Abstract
Hypertrophic cardiomyopathy (HCM) is one of the most common genetic heart diseases. Its features include abnormal cardiomyocyte hypertrophy, microvascular dysfunction, and increased accumulation of intercellular matrix. We aim to unravel genes associated with the pathogenesis of HCM and provide a potential target for diagnosis and treatment. Key modules were identified by weighted gene co-expression network analysis (WGCNA). A miRNA-mRNA network was constructed with the predicted miRNA and the most likely hub gene was screened out for gene set enrichment analysis (GSEA). The diagnostic capacity of hub gene was verified by receiver operating characteristic (ROC) curves. Single-cell sequencing (sc-RNA seq) data of normal adult hearts were used to further predict the specific cell types expressing the hub gene. WGCNA assigned genes into different modules and found that the genes contained in the red module had the strongest positive correlation with HCM disease. 2.5% of the genes were common between DEG and hub genes. With the miRNA-mRNA network, osteomodulin (OMD) was identified as the most potential hub gene. GSEA showed that OMD was mainly involved in the synthesis of extracellular matrix and had a certain inhibitory effect on the immune system. The expression of OMD in HCM was validated and ROC curve analysis showed that OMD could distinguish HCM from controls with the area under the curve (AUC) > 0.7. The sc-RNA seq revealed that OMD was mainly expressed in the later stages of cardiac fibroblasts, suggesting that OMD may have an effect on fibroblasts, participating in the pathogenesis of HCM. OMD may serve as a biomarker and therapeutic target for HCM in the future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hypertrophic cardiomyopathy (HCM) is a common myocardial disease with asymmetric septal hypertrophy as the most important feature, which can induce heart failure, arrhythmia, and sudden cardiac death (Ullal et al. 2016; Maron et al. 2017). With the development of better diagnostic techniques, the prevalence of HCM is estimated to increase from 0.2 to 0.5% or even higher (Semsarian et al. 2015). Currently, there are several treatment strategies for HCM available. For patients with exertional dyspnea, pharmacological treatments including β-blocker, verapamil, and disopyramide are considered appropriate (Soullier et al. 2012; Wijnker and Velden 2020). A septal myotomy-myectomy operation is considered a standard care for patients with severe refractory symptoms related to marked outflow obstruction (Kobayashi et al. 2014; Kim et al. 2016). Implantable cardioverter-defibrillator was also used for the prevention of sudden cardiac death for high-risk patients (Sidhu et al. 2020). Moreover, alcohol septal ablation and pacing are also considered appropriate options (Liebregts et al. 2017; Daubert et al. 2018).
HCM is often associated with mutations in genes encoding proteins for myocardial sarcomere, calcium-handling, and Z-disk (Maron 2002), with the mutations in β-myosin heavy chain, cardiac troponin T, and myosin-binding protein C accounting for the majority of HCM cases (Hossain et al. 2019; Cohn et al. 2019; Coppini et al. 2020). However, due to the improvement in genetic technology, new genes involved in the pathogenesis of HCM are being continually discovered, which need to be further studied. Histopathology of HCM patients has revealed the presence of microvascular dysfunction, excessive fibrosis, and thickening of vascular intima and media, which were also associated with myocardial ischemia (Maron et al. 2009). Moreover, myofibril disruption, a massive increase in the extracellular matrix and the subsequent appearance of myocardial fibrosis, are also the main pathological changes leading to malignant ventricular arrhythmia, ventricular dilation dysfunction, and heart failure (Tsoutsman et al. 2013; Harris et al. 2006). Tissue-level inflammation also plays an important role in HCM (Becker et al. 2020). Taken together, it is clear that the etiology of HCM can be varied, and hence, it is important to uncover and explore the genes related to its pathogenesis for precise diagnosis and treatment.
Weighted gene co-expression network analysis (WGCNA) is a common tool to explore the correlation of gene modules with disease status and it has been widely applied in multiple types of cancer, autoimmune diseases, and brain imaging analysis (Anaya et al. 2016; Liu et al. 2019; Tang et al. 2020). By transforming gene expression data into co-expression gene modules and relationship between gene and clinical traits, WGCNA is useful for the identification of potential biomarkers or therapeutic targets (Langfelder and Horvath 2008).
