Abstract
Gobioid larvae collected from the coast of Okinawa Island, southern Japan, were identified on the basis of a combination of morphological characters and a sequence of a hypervariable region of mitochondrial 12S ribosomal RNA (12S rRNA) gene (138–145 base pairs). A short-term formalin fixation technique enabled identification using both morphological and genetic methods. Thirteen of the 21 types of gobioid larvae assessed were identified to the species level. Additionally, we described the morphology of the postflexion larvae of wormfish Paragunnellichthys sp. and shrimp-associated goby Ctenogobiops feroculus, identified for the first time in their respective genera.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gobioid fish, suborder Gobioidei (Teleostei), is an important constituent of larval fish assemblages in tropical and subtropical coastal waters (e.g. Clarke 1991; Maeda and Tachihara 2014; Isari et al. 2017). Nevertheless, the understanding of early life history of gobies is poorly known (Borges et al. 2011). This lack of current knowledge can largely be attributed to the fact that gobioid larvae are difficult to identify.
Identification of fish larvae has been traditionally based on morphological characters, which are often not sufficient to identify larval fish to the species level (Ko et al. 2013). Recently, genetic methods called DNA barcoding have been used to identify fish eggs and larvae (e.g. Victor et al. 2009; Kawakami et al. 2010; Marancik et al. 2010; Kimmerling et al. 2018). This technique can distinguish even closely related species (e.g. Baldwin et al. 2009; Tawa et al. 2014). These genetic methods enable any researcher with expertise in molecular biology to achieve results of similar precision. However, there are certain limitations in identifying fish eggs and larvae by genetic methods such as mitochondrial introgression, the lack of reference sequences in the database, and misidentification of reference data (Kawakami et al. 2010; Leis 2015). Therefore, it is preferable to identify fish eggs and larvae using a combination of morphological and genetic methods.
The larval fixation technique substantially influences the accuracy and reliability of the identification results. Previous studies successfully applied both morphological and genetic methods using larvae fixed in 70–99% ethanol (e.g. Tawa et al. 2014; Leis et al. 2015) because formalin damages DNA (Hykin et al. 2015). However, larval fixation in highly concentrated ethanol may cause more severe sample tissue shrinkage than 5–10% buffered formalin (Cunningham et al. 2000), resulting in specimens unsuitable for morphological observation. Previous studies demonstrated that by appropriately shortening the formalin fixation step, mitochondrial DNA (mtDNA) of sufficient quality can be obtained for genetic analysis (Díaz-Viloria et al. 2005; Chakraborty et al. 2006). However, short-term formalin fixation method has not been applied to identify fish larvae using a combination of morphological and genetic approaches.
For genetic identification of fish species, partial sequences of mtDNA, such as cytochrome c oxidase subunit 1 (COI), 12S ribosomal RNA gene (12S rRNA), 16S ribosomal RNA gene (16S rRNA), and cytochrome b gene (cyt b), have been used (Teletchea 2009). Among these DNA markers, a hypervariable region of 12S rRNA is a short fragment (less than 200 bp) containing sufficient interspecific differences to identify fishes at the species level except for some closely related congeners (Miya et al. 2015), and has been used for metabarcoding environmental DNA (e.g. Yamamoto et al. 2017). The region capable of species identification with a short fragment would be suitable for genetic identification of specimens in which the DNA may have been damaged due to formalin fixation. Moreover, a large amount of 12S rRNA sequence data for Japanese gobioid species is available through the DDBJ/EMBL/GenBank (12S rRNA: 293 species; COI: 270 species; cyt b: 177 species; 16S rRNA: 144 species, https://www.ncbi.nlm.nih.gov/nuccore).
The aim of the present study was to identify gobioid larvae based on a combination of morphological characters and a sequence of a hypervariable region of 12S rRNA. To enable identification using morphological and genetic methods, we applied short-term formalin fixation to the larvae. We also describe in detail the morphologies of larvae from two gobioid genera identified for the first time.
