Abstract
The shikimate pathway is indispensable for the biosynthesis of natural products with aromatic moieties. These products have wide current and potential applications in food, cosmetics and medicine, and consequently have great commercial value. However, compounds extracted from various plants or synthesized from petrochemicals no longer satisfy the requirements of contemporary industries. As a result, an increasing number of studies has focused on this pathway to enable the biotechnological manufacture of natural products, especially in E. coli. Furthermore, the development of synthetic biology, systems metabolic engineering and high flux screening techniques has also contributed to improving the biosynthesis of high-value compounds based on the shikimate pathway. Here, we review approaches based on a combination of traditional and new metabolic engineering strategies to increase the metabolic flux of the shikimate pathway. In addition, applications of this optimized pathway to produce aromatic amino acids and a range of natural products is also elaborated. Finally, this review sums up the opportunities and challenges facing this field.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
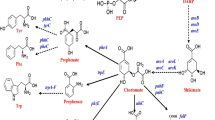
The shikimate pathway is indispensable for maintaining the normal metabolism of cells and is ubiquitously present in plants, animals and microorganisms [6, 14]. In E. coli, various carbon sources are converted into phosphoenolpyruvate (PEP) and erythrose‐4‐phosphate (E4P) through the glycolysis pathway (EMP pathway) and the pentose phosphate pathway (PP pathway), respectively. Subsequently, both PEP and E4P are transformed into 3‐deoxy‐d‐arabino‐heptulosonate‐7‐phosphate (DAHP), which is the initial substrate of the shikimate pathway. The condensation of PEP and E4P is catalyzed by DAHP synthetase in E. coli, which is coded by aroF, aroG, and aroH [59], which makes it challenging to increase the reaction flux toward the shikimate pathway. DAHP is subsequently transformed into chorismic acid (CHOR) through six reactions [20, 61, 64, 32] (Fig. 1). In addition, there are also bypass pathways related to shikimate formation, such as the quininic acid pathway and the gallic acid pathway (Fig. 1) [34, 36]. These fundamental pathways are crucial for the synthesis of many high-value compounds. For example, chorismic acid, quininic acid and gallic acid deriving from these pathways act as important intermediates or precursors for the manufacturing of a number of natural products, which are usually extracted from various plants or are synthesized chemically. However, plant extraction and chemical synthesis have many disadvantages such as environment contamination, high energy consumption, and low yields [6]. Hence, the development of alternative microbial routes is an increasingly important trend in the large-scale production of natural products. With the development of metabolic engineering and synthetic biology strategies, great progress has been made in the construction of microbial cell factories for the production of increasingly complex natural products. In many cases, a well-defined model organism such as E. coli is preferable as a cell factory for the synthesis of certain natural products. It is considered a perfect platform host for the development of industrial production due to its genetic tractability, favorable growth conditions and availability of versatile genetic manipulation tools [57]. This review mainly summarizes recent advances in increasing the flux through the shikimate pathway by diverse metabolic engineering strategies and the utilization of this pathway for the synthesis of high-value compounds in E. coli.
a Schematic representation of approaches used to adapt the shikimate pathway for biotechnological purposes. TktA transketolase 1, PpsA phosphoenolpyruvate synthetase, PykAF pyruvate kinases I and p II, AroF AroG and AroH 3-deoxy-7-phosphoheptulonate synthase, AroB 3-dehydroquinate synthase, AroD 3-dehydroquinate dehydratase, YdiB shikimate dehydrogenase/quinate dehydrogenase, AroE shikimate dehydrogenase, AroL and AroK shikimate kinases 1 and 2, AroA 3-phosphoshikimate 1-carboxyvinyltransferase, AroC chorismate synthase, UbiC chorismate lyase, PobA p-hydroxybenzoate hydroxylase, EntC isochorismate synthase, PchB isochorismate pyruvate lyase, EntAB 2,3-dihydro-2,3-dihydroxybenzoate dehydrogenase and enterobactin synthase component B, PprA PhPA reductase from the fungus Wickerhamia fluorescens, DHBA-1-M 2,3-dihydroxybenzoic acid 1-monoxygenase, HpaBC 4-hydroxyLphenylacetate 3-hydrolase, SMO and CDO salicylate 1-monoxygenase and catechol 1,2-dioxygenase, HQT hydroxycinnamoyl-CoA quinate transferase from Nicotiana tabacum, FldABCI L-phenyllactate dehydratase components, FevV L-tyrosine ammonia lyase from Streptomyces sp. WK-5344, 4CL 4-coumarate-CoA ligase, CHS chalcone synthase from Petunia hybrid, CHI chalcone isomerase from Medicago sativa, CPR cytochrome P450 reductase from Catharanthus roseus, F3′H flavonoid 3′-hydroxylase from G. hybrid, CAA caffeic acid, CA cinnamic acid, CHA chlorogenic acid, HT hydrolysable tannins, MA muconic acid, PG pyrogallol, PhLA phenyllactate, SA salicylic acid, AN arbutin, ED eriodictyol, NG naringenin, NGC naringenin chalcone, CatA catechol 1,2-dioxygenase from Klebsiella pneumoniae strain CICIM B7001, EntX 2,3-dihydroxybenzoate decarboxylase from Pseudomonas putida strain KT2440. ldh-lpox L-phenylalanine oxidase and L-lactate dehydrogenase, PAL phenylalanine ammonia lyase, FDC1 ferulic acid decarboxylase, tCA ferulic acid, SR Styrene. b, c and d include chemical structure mentioned compounds in (a)
Strategies for enhancing the metabolic flux through the shikimate pathway
A prerequisite for the industrial biosynthesis of aromatic compounds via the shikimate pathway is the enhancement of its flux. The main metabolic engineering strategies include the engineering of transport systems, the trim of competing pathways and the optimization of critical enzymes.