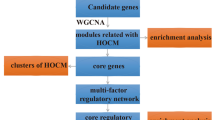
In this study, WGCNA was used to analyze gene expression data from GEO. To identify the key module, a functional enrichment analysis was carried out to explore potential functions of genes related to HCM. Furthermore, a gene set enrichment analysis was performed for the selected hub gene to find the relevant gene sets.
Methods
Data Collection
The GSE36961, GSE89714, and GSE134355 datasets were downloaded from the NCBI Gene Expression Omnibus (GEO) (https://www.ncbi.nlm.nih.gov/geo/). GSE36961 consisted of 106 HCM samples and 39 controls, the platform was Illumina HumanHT‐12 V3.0 expression bead chip (GPL15389), clinical information included age, gender and disease status (Bos et al. 2020). GSE89714 contained 5 HCM samples and 4 controls, the platform was Illumina HiSeq 2000 (GPL11154). GSE134355 contained single-cell sequencing data of human organs, the platform was HiSeq X Ten (GPL20795), and the data on adult hearts were selected for further analysis.
Weighted Gene Co-Expression Network Analysis
The top 5000 genes from the GSE36961 dataset, ranked by standard deviation (SD) from large to small, were selected for WGCNA. The appropriate soft-threshold power for network construction was provided by calculating the scale-free topology fit index for several powers using R package “WGCNA” (Langfelder and Horvath 2008). The adjacent matrix was transformed into TOM (Topological Overlap Matrix) and a hierarchical clustering tree was constructed to assign genes with similar expression profiles into the same module. The correlation between modules and clinical traits was calculated. The gene significance(GS) was defined as the log10 transformation of the P value of the linear regression between the gene expression and clinical traits. The definition of module significance(MS) was the average GS for all genes included in the same module. The module with the highest absolute MS was considered the key module most relevant to the clinical traits.
Identification of Hub Genes in the Key Module
The Search Tool for the Retrieval of Interacting Genes (STRING) (http://string-db.org/) is a database that can search and predict the interactions between known proteins, which was used to analyze the interactions in this study. Genes that interacted highly with others in the same module were considered as hub genes. Cytoscape was used for the visualization of genes from the key module and the top 20 genes were chosen for further analysis based on the result of the main connected component(MCC) calculation. The selection of differential expressed genes (DEGs) were done with R package “limma”, with log2|FC|> 0.6 and adjusted P value < 0.01 by Benjamini–Hochberg as the cutoff value. Volcano plots and hierarchical clustering analysis were performed on R “ggplot2” and “pheatmap”, respectively. Venn diagrams of overlapping hub genes in the key module and DEGs were created using the online tool “Venny” (http://bioinfogp.cnb.csic.es/tools/venny/) (Bardou et al. 2014).
Functional Enrichment Analysis
The R package “clusterprofiler” was used for Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis of the key module and differentially expressed genes (DEGs). The GO terms included biological process (BP), molecular function (MF), and cellular component (CC) (Yu et al. 2012). Adjusted P value of < 0.05 was regarded significant.
miRNA-mRNA Network
To refine the potential target gene, miRNAs upstream of the hub genes were predicted by “TargetScan” and “miRTarBase” (Agarwal et al. 2015; Chou et al. 2018). The intersection was used for the construction of the miRNA-mRNA network using Cytoscape. Based on the Human microRNA Disease Database (HMDD) (http://www.cuilab.cn/hmdd) and published studies, we identified the miRNAs which had been reported to be associated with HCM pathology and the hub genes regulated by those miRNAs were selected for further analysis and validation (Lu et al. 2008; Kuster et al. 2013; Fang et al. 2015).
Validation and Efficacy Evaluation of Hub Gene
Hub gene expression was validated using the GSE89714 dataset. The correlation between hub genes and other genes was applied by the Pearson correlation coefficient. Moreover, ROC curves and AUC of the hub gene and genes highly connected with the hub gene were calculated using R package “pROC” to evaluate the capacity to distinguish HCM patients and controls. GSE36961 and GSE89714 datasets were normalized and integrated. 70% of the samples were randomly selected as a training cohort and 30% were selected as a validation cohort. The optimal cutoff values of studied genes were obtained by ROC analysis in the training cohort and applied to the validation cohort to evaluate the diagnostic ability of studied genes. The indexes included sensitivity, specificity, positive predictive value(PPV) and negative predictive value(NPV).