Materials and methods
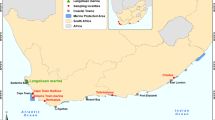
Specimens. Larval fish samples were collected with a light trap at Shinzato Fishing Port (26°42′08″N, 127°54′00″E) on the northern coast of Okinawa Island, southern Japan (see Oka and Miyamoto 2014). Sampling was conducted once or twice a month between April 2011 and October 2012. All samples were fixed in 10% seawater formalin immediately after collection. They were sorted and transferred into 70% ethanol within 24 hours. The larval specimens were preserved for 3–7 years without replacing the storage solution. Immature gobioid fish with scales and various cryptic Schindleria fish species (see Kon et al. 2007) were excluded from the present study.
Morphological identification. Gobioid larvae were distinguished on the basis of their morphological characters and identified to the lowest possible taxon according to the descriptions of Neira et al. (1998), Smith and Thacker 2000, Harada and Suharti (2000), Leis and Carson-Ewart (2004), Maeda and Tachihara (2005), Maeda (2008), Okiyama (2014), and Tran et al. (2018). Taxonomic nomenclature was according to Akihito et al. (2013). When a larva was identified only to the genus or family level, we used “sp.” after the genus or family name. In case of more than one species, they were distinguished with different letters (e.g. sp. A and sp. B). Terminology for morphological characters and measurements followed that of Leis and Carson-Ewart (2004). Standard lengths (SL) were measured using a digital microscope (Keyence, VHX-1000) or a vernier caliper under a stereomicroscope (Nikon, SMZ645). The following characters of the specimens described were measured based on photographs taken using digital camera (Olympus, E-M1): body depth at the pectoral-fin base (BD), head length (HL), snout length (SnL), eye diameter (ED), pre-dorsal-fin length, preanal length, caudal-peduncle length (horizontal distance from posterior end of the dorsal-fin base to caudal-fin base) and caudal-peduncle depth (least vertical distance of caudal peduncle). Dorsal-, anal-, pectoral-, and pelvic-fin rays were counted under the stereomicroscope. Measurements were made the distances along the body axis on the two dimensional images.
Genetic identification. DNA sequencing was performed on at least one individual per type distinguished by its morphological characters. Total DNA was extracted from the right eyeball of each specimen by the HotSHOT method (Truett et al. 2000). A partial sequence of mtDNA 12S rRNA region was amplified by polymerase chain reaction (PCR) in a thermal cycler (Gene Atlas 322, Astec, Fukuoka, Japan) with the primer pairs MiFishUF (5’-GTCGGTAAAACTCGTGCCAGC-3’) and MiFishUR (5’-CATAGTGGGGTATCTAATCCCAGTTTG-3’) (Miya et al., 2015). PCR was conducted with a KOD FX Neo DNA polymerase Kit (Toyobo, Osaka, Japan) in a 12.5-μl total volume: 6.25 μl of 2× PCR buffer, 0.5 μl of 10 μM of each primer, 2.5 μl of 2 mM dNTPs, 0.25 μl of KOD FX Neo, 0.25 μl of template DNA, and 2.25 μl dH2O. The PCR cycling conditions were as follows: 94 °C for 2 min, 10 cycles of denaturation at 98 °C for 10 s, annealing at 55 °C for 15 s, extension at 68 °C for 20 s, 25 cycles of denaturation at 98 °C for 10 s, annealing at 53 °C for 15 s, extension at 68 °C for 20 s, and a final extension at 68 °C for 2 min. Sequences were determined by Sanger Sequencing Service (FASMAC, Kanagawa, Japan). All sequences were deposited in DDBJ/EMBL/GenBank (Accession numbers LC515643–LC515665). Voucher specimens were deposited in the collection of the Okinawa Churashima Foundation Research Center (OCF).
The DNA sequences obtained for all representative larvae were compared with sequences deposited in DDBJ/EMBL/GenBank using BLAST (https://blast.ncbi.nlm.nih.gov/Blast.cgi) using the default parameters. Identification to the species level was considered as valid if more than 98.0% of the sequence corresponded to the sequence(s) of a single species. If more than 98.0% of sequence corresponded to those of more than two species, the larva was identified as belonging to a taxon that included all of these species.
