Abstract
Purpose
This study aims to characterize antimicrobial resistance (AMR) of all the non-duplicated Acinetobacter baumannii strains isolated from an intensive care unit in a tertiary hospital during the period of January 1 to December 31, 2015.
Methods
A. baumannii (n = 95 strains) isolated from patients was subjected to antimicrobial susceptibility test (AST) by Vitek 2 Compact system to determine minimum inhibitory concentrations, followed by genotyping by enterobacterial repetitive intergenic consensus-PCR (ERIC-PCR). Resistance genes of interest were PCR amplified and sequenced.
Results
All isolates were qualified as MDR, with a resistance rate of > 80% to 8 antimicrobials tested. In terms of beta-lactamase detection, the blaOXA23, blaTEM-1, and armA genes were detected frequently at 92.63%, 9 1.58%, and 88.42%, respectively. The metallo-β-lactamase genes blaIMP and blaVIM were undetected. Aph (3’)-I was detected in 82 isolates (86.32%), making it the most prevalent aminoglycoside-modifying enzyme (AMEs) encoding gene. In addition, ant (3″)-I was detected at 30.53%, while 26.32% of the strains harbored an aac (6')-Ib gene. ERIC-PCR typing suggested moderate genetic diversity among the isolates, which might be organized into 10 distinct clusters, with cluster A (n = 86 isolates or 90.53%) being the dominant cluster.
Conclusions
All of the A. baumannii strains detected in the ICU were MDR clones exhibiting extremely high resistance to carbapenems and aminoglycosides as monitored throughout the study period. They principally belonged to a single cluster of isolates carrying blaOXA23 and armA co-producing different AMEs genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Acinetobacter baumannii (A. baumannii) is an opportunistic pathogen adept at colonizing and thriving in the hospital environment. In the recent decade, carbapenemase-producing multidrug-resistant (MDR) A. baumannii has emerged as a prominent cause of healthcare-associated infections (HAIs) notably at intensive care units (ICUs), and its incidence seems to be ascending alarmingly in parts of China (He et al. 2011; Li et al. 2018; Bitrian et al. 2012; Behdad et al. 2020). Patients undergoing invasive procedures, immunosuppressive therapy, or treatment with broad-spectrum antibiotics are vulnerable to HAIs caused by A. baumannii, particularly in the contexts of ventilator-associated pneumonia, bacteremia, septicemia, urinary tract, and wound infections (Bitrian et al. 2012; del Mar et al. 2005; Freire et al. 2016; Gomez-Arrebola et al. 2021).
By virtue of its extraordinary aptitude to survive in the hospital environment and to develop extremely high resistance to an array of common antibiotics including aminoglycoside and carbapenem classes of antibiotics, A. baumannii has become a major challenge to medical care at the ICU (Shimose et al. 2016; Molter et al. 2016; Shamsizadeh et al. 2017). One of the most prevalent sequence types (ST) of epidemic clones in China is ST208, which has gained notoriety for causing outbreaks in local ICUs (Bahador et al. 2015). Analysis on genomic relatedness among clinical isolates can help detect an epidemic strain, which can also offer information on infection diagnosis and anti-infection treatment.
Although substantial efforts have been made over the years in monitoring the epidemicity and AMR trends of A. baumannii in China, the scope of previous studies tends to be limited to highly populous urban centers in northern and eastern China (Ning et al. 2017; Zhou et al. 2018). In southern China including Guangdong province (population 108.5 million), where the humid subtropical climate indeed favors microbial growth, epidemiological surveys on A. baumannii in HAIs were only with moderate frequencies and again covered only very large urban centers such as Guangzhou (population 14.9 million) (Zhou et al. 2015; Li et al. 2013). In contrast, studies on HAIs by A. baumannii and underlying mechanisms of AMR are otherwise scant in other Chinese regions overlooked in epidemiological survey. In this regard, we undertook the current study to examine the AMR traits, molecular determinants of AMR, and clonal relationship of A. baumannii strains isolated from an ICU of a teaching tertiary hospital in the Chaoshan metropolitan area (13.93 million residents) in Guangdong province, southern China. We found that the isolates (n = 95) generally exhibited very high resistance to most of the commonly used clinical antibiotics including aminoglycosides and carbapenems, carried blaOXA23, blaTEM-1, and aph (3’)-I as the most frequently detected resistance genes, and consisted mostly of strains belonging to a single dominant cluster (cluster A in ERIC-PCR analysis).