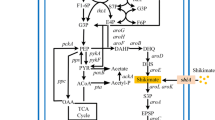
E. coli can assimilate several carbon resources through different specific transport systems, whereby some substances utilize more than two transport routes (Fig. 2A) [51, 57]. Different transport routes have crucial effects on the final distribution of carbon flux between the shikimate pathway and cell growth, and researchers used diverse carbon sources such as glucose, glycerol, xylose, fructose, and sucrose and engineered their transport routes in E. coli [1, 24, 25, 70]. Glucose enters into the cell by either PTS or the non-PTS route (Fig. 2B), whereby the former utilizes a phosphate from PEP in the process of glucose transport [27]. However, PEP is an important precursor of metabolic routes that converge into the shikimate pathway. Therefore, reducing the consumption of PEP through PTS modification is an important strategy to increase the metabolic flux into the shikimate pathway. The glucose PTS is composed of IICBGlc, enzyme IIAGlc, HPr and enzyme I encoded by ptsI, ptsG, ptsH, and crr, respectively. Accordingly, ptsH was knocked out to decrease the consumption of PEP for the improvement of L-tryptophan production [42]. In a different approach, PTS was replaced by non-PTS that consists of glucokinase (encoded by glk), galactose permease (encoded by galP) and H+ [4,70] (Fig. 2B). This strategy can effectively decrease PEP consumption. In addition, different carbon sources were employed as substrates to increase the shikimate pathway flux as well [37]. The highest yield and production rate of shikimate were obtained in glucose and glycerol medium, respectively. Xylose and glucose, as fermentative substrates, enter cells through different transport systems. Consequently, Ranjan et al. [59] used them to synthesize the first compound of the shikimate pathway, DAHP.
(A) Import pathways of some common carbohydrate substrates into E. coli cells. ① a XylFGH is a high-affinity ABC transporter that imports xylose into the cell. b XylE is a low affinity module that conveys xylose into the cell. c Xylose enters the cell through hyperosmotic diffusion, which is controlled by XylAB ② E. coli absorbs xylose via three different routes. a Fructose enters E. coli via the membrane-spanning enzyme E2fru that is encoded by fruAB. b Fructose is transported by the mannose-transport system encoded by the manXYZ operon. c Fructose diffuses into the cell via an isoform of the major glucose permease of the PTS system (PtsG-F) ③ Sucrose enters the cell via either the PTS or a non-PTS route specific for sucrose. a Sucrose enters the cell via ScrY and EIIscr (encoded by scrA). b CscB permease encoded b assists the passive transport of sucrose ④ L-arabinose is transported into the cell through two independent inducible pathways. a A low-affinity system consists of AreE. b A high-affinity system consists of AreFGH.⑤ Glycerol is metabolized in E. coli in three ways to form dihydroxyacetone phosphate (DHAP): a The glycerol kinase (GlpK) and aerobic glycerol 3-phosphate dehydrogenase (GlpD) route. b The GlpK and anaerobic glycerol 3-phosphate dehydrogenase (GlpABC) route. c The fermentative glycerol dehydrogenase (GldA) and DHA kinase (DhaKLM) route. (B) Overview of the glucose transport routes of E. coli. After glucose diffuses into the periplasm through OmpC, OmpF, or LamB, it can enter the cell via the enzyme IICBGlc, enzyme IIAGlc, HPr and enzyme I encoded by ptsI, ptsG, ptsH, crr. They constitute the glucose PTS (phosphoenolpyruvate dependent carbohydrate phosphotransferase system); GalP imports glucose using the proton motive force. Subsequently, glucose is phosphorylated by Glck; MglABC also mediates glucose import and then Glck phosphorylates glucose [39, 51]
The trim of competing pathways is also an important strategy for improving shikimate flux. This approach can effectively reduce byproduct synthesis and carbon resource depletion. For example, pta (encoding acetate kinase) and ackA (encoding acetate kinase) were deleted to reduce the accumulation of acetate and waste of carbon source [68]. In addition, the overexpression of ppsA (encoding phosphoenolpyruvate synthetase) (Fig. 1) and the knockdown of pykAF (encoding pyruvate kinases I and II) (Fig. 1) are usually chosen to convert more pyruvate into PEP [12], and overexpressing tktA (encoding transketolase 1) [38, 47, 48] (Fig. 1) can increase the pool of E4P, which is also an important intermediate. In recent years, the development of biotechnology has made it possible to switch off relevant genes at desirable time points using biosensors to balance cell growth and product accumulation. Gupta et al. [29] used pathway-independent quorum sensing circuits to control the metabolic flow by regulating the expression of pfkA (encoding 6-phosphofructokinase I) and aroK (encoding shikimate kinase 1) at proper times. This method is superior to the traditional method of adding an inducer and realizes dynamic control, since the inducer may reduce the growth or productivity of the strain and increase the cost of fermentation.
The tuning of enzymes and proteins, including modifying and optimizing enzyme structure [47] and introducing heterologous enzymes [38], is also an effective approach to further increase the flow through the shikimic acid pathway. These modification strategies can release feedback inhibition of specific enzymes or accelerate the channeling of substrates.
With the rapid development of bioinformatics and metabolomics, large amounts of metabolic information can be collected in a specific database, which in turn allows the simulation of intracellular metabolic pathways and enables the rational engineering of the shikimate pathway. A widely used, albeit expensive method is 13C metabolic flux analysis, which enables the accurate measurement of intracellular fluxes [66, 74]. What’s more, to verify the successful modification of target genes and understand the physiological and metabolic state of the production strains, several mathematical models have been built. For example, the combination of dynamic metabolic modeling and response surface methodology was employed to construct a model for improving shikimate production in E. coli [53]. In this study, fermentation parameters of time course of biomass, glucose concentration, yeast extract concentration and shikimic acid concentration were collected and then these parameters were optimized using response surface model. This model facilitated improvement of fed-batch process.
Application of the shikimate pathway for the biosynthesis of aromatic compounds
To manufacture diverse value-added compounds applied in cosmetics, foods, pharmaceuticals and related fields, many studies focused on the shikimate pathway. In this section, we summarized some successful cases of using the afore-mentioned methods to modify the shikimate pathway of E. coli to synthesize high-value-added compounds. The product spectrum includes aromatic amino acids, chorismic acid, quininic acid (QA), gallic acid (GA) and their derivatives (Fig. 1 and Table 1).
3-dehydroquinc acid, 3-dehydroshikimic acid and their derivatives
In E. coli, QA and GA are derived from 3-dehydroquinc acid (DHQ) and 3-dehydroshikimic acid (DHS), respectively [22] (Fig. 1). From the point of view of their structures, both have multiple hydroxyl groups and one carboxyl group on the benzene ring, which makes them potential precursors of high-value-added compounds.
The QA present in fruits is an intermediate in the synthesis of chlorogenic acid. Cha et al. employed glucose as a carbon resource to synthesize QA in engineered E. coli using a co-culture system. In this approach, one strain was used to synthesize caffeic acid from L-tyrosine (Tyr), and another strain transformed caffeic acid and quinate into chlorogenic acid (CHA) using hydroxycinnamoyl-CoA quinate transferase (HQT) from Nicotiana tabacum. However, the production of caffeic acid from L-Tyr was subject to inhibition by high concentrations of the substrate, which directly influenced chlorogenic acid production [8].
