Abstract
Ethyl acetate has attracted much attention as an important chemical raw material and a flavor component of alcoholic beverages. In this study, the biosynthetic pathway for the production of ethyl acetate in Chinese liquor yeast was unblocked. In addition to engineering Saccharomyces cerevisiae to increased intracellular CoA and acetyl-CoA levels, we also increased the combining efficiency of acetyl-CoA to ethanol. The genes encoding phosphopantothenate-cysteine ligase, acetyl-CoA synthetase, and alcohol acetyltransferase were overexpressed by inserting the strong promoter PGK1p and the terminator PGK1t, respectively, and then combine them. Our results finally showed that the ethyl acetate levels of all engineering strains were improved. The final engineering strain CLy12a-ATF1-ACS2-CAB2 had a significant increase in ethyl acetate yield, reaching 610.26 (± 14.28) mg/L, and the yield of higher alcohols was significantly decreased. It is proved that the modification of ethyl acetate metabolic pathway is extremely important for the production of ethyl acetate from Saccharomyces cerevisiae.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Carboxylate esters such as ethyl acetate and isoamyl acetate are lipophilic molecules that generally have low odor thresholds. Ethyl acetate is widely present in many types of fermented alcoholic beverages, providing a pleasant fruit aroma that affects sensory quality and flavor characteristics [10]. In the freshly brewed liquor of Chinese Fenjiu, the content of ethyl acetate exceeds 2000 mg/L [11]. How to effectively improve and control the ethyl acetate production have always been the research focus of alcohol beverage improvement [16]. In industrial production, the synthesis of ethyl acetate is mainly accomplished by standard chemical means [9]. However, a recent bioprocessing pilot plant using K. marxianus produced 10.9 g/L ethyl acetate titer from waste whey feed, providing a new idea for industrial production of ethyl acetate [8]. Ethyl acetate is produced as a secondary metabolite in the alcohol fermentation process through three metabolic pathways [23]. The use of yeast to produce ethyl acetate is a potential option in the future [17].
Ethyl acetate production can be improved to a certain extent by improving and optimizing the production process, but the degree of improvement is not obvious [3]. Genetic engineering technology has brought us new thoughts. One possible limiting factor is the lack of supply of reactive precursor materials [4, 5]. A significant increase in CoA level was observed following overexpression of the rate-controlling enzyme pantothenate kinase and supplementation of the precursor pantothenic acid simultaneously [19]. In yeast, YIL083C encoding a phosphopantothenate-cysteine ligase has been identified (gene designation: CAB2) [12]. In the metabolic pathway from pantothenic acid to coenzyme A, gene CAB2 responsible for late CoA biosynthesis is turned out to be essential [13]. The ACS2 knockout strain lost its abilities to grow on glucose media, demonstrating that the ACS2 gene is essential for the utilization of glucose in the fermentation medium [20]. Previous studies have demonstrated that overexpression of the ACS2 gene increases the ability of Saccharomyces cerevisiae to synthesize acetyl-CoA, while acetic acid tolerance is also enhanced [2]. The alcohol acetyltransferases are encoded by the ATF1, ATF2, and Lg-ATF1 genes and have been widely recognized as the most important enzymes in the production of ethyl acetate by yeast [7, 22]. Compared with the control, the expression intensity of ATF1 gene significantly increased the concentration of ethyl acetate, while ATF2 and Lg-ATF1 demonstrate lower degree of effect [7, 22]. The activity of AATase and the ethyl acetate production was dramatically reduced due to the null mutation of ATF1 [21].