GSEA Analysis of Hub Gene
Gene Set Enrichment Analysis (GSEA) for the single hub gene was performed as described previously (Subramanian et al. 2005) using the R “clusterprofiler” and c2.cp.kegg.v7.1.symbols.gmt, the c5.all.v7.1.symbols.gmt from Molecular Signatures Database (MSigDB) were chosen as reference gene sets. According to the median expression level of the hub gene, 145 samples were divided into a high expression group or a low expression group. The expression levels of all genes between the two groups (high/low) were compared and sorted from upregulated genes to downregulated genes. By comparing with the GSEA reference gene set, the signal pathways that were activated or suppressed in each group were obtained. Adjusted P value of < 0.05 was selected as a cutoff.
Single-Cell Sequencing
To further explore the possible mechanism of the hub gene in the heart tissue, we extracted the single-cell sequencing data on adult hearts from GSE134355. The R package “Seurat” was used for data integration, elimination of batch effect, and quality control (Tran et al. 2020). Cells were divided into various clusters using t-SNE analysis and cluster annotation was performed by R package “scHCL” (Han et al. 2020). Moreover, single-cell pseudotime trajectories were carried out with the R package “monolce2” for further exploration of the dynamic changes in the hub gene (Qiu et al. 2017).
Results
Expression Profiling
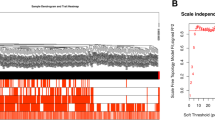
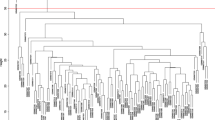
The dataset GSE36961 included 106 HCM and 39 controls. For genes matched by multiple probes, the median was considered as the expression value. Finally, a total of 37,846 genes were extracted and the top 5000 genes selected for WGCNA were ranked based on standard deviation in descending order. Based on a cluster analysis of these 5000 genes, the cohort of 145 samples was further divided into two clusters (Supplementary Fig. 1). To identify the relatively balanced scale independence and mean connectivity of WGCNA, a network topology analysis was carried out to select the suitable soft-thresholding power from 1 to 20. A power value of 6 was chosen for the hierarchical clustering tree of the selected 5000 genes with 0.9 as the correlation coefficient threshold (Fig. 1a, b).
Weighted Co-Expression Network (WGCNA)
Based on the degree of connectivity, genes were assigned to 14 different modules. There were 191 genes in the black module, 916 genes in the blue module, 776 genes in the brown module, 43 genes in the cyan module, 248 genes in the green module, 102 genes in the green-yellow module, 143 genes in the magenta module, 181 genes in the pink module, 116 in the purple module, 220 genes in the red module, 50 genes in the salmon module, 54 genes in the tan module, 1189 genes in the turquoise module and 454 genes in the yellow module. The genes that could not be assigned into any module were assigned to the gray module, and removed from subsequent analysis (Fig. 2a).
Correlations and Identification of the Key Module
Intermodular association analysis showed that the modules were independent of each other, indicating relative independence of gene expression in each module as shown in the heat map in Fig. 3a. The eigengenes were calculated and clustered based on the correlation to explore the co-expression similarity of all modules. The red module showed the highest positive correlation, while the brown module showed the highest negative correlation with disease status (HCM and control), suggesting that the red module was the most positively associated with the disease status of HCM (Fig. 3b). Also, The result revealed that the 14 modules was mainly separated into 3 clusters, which was similar to the result of the heat map (Fig. 3c, d).