GA is an important intermediate in the synthesis of hydrolysable tannins [55] which is a polyphenol substance, and also acts as a primary anti-inflammatory agent and an active component responsible for reducing coronary arterial disease (CAD) and arterial thrombosis [3]. Shikimate dehydrogenase (encoded by aroE) is essential for GA synthesis in E. coli. Muir et al. [55] overexpressed aroE to enhance the yield of GA. In addition, GA can also be synthesized from chorismic acid. The pobA gene (encoding p-hydroxybenzoate hydroxylase) from Pseudomonas aeruginosa was introduced into E. coli, after which a new mutant of this gene with improved activity was developed and used to synthesize GA [10]. In this report, PobA was rationally engineered by the structural analysis and rational design. They found that the enzyme with Y385F and T294A has higher catalytic activity which is a fourfold increase, compared with the reported Y385F mutant.
Chorismic acid derivatives
Chorismic acid, the end product of the shikimate pathway, plays vital roles as a key intermediate in the metabolism of aromatic compounds (Fig. 1). Salicylic acid (SA) is one of the most essential aromatic monomers derived from chorismic acid and a precursor in the synthesis of aspirin (AN) [56] and muconic acid (MA) [40]. For example, Qian et al. used high-throughput screening technology based on a biosensor to identify SA high-producer strains [58]. In this report, a combinatorial library was constructed by comprising a series of ribosome binding site sequences corresponding to a range of predicted protein translation initiation rates, including entC (encoding isochorismate synthase), pchB (encoding isochorismate pyruvate lyase), aroL (encoding shikimate kinases 1), ppsA, tktA and aroGfbr. Then a biosensor with AraC-PBAD-lacZ sensor-reporter system was used to screen high SA production strains. Compared with conventional methods, this method can be used to quickly and accurately select target strains from large libraries. Moreover, Lin et al. produced SA in engineered E. coli by expressing isochorismate synthase and introducing isochorismate pyruvate lyase from Pseudomonas fluorescens. SA production of the engineered strain reached 1.2 g/L [40]. Subsequently, they used SA as precursor for the synthesis of MA via a partial degradation pathway composed with salicylate 1-monoxygenase (SMO) and catechol 1,2-dioxygenase (CDO). In the report, SMO (encoded by nahGop) with the highest activity had been identified.
Besides, Arbutin (AT), which is widely used in the pharmaceutical and cosmetic industries due to its well-known skin-lightening effect, as well as antioxidant, antimicrobial, and anti-inflammatory activities, was synthesized from chorismic acid. In one metabolic engineering strategy, the flavin adenine dinucleotide-dependent 4-hydroxybenzoate 1-hydroxylase (MNX1) from Candida parapsilosis CBS604 and arbutin synthase (AS) from Rauvolfia serpentina were introduced into E. coli, and then enhancing shikimate pathway was used to further increase the AT production, resulting in 3.29 g/L AT [67]. Pyrogallol (PG) also was synthesized from chorismic acid in E. coli via expressing an efficient 2,3-dihydroxybenzoic acid (2,3-DHBA) 1-monoxygenase that was identified among a series of oxygenases and hydroxylases based on structural similarity of their the substrates and products [65]. Of note, this work introduced a biological reaction that was not known in nature—the conversion of 2,3-DHBA into PG.
It is prerequisite for overproduction of AN, MA, SA, PG and AT to increase shikimate pathway flux. The construction of minimum chassis cells is considered as a promising strategy to effectively synthesize these natural compounds. For example, free-chromosome engineering could offer a reference to construct a minimum chassis cells [18]. This engineering was the removal of native chromosome by double-stranded breaks made by heterologous I-CeuI endonuclease and the degradation activity of endogenous nucleases, which resulted in simple cells. Synthetic genetic circuits were introduced into these simple cells to express target genes and then obtain target metabolites. These simple cells maintained the ability of expression of target genes, and the expression of these genes was not disturbed by native chromosome. Besides, genes of glycolysis pathway which are able to regenerate ATP and NADH/NADPH were assembled to a plasmid with tight regulatory system. Then the plasmid was introduced into these cells to extend their longevity. For this strategy, there are two advantages: elimination of disruption from chromosome and artificial design of genetic circuits.
Aromatic amino acids
The condensation of E4P and PEP yields DAHP, which is converted into chorismic acid through the shikimate pathway, and further into aromatic amino acids including L-Tyr, L-phenylalanine (L-Phe) and L-tryptophan (L-Trp). These are crucial building blocks for many high-value-added compounds. For example, in addition to being an essential amino acid in the human body, L-Trp is the starting material for the synthesis of numerous pharmaceuticals such as antidepressants and antitumor drugs. L-Phe is also a vital ingredient of the widely used artificial sweetener aspartame, whereas L-Tyr is an essential dietary component for patients with L-phenylketonuria and is also the starting material for the synthesis of L-DOPA [21]. Because aromatic amino acids have such a wide range of uses, there is a market in excess of 14,000 ton/year for L-tryptophan, and the production of L-Phe exceeds 30,000 tons/year [61]. To achieve more efficient production of the three aromatic amino acids, increasing numbers of studies employ renewable materials to produce them by microbial fermentation. Various genetic editing tools such as clustered regularly interspaced short palindromic repeats (CRISPR) and error-prone PCR were used to generate diverse chassis cells. Then, according to the need of specific products, these chassis cells need to be modified by specific biological strategies, including reducing overflow metabolism, modifying transport systems, de-inhibition of enzymes, and construction of co-expression systems.
For instance, the effects of genetic manipulation of phosphate acetyltransferase (encoded by pta), high affinity tryptophan transporter (encoded by mtr) and aromatic amino acid exporter (encoded by yddG) on L-Trp production were studied with the aim to reduce acetate synthesis and accelerate tryptophan export [28, 43]. The combination of reducing overflow metabolism and modifying tryptophan transport system was a strategy to improve tryptophan production in this study. Furthermore, due to polyhydroxybutyrate (PHB) biosynthesis pathway diverting the acetyl-CoA, a precursor of acetate, to PHB biosynthesis and reducing acetate secretion, the pathway was introduced into E. coil for improving L-Trp production as well [26, 50, 63, 68]. Feedback inhibition and repression are ubiquitously present in E. coli. DHAP is synthesized by enzymes encoded by the three genes aroF, aroG, and aroH, which are, respectively, inhibited by high intracellular concentrations of L-Trp, L-Phe and L-Tyr. To circumvent this problem, aroF and aroH were knocked out and aroG was rationally engineered to relieve the feedback inhibition by the three aromatic amino acids, which increased the production of L-Trp [9]. Much work was done on enhancing the supply of available PEP, which is the crucial precursor for L-Trp synthesis in E. coli. Common approaches include knocking out or knocking down the PTS system [42].
