Abstract
Background
Accurate prediction of pathologic results for early gastric cancer (EGC) based on endoscopic findings is essential in deciding between endoscopic and surgical resection. This study aimed to develop an artificial intelligence (AI) model to assess comprehensive pathologic characteristics of EGC using white-light endoscopic images and videos.
Methods
To train the model, we retrospectively collected 4,336 images and prospectively included 153 videos from patients with EGC who underwent endoscopic or surgical resection. The performance of the model was tested and compared to that of 16 endoscopists (nine experts and seven novices) using a mutually exclusive set of 260 images and 10 videos. Finally, we conducted external validation using 436 images and 89 videos from another institution.
Results
After training, the model achieved predictive accuracies of 89.7% for undifferentiated histology, 88.0% for submucosal invasion, 87.9% for lymphovascular invasion (LVI), and 92.7% for lymph node metastasis (LNM), using endoscopic videos. The area under the curve values of the model were 0.992 for undifferentiated histology, 0.902 for submucosal invasion, 0.706 for LVI, and 0.680 for LNM in the test. In addition, the model showed significantly higher accuracy than the experts in predicting undifferentiated histology (92.7% vs. 71.6%), submucosal invasion (87.3% vs. 72.6%), and LNM (87.7% vs. 72.3%). The external validation showed accuracies of 75.6% and 71.9% for undifferentiated histology and submucosal invasion, respectively.
Conclusions
AI may assist endoscopists with high predictive performance for differentiation status and invasion depth of EGC. Further research is needed to improve the detection of LVI and LNM.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Gastric cancer is the fifth most common malignancy and the fourth leading cause of cancer-related death worldwide [1]. Although radical surgery was traditionally the only curative treatment for gastric cancer, recent advances in endoscopic resection have demonstrated favorable clinical outcomes in early gastric cancer (EGC), concurrently improving quality of life for patients by preserving the stomach [2].
Endoscopic submucosal dissection (ESD) is considered curative for EGC without lymph node metastasis (LNM). Owing to the lack of reliable imaging methods to precisely detect LNM in EGC [3, 4], current guidelines recommend curative criteria for ESD based on pathologic features in resected specimens associated with a minimal risk of LNM [5, 6]. These factors include the differentiation status, invasion depth, and lymphovascular invasion (LVI) of the tumor. Since these characteristics are confirmed postoperatively, the accurate prediction of pathologic outcomes before treatment is essential to select the optimal curative approach between endoscopic and surgical resection.
Endoscopists perform forceps biopsies with assistance of magnifying endoscopy with narrow-band imaging (ME-NBI) to evaluate differentiation status, and endoscopic ultrasonography (EUS) to detect submucosal invasion before deciding the treatment strategy for EGC. However, previous studies have revealed significant histologic discrepancies between biopsies and resected specimens, potentially leading to non-curative ESD or missed opportunities for ESD in surgical cases [7,8,9]. In addition, EUS is not superior to conventional endoscopy in determining the invasion depth of EGC, with an accuracy of approximately 70% [10,11,12]. Therefore, a detailed assessment of endoscopic features by physicians is essential for predicting pathologic results in EGC.
With advancements in deep learning methods, recent studies have proposed artificial intelligence (AI) models for detecting and characterizing EGC in endoscopic images, aiming to assist physicians in evaluating endoscopic features [13]. This includes our previous study, where we developed an AI model which can detect EGC in endoscopic videos [14]. Although several models have been developed to assess the invasion depth of EGC using endoscopic images, there remains a need for further research into AI-assisted pathologic prediction for EGC to enhance the performance in video analysis [15, 16]. Moreover, to the best of our knowledge, no previous study has explored the capability of AI in predicting LVI or LNM based on endoscopic images or videos.
Therefore, this study aimed to develop and evaluate an AI model that comprehensively predicts the postoperative pathologic results of EGC, including the differentiation status, invasion depth, LVI, and LNM, based on preoperative white-light endoscopic images and videos.