a Interaction relationship analysis of co-expression genes. Different colors of vertical axis and horizontal axis represent different modules. The density of yellow in the middle represents the degree of connectivity of different modules. b Heatmap of the correlation between module eigengenes and disease status of HCM. The red module showed to be the most positively correlated with the disease status. c Hierarchical clustering of module hub genes. d Heatmap plot of the adjacencies in the hub gene network
Validation and Efficacy Evaluation of Hub Gene
According to the result of WGCNA, a correlation coefficient of 0.78 between the module membership and GS in the red module was found to be the most relevant to the disease status (Fig. 4a). Cytoscape was used to select the top 20 genes from the red module based on MCC calculation (Fig. 4b). With the cutoff value at log2|FC|> 0.6 and adjusted P value at < 0.01, a total of 648 DEGs were screened out, which included 254 upregulated and 394 downregulated genes (Fig. 4c, d). Figure 4e demonstrates a Venn Diagram of 16 genes common between the DEGs and hub genes in the red module. The common genes were DIO2, IRX6, LUM, PDE4D, PROS1, PDE5A, OMD, THBS4, COMP, ACE2, SYNPO2L, TMEM30B, PLCE1, APCDD1L, SFRP4 and IL17D (Fig. 4e). We speculated that these 16 common genes may be more relevant to the pathogenesis of HCM than the other genes in the red module. The miRNAs-mRNA network showed there were 27 miRNAs that could regulate more than one of these gene (Supplementary Fig. 2). Searching the HMDD database and published literature, we found that the downregulation of hsa-miR-34b and hsa-miR-373 was related to the development of HCM. Further analysis revealed that the downstream target of hsa-miR-34b was PLCE1 and TMEM30B; meanwhile, hsa-miR-373 targeted SYNPO2L and OMD. Transcript abundance of the four genes-PLCE1, TMEM30B, SYNPO2L, and OMD was significantly upregulated in the HCM cohort (Fig. 5a). As the association between the upregulation of SYNPO2L and HCM was well established, we chose OMD as our target for further analysis and validation. Validation with GSE89714 concurred with the findings observed with GSE36961- that OMD was highly expressed in HCM (Fig. 5b, c). In order to further evaluate the predictive ability of OMD, GSE36961 dataset and GSE89714 dataset were normalized and integrated, 70% of samples were randomly chosen as a training cohort and 30% of the samples were chosen as a validation cohort. The ROC analysis was applied to OMD and several genes closely related to OMD including LUM and MFAP4 in the training cohort (Fig. 5d–f). Based on ROC, the optimal cutoff values for OMD, LUM and MFAP4 were obtained.Applying these cutoff values to the validation cohort, the sensitivity, specificity, positive predictive value(PPV), and negative predictive value(NPV) in predicting HCM were calculated and shown in Table 1. For HCM, high expression of OMD, LUM and MFAP4 had PPV of 96.15%, 91.67% and96.30%,respectively. The low expression of OMD, LUM, and MFAP4 had NPV of 42.86%, 63.64%, and 45.00%, respectively.
a The gene significance for HCM in the red module (One dot represents one gene in the red module). b The top 20 hub genes in the red module. The color of nodes represents the degree of gene interaction, the darker the color,the higher connectivity of the genes. The edge represents the interaction between genes. c Volcano plot visualizing DEGs in GSE36961. d Heat map hierarchical clustering shows DEGs in HCM samples compared with normal samples. e Identification of common genes between DEGs and red module
a The extracted miRNA-mRNA regulatory network, the green nodes represent the downregulated disease-related miRNAs, and the red circular nodes represent the upregulated hub genes in the red module. b Expression of OMD in GSE36961. c Validation of OMD in GSE89714. d ROC curve of OMD. e ROC curve of LUM. f ROC curve of MFAP4
Functional Enrichment Analysis
The KEGG pathway analysis demonstrated that the red module was primarily enriched in the TGF-beta signaling pathway, ECM-receptor interaction, Complement and coagulation cascades (Fig. 6a). The GO enrichment analysis showed that the genes were mainly enriched in extracellular structure organizations, extracellular matrix organizations, and ossifications (Fig. 6b). Moreover, DEGs were mainly enriched in metabolic pathways, phagosomes, and pathways in cancer after KEGG pathway analysis (Fig. 7a). The GO enrichment analysis showed that the DEGs were enriched in extracellular structure organizations, extracellular matrix organizations, and neutrophil-mediated immunities (Fig. 7b). Hence, this indicated that both the hub genes from WGCNA and DEGs showed strong correlation with the extracellular matrix.
