2018). The total number of species present in the community under study is called species richness. By and large, species richness is straightforward as taxonomic identification and description of a new species is well described for higher plants. However, in the case of microalgae and cyanobacteria, both the species concept and the criteria for taxonomic identification of a new species are ambiguous (Gupta et al. 2013; Chaurasia 2015; Dvořák et al. 2015; Komárek 2016). Identification of microalgal and cyanobacterial species based only on morphological features is not appreciated nowadays because of phenotypic plasticity (to different environments and the culture media) and the presence of cryptic species (Hadi et al. 2016). As cyanobacteria reproduce asexually, their different identified forms do not also fully satisfy the criteria of the biological species concept. Rippka et al. (1979) used to classify cyanobacterial forms into different groups, but, this grouping is inadequate in view of the taxonomic species concept.
Classification based on 16S rRNA gene sequence is used for molecular identification of cyanobacteria (Hoffmann et al. 2005). Komárek (2006) advocated the use of molecular criteria for the identification of cyanobacterial species. However, the use of this approach becomes debatable when the outcome is correlated with morphological features, particularly considering the adaptations of cyanobacteria to the changing environmental conditions (Gupta et al. 2013). Moreover, molecular methods used for identification also do not fully justify the biological species concept. This approach may also reveal variable species identification of the same specimen by employing different molecular markers. Recently, DNA barcoding was employed by some researchers for the identification of microalgal and cyanobacterial species (Dvořák et al. 2015; Ballesteros et al. 2021). However, Eckert and his team reported barcoding gaps in more than half of the studied cases (Eckert et al. 2015). Hence barcoding needs proper validation before using it as a tool for the identification of microalgal or cyanobacterial species.
The polyphasic approach that takes into account morphological, genetic and ecological attributes of cyanobacteria and microalgae for species characterization has been employed by many researchers (Komárek and Kaštovský 2003; Zapomělová et al. 2013; Hauer et al. 2014; Komárek 2016; Sciuto and Moro 2015; Renuka et al. 2018). Several recent taxonomic revisions of cyanobacteria are based on this approach. However, some serious concerns are associated with this approach too. Dvořák and co-workers have provided an elegant discussion on the species concept and taxonomic diversity of cyanobacteria in their review (Dvořák et al. 2015). Moreover, the status of species characterization in the case of cyanobacteria and microalgae is still puzzling. Thus, this aspect demands sincere efforts as without framing a sound basis of species concept and identification of cyanobacteria and microalgae, it would not be possible to realize their actual diversity in any habitat or community, including agricultural ecosystems (Palinska and Surosz 2014; Chaurasia 2015; Komárek 2016).
Species evenness
The equitability of distribution of species inhabiting the community of interest is mentioned as its evenness in the field of ecology. If all the species inhabiting the community are present in equal proportions, it is called even. In contrast, if species are disproportionately present with one or two species dominating the community, it is referred to as uneven (Wittebolle et al. 2009). This concept is straightforward in the case of the diversity of higher plants, but it becomes yet again complicated for cyanobacteria and microalgae due to the imprecise nature of species concept and their taxonomic identification. Evenness is a key factor that regulates the functional stability of ecosystems. It is also important for understanding the representation of functional traits of each species. The communities with uneven distribution of species are often believed to be susceptible to invasion and are not resilient to stresses and disturbances (Wittebolle et al. 2009; Daly et al. 2018).
Species disparity
The third but ignored component of diversity is disparity (Daly et al. 2018). The species richness and evenness are based on species-neutral diversity. This means that distinct species have nothing in common. These components do not account for any disparity between species. According to this, a community of five markedly different species is not considered more diverse than a community of five species of the same genus. However, this might not always be the case in a natural community, particularly in the case of microbial ones. Various species of the same genus may possess several common attributes and thus might greatly influence the functional stability of the community. Thus, the disparity is somehow accounting both the similarity and dissimilarity that exist between similar kinds of species. The measurement of similarity or dissimilarity between species can be done considering genetic, functional, morphological and phylogenetic grounds.
Functional diversity
Villéger et al. (2008) introduced the concept of enumerating functional diversity. This idea involves an inclusive approach and integrally involve the issue of disparity. These authors recommended enumeration of functional richness, functional evenness and functional divergence. Since these indices are independently calculated, they do not influence each other similar to Whittakerian measures like alpha, beta and gamma diversity. In addition, functional diversity measures are complementary indices. The functional richness has merit to consider the niche and the niche volume of a particular species in a community (Mason and Mouillot 2013). Functional evenness gives weightage to species abundance when functional space is filled by species (Villéger et al. 2008). Divergence of species in their functional space from the centre of gravity is analyzed by functional divergence, which also prioritizes abundance. Thus, these indices independently dispense arrangement of species (relative abundance and orientation) in a multidimensional functional space and bring into light biodiversity–environment–ecosystem relationships (Villéger et al. 2008). The mathematical expressions and other details of functional diversity indices can be found elsewhere (Villéger et al. 2008; Mason and Mouillot 2013). Moreover, the concept of functional diversity is largely unexplored for microbial communities and thus demands adequate attention.
Biodiversity measurement within the community
Any important study of biodiversity, no matter which aspect is in focus, must include a quantitative evaluation. However, it is a complicated task both theoretically and practically. Biodiversity is enumerated by developing mathematical functions, usually known as biodiversity indices. The use of such indices allows comparison between spatial regions, temporal periods, taxa, niches or trophic levels. Biodiversity indices measure the taxonomical and phylogenetical relationship of the species and are the numerical, partial inter-changeable tools to quantify diversity (Clarke and Warwick 2001; Contoli and Luiselli 2015). After employing molecular tools for the identification of cyanobacteria and microalgae, indices are applied to enumerate the diversity of the region under study. For the metagenomics approach of diversity analysis, the indices may be calculated by counting the Operational Taxonomic Units (Hill et al. 2003; Rasheed et al. 2013). In literature, various kinds of diversity indices, such as Simpson (1949), Shannon and Weaver (1949) and Margalef (1958), have been suggested. The data collected either in binary form (i.e., presence or absence of species at a study site) or in a quantitative form, which contain many zero values for absent species, are required for the calculation of the indices. A meaningful discussion on the mathematical formulation of different indices and their grouping as classical, effective numbers, similarity sensitive and parametric families can be found elsewhere (Daly et al. 2018). Moreover, all these indices comprise certain strengths and may suffer from some kinds of constraints as well. Mathematical expression and parameter details of some useful biodiversity indices are listed in Table 2.
Species richness is the measure of the number of species present in a community and does not emphasize the number of individuals of a species present in the community. With the involvement of spatial diversity, species richness is regarded as the key measure of biodiversity (Elo et al. 2018). The most commonly used species richness indices are Margalef’s and Menhinick’s. These indices are easy to calculate and have a direct relationship with the number of species and sample size (Magurran 2004). The total number of species generally increases with increasing the sampling area. Nevertheless, species richness is the simplest index and is still being used by ecologists as a measure of diversity even though it does not throw any light on the relative abundance of documented species. Species abundance distribution can give an insight into the processes that decide the biological diversity of the communities. It reflects the competition for limiting resources among species (Magurran 2004). A precise study of temporal and spatial changes in a community should be done as it can provide information regarding variations in species abundance. Certain models such as the log-normal, the log-series, the broken-stick model and the geometric-series are used by researchers for such purposes (Tokeshi 1993; Hill et al. 2003). However, these models have been generally applied to higher plant communities and are rarely explored for microbial systems like cyanobacterial and microalgal communities.