In this work, we aimed to modify the expression intensity of target genes by inserting the PGK1 promoter and terminator, and have successfully constructed overexpression engineering strains of CAB2, ACS2 and ATF1 separately, and constructed the combined strains simultaneously. Our target was to enhance catalysis and combination efficiency while increase the production of CoA and acetyl-CoA which directed the metabolism to the ethyl acetate pathway. The interaction between CAB2, ACS2, and ATF1 genes and their effects on ethyl acetate production were studied by fermentation experiments in corn hydrolysate. The results showed that the positive change of ethyl acetate production can be detected in all engineering strains, and the combined overexpression of ACS2 and ATF1 had a more obvious synergistic effect. This strategy resulted in a significant increase in ethyl acetate titers and further explored the role of related genes in the production of Saccharomyces cerevisiae flavors. In addition, this study laid the foundation for industrial production of ethyl acetate and has certain reference value for the future optimization of other alcoholic beverages.
Materials and methods
Strains and growth conditions
All yeast strains and plasmids used in this study are summarized in Table 1. Escherichia coli strain DH5a was utilized for plasmid propagation. Haploid strain CLy12a was used as the background strain in this work. Lysogeny broth (LB) medium (5 g/L yeast extract, 5 g/L NaCl and 10 g/L peptone, pH 7.0) at 37 °C, with 100 mg/L ampicillin added for plasmid selection. Yeast strains were cultured in YEPD medium (20 g/L peptone and 20 g/L glucose, 10 g/L yeast extract, pH 7.0) at 30 °C. The YEPD medium was added with a 400 mg/L G418 during the screening of positive yeast transformants. To select Zeocin-resistant yeast strains, 500 mg/L Zeocin was added to the YEPD plates. Then, the galactose medium (20 g/L peptone and 20 g/L galactose, 10 g/L yeast extract, pH 7.0) was used for the Cre recombinase expression in the yeast. All solid media used in this study contained 2% agar powder. Liquid YEPD medium was used for cell culture, transcription level analysis and enzyme activity assay.
Yeast strain construction
The polymerase chain reaction (PCR) primers used in this study are listed in Table 2. The fragment of upstream flank of CAB2 (U) was amplified using primer pair 1-KCAB2-U and 1-KCAB2-D. 6-KCAB2-U and 6-KCAB2-D were used to obtain the downstream flanking fragment CAB2 (D). The PGK1p promoter and the PGK1t terminator were amplified from primer pairs 2-KCAB2-U and 2-KCAB2-D, 4-KCAB2-U and 4-KCAB2-D, respectively. The gene CAB2 was amplified from genomic DNA isolated from the strain CLy12a using primer pair 3-KCAB2-U and 3-KCAB2-D. To obtain KanMX resistance, the PUG6 plasmid was used as a template, and the primer pair 5-KCAB2-U and 5-KCAB2-D were used for PCR amplification [1]. An overlapping sequence of approximately (20–25 bp) was added to the ligation terminus of each fragment to complete chimerization during intracellular homologous recombination. The six fragments verified by agarose gel electrophoresis were purified and recovered, and then all fragments were transferred into CLy12a by lithium acetate conversion to complete intracellular recombination, and the strain CLy12a-CAB2 was constructed. A similar procedure was applied to the construction of strains CLy12a-ACS2 and CLy12a-ATF1. The Cre-LoxP system was used to remove the KanMX resistance markers of the strains CLy12a-ACS2 and CLy12a-ATF1 to obtain strains CLy12a-ACS2-ΔKanMX and CLy12a-ATF1-ΔKanMX. The strain CLy12a-ACS2-CAB2 was constructed by the above method using CLy12a-ACS2-ΔKanMX as a starting strain. At the same time, CLy12a-ATF1-ΔKanMX was used as the starting strain, CLy12a-ATF1-CAB2 and CLy12a-ATF1-ACS2 were also obtained. The KanMX resistance marker of the CLy12a-ATF1-ACS2 strain was removed, and the strain CLy12a-ATF1-ACS2-ΔKanMX was obtained. On this basis, the CAB2 gene was overexpressed to obtain the final engineering strain CLy12a-ATF1-ACS2-CAB2.