Gene Set Enrichment Analysis
The KEGG pathway analysis showed that high expression of OMD could activate valine, leucine, and isoleucine degradation, Huntington disease, and the TGF-beta signaling pathway, and primarily suppressed B cell receptors, Toll-like receptors, MAPK signaling pathways, and cytokine receptor interactions (Fig. 8a). By GO, OMD was found to be enriched in extracellular matrix structural constituents, collagen-containing extracellular matrices, microtubule-based movements, and extracellular structural organizations, and suppressed in the cellular response to biotic stimuluses, B cell-mediated immunities, activation of innate immune responses, antigen receptor-mediated signaling pathways, and B cell activation (Fig. 8b). This GSEA suggested that higher expression of OMD promoted extracellular matrix remodeling and inhibited immune function to a certain degree.
Single-Cell Sequencing
The sc-RNA sequencing data from normal adult hearts were extracted from GSE134355, and after normalization, data quality control, and removal of batch effect, a total of 19,996 cells were selected for analysis. Cells were divided into 8 clusters and cell types were annotated according to the marker genes, including cardiomyocyte, endothelial cell, fibroblast, macrophage, monocyte, neutrophil, natural killer(NK) cell, and smooth muscle cell (Fig. 9a–c). Our results showed that within the cardiac tissue, OMD was mainly expressed in fibroblasts (Fig. 9d). We extracted the expression matrix of fibroblasts and based on the variable genes, fibroblasts were further divided into 4 subgroups and OMD was found to be primarily expressed in clusters 1 and 2 (Fig. 9e–f). A pseudotime trajectory of the fibroblasts using the R package “monocle2”, showed 8 branches, and the cells were grouped into 9 states (Fig. 9g–i). The root of trajectory was populated by the majority of cluster 0, while the end of the trajectory was mainly populated by clusters 1 and 2. According to the estimated pseudotime trajectory, we considered that the cells may travel from cluster 0 to 1 and 2. Also, the dynamic changes in the relative expression of OMD indicated that OMD was lower in the early states of the fibroblasts than the later states where it was higher (Fig. 9j–l). These results implied that the increased expression of OMD may affect the function of fibroblasts at later states.
a Integration of sc-RNA seq datasets of adult hearts in GSE134355, in which the batch effect has been removed. b t-SNE plot showing the cell types of 19,996 single cells isolated from adult hearts. c Heatmap showing the DEGs of each cell type. d t-SNE plot showing that OMD was mainly expressed in fibroblasts. e t-SNE plot showing that fibroblasts were grouped into four clusters. f t-SNE plot showing that OMD was mainly expressed in cluster 1 and 2 in fibroblasts. g–i Monocle analysis showing fibrobalsts in pseudotime. Cells are colored based on cluster (g), state (h), and pseudotime (i). j–l Monocle analysis showing the expression of OMD in fibroblast in pseudotime. Cells are colored based on cluster (j), state (k), and pseudotime (l)
Discussion
HCM is one of the most common nontraumatic causes of cardiac death. Histopathological findings have revealed that a variety of pathological processes are involved in the development of HCM, including microvascular dysfunction, increased interstitial space due to extracellular matrix formation, myofibrillar disruption, and inflammatory responses at the tissue level. In this study, we used a global approach to construct co-expression network of HCM to predict the clusters of genes involved in the pathogenesis of HCM. We aimed to find new and critical biomarkers and to understand the molecular mechanism of HCM, which might contribute to the diagnosis and treatment of the disease.
We utilized WGCNA to construct a HCM-related gene co-expression network that was grouped into 14 modules. Among these, the red module containing 220 genes was most relevant to the disease status. A GO enrichment analysis demonstrated that the 648 DEGs and hub genes in the red module were mainly enriched in extracellular structure organizations, extracellular matrix organizations, and ossifications. These findings were in accordance to previously reported changes in the extracellular matrix in interstitial spaces in HCM.