Materials and methods
Research settings and bacterial isolates
This study was conducted at the ICU of a tertiary-level teaching hospital affiliated to the Shantou University Medical College (SUMC) in Shantou City in Guangdong, a populous province in southern China. The hospital (1816 inpatient beds) serves the Chaoshan metropolitan area in eastern Guangdong. A total of 95 non-duplicated A. baumannii isolates were systematically collected from patients’ samples during the period of January 1 to December 31, 2015. This study had been reviewed and approved by the Research Ethics Committee of the First Affiliated Hospital of Shantou University Medical College. The study was given a waiver of informed consent on the ground that it focuses only on characterizing bacterial isolates and involves no patient’s information.
Antimicrobial susceptibility
All isolates were first identified by using Vitek 2 Compact system (bioMérieux, France) and their antimicrobial susceptibility profiles obtained by using the Gram negative susceptibility cards (GN16 cards), according to the manufacturer’s instructions. Antimicrobial susceptibility test (AST) results for MICs (minimum inhibitory concentrations) were interpreted according to the criteria recommended by the Clinical and Laboratory Standards Institute (CLSI 2015). Confirmed A. baumannii isolates were stored at − 80 °C for subsequent experiments.
Detection of antimicrobial resistance genes
Whole genomic DNA was extracted by using TIANamp Bacteria DNA kit (Tiangen Biotech, China), according to the manufacturer’s instructions. Detection of antimicrobial resistance (AMR) genes by PCR amplification was carried out with specific primers (Lin et al. 2015; Chen et al. 2010) (see details in Table 1) to screen for the following genes of interest: extended-spectrum β-lactamases (ESBLs) encoding gene (blaTEM-1, blaSHV), metallo-β-lactamases encoding genes (blaIMP, blaVIM-2, blaNDM-1), OXA carbapenemases encoding genes (blaOXA23, blaOXA24, blaOXA58), aminoglycoside-modifying enzyme (AME) encoding genes (aac(6')-Ib, ant(3″)-I, aph(3’)-I), and 16 s rRNA methylase encoding gene (armA) were detected. For PCR amplification, the following thermal cycling conditions were adopted: initial denaturation at 94 °C for 3 min, followed by 30 cycles (94 °C for 1 min, 58–62 °C for 1 min, and 72 °C for 1 min), and a final extension step of 8 min at 72 °C. PCR products were separated by electrophoresis (at 100 V through a 1% agarose gel in 0.5 × TBE running buffer), stained with ethidium bromide, and observed under ultraviolet light. Identity of all PCR products was confirmed by DNA sequencing (Beijing Genomics Institute, BGI).
Genotyping of isolates
For determination of genetic relatedness of the isolates, enterobacterial repetitive intergenic consensus-PCR (ERIC-PCR) was performed with primer ERIC2 (5′-AAGTAAGTGACTGGGGTGAGCG-3′) (Bahador et al. 2015) to amplify the conserved sequences of bacterial strains, by using the following thermal cycling conditions: initial denaturation at 94 °C for 5 min, 4 cycles (94 °C for 1 min, 26 °C for 1 min, 72 °C for 1 min), then 40 cycles (94 °C for 30 s, 40 °C for 30 s, and 72 °C for 1 min), and extension at 72 °C for 5 min. To resolve the PCR products, each PCR product was analyzed by electrophoresis in a 1.5% agarose gel stained with ethidium bromide. Results for ERIC-PCR banding patterns were appraised by the software Quantity One (version 4.6.2) and scored as absent (0) or present (1) to construct a dendrogram according to the unweighted pair group (UPGMA) method, using the software NTSYS-pc (version 2.10e). Isolates with more than 90% similarity were considered as belonging to the same cluster.
