Abstract
A novel marine hydrogen- and sulfur-oxidizing bacterium, designated strain S2-6 T, was isolated from the deep-sea sediment samples at the Longqi hydrothermal system, southwestern Indian Ocean. Cells were Gram-stain-negative, motile, short rods with a single polar flagellum. Growth was observed at 10–45 °C (optimum 33 °C), pH 5.0–8.0 (optimum pH 7.0) and 1.5 to 6.0% (w/v) NaCl with an optimum at 3.0% (w/v). The isolate was an obligate chemolithoautotroph capable of growth using thiosulfate, tetrathionate, elemental sulfur or sodium sulfide as the energy source, and oxygen or nitrate as the sole electron acceptor. When hydrogen was used as the energy source, strain S2-6 T could respire oxygen, nitrate or element sulfur. The major cellular fatty acids of strain S2-6 T were summed feature 3 (C16:1ω7c and/or C16:1ω6c), C16:0 and summed feature 8 (C18:1ω7c and/or C18:1ω6c). The total size of its genome was 2,320,257 bp and the genomic DNA G + C content was 37.3 mol%. Phylogenetic analysis based on 16S rRNA gene sequences and core genes showed that the novel isolate belonged to the genus Sulfurimonas and was most closely related to Sulfurimonas paralvinellae GO25T (96.8% sequence identity) and Sulfurimonas autotrophica OK10T (95.8% sequence identity). The average nucleotide identity and DNA-DNA hybridization values between strain S2-6 T and S. paralvinellae GO25T and S. autotrophica OK10T were 74.6%-81.2% and 19.1%-24.6%, respectively. Based on the polyphase taxonomical data, strain S2-6 T represents a novel species of the genus Sulfurimonas, for which the name Sulfurimonas sediminis sp. nov. is proposed, with the type strain S2-6 T (= MCCC 1A14513T = KCTC 15854 T).
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Deep-sea hydrothermal vent environments are characterized by the steep gradients of physical and chemical parameters in the mixing zones between hot vent fluids and cold deep-sea water (Dick 2019). Chemosynthetic bacteria and archaea inhabited there utilize chemical energy to fix inorganic carbon into organic carbon for microbial growth, and constitute the foundation of vent ecosystems (McNichol et al. 2018; Nadine et al. 2018). Chemolithoautotrophs are an important component of the microbial community in the deep-sea hydrothermal vent, and play critical roles in deep-sea carbon, nitrogen and sulfur cycling (Cao et al. 2014; Ding et al. 2017).
The genus Sulfurimonas was first proposed by Inagaki et al. (2003) and belonged to the class Campylobacteria (formerly Epsilonproteobacteria) within the phylum of Epsilonbacteraeota (formerly Proteobacteria) (Inagaki et al. 2003; Waite et al. 2017, 2018). Members of the genus Sulfurimonas are ubiquitous in diverse habitats, including deep-sea hydrothermal vents, marine sediments, pelagic water column redoxclines and terrestrial soils (Han and Perner 2015). They are characterized as small sulfur-oxidizing bacteria, which are capable of growing chemoautotrophically with reduced sulfur compounds, such as sulfide, thiosulfate and elemental sulfur (Han and Perner 2015). At the time of writing, the genus contains eight validly published species with Sulfurimonas autotrophica as type species and three Candidatus species including Ca. Sulfurimonas hongkongensis AST-10, Ca. Sulfurimonas marisnigri SoZ1 and Ca. Sulfurimonas baltica GD2. Among them, S. autotrophica OK10T, Sulfurimonas denitrificans DSM1251T, Ca. S. hongkongensis AST-10, Sulfurimonas xiamensis 1-1NT and Sulfurimonas lithotrophica GYSZ_1T were isolated from coastal or deep-sea marine sediments. Sulfurimonas paralvinellae GO25T was isolated from a hydrothermal polychaete nest, Sulfurimonas indica NW8NT was isolated from a hydrothermal sulfide chimney, Sulfurimonas crateris SN118T was isolated from a terrestrial mud volcano, and Sulfurimonas gotlandica GD1T, Ca. S. baltica GD2 and Ca. S. marisnigri SoZ1 were isolated from the pelagic redoxcline of the Baltic Sea and Black Sea (Hoor 1975; Takai et al. 2006; Labrenz et al. 2013; Cai et al. 2014; Ratnikova et al. 2019; Hu et al. 2020; Wang et al. 2020b; Henkel et al. 2020). In this study, we report a novel strain, designated S2-6 T, isolated from the deep-sea sediment samples at the Longqi hydrothermal system, southwestern Indian Ocean. Comparative 16S rRNA gene sequence analysis and physiological properties indicated that strain S2-6 T belonged to the genus Sulfurimonas. The aim of the present work is to determine the exact taxonomic position of strain S2-6 T by using a polyphasic approach.