The biosynthesis pathways of L-Phe and L-Tyr share some similarities. Chorismic acid is converted to prephenate by enzymes encoded by pheA/tyrA (encoding fused chorismate mutase/prephenate dehydrogenase), and further to L-Tyr or L-Phe by specific enzymes. L-Phe is an important precursor for the low-calorie sweetener aspartame. To achieve an improvement of L-Phe titer, the overflow metabolism was reduced by moderating the glucose uptake rate by inactivating the crr gene that encodes the IIAGlc glucose-specific transporter [45]. In one study, PTS was inactivated, after which glucose facilitator and glucokinase (glf and glk) from Zymomonas mobilis were introduced into E. coli to replace PTS and increase the shikimate pathway flux [49]. In addition, AroK, AroL, AroA, AroC, PheA and TyrB, which are involved in the shikimate pathway, were analyzed using the Multi-Enzyme Reaction System to identify the key enzyme for the improvement of L-Phe yield [15]. In this study, the simplified steps about the identification of key enzymes were as follows: Firstly, relative proportion of intracellular enzyme concentration of a strain was obtained by proteomics. Secondly, crude enzyme extract of the strain was prepared. Then a sufficient amount of substrate was added to the crude enzyme extract. Finally, a pure enzyme with known concentration was added to the reaction system of the previous step, and then the yield of the final product in the reaction solution was detected to determine whether the enzyme was a key enzyme. Their results demonstrated that the concentration of AroA increasing in vivo was favorable to improve L-Phe production. Besides, because ydiB (encoding shikimate dehydrogenase) and aroK (encoding shikimate kinases) are also key genes that convert DHS to form SHK, they were selected as the target genes for improving the L-Phe yield [44]. In this report, ydiB and aroK were overexpressed, which leads to the improvement of L-Phe yield and the short of lag phase. Genetic modification of pathways for the production of the third aromatic amino acid, L-Tyr, is relatively simple. For example, the two major genes, aroK and ydiB, can be overexpressed to directly increase L-Tyr production [52]. Besides, to enable precise regulation of L-Tyr synthesis, a constitutive promoter and 5-untranslated region (5′-UTR) were artificially designed, resulting in the 3 g/L L-Tyr in shake flasks [11]. Contrasted by the overexpression and knock out or down of genes, this strategy, artificially designed 5-untranslated region (5′-UTR), could better precisely regulate expression level of genes. In addition, Santos et al. [62] utilized a combinatorial metabolic engineering approach for optimizing cellular phenotype to further increase L-Tyr production based on engineered strain. The combination of global transcriptional machinery engineering and high-throughput screening was used to improve L-tyrosine production. In this report, two separate global transcription machinery engineering (gTME) libraries of rpoA(encoding RNA polymerase subunit α) and rpoD (encoding RNA polymerase, sigma D) were constructed based on the mutagenesis of the RNA polymerase subunits and then three strains with high-level production of L-Tyr were screened via the tyrosinase-mediated conversion of L-Tyr to the dark and diffusible pigment melanin.
Aromatic amino acid derivatives
A number of valuable natural products contain benzene rings. Examples include L-DOPA, mandelic acid, violacein, avenanthramides and resveratrol. To biosynthesize these compounds, many researchers used synthetic biology to introduce heterologous genes and pathways into E. coli. Several successful studies that utilized aromatic amino acids to synthesize useful natural products are reviewed in this section.
L-Trp is usually used as the starting material for the synthesis of deoxyviolacein and violacein, which have multiple pharmaceutical activities, such as antibacterial [56], antitumoral [17], antiviral [2], antifungal [5], antiprotozoan [33], and antioxidant [7] effects. Several natural strains were engineered to produce these compounds. The vioABCE operon from Chromobacterium violaceum and vioD (encoding VioD) from Janthinobacterium lividum were introduced into E.coli to enable the synthesis of deoxyviolacein (Devio) and violacein (Vio) from L-Trp (Fig. 1) [19]. However, before the vioABCDE operon was introduced into E. coli, a chassis cell that can efficiently produce tryptophan first had to be constructed. To solve the problem of low efficiency when using plasmid expression, vioD was integrated into the genome. At the same time, the supply of building blocks including serine, chorismic acid and L-Trp was engineered by introducing the optimized serAfr (encoding mutated phosphoglycerate dehydrogenase), aroGfr and trpEfr (encoding mutated anthranilate synthase subunit) genes [60]. Subsequently, vioE (encoding VioE) was overexpressed to improve the violacein yield, because VioE catalyzes the rate-limiting step [72].
Similarly, L-Phe from chorismate can serve as a precursor for the synthesis of a series of important compounds such as phenyllactate, phenylacetate, phenylethanol, cinnamic acid (CA) and styrene. L-phenylalanine oxidase from Coprinopsis cinereus and L-lactate dehydrogenase from Lactobacillus plantarum were co-expressed to produce L-phenyllactate, which has antimicrobial activity. Then a two-stage fermentation process with a temperature shift was used to increase the yield of phenyllactate (PhlA) [70]. In addition, E. coli expressing pprA (encoding phenylpyruvate reductase, PprA) from W. fluorescens TK1 was also able to synthesize (PhlA) [30]. In other study, not only PhlA was synthesized by PprA but also was it used as an intermediate to produce CA via FidABCI from C. sporogenes (Fig. 1) [54].
L-Tyr can be converted into caffeic acid (CAA) by simultaneously co-expressing a fusion protein of TAL (L-Tyrosine ammonia lyase), 4CL (4-coumarate-CoA ligase), and 4-hydroxyL-phenyllactate 3-hydroxylase (4HPA3H) (Fig. 1). CAA can be sued as a building block for thermoplastics and a precursor for biologically active compounds [35]. Furthermore, L-Tyr can be transformed into eriodictyol (ED) via six heterologous enzymes—tyrosine ammonia lyase (TAL), 4-coumarate-CoA ligase (4CL), chalcone synthase (CHS), chalcone isomerase (CHI), flavonoid 3′-hydroxylase (F3′H) and CPR, encoding cytochrome P450 reductase (CPR) from R. glutinis, P. crispum, P. hybrida, M. sativa, G. hybrida and C. roseus, respectively [73]. In this study, an appropriate expression of these enzymes is paid attention to improve ED yield. The genes encoding these enzymes were codon-optimized for expression in E. coli (https://www.jcat.de/). In addition, L-Tyr can also be converted into L-DOPA by HpaBC (Fig. 1). A chassis cell that can effectively produce L-Tyr was constructed by engineering aroC, aroL, aroG and tyrB. Subsequently, hpaBC (encoding 4-hydroxyphenylacetate-3-hydrolase, HpaBC) was introduced into this chassis cell to produce L-DOPA from glycerol [13]. This resulting strain could produce 12.5 g/L L-DOPA in fed batch.