Statistics analysis
Statistical analysis on antimicrobial susceptibility rates was analyzed by WHONET 5.6 software.
Results
Isolate characteristics and resistance rates
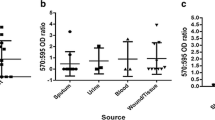
In this study, a total of 95 non-duplicative A. baumannii strains were isolated from ICU patients. Strains from male patients evidently outnumbered those from females at a ratio of 65 (68.42%) to 30 (31.58%). Affected patients had a mean age of 61.93 ± 1.87 years (range of 7 to 89 years old). The major isolation sites were sputum (n = 91), puncture fluid (n = 2), and stool (n = 2). As shown in Table 2, AST results suggested that all isolates could be qualified as multidrug-resistant (MDR) A. baumannii, which were highly resistant to 8 antibiotics including cefepime (FEP), ceftriaxone (CRO), imipenem (IPM), gentamycin (GEN), tobramycin (TOB), ciprofloxacin (CIP), trimethoprim/sulfamethoxazole (SXT), and piperacillin/tazobactam (TZP) while levofloxacin (LVX) might not be deemed any more efficient than the abovementioned agents against A. baumannii, as it had a notable rate of intermediate-level resistance (37.90%) as shown in Fig. 1.
Genotypic patterns in ERIC-PCR analysis
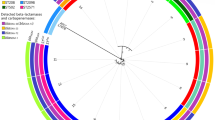
ERIC-PCR was used to compare the genetic relatedness among the A. baumannii isolates. All PCR banding patterns ranging from 550 to 2000 bp were analyzed by the NTSYS software to construct a dendrogram, as shown in Fig. 2. In general, 86 (or 90.53%) of the analyzed A. baumannii strains belonged to a major cluster A, while the remaining 9 isolates exhibited substantially different banding patterns, additionally designated as isolates B, C, D, E, F, G, H, J, and K. In a longitudinal analysis, strains belonging to cluster A were detectable throughout the study period in 2015, indicating that members of this cluster correspond to the major clone causing the epidemic of MDR A. baumannii at our hospital.
Determination of antimicrobial resistance genes
Analysis on AMR genes suggests that the A. baumannii isolates included in this study had high carriage rates for some specific AMR genes. Among the 95 strains, 87 (91.58%) were tested positive for the ESBL encoding gene blaTEM-1. In terms of detection of carbapenemase genes, 88 strains (92.63%) were found to harbor the blaOXA23 gene. Gene armA, a member of 16S rRNA methylases, was detected in 84 isolates (88.42%), while the most prevalent AME encoding gene aph (3’)-I was found in 82 isolates (86.32%). In comparison, 29 (30.53%) and 25 (26.32%) of the isolates harbored the ant (3″)-I and aac (6')-Ib genes, respectively. In further analysis, the genes blaSHV, blaIMP, blaVIM-2, blaNDM-1, blaOXA24, and blaOXA58 were undetected in any of the A. baumannii strains (Table 3).
Through genotyping and detection of resistance genes, we classified the 95 isolates of A. baumannii in this study, as shown in Table 3. Cluster A could be categorized into 9 subtypes on the basis of different AMR gene combinations. The most prevalent subtype in cluster A was subtype Ai, comprising 55 (63.95%) isolates expressing the genes blaOXA23, blaTEM-1, armA, and aph (3’)-I. Among subtype Ai isolates, 38 (44.19%) were non-susceptible to all of the antibiotics tested. Twenty-two (25.58%) isolates in group Aii harbored 6 different genes, non-susceptible to cefepime and ceftriaxone. The aminoglycoside resistance genes aph (3’)-I and ant (3″)-I were only detected in subtypes Aviii and Aix, but surprisingly they were susceptible to gentamycin and tobramycin. The rest of the isolates showed various combinations of AMR genes, presumably giving rise to different resistance patterns.