Materials and methods
Bacterial isolation and culture condition
The sediment sample was collected from the Longqi hydrothermal system in the southwestern Indian Ocean (49°64′E, 37°78′S; Site 49I-SWIR-S11TVG04) in January 2018. For isolation, 1 g of each sediment sample was transferred into 50 ml serum bottles containing 10 ml MMJS (Takai et al. 2006) medium under a gas phase mixture of 78% N2/18% CO2/2% O2 (200 kPa) and incubated at 28 °C according to the previous description (Jiang et al. 2017). MMJS medium consisted of (per litre of distilled, deionized water) 30.0 g NaCl, 0.14 g K2HPO4, 0.14 g CaCl2·2H2O, 0.25 g NH4Cl, 4.18 g MgCl·6H2O, 0.33 g KCl, 0.5 mg NiCl·6H2O, 0.5 mg Na2SeO3·5H2O, 0.01 g Fe(NH4)2(SO4)2·6H2O, 10 mM Na2S2O3·5H2O, 1 g NaHCO3, 10 ml trace mineral solution and 1 ml vitamin solution (Balch et al. 1979). After incubation for 7 days, growth of motile and rod-shaped cells in MMJS medium was observed. The well-grown culture was further purified using the dilution-to-extinction method with the same medium (Takai and Horikoshi 2000). After repeating the dilution-to-extinction for three times from the well-grown culture at the maximum dilution series (10–7), a sulfur-oxidizing bacterium, designated strain S2-6 T, was obtained. The purity of the isolate was verified by microscopic observation, cultivation on heterotrophic media and determination of the 16S rRNA gene sequence.
DNA extraction, genomic and phylogenetic analyses
The genomic DNA of strain S2-6 T was extracted according to a previously described method (Jiang et al. 2010), and the 16S rRNA gene was amplified by PCR primers described previously (Liu and Shao 2005). The 16S rRNA gene sequence was identified using global alignment algorithm implemented at the EzBioCloud server (https://www.ezbiocloud.net/; (Yoon et al. 2017)). Phylogenetic trees were performed using MEGA software package version 6.0 (Tamura et al. 2013) using neighbor-joining (Saitou and Nei 1987), maximum-likelihood (Felsenstein 1981) and minimum evolution (Rzhetsky and Nei 1992) methods after multiple alignments of the data by CLUSTAL_W. Evolutionary distances were calculated using Kimura’s two-parameter model (Kimura 1980) and bootstrap values were determined based on 1000 replications (Felsenstein 1985).
The complete genome sequence of strain S2-6 T was sequenced by Tianjin Biochip Corporation (Tianjin, PR China), using the single molecule real-time (SMRT) technology on the Pacific Biosciences (PacBio) sequencing platform. The sequenced reads were filtered, and high quality paired-end reads were assembled to construct a circular genome using SOAPdonovo software (Luo et al. 2012). The G + C content of the chromosomal DNA was determined according to the genome sequence. tRNAscan-SE (Lowe and Chan 2016) and rRNAmmer (Lagesen et al. 2007) were used to predict the tRNA and rRNA contained in the genome. Gene prediction by using Glimmer v3.02 (Delcher et al. 2007). The average nucleotide identity (ANI) values of strain S2-6 T and the relatives were estimated using OrthoANI computation (Yoon et al. 2017). Digital DNA-DNA hybridization (DDH) estimates were calculated on the GGDC website (https://ggdc.dsmz.de/). The phylogenomic tree was constructed based on an up-to-date 92 bacterial core gene sets by UBCG version 3.0 (Na et al. 2018).
Morphology, physiology and chemotaxonomic analysis
The cell morphology of strain S2-6 T was observed with the cells cultured in MMJS liquid medium at 28 °C for 1 day with a transmission electron microscopy (Model JEM-1230; JEOL). The Gram-stain test was conducted with a Gram staining kit (Hangzhou Tianhe Microorganism Reagent Co.). Growth characteristics were determined by direct cell counting using a phase contrast microscope (Eclipse 80i, Nikon, Japan) according to the previous method (Jiang et al. 2017). All experiments described below were conducted in triplicate. The growth temperature was determined under various temperatures for 1 day (15, 20, 25, 28, 30, 33, 35, 37, 40, 45, 50 and 60 °C) or 4 days (4 and 10 °C) to observe its growth. The growth salinity range was examined by adjusting the concentrations of 0 to 9.0% (w/v) NaCl, at 0.5% (w/v) intervals. The pH range of growth was tested in MMJS medium by adjusting pH with different buffers, including 10 mM acetate/acetic acid buffer (pH 3.0–5.5), MES (pH 5.0–6.5), PIPES (pH 6.5–7.0), HEPES (pH 7.0–8.0), Tris and CAPSO (pH 8.0 and above). Oxygen sensitivity was examined using MMJS basal medium without nitrate with different O2 concentrations gradients (0%, 1%, 2%, 4%, 6%, 8%, 10% at 200 kPa and 20% at 100 kPa) in the headspace gas. In the case of oxygen absence, 10 mM nitrate was added as a potential electron acceptor.