To synthesize these natural products, numerous heterologous genes were introduced into E. coli. This can lead to compatibility problems, such as low or no expression of the introduced enzyme, or low enzyme activity in spite of good expression. These problems urgently need to be solved before industrialization of the related bioprocesses becomes a viable option. To the best of our knowledge, optimization of gene sequences, replacement of expression vector or truncation of peptide chain are commonly selected to solve the successful expression of enzymes; strong promoter and ribosome binding site (RBS) or high copy plasmid are usually used to overcome low expression of enzymes; the improvement of enzyme activity mainly relies on rational design (by Rosetta) or saturation mutation (by specific primer).
Concluding remarks and future perspectives
Throughout the last several decades, a multitude of natural products derived from intermediates of the shikimate pathway have been studied and produced in E. coli. These natural compounds have complex structures, so that chemical synthesis or plant extraction cannot meet the requirements of modern industry. As a result, microbial fermentation using advanced cell factories has gradually replaced them. An important advantage of biotechnological methods is the possibility to use renewable and economical raw materials, and the rapid development of metabolic engineering in the last ten years makes it possible to rationally and quickly construct production strains. The shikimate pathway of E. coli was studied by numerous researchers aiming to manufacture the natural products mentioned above. After several generations of efforts, the envisioned production processes have been achieved, but there are still some challenges. For example, due to the complexity of metabolic pathways such as L-tryptophan synthesis, the low yield of some products has not been solved. If this obstacle is not removed, it will lead to a waste of resources and an increase in costs. In successful studies, this problem was solved using non-PTS transport and changing the culture media. However, PTS modification can lead to adverse effects, such as growth retardation and low productivity, and there still is no effective way to replace PTS. Presently, a non-PTS, MglCBA-Glk transport system, has not yet been used to increase shikimate flux. The transport system can work when sugar is present at a very low concentration [23]. It is possible that a combined use of the transport system and other transport systems will be developed to solve this problem. On the other hand, the synthesis of most natural products requires the introduction of heterologous genes into E. coli, and their expression can potentially disrupt the native metabolism. To overcome this problem, researchers developed new tools such as the RBS calculator (https://www.denovodna.com/software/) for tuned expression, biosensors to regulate expression in real time, and the use of multi-enzymes systems. However, there is no universal and effective method to resolve this challenge. Nevertheless, some important global regulators such as TyrR [69] and TrpR [31], as well as the regulatory factors that control energy metabolism, have been paid increasing attention by researchers, with the aim to achieve precise regulation of the shikimate pathway. In view of the development of metabolic engineering E. coli to date, it is not difficult to see that diverse omics approaches, new enzyme design strategies and bioinformatic methods gradually become more widespread. In the near future, the development of molecular tools will facilitate further research on the efficient biotechnological production of aromatic amino acids and valuable natural compounds. Throughout, the increase of shikimate pathway flux is a key factor to synthesize compounds deriving from this pathway. Thus, we can well engineer this pathway via some strategies that are shown below. Firstly, E4P and PEP amounts are increased from heterologous pathways constructed or mixed carbon added as substrate. Secondly, appropriate expression level of pathway enzyme can be identified based on dynamical parameter of chemical reactions existing in this pathway, not just the inactivation, knockdown and overexpression of certain genes. Thirdly, due to the imbalance of reducing equivalent being usually a limited step, maintaining cofactor balance may be also an important strategy. Actually, a prominent engineering strain constructed depends on the integrated use of several proper strategies not just one. This review offers a broad overview for all researchers who wish to harness the shikimate pathway of E. coli for biotechnological purposes.
References
Ahn J, Chung BKS, Lee D, Park M, Karimi IA, Jung J, Lee H (2011) NADPH-dependent pgi-gene knockout Escherichia coli metabolism producing shikimate on different carbon sources. FEMS Microbiol Lett 324(1):10–16. https://doi.org/10.1111/j.1574-6968.2011.02378.x
Andrighetti-Fröhner CR, Antonio R, Creczynski-Pasa t, Barardi CRM (2003) Cytotoxicity and potential antiviral evaluation of violacein produced by chromobacterium violaceum. Mem Inst Oswaldo Cruz 98(6):843–848. https://doi.org/10.1590/S0074-02762003000600023
Appeldoorn CCM, Bonnefoy A, Lutters BCH, Daenens K, van Berkel TJC, Hoylaerts MF, Biessen EAL (2005) Gallic acid antagonizes P-selectin–mediated platelet-leukocyte interactions. Circulation 111(1):106–112. https://doi.org/10.1161/01.CIR.0000151307.10576.02
Balderas-Hernandez VE, Sabido-Ramos A, Silva P, Cabrera-Valladares N, Hernandez-Chavez G, Baez-Viveros JL, Martinez A, Bolivar F, Gosset G (2009) Metabolic engineering for improving anthranilate synthesis from glucose in Escherichia coli. Microb Cell Fact 8:19. https://doi.org/10.1186/1475-2859-8-19