Fermentation experiments
To prepare a liquid fermentation medium, 3000 g of corn flour was weighed, mixed with 9 L of water, and kept at 60–70 °C for 20 min. A high temperature resistant α-amylase 1.8 mL (2 × 105 U/mL; Novozymes Biotech Co., Ltd., Tianjin, China) was added to the mixture and incubated at 90 °C for 90 min in a water bath. Stirring was carried out periodically during the liquefaction process, and the liquefaction end point was based on iodine test blue without reddish brown. Then, the temperature of the mixture was lowered to 65 °C, and 5 M H2SO4 was used to adjust pH to 4.2–4.4, and saccharification enzyme 3 mL (200 U/mL; Novozymes Biotechnology Co. Ltd., Tianjin, China) was added at 55–60 °C for 20 h to complete the saccharification. After the mixture was cooled to room temperature, the residue was removed by filtration through gauze and the supernatant was retained. Yeast cells were cultured in 5 mL of eight Brix medium (additional 0.5% yeast extract) at 30 °C for 24 h and were transferred to 45 mL of 12 Brix medium (additional 0.5% yeast extract) at 30 °C for 16 h. A total of 15 mL of second precultured yeast cells were dispensed into 135 mL of 20 Brix medium while supplementing 1 mL of nutrient salt. The erlenmeyer flask (250 mL) was tightly sealed with parafilm for oxygen-limited fermentation and a carbon dioxide-releasing channel was created on parafilm with a 0.5 mm injection needle. The fermentation temperature was 30 °C. The weight loss results were measured every 12 h, and if the difference of two weighing results were less than 1 g comparably then the fermentation was completed. All fermentations were performed in triplicate. The CO2 loss was measured by an analytical balance.
Gas chromatography analysis
Gas chromatography analysis has been widely used in the detection of volatile compounds in liquor. The contents of ethyl acetate and higher alcohols were analyzed by gas chromatography analysis. Distillation was carried out after the fermentation of the hydrolysate was completed, and sample analysis was performed on an Agilent 7890C gas chromatograph equipped with an Agilent G4513A auto sampler, injector and flame ionization detector (FID) and an AT.LZP-930 column (50 m × 320 μm internal diameter and 1 μm coating thickness). The GC was used as follows: nitrogen was used as carrier gas at a constant flow rate of 1 mL/min. The injector temperature was 200 °C, split ratio was 5:1, and injection volume was 1 μL. The oven temperature program used was 50 °C (8 min), then raised to 120 °C at the speed of 5 °C/min and held at the final temperature for 8 min. The standard used in the measurement was purchased from Merck Serono Co. Ltd.
High performance liquid chromatography analysis
At the end of the fermentation, the hydrolysates were separately collected and subjected to centrifugation. The contents of residual acetic acid were analyzed by high performance liquid chromatography. Each sample was filtered through a 0.22 μm aqueous phase filter before liquid chromatography. Using 1200SL liquid chromatograph, the column was HPX-87H (300 mm × 7.8 mm × 9 μm), DAD detector, 5 mmol/L dilute sulfuric acid solution as mobile phase, oven temperature 60 °C, flow rate 0.6 mL/min, The detection time was 23 min, and the injection amount per sample was 20 μL.
Real-time quantitative PCR (RT-qPCR)
Each strain was cultured in YEPD medium to logarithmic growth phase, and mRNA was isolated using a UNIQ-10 column Trizol total RNA extraction kit. Reverse transcription of mRNA into cDNA (RevertAid Premium Reverse Transcriptase). Real-time PCR was performed using a SG Fast qPCR Master Mix and an ABI StepOne plus type real-time PCR instrument, and UBC6 gene was used as a reference gene. The final data were calculated using the threshold cycle (2−△△CT) method.