Discussion
ERIC-PCR, a genotyping method premised on amplification of conserved regions of genomic DNA, has the advantage of facile instrumentation and reliability comparable to pulsed field gel electrophoresis (PFGE). It has been proven useful for determining genomic relationship across strains with heterogeneous backgrounds (Cartelle Gestal et al. 2016; Ece et al. 2015). In the present study, a dendrogram based on ERIC-PCR results identifies 1 cluster (cluster A) and other 9 distinct isolates, suggesting that a single dominant clone of MDR A. baumannii prevailed in the ICU in 2015 (Jan. to Dec.). In terms of resistance phenotype, strains in cluster A were consistently more non-susceptible to all tested antibiotics than other strains. By using ERIC-PCR as a genotyping method, Ning and coworkers reported carbapenem-resistant clones of A. baumannii spreading at an ICU in western China (Ning et al. 2017). Chen and coworkers also described a major epidemic strain spreading at different hospital units in Hunan province of southern China (Chen et al. 2016). In our study, the spread of A. baumannii strains in the ICU lasted for a substantial period and their resistance rates to antibiotics were extremely high. We found that among the 9 subtypes of strains within cluster A, the most frequent type of AMR gene combination was blaOXA23 -blaTEM-1-aph (3’)-I-armA. Strains harboring this gene combination could be routinely isolated throughout the study period, suggesting the existence of entrenched extrinsic factors favoring their spread. Cross-transmission and contamination within the ward environment might underpin this process, which calls for greater awareness for monitoring and timely disinfection of the ward environment (Protano et al. 2019).
In our study, we found that multidrug-resistant Acinetobacter baumannii (MDRAB) strains simultaneously carrying the blaOXA23 gene and multiple aminoglycoside resistance genes are apparently spreading in southern China. The carriage of blaOXA23 carbapenemases in A. baumannii has been documented worldwide and blaOXA23 was one of the most prevalent carbapenemase genes detected in Chinese hospitals (Ruan et al. 2013; Shoja et al. 2017). While the prevalence of A. baumannii co-expressing aminoglycoside resistance genes and carbapenemase genes has been reported in eastern China (Wang et al. 2016), to the best of our knowledge, there have been no studies on the epidemicity of A. baumannii co-carrying AMR genes against aminoglycosides and carbapenems in southern China. It is noteworthy that the blaTEM-1 gene was the most prevalent ESBL gene in the present study, which differs from a previous study, where blaCTX-M was reported to be the predominant ESBL gene (Mahamat et al. 2016).
Aminoglycoside resistance of A. baumannii has been reported with increasing frequency in China in recent years (Gao et al. 2017; Jiang et al. 2014; Lin et al. 2015). In a study on A. baumannii from Jiangsu province, China, the most prevalent AMEs were identified as aac(3')-I and aac(6')-Ib (Wen et al. 2014). The resistance rates for GEN and TOB in this study were 89.47% and 86.32%, respectively (Table 2), and the most representative aminoglycoside resistance gene combination in the present study was armA-aph(3’)-I (58.95%). Interestingly, the isolate exhibited susceptibility to both GEN and TOB with only armA gene being detected. The most prevalent of the AMEs was aph(3’)-I (86.32%), followed by ant(3″)-I (30.53%), with 84 (88.42%) of the strains carrying armA. In addition, high levels of aminoglycoside resistance co-occurring with carbapenem resistance have been reported in epidemic clones of A. baumannii from western China (Lin et al. 2015). The imipenem resistance rates of A. baumannii were extremely high in China and numerous studies have raised concerns over the emergence and spread of imipenem-resistant A. baumannii in hospitals (Neves FC et al. 2016). Resistance rates for imipenem reported in different Chinese ranged from 58 to 100% (Jiang et al. 2016; Zong et al. 2008; Ji et al. 2014; Wu et al. 2015). Our current results suggested that efficacy of carbapenems as treatment for MDR-AB infections seemed to be fast diminishing, especially in ICU contexts. A growing body of literature documents blaOXA23 as a predominant carbapenemase genotype among epidemic clones in China (Chen et al. 2017; Thummeepak et al. 2016) and outbreaks caused by blaOXA23 producing A. baumannii paralleled those occurring worldwide (Neves et al. 2016; Hammoudi et al. 2015; Novovic et al. 2015; Koh et al. 2007; Martins et al. 2009). In this present study, we found that 88 of the A. baumannii strains (92.63%) harbored a blaOXA23 gene, suggestive of a level of prevalence seen in other parts of China (Ana Kovacic et al. 2017). Collectively, we proposed that the presence of blaOXA23 gene could be a cardinal molecular determinant of carbapenem resistance in our study.