Becker MH, Brucker RM, Schwantes CR, Harris RN, Minbiole KPC (2009) The bacterially produced metabolite violacein is associated with survival of amphibians infected with a lethal fungus. Appl Environ Microb 75(21):6635–6638. https://doi.org/10.1128/AEM.01294-09
Bilal M, Wang S, Iqbal HMN, Zhao Y, Hu H, Wang W, Zhang X (2018) Metabolic engineering strategies for enhanced shikimate biosynthesis: current scenario and future developments. Appl Microbiol Biotechnol 102(18):7759–7773. https://doi.org/10.1007/s00253-018-9222-z
Bromberg N, Durán N (2001) Violacein transformation by peroxidases and oxidases: implications on its biological properties. J Mol Catal B Enzym 11(4):463–467. https://doi.org/10.1016/S1381-1177(00)00171-5
Cha MN, Kim HJ, Kim BG, Ahn J (2014) Synthesis of chlorogenic acid and p-coumaroyl shikimates from glucose using engineered Escherichia coli. J Microbiol Biotechnol 24(8):1109–1117. https://doi.org/10.4014/jmb.1403.03033
Chen L, Zeng A (2017) Rational design and metabolic analysis of Escherichia coli for effective production of L-tryptophan at high concentration. Appl Microbiol Biotechnol 101(2):559–568. https://doi.org/10.1007/s00253-016-7772-5
Chen Z, Shen X, Wang J, Wang J, Yuan Q, Yan Y (2017) Rational engineering of P-hydroxybenzoate hydroxylase to enable efficient gallic acid synthesis via a novel artificial biosynthetic pathway. Biotechnol Bioeng 114(11):2571–2580. https://doi.org/10.1002/bit.26364
Cheol Kim S, Eun Min B, Gyu Hwang H, Woo Seo S, Jung GY (2015) Pathway optimization by re-design of untranslated regions for L-tyrosine production in Escherichia coli. Sci Rep. https://doi.org/10.1038/srep13853
Cui Y, Ling C, Zhang Y, Huang J, Liu J (2014) Production of shikimic acid from Escherichia coli through chemically inducible chromosomal evolution and cofactor metabolic engineering. Microb Cell Fact 13(1):21. https://doi.org/10.1186/1475-2859-13-21
Das A, Tyagi N, Verma A, Akhtar S, Mukherjee KJ (2018) Metabolic engineering of Escherichia coli W3110 strain by incorporating genome-level modifications and synthetic plasmid modules to enhance L-Dopa production from glycerol. Prep Biochem Biotechnol 48(8):671–682. https://doi.org/10.1080/10826068.2018.1487851
Díaz-Quiroz DC, Cardona-Félix CS, Viveros-Ceballos JL, Reyes-González MA, Bolívar F, Ordoñez M, Escalante A (2018) Synthesis, biological activity and molecular modelling studies of shikimic acid derivatives as inhibitors of the shikimate dehydrogenase enzyme of Escherichia coli. J Enzyme Inhib Med Chem 33(1):397–404. https://doi.org/10.1080/14756366.2017.1422125
Ding D, Liu Y, Xu Y, Zheng P, Li H, Zhang D, Sun J (2016) Improving the production of L-phenylalanine by identifying key enzymes through multi-enzyme reaction system in vitro. Sci Rep. https://doi.org/10.1038/srep32208
Durán N, Menck CF (2001) Chromobacterium violaceum: a review of pharmacological and industiral perspectives. Crit Rev Microbiol 27(3):201–222. https://doi.org/10.1080/20014091096747
Durán N, Justo GZ, Ferreira CV, Melo PS, Cordi L, Martins D (2007) Violacein: properties and biological activities. Biotechnol Appl Biochem 48(3):127. https://doi.org/10.1042/BA20070115
Fan C, Davison PA, Habgood R, Zeng H, Decker CM, Gesell SM, Lueangwattanapong K, Townley HE, Yang A, Thompson IP, Ye H, Cui Z, Schmidt F, Hunter CN, Huang WE (2020) Chromosome-free bacterial cells are safe and programmable platforms for synthetic biology. Proc Natl Acad Sci USA 117(12):6752–6761. https://doi.org/10.1073/pnas.1918859117
Fang M, Zhang C, Yang S, Cui J, Jiang P, Lou K, Wachi M, Xing X (2015) High crude violacein production from glucose by Escherichia coli engineered with interactive control of tryptophan pathway and violacein biosynthetic pathway. Microb Cell Fact. https://doi.org/10.1186/s12934-015-0192-x
Floras N, Xiao J, Berry A, Bolivar F, Valle F (1996) Pathway engineering for the production of aromatic compounds in Escherichia coli. Nat Biotechnol 14(5):620–623. https://doi.org/10.1038/nbt0596-620
Fordjour E, Adipah FK, Zhou S, Du G, Zhou J (2019) Metabolic engineering of Escherichia coli BL21 (DE3) for de novo production of l-DOPA from d-glucose. Microb Cell Fact. https://doi.org/10.1186/s12934-019-1122-0
García S, Flores N, De Anda R, Hernández G, Gosset G, Bolívar F, Escalante A (2017) The role of the ydiB Gene, which encodes Quinate/Shikimate dehydrogenase, in the production of Quinic, Dehydroshikimic and Shikimic acids in a PTS- strain of Escherichia coli. J Mol Microb Biotechnol 27(1):11–21. https://doi.org/10.1159/000450611
Gosset G (2005) Improvement of Escherichia coli production strains by modification of the phosphoenolpyruvate:sugar phosphotransferase system. MIcrob Cell Fact 4(1):14. https://doi.org/10.1186/1475-2859-4-14
Gottlieb K, Albermann C, Sprenger GA (2014) Improvement of L-phenylalanine production from glycerol by recombinant Escherichia coli strains: the role of extra copies of glpK, glpX, and tktA genes. Microb Cell Fact 13(1):96. https://doi.org/10.1186/s12934-014-0096-1
Gu P, Fan X, Liang Q, Qi Q, Li Q (2017) Novel technologies combined with traditional metabolic engineering strategies facilitate the construction of shikimate-producing Escherichia coli. Microb Cell Fact. https://doi.org/10.1186/s12934-017-0773-y
Gu P, Kang J, Yang F, Wang Q, Liang Q, Qi Q (2013) The improved l-tryptophan production in recombinant Escherichia coli by expressing the polyhydroxybutyrate synthesis pathway. Appl Microbiol Biotechnol 97(9):4121–4127. https://doi.org/10.1007/s00253-012-4665-0
Gu P, Yang F, Kang J, Wang Q, Qi Q (2012) One-step of tryptophan attenuator inactivation and promoter swapping to improve the production of L-tryptophan in Escherichia coli. Microb Cell Fact 11(1):30. https://doi.org/10.1186/1475-2859-11-30
Gu P, Yang F, Li F, Liang Q, Qi Q (2013) Knocking out analysis of tryptophan permeases in Escherichia coli for improving L-tryptophan production. Appl Microbiol Biotechnol 97(15):6677–6683. https://doi.org/10.1007/s00253-013-4988-5