The ability for sulfur oxidation was tested in the MMJS medium using various sulfur compounds other than thiosulfate as the sole energy source, including sulfite (5 mM), tetrathionate (5 mM), thiocyanate (5 mM), elemental sulfur (1% w/v) or sodium sulfide (50, 100, 200, 400, 800 µM, 1, 2 mM) under a gas phase of 76% N2/20% CO2/4% O2 (200 kPa). Molecular hydrogen was also examined in MMJH medium in the absence of thiosulfate under a gas phase of 76% H2 /20% CO2 /4% O2 (200 kPa). In an attempt to determine the utilization of electron acceptors, sulfate (5 mM), sulfite (5 mM), elemental sulfur (1%, w/v), nitrate (10 mM) and nitrite (1 or 5 mM) were tested with H2 as the sole electron donor under 80% H2 /20% CO2 (200 kPa) and ferric citrate (20 mM), ferrihydrite (20 mM), manganese(IV) oxide (200 μM), nitrate (10 mM) and nitrite (1 or 5 mM) were tested with thiosulfate as the sole electron donor under 80% N2/20% CO2 (200 kPa). Utilization of inorganic nitrogen source, such as ammonium chloride (5 mM), sodium nitrate (5 mM), sodium nitrite (5 mM) or molecular nitrogen (N2), which was added to MMJHS medium lacking all nitrogen source under a gas phase of 76% H2 /20% CO2 /4% O2 (200 kPa), was examined. The potential nutrients required for growth such as selenite, tungstate and vitamins were also examined with MMJS medium with and without the specified nutrients.
Heterotrophic growth was examined in a MMJS medium without NaHCO3 under a gas phase of 96% N2 /4% O2 (200 kPa), containing the following organic carbon sources: 0.1% (w/v) peptone, yeast extract, tryptone, starch, casein and casamino acids, 5 mM acetate, formate, citrate, tartrate, succinate, propionate and pyruvate, 5 mM each of 20 amino acids, 0.02% (w/v) sucrose, galactose, glucose, lactose, fructose, maltose and trehalose. Utilization of these organic compounds as alternative energy sources was also examined in MMJ medium in the absence of thiosulfate under a gas phase of 76% N2 /20% CO2 /4% O2 (200 kPa).
The cellular fatty acid composition was analyzed from cells grown in MMJS medium at 33 °C for 24 h. Fatty acids were saponified, extracted, and methylated using the standard protocol of the Microbial Identification System (MIDI, Sherlock Microbial Identification System, version 6.0B). The fatty acids were analyzed by gas chromatography (GC, Agilent Technologies 6850) and identified by using the TSBA 6.0 database of the Microbial Identification System (Frolov et al. 2017).
CRISPRs and genomic islands
Identification of the CRISPR arrays were analyzed by using the CRISPRCas Finder webserver, with default parameters (https://crispr.i2bc.paris-saclay.fr/) (Grissa et al. 2007). Presence of genomic islands was investigated by using the IslandViewer4 webserver (http://www.pathogenomics.sfu.ca/islandviewer/) (Bertelli et al. 2017).
Results and discussion
Morphology
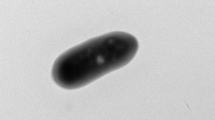
Cells of strain S2-6 T were Gram-stain-negative, rod-shaped (0.5–1.0 × 0.8–2.0 μm) and motile with a polar flagellum (Fig. 1). Morphological features of strain S2-6 T were similar to those of S. autotrophica OK10T, S. paralvinellae GO25T and S. indica NW8NT (Table 1).
Phylogenetic and phylogenomic analyses
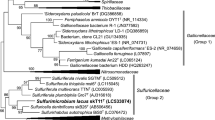
Comparison of the 16S rRNA gene sequences obtained from PCR amplification showed that strain S2-6 T is most closely related to S. paralvinellae GO25T (96.8% sequence similarities), S. autotrophica OK10T (95.8%) and S. indica NW8NT (95.1%). Phylogenetic tree based on the neighbour-joining method indicated that strain S2-6 T formed a distinct cluster with S. paralvinellae GO25T, S. autotrophica OK10T and S. indica NW8NT (Fig. 2). This topology was confirmed by the phylogenetic trees reconstructed with the maximum-likelihood and minimum evolution methods (Fig. S1; S2), which indicated that strain S2-6 T belonged to the genus Sulfurimonas and represent a separate species.