Conclusion
In this study, we described the resistance traits and genetic relatedness of MDR A. baumannii strains with high resistance that prevailed at the ICU of a teaching tertiary hospital in the Chaoshan area of Guangdong province, a populous yet epidemiologically overlooked region in southern China. Those strains highly resistant to carbapenem and aminoglycoside including imipenem and gentamycin/tobramycin may be associated with the carriage of blaOXA23 and AME genes as determined in PCR assays. A single cluster A of epidemic clones seemed to dominate the spread of MDR A. baumannii in the ICU of our hospital. Surveillance work in this study represents a first step towards a better understanding of MDR A. baumannii as a causative agent in ICUs, which calls for greater attention to continued monitoring and rational use of antibiotics.
Availability of data and materials
Not applicable.
Code availability
Not applicable.
References
Bahador A, Raoofian R, Pourakbari B, Taheri M, Hashemizadeh Z, Hashemi FB (2015) Genotypic and antimicrobial susceptibility of carbapenem-resistant Acinetobacter baumannii: analysis of is aba elements and blaOXA23-like genes including a new variant. Front Microbiol 6:1249. https://doi.org/10.3389/fmicb.2015.01249
Behdad R, Pargol M, Mirzaie A, Karizi SZ, Noorbazargan H, Akbarzadeh I (2020) Efflux pump inhibitory activity of biologically synthesized silver nanoparticles against multidrug-resistant Acinetobacter baumannii clinical isolates. J Basic Microbiol 60:494–507. https://doi.org/10.1002/jobm.201900712
Bitrian M, Solari CM, González RH, Nudel CB (2012) Identification of virulence markers in clinically relevant strains of Acinetobacter genospecies. Int Microbiol 15:79–88. https://doi.org/10.2436/20.1501.01.161
Carmen Gomez-Arrebola, Cristina Solano, Iñigo Lasa (2021) Regulation of gene expression by non-phosphorylated response regulators. International microbiology published online: 13 May 2021. https://doi.org/10.1007/s10123-021-00180-2
Cartelle Gestal M, Zurita J, Gualpa G, Gonzalez C, Paz YMA (2016) Early detection and control of an Acinetobacter baumannii multi-resistant outbreak in a hospital in Quito. Ecuador J Infect Dev Ctries 10(12):1294–1298. https://doi.org/10.3855/jidc.7544
Chen D, Xie J, Chen H, Yang Y, Zhan Z, Liang L et al (2016) Infection in southern Chinese patients with systemic lupus erythematosus: spectrum, drug resistance, outcomes, and risk factors. J Rheumatol 43(9):1650–1656. https://doi.org/10.3899/jrheum.151523
Chen TL, Chang WC, Kuo SC, Lee YT, Chen CP, Siu LK, Cho WL, Fung CP et al (2010) Contribution of a plasmid-borne blaOXA-58 gene with its hybrid promoter provided by IS1006 and an ISAba3-like element to beta-lactam resistance in Acinetobacter genomic species 13TU. Antimicrob Agents Chemother 54(8):3107–3112. https://doi.org/10.1128/AAC.00128-10
Chen Y, Gao J, Zhang H, Ying C (2017) Spread of the blaOXA23-containing Tn2008 in carbapenem-resistant Acinetobacter baumannii isolates grouped in CC92 from China. Front Microbiol 8:163. https://doi.org/10.3389/fmicb.2017.00163