Gupta A, Reizman IMB, Reisch CR, Prather KLJ (2017) Dynamic regulation of metabolic flux in engineered bacteria using a pathway-independent quorum-sensing circuit. Nat Biotechnol 35(3):273–279. https://doi.org/10.1038/nbt.3796
Hideo Kawaguchi HMSN, Chiaki Ogino AAK (2019) Enhanced phenyllactic acid production in Escherichia coli via oxygen limitation and shikimate pathway gene expression. Biotechnol J. https://doi.org/10.1002/biot.201800478
Imamoto GBLS (1984) Transcription of the trpR gene of Escherichia coil: an autogeneously regulated system studied by direct measurements of mRNA levels in vivo. Mol Gen Genet 193:244–250. https://doi.org/10.1007/bf00330675
Jiang M, Zhang H (2016) Engineering the shikimate pathway for biosynthesis of molecules with pharmaceutical activities in E. coli. Curr Opin Biotech 42:1–6. https://doi.org/10.1016/j.copbio.2016.01.016
Jiang P, Wang H, Zhang C, Lou K, Xing X (2010) Reconstruction of the violacein biosynthetic pathway from Duganella sp. B2 in different heterologous hosts. Appl Microbiol Biotechnol 86(4):1077–1088. https://doi.org/10.1007/s00253-009-2375-z
Johansson L, Lindskog A, Silfversparre G, Cimander C, Nielsen KF, Lidén G, Department OCE, Lund U, Institutionen FK, Lunds U (2005) Shikimic acid production by a modified strain of E. coli (W3110.shik1) under phosphate-limited and carbon-limited conditions. Biotechnol Bioeng 92(5):541. https://doi.org/10.1002/bit.20546
Kawaguchi H, Katsuyama Y, Danyao D, Kahar P, Nakamura-Tsuruta S, Teramura H, Wakai K, Yoshihara K, Minami H, Ogino C, Ohnishi Y, Kondo A (2017) Caffeic acid production by simultaneous saccharification and fermentation of kraft pulp using recombinant Escherichia coli. Appl Microbiol Biotechnol 101(13):5279–5290. https://doi.org/10.1007/s00253-017-8270-0
Krämer M, Bongaerts J, Bovenberg R, Kremer S, Müller U, Orf S, Wubbolts M, Raeven L (2003) Metabolic engineering for microbial production of shikimic acid. Metab Eng 5(4):277–283. https://doi.org/10.1016/j.ymben.2003.09.001
Lee D (2008) Exploring the effects of carbon sources on the metabolic capacity for exploring the effects of carbon sources on the metabolic capacity for predictions. J Microbiol Biotechnol. https://doi.org/10.4014/jmb.0700.705
Lee M, Hung W, Tsai S (2017) Improvement of shikimic acid production in Escherichia coli with growth phase-dependent regulation in the biosynthetic pathway from glycerol. World J Microbiol Biotechnol. https://doi.org/10.1007/s11274-016-2192-3
Li Q, Wu H, Li Z, Ye Q (2016) Enhanced succinate production from glycerol by engineered Escherichia coli strains. Bioresour Technol 218:217–223. https://doi.org/10.1016/j.biortech.2016.06.090
Lin Y, Sun X, Yuan Q, Yan Y (2014) Extending shikimate pathway for the production of muconic acid and its precursor salicylic acid in Escherichia coli. Metab Eng 23:62–69. https://doi.org/10.1016/j.ymben.2014.02.009
Liu C, Men X, Chen H, Li M, Ding Z, Chen G, Wang F, Liu H, Wang Q, Zhu Y, Zhang H, Xian M (2018) A systematic optimization of styrene biosynthesis in Escherichia coli BL21(DE3). Biotechnol Biofuels. https://doi.org/10.1186/s13068-018-1017-z
Liu L, Chen S, Wu J (2017) Phosphoenolpyruvate:glucose phosphotransferase system modification increases the conversion rate during l-tryptophan production in Escherichia coli. J Ind Microbiol Biotechnol 44(10):1385–1395. https://doi.org/10.1007/s10295-017-1959-3
Liu Q, Cheng Y, Xie X, Xu Q, Chen N (2012) Modification of tryptophan transport system and its impact on production of l-tryptophan in Escherichia coli. Bioresour Technol 114:549–554. https://doi.org/10.1016/j.biortech.2012.02.088
Liu S, Xiao M, Zhang L, Xu J, Ding Z, Gu Z, Shi G (2013) Production of l-phenylalanine from glucose by metabolic engineering of wild type Escherichia coli W3110. Process Biochem 48(3):413–419. https://doi.org/10.1016/j.procbio.2013.02.016
Liu SP, Liu RX, Xiao MR, Zhang L, Ding ZY, Gu ZH, Shi GY (2014) A systems level engineered E. coli capable of efficiently producing L-phenylalanine. Process Biochem 49(5):751–757. https://doi.org/10.1016/j.procbio.2014.01.001
Liu X, Lin J, Hu H, Zhou B, Zhu B (2014) Metabolic engineering of Escherichia coli to enhance shikimic acid production from sorbitol. World J Microbiol Biotechnol 30(9):2543–2550. https://doi.org/10.1007/s11274-014-1679-z
Liu X, Lin J, Hu H, Zhou B, Zhu B (2016) Enhanced production of shikimic acid using a multi-gene co-expression system in Escherichia coli. Chin J Nat Med 14(4):286–293. https://doi.org/10.1016/S1875-5364(16)30029-2
Liu X, Lin J, Hu H, Zhou B, Zhu B (2016) Site-specific integration and constitutive expression of key genes into Escherichia coli chromosome increases shikimic acid yields. Enzyme Microb Technol 82:96–104. https://doi.org/10.1016/j.enzmictec.2015.08.018
Liu Y, Xu Y, Ding D, Wen J, Zhu B, Zhang D (2018) Genetic engineering of Escherichia coli to improve L-phenylalanine production. BMC Biotechnol 18(1):5–12. https://doi.org/10.1186/s12896-018-0418-1
Luo W, Huang J, Zhu X, Huang L, Cai J, Xu Z (2013) Enhanced production of l-tryptophan with glucose feeding and surfactant addition and related metabolic flux redistribution in the recombinant Escherichia coli. Food Sci Biotechnol 22(1):207–214. https://doi.org/10.1007/s10068-013-0029-5
Luo Y, Zhang T, Wu H (2014) The transport and mediation mechanisms of the common sugars in Escherichia coli. Biotechnol Adv 32(5):905–919. https://doi.org/10.1016/j.biotechadv.2014.04.009
Lütke-Eversloh T, Stephanopoulos G (2008) Combinatorial pathway analysis for improved L-tyrosine production in Escherichia coli: identification of enzymatic bottlenecks by systematic gene overexpression. Metab Eng 10(2):69–77. https://doi.org/10.1016/j.ymben.2007.12.001