We identified 16 hub genes common between DEGs and the red module, and upstream miRNAs regulating them were predicted for the construction of a miRNA-mRNA network. Based on the results of HMDD and published studies related to HCM, OMD was finally chosen as a potential target gene for the disease. OMD is a leucine- and aspartic acid-rich keratan sulfate proteoglycan, which belongs to the small leucine-rich proteoglycan family (SLRP) family (Juchtmans et al. 2015). SLRPs are produced and function in a tissue-specific manner, and are known to be involved in the regulation of matrix assembly and host immunity. SLRP in bone is related to collagen fibrillogenesis and bone mineralization, while in renal disease, it plays an important role in signal transduction and immune response (Zvackova et al. 2017; Zou et al. 2019). OMD was first isolated from the mineralized matrix of bone and a recent study has shown that OMD could directly bind to Type I collagen, further regulating the diameter and shape of collagen fibrils (Tashima et al. 2015). Moreover, GSEA revealed that OMD was enriched in extracellular matrix structural constituents, collagen-containing extracellular matrix, microtubule-based movements, and extracellular structural organizations. Taken together, these results provided an implication that OMD was associated with the regulation of the extracellular matrix.
The upregulation of OMD in HCM samples when compared with controls was confirmed and validated with GSE89714. Our findings were in line with those reported on distinct populations, using different platforms for gene expression analysis, suggesting that the upregulation of OMD in HCM was universal. The ROC curves of OMD and genes highly connected with OMD based on GSE36961 generated AUCs greater than 0.7, suggesting that OMD had a good diagnostic value for HCM.
Single-cell sequencing data of normal adult hearts from GSE134355 showed that OMD was mainly expressed in fibroblasts. The fibroblasts were divided into 4 clusters and OMD was mainly expressed in cluster 1 and cluster 2. A pseudotime trajectory analysis and dynamic changes in the expression showed that the expression of OMD was on the rise at the later stage of fibroblasts. Based on these results, we speculated that the increased expression of OMD may regulate the synthesis of extracellular matrix by fibroblasts at the later stages of the disease. Besides, GSEA also showed that OMD could suppress the cellular response to Biotic stimulus, B cell-mediated immunity, activation of an innate immune response, antigen receptor-mediated signaling pathway, and B cell activation. These results implied that the higher expression of OMD may influence not only fibroblasts but also immune cells in HCM.
There were some limitations to our study. First, this study was primarily in silico data analysis and data mining, and the results were not validated in vitro and in vivo. Second, the exact molecular mechanisms of OMD in HCM was not further explored. Hence, studies to elucidate the molecular mechanisms of OMD in HCM is necessary in the future.
Abbreviations
- HCM:
-
Hypertrophic cardiomyopathy
- GEO:
-
Gene expression omnibus
- STRING:
-
The search tool for the retrieval of interacting genes
- sc-RNA seq:
-
Single-cell RNA sequencing
- OMD:
-
Osteomodulin
- WGCNA:
-
Weighted gene co-expression network analysis
- SD:
-
Standard deviation
- GS:
-
Gene significance
- MS:
-
Module significance
- GO:
-
Gene ontology
- CC:
-
Cellular component
- BP:
-
Biological process
- MF:
-
Molecular function
- MCC:
-
Main connected component
- HMDD:
-
Human microRNA disease database
- PPV:
-
Positive predictive value
- NPV:
-
Negative predictive value
- LUM:
-
Lumican
- MFAP4:
-
Microfibrillar associated protein 4
- GSEA:
-
Gene Set Enrichment Analysis
- MsigDB:
-
Molecular Signatures Database
- SLRP:
-
Small leucine-rich proteoglycan family
References
Agarwal V, Bell GW, Nam JW, Bartel DP (2015) Predicting effective microRNA target sites in mammalian mRNAs. eLife 4.