Final identification. The final identification results for each larva were determined as follows. Case I: When the larva was identified to the same species or genus by the morphological and genetic methods, it was considered that there was no discrepancy, and the taxon at the lowest taxonomic level was used as the final identification; Case II: When no highly matched sequences (> 98.0%) were detected by BLAST, the taxon identified by the morphological method was used as the final identification; Case III: When the larva was identified to the family level using the morphological method and to the species level using the genetic method, the species identified using the genetic method was used as the final identification; Case IV: When there was any inconsistency between the morphological and genetic identifications, both candidates were inspected based on the larval morphology, sequencing data, and other available information. The most likely taxon was used as the final identification.
Results
Morphological identification. A total of 133 specimens of the postflexion gobioid larvae were distinguished into 21 types based on the morphological characters. Five types (Eleotris melanosoma, Eviota japonica, Pleurosicya bilobata, Redigobius bikolanus, and Ptereleotris microlepis) were identified to the species level, 14 to the genus level, and two to the family level (Table 1).
Genetic identification. DNA segments of 138–145 base pairs (bp) of the 12S rRNA gene were sequenced using DNA extracted from the larvae fixed in formalin for 1 day and preserved in ethanol for 3–7 years thereafter. The sequences of 13 types of the larvae were matched > 98.0% with single species (Table 1). The sequences of larvae OCF-P4150 and OCF-P4151 matched those of Oxyurichthys visayanus, but also presented with high similarity (> 98.0%) to that of Oxyurichthys sp. and Oxyurichthys cornutus (Table 1). Therefore, we genetically identified these larvae as Oxyurichthys sp. The sequence of larva OCF-P4145 showed high similarity with those of Trypauchenopsis intermedia (100%) and Trypauchenopsis sp. (99.3%) (Table 1). Thus, we genetically identified this larva as Trypauchenopsis sp. The sequences of larva OCF-P4144 matched with those of Bunaka gyrinoides (100%) and E. melanosoma (99.3%) (Table 1). Therefore, we genetically identified it as Eleotridae sp. No sequences were matched for the remaining five larval types (< 95.0% accuracy) (Table 1). The sequences of OCF-P4150 and OCF-P4159 were the same as those of OCF-P4151 and OCF-P4160, respectively, and were identified as the same respective types based on their morphological characters.
Final identification. Morphological and genetic methods identified 13 types to the species level and seven types to the genus level (Table 1; Figs. 1–3). Only one larval type (OCF-P4144) was inconsistently matched by both methods (Case IV). Bunaka gyrinoides and E. melanosoma were initially considered candidate species. The sequence of OCF-P4144 matched > 98.0% with those of B. gyrinoides (accession No. LC499417) and E. melanosoma (accession No. LC506283). Moreover, it also matched 97.2% with the sequence of E. melanosoma (accession No. AF265372). However, no other sequences for B. gyrinoides were found. The morphology of the larva OCF-P4144 was closely similar to that of E. melanosoma of Maeda and Tachihara (2005) but differed from that of B. gyrinoides of Okiyama (2014). Thus, we identified larva OCF-P4144 as E. melanosoma.
Photograph of preserved gobioid larval specimens collected on the northern coast of Okinawa Island. a Eleotris melanosoma (OCF-P4144; 10.8 mm SL), b Trypauchenopsis sp. (OCF-P4145; 8.6 mm SL), c Eviota japonica (OCF-P4146; 8.0 mm SL), d Pleurosicya bilobata (OCF-P4147; 7.4 mm SL), e Redigobius bikolanus (OCF-P4148; 5.1 mm SL), f Ptereleotris microlepis (OCF-P4149; 17.7 mm SL), g Oxyurichthys sp. (OCF-P4150; 13.0 mm SL), h Amblygobius phalaena (OCF-P4152; 9.6 mm SL), i Amblygobius nocturnus (OCF-P4153; 10.6 mm SL), j Favonigobius gymnauchen (OCF-P4154; 7.6 mm SL), k Favonigobius sp. B (OCF-P4155; 6.3 mm SL), l Valenciennea longipinnis (OCF-P4156; 16.6 mm SL), m Eviota abax (OCF-P4157; 8.5 mm SL), n Eviota distigma (OCF-P4158; 6.6 mm SL), o Eviota sp. C (OCF-P4159; 7.7 mm SL), p Eviota sp. D (OCF-P4161; 7.0 mm SL), q Eviota sp. E (OCF-P4162; 8.4 mm SL), r Gunnellichthys pleurotaenia (OCF-P4163; 22.6 mm SL), s Gobiidae sp. B (OCF-P4166; 12.0 mm SL)
Descriptions of larvae. The present study is the first to clarify the larval morphologies of two gobioid taxa. We describe the morphologies of Paragunnellichthys sp. and Ctenogobiops feroculus larvae in some detail, as these were the first described species in their respective genera. Range and mean of SL and sampling month for each larval type are listed in Electronic Supplementary Materials (ESM) Table S1.