Results
Characterization of engineering strains
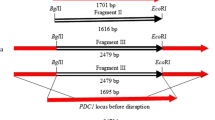
The genes of CAB2, ACS2, and ATF1 were overexpressed individually and collectively as described in materials and methods. The strains and plasmids are shown in Table 1, and the primers are listed in Table 2, which are involved in this study. The genetic manipulation of the intracellular homologous recombination method was used to complete the targeted transformation of the strains. The Cre-LoxP system facilitated the removal of the KanMX screening marker to assist the subsequent operations (the specific implementation strategy is shown in Fig. 1). The engineering strains and the parent strain were separately cultured in a YEPD liquid medium at 30 °C to confirm whether the growth rate of the strains was consistent and not damaged.
Fermentation performance
The fermentation properties of engineering strains were compared with CLy12a in the corn hydrolysate. The relevant fermentation attributes of all strains were recorded to assess their abilities to ferment in corn hydrolysate (Table 3). The fermentation time of all the strains was consistent, and the total weight loss of CO2 did not change significantly. The results of high performance liquid chromatography analysis showed that during the fermentation, all strains produced approximately the same levels of ethanol, and glucose was almost depleted.
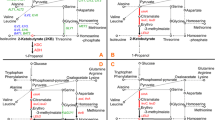
As shown in Fig. 2a, the ethyl acetate production of all engineering strains was detected to have different degrees of increase. The ethyl acetate yield of CLy12a-ATF1 strain was 414.40 mg/L, which was most obvious in the single-gene overexpressing strains CLy12a-CAB2, CLy12a-ACS2, and CLy12a-ATF1. The ethyl acetate production of the multi-gene overexpression strains constructed on the basis of the single-gene overexpression strains were further improved. The strain CLy12a-ATF1-ACS2-CAB2 had the highest ethyl acetate content of 610.26 mg/L, which was more than 60-fold higher than the 9.22 mg/L of the original strain CLy12a. The residual acetic acid in the fermentation broth of the strain CLy12a-ATF1-ACS2-CAB2 was almost completely consumed, and only 52.80 mg/L was detected, which was more than 80% lower than CLy12a. For all engineering strains, the higher the ethyl acetate yield is, the lower is the residual acetic acid, which is consistent with the expected experimental results. The higher alcohol contents are shown in Fig. 2b. Compared with CLy12a, there was no significant change in the higher alcohol contents of the engineering strains CLy12a-CAB2, CLy12a-ACS2, and CLy12a-CAB2-ACS2. The strains of CLy12a-ATF1 had the highest decrease in n-propanol, isobutanol, and phenylethanol content, which decreased by 26, 33, and 27%, respectively. The strains overexpressing CAB2 and ACS2 genes based on CLy12a-ATF1 strain (CLy12a-ATF1-CAB2, CLy12a-ATF1-ACS2 and CLy12a-ATF1-ACS2-CAB2) have lower n-propanol, isobutanol and phenylethanol contents than the original strain CLy12a but higher than the strain CLy12a-ATF1. All engineering strains showed that the higher the ethyl acetate production is,the is lower the isoamyl alcohol contents. Among them, the contents of isoamyl alcohol in CLy12a-ATF1-ACS2-CAB2 decreased the most, reaching 62%.