Shannon’s index is the most commonly used measure, for the estimation of ecological diversity (Tandon et al. 2007; Pandey and Kulkarni 2006). It is a mathematical measurement to define community composition, i.e., the number of species and commonness of species in a community. It measures the degree of uncertainty in predicting the species of a random individual from a community with S species and N individuals. It is highly regulated by rare species and species richness. Since this index is susceptible to slight variations in diversity representing the actual state of the environment, it is preferred over the other available indices. Yadav et al. (2018) used the Shannon diversity index for evaluating the effect of nutrient enrichment on the species composition of periphytic algal communities colonizing chemical diffusing substrates. Likewise, Ikram’s group successfully employed Shannon diversity to study changes in microalgal and cyanobacterial communities along the gradients of temperature and other physicochemical factors in two hot springs of Garhwal Himalaya, India (Ikram et al. 2021a). These latter authors showed that the Shannon diversity decreased considerably as water temperature exceeded 50 °C in the studied hot springs.
The other important index to measure biodiversity is the Simpson index. This particular index attaches importance to the evenness of common species and picks up the species that are dominant or eminent in the community (Simpson 1949). However, as higher values indicate lower diversity, this index is not considered a very natural measure of biodiversity. The reciprocal form of Simpson original index measures evenness but suffers from the constraint that the index varies with the species richness. Gini-Simpson’s diversity index, also known as the probability interspecific encounter, gets an upper hand among various indices derived from Simpson’s original index as it is less sensitive to species richness and emphasizes the most abundant species in a community (Daly et al. 2018).
Biodiversity between communities
The functioning of ecosystems for the conservation of biodiversity and ecosystem management can be better understood by measuring beta diversity. Beta diversity represents the dissimilarity in species composition between sites in a landscape or geographical region (Whittaker 1960). Beta diversity can be estimated by computing diversity indices for each site and testing hypotheses about the environmental factors which may offer a suitable explanation for the variations existing among sites. An alternative approach may involve a direct analysis of the community composition data over the study sites concerning the sets of environmental and spatial variables (Legendre et al. 2005). The statistical methods of partitioning the variation of the diversity indices or the community composition data to the environmental and spatial variables are very useful for accomplishing such tasks (Peres-Neto et al. 2006). Bray Curtis dissimilarity and Jaccard’s index are two popular statistic-based tools, which are used by researchers for comparing diversity between two communities (Schroeder and Jenkins 2018). Moreover, these concepts have sporadically been explored for diversity assessment of microalgae and cyanobacteria.
Diversity of a landscape
Ecologists term the overall species richness of a landscape or geographical area as gamma diversity. According to Whittaker (1972), alpha and beta diversity are the two independent components of it. However, the modern ecologists prefer to use the term landscape diversity that not merely includes total species richness but also takes into account the patch diversity, such as patch number, patch shape, landscape fragmentation, patch edge, and diverse functional aspects of inhabiting species (Bojie and Liding 1996). Thus, landscape diversity involves a holistic approach for exploring biodiversity and ecosystem functioning of a geographical area. It is an imperative concept for restoring and maintaining the sustainable and resilient features of agriculture landscapes (Schaller et al. 2018). The microalgal and cyanobacterial diversity, which has been hitherto ignored by plant ecologists despite its valuable ecological functions, need to be given due emphasis in any program aimed at measuring landscape diversity.
Zeta diversity
Beta diversity, whether derived through the multiplicative or additive partitioning approach, is commonly used by researchers to understand similarity in species composition of two different sites. However, beta diversity does not present the holistic view regarding the actual pattern of diversity if the study area involves multiple sites. Therefore, some other kinds of relationships like species-area curve and interspecific distribution and rarity and endemism pattern are required to understand the phenomenon in totality (Gaston and Blackburn 2000; McGill 2010). This diversity measure determines the total set of biodiversity ingredients and systematically provides the spatial distribution of multispecies groups. It simultaneously provides information regarding the species-area relationship, multispecies dwelling patterns and ranking of species endemism. The exponential and power-law expressions of zeta diversity are also capable of deducing the information regarding niche assembly processes. Thus, zeta diversity is regarded as a pertinent measure for providing all-inclusive insights into biodiversity distribution patterns and the processes that regulate them and their response to the changing environmental factors (Hui et al. 2014, 2018). However, this measure of diversity has received meagre attention from researchers working in the area of microbial ecology.
Useful statistical tools
Sampling methods and collection of basic data
The collection of data relating to the abundance of various species is essential for calculating all kinds of biodiversity indices. Such data provide primary information of the community under the study. Since all kinds of further analyses are based on data collected from sampling, it must be done with utmost care. The size and strategical procedure of sample collection need to be decided very carefully keeping in view the statistical concepts and apparent features of the study area. Of the different sampling techniques prescribed by statisticians, such as simple random sampling, stratified sampling, cluster sampling, multi-stage sampling, etc., the most suitable one can be selected.
During the biodiversity estimation of higher plants in forests, grasslands or shrublands, the widely used sampling approaches are quadrat, transect, and plotless methods (Misra 1968; Kershaw 1973; Elzinga et al. 2001). However, a similar standardization of sampling procedure is lacking for microalgae and cyanobacteria. A majority of studies focusing on the biodiversity of microalgae and cyanobacteria do not appropriately describe the sampling procedure used. Some researchers have used a quadrat size of 400 cm2 for sampling microalgae from thermal springs (Sompong et al. 2005), while others have advocated using 100 cm2 size, without mentioning any valid reason for this choice (Ikram et al. 2021a). If we consider the micrscopic size of cyanobacteria and microalgae, the above-mentioned quadrat sizes could be considered unreasonably large. But the quadrat size should neither be very large nor very small. While the former makes the study tiresome, the latter may provide imprecise results.
For estimating relative abundance and calculating various diversity indices, the primary requisite is to count the number of individuals of a species present in the sampled quadrats. In the case of microalgae and cyanobacteria, the counting of individuals can be done with the help of a haemocytometer or any other similar device. However, it is not as simple as we think. Due to the unicellular and multi-cellular morphology of microalgal and cyanobacterial forms, deciding individual representation often becomes difficult. In the case of unicellular algae and diatoms, each cell can be taken as an individual unit. However, this practice can not be applied per se for the large filamentous algal forms. Earlier researchers preferred to count each cell of a filament as a unit. But then the variable length of filaments as also their curvature create trouble in haemocytometer-based counting. The statistical tools can play an important role in standardizing such procedures. Lawton et al. (1999) and Olson (1950) suggested some statistical corrections that should be taken into account during counting filamentous algal forms. A definite length of the filament is considered as a unit for such small filamentous cyanobacterial forms in which septa are hardly visible (DeNicola et al. 2006; Passy and Larson 2011; Yadav et al. 2018). The colonial and aggregate forming taxa also create difficulties during counting individual representation in the community. Some researchers considered a specified area as a representation of an algal cell unit. However, such methodology cannot be considered as a true representation of individual share in the community and hence needs biological and statistical justification before generalization.
Sometimes, a very small amount of microalgal and cyanobacterial samples may comprise a large number of cells and it virtually becomes difficult to count cells under the microscope. Researchers generally dilute the samples to overcome such problems. Depending on the density of algal cells, some researchers counted 100 to 500 cells per sample (Yadav et al. 2018). But, it is an intuitional choice and the minimum number of counting of algal cells that could be reasonable and appropriate for representing the share of an individual species in the community should be justified statistically. A variety of statistical tools are available to help in this context. Generally, it is expected to have enough cell counts so that the standard error of data remains < 10% (Gotelli and Ellison 2004).
Plotless or distance-based sampling techniques, such as point-centred quarter, nearest neighbours and closest individual methods have been thoroughly worked out in the case of higher plants (Elzinga and Salzer 1998; Hijbeek et al. 2013). These methods are generally applied in forests but can be used in grasslands and shrublands as well. These techniques are used to estimate the density and distribution of plants considering the average space occupied by an individual in the study area. Plotless techniques comprise several advantages over quadrat-based sampling. It is usually prompt, need less equipment and does not involve determination or adjustment in quadrat size and numbers. However, these techniques have hardly been employed or adequately modified and optimized in exploring the microbial diversity of soil ecosystems.