Neighbor-joining tree based on 16S rRNA gene sequences showing the the relationship between strain Sulfurimonas sediminis S2-6 T and other members within the genus Sulfurimonas. Bootstrap values based on 1000 replicates are shown at branch nodes. Branch node values below 50% are not shown. The tree is drawn to scale, with branch lengths measured in the number of substitutions per site. Bar = 0.01 substitutions per nucleotide position
Pairwise ANI values between strain S2-6 T and its closest relative organism, S. paralvinellae GO25T, was 77.8%. The DDH values between strains S2-6 T and S. paralvinellae GO25T was 21.1%. The ANI and DDH values between strains S2-6 T and S. autotrophica OK10T were 81.2% and 24.6%, respectively. All these values are much lower than the threshold criterion for prokaryotic species delineation, which is 95–96% for ANI and 70% for DDH (Chun et al. 2018), indicating that strain S2-6 T represent a novel species in genus Sulfurimonas. The phylogenetic tree based on the 92 core gene sequences indicated that strain S2-6 T formed a branch with strains S. autotrophica OK10T, S. paralvinellae GO25T and S. indica NW8NT (Fig. 3), which supported that strain S2-6 T should be assigned to one novel species of the genus Sulfurimonas. This result was in agreement with the result of 16S rRNA gene phylogeny.
Phylogenetic tree inferred using UBCGs showing the position of Sulfurimonas sediminis S2-6 T and closely related taxa within the genus Sulfurimonas using the maximum-likelihood algorithm. The node is labeled with Gene Support Index (GSI) values. The accession numbers of the genomes are shown in parentheses. Bar, 0.05 substitutions per position
Physiological characteristics
Growth tests revealed that strain S2-6 T can grow at 10–45 °C (optimum 33 °C), pH 5.0–8.0 (optimum pH 7.0) and 1.5–6.0% (w/v) NaCl with an optimum at 3.0% (w/v). In addition, strain S2-6 T grew over a range of 0–20% O2 and the optimum growth occurred at 4% O2. Under optimal conditions, the cell doubling time of strain S2-6 T was 5.3 h (Table 1).
The chemoautotrophic growth tests showed that strain S2-6 T could grow with thiosulfate, tetrathionate, elemental sulfur or sodium sulfide as the sole energy source, and oxygen or nitrate as the sole electron acceptor. Molecular hydrogen could also be used as an energy source. When hydrogen was used as the sole energy source, strain S2-6 T could grow with oxygen, nitrate or element sulfur as electron acceptors. The end product of nitrate reduction was N2, and nitrite or ammonia did not accumulate; the product of sulfur compounds oxidation was sulfate; the product of elemental sulfur reduction was hydrogen sulfide. Inorganic nitrogen sources tests showed strain S2-6 T could grow with ammonium as the nitrogen sources, but not use sodium nitrate, sodium nitrite and molecular nitrogen. Selenium, tungsten and vitamins supplementation were not required for growth. Heterotrophic growth tests showed that none of the organic compounds sustained the growth of strain S2-6 T as the carbon or energy source. These results confirmed that strain S2-6 T is a typical chemolithoautotroph.
The major cellular fatty acids of strain S2-6 T were summed feature 3 (C16:1ω7c and/or C16:1ω6c), C16:0 and summed feature 8 (C18:1ω7c and/or C18:1ω6c) (Table 1), which were similar to those of S. paralvinellae GO25T, S. indica NW8NT and S. xiamenensis 1-1NT and distinctly different from those of S. autotrophica OK10T and Ca. S. hongkongensis AST-10. The significant differences between strains S2-6 T and S. paralvinellae GO25T were that the fatty acid C16:1 ω5c was observed at 1.3% in strain S2-6 T, but it was not found in strain S. paralvinellae GO25T. In addition, the fatty acid C18:0 was detected at 1.5% in strain S2-6 T, while it was only trace level of 0.7% in strain S. paralvinellae GO25T.
Genome annotation and analysis
The complete genome of strain S2-6 T consisted of a single circular chromosome with a total length of 2,320,257 bp long with a G + C content of 37.3 mol%. No plasmids were detected. Genome contained 2,445 predicted genes including 2,344 protein-coding genes, 59 rRNAs (four 5S rRNA, four 16S rRNA, and four 23S rRNA), 44 tRNAs, and 3 ncRNAs. Among the all protein-coding gene sequences, 1,755 genes were mapped over to 20 functional categories of COG database, while the remaining 589 genes could not be assigned to any categories. As shown in Fig. S3, the major categories were energy production and conversion (8.43%); replication, recombination and repair (8.21%); translation, ribosomal structure and biogenesis (7.75%); amino acid transport and metabolism (7.58%); cell wall/membrane/envelope biogenesis (6.61%); function unknown (6.04%); coenzyme transport and metabolism (5.87%); signal transduction mechanisms (5.75%); posttranslational modification, protein turnover, chaperones (5.07%) and inorganic ion transport and metabolism (5.01%).