a The bar graph represents the average concentration of ethyl acetate and acetic acid produced by the strains, and the high yield of ethyl acetate is accompanied by low yields of acetic acid residues. Data are presented as means and standard deviations of three independent biological replicates. b The difference in higher alcohol production between strains. The maximum decrease of n-propanol was 26.35%, the maximum decrease of isobutanol was 33.53%, the maximum decrease of isoamyl alcohol was 55.93%, and the maximum decrease of phenylethanol was 26.76%. Data are presented as means and standard deviations of three independent biological replicates
Analysis of gene expression
It is noteworthy that there were significant differences in ethyl acetate and higher alcohol productions among the engineering strains. We quantified the mRNA expression levels of CAB2, ACS2, and ATF1 to clarify the relationship among these genes (Fig. 3). The results of RT-qPCR suggest that the CAB2 expression level of CLy12a-CAB2, the ACS2 expression level of CLy12a-ACS2, and the ATF1 expression level of CLy12a-ATF1 were 11.1-, 2.3-, and 5.8-fold higher than that of CLy12a, respectively. The above results confirmed that the target gene mRNA levels of the engineering strains were improved under the action of the PGK1p promoter and the PGK1t terminator. A new change in mRNA levels was detected for the combined overexpressed strains. For the final engineering strain CLy12a-CAB2-ACS2-ATF1, the mRNA level of CAB2 gene was 1.5-fold higher than that of CLy12a-CAB2, the mRNA level of ACS2 gene was 3.4-fold higher than that of CLy12a-ACS2, and the mRNA level of ATF1 gene was 3.0-fold higher than that of CLy12a-ATF1. Similar experimental results were also detected in the remaining engineering strains. The above results have demonstrated that the combined overexpression of the CAB2, ACS2, and ATF1 genes in the metabolic pathway results in a superimposed effect of mutual promotion. Overexpression of the CAB2, ACS2, and ATF1 genes are effective for the ability of Saccharomyces cerevisiae to produce ethyl acetate. The mRNA levels of all engineering strains were positively correlated with their ethyl acetate contents, which was consistent with the expected experimental results.
Differences in mRNA levels between strains. All of the overexpressed genes showed a significant increase in the corresponding mRNA levels. The production of multi-gene overexpression strains based on single-gene overexpression strains showed higher mRNA levels, confirming the interaction between genes. Data are presented as means and standard deviations of three independent biological replicates
Discussion
Due to its wide range of functions, some studies have begun to explore the ethyl acetate pathway of Saccharomyces cerevisiae [15, 17]. It is very important and necessary to find a yeast strain capable of producing a large amount of ethyl acetate. Previous related research results have not been sufficient for the production improvement of ethyl acetate, and efforts in the direction of metabolic engineering have not been perfected. The focus of this study was to regulate the expression levels of genes involved in the pathway of ethyl acetate metabolism.
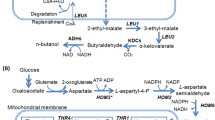
CAB2, ACS2, and ATF1 are important genes in the ethyl acetate metabolic pathway. The phosphonantothenate encoded by the CAB2 gene affects the contents of the precursor substance coenzyme A. The ACS2 gene encodes a “non-aerobic” type acetyl-CoA synthetase. Ethyl acetate is produced by the combination of ethanol and acetyl-CoA under the catalysis of an alcohol acetyltransferase encoded by the ATF1 gene. Previous studies have shown that the strength of the promoter and terminator largely determines the transcriptional activity of genes [14]. The main method in this study was to increase their expression levels by inserting the strong promoter PGK1p and the strong terminator PGK1t before and after the relevant metabolic genes. Different from previous studies, to obtain more stable strains performance, the construction of plasmids was omitted, and the genetic modification processes were directly integrated into the yeast genome, which greatly simplified the frequency of molecular manipulation and reduced the possibility of strain damage. Based on the basic fermentation data (Table 3), it was confirmed that the basic fermentation characteristics of the engineering yeast strains were not affected by the genetic modification. Overexpression of the ATF1 gene resulted in a significant increase in ethyl acetate production, which was consistent with the previous studies. Because of the extra addition of the PGK1 terminator and the synergetic effect of simultaneous overexpression of the three genes, the ethyl acetate production of the final engineering strain increased more than 60-fold compared to the original strain. This result is far superior to the ethyl acetate production (tenfold to 20-fold) in previous studies [6, 7]. At the same time, the promotion and superposition effects of CAB2, ACS2, and ATF1 genes were confirmed for the first time. The improvement of mRNA levels also provided strong evidence for the validity of this hypothesis (Fig. 3). In addition, the total production of higher alcohols showed a different degree of decline due to the unblocking of the ethyl acetate metabolic pathway, which is consistent with the results of other studies [16].