Detecting outliers and errors in collected data
Outliers are recorded values of measurements or observations that are outside the range of the bulk of the data (Gotelli and Ellison 2004). On the other hand, errors are recorded values that do not represent the original measurements or observations (Gotelli and Ellison 2004; Osborne and Overbay 2004). Some, but not all, errors are also outliers. Conversely, not all outliers in a dataset are errors. Detection of errors and outliers in the biodiversity data set is another important task of an ecologist as they can have a prominent influence on the results of statistical tests by increasing variance in the data (Gotelli and Ellison 2004). Some researchers consider outliers as noise, but outliers may be more than that in the case of biodiversity-based studies. They can reflect the key biological functions of species in an ecosystem and sincere thought over their presence may lead to new hypotheses, ideas, or discovery of an entirely new species. In some cases, a few data values appear outliers just because of the forced normalization of the dataset, but the appropriate transformation of data can be used to resolve such issues. Three simple techniques, calculating column statistics, checking ranges and precision of column values, and graphical exploration of data can be employed for this purpose. The simple column statistics is a straightforward way to find out the unusual low or high values in the spreadsheet data. The measurements of simple statistical parameters like mean, median, standard deviation, and variance provide a quick overview of the range of the values in the data set. The suspicious minimum and maximum values can easily be identified. Most of the spreadsheet software packages comprise functions that calculate these values. Further, the spreadsheet functions can also be used to check that all the data values in a column are within reasonable boundaries. Graphical exploratory data analysis (Graphical EDA) is another way to hunt for outliers and errors (Gotelli and Ellison 2004). Three types of graphs namely (i) box plots, (ii) stem-and-leaf plots and (iii) scatter plots are very popular to find out the unexpected trends or patterns in the data sets. The first two are used for the plotting of a single variable. On the other hand, scatter plots are used for bivariate or multivariate data.
Transformation of biodiversity data
When the purpose is to know how the different environmental factors influence the distribution of various genera and species inhabiting a community or landscape along space and time, we need to normalize the whole data set for analyses. The transformation makes it possible and converts the data in such a form that becomes more understandable, communicable and appropriate for meeting the assumptions. For transforming data, a mathematical function is simply applied to all the observations of a particular variable (Gotelli and Ellison 2004; Legendre and Legendre 2012). Most transformations comprise simple algebraic continuous monotonic functions. This valuable tool is also important for exploring variations in species composition of microalgal and cyanobacterial communities along the gradients of diverse physicochemical factors (Sompong et al. 2005; Ikram et al. 2021a, b). Because of the involvement of monotonic functions, transformation does not change the rank order of the data but does affect the variance and shape of the probability distribution. Transformations are often used to convert non-linear relationships into linear relationships as they are more comprehensible. The logarithmic, square-root, reciprocal, Box-Cox are examples of some other useful transformations.
Analysis of biodiversity data
Regression
Regression is a powerful tool in studying and modelling the spatial distribution of species in relation to various environmental factors (Gotelli and Ellison 2004). The most basic regression describes the linear relationship between an independent variable and a dependent variable. From a statistician’s point of view, regression and correlation are different tools as the former reveals the association between dependent and independent variables based on cause-and-effect relationship, while the latter comprises variables that are found merely associated without any such cause-and-effect. Although different models in statistics have been developed for regression and correlation, some researchers consider that the distinction is arbitrary and often just semantic. Moreover, environmentalists do not pursue correlations between variables unless they think or suspect a certain kind of cause-and-effect relationship. The non-parametric extension and several other kinds of modifications of classical regression are widely used in biodiversity studies for developing a variety of ecological models for different responses, such as species richness, abundance classes, and presence-absence data (Lehmann et al. 2002).
Multivariate analysis
Multivariate techniques are used for studying the relationship between the distribution pattern of species in relation to environmental parameters. For this, the similarity coefficients are calculated and subsequently data are classified by clustering or mapped into two- or three-dimensional plots, known as ordination plots. The ordination plots represent the relative dissimilarity of species composition. The arrangement of data of samples in these plots is positioned on the distances between the pair of samples of the communities. The points placed near each other on an ordination map are said to have similar communities. Attempts have been made to obtain information about the species-environment relations from the data of the communities obtained from the field surveys by ecologists (ter Braak 1988). The data of biological communities always show a skewed distribution. And the non-linear relationship is present between the environmental variables and the species. A unimodal function of the environmental variables is observed for species in a community (Whittaker 1956, 1967). The clustering and ordination techniques are used to summarize the multi-species data to similar clusters or ordination axes and interpreted according to the known data about the environment and the species of the area under study (ter Braak 1988). The interpretation of the relationship between the species and the environment by cluster and ordination techniques has been termed as indirect gradient analysis by Whittaker (1967). Hence, a technique of direct gradient analysis was put forward by Whittaker (1967), which is also known as regression, to describe the relationship of environmental variables with the species.
The common multivariate techniques used by ecologists are ANOVA, ANOSIM, cluster analysis, principal component analysis, canonical correspondence analysis, multidimensional scaling. Moreover, multiple correlations could also be applied to find out the interdependency of different environment variables.
Analysis of variance (ANOVA)
ANOVA is used to test the difference between the means of the studied samples. For ecological studies, it checks the null hypothesis of no difference in mean of diversity between the two sites (Gotelli and Ellison 2004). The result is said to be statistically significant when the p-value (probability) is less than the significance level (usually taken as 0.05) or 95% confidence level. The null hypothesis is rejected denoting differences in the mean of diversity between the sites. But the ANOVA test is not preferred much for testing the difference between the sites or samples because of the intricacy of reliance between species and the general ineptness of the normality assumption. When species data contains a large number of zero values, its transformation for getting normality is not possible. In such circumstances, other multivariate techniques are applied for the interpretation of data as also to determine the relationship of species with environmental parameters.
Analysis of similarities (ANOSIM)
It is a non-parametric permutation technique, which has been aptly described by Clarke and Green (1988). It is employed to the (rank) similarity matrix underlying the ordination or classification of samples. It is a more valid and informative test as a large number of replicates and permutations are made in it. Interpretations are drawn if there are significant differences between the groups. The R-value is observed for each pair-wise comparison. The pairwise test between the samples gives a p-value which denotes how significantly samples are different, and the R-value shows how strongly they are different from each other. The indicator R* shows complete dissimilarity in communities when equal to 1, while reveals a close similarity if values tend to be 0.
Multidimensional scaling (MDS)
This technique is based on the similarity or dissimilarity between the samples. The non-metric MDS ordination is a visual display of the pattern of proximities. It utilizes the rank of similarity to display the samples in the plot. The most similar samples are placed together in the ordination plot, while the widely apart samples reflect the variations among them. The goodness of fit is known as stress in the case of MDS. The stress value ranges between 0 and 1. The stress value near zero represents a good fit of the model for MDS. The MDS is used by the ecologists to illustrate the similarities between the samples in a smaller number of dimensions.
Cluster analysis
The cluster analysis is the method employed for presenting variance amongst the communities and the samples. It intends to detect the “natural grouping” of samples based on their similarity to each other. When comparing different sites (or subsites of sites), the similarity matrix of species is maneuvered to elucidate the species that analogously co-occur across the sites or subsites (Fig. 1). The hierarchical agglomerative method is the most commonly used technique of clustering. The data or the samples are fused to form clusters based on a similarity matrix (such as Bray-Curtis), till a single cluster is formed. It is generally represented by a dendrogram (i.e., tree diagram) (Clarke and Warwick 2001).
Principal component analysis (PCA)
PCA is used to decrease the number of factors from the sample and identify the significant one from the big data pool of samples. It focuses on conserving as much data as possible while diminishing the multi-dimension data to lower dimensions. The orthogonal transformations are used to convert the feasibly correlated variables into linearly uncorrelated variables. These linearly uncorrelated variables are known as “principal components” that record for the most variance in the sample. The first principal component represents the highest possible variance. The ordination plot of PCA represents how close/profoundly associated two factors are. The transformed data (using eigenvectors) are used by PCA as number in the ordination plot rather than the real data. Because by plotting the real data, the relationship and the pattern between the points cannot be interpreted (Clarke and Warwick 2001). PCA is a more apt test for environmental variables as ecological data (specifically in the case of microbes) have more zero counts that do not need any special treatment (data normality test).