Paragunnellichthys sp.
Material examined. OCF-P4164, postflexion stage, 14.6 mm SL, 20 October 2012.
Description. Body very elongate, depth 7.1% of SL. Head small, length 13.8% of SL. Eye small, diameter 20.2% of HL. Myomeres 22 + 26 = 48. Mouth large; posterior margin of the maxilla reaching vertically through middle of pupil; lower jaw protruding anteriorly. Dorsal-fin base single and long, length 80.5% of SL. Anal-fin base long, length 42.1% of SL. Pectoral fin not reaching to anus. Pelvic fin small, 15.2% of HL.
Body less pigmented; four melanophores along lateral midline on caudal peduncle; two melanophores along each dorsal and ventral midline on caudal peduncle; internal melanophores on dorsal part of gas bladder.
Remarks. Larva OCF-P4164 had a very elongate body and long dorsal- and anal-fin bases, consistent with the characteristics of the family Microdesmidae (Smith and Thacker 2000; Leis and Carson-Ewart 2004). Of Microdesmidae, Gunnellichthys and Paragunnellichthys are known from the Indo-Pacific (Leis and Carson-Ewart 2004). Although the number of fin rays of OCF-P4164 closely matched that of Paragunnellichthys seychellensis (see Dawson 1967), this species is not known from Okinawa Island. Paragunnellichthys sp. was recorded with a voucher specimen (HMNH-P6154) from Irabu Island (Yoshigou 2014) and with photographs from Okinawa and Iriomote Islands in southern Japan (http://fishpix.kahaku.go.jp/fishimage-e/index.html). Thus, OCF-P4164 was identified as a member of the genus Paragunnellichthys, but further study is needed to determine whether it is the same species as HMNH-P6154 and the other photographed specimens.
Among the five genera of the family Microdesmidae (Clarkichthys, Cerdale, Microdesmus, Gunnellichthys, and Paragunnellichthys), the postflexion larvae of four genera, excluding Clarkichthys, have been described in previous studies (Smith and Thacker 2000; Leis and Carson-Ewart 2004; Okiyama 2014). All were characterized by very elongate body, small pelvic fins, and long dorsal- and anal-fin bases. The postflexion larvae of Gunnellichthys and Paragunnellichthys were distinguished from those of Cerdale and Microdesmus by the lack of melanophores around pelvic-fin base and gut. The postflexion larvae of Paragunnellichthys were distinguished from those of Gunnellichthys by the lack of paired melanophores along each side of anal-fin base.
Ctenogobiops feroculus
Material examined. OCF-P4165, postflexion stage, 8.0 mm SL, 20 May 2011.
Description. Body elongate, depth 18.1% of SL. Head moderate, length 26.5% of SL. Eye small, diameter 23.5% of HL. Myomeres 7 + 17 = 24. Second dorsal- and anal-fin base moderate, length 27.7% and 28.5% of SL. Pelvic fins connected by membrane between last rays; lacking frenum (possibly undeveloped). Pectoral and pelvic fins not reaching to anus.
Body lightly pigmented; melanophores at angle of lower jaw, along isthmus, and on pelvic-fin base; unpaired melanophores along each side of anal-fin base; single row of melanophores along ventral midline of caudal peduncle; single row of faint melanophores along lower caudal-fin ray; internal melanophores around otic capsule, dorsal part of gas bladder, and above anus.
Remarks. The postflexion larvae of C. feroculus could be distinguished from previously reported gobioid larvae by the combination of fin-ray count, body size, and pigment pattern. The fin-ray count and body size of the postflexion larvae of C. feroculus were similar to those of the larvae of Oligolepis sp. of Okiyama (2014). However, C. feroculus differ from Oligolepis sp. in lacking internal pigmentation above posterior end of anal-fin base (vs. having conspicuous internal pigmentation above posterior end of anal-fin base), having single row of melanophores along ventral midline of caudal peduncle (vs. no melanophores along ventral midline of caudal peduncle), and having unpaired melanophores along anal-fin base (vs. paired melanophores along anal-fin base).