Martínez JA, Rodriguez A, Moreno F, Flores N, Lara AR, Ramírez OT, Gosset G, Bolivar F (2018) Metabolic modeling and response surface analysis of an Escherichia coli strain engineered for shikimic acid production. BMC Syst Biol. https://doi.org/10.1186/s12918-018-0632-4
Masuo S, Kobayashi Y, Oinuma K, Takaya N (2016) Alternative fermentation pathway of cinnamic acid production via phenyllactic acid. Appl Microbiol Biotechnol 100(20):8701–8709. https://doi.org/10.1007/s00253-016-7623-4
Muir RM, Ibáñez AM, Uratsu SL, Ingham ES, Leslie CA, McGranahan GH, Batra N, Goyal S, Joseph J, Jemmis ED, Dandekar AM (2011) Mechanism of gallic acid biosynthesis in bacteria (Escherichia coli) and walnut (Juglans regia). Plant Mol Biol 75(6):555–565. https://doi.org/10.1007/s11103-011-9739-3
Noda S, Shirai T, Oyama S, Kondo A (2015) Metabolic design of a platform Escherichia coli strain producing various chorismate derivatives. Metab Eng 33:119–129. https://doi.org/10.1016/j.ymben.2015.11.007
Pontrelli S, Chiu T, Lan EI, Chen FYH, Chang P, Liao JC (2018) Escherichia coli as a host for metabolic engineering. Metab Eng 50:16–46. https://doi.org/10.1016/j.ymben.2018.04.008
Qian S, Li Y, Cirino PC (2019) Biosensor-guided improvements in salicylate production by recombinant Escherichia coli. Microb Cell Fact. https://doi.org/10.1186/s12934-019-1069-1
Ranjan Patnaik RGSA (1995) Pathway engineering for production of aromatics in Escherichia coli: confirmation of stoichiometric analysis by independent modulation of AroG, TktA, and Pps activities. Biotechnol Bioeng 46(4):361–370. https://doi.org/10.1002/bit.260460409
Rodrigues AL, Trachtmann N, Becker J, Lohanatha AF, Blotenberg J, Bolten CJ, Korneli C, de Souza Lima AO, Porto LM, Sprenger GA, Wittmann C (2013) Systems metabolic engineering of Escherichia coli for production of the antitumor drugs violacein and deoxyviolacein. Metab Eng 20:29–41. https://doi.org/10.1016/j.ymben.2013.08.004
Rodriguez A, Martínez JA, Flores N, Escalante A, Gosset G, Bolivar F (2014) Engineering Escherichia coli to overproduce aromatic amino acids and derived compounds. Microb Cell Fact 13(1):126. https://doi.org/10.1186/s12934-014-0126-z
Santos CNS, Xiao W, Stephanopoulos G (2012) Rational, combinatorial, and genomic approaches for engineering L-tyrosine production in Escherichia coli. Proc Natl Acad Sci 109(34):13538–13543. https://doi.org/10.1073/pnas.1206346109
Wang J, Cheng L, Wang J, Liu Q, Shen T, Chen N (2013) Genetic engineering of Escherichia coli to enhance production of l-tryptophan. Appl Microbiol Biotechnol 97(17):7587–7596. https://doi.org/10.1007/s00253-013-5026-3
Wang J, Shen X, Rey J, Yuan Q, Yan Y (2018) Recent advances in microbial production of aromatic natural products and their derivatives. Appl Microbiol Biotechnol 102(1):47–61. https://doi.org/10.1007/s00253-017-8599-4
Wang J, Shen X, Yuan Q, Yan Y (2018) Microbial synthesis of pyrogallol using genetically engineered Escherichia coli. Metab Eng 45:134–141. https://doi.org/10.1016/j.ymben.2017.12.006
Wolfsberg E, Long CP, Antoniewicz MR (2018) Metabolism in dense microbial colonies: 13C metabolic flux analysis of E. coli grown on agar identifies two distinct cell populations with acetate cross-feeding. Metab Eng 49:242–247. https://doi.org/10.1016/j.ymben.2018.08.013
Xiaolin Shen JW, Qipeng Yuan YY (2017) High-level de novo biosynthesis of arbutin in engineered Escherichia coli. Metab Eng. https://doi.org/10.1016/j.ymben.2017.06.001
Xu Q, Bai F, Chen N, Bai G (2017) Gene modification of the acetate biosynthesis pathway in Escherichia coli and implementation of the cell recycling technology to increase L-tryptophan production. PLoS ONE 12(6):e179240. https://doi.org/10.1371/journal.pone.0179240
Yang J, Camakaris H, Pittard AJ (1993) Mutations in the tyrR gene of Escherichia coli which affect tyrR-mediated activation but not tyrR-mediated repression. J Biotechnol 175(19):6372–6375. https://doi.org/10.1128/jb.175.19.6372-6375.1993
Yi J, Draths KM, Li K, Frost JW (2003) Altered Glucose Transport and Shikimate Pathway Product Yields in E. coli. Biotechnol Prog 19(5):1450–1459. https://doi.org/10.1021/bp0340584
Zhang J, Li X (2018) Novel strategy for phenyllactic acid biosynthesis from phenylalanine by whole cell recombinant Escherichia coli coexpressing l-phenylalanine oxidase and l-lactate dehydrogenase. Biotechnol Lett 40(1):165–171. https://doi.org/10.1007/s10529-017-2456-5
Zhou Y, Fang M, Li G, Zhang C, Xing X (2018) Enhanced production of crude violacein from glucose in Escherichia coli by overexpression of rate-limiting key enzyme(s) involved in violacein biosynthesis. Appl Biochem Biotechnol 186(4):909–916. https://doi.org/10.1007/s12010-018-2787-2
Zhu S, Wu J, Du G, Zhou J, Chen J (2014) Efficient synthesis of eriodictyol from l-tyrosine in Escherichia coli. Appl Environ Microb 80(10):3072–3080. https://doi.org/10.1128/AEM.03986-13
Zhucui Li YLYJ (2018) Exploiting high-resolution mass spectrometry for targeted metabolite quantification and 13C-labeling metabolism analysis. Microb Metab. https://doi.org/10.1007/978-1-4939-8757-3_9
Acknowledgements
This work was supported by the National Key R&D Program of China (2018YFA0900302), the National Natural Science Foundation of China (21621004, 31900052), the Tianjin Science Fund for Distinguished Young Scholars (17JCJQJC45300), the Science and Technology Service Network (STS) Initiative of the Chinese Academy of Sciences (KFJ-STS-ZDTP-065), and he National Key R&D Program of China (2018YFA0901600).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Li, Z., Wang, H., Ding, D. et al. Metabolic engineering of Escherichia coli for production of chemicals derived from the shikimate pathway. J Ind Microbiol Biotechnol 47, 525–535 (2020). https://doi.org/10.1007/s10295-020-02288-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-020-02288-2