Anaya JM, Rojas-Villarraga A, Mantilla RD, Arcos-Burgos M, Sarmiento-Monroy JC (2016) Polyautoimmunity in Sjögren Syndrome. Rheumat Dis Clin North Am 42:457–472
Bardou P, Mariette J, Escudié F, Djemiel C, Klopp C (2014) jvenn: an interactive Venn diagram viewer. BMC Bioinform 15:293
Bos JM, Hebl VB, Oberg AL et al (2020) Marked Up-Regulation of ACE2 in Hearts of patients with obstructive hypertrophic cardiomyopathy: implications for SARS-CoV-2-Mediated COVID-19. Mayo Clinic Proceed
Becker RC, Owens AP, Sadayappan S (2020) Tissue-level inflammation and ventricular remodeling in hypertrophic cardiomyopathy. J Thromb Thrombol 49:177–183
Chou CH, Shrestha S, Yang CD et al (2018) miRTarBase update 2018: a resource for experimentally validated microRNA-target interactions. Nucleic Acids Res 46:D296–D302
Cohn R, Thakar K, Lowe A et al (2019) A contraction stress model of hypertrophic cardiomyopathy due to sarcomere mutations. Stem Cell Rep 12:71–83
Coppini R, Santini L, Olivotto I, Ackerman Michael J, Cerbai E (2020) Abnormalities in Sodium Current and Calcium Homeostasis as Drivers of Arrhythmogenesis in Hypertrophic Cardiomyopathy. Cardiovasc Res
Daubert C, Gadler F, Mabo P, Linde C (2018) Pacing for hypertrophic obstructive cardiomyopathy: an update and future directions. Europace : Eur Pacing, Arrhythmias, Cardiac electrophysiol: J work Groups Cardiac Pacing, Arrhythmias, Cardiac Cell Electrophysiol Soc Cardiol 20:908–920
Fang L, Ellims AH, Moore X et al (2015) Circulating microRNAs as biomarkers for diffuse myocardial fibrosis in patients with hypertrophic cardiomyopathy. J Transl Med 13:314
Harris KM, Spirito P, Maron MS et al (2006) Prevalence, clinical profile, and significance of left ventricular remodeling in the end-stage phase of hypertrophic cardiomyopathy. Circulation 114:216–225
Hossain MB, Elbeck Z, Siga H, Knöll R (2019) Myosin binding protein-C and hypertrophic cardiomyopathy: role of altered C10 domain. Cardiovasc Res 115:1943–1945
Han X, Zhou Z, Fei L et al (2020) Construction of a human cell landscape at single-cell level. Nature 581:303–309
Kuster DWD, Mulders J, Cate FJT et al (2013) MicroRNA transcriptome profiling in cardiac tissue of hypertrophic cardiomyopathy patients with MYBPC3 mutations. J Mol Cell Cardiol 65:59–66
Ne J, Dhollander AAM, Coudenys J et al (2015) Distinct dysregulation of the small leucine-rich repeat protein family in osteoarthritic acetabular labrum compared to articular cartilage. Arthritis Rheumatol 67:435–441
Kobayashi T, Dhillon A, Popovic Z et al (2014) Differences in global and regional left ventricular myocardial mechanics in various morphologic subtypes of patients with obstructive hypertrophic cardiomyopathy referred for ventricular septal myotomy/myectomy. Am J Cardiol 113:1879–1885
Kim LK, Swaminathan RV, Looser P et al (2016) Hospital volume outcomes after septal myectomy and alcohol septal ablation for treatment of obstructive hypertrophic cardiomyopathy: us nationwide inpatient database, 2003–2011. JAMA Cardiol 1:324–332
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinform 9:559
Lu M, Zhang Q, Min D et al (2008) An analysis of human microRNA and disease associations. PLoS ONE 3:e3420
Liebregts M, Faber L, Jensen MK et al (2017) Outcomes of alcohol septal ablation in younger patients with obstructive hypertrophic cardiomyopathy. JACC Cardiovasc Intervent 10:1134–1143
Liu Z, Wu K, Wu J, Tian D, Chen Y, Yang Z, Wu A (2019) NEAT1 is a potential prognostic biomarker for patients with nasopharyngeal carcinoma. J Cell Biochem 120:9831–9838
Maron BJ (2002) Hypertrophic cardiomyopathy: a systematic review. JAMA 287:1308–1320
Maron MS, Olivotto I, Maron BJ et al (2009) The case for myocardial ischemia in hypertrophic cardiomyopathy. J Am College Cardiol 54:866–875
Maron BJ, Maron MS, Rowin EJ (2017) Perspectives on the overall risks of living with hypertrophic cardiomyopathy. Circulation 135:2317–2319
Qiu X, Hill A, Packer J, Lin D, Ma YA, Trapnell C (2017) Single-cell mRNA quantification and differential analysis with Census. Nat Methods 14:309–315
Subramanian A, Tamayo P, Mootha VK et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proceed Natl Acad Sci USA 102:15545–15550
Soullier C, Obert P, Doucende G et al (2012) Exercise response in hypertrophic cardiomyopathy: blunted left ventricular deformational and twisting reserve with altered systolic-diastolic coupling. Circul Cardiovasc Imaging 5:324–332
Semsarian C, Ingles J, Maron MS, Maron BJ (2015) New perspectives on the prevalence of hypertrophic cardiomyopathy. J Am College Cardiol 65:1249–1254
Sidhu K, Han L, Picard KCI, Tedrow UB, Lakdawala NK (2020) Ventricular Tachycardia in cardiolaminopathy: characteristics and considerations for device programming. Heart Rhythm.