No CRISPR loci were found, while 13 genomic islands (GI) of a total length of 173.8 kb were detected (Table 2). The size of the 13 putative islands ranged from 3,460 bp (GI 7) to 28,629 bp (GI 6). The largest GI 6 contained 44 genes, whereas the smallest GI 7 had 7 genes. The vast majority of the genes located on the genomic islands encode proteins annotated as hypothetical proteins. In these 13 GIs, 269 CDS were identified, including CDS encoding mobile element protein, transcriptional regulator, ribosomal proteins, ABC transport family systems, glycosyltransferase protein family, DNA polymerase, type I (II) restriction enzyme, two-component system, carbohydrate metabolism, thioredoxin reductase, iron-sulfur assembly protein, oxidoreductases for metabolism and so on. Among these GIs, seven contain mobile genetic elements, such as integrase and transposase genes, suggesting that these GIs can self-mobilize and could also support potential active horizontal gene transfer in the strain. Meanwhile, one of genomic islands carried genes encoding assimilatory sulfate reduction pathway including sulfate adenylyltransferase (Sat and CysDN) and adenylylsulfate kinase (CysC) (Table 2). The detection of sulfate reducing genes on GI indicates the contribution of mobile elements in the adaptation of bacteria to the environment and in active participation in the sulfur cycle.
The whole-genome characteristics were generally accordant with the main metabolic features experimentally demonstrated in strain S2-6 T. The genome of strain S2-6 T possessed all genes essential for carbon fixation via the reductive citric acid cycle (rTCA), such as ATP-dependent citrate lyase (aclAB, EC 2.3.3.8), 2-oxoglutarate:ferredoxin oxidoreductase (oorABCD, EC 1.2.7.3) and pyruvate:ferredoxin oxidoreductase (porABCD, EC 1.2.7.1) but not for the Calvin-Benson cycle. Strain S2-6 T could grow with hydrogen as an energy source. Genomic analysis revealed that strain S2-6 T contained a complete gene cluster encoding group 1, group 2 and group 4 [NiFe]-Hydrogenases. The ability to use oxygen as a terminal electron acceptor can be ensured by the presence of a gene cluster of cytochrome c oxidases cbb3-type that encode the FixNOQP proteins. Notably, the genome contained all genes required for the complete reduction of nitrate to N2 including nitrate reductases (napAGHBFLD), nitrite reductases (nirS, EC:1.7.2.1), nitric oxide reductases (norBC, EC:1.7.2.5) and nitrous oxide reductases (nosZ, EC:1.7.2.4).
As for sulfur metabolism, the genome of strain S2-6 T contained homologs for genes encoding soxAZYXBCD, all of which are required for assembling a fully functional complex that oxidizes reduced sulfur compounds. In addition, the sox genes in strain S2-6 T were present in two clusters, soxXYZAB (soxX, EC:2.8.5.2; soxA, EC:2.8.5.2; soxB, EC:3.1.6.20) and soxZYCD (soxD, EC:1.8.2.6). The genes encoding sulfide: quinone oxidoreductase (sqr, EC:1.8.5.4), participating in oxidizing sulfide to elemental sulfur, was also found in the genome. In addition, homologs of genes encoding known enzyme systems of reduced sulfur compounds oxidation such as sulfur oxygenase/re-ductase (Sor, EC:1.8.2.1) was present in genome of strain S2-6 T. No genes involved in dissimilatory sulfate reduction, such as adenylylsulfate reductase (aprAB, EC:1.8.99.2) and sulfite reductase (dsrAB, EC:1.8.99.5), and assimilatory sulfate reduction, such as phosphoadenosine phosphosulfate reductase (cysH, EC:1.8.4.10) and sulfite reductase (cysI, EC:1.8.1.2), were detected in this bacterium. In addition, strain S2-6 T can grow with elemental sulfur as the elector acceptor, which was consistent with the observe of polysulfide reductase (psr) genes found in the genome (Wang et al. 2020a).