Methods
The AI model developed in this study is an extension of the ENAD CAD-G, a convolutional neural network (CNN)-based model for detecting and classifying gastric lesions in endoscopic videos, as demonstrated in our previous study [14].
Study design and datasets
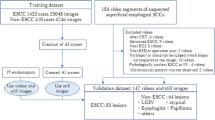
Figure 1 shows an overview of the study design and datasets. The total dataset of endoscopic images and videos was divided into an internal dataset used for training, internal validation, and testing and an external dataset employed for the external validation of the AI model.
For the internal dataset, we retrospectively collected 4,596 preoperative white-light endoscopic images of EGC from patients who underwent ESD or radical surgery between January 2018 and December 2022 at Seoul National University Hospital (SNUH), a tertiary hospital in the Republic of Korea. To assess the performance of the AI model in videos, we prospectively included 163 white-light endoscopic videos of patients referred from community clinics who underwent ESD or surgical resection for EGC between April 2022 and April 2023. For the external dataset, we used 436 images retrospectively collected from patients who underwent surgery for EGC between January 2020 and June 2020, and 89 videos prospectively collected from patients who underwent ESD for EGC between April 2022 and October 2022 at another tertiary hospital, Seoul National University Bundang Hospital (SNUBH), Republic of Korea.
This study was conducted in accordance with the Declaration of Helsinki and was approved by the ethics committees of the participating hospitals (IRB No. 2109–048-1253 at SNUH and IRB No. 2201–735-405 at SNUBH). Written informed consent was obtained from all prospectively enrolled patients who provided endoscopic videos. The requirement for informed consent was waived for the patients whose retrospective images were included in this study.
Preparation of endoscopic images
Supplementary Figure S1 shows the process of preparing endoscopic images before training the model. We retrospectively investigated the medical records of 1,617 patients who underwent ESD and 1,641 patients who underwent radical surgery for EGC at SNUH between 2018 and 2022. All preoperative white-light endoscopic images of the patients were reviewed by five endoscopists from SNUH, who selected images that best characterized the target lesions (two or three images per lesion) and excluded images with low resolution or blurring. Patients who underwent additional surgery after non-curative ESD were considered as surgical patients. Patients who had received any previous endoscopic treatment for the target lesions before ESD or had undergone ESD at another hospital before surgery were excluded. Patients with inconclusive pathologic results or those who did not undergo preoperative endoscopies at SNUH were also excluded. Finally, 2,453 images from 1,031 (51.4% of total patients) patients who underwent ESD and 2,143 images from 975 (48.6% of total patients) patients who underwent surgery were included in the internal dataset.
Patient enrollment of endoscopic videos
We prospectively enrolled patients who were diagnosed with gastric dysplasia or EGC on initial biopsies, underwent ESD or radical surgery, and were confirmed with EGC based on the pathological reports of the resected specimens. The indication for ESD was one of the following conditions: i) differentiated-type EGC with tumor size ≤ 2 cm and endoscopically suspected mucosal cancer without ulceration or ii) high-grade dysplasia [17]. We excluded patients who had previously undergone gastrectomy and those with contraindications for biopsy due to bleeding tendency or anticoagulant use. All endoscopic examinations were performed preoperatively using standard video endoscopes (GIF-Q260, GIF-H260, or GIF-H290; Olympus Medical Systems, Tokyo, Japan). Consequently, 163 (50 ESD patients and 113 surgery patients) patients from SNUH and 89 ESD patients from SNUBH were included in the study, and their endoscopic videos were provided. All ESD procedures were performed by experienced endoscopists following a standardized protocol [18]. The surgical procedures were based on standard gastrectomy with D1 + or D2 lymph node dissection [5].