Tsoutsman T, Wang X, Garchow K, Riser B, Twigg S, Semsarian C (2013) CCN2 plays a key role in extracellular matrix gene expression in severe hypertrophic cardiomyopathy and heart failure. J Mol Cell Cardiol 62:164–178
Tashima T, Nagatoishi S, Sagara H, Ohnuma SI, Tsumoto K (2015) Osteomodulin regulates diameter and alters shape of collagen fibrils. Biochem Biophys Res Commun 463:292–296
Tran HTN, Ang KS, Chevrier M et al (2020) A benchmark of batch-effect correction methods for single-cell RNA sequencing data. Genome Biol 21:12
Tang J, Yang Q, Cui Q et al (2020) Weighted gene correlation network analysis identifies RSAD2, HERC5, and CCL8 as prognostic candidates for breast cancer. J Cell Physiol 235:394–407
Ullal AJ, Abdelfattah RS, Ashley EA, Froelicher VF (2016) Hypertrophic cardiomyopathy as a cause of sudden cardiac death in the young: a meta-analysis. Am J Med 129:486-496.e2
Wijnker PJM, Velden J (2020) Mutation-specific pathology and treatment of hypertrophic cardiomyopathy in patients, mouse models and human engineered heart tissue. Biochimica et biophysica acta. Mol Basis Dis 1866:165774.
Yu G, Wang LG, Han Y, He QY (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. Omics: J Integrat Biol 16:284–287
Zou W, Wan J, Li M et al (2019) Small leucine rich proteoglycans in host immunity and renal diseases. J Cell Commun Signal 13:463–471
Zvackova I, Matalova E, Lesot H (2017) Regulators of collagen fibrillogenesis during molar development in the mouse. Front Physiol 8:554
Acknowledgements
We would like to acknowledge the reviewers for their helpful comments on this paper.
Funding
This study was supported by the National Nature Science Foundation of China (Grant Numbers 31771099, 81971765, 31771060, 81871504 and 31671025).
Author information
Authors and Affiliations
Contributions
CO and MC finished study design, WG, WF, and JH finished experimental studies, WF, XF, and JH finished data analysis, and WG, WF, and XF finished manuscript editing. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10528_2021_10050_MOESM1_ESM.tif
Supplementary Figure 1 The clustering was based on the expression data of GSE36961, which contained 106 HCM and 39 controls. The top 5000 genes with highest SD values were used for the analysis of WGCNA. The color was related to the disease status(HCM and control), age and gender.(The red in status represents HCM and the white represents the control group. The red in gender represents male and the white represents female. The red intensity was related to the age, with the age increases the color deepens.) (TIF 34596 KB)
10528_2021_10050_MOESM2_ESM.pdf
Supplementary Figure 2 miRNA-mRNA regulatory network. The pink circular nodes represents the hub genes, the blue nodes represents the predicted miRNA regulating hub genes, and the yellow nodes represents the common miRNAs among hub genes (PDF 18 KB)
Rights and permissions
About this article
Cite this article
Guo, W., Feng, W., Fan, X. et al. Osteomodulin is a Potential Genetic Target for Hypertrophic Cardiomyopathy. Biochem Genet 59, 1185–1202 (2021). https://doi.org/10.1007/s10528-021-10050-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-021-10050-1