del Mar TM, Cartelle M, Pertega S, Beceiro A, Llinares P, Canle D et al (2005) Hospital outbreak caused by a carbapenem-resistant strain of Acinetobacter baumannii: patient prognosis and risk-factors for colonisation and infection. Clin Microbiol Infect 11(7):540–546. https://doi.org/10.1111/j.1469-0691.2005.01184.x
Ece G, Erac B, Yurday Cetin H, Ece C, Baysak A (2015) Antimicrobial susceptibility and clonal relation between Acinetobacter baumannii strains at a tertiary care center in Turkey. Jundishapur J Microbiol 8(2):e15612. https://doi.org/10.5812/jjm.15612
Freire MP, de Oliveira GD, Garcia CP, Campagnari Bueno MF, Camargo CH, Kono Magri ASG et al (2016) Bloodstream infection caused by extensively drug-resistant Acinetobacter baumannii in cancer patients: high mortality associated with delayed treatment rather than with the degree of neutropenia. Clin Microbiol Infect 22(4):352–358. https://doi.org/10.1016/j.cmi.2015.12.010
Gao L, Lyu Y, Li Y (2017) Trends in drug resistance of Acinetobacter baumannii over a 10-year period: nationwide data from the China surveillance of antimicrobial resistance program. Chin Med J 130(6):659–664. https://doi.org/10.4103/0366-6999.201601
Hammoudi D, Moubareck CA, Hakime N, Houmani M, Barakat A, Najjar Z et al (2015) Spread of imipenem-resistant Acinetobacter baumannii co-expressing OXA-23 and GES-11 carbapenemases in Lebanon. Int J Infect Dis 36:56–61. https://doi.org/10.1016/j.ijid.2015.05.015
He C, Xie Y, Fan H, Kang M, Tao C, Zhang R et al (2011) Spread of imipenem-resistant Acinetobacter baumannii of European clone II in western China. Int J Antimicrob Agents 38(3):257–260. https://doi.org/10.1016/j.ijantimicag.2011.04.015
Jiang M, Liu L, Ma Y, Zhang Z, Li N, Zhang F et al (2016) Molecular epidemiology of multi-drug resistant Acinetobacter baumannii isolated in Shandong. China Front Microbiol 7:1687. https://doi.org/10.3389/fmicb.2016.01687
Jiang M, Zhang Z, Zhao S (2014) Epidemiological characteristics and drug resistance analysis of multidrug-resistant Acinetobacter baumannii in a China hospital at a certain time. Pol J Microbiol 63(3):275–281
Ji S, Chen Y, Ruan Z, Fu Y, Ji J, Fu Y et al (2014) Prevalence of carbapenem-hydrolyzing class D beta-lactamase genes in Acinetobacter spp isolates in China. Eur J Clin Microbiol Infect Dis 33(6):989–997. https://doi.org/10.1007/s10096-013-2037-z
Koh TH, Sng LH, Wang GC, Hsu LY, Zhao Y (2007) IMP-4 and OXA beta-lactamases in Acinetobacter baumannii from Singapore. J Antimicrob Chemother 59(4):627–632. https://doi.org/10.1093/jac/dkl544
Kovacic A, Dekic MSMS, Tonkic M, Novak A, Rubic Z et al (2017) Transmission and survival of carbapenem resistant Acinetobacter baumannii outside hospital setting. Int Microbiol 20(4):165–169. https://doi.org/10.2436/20.1501.01.299
Lin T, Tang CG, Li QH, Ji J, Ge HY, Zhang XY et al (2015) Identification of aac(2’)-I type b aminoglycoside-modifying enzyme genes in resistant Acinetobacter baumannii. Genet Mol Res 14(1):1828–1835. https://doi.org/10.4238/2015.March.13.11
Li Y, Cao X, Ge H, Jiang Y, Zhou H, Zheng W (2018) Targeted surveillance of nosocomial infection in intensive care units of 176 hospitals in Jiangsu province. China the Journal of Hospital Infection 99(1):36–41. https://doi.org/10.1016/j.jhin.2017.10.009