Pathologic definitions
The pathologic characteristics of EGC in the images and videos were obtained from the pathological reports of specimens resected by ESD or surgery based on the 2022 Korean gastric cancer treatment guidelines [5]. Expert pathologists assessed all resected specimens. Differentiated-type EGC includes papillary, well, or moderately differentiated tubular adenocarcinoma, whereas undifferentiated-type EGC includes poorly differentiated adenocarcinoma, signet ring cell carcinoma, and mucinous carcinoma. In cases of mixed-type gastric cancer, the classification was determined by the histological type of the predominant lesion [19]. Submucosal invasion < 500 µm was defined as SM1 and submucosal invasion ≥ 500 µm was defined as SM2. The status of LNM in the resected specimens was also investigated in the surgical cases.
Distribution of target lesions in datasets
Table 1 summarizes the pathologic characteristics of EGC in the datasets. The training set comprised 19.6% (850/4,336) of images and 31.3% (48/153) of videos for undifferentiated histology, 23.7% (1,027/4,336) of images and 27.5% (42/153) of videos for submucosal invasion, 7.5% (327/4,332) of images and 6.5% (10/153) of videos for LVI, and 9.5% (187/1,962) of images and 5.7% (6/105) of videos for LNM.
The test set comprised 43.5% (113/260) of images and 50% (5/10) of videos for undifferentiated histology, 52.3% (136/260) of images and 40% (4/10) of videos for submucosal invasion, 19.2% (50/260) of images and 10% (1/10) of videos for LVI, and 15.9% (32/201) of images and 37.5% (3/8) of videos for LNM. In addition, the external dataset included 53.0% (231/436) of images and 1.1% (1/89) of videos for undifferentiated histology, 43.1% (188/436) of images and 12.4% (11/89) of videos for submucosal invasion, 11.5% (50/436) of images and 4.5% (4/89) of videos for LVI, and 13.1% (57/436) of images for LNM.
Training and internal validation of the AI model
The training set comprised 4,336 images and 153 videos, which were used to train and internally validate the AI model (Fig. 1). For the internal validation, the images and videos in the dataset were randomly divided into five subsets. Four subsets were used for training, and the remaining subset, was used for validation to calculate the predictive performance of the trained model. This cross-validation process was conducted five times to ensure comprehensive evaluation of all images and videos in the set.
The AI model was based on CNN architecture and utilized Efficientnetb0 to evaluate the pathologic characteristics of target lesions in endoscopic images [20]. The model employed a soft voting method to categorize these lesions into distinct predictive classes: differentiation status (differentiated or undifferentiated), invasion depth (mucosal or submucosal), LVI (positive or negative), and LNM (positive or negative). A generative model using Stylegan2 was integrated to enhance predictive performance of the model and increase its sensitivity [21]. Representative images analyzed by the model are presented in Fig. 2.
Representative examples of pathologic predictions by the AI model in endoscopic images. Each endoscopic image contains one lesion of EGC with the following pathologic characteristics. a Differentiated-type EGC of mucosal invasion without both LVI and LNM. b Differentiated-type EGC of submucosal invasion with positive LVI and negative LNM. c Undifferentiated-type EGC of mucosal invasion without both LVI and LNM. d Undifferentiated-type EGC of submucosal invasion with both LVI and LNM. The provided confidence levels represent the probabilistic evaluation of the AI model for predicting postoperative pathologic results of EGC in the images. AI artificial intelligence, EGC early gastric cancer, LVI lymphovascular invasion, LNM lymph node metastasis
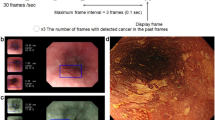
Figure 3 shows a schematic diagram of the evaluation of the endoscopic videos. Initially, gastric lesions within the videos were recognized and outlined with boundaries (cropped), using a lesion detection model based on YOLOv5 developed in our previous study [14]. Subsequently, the cropped images were categorized as cancer, adenoma, or non-neoplastic lesions using a lesion classification model that employs EfficientNETB0 [14]. The AI model then calculated the confidence levels for pathologic predictions of the identified cancers. Finally, the model utilized a soft voting method to determine the pathologic classifