Abstract
Gracilibacillus dipsosauri is a moderately-halophilic Gram-positive bacterium which forms an extracellular α-amylase that is induced by starch, repressed by d-glucose, and active in 2.0 M KCl. Previous studies showed that while enzyme activity could be measured with the synthetic substrate 2-chloro-4-nitrophenyl-α-d-maltotrioside (CNPG3), other assays were inconsistent and the protein showed aberrant mobility during nondenaturing gel electrophoresis. To clarify the properties of this enzyme, the genome of G. dipsosauri was sequenced and was found to be 4.19 Mb in size with an overall G+C content of 36.9%. A gene encoding an α-amylase composed of 691 amino acids was identified. The protein was a member of the glycosyl hydrolase 13 family, which had a molecular mass of 77,396 daltons and a pI of 4.39 due to an unusually large number of aspartate and glutamate residues (95/691 or 13.7%). BLAST analysis of the amino acid sequence revealed significant matches to other proteins with cyclodextrin glycosyltransferase activity. Partial purification of the protein from G. dipsosauri showed that fractions catalyzing the hydrolysis of CNPG3 and p-nitrophenyl-d-maltoheptoside also catalyzed the formation of β-cyclodextrin but not α-cyclodextrin or γ-cyclodextrin. Formation of β-cyclodextrin was not stimulated by high salt concentrations but did occur with rice, potato, wheat, and corn starches and amylopectin. These studies explain the unusual features of the α-amylase from G. dipsosauri and indicate it should be classified as EC 2.4.1.19. The availability of the complete genomic sequence of G. dipsosauri will provide the basis for studies on other enzymes from this halophile which may be useful for biotechnology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microorganisms that have evolved to grow in habitats characterised by high temperatures, extremely acidic or alkaline pH, high salt concentrations, and high pressures have been termed extremophiles (Madigan and Marrs 1997; Horikoshi and Grant 1998; Gerday and Glansdorff 2007; Canganella and Wiegel 2011). They have been investigated as models for life outside of the planet Earth (Rothschild and Mancinelli 2001; Merino et al. 2019), as potential contributors to bioremediation on the Earth (Orellana et al. 2018), and as sources of proteins for biotechnology (Hough and Danson 1999; Niehaus et al. 1999; Schiraldi and De Rosa 2002; Dalmaso et al. 2015; Krüger et al. 2018). Among the enzymes isolated and purified from extremophilic bacteria and archaea are amylases, cellulases, chitinases, proteases, and DNA polymerases.
Gracilibacillus dipsosauri strain DD1 is a moderately halophilic bacterium that was originally isolated from the nasal cavity of a desert iguana (Deutch 1994). Strain DD1 was identified as the type strain of new species of Bacillus based on its 16S rRNA sequence and shown to be loosely associated with other salt-tolerant bacteria such as Sporosarcina halophila and Marinococcus albus (Lawson et al. 1996). It was later reclassified as Gracilibacillus dipsosauri after reorganization of several halotolerant genera (Wainø et al. 1999). Other species of Gracilibacillus have subsequently been identified and linked phylogenetically to G. dipsosauri including G. orientalis (Carrasco et al. 2006), G. boraciitolerans (Ahmed et al. 2007), G. quingheiensis (Chen et al. 2008a), G. lacisalsi (Jeon et al. 2008), G. halophilus (Chen et al. 2008b), G. saliphilus (Tang et al. 2009), G. thailandensis (Chamroensaksri et al. 2010), G. ureilyticus (Huo et al. 2010), G. kekensis (Gao et al. 2012), G. bigeumensis (Kim et al. 2012), G. xinjiangensis (Yang et al. 2013), G. marinus (Huang et al. 2013), G. alcaliphilus (Hirota et al. 2014), G. kimchi (Oh et al. 2016), G. massiliensis (Diop et al. 2016), G. aidingensis (Guan et al. 2017), G. phocaeensis (Senghor et al. 2017), G. eburneus (Guan et al. 2018), and G. timonensis (Diop et al. 2018). The current edition of Bergey’s Manual of Systematics of Archaea and Bacteria Online (Editorial Board 2015) contains a summary of the genus Gracilibacillus and shows the relationship of these bacteria to others in the order Bacillales. Several recent papers illustrate the phylogenetic relationships of G. dipsosauri and other species in this genus based on their 16S rRNA sequences (Senghor et al. 2017; Diop et al. 2018; Guan et al. 2018).
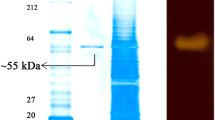
In its initial characterisation, G. dipsosauri strain DD1 gave positive results for the hydrolysis of starch, triacylglycerides, esculin, and o-nitrophenyl-β-d-galactoside (Deutch 1994). As part of a long-term project to purify enzymes from this microorganism which can function in high salt concentrations, an extracellular enzyme capable of degrading starch was isolated and described as an α-amylase (Deutch 2002). The enzyme produced clear zones of hydrolysis on agar plates containing soluble starch, was induced by starch but repressed by d-glucose, and showed its highest activity after growth of the bacterium in 1.0 M KCl. The enzyme had a pH optimum of 6.5, was stable at 60 °C, and was stimulated by 1.0 M Na2SO4. While the enzyme resembled other α-amylases, it had several unusual properties. First, although its activity could be measured quantitatively using 2-chloro-4-nitrophenyl-α-d-maltotrioside (CNPG3) as the substrate (Gella et al. 1997; Suganuma et al. 1997; Foo and Bais 1998; Lorentz et al. 1999), assays based the formation of reducing sugars like d-glucose (Bernfeld 1955), binding of iodine to starch (Laderman et al. 1993), or hydrolysis of dye-labeled polymers (McCleary 1988) were inconsistent. Second, the purified enzyme had a denatured molecular mass in SDS-polyacrylamide gel electrophoresis of about 80 kDa but behaved in non-denaturing gels as if it had a mass of about 30 kDa. Third, the purified enzyme was active in high concentrations of KCl or NaCl (up to 2.0 M) and was stable in salt for long periods of time. Attempts to clone the gene for the enzyme from G. dipsosauri genomic DNA into plasmids such as pGEM-3Zf(+) for E. coli or to amplify the gene using primers based on salt-tolerant α-amylase sequences from Halobacillus halophilus, Oceanobacillus iheyensis, and Gracilibacillus halophilus between 2005 and 2013 were unsuccessful. To clarify the properties of this novel α-amylase, the sequence of the G. dipsosauri genome has now been determined and the properties of the gene encoding this enzyme analyzed. We now report that the α-amylase activity in strain DD1is associated with a protein that can also function as a cyclodextrin glycosyltransferase (CGTase).
Materials and methods
Sequencing of the G. dipsosauri genome
The DNA of G. dipsosauri strain DD1 was sequenced and assembled using a set of basic methods that have now become widely used (Ekblom and Wolf 2014; Simpson and Pop 2015). Bacteria were grown overnight in 25 ml of Difco™ tryptic soy broth containing 1.0 M KCl at 37 °C with aeration in a shaker-incubator at 250 rpm. Genomic DNA was extracted using the reagents in a Zymo Research Fungal/Bacterial Mini-prep kit as directed by the manufacturer (Zymo Research, Tustin, CA USA). The preparation had a DNA concentration of 182 ng/µl with a A260/A280 ratio of 1.85. The sample was sent to the Center for Genomics and Bioinformatics at Indiana University, where sequencing was done with an Illumina MiSeq system. There were 986,585 paired reads, with a total of about 592,154,880 bases. The raw Illumina MiSeq 2x300 bp reads were quality checked using FastQC v0.10.1 (https://www.bioinformatics.babraham.ac.uk/projects/fastqc), followed by adapter trimming and quality clipping by Trimmomatic 0.35 (http://www.usadellab.org/cms/?page=trimmomatic). Any reads shorter than 150 bp were dropped. Any reads with start, end, or the average quality within a 4 bp window falling below quality scores of 15 were trimmed. A clean set of 698,667 read pairs was saved for further insert size estimation. Kmer analysis was run by Jellyfish 2.2.4 (https://www.cbcb.umd.edu/software/jellyfish/) over both the entire 986,933 read pairs and the clean 698,667 read pairs for genome size estimation. Clean reads were aligned to the G. orientalis reference genome (https://www.ncbi.nlm.nih.gov/assembly/GCF_900114645.1/) by bwa mem 0.7.15 for insert size estimation. Spades 3.11.1 (http://cab.spbu.ru/software/spades/) with mismatch corrector mode was applied for whole-genome assembly with kmer sizes of 21, 33, 55, and 77. The best whole genome assembly with kmer size 77 was evaluated by comparing it to G. orientalis reference genome by Quast 4.5 (http://bioinf.spbau.ru/quast). When sorting contigs from largest to smallest, the first 28 contigs with minimum length of 1000 bp were extracted. CAR, a novel reference-based contig assembly and scaffolding tool (http://genome.cs.nthu.edu.tw/CAR/), was applied to the 28 contigs for scaffolds. 21 contigs were kept in the final four scaffolds. Quast was used again to evaluate the assembly with 28 contigs and 21 contigs. BUSCO, a tool for assessing genome assembly and annotation completeness with benchmarking universal single copy orthologs (http://busco.ezlab.org/), indicated 98.7% genome completeness with 146 complete BUSCOs out of 148 total BUSCOs defined in bacteria database. A total of 1602 genes were predicted de novo by Glimmer (Gene Locator and Interpolated Markov Model ER https://ccb.jhu.edu/software/glimmer/). Additional genome annotation was performed by a protein homology-based tblastn (blast + 2.3.0) approach using protein sequences of the Gracilibacillus orientalis reference genome. Genes predicted de novo were compared to genes annotated by tblastn, of which 1311 matched. The assembly gave an average coverage across the genome of 141x. The complete genome has been deposited at the National Center for Biological Information (NCBI Reference Sequence: NZ_QGTD00000000.1).
Assays for α-amylase and cyclodextrin glycosyltransferase activities
The α-amylase activity of the G. dipsosauri enzyme with 2-chloro-4-nitrophenyl-α-d-maltotrioside (CNPG3) as the substrate was measured as previously described (Deutch 2002). Briefly, 1.0 ml reaction mixtures contained 50 mM potassium MES buffer, pH 6.1, 5 mM CaCl2·2H2O, 200 mM KSCN, 1.0 M Na2SO4, 2.25 mM CNPG3 (Toronto Research Chemicals, Toronto Canada), and varying volumes of an enzyme solution. The reactions were incubated at room temperature in a Shimadzu U-160 spectrophotometer and the absorbance at 405 nm measured continuously for 10–30 min. Activity was expressed as nmol/min ml using a molar extinction coefficient of 14.6 × 103 for 2-chloro-4-nitrophenol (Winn-Deen et al. 1988). Amylase activity with p-nitrophenyl-d-maltoheptoside (PNPG7) as the substrate (Sheehan and McCleary 1988) was measured using an assay kit from TECO Diagnostics (Anaheim, CA USA). The reagent powder was dissolved in 6.0 ml distilled water and 1.0 ml portions combined with varying volumes of an enzyme solution. The absorbance was measured at 405 nm in a Shimadzu U-160 spectrophotometer for 10 min and the activity in International Units/liter was calculated as directed by the manufacturer. Cyclodextrin glycosyltransferase activity leading to the formation of β-cyclodextrin was measured by a modification of the procedures described by Goel and Nene (1993) and Basappa et al. (1998). Enzyme solutions were diluted 1/10 into a substrate mixture containing 50 mM potassium MES, pH 6.1, 5 mM CaCl2.2H2O, 1.0 M KCl, and 1% (w/v) soluble starch (Difco Laboratories, Detroit, MI USA). The reactions were incubated at room temperature for up to 60 min and terminated by heating in a heat block at 95 °C for 5 min. Samples were stored at − 20 °C, thawed, and tested for β-cyclodextrin by adding 100 µl portions to 900 µl of a working phenolphthalein solution containing 0.1 mM phenolphthalein (Biopharm Inc., Hatfield, AR USA) in 125 mM Na2CO3. The decrease in absorbance relative to the absorbance of the stock substrate solution was measured immediately at 550 nm in the Shimadzu U-160 spectrophotometer. The amount of β-cyclodextrin was calculated using a linear standard curve covering the range of 0–300 µg of pure β-cyclodextrin (L’eternel LLC, Aurora, OH USA). For some assays, the KCl in the substrate mixture was omitted or replaced by other salts. For other assays, the 1% soluble starch was replaced by other purified starch samples including rice, potato, wheat, and corn. Reactions were also tested for the formation of α-cyclodextrin with methyl orange (Lejeune et al. 1989) and for the formation of γ-cyclodextrin with bromcresol green (Kato and Horikoshi 1984) using standard curves prepared from purified samples of α-cyclodextrin and γ-cyclodextrin (L’eternel LLC, Aurora OH USA). All assays were done in triplicate independently at least twice and varied by < 10%.
Partial purification of the G. dipsosauri strain DD1 enzyme
The α-amylase activity from G. dipsosauri strain DD1 was partially purified from culture fluid by a modification of the procedure previously described (Deutch 2002). The bacteria were grown overnight in 10 ml of Bacto tryptic soy broth without dextrose (Becton, Dickinson and Company, Sparks, MD USA) containing 1.0 M KCl and 0.1% (w/v) soluble starch (Difco Laboratories, Detroit, MI USA) at 37 °C with aeration in a shaker incubator at 250 rpm. The culture was diluted 1/50 into 300 ml of fresh medium and grown overnight again at 37 °C with aeration. The bacteria were removed by centrifugation at 10,000×g in a Bio-Lion XC-H165 centrifuge and the culture fluid stored at 4 °C. Two volumes of ice-cold 95% ethanol were added slowly to 100 ml of the culture fluid and the mixture stored at − 20 °C for 2 h. The proteins were collected by centrifugation at 10,000 x g in a Bio-Lion XC-H165 centrifuge and resuspended in 10 ml of 50 mM potassium MES buffer, pH 6.1. The suspension was stored overnight at 4 °C and centrifuged again to remove any residual starch and the supernatant saved as the concentrated culture fluid.
To further purify the protein, a 10 ml portion of the concentrated culture fluid was applied to a 2.5 × 20 cm column of Bio-Gel P-6 desalting gel (Bio-Rad Laboratories, Hercules, CA USA) and 200-drop fractions collected during elution with 50 mM potassium MES buffer, pH 6.1. The fractions were tested for absorbance at 280 nm. Those in the void volume, as determined with a solution of blue dextran, were tested for α-amylase activity with CNPG3 and PNPG7 as substrates and for the formation of β-cyclodextrin. The four most active fractions were pooled and retested for all three activities. The solution was concentrated by centrifugation at 5000×g through a Millipore 3000 MWCO filter (Millipore Corporation, Cork Ireland) and 2.0 ml applied to a 1.5 × 15 cm column of Fast Flow DEAE-Sepharose (GE Healthcare, Pittsburgh, PA USA). Fractions (200-drop) were collected during elution with 50 mM potassium MES buffer, pH 6.1 and tested for absorbance at 280 nm, for α-amylase activity with CNPG3 and PNPG7 as substrates, and for the formation of β-cyclodextrin. The two most active fractions were pooled and retested for all three activities.
Protein assays
Protein concentrations were determined by the bicinchoninic acid method (Smith et al. 1985) using reagents from Gene and Cell Technologies (Richmond, CA USA) and bovine serum albumin as the standard.
Results and discussion
General features of the G. dipsosauri genome
The genome of G. dipsosauri was observed to be 4.19 Mb in size with an overall G+C content of 36.9%. It was composed 4052 genes, 3941 of which appeared to code for proteins. Table 1 shows a summary of all of the Gracilibacillus genomes currently available at the NCBI. Phylogenetic analyses of the various Gracilibacillus species based on 16S rRNA sequences have indicated that G. dipsosauri is most closely related to G. ureilyticus (Senghor et al. 2017; Diop et al. 2018; Guan et al. 2018). The next most-closely related lineage includes G. orientalis and G. phocaeensis. The genome of G. dipsosauri is similar in size and composition to that of G. ureilyticus, but both of these genomes are smaller than those of G. orientalis and G. phocaeensis.
Characterization of the G. dipsosauri α-amylase gene and protein
Analysis of the complete genome of G. dipsosauri revealed the presence of a genetic locus (DLJ74_06065) in node 2 (nucleotides 438287-440362) which appeared to code for an α-amylase of 691 amino acids. It is described as PWU69658.1 in the NCBI Protein database (https://www.ncbi.nlm.nih.gov/protein/PWU69658.1) and as A0A317L430 in the UniProtKB database (https://www.uniprot.org/uniprot/?query=accession:A0A317L430). The complete amino acid sequence is shown in Fig. 1 along with a graphical representation of the conserved domains and a model of its three-dimensional structure. The protein appeared to be a member of the glycosyl hydrolase 13 family. The predicted molecular mass of the protein was 77,396 daltons, the predicted pI was 4.39, and the protein was hypothesized to fold into a compact structure with 59.9% α-helixes and 43.6% β-strands. BLAST analysis of the amino acid sequence revealed significant matches to an α-amylase from Paraliobacillus ryukyuensis (87% identity), an α-amylase from Bacillus sp. FJAT-45505 (73% identity), an α-amylase from Gracilibacillus orientalis (70% identity), and a cyclodextrin glycosyltransferase (CGTase) from Bacillus cereus (66% identity). There were many matches to other CGTases in the range of 55% to 65% identity. Of the other Gracilibacillus species that have been identified, only some show a positive reaction for starch hydrolysis including G. kekensis, G. boracitolerans, G. ureilyticus, G. saliphilus, G. orientalis, G. halophilus, and G. halotolerans (Gao et al. 2012) and are predicted to contain α-amylases. Supplementary figure S1 shows a multiple alignment of the α-amylase sequences from G. dipsosauri, G. halophilus, G. orientalis, G. kekensis, and G. ureilyticus. There are large variations in the sequences and only 4.27% overall identity.
Characteristics of the α-amylase protein from Gracilibacillus dipsosauri strain DD1. Panel a shows the amino acid sequence of the protein, which is described in more detail as PWU69658.1 in the NCBI Protein database (https://www.ncbi.nlm.nih.gov/protein/PWU69658.1) and as A0A317L430 in the UniProtKB database at the Swiss Institute of Bioinformatics (https://www.uniprot.org/uniprot/?query=accession:A0A317L430). Panel b shows an image of the conserved domains of the protein from the NCBI site. Triangles show conserved residues at the active site or for the binding of calcium ions or starch. Panel c shows a diagram of the possible three-dimensional structure of the protein as created by the Phyre2 program based similarities of the amino acid sequence to one from Thermoanerobacterium thermosulfuruigenes (Kelley et al. 2015)
Properties of the G. dipsosauri α-amylase protein
We used the predicted amino acid sequence of the G. dipsosauri α-amylase to answer three unresolved questions about this enzyme. First, why did the enzyme show good activity with 2-chloro-4-nitrophenylmaltotrioside (CNPG3) as the substrate but poor activity in other assays for α-amylase activity? BLAST analysis of the amino acid sequence revealed sequence similarities to other enzymes that have CGTase activity. These enzymes are members of the glycosyl hydrolase 13 family which catalyze the breakdown of starches to yield cyclic molecules with 6 (α-cyclodextrins), 7 (β-cyclodextrins), or 8 (γ-cyclodextrins) glycosyl units (Qi and Zimmermann 2005; Stam et al. 2006; Leemhuis et al. 2010). β-Cyclodextrin is the most common product and has been extensively investigated for its possible use in drug delivery (Duchêne and Bochot 2016; Jansook et al. 2018). Because cyclodextrins are internally linked, they behave as nonreducing sugars and do not react with reagents such as 3,5-dinitrosalicylic acid (Bernfeld 1955) that are often used in assays of α-amylase activity.
To determine if the enzyme from G. dipsosauri is a CGTase, the culture fluid from bacteria grown in tryptic soy broth without dextrose containing 1.0 M KCl and 0.5% soluble starch was tested for the formation of β-cyclodextrin using phenolphthalein as the colorimetric reagent and found to give a positive result. The same culture fluid also gave positive tests for the hydrolysis of 2-chloro-4-nitrophenyl-α-d-maltotrioside (CNPG3) and for the hydrolysis of p-nitrophenyl-d-maltoheptoside (PNGP7). The proteins in the culture fluid were concentrated by ethanol precipitation and retested for each enzyme activity. Although the units of activity with the three assays were different, the increase in activity after concentration was about 6-fold with each assay (Table 2). There was an increase in the specific activity with all three assays as peptides in the culture medium were removed. When the proteins in the concentrated culture fluid were separated on a column of Bio-Gel P-6 desalting gel, all three activities eluted in the same fractions (Fig. 2a). These fractions corresponded to the void volume, which would be the predicted location for a protein with a mass of about 80 kDa. When the proteins in the concentrated culture fluid were further purified on a column of Fast Flow DEAE-Sepharose, all three activities eluted in the same fractions (Fig. 2b). These results were consistent with the hypothesis that α-amylase and cyclodextrin glycosyl transferase activity are associated with the same protein.
Elution of proteins with α-amylase activity from G. dipsosauri strain DD1 during column chromatography. Panel a shows the elution of proteins in the ethanol precipitate of culture fluid from a column of Bio-Gel P-6 desalting gel. Panel b shows the elution of proteins in the desalting column pool from a column of DEAE-Sepharose. In each case, (○) indicates absorbance at 280 nm, (△) indicates activity as measured by hydrolysis of CNGP3, (□) indicates activity as measured by hydrolysis of PNPG7, and (◊) indicates activity as measured by the formation of β-cyclodextrin. Activities were determined as described in Materials and methods
Second, why did the protein with a predicted molecular mass of about 80 kDa, which behaved as expected during gel filtration chromatography on columns of Bio-Gel P-6 desalting gel and Sephadex G-100 or during electrophoresis in SDS-PAGE denaturing gels, exhibit an apparent molecular mass of about 30,000 in nondenaturing polyacrylamide gels? The explanation for this is that the predicted isoelectric point (pI) of the α-amylase was 4.39 and that 95/691 or 13.7% of the amino acids in its sequence were aspartate or glutamate. Under non-denaturing conditions in the standard Laemmli buffer system (pH 8.8 in running gel, pH 6.8 in stacking gel), the protein would be negatively-charged and would be expected to move rapidly towards the positive electrode with a smaller than expected apparent mass. Similar aberrations in mobility during gel electrophoresis have been observed before (Hames 1998).
Third, why did the α-amylase from G. dipsosauri have such a high tolerance to salts? It retained high activity in 2.0 M KCl, and was strongly stimulated by 1.0 M Na2SO4. Previous studies have shown that salt-tolerance is associated with a high ratio of acidic to basic residues and a small number of hydrophobic residues (Lanyi 1974; Madern et al. 2000; Graziano and Merlino 2014)). As noted above, 95/691 (13.7%) of the residues are aspartate or glutamate. By contrast, only 39/691 (5.6%) of the residues are arginine and lysine. Similarly, only 115 (16.6%) are phenylalanine, leucine, or isoleucine. It is interesting to note that the α-amylase of G. dipsosauri has only one cysteine residue and so disulfide bonds play no role in the stabilisation of the protein’s structure.
Characterisation of the G. dipsosauri cyclodextrin glycosyltransferase activity
To determine if the enzyme from G. dipsosauri can also catalyze the formation of α-cyclodextrin or γ-cyclodextrin in addition to β-cyclodextrin, the purified protein was also tested for the other activities using methyl orange (Lejeune et al. 1989) or bromcresol green (Kato and Horikoshi 1984) as the colorimetric reagent. There was no detectable α-cyclodextrin or γ-cyclodextrin after 60 min of incubation in a reaction mixture containing 1% soluble starch and 1.0 M KCl. There was no hydrolysis of p-nitrophenyl-α-d-glucopyranoside as a substrate in either the standard reaction mixture used for CNPG3 hydrolysis or in reaction mixtures containing just 0, 0.5 M, or 1.0 M KCl.
Previous studies of the α-amylase activity using CNPG3 as the substrate indicated that the enzyme was stimulated by increasing concentrations of KCl and almost four-fold by 1.0 M Na2SO4 (Deutch 2002). To determine if the formation of β-cyclodextrin is also stimulated by various salts, the CGTase activity of the purified protein was tested in reactions containing 1% soluble salt (Fig. 3a). Unlike the hydrolysis activity with CNPG3 as the substrate, the activity gradually decreased as the KCl concentration increased and was inhibited about 50% by 1.0 M Na2SO4. Studies of other CGTases has shown that these enzymes can act on a variety of different starches. To further characterise the enzyme from G. dipsosauri strain DD1, the partially purified protein was incubated in reaction mixtures containing without or with 1.0 M KCl and 1% (w/v) concentrations of Difco soluble starch, Sigma rice starch, Sigma potato starch, Sigma wheat starch, or Sigma corn starch. The activities with the purified starches as measured by the formation of β-cyclodextrin were about two times higher than that seen with the Difco soluble starch (Fig. 3b). 1n each case, addition of 1.0 M KCl caused a reduction in the activity of about 10%. The activity of the partially purified enzyme also was measured without or with 1.0 M KCl and 1% potato amylose or potato amylopectin as the substrate (Fig. 3c). The formation of β-cyclodextrin was much higher with amylopectin than with amylose, although this appeared to be related in part of the greater solubility of the latter compound. Again, addition of 1.0 M KCl caused a slight reduction in β-cyclodextrin formation. Previous studies did show large zone of hydrolysis with G. dipsosauri strain DD1 was streaked on agar plates containing amylopectin (Deutch 2002).
Formation of β-cyclodextrin by the purified α-amylase from G. dipsosauri strain DD1. Panel a shows the effect of different salt concentrations on the enzyme activity. Panel b shows the enzyme activity with different starches in the absence (light bars) and presence (dark bars) of 1.0 M KCl. Panel C shows the enzyme activity with 1% amylose or 1% amylopectin in the absence and presence of 1.0 M KCl. In each case, the bars shows the means ± one standard deviation based on three replicate assays in each experiment. The protein concentration of the purified fraction was 78 µg/ml
Conclusions
These studies clarify the properties of the salt-tolerant α-amylase from G. dipsosauri. Although the enzyme’s activity could be clearly demonstrated on agar plates containing soluble starch and measured quantitatively with the synthetic substrate CNPG3, other assays including those based on the formation of reducing sugars were inconsistent. The data presented here from an analysis of the gene sequence and from various biochemical tests indicate that the enzyme functions primarily as a cyclodextrin glycosyltransferase. It can degrade a variety of starches, but unlike the α-amylase activity measured with CNPG3 as the substrate, the CGTase activity is not stimulated by high salt concentrations. The protein has an actual molecular mass of about 78,000 daltons and a strongly acidic pI. It falls into subfamily GH13_2 as described by Stam et al. (2006) and thus should be classified as an example of EC 2.4.1.19. With the complete genomic sequence of G. dipsosauri now available, we are in a position to characterize other salt-tolerant enzymes from this bacterium with potential biotechnological applications.
References
Ahmed I, Yokota A, Fugiwara T (2007) Gracilibacillus boraciitolerans sp. nov., a highly boron tolerant and moderately halotolerant bacterium isolated from soil. Int J Sys Evol Microbiol 57:796–802
Basappa C, Rao P, Rao DN, Divakar S (1998) A modified colorimetric method for the estimation of β-cyclodextrin using phenolphthalein. Int J Food Sci Technol 33:517–520
Bernfeld P (1955) Amylases, α and β. Methods Enzymol 1:149–158
Canganella F, Wiegel J (2011) Extremophiles: from abyssal to terrestrial ecosystems and possibly beyond. Naturwissenschaften 98:253–279
Carrasco IJ, Márquez MC, Yanfen X, Ma Y, Cowan DA, Jones BE, Grant WD, Ventosa A (2006) Gracilibacillus orientalis sp. nov., a novel moderately halophilic bacterium from a salt lake in Inner Mongolia. China. Int J Syst Evol Microbiol 56:599–604
Chamroensaksri N, Tanasupawat S, Akaracharanya A, Visessanguan W, Kudo T, Ito T (2010) Gracilibacillus thailandensis sp. nov., from fermented fish (pla-ra). Int J Syst Evol Microbiol 60:944–948
Chen YG, Cui XL, Zhang YQ, Li WJ, Wang YX, Xu LH, Peng Q, Wen ML, Jiang CL (2008a) Gracilibacillus quinghaiensis sp. nov., isolated from salt-lake sediment in the Qaidam Basin, north-west China. Sys Appl Microbiol 31:181–189
Chen YG, Cui XL, Zhang YQ, Li WJ, Wang YX, Xu LH, Peng Q, Wen ML, Jiang CL (2008b) Gracilibacillus halophilus sp. nov., a moderately halophilic bacterium isolated from saline soil. Int J Sys Evol Microbiol 58:2403–2408
Dalmaso GZL, Ferreira D, Vermelho AB (2015) Marine extremophiles: a source of hydrolases for biotechnological applications. Mar Drugs 13:1925–1965
Deutch CE (1994) Characterization of a novel salt-tolerant Bacillus sp. from the nasal cavity of desert iguanas. FEMS Microbiol Lett 121:55–60
Deutch CE (2002) Characterization of a salt-tolerant extracellular α-amylase from Bacillus dipsosauri. Lett Appl Microbiol 35:78–84
Diop A, Khelaifia S, Armstrong N, Labs N, Fourrnier PE, Raoult D, Million M (2016) Microbial culturonics unravels the halophilic microbiota repertoire of table salt: description of Gracilibacillus massiliensis sp. nov. Microb Ecol Health Dis 27:32049. https://doi.org/10.3402/mehd.v27.32049
Diop A, Seck E, Dubourg G, Armstrong N, Blanc-Tailleur C, Raoult D, Fournier PE (2018) Genome sequence and description of Gracilibacillus timonensis sp. nov. strain Marseille- P2481T, a moderate halophilic bacterium isolated from the human gut microflora. MicrobiologyOpen 8:e638. https://doi.org/10.1002/mbo3.638
Duchêne D, Bochot A (2016) Thirty years of cyclodextrins. Int J Pharm 514:58–72
Editorial Board (2015) Gracilibacillus. Bergey’s Manual of Systematics of Archaea and Bacteria Online. https://doi.org/10.1002/9781118960608.gbm00534
Ekblom R, Wolf JBW (2014) A field guide to whole genome sequencing, assembly and annotation. Evol Appl 7:1026–1042
Foo AY, Bais R (1998) Amylase measurement with 2-chloro-4-nitrophenyl-maltotrioside as substrate. Clin Chim Acta 272:137–147
Gao M, Li ZZ, Zhou YG, Liu HC, Ma YC, Wang L, Chen SF, Ji XC (2012) Gracilibacillus kekensis sp. nov., a moderate halophile isolated from Keke Salt Lake. Int J Syst Evol Microbiol 62:1032–1036
Gella FJ, Gubern G, Vidal R, Canalias F (1997) Determination of total and pancreatic α-amylase in human serum with 2-chloro-4-nitrophenyl-α-d-maltotrioside as substrate. Clinica Chemica Acta 259:147–160
Gerday C, Glansdorff N (2007) Physiology and biochemistry of extremophiles. ASM Press, Washington
Goel A, Nene SN (1993) Modifications in the phenolphthalein method for spectrophotometric estimation of beta cyclodextrin. Starch (Stäke) 47:399–400
Graziano G, Merlino A (2014) Molecular basis of protein halotolerance. Biochim Biophys Acta 1844:850–858
Guan TW, Tian L, Li EY, Tang SK, Zhang XP (2017) Gracilibacillus aidingensis sp. nov., a novel moderately halophilic bacterium isolated from Aiding salt lake. Arch Microbiol 199:1277–1281
Guan HL, Zhang YJ, Lu XJ, Jia M, Zhang ZY, Gao XH, Ma YC, Tian F, Tang SK (2018) Gracilibacillus eburneus sp. nov., a moderately halophilic bacterium isolated form Xinjiang province, China. Archiv Microbiol 200:423–429
Hames BD (1998) Gel electrophoresis of proteins: a practical approach (3/e). Oxford University Press, Oxford
Hirota K, Hanaoka Y, Nodasaka Y, Yumoto I (2014) Gracilibacillus alcaliphilus sp. nov., a facultative alkaliphile isolated from indigo fermentation liquor for dyeing. Int J Syst Evol Microbiol 64:3174–3180
Horikoshi K, Grant WD (1998) Extremophiles: microbial life in extreme environments. Wiley-Liss, New York
Hough DW, Danson MJ (1999) Extremozymes. Curr Opin Chem Biol 3:39–46
Huang HQ, Wang Y, Yuan WD, Xiao C, Ye JJ, Liu M, Zhu J, Sun QG, Bao SX (2013) Gracilibacillus marinus sp. nov., isolated from the northern South China Sea. Antonie Van Leeuwenhoek 104:695–701
Huo YY, Xu XW, Cui HL, Wu M (2010) Gracilibacillus ureilyticus sp. nov., a halotolerant bacterium from a saline-alkaline soil. Int J Syst Evol Microbiol 60:1383–1386
Jansook P, Ogawa N, Loftsson T (2018) Cyclodextrins: structure, physicochemical properties and pharmacological applications. Int J Pharm 535:272–284
Jeon CO, Lim JM, Jang HH, Park DJ, Xu LH, Jiang CL, Kim CJ (2008) Gracilibacillus lacisalsi sp. nov., a halophilic Gram-positive bacterium from a salt lake in China. Int J Sys Evol Microbiol 58:2282–2286
Kato T, Horikoshi K (1984) Colorimetric determination of γ-cyclodextrin. Anal Chem 56:1738–1740
Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJE (2015) The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc 10:845–858
Kim P, Lee JC, Park DJ, Shin KS, Kim JY, Kim CJ (2012) Gracilibacillus bigeumensis sp. nov., a moderately halophilic bacterium from solar saltern soil. Int J Syst Evol Microbiol 62:1857–1863
Krüger A, Schäfers C, Schröder C, Antranikian G (2018) Towards a sustainable biobased industry—highlighting the impact of extremophiles. New Biotechnol 40:144–153
Laderman KA, Davis BR, Krutzsch HC, Lewis MS, Griko YV, Privalov PL, Anfinsen CB (1993) The purification and characterization of an extremely thermostable α-amylase from the hyperthermophilic archaebacterium Pyrococcus furiosus. J Biol Chem 268:24394–24401
Lanyi JK (1974) Salt-dependent properties of proteins from extremely halophilic bacteria. Bacteriol Rev 38:272–290
Lawson PA, Deutch CE, Collins MD (1996) Phylogenetic characterization of a novel salt- tolerant Bacillus species: description of Bacillus dipsosauri sp. nov. J Appl Bacteriol 81:109–112
Leemhuis H, Kelly RM, Dijkhuizen L (2010) Engineering of cyclodextrin glucanotransferases and the impact for biotechnological applications. Appl Microbiol Biotechnol 85:823–835
Lejeune A, Sakaguchi K, Imanaka T (1989) A spectrophotometric assay for the cyclization activity of cyclomaltohexaose (α-cyclodextrin) glucanotransferase. Anal Biochem 181:6–11
Lorentz K, Gütschow B, Renner F (1999) Evaluation of direct α-amylase assay using 2-chloro-4- nitrophenyl-α-d-maltotrioside. Clin Chem Lab Med 37:1053–1062
Madern D, Ebel C, Zaccai G (2000) Halophilic adaptation of enzymes. Extremophiles 4:91–98
Madigan MT, Marrs BL (1997) Extremophiles. Sci Am 276:82–87
McCleary BV (1988) Soluble dye-labeled polysaccharides for the assay of endohydrolases. Methods Enzymol 160:74–86
Merino N, Aronson HS, Bojanova DP, Feyhl-Buska J, Wong ML, Zhang S, Giovannelli D (2019) Living at the extremes: extremophiles and the limits of life in a planetary context. Front Microbiol 10:780. https://doi.org/10.3389/fmicb.2019.00780
Niehaus F, Bertoldo C, Kähler Antrankian G (1999) Extremophiles as a source of novel enzymes for industrial application. Appl Microbiol Biotechnol 51:711–729
Oh YJ, Lee HW, Kim SK, Kwon MS, Lee J, Jang JY, Park HW, Nam YD, Seo MJ, Choi HJ (2016) Gracilibacillus kimchii sp. nov., a halophilic bacterium isolated from kimchi. J. Microbiol 54:588–593
Orellana R, Macaya C, Bravo G, Dorochesi F, Cumsille A, Valencia R, Rojas C, Seeger M (2018) Living at the frontiers of life: extremophiles in Chile and their potential for bioremediation. Front Microbiol 9:2309. https://doi.org/10.3369/fmicb.2018.02309
Qi Q, Zimmermann W (2005) Cyclodextrin glucanotransferase: from gene to applications. Appl Microbiol Biotechnol 66:474–484
Rothschild LJ, Mancinelli RL (2001) Life in extreme environments Nature 409:1092–1101
Schiraldi C, De Rosa M (2002) The production of biocatalysts and biomolecules from extremophiles. Trends Biotechnol 20:515–521
Senghor B, Khelaifia S, Bassêne H, Seck EH, Fournier PE, Sokhna C, Raoult D, Lagier JC (2017) ‘Gracilibacillus phocaeensis’ sp. nov., ‘Sediminibacillus massiliensis’ sp. nov., and ‘Virgibacillus ndiopensis’ sp. nov., three halophilic species isolated from salty human stools by culturomics. New Microb New Infect 20:51–54
Sheehan H, McCleary BV (1988) A new procedure for the measurement of fungal and bacterial α-amylase. Biotechnol Tech 2:289–292
Simpson JT, Pop M (2015) The theory and practice of genome sequence assembly. Ann Rev Genomcs Hum Genet 16:151–172
Smith P, Krohn RI, Hermanson GT, Malina K, Garner FH, Provensano MD, Fugimoto EK, Goeke NM, Olson GJ, Klenk DC (1985) Measurement of protein using bicinchoninic acid. Anal Biochem 150:76–85
Stam MR, Danchin EGJ, Rancurel C, Coutinho PM, Henrissat B (2006) Dividing the large glycoside hydrolase family 13 into subfamilies: towards improved functional annotations of α-amylase-related proteins. Protein Eng Des Sel 19:555–562
Suganuma T, Maeda Y, Kitahara K, Nagahama T (1997) Study of the action of human salivary alpha-amylase on 2-chloro-4-nitrophenyl-α-maltotrioside in the presence of potassium thiocyanate. Carbohydr Res 303:219–227
Tang SK, Wang Y, Lou K, Mao PH, Jin X, Jiang CL, Xu LH, Li WJ (2009) Gracilibacillus saliphilus sp. nov., a moderately halophilic bacterium isolated from a salt lake. Int J Syst Evol Microbiol 59:1620–1624
Wainø M, Tindall BJ, Schumann P, Ingvorsen K (1999) Gracilibacillus gen. nov., with description of Gracilibacillus halotolerans gen. nov., sp. nov.; transfer of Bacillus dipsosauri to Gracilibacillus dipsosauri comb. nov., and Bacillus salexigens to the genus Salibacillus gen. nov., as Salibacillus salexigens comb. nov. Int J Syst Bacteriol 49:821–831
Winn-Deen ES, David H, Sigler G, Chavez R (1988) Development of a direct assay for α- amylase. Clin Chem 34:2005–2008
Yang N, Ren B, Dai H, Liu Z, Zhou Y, Song F, Zhang L (2013) Gracilibacillus xinjiangesnsis sp. nov., a new member of the genus Gracilibacillus isolated from the Xinjiang region, China. Antonie Van Leeuwenhoek 104:809–816
Acknowledgements
We thank Maria Dolores Sotelo for her efforts to clone the α-amylase gene from G. dipsosauri and Drs. Pamela Marshall and Jeffrey Newman for facilitating the sequencing of the G. dipsosauri genome at Indiana University.
Funding
There was no external funding source for this work.
Author information
Authors and Affiliations
Contributions
Charles E. Deutch did all of the microbiological and biochemical experiments and wrote the manuscript. Shanshan Yang did the assembly and analysis of the Gracilibacillus dipsosauri genome.
Corresponding author
Ethics declarations
Conflicts of interest
We declare that we have no conflicts of interest in publishing this work.
Human or animal participants
No humans or animals were used in this project.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Multiple sequence alignment of the α-amylases from G. dipsosauri (PWU69658.1), G. halophilus (N4WMC6), G. orientalis (A0A1H9MMU5), G. kekensis (A0A1M7K410), and G. ureilyticus (A0A1I4H7R9). Fully conserved residues are indicated by (*); similar residues are indicated by (:) or (.) (PNG 146 kb)
Rights and permissions
About this article
Cite this article
Deutch, C.E., Yang, S. Genomic sequencing of Gracilibacillus dipsosauri reveals key properties of a salt-tolerant α-amylase. Antonie van Leeuwenhoek 113, 1049–1059 (2020). https://doi.org/10.1007/s10482-020-01417-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-020-01417-2