Canonical correspondence analysis (CCA)
CCA is used when there are a large number of species present in the community and a great intrinsic variability may prevail in the system. The ecological data are either quantitative (abundance i.e., number of individuals) or incidence types (presence/absence), and the species to environment relationship is non-linear and non-monotonic. Thus, in the light of such characteristics, CCA is more appropriate than other traditional linear-based multivariate techniques (ter Braak and Verdonschot 1995). This method helps ecologists to decipher the response of species to the environmental variables or distribution patterns along the environmental gradients. CCA can also be used for examining spatial and seasonal disparities in the communities (Snoeijs and Prentice 1989; Bakker et al. 1990; Anderson et al. 1994). In the biplot of CCA (Fig. 2), arrows indicate the quantitative environmental variables. The length of the arrow indicates variable importance and their positive or negative association with the axis (Abrantes et al. 2006). The angle between vectors indicates a correlation among environmental variables. The locations of points (sites or species) in the plot represent their compositional similarity to each other and are dominated by species that are projected near them in the CCA plot. The location of species indicates their distributional similarity to each other.
Multiple correlations
It is a statistical technique applied to estimate the correlation and interdependency of different environmental parameters. The effect of different factors on one factor and the strength of the relationship between them can be inferred by the multiple correlation method. A strong correlation represents a prominent effect of different factors on a single factor. Conversely, a poor correlation reveals that the effect of other factors on the factor under consideration is unimportant.
Software packages
Nowadays, a variety of statistical software packages are available, namely, ANALYTICA, IBM-SPSS, STATISTICA, STATA, SIGMA PLOT, MATLAB, OriginPro, XLSTAT, R package, BIOTA, CANOCO, PAleontological STatistics (PAST) and PC-ORD. Of these, some packages are exclusively used for mathematical and statistical operations, while others are meant for biodiversity assessment. Researchers have employed different software for determining microalgal and cyanobacterial diversity (Omelon et al. 2007; Barinova et al. 2011; Zhan and Sun 2012; Kühl et al. 2012; Roy et al. 2015; Schulz et al. 2016; Gaikwad et al. 2016; Mogul et al. 2017; Zhang et al. 2021). The development of software packages has made the use of statistical tools very easy and comfortable. Considerable caution needs to be exercised while using these software as ignorance of basic statistical concepts may lead to incorrect interpretation of the results. Ignorance of statistical concepts also leads to an unsound experimental design. Hence, care should be taken while employing statistical design or analyzing results through software packages. It would be still better if a proficient statistician is directly involved right from the time of designing the experiment. Some software packages are not user-friendly because of their complex ways of data feeding. Table 3 lists the advantages and drawbacks of some statistical packages. Since the selection of statistical software depends mainly on its user-friendly features, the focus should now be given to developing subject-specific statistical software packages for the assessment of microbial biodiversity.
Conclusions and future perspectives
Microalgae and cyanobacteria are extremely important for maintaining the vitality of the agroecosystems. However, they have remained a neglected component in biodiversity studies. The concept of functional diversity has several merits. However, it has not been effectively explored for microalgal and cyanobacterial communities. A majority of previous research in the field of microalgae and cyanobacteria is focused on identifying species and phylogenetic diversity. However, species concept and the criteria for taxonomic identification of a new species in microalgae and cyanobacteria is very confusing. Thus, this particular aspect demands precise and critical efforts. Landscape diversity assessment is an imperative proposition for restoring and maintaining the sustainable and resilient features of agriculture landscapes. Thus, any program developed to estimate landscape diversity needs sufficient attention to include work components for measuring the microalgal and cyanobacterial diversity. A majority of studies dealing with microalgal and cyanobacterial diversity do not appropriately describe the sampling procedure details. Detection of errors and outliers in the biodiversity data set is crucial to get a better insight of it and also for performing various statistical analyses. While regression and multiple correlations help in realizing the relations of different environmental factors, ANOVA, ANOSIM, MDS and cluster analysis are powerful techniques for understanding the compositional differences of different microbial communities. PCA and CCA are effective in interpreting the influence of environmental factors on the distribution of microalgal and cyanobacterial species in a study area. Statistical software packages are the backbone of the current research activities. Some of the software packages available are simplistic, while others incorporate operational complexities. Thus, before employing such software programmes in biodiversity-based investigations, gaining a sound understanding of them is strongly recommended. As the selection of statistical software depends mainly on its user-friendly features, it is high time to shift the major focus to developing biodiversity statistical packages specific to microorganisms.
Availability of data and materials
Not applicable.
References
Abdel-Raouf N, Al-Homaidan AA, Ibraheem IBM (2012) Agricultural importance of algae. Afr J Biotechnol 11(54):1648–11658
Abinandan S, Subashchandrabose SR, Venkateswarlu K, Megharaj M (2019) Soil microalgae and cyanobacteria: the biotechnological potential in the maintenance of soil fertility and health. Crit Rev Biotechnol 39(8):981–998
Abrantes N, Antunes SC, Pereiran MJ, Goncalves F (2006) Seasonal succession of cladocerans and phytoplankton and their interactions in a shallow eutrophic lake (Lake Vela, Portugal). Acta Oecol 29:54–64
Ahlesaadat M, Riahi H, Shariatmadari Z, Hakimi Meybodi MH (2017) A taxonomic study of cyanobacteria in wheat fields adjacent to industrial areas in Yazd province (Iran). Rostaniha 18(2):107–121
Ahmed M, Stal LJ, Hasnain S (2010) Association of non-heterocystous cyanobacteria with crop plants. Plant Soil 336(1):363–375
Akins LN, Ayayee P, Leff LG (2018) Composition and diversity of cyanobacteria-associated and free-living bacterial communities during cyanobacterial blooms. Ann Microbiol 68(8):493–503
Alghanmi HA, Jawad HM (2019) Effect of environmental factors on cyanobacteria richness in some agricultural soils. Geomicrobiol J 36(1):75–84
Allison SD, Martiny JB (2008) Resistance, resilience, and redundancy in microbial communities. Proc Natl Acad Sci 105(Supplement 1):11512–11519
Alvarez AL, Weyers SL, Goemann HM, Peyton BM, Gardner RD (2021) Microalgae, soil and plants: A critical review of microalgae as renewable resources for agriculture. Algal Res 54:102200
Ampe F, Miambi E (2000) Cluster analysis, richness and biodiversity indexes derived from denaturing gradient gel electrophoresis fingerprints of bacterial communities demonstrate that traditional maize fermentations are driven by the transformation process. Int J Food Microbiol 60(1):91–97
Anderson NJ, Korsman T, Renberg I (1994) Spatial heterogeneity of diatom stratigraphy in varved and non-varved sediments of a small, boreal-forest lake. Aquat Sci 56:40–58
Armitage DW, Gallagher KL, Youngblut ND, Buckley DH, Zinder SH (2012) Millimeter-scale patterns of phylogenetic and trait diversity in a salt marsh microbial mat. Front Microbiol 3:293
Ashraf MA, Asif M, Zaheer A, Malik A, Ali Q, Rasool M (2013) Plant growth promoting rhizobacteria and sustainable agriculture: a review. Afr J Microbiol Res 7(9):704–709
Badger MR, Price GD, Long BM, Woodger FJ (2006) The environmental plasticity and ecological genomics of the cyanobacterial CO2 concentrating mechanism. J Exp Bot 57:249–265
Bakker C, Herman RMJ, Vink M (1990) Changes in seasonal succession of phytoplankton induced by the storm-surge barrier in the Oosterschelde (SW Netherlands). J Plankton Res 12:947–972
Ballesteros I, Terán P, Guamán-Burneo C, González N, Cruz A, Castillejo P (2021) DNA barcoding approach to characterize microalgae isolated from freshwater systems in Ecuador. Neotropical Biodivers 7(1):170–183
Banerjee S, Misra A, Sar A, Pal S, Chaudhury S, Dam B (2020) Poor nutrient availability in opencast coalmine influences microbial community composition and diversity in exposed and underground soil profiles. App Soil Ecol 152:103544
Barinova S, Kukhaleishvili L, Nevo E, Janelidze Z (2011) Diversity and ecology of algae in the Algeti National Park as a part of the Georgian system of protected areas. Turk J Botany 35(6):729–774
Berger WH, Parker FL (1970) Diversity of planktonic Foraminifera in deep sea sediments, vol 168. Science, pp 1345–1347
Bharadwaj N, Baruah PP (2013) Diversity and abundance of N2-fixing cyanobacterial population in rice field soil crusts of Lower Brahmaputra Valley agro-climatic zone. J Algal Biomass Util 4:23–33
Bodor A, Bounedjoum N, Vincze GE, Kis ÁE, Laczi K, Bende, Szilaǵyi Á, Kovács T, Perei K, Rákhely G (2020) Challenges of unculturable bacteria: environmental perspectives. Rev Environ Sci Biotechnol 19(1):1–22
Bojie F, Liding C (1996) Landscape diversity types and their ecological significance. Di li xue bao 51(5):454–462
Chao A, Gotelli NJ, Hsieh TC, Sander EL, Ma KH, Colwell RK, Ellison AM (2014) Rarefaction and extrapolation with Hill numbers: a framework for sampling and estimation in species diversity studies. Ecol Monogr 84(1):45–67
Chaudhary V, Prasanna R, Bhatnagar AK (2013) Influence of phosphorus and pH on the fungicidal potential of Anabaena strains. J Basic Microbiol 53(3):201–213
Chaurasia A (2015) Cyanobacterial biodiversity and associated ecosystem services: introduction to the special issue. Biodivers Conserv 24:707–710
Chiarucci A, Bacaro G, SM Scheiner (2011) Old and new challenges in using species diversity for assessing biodiversity. Philos Trans R Soc Lond B Biol Sci 366(1576):2426–2437
Chittora D, Meena M, Barupal T, Swapnil P, Sharma K (2020) Cyanobacteria as a source of biofertilizers for sustainable agriculture. Biochem Biophys Rep 22:100737
Chunleuchanon S, Sooksawang A, Teaumroong N, Boonkerd N (2003) Diversity of nitrogen-fixing cyanobacteria under various ecosystems of Thailand: population dynamics as affected by environmental factors. World J Microbiol Biotechnol 19(2):167–173
Clarke KR, Green RH (1988) Statistical design and analysis for a ‘biological effects’ study. Mar Ecol Prog Ser 46:213–226
Clarke KR, Warwick RM (2001) Change in marine communities: an approach to statistical analysis and interpretation, 2 edn. Plymouth Marine laboratory, UK: PRIMER-E Ltd
Contoli L, Luiselli L (2015) Contributions to biodiversity theory: the importance of formal rigor. Web Ecol 15:33–37
Costanza R, Fisher B, Mulder K, Liu S, Christopher T (2007) Biodiversity and ecosystem services: A multi-scale empirical study of the relationship between species richness and net primary production. Ecol Econ 61(2–3):478–491
Cox G (1990) Laboratory manual of general ecology. In Brown WC, 6th edn. Dubuque, Iowa
Daly AJ, Baetens JM, De Baets B (2018) Ecological diversity: measuring the unmeasurable. Mathematics 6(7):119
Dash MC (2001) Fundamentals of Ecology. Tata McGraw-Hill Publication, p 525
DeNicola DM, Eyto ED, Wemaere A, Irvine K (2006) Periphyton response to nutrient addition in three lakes of different benthic productivity. J North Am Benthol Soc 25(3):616–631
Dey HS, Tayung K, Bastia AK (2010) Occurrence of nitrogen-fixing cyanobacteria in local rice fields of Orissa, India. Ecoprint: An Int J Ecol 17:77–85
Dvořák P, Poulíčková A, Hašler P, Belli M, Casamatta DA, Papini A (2015) Species concepts and speciation factors in cyanobacteria, with connection to the problems of diversity and classification. Biodivers Conserv 24(4):739–757
Eckert EM, Fontaneto D, Coci M, Callieri C (2015) Does a barcoding gap exist in prokaryotes? Evidences from species delimitation in cyanobacteria. Life 5:50–64
Ehrlich P, Ehrlich A (1981) Extinction:the causes and consequences of the disappearance of species
El-Bestawy EA, Abd El-Salam, Mansy HAR (2007) Potential use of environmental cyanobacterial species in bioremediation of lindane contaminated effluents. Int Biodeterior Biodegradation 59:180–192
Elo M, Alahuhta J, Kanninen A, Meissner KK, Seppälä K, Mönkkönen M (2018) Environmental characteristics and anthropogenic impact jointly modify aquatic macrophyte species diversity. Front Plant Sci 9:1001
Elzinga CL, Salzer DW, Willoughby JW, Gibbs JP (2001) Monitoring plant and animal populations. Blackwell Science, Malden, MA, USA, p 360
Elzinga CL, Salzer DW (1998) Measuring & monitoring plant populations. US Department of the Interior, Bureau of Land Management
Emerson JB, Adams RI, Román CMB, Brooks B, Coil DA, Rothschild LJ (2017) Schrödinger’s microbes: tools for distinguishing the living from the dead in microbial ecosystems. Microbiome 5(1):86
Fattorini L (2003) Statistical analysis of ecological diversity. In: El-Shaarawi AH, Jureckova J (ed) Environmetrics 18–29
Fioravante IA, Barbosa FAR, Augustic R, Magalhães SMS (2010) Removal of methyl parathion by cyanobacteria Microcystis novacekii under culture conditions. J Environ Monit 12:1302–1306
Gaikwad S, Shouche YS, Gade WN (2016) Microbial community structure of two freshwater sponges using Illumina MiSeq sequencing revealed high microbial diversity. AMB Express 6(1):1–9
Gaston KJ, Blackburn TM (2000) Pattern and process in macroecology. Blackwell Science, Oxford
Ghadage SJ, Karande VC (2019) The distribution of blue-green algae (Cyanobacteria) from the paddy fields of Patan and Karad tehsils of Satara District, Maharashtra, India. J Threat Taxa 11(14):14862–14869
Ghosh TK, Saha KC (1997) Effects of inoculation of cyanobacteria on nitrogen status and nutrition of rice (Oryza sativa L.) in an entisol amended with chemical and organic sources of nitrogen. Biol Fertil Soils 24:123–128