To obtain better and superior performance, it is necessary to further optimize the metabolic pathway. The low residual acetic acid production of the engineering strains (Fig. 2) provides an idea for further increasing the ethyl acetate level. Another strategy that may be of concern is the blocking of related branched metabolic pathways such as butanol and glycerol [4, 18]. In this study, we confirmed the interaction between CAB2, ACS2, and ATF1 by our engineering strategy, which is of great significance for studying the metabolic mechanism of ethyl acetate. The results of this study provide an effective method for the optimization of yeast strains in the future, which has a good reference value for improving the quality and flavor of wine, and provides a possibility for the new industrialized production mode of ethyl acetate.
References
Agaphonov M, Romanova N, Choi ES, Ter-Avanesyan M (2010) A novel kanamycin/G418 resistance marker for direct selection of transformants in Escherichia coli and different yeast species. Yeast 27:189–195. https://doi.org/10.1002/yea.1741
Ding J, Holzwarth G, Penner MH, Pattonvogt J, Bakalinsky AT (2015) Overexpression of acetyl-CoA synthetase in Saccharomyces cerevisiae increases acetic acid tolerance. FEMS Microbiol Lett 362:1. https://doi.org/10.1093/femsle/fnu042
Hiralal L, Olaniran AO, Pillay B (2014) Aroma-active ester profile of ale beer produced under different fermentation and nutritional conditions. J Biosci Bioeng 117:57–64. https://doi.org/10.1016/j.jbiosc.2013.06.002
Krivoruchko A, Serrano-Amatriain C, Chen Y, Siewers V, Nielsen J (2013) Improving biobutanol production in engineered Saccharomyces cerevisiae by manipulation of acetyl-CoA metabolism. J Ind Microbiol Biotechnol 40:1051–1056. https://doi.org/10.1007/s10295-013-1296-0
Krivoruchko A, Zhang Y, Siewers V, Chen Y, Nielsen J (2015) Microbial acetyl-CoA metabolism and metabolic engineering. Metab Eng 28:28–42. https://doi.org/10.1016/j.ymben.2014.11.009
Li W, Wang JH, Zhang CY, Ma HX, Xiao DG (2017) Regulation of Saccharomyces cerevisiae genetic engineering on the production of acetate esters and higher alcohols during Chinese Baijiu fermentation. J Ind Microbiol Biotechnol 44:1–12. https://doi.org/10.1007/s10295-017-1907-2
Lilly M, Bauer FF, Lambrechts MG, Swiegers JH, Cozzolino D, Pretorius IS (2006) The effect of increased yeast alcohol acetyltransferase and esterase activity on the flavour profiles of wine and distillates. Yeast 23:641–659. https://doi.org/10.1002/yea.1382
Löser C, Urit T, Stukert A, Bley T (2013) Formation of ethyl acetate from whey by Kluyveromyces marxianus on a pilot scale. J Biotechnol 163:17–23. https://doi.org/10.1016/j.jbiotec.2012.10.009
Löser C, Urit T, Bley T (2014) Perspectives for the biotechnological production of ethyl acetate by yeasts. Appl Microbiol Biotechnol 98:5397–5415. https://doi.org/10.1007/s00253-014-5765-9
Loviso CL, Libkind D (2018) Síntesis y regulación de compuestos del aroma y el sabor derivados de la levadura en la cerveza: ésteres. Revista Argentina De Microbiología 50:436–446. https://doi.org/10.1016/j.ram.2017.11.006
Ma Y, Hua Q, Wei W, Chen T, Du X, Zhai X, Zhang S (2014) Variations in physicochemical properties of Chinese Fenjiu during storage and high-gravity technology of liquor aging. Int J Food Prop 17:923–936. https://doi.org/10.1080/10942912.2012.678536