Discussion
We revealed the morphology of the postflexion larvae of Paragunnellichthys sp. (Microdesmidae) and C. feroculus (Gobiidae) for the first time in their respective genera. These postflexion larvae were in the developmental stage with poor pigmentation and almost complete fins, and they would be at the stage just before settlement. The microdesmid postflexion larvae at the settlement stage have been reported from both Indo-Pacific and Atlantic Oceans. The settlement size of Indo-Pacific Gunnellichthys larvae is 22.6–28.0 mm SL (Leis and Carson-Ewart 2004; present study), and the size of the larvae of the Atlantic microdesmid Cerdale and Microdesmus is 15–36 mm SL (Smith and Thacker 2000). The size of the postflexion larva of Paragunnellichthys sp. is 14.6 mm SL; thus this is similar to the smallest settlement size of Atlantic microdesmid species. The larval morphology of C. feroculus was the first to be described among the 12 genera (Karplus and Thompson 2011) of the shrimp-associated gobies. The size of the postflexion larvae of C. feroculus (8.0–8.4 mm SL) was included in the typical size at settlement of Gobiidae members, 6–9 mm SL (Victor et al. 2010).
Using a combination of morphological and genetic methods, we identified 13 of the 21 types (61.9%) of gobioid larvae to the species level. We were unable to fulfil our objective of identifying all larvae to the species level, which indicates that there are still certain limitations in identifying gobioid larvae. To enable the identification of all larvae to species level, more comprehensive larval descriptions and a large number of databases containing DNA reference sequences, which constitute the basis of species identification, are required. To date, many identification guidebooks of fish larvae have been published (Leis 2015) and some large-scale sequencing projects are being undertaken to identify fish (e.g. Ward et al. 2005; Miya et al. 2015). However, neither larval descriptions nor reference sequences suffice (Leis 2015; Steinke et al. 2017). Among the 518 species in 127 gobioid fish genera found in Japanese waters (Akihito et al. 2013), the early stages of 142 taxa in 71 genera (including fishes identified to the genus or family level) are described in an atlas of early stage fishes in Japan (Okiyama 2014). The number of reference sequences of 12S rRNA regions used in the present study was 293 gobioid species in 96 genera (https://www.ncbi.nlm.nih.gov/nuccore). The identification of gobioid larvae should dramatically improve as the numbers of larval descriptions and reference sequences increase. The information and data derived therefrom should help to elucidate larval ecology, which has been heretofore obscure as it has been difficult to identify fish species based solely on their morphological characters.
References
Akihito, Sakamoto K, Ikeda Y, Aizawa M (2013) Gobioidei. In: Nakabo T (ed) Fishes of Japan with pictorial keys to the species, 3rd edition. Tokai University Press, Hadano, pp 1347–1608, 2109–2211
Borges R, Faria C, Gil F, Gonçalves EJ (2011) Early development of gobies. In: Patzner R, Van Tassell JL, Kovacic M, Kapoor BG (eds) The biology of gobies. CRC Press, Boca Raton, USA, pp 403–464
Baldwin CC, Mounts JH, Smith DG, Weigt LA (2009) Genetic identification and color descriptions of early life-history stages of Belizean Phaetoptyx and Astrapogon (Teleostei: Apogonidae) with comments on identification of adult Phaetoptyx. Zootaxa:1–22
Chakraborty A, Sakai M, Iwatsuki Y (2006) Museum fish specimens and molecular taxonomy: A comparative study on DNA extraction protocols and preservation techniques. J Appl Ichthyol 22:160–166
Clarke TA (1991) Larvae of nearshore fishes in oceanic waters near Oahu, Hawaii. NOAA, Tech Report NMFS 101:1–19
Cunningham MK, Granberry WF Jr, Pope KL (2000) Shrinkage of inland silverside larvae preserved in ethanol and formalin. N Am J Fish Manage 20:816–818
Dawson CE (1967) Paragunnellichthys seychellensis, a new genus and species of gobioid fish (Microdesmidae) from the Western Indian Ocean. Proc Biol Soc Wash 80:73–82
Díaz-Viloria N, Sánchez-Velasco L, Pérez-Enríquez R (2005) Inhibition of DNA amplification in marine fish larvae preserved in formalin. J Plankton Res 27:787–792
Harada S, Suharti SR (2000) Gobiidae. In: Matsuura K, Sumadhiharga OK, Tsukamoto K (eds) Field guide to Lombok Island: identification guide to marine organisms in seagrass beds of Lombok Island, Indonesia. Ocean Research Institute, University of Tokyo, Tokyo, pp 352–369
Hykin SM, Bi K, McGuire JA (2015) Fixing formalin: a method to recover genomic-scale DNA sequence data from formalin-fixed museum specimens using high-throughput sequencing. PLoS One 10:e0141579
Isari S, Pearman JK, Casas L, Michell CT, Curdia J, Berumen ML, Irigoien X (2017) Exploring the larval fish community of the central Red Sea with an integrated morphological and molecular approach. PLoS One 12:e0182503
Karplus I, Thompson AR (2011) The partnership between gobiid fishes and burrowing alpheid shrimps. In: Patzner R, Van Tassell JL, Kovacic M, Kapoor BG (eds) The Biology of Gobies. CRC Press, Boca Raton, USA, pp 559–607
Kawakami T, Aoyama J, Tsukamoto K (2010) Morphology of pelagic fish eggs identified using mitochondrial DNA and their distribution in waters west of the Mariana Islands. Environ Biol Fishes 87:221–235
Kimmerling N, Zuqert O, Amitai G, Gurevich T, Armoza-Zvuloni R, Kolesnikov I, Berenshtein I, Melamed S, Gilad S, Benjamin S, Rivlin A, Ohavia M, Paris CB, Holzman R, Kiflawi M, Sorek R (2018) Quantitative, species-level ecology of reef fish larvae via metabarcoding. Nat Ecol Evol 2:306–316
Ko HL, Wang YT, Chiu TS, Lee MA, Leu MY, Chang KZ, Chen WY, Shao KT (2013) Evaluating the accuracy of morphological identification of larval fishes by applying DNA barcoding. PLoS One 8:e53451
Kon T, Yoshino T, Mukai T, Nishida M (2007) DNA sequences identify numerous cryptic species of the vertebrate: a lesson from the gobioid fish Schindleria. Mol Phylogenet Evol 44:53–62
Leis JM (2015) Taxonomy and systematics of larval Indo-Pacific fishes: a review of progress since 1981. Ichthyol Res 62:9–28
Leis JM, Carson-Ewart BM (eds) (2004) The larvae of Indo-Pacific coastal fishes: a guide to identification (Fauna Malesiana Handbook 2, 2nd edition). Brill, Leiden
Leis JM, Meyer O, Hay AC, Gaither MR (2015) A coral-reef fish with large, fast conspicuous larvae and small, cryptic adults. Copeia 103:78–86
Maeda K (2008) Dispersal strategy of gobioid larvae estimated by morphology and age at recruitment. PhD dissertation. University of the Ryukyus, Okinawa
Maeda K, Tachihara K (2005) Recruitment of amphidromous sleepers Eleotris acanthopoma, Eleotris melanosoma, and Eleotris fusca into the Teima River, Okinawa Island. Ichthyol Res 52:325–335
Maeda K, Tachihara K (2014) Larval fish fauna of a sandy beach and an estuary on Okinawa Island, focusing on larval habitat utilization by the suborder Gobioidei. Fish Sci 80:1215–1229
Marancik KE, Richardson DE, Lyczkowski-Shultz J, Konieczna M, Cowen RK (2010) Evaluation of morphological characters to identify grouper (Serranidae: Epinephelini) larvae in the Gulf of Mexico using genetically identified specimens. Bull Mar Sci 86:571–624
Miya M, Sato Y, Fukunaga T, Sado T, Poulsen JY, Sato K, Minamoto T, Yamamoto S, Yamanaka H, Araki H, Kondoh M, Iwasaki W (2015) MiFish, a set of universal PCR primers for metabarcoding environmental DNA from fishes: detection of more than 230 subtropical marine species. Roy Soc Open Sci 2, 150088
Neira FJ, Miskiewicz AG, Trnski T (eds) (1998) Larvae of temperate Australian fishes: laboratory guide for larval fish identification. University of Western Australia Press, Nedlands
Oka S, Miyamoto K (2014) Larvae and juvenile fishes collected by light-trap sampling at Shinzato Port, Okinawa Island, southern Japan. Fauna Ryukyuana 16:1–11
Okiyama M (ed) (2014) An atlas of early stage fishes in Japan second edition. Tokai University Press, Hadano
Smith DG, Thacker CE (2000) Late-stage larvae of the Family Microdesmidae (Teleostei, Gobioidei) from Belize, with notes on systematics and biogeography in the western Atlantic. Bull Mar Sci 67:997–1012
Steinke D, deWaard JR, Gomon MF, Johnson JW, Larson HK, Lucanus O, Moore GI, Reader S, Ward RD (2017) DNA barcoding the fishes of Lizard Island (Great Barrier Reef). Biodivers Data J 5:e12409
Tawa A, Aoyama J, Yoshimura T, Wouthuyzen S, Mochioka N (2014) Leptocephalus larvae of two moray eels (Anguilliformes; Muraenidae), Gymnothorax sagmacephalus and Gymnothorax albimarginatus, identified from morphometric and genetic evidence. Ichthyol Res 61:32–41
Teletchea F (2009) Molecular identification methods of fish species: reassessment and possible applications. Rev Fish Biol Fisher 19:265–293
Tran TT, Tran HD, Nguyen HX (2018) Larval description and habitat utilization of an amphidromous goby, Redigobius bikolanus (Gobiidae). Anim Biol 68:1–12
Truett GE, Heeger P, Mynatt RL, Walker JA, Warman ML (2000) Preparation of PCR-quality mouse genomic DNA with hot sodium hydroxide and tris (HotShot). BioTechniques 29:52–54
Victor BC, Hanner R, Shivji M, Hyde J (2009) Identification of the larval and juvenile stages of the Cubera Snapper, Lutjanus cyanopterus, using DNA barcoding. Zootaxa 2215:24–36
Victor BC, Vasquez-Yeomans L, Valdez-Moreno M, Wilk L, Jones DL, Lara MR, Caldow C, Shivji M (2010) The larval, juvenile, and adult stages of the Caribbean goby, Coryphopterus kuna (Teleostei: Gobiidae): a reef fish with a pelagic larval duration longer than the post-settlement lifespan. Zootaxa 2346:53–61
Ward RD, Zemlak TS, Innes BH, Last PR, Hebert PDN (2005) DNA barcoding Australia’s fish species. Phil Trans Royal Soc B: Biol Sci 360:1847–1857
Yamamoto S, Masuda R, Sato Y, Sado T, Araki H, Kondoh M, Miya M (2017) Environmental DNA metabarcoding reveals local fish communities in a species-rich coastal sea. Sci Rep 7:40368
Yoshigou H (2014) Annotated checklist and bibliographic records of inland water fishes of the Ryukyu Archipelago, Japan. Fauna Ryukyuana 9:1–153
Acknowledgments
We thank Takeshi Kon (University of the Ryukyus), Ken Maeda (Okinawa Institute of Science and Technology Graduate University), and Taiki Ishihara (Japan Fisheries Research and Education Agency) for providing information on DNA barcoding and gobioid fishes. This work was supported by the admission fees for the Okinawa Churaumi Aquarium.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article was registered in the Official Register of Zoological Nomenclature (ZooBank) as 3CCEE403-456B-473B-A4A8-061E086CB780.
This article was published as an Online First article on the online publication date shown on this page. The article should be cited by using the doi number.
Electronic supplementary material
Below is the link to the electronic supplementary material.
About this article
Cite this article
Hanahara, N., Miyamoto, K. & Oka, Si. Morphological and genetic identification of formalin-fixed gobioid larvae and description of postflexion larvae of Paragunnellichthys sp. and Ctenogobiops feroculus. Ichthyol Res 68, 182–190 (2021). https://doi.org/10.1007/s10228-020-00769-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10228-020-00769-z