Li Y, Pan C, Zhao Z, Zhao Z, Chen H, Lu W (2013) Effects of a combination of amlodipine and imipenem on 42 clinical isolates of Acinetobacter baumannii obtained from a teaching hospital in Guangzhou. China BMC Infect Dis 13:548. https://doi.org/10.1186/1471-2334-13-548
Mahamat A, Bertrand X, Moreau B, Hommel D, Couppie P, Simonnet C et al (2016) Clinical epidemiology and resistance mechanisms of carbapenem-resistant Acinetobacter baumannii, French Guiana, 2008–2014. Int J Antimicrob Agents 48(1):51–55. https://doi.org/10.1016/j.ijantimicag.2016.03.006
Martins AF, Kuchenbecker R, Sukiennik T, Boff R, Reiter KC, Lutz L et al (2009) Carbapenem-resistant Acinetobacter baumannii producing the OXA-23 enzyme: dissemination in Southern Brazil. Infection 37(5):474–476. https://doi.org/10.1007/s15010-009-9003-9
Molter G, Seifert H, Mandraka F, Kasper G, Weidmann B, Hornei B et al (2016) Outbreak of carbapenem-resistant Acinetobacter baumannii in the intensive care unit: a multi-level strategic management approach. J Hosp Infect 92(2):194–198. https://doi.org/10.1016/j.jhin.2015.11.007
Neves FC, Clemente WT, Lincopan N, Paiao ID, Neves PR, Romanelli RM et al (2016) Clinical and microbiological characteristics of OXA-23- and OXA-143-producing Acinetobacter baumannii in ICU patients at a teaching hospital. Brazil Braz J Infect Dis 20(6):556–563. https://doi.org/10.1016/j.bjid.2016.08.004
Ning NZ, Liu X, Bao CM, Chen SM, Cui EB, Zhang JL et al (2017) Molecular epidemiology of blaOXA23-producing carbapenem-resistant Acinetobacter baumannii in a single institution over a 65-month period in North China. BMC Infect Dis 17(1):14. https://doi.org/10.1186/s12879-016-2110-1
Novovic K, Mihajlovic S, Vasiljevic Z, Filipic B, Begovic J, Jovcic B (2015) Carbapenem-resistant Acinetobacter baumannii from Serbia: revision of CarO classification. PLoS ONE 10(3):e0122793. https://doi.org/10.1371/journal.pone.0122793
Protano C, Cammalleri V, Romano Spica V, Valeriani F, Vitali M (2019) Hospital Environment as a Reservoir for Cross Transmission: Cleaning and Disinfection Procedures. Ann Ig 31(5):436–448. https://doi.org/10.7416/ai.2019.2305
Thummeepak R, Kongthai P, Leungtongkam U, Sitthisak S (2016) Distribution of virulence genes involved in biofilm formation in multi-drug resistant Acinetobacter baumannii clinical isolates. Int Microbiol 19:121–129. https://doi.org/10.2436/20.1501.01.270
Ruan Z, Chen Y, Jiang Y, Zhou H, Zhou Z, Fu Y et al (2013) Wide distribution of CC92 carbapenem-resistant and OXA-23-producing Acinetobacter baumannii in multiple provinces of China. Int J Antimicrob Agents 42(4):322–328. https://doi.org/10.1016/j.ijantimicag.2013.06.019
Shamsizadeh Z, Nikaeen M, Nasr Esfahani B, Mirhoseini SH, Hatamzadeh M, Hassanzadeh A (2017) Detection of antibiotic resistant Acinetobacter baumannii in various hospital environments: potential sources for transmission of Acinetobacter infections. Environ Health Prev Med 22(1):44. https://doi.org/10.1186/s12199-017-0653-4