Goswami M, Bhattacharyya P, Mukherjee I, Tribedi P (2017) Functional Diversity: An Important Measure of Ecosystem Functioning. Adv Microbiol 7:82–93
Gotelli NJ, Ellison AM (2004) A primer of ecological statistics, vol 1. Sinauer Associates, Sunderland
Guisan A, Edwards TC Jr, Hastie T (2002) Generalized linear and generalized additive models in studies of species distributions: setting the scene. Ecol modell 157(2–3):89–100
Gupta V, Prasanna R, Srivastava AK, Sharma J (2012) Purification and characterization of a novel antifungal endo-type chitosanase from Anabaena fertilissima. Ann Microbiol 62:1089–1092
Gupta V, Ratha SK, Sood A, Chaudhary V, Prasanna R (2013) New insights into the biodiversity and applications of cyanobacteria (blue-green algae)—prospects and challenges. Algal res 2(2):79–97
Hadi SIIA, Santana H, Brunale PPM, Gomes TG, Oliveira MD, Brasil BSAF (2016) DNA barcoding green microalgae isolated from neotropical inland waters. PLoS ONE 11(2):e0149284
Hashtroudi MS, Ghassempour A, Riahi H, Shariatmadari Z, Khanjir M (2013) Endogenous auxins in plant growth-promoting cyanobacteria Anabaena vaginicola and Nostoc calcicole. J Appl Phycol 25:379–386
Hauer T, Bohunická M, Johansen JR, Mareš J, Berrendero-Gomez E (2014) Reassessment of the cyanobacterial family Microchaetaceae and establishment of new families Tolypothrichaceae and Godleyaceae. J Phycol 50(6):1089–1100
Hendrayanti D, Khoiriyah I, Fadilah N, Salamah A (2018) Diversity of N2-fixing cyanobacteria in organic rice field during the cycle of rice crops. In AIP Conference Proceedings (Vol. 2002: No. 1, p. 020011). AIP Publishing LLC
Hijbeek R, Koedam N, Khan MNI, Kairo JG, Schoukens J, Dahdouh-Guebas F (2013) An evaluation of plotless sampling using vegetation simulations and field data from a mangrove forest. PLoS ONE 8(6):e67201
Hill TC, Walsh KA, Harris JA, Moffett BF (2003) Using ecological diversity measures with bacterial communities. FEMS Microbiol Ecol 43(1):1–11
Hoffmann L, Komárek J, Kaštovský J (2005) System of cyanoprokaryotes (cyanobacteria) state in 2004. Arch Hydrobiol Suppl Algol Stud 117:95–115
Hui C, McGeoch MA (2014) Zeta diversity as a concept and metric that unifies incidence-based biodiversity patterns. Am Nat 184:684–694
Hui C, Vermeulen W, Durrheim G (2018) Quantifying multiple-site compositional turnover in an Afrotemperate forest, using zeta diversity. For Ecosyst 5(1):1–9
Hussain A, Hasnain S (2011) Phytostimulation and biofertilization in wheat by cyanobacteria. J Ind Microbiol Biotechnol 38:85–92
Ikram SF, Kumar D, Singh V, Tripathi BN, Kim BH (2021a) Microalgal and cyanobacterial diversity of two selected hot springs of Garhwal Himalaya, Uttarakhand, India. Fundam Appl Limnol 111–127
Ikram SF, Uniyal V, Kumar D (2021b) Changes in species composition of cyanobacterial and microalgal communities along a temperature gradient in Tapovan Hot Spring, Garhwal Himalaya, Uttarakhand, India. Aquat Ecol 1–12
Irissari P, Gonnet S, Monza J (2001) Cyanobacteria in Uruguayan rice fields: diversity, nitrogen fixing ability and tolerance to herbicides and combined nitrogen. J Biotechnol 91(2–3):95–103
Jost L, Chao A (2008) Diversity Analysis. Taylor & Francis, Milton Keynes, UK
Karthikeyan N, Prasanna R, Sood A, Jaiswal P, Nayak S, Kaushik BD (2009) Physiologicl characterization and electron microscopic investigations of cyanobacteria associated with wheat rhizosphere. Folia Microbiol 54:43–51
Kershaw KA (1973) Quantitative and Dynamic Plant Ecology, 2nd edn. Edward Arnold, London, p 308
Khare P, Singh A, Prabha C, Kumari S (2014) Study of cyanobacterial biodiversity in rice fields of central Bihar. In 3rd World Conference on Applied Science Engineering and Technology, Kathmandu, Nepal
Komárek J (2006) Cyanobacterial taxonomy: current problems and prospects for the integration of traditional and molecular approaches. Algae 21(4):349–375
Komárek J (2016) A polyphasic approach for the taxonomy of cyanobacteria: principles and applications. Eur J Phycol 51(3):346–353
Komárek J, Kaštovský J (2003) Coincidences of structural and molecular characters in evolutionary lines of cyanobacteria. Algol Stud 148:305–325
Kühl M, Behrendt L, Trampe ECLE, Qvortrup K, Schreiber U, Borisov SM, Larkum AW (2012) Microenvironmental ecology of the chlorophyll b-containing symbiotic cyanobacterium Prochloron in the didemnid ascidian Lissoclinum patella. Front Microbiol 3:402
Kushwaha M, Marwa N, Pandey V, Singh N (2020) Advanced Tools to Assess Microbial Diversity and Their Functions in Restoration of Degraded Ecosystems. In: Singh JS, Vimal SR (ed) Microbial Services in Restoration Ecology. Elsevier, pp 83–97
Laureto LMO, Cianciaruso MV, Samia DSM (2015) Functional diversity: an overview of its history and applicability. Nat Conserv 13(2):112–116
Lawton JH (1994) What do species do in ecosystems? Oikos 367–374
Lawton L, Marsalek B, Padisa J, Chorus I (1999) Determination of cyanobacteria in the laboratory. In: Chorus I, Bartram J (ed) Toxic cyanobacteria in water: A guide to their public health consequences, monitoring and management, WHO, pp 334–361
Lear G, Lau K, Perchec AM, Buckley HL, Case BS, Neale M, Fierer N, Leff JW, Kim M, Handley KM, Lewis G (2017) Following Rapoport’s Rule: the geographic range and genome size of bacterial taxa decline at warmer latitudes. Environ Microbiol 19(8):3152–3162
Legendre P, Borcard D, Peres-Neto PR (2005) Analyzing beta diversity: partitioning the spatial variation of community composition data. Ecol Monogr 75(4):435–450
Legendre P, Legendre L (2012) Numerical Ecology, 3rd English edn. Elsevier Science BV, Amsterdam
Lehmann A, Overton JM, Leathwick JR (2002) GRASP: generalized regression analysis and spatial prediction. Ecol Model 157(2–3):189–207
Mader P, Kaiser F, Adholeya A, Singh R, Uppal HS, Sharma AK, Liried PM (2011) Inoculation of root microorganisms for sustainable wheat rice and wheat black gram rotations in India. Soil Biol Biochem 43:609–619
Magurran AE (2004) Measuring biological diversity. Blackwell Science Ltd
Malavasi V, Soru S, Cao G (2020) Extremophile Microalgae: the potential for biotechnological application. J Phycol 56(3):559–573
Manjunath M, Prasanna R, Lata PD, Singh R, Kumar A, Jaggi S, Kaushik BD (2010) Biocontrol potential of cyanobacterial metabolites against damping off disease caused by Pythium aphanidermatum in solanaceous vegetables. Arch Phytopathol Pflanzenschutz 43:666–677
Margalef R (1958) Temporal succession and spatial heterogeneity in phytoplankton. In: Buzzati T (ed) Perspectives in Marine biology. University of California Press, Berkeley, pp 323–347
Mason NWH, Mouillot D (2013) Functional Diversity Measures. In: Levin SA (ed) Encyclopedia of Biodiversity, vol 3, 2nd edn. Waltham MA, Academic Press, pp 597–608
May RM (1975) Patterns of species abundance and diversity. In: Cody ML, Diamond JM (eds) Ecology and evolution of communities. Harvard University Press, Cambridge, MA, pp 81–120
Mazhar S, Hasnain S (2011) Screening of native plant growth promoting cyanobacteria and their impact on Triticum aestivum var. Uqab 2000 growth. Afr J Agric Res 5:3988–3993
McGill BJ (2010) Towards a unification of unified theories of biodiversity. Ecol Lett 13:627–642
McIntosh RP (1967) An index of diversity and the relation of certain concepts to diversity. Ecology 48:392–404
Misra R (1968) Ecology Work Book. Oxford and IBH Publication, New Delhi, p 244
Mogul R, Vaishampayan P, Bashir M, McKay CP, Schubert K, Bornaccorsi R, Wilhelm MB (2017) Microbial community and biochemical dynamics of biological soil crusts across a gradient of surface coverage in the Central Mojave Desert. Front Microbiol 8:1974
Nabout JC, da Silva Rocha B, Carneiro FM, Sant’Anna CL (2013) How many species of Cyanobacteria are there? Using a discovery curve to predict the species number. Biodivers Conserv 22(12):2907–2918
Najdenski HM, Gigova LG, Iliev II, Pilarski PS, Lukavsky J, Tsvetkova IV, Ninova MS, Kussovski VK (2013) Antibacterial and antifungal activities of selected microalgae and cyanobacteria. Int J Food Sci Technol 48:1533–1540
Natarajan C, Prasanna R, Gupta V, Dureja P, Nain L (2012) Dissecting the fungicidal activity of Calothrix elenkinii using chemical analyses and microscopy. Appl Biochem Microbiol 48:51–57
O’Brien SL, Gibbons SM, Owens SM, Hampton-Marcell J, Johnston ER, Jastrow JD, Gilbert JA, Meyer F, Antonopoulos DA (2016) Spatial scale drives patterns in soil bacterial diversity. Environ Microbiol 18(6):2039–2051
Oliveira BG, Mendes LW, Smyth EM, Tsai SM, Feigl BJ, Mackie RI (2020) Assessment of microbial diversity associated with CH4 emission from sugarcane vinasse storage and transportation systems. J Environ Manage 269:110748
Olson FCW (1950) Quantitative estimates of filamentous algae. Trans Am Micros Soc 59:272–279
Omelon CR, Pollard WH, Ferris FG (2007) Inorganic species distribution and microbial diversity within high Arctic cryptoendolithic habitats. Microb Ecol 54(4):740–752
Osborne JW, Overbay A (2004) The power of outliers (and why researchers should always check for them). Pract Assess Res Eval 9(1):6
Pagliarella MC, Corona P, Fattorini L (2018) Spatially-balanced sampling versus unbalanced stratified sampling for assessing forest change: evidences in favour of spatial balance. Environ Ecol Stat 25(1):111–123
Palinska KA, Surosz W (2014) Taxonomy of cyanobacteria: a contribution to consensus approach. Hydrobiologia 740:1–11
Pandey BN, Kulkarni GK (2006) Biodiversity and environment. Nangia SB. APH Publishing Corporation
Passy SI, Larson CA (2011) Succession in stream biofilms is an environmentally driven gradient of stress tolerance. Microb Ecol 62(2):414–424
Pereira I, Ortega R, Barrientos L, Moya M, Reyes G, Kramm V (2009) Development of a biofertilizer based on filamentous nitrogen-fixing cyanobacteria for rice crops in Chile. J Appl Phycol 21:135–144
Peres-Neto PR, Legendre P, Dray S, Borcard D (2006) Variation partitioning of species data matrices: estimation and comparison of fractions. Ecology 87(10):2614–2625
Petchey OL, Gaston KJ (2006) Functional diversity: back to basics and looking forward. Ecol Lett 9(6):741–758
Pielou EC (1977) Mathematical ecology (No. 574.50151 P613 1977). Wiley
Prasanna R, Chaudhary V, Gupta V, Babu S, Kumar A, Singh R, Shivay YS, Nain L (2013a) Cyanobacteria mediated plant growth promotion and bioprotection against Fusarium wilt in tomato. Eur J Plant Pathol 136:337–353
Prasanna R, Jaiswal P, Shrikrishna J, Joshi M, Nain L, Rana A, Shivay YS (2012) Evaluating the potential of rhizo-cyanobacteria as inoculants for rice and wheat. J Agric Sci Technol 8:157–171
Prasanna R, Nayak S (2007) Influence of diverse rice soil ecologies on cyanobacterial diversity and abundance. Wetl Ecol Manag 15(2):127–134
Prasanna R, Sharma E, Sharma P, Kumar A, Kumar R, Gupta V, Pal RK, Shivay YS, Nain L (2013b) Soil fertility and establishment potential of inoculated cyanobacteria in rice crop grown under non flooded conditions. Paddy Water Environ 11:175–183
Rai AN, Singh AK, Syiem MB (2019) Plant growth-promoting abilities in cyanobacteria. In: Mishra AK, Tiwari DN, Rai AN (ed) Cyanobacteria. Academic Press, pp 459–476
Rai PK, Rai A, Sharma NK, Singh S (2018) Study of soil cyanobacteria along a rural-urban gradient. Algal Res 35:142–151
Rasheed Z, Rangwala H, Barbará D (2013) 16S rRNA metagenome clustering and diversity estimation using locality sensitive hashing. BMC Syst Biol 7(4):1–13
Rejmánková E, Komárek J, Komárková J (2004) Cyanobacteria—a neglected component of biodiversity: patterns of species diversity in inland marshes of northern Belize (Central America). Divers Distrib 10(3):189–199
Renuka N, Guldhe A, Prasanna R, Singh P, Bux F (2018) Microalgae as multi-functional options in modern agriculture: current trends, prospects and challenges. Biotechnol Adv 36(4):1255–1273
Rippka R, Deruelles J, Waterbury JB, Herdman M, Stanier RY (1979) Generic assignments, strain histories and properties of pure cultures of cyanobacteria. Microbiology 111(1):1–61
Roy S, Bhattacharya S, Debnath M, Ray S (2015) Diversity of cyanobacterial flora of Bakreswar geothermal spring, West Bengal, India-II. Algol Stud 147:29–44
Schaller L, Targetti S, Villanueva AJ, Zasada I, Kantelhardt J, Viaggi D (2018) Agricultural landscapes, ecosystem services and regional competitiveness—Assessing drivers and mechanisms in nine European case study areas. Land Use Policy 76:735–745
Schleifer KH (2004) Microbial diversity: facts, problems and prospects. Syst Appl Microbiol 27(1):3
Schroeder PJ, Jenkins DG (2018) How robust are popular beta diversity indices to sampling error? Ecosphere 9(2):e02100
Schulz K, Mikhailyuk T, Dreßler M, Leinweber P, Karsten U (2016) Biological soil crusts from coastal dunes at the Baltic Sea: cyanobacterial and algal biodiversity and related soil properties. Microb Ecol 71(1):178–193
Sciuto K, Moro I (2015) Cyanobacteria: the bright and dark sides of a charming group. Biodivers Conserv 24(4):711–738
Shannon CE, Weaver W (1949) The Mathematical Theory of Communication. University of Illinois Press, Urbana
Sharma NK (2015) From natural to human-impacted ecosystems: rationale to investigate the impact of urbanization on cyanobacterial diversity in soils. Biodivers Conserv 24(4):1007–1015
Sharma R, Khokhar MK, Jat RL, Khandelwal SK (2012) Role of algae and cyanobacteria in sustainable agriculture system. Wudpecker J Agric Res 1(9):381–388
Shen C, Ni Y, Liang W, Wang J, Chu H (2015) Distinct soil bacterial communities along a small-scale elevational gradient in alpine tundra. Front Microbiol 6:582
Simpson EM (1949) Measurement of diversity. Nature, London 163:688
Singh DP, Khattar JIS, Kaur M, Kaur G, Gupta M, Singh Y (2013) Anilofos tolerance and its mineralization by the cyanobacterium Synechocystis sp. strain PUPCCC 64. PLoS ONE 8(1):1–10
Singh DP, Khattar JIS, Nadda J, Singh Y, Garg A, Kaur N, Gulati A (2011) Chlorpyrifos degradation by the cyanobacterium Synechocystis sp. strain PUPCCC 64. Environ Sci Pollut Res Int 18:1351–1359
Singh HS, Khattar JS, Ahluwalia AS (2014) Cyanobacteria and agricultural crops. Vegetos 27(1):37–44
Singh JS, Kumar A, Rai AN, Singh DP (2016) Cyanobacteria: a precious bio-resource in agriculture, ecosystem, and environmental sustainability. Front Microbiol 7:529
Singh RN (1950) Reclamation of “usar” lands in India through blue green algae. Nature 165:325–326
Singh RN (1961) Reclamation of usar lands. Role of Blue Green Algae in Nitrogen Economy of Indian Agriculture New Delhi. ICAR 83–98
Singh P, Kumar N, Jethva M, Yadav S, Kumari P, Thakur A, Kushwaha HR (2018) Riboswitch regulation in cyanobacteria is independent of their habitat adaptations. Physiol Mol Biol Plants 24:315–324
Snoeijs RJM, Prentice IC (1989) Effects of cooling water discharge on the structure and dynamics of epilithic algal communities in the northern Baltic. Hydrobiologia 184:99–123
Sommer U, Adrian R, De Senerpont Domis L, Elser JJ, Gaedke U, Ibelings B, Erik Jeppesen E, Lürling M, Molinero JC, Mooij WM, van Donk E, Winder M (2012) Beyond the Plankton Ecology Group (PEG) model: mechanisms driving plankton succession. Annu Rev Ecol Evol Syst 43:429–448
Sommer U, Gliwicz ZM, Lampert W, Duncan A (1986) The PEG model of a seasonal succession of planktonic events in fresh waters. Arch Hydrobiol 106:433–471
Sompong U, Hawkins PR, Besley C, Peerapornpisal Y (2005) The distribution of cyanobacteria across physical and chemical gradients in hot springs in northern Thailand. FEMS Microbiol Ecol 52(3):365–376
Song T, Mårtensson L, Eriksson T, Zheng W, Rasmussen U (2005) Biodiversity and seasonal variation of the cyanobacterial assemblage in a rice paddy field in Fujian, China. FEMS Microbiol Ecol 54(1):131–140
Srinivas M, Aruna M (2016) Diversity of Blue Green Algae from Paddy Fields. Int J Sci 5(5):27–30
Srivastava AK, Bhargava P, Kumar A, Rai LC, Neilan BA (2009) Molecular characterization and the effect of salinity on cyanobacterial diversity in the rice fields of Eastern Uttar Pradesh, India. Saline syst 5(1):1–17
Subramanian G, Sekar S, Sampoornam S (1994) Biodegradation and utilization of organophosphorus pesticides by cyanobacteria. Int Biodeter Biodegr 33:129–143
Suganya T, Varman M, Masjuki HH, Renganathan S (2016) Macroalgae and microalgae as a potential source for commercial applications along with biofuels production: a biorefinery approach. Renew Sust Energ Rev 55:909–941
Swarnalakshmi K, Prasanna R, Kumar A, Pattnaik S, Chakravarty K, Shivay YS, Singh R, Saxena AK (2013) Evaluating the influence of novel cyanobacterial bio filmed biofertilizers on soil fertility and plant nutrition in wheat. Eur J Soil Biol 55:107–116
Tandon P, Abrol YP, Kumaria S (2007) Biodiversity and its Significance. I. K. International Publishing House
ter Braak CJF (1988) Unimodal Models to Relate Species to Environment. Biometrics 44(2):631
ter Braak CJF, Verdonschot PFM (1995) Canonical correspondence analysis and related multivariate methods in aquatic ecology. Aquat Sci 57:255–289
Thukral AK (2017) A review on measurement of Alpha diversity in biology. Agric Res J 54(1):1–10
Tokeshi M (1993) Species abundance patterns and community structure. Adv Ecol Res 24:111–186
Tuomisto H (2012) An updated consumer’s guide to evenness and related indices. Oikos 121:1203–1218
Van Der Putten WH (2017) Belowground drivers of plant diversity. Science 355(6321):134–135
Venkataraman GS (1993) Blue-green algae (Cyanobacteria). In Tata SN, Wadhwani AM, Mehdi MS (ed) Biological Nitrogen Fixation. (New Delhi: Indian Council Agric Res), 45–76
Vijayan D, Ray JG (2015) Ecology and diversity of Cyanobacteria in Kuttanadu paddy wetlands, Kerala, India. Am J Plant Sci 6(18):2924
Villéger S, Mason NW, Mouillot D (2008) New multidimensional functional diversity indices for a multifaceted framework in functional ecology. Ecology 89(8):2290–2301
Ward CS, Yung CM, Davis KM, Blinebry SK, Williams TC, Johnson ZI, Hunt DE (2017) Annual community patterns are driven by seasonal switching between closely related marine bacteria. ISME J 11(6):1412–1422
Ward DM, Castenholz RW, Miller SR (2012) Cyanobacteria in geothermal habitats. In: Potts M, Whitton BA (eds) Ecology of cyanobacteria: their diversity in time and space. Kluwer Academic Publishers, Dordrecht, pp 37–59
Whittaker RH (1956) Vegetation of the Great Smoky Mountains. Ecol Monogr 26:1–80
Whittaker RH (1960) Vegetation of the Siskiyou mountains, Oregon and California. Ecol Monogr 30(3):279–338
Whittaker RH (1967) Gradient analysis of vegetation. Biol Rev 49:207–264
Whittaker RH (1972) Evolution and measurement of species diversity. Taxon 21(2–3):213–251
Whittaker RH (1977) Evolution of species diversity in land communities. In: Hecht MK, Steere WC, Wallace B (eds) Evolutionary biology, vol 10. Plenum Press, New York, pp 250–268
Wilson EO (1988) Biodiversity. National Academy Press, Washington, DC
Wittebolle L, Marzorati M, Clement L, Balloi A, Daffonchio D, Heylen K, De Vos P, Verstraete W, Boon N (2009) Initial community evenness favours functionality under selective stress. Nature 458(7238):623–626
Yadav A, Kumar D, Singh RS, Pandey LK, Rai J (2018) Seasonal variations in response of periphytic algal community to nutrient enrichment in the river Ganga (Varanasi, India). Ann Limnol - Int J Lim 54:32
Yandigeri MS, Kashyap S, Pabbi S (2011) Studies on mineral phosphate solubilization by cyanobacteria Westiellopsis and Anabaena. Microbiology 80(4):58–565
Yandigeri MS, Pabbi S (2005) Response of diazotrophic cyanobacteria to alternative sources of phosphorus. Indian J Microbiol 45(2):132–134
Yandigeri MS, Yadav AK, Meena KK, Pabbi S (2010) Effect of mineral phosphate on growth and nitrogen fixation of diazotrophic cyanobacteria Anabaena variabilis and Westiellopsis prolifica. Antonie Van Leeuwenhoek 97(3):297–306
Zancan S, Trevisan R, Paoletti MG (2006) Soil algae composition under different agro-ecosystems in North-Eastern Italy. Agric Ecosyst Environ 112(1):1–12
Zapomělová E, Mareš J, Lukešová A, Skácelová O, Řeháková K, Kust A (2013) Extensive polyphyly of non-gas-vacolate Anabaena species (Nostocales, Cyanobacteria) – a challenge for modern cyanobacterial taxonomy. Proceedings of 19th Symposium of IAC. Cleveland OH. p. 51
Zhan J, Sun Q (2012) Diversity of free-living nitrogen-fixing microorganisms in the rhizosphere and non-rhizosphere of pioneer plants growing on wastelands of copper mine tailings. Microbiol Res 167(3):157–165
Zhang H, Hu C, Jia X, Xu Y, Wu C, Chen L, Wang F (2012) Characteristics of γ hexachlorocyclohexane biodegradation by a nitrogen-fixing cyanobacterium, Anabaena azotica. J Appl Phycol 24:221–225
Zhang H, Zong R, He H, Liu K, Yan M, Miao Y, Miao Y, Ma B, Huang X (2021) Biogeographic distribution patterns of algal community in different urban lakes in China: insights into the dynamics and co-existence. J Environ Sci 100:216–227
Acknowledgements
SFI thanks the University Grants Commission, New Delhi for financial support in the form of a University Research Fellowship. LS thanks the Head, Department of Statistics, H.N.B. Garhwal University, for generous support. DK thanks the University Grants Commission for awarding a start-up project.
Author information
Authors and Affiliations
Contributions
SFI performed literature searches, data analysis and prepared the first draft of the manuscript. LS contributed to writing the statistical portions of this review. DK was involved in conceiving the idea and finalizing the manuscript for submission. CMS refined the discussion and fine-tuned the entire draft. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Anurag chaurasia.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ikram, S.F., Singh, L., Kumar, D. et al. Prospects and constraints in studying the biodiversity of agriculturally important microalgae and cyanobacteria and useful statistical tools. Biodivers Conserv 31, 1095–1124 (2022). https://doi.org/10.1007/s10531-022-02388-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10531-022-02388-8