Taxonomic conclusion
Characteristics of S2-6 T and species of the genus Sulfurimonas with validly published names are summarized in Table 1. There are many similar features between strain S2-6 T and the closely relatives of genus Sulfurimonas including Gram-stain negative and rod-shaped, with chemotaxonomic characteristics such as summed feature 3 (C16:1ω7c and/or C16:1ω6c), C16:0 and summed feature 8 (C18:1ω7c and/or C18:1ω6c) as the major fatty acid. The DNA G + C content of strain S2-6 T was 37.3 mol% and falls within the range (33.2–38.8 mol%) reported for members of the genus Sulfurimonas. Strain S2-6 T fell within the cluster comprising the genus Sulfurimonas and was most closely related to S. paralvinellae GO25T (96.8% sequence identity) and S. autotrophica OK10T (95.8% sequence identity). Strain S2-6 T exhibited many phenotypic similarities to S. paralvinellae GO25T, including morphology and the ability to grow under atmospheric concentrations of oxygen. However, there were many differences between strain S2-6 T and S. paralvinellae GO25T in some aspects (Table 1). The utilization patterns of electron donors are significantly different because that strain S2-6 T can use sulfide and tetrathionate as electron donors, whereas strain S. paralvinellae GO25T can not. Strain S2-6 T can grow well even at 20% (v/v) oxygen concentrations in the headspace gas, while strain S. paralvinellae GO25T only grow up to 10% partial pressure of O2. In addition, strain S. paralvinellae GO25T is a cold-tolerant bacterium and can grow well at 4 °C, while strain S2-6 T can not. The ANI and DDH values are below the species threshold. These data clearly indicate that strain S2-6 T can be differentiated from S. paralvinellae GO25T at the species level. In conclusion, strain S2-6 T should represent a novel species of the genus Sulfurimonas, for which the name Sulfurimonas sediminis sp. nov. is proposed.
Description of Sulfurimonas sediminis sp. nov.
Sulfurimonas sediminis (se. di’mi. nis, L. gen. n. sediminis of sediment).
Cells are Gram-stain-negative, rod shaped (0.8–2.0 µm long and 0.5–1.0 µm in width) and motile by a polar flagellum. Facultatively anaerobic and microaerophilic. Growth occurs at 10–45 °C (optimum 33 °C), pH 5.0–8.0 (optimum pH 7.0), and 1.5–6.0% (w/v) NaCl (optimum 3.0% (w/v)). Obligate chemolithoautotrophic growth occurs with H2, So, thiosulfate, tetrathionate and sulfide as electron donors, and nitrate, O2 or So as the sole electron acceptor. The isolate was an obligate chemolithoautotroph capable of growth using thiosulfate, tetrathionate, elemental sulfur or sodium sulfide as the sole energy source, and oxygen or nitrate as the sole electron acceptor. When hydrogen was used as the energy source, strain S2-6 T could respire oxygen, nitrate or element sulfur. Organic substrates are not utilized as carbon sources and energy sources. Ammonium is utilized as nitrogen source. Vitamins, selenium and tungsten are not required for growth. Major cellular fatty acids are summed feature 3 (C16:1ω7c and/or C16:1ω6c), C16:0 and summed feature 8 (C18:1ω7c and/or C18:1ω6c).
The type strain, S2-6 T (= MCCC 1A14513T = KCTC 15854 T) was isolated from the deep-sea sediment samples at the Longqi hydrothermal system, southwestern Indian Ocean. The DNA G + C content of the type strain is 37.3 mol%. The GenBank accession number for the 16S rRNA gene sequence is MN080871 and the complete genome accession number is CP041235.
Data availability statement
The GenBank accession numbers for the 16S rRNA gene sequence and complete genome sequence of Sulfurimonas sediminis S2-6 T are MN080871 and CP041235, respectively.
Abbreviations
- MCCC:
-
Marine Culture Collection of China
- KCTC:
-
Korean Collection for Type Cultures
- ANI:
-
Average nucleotide identity
- DDH:
-
DNA-DNA hybridization
References
Balch WE, Magrum LJ, Fox GE, Woese CR, Wolfe RS (1979) Methanogens: a re-evaluation of a unique biological group. Microbiol Rev 43:260–296
Bertelli C, Laird MR, Williams KP (2017) IslandViewer 4: expanded prediction of genomic islands for larger-scale datasets. Nucleic Acids Res 45:W30–W35
Cai L, Shao MF, Zhang T (2014) Non-contiguous finished genome sequence and description of Sulfurimonas hongkongensis sp. nov., a strictly anaerobic denitrifying, hydrogen- and sulfur-oxidizing chemolithoautotroph isolated from marine sediment. Stand Genomic Sci 9:1302–1310
Cao H, Wang Y, Lee OO, Zeng X. Shao Z, Qian P (2014) Microbial sulfur cycle in two hydrothermal chimneys on the Southwest Indian Ridge. Mbio 5.
Chun J, Oren A, Ventosa A, Christensen H, Arahal DR, da Costa MS, Rooney AP, Yi H, Xu X, Meyer SD, Trujillo ME (2018) Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol 68:461–466
Delcher AL, Bratke KA, Powers EC, Salzberg SL (2007) Identifying bacterial genes and endosymbiont DNA with Glimmer. Bioinformatics (Oxford, England) 23:673–679
Dick GJ (2019) The microbiomes of deep-sea hydrothermal vents: distributed globally, shaped locally. Nat Rev Microbiol 17:271–283
Ding J, Zhang Y, Wang H, Jian H, Leng H, Xiao X (2017) Microbial community structure of deep-sea hydrothermal vents on the ultraslow spreading Southwest Indian Ridge. Front Microbiol 8:1012
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evo 17:368–376
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Frolov EN, Kublanov IV, Toshchakov SV, Samarov NI, Novikov AA, Lebedinsky AV, Bonch-Osmolovskaya EA, Chernyh NA (2017) Thermodesulfobium acidiphilum sp. nov., a thermoacidophilic, sulfate-reducing, chemoautotrophic bacterium from a thermal site. Int J Syst Evol Microbiol 67:1482–1485
Grissa I, Vergnaud G, Pourcel C (2007) CRISPRFinder: a web tool to identify clustered regularly interspaced short palindromic repeats. Nucleic Acids Res 35:W52–W57
Han Y, Perner M (2015) The globally widespread genus Sulfurimonas: versatile energy metabolisms and adaptations to redox clines. Front Microbiol 6:989
Henkel JV, Vogts A, Werner J, Neu TR, Spröer C, Bunk B, Schulz-Vogt HN (2020) Candidatus sulfurimonas marisnigri sp. nov. and candidatus sulfurimonas baltica sp. nov. thiotrophic manganese oxide reducing chemolithoautotrophs of the class campylobacteria isolated from the pelagic redoxclines of the black sea and the baltic sea. Syst Appl Microbiol 44.
Hoor TTA (1975) A new type of thiosulphate oxidizing, nitrate reducing microorganism: Thiomicrospira denitrificans sp. nov. Net J Sea Res 9:344–351
Hu QT, Wang SS, Lai QL, Shao ZZ, Jiang LJ (2020) Sulfurimonas indica sp. nov., a hydrogen- and sulfur- oxidizing chemolithoautotroph isolated from a hydrothermal sulfide chimney in the Northwest Indian Ocean. Int J Syst Evol Microbiol.
Inagaki F, Takai K, Kobayashi H, Nealson KH, Horikoshi K (2003) Sulfurimonas autotrophica gen. nov., sp. nov., a novel sulfur-oxidizing ε-proteobacterium isolated from hydrothermal sediments in the Mid-Okinawa Trough. Int J Syst Evol Microbiol 53:1801–1805
Jiang LJ, Zheng YP, Peng XT, Zhou HY, Zhang CL, Xiao X, Wang FP (2010) Vertical distribution and diversity of sulfate-reducing prokaryotes in the Pearl River estuarine sediments, Southern China. Fems Microbiol Ecol 70:93–106
Jiang LJ, Lyu J, Shao ZZ (2017) Sulfur metabolism of Hydrogenovibrio thermophilus strain S5 and its adaptations to deep-sea hydrothermal vent environment. Front Microbiol 8:2513
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Labrenz M, Grote J, Mammitzsch K, Boschker HTS, Laue M, Jost G, Glaubitz S, Jurgens K (2013) Sulfurimonas gotlandica sp. nov., a chemoautotrophic and psychrotolerant epsilonproteobacterium isolated from a pelagic redoxcline, and an emended description of the genus Sulfurimonas. Int J Syst Evol Microbiol 63:4141–4148
Lagesen K, Hallin P, Rødland EA, Staerfeldt HH, Rognes T, Ussery DV (2007) RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res 35:3100–3108
Liu C, Shao Z (2005) Alcanivorax dieselolei sp. nov., a novel alkane-degrading bacterium isolated from sea water and deep-sea sediment. Int J Syst Evol Microbiol 55:1181–1186
Lowe TM, Chan PP (2016) tRNAscan-SE On-line: integrating search and context for analysis of transfer RNA genes. Nucleic Acids Research W1
Luo R, Liu B, Xie Y, Li Z, Huang W, Yuan J, He G, Chen Y, Pan Q, Liu Y, Tang J, Wu G, Zhang H, Shi Y, Liu Y, Yu C, Wang B, Lu Y, Han C, Cheung D W, Yiu S M, Peng S, Zhu X, Liu G, Liao X, Li Y., Yang H, Wang J, Lam T W, Wang J (2012) SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1
McNichol J, Stryhanyuk H, Sylva SP, Thomas F, Musat N, Seewald JS, Sievert SM (2018) Primary productivity below the seafloor at deep-sea hot springs. J Proc N atl Acad Sci USA 115:6756–6761
Na SI, Kim YO, Yoon SH, Ha SM, Baek I, Chun J (2018) UBCG: up-to-date bacterial core gene set and pipeline for phylogenomic tree reconstruction. J Microbiol 56:280–285
Nadine LB, Yucel M, Das A, Sievert SM, Girguis PR (2018) Hydrothermal energy transfer and organic carbon production at the deep seafloor. Front Microbiol 5:392–531
Ratnikova NM, Slobodkin AI, Merkel AY, Kopitsyn DS, Kevbrin VV, Bonch-Osmolovskaya EA, Slobodkina GB (2019) Sulfurimonas crateris sp. nov., a facultative anaerobic sulfur-oxidizing chemolithoautotrophic bacterium isolated from a terrestrial mud volcano. Int J Syst Evol Microbiol, 1–6
Rzhetsky A, Nei M (1992) Statistical properties of the ordinary least-squares, generalized least-squares, and minimum-evolution methods of phylogenetic inference. J Mol Evol 35:367–375
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Takai K, Horikoshi K (2000) Thermosipho japonicus sp. nov., an extremely thermophilic bacterium isolated from a deep-sea hydrothermal vent in Japan. Extremophiles 4:9–17
Takai K, Suzuki MT, Nakagawa S, Miyazaki M, Suzuki Y, Inagaki F, Horikoshi K (2006) Sulfurimonas paralvinellae sp. nov., a novel mesophilic, hydrogen- and sulfur-oxidizing chemolithoautotroph within the Epsilonproteobacteria isolated from a deep-sea hydrothermal vent polychaete nest, reclassification of Thiomicrospira denitrificans as Sulfurimonas denitrificans comb. nov. and emended description of the genus Sulfurimonas. Int J Syst Evol Microbiol 56:1725–1733
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol Biol Evol, pp 2725–2729
Waite DW, Vanwonterghem I, Rinke C, Parks DH, Zhang Y, Takai K, Sievert SM, Simon J, Campbell BJ, Hanson TE, Woyke T, Klotz MG, Hugenholtz P (2017) Comparative genomic analysis of the class Epsilonproteobacteria and proposed reclassification to Epsilonbacteraeota (phyl. nov.). Front Microbiol 8:682
Waite DW, Vanwonterghem I, Rinke C, Parks DH, Zhang Y, Takai K, Sievert SM, Simon J, Campbell BJ, Hanson TE, Woyke T, Klotz MG, Hugenholtz P (2018) Addendum: Comparative genomic analysis of the class Epsilonproteobacteria and proposedreclassification to Epsilonbacteraeota (phyl. nov.). Front Microbiol 9:772
Wang SS, Jiang LJ, Liu XW, Yang SP, Shao ZZ (2020) Sulfurimonas xiamenensis sp. nov. and Sulfurimonas lithotrophica sp. nov., hydrogen- and sulfur-oxidizing chemolithoautotrophs within the Epsilonproteobacteria isolated from coastal sediments, and an emended description of the genus Sulfurimonas. Int J Syst Evol Microbiol 70:2657–2663
Wang S, Jiang L, Hu Q, Liu X, Yang S, Shao Z (2020a) Elemental sulfur reduction by a deep-sea hydrothermal vent Campylobacterium Sulfurimonas sp. NW10. Environ Microbiol
Yoon SH, Ha SM, Lim J, Kwon S, Chun J (2017) A large-scale evaluation of algorithms to calculate average nucleotide identity. Anton Van Leeuwen 110:1281–1286
Acknowledgements
This work was financially supported by the National Key R&D Program of China (No. 2018YFC0310701), National Natural Science Foundation of China (No. 41672333) and COMRA program (No. DY135-B2-01).
Funding
This work was financially supported by the National Key R&D Program of China (No. 2018YFC0310701), National Natural Science Foundation of China (No. 41672333) and COMRA program (No. DY135-B2-01).
Author information
Authors and Affiliations
Contributions
LJ and SY supervised the project. SW, XL and SX carried out the experiments. SW, QL and LJ analyzed the data. SW, SY and ZS wrote and revised the manuscript. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, S., Shao, Z., Lai, Q. et al. Sulfurimonas sediminis sp. nov., a novel hydrogen- and sulfur-oxidizing chemolithoautotroph isolated from a hydrothermal vent at the Longqi system, southwestern Indian ocean. Antonie van Leeuwenhoek 114, 813–822 (2021). https://doi.org/10.1007/s10482-021-01560-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-021-01560-4