Olzhausen J, Schübbe S, Schüller HJ (2009) Genetic analysis of coenzyme A biosynthesis in the yeast Saccharomyces cerevisiae: identification of a conditional mutation in the pantothenate kinase gene CAB1. Curr Genet 55:163–173. https://doi.org/10.1007/s00294-009-0234-1
Olzhausen J, Moritz T, Neetz T, Schüller HJ (2013) Molecular characterization of the heteromeric coenzyme A-synthesizing protein complex (CoA-SPC) in the yeast Saccharomyces cerevisiae. FEMS Yeast Res 13:565–573. https://doi.org/10.1111/1567-1364.12058
Partow S, Siewers V, Bjorn S, Nielsen J, Maury J (2010) Characterization of different promoters for designing a new expression vector in Saccharomyces cerevisiae. Yeast 27:955–964. https://doi.org/10.1002/yea.1806
Pires EJ, Teixeira JA, Brányik T, Vicente AA (2014) Yeast: the soul of beer’s aroma—a review of flavour-active esters and higher alcohols produced by the brewing yeast. Appl Microbiol Biotechnol 98:1937–1949. https://doi.org/10.1007/s00253-013-5470-0
Procopio S, Qian F, Becker T (2011) Function and regulation of yeast genes involved in higher alcohol and ester metabolism during beverage fermentation. Eur Food Res Technol 233:721–729. https://doi.org/10.1007/s00217-011-1567-9
Saerens SM, Delvaux FR, Verstrepen KJ, Thevelein JM (2010) Production and biological function of volatile esters in Saccharomyces cerevisiae. Microb Biotechnol 3:165–177. https://doi.org/10.1111/j.1751-7915.2009.00106.x
Turner TL, Zhang GC, Kim SR, Subramaniam V, Steffen D, Skory CD, Jang JY, Yu BJ, Jin YS (2015) Lactic acid production from xylose by engineered Saccharomyces cerevisiae without PDC or ADH deletion. Appl Microbiol Biotechnol 99:8023–8033. https://doi.org/10.1007/s00253-015-6701-3
Vadali RV, Bennett GN, San KY (2004) Applicability of CoA/acetyl-CoA manipulation system to enhance isoamyl acetate production in Escherichia coli. Metab Eng 6:294–299. https://doi.org/10.1016/j.ymben.2004.02.006
Van den Berg MA, Steensma HY (1995) ACS2, a Saccharomyces cerevisiae gene encoding acetyl-coenzyme A synthetase, essential for growth on glucose. Eur J Biochem 231:704–713. https://doi.org/10.1111/j.1432-1033.1995.0704d.x
Verstrepen KJ, Van Laere SD, Vanderhaegen BM, Derdelinckx G, Dufour JP, Pretorius IS, Winderickx J, Thevelein JM, Delvaux FR (2003) Expression levels of the yeast alcohol acetyltransferase genes ATF1, Lg-ATF1, and ATF2 control the formation of a broad range of volatile esters. Appl Environ Microbiol 69:5228. https://doi.org/10.1128/aem.69.9.5228-5237.2003
Verstrepen KJ, Derdelinckx G, Dufour JP, Winderickx J, Pretorius IS, Thevelein JM, Delvaux FR (2010) The Saccharomyces cerevisiae alcohol acetyl transferase gene ATF1 is a target of the cAMP/PKA and FGM nutrient-signalling pathways. FEMS Yeast Res 4:285–296. https://doi.org/10.1049/ip-vis:20010314
Yong CP, Shaffer CEH, Bennett GN (2009) Microbial formation of esters. Appl Microbiol Biotechnol 85:13. https://doi.org/10.1007/s00253-009-2170-x
Acknowledgements
This work was funded by The National Natural Science Foundation of China (Grant no. 31471724), the Project Supported by the Foundation (No. ZXKF20180102) of Tianjin Engineering Research Center of Microbial Metabolism and Fermentation Process Control, P. R. China.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Dong, J., Wang, P., Fu, X. et al. Increase ethyl acetate production in Saccharomyces cerevisiae by genetic engineering of ethyl acetate metabolic pathway. J Ind Microbiol Biotechnol 46, 801–808 (2019). https://doi.org/10.1007/s10295-019-02142-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-019-02142-0