Shimose LA, Masuda E, Sfeir M, Berbel Caban A, Bueno MX, dePascale D et al (2016) Carbapenem-resistant Acinetobacter baumannii: concomitant contamination of air and environmental surfaces. Infect Control Hosp Epidemiol 37(7):777–781. https://doi.org/10.1017/ice.2016.69
Shoja S, Moosavian M, Rostami S, Farahani A, Peymani A, Ahmadi K et al (2017) Dissemination of carbapenem-resistant Acinetobacter baumannii in patients with burn injuries. J Chin Med Assoc 80(4):245–252. https://doi.org/10.1016/j.jcma.2016.10.013
Wang Y, Shen M, Yang J, Dai M, Chang Y, Zhang C et al (2016) Prevalence of carbapenemases among high-level aminoglycoside-resistant Acinetobacter baumannii isolates in a university hospital in China. Exp Ther Med 12(6):3642–3652. https://doi.org/10.3892/etm.2016.3828
Wen JT, Zhou Y, Yang L, Xu Y (2014) Multidrug-resistant genes of aminoglycoside-modifying enzymes and 16S rRNA methylases in Acinetobacter baumannii strains. Genet Mol Res 13(2):3842–3849. https://doi.org/10.4238/2014.May.16.9
Wu W, He Y, Lu J, Lu Y, Wu J, Liu Y (2015) Transition of blaOXA-58-like to blaOXA-23-like in Acinetobacter baumannii clinical isolates in southern China: an 8-year study. PLoS ONE 10(9):e0137174. https://doi.org/10.1371/journal.pone.0137174
Zhou K, Tang X, Wang L, Guo Z, Xiao S, Wang Q et al (2018) An emerging clone (ST457) of Acinetobacter baumannii clonal complex 92 with enhanced virulence and increasing endemicity in South China. Clinical infectious diseases 67(suppl_2):S179-s188. https://doi.org/10.1093/cid/ciy691
Zhou Y, Wu X, Zhang X, Hu Y, Yang X, Yang Z et al (2015) Genetic characterization of ST195 and ST365 carbapenem-resistant Acinetobacter baumannii harboring blaOXA-23 in Guangzhou. China Microb Drug Resist 21(4):386–390. https://doi.org/10.1089/mdr.2014.0183
Zong Z, Lu X, Valenzuela JK, Partridge SR, Iredell J (2008) An outbreak of carbapenem-resistant Acinetobacter baumannii producing OXA-23 carbapenemase in western China. Int J Antimicrob Agents 31(1):50–54. https://doi.org/10.1016/j.ijantimicag.2007.08.019
Acknowledgements
Nai-Kei Wong modified the manuscript.
Funding
This work was supported by 2020 Li Ka Shing Foundation Cross-Disciplinary Research Grant (2020LKSFG06A), a grant from Medical Science and Technology Foundation of Guangdong Province (B2018168), China, and Scientific Research Project of Hunan Provincial Department of Education (18C1145).
Author information
Authors and Affiliations
Contributions
Yuan-Chun Huang designed the experiments, carried out the study, interpreted the data, and reviewed the manuscript. Zhuo-Ran Chen and Hui-Wu Guo designed and performed the experiments and wrote the paper. Jun Liu, Mao-Zhang Fu, Ying-Kun Qiu, and Qing Pan collected and analyzed the data. All the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Hui-Wu Guo is the co-first author.
Rights and permissions
About this article
Cite this article
Chen, ZR., Guo, HW., Liu, J. et al. Resistance traits and molecular characterization of multidrug-resistant Acinetobacter baumannii isolates from an intensive care unit of a tertiary hospital in Guangdong, southern China. Int Microbiol 25, 471–479 (2022). https://doi.org/10.1007/s10123-022-00233-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10123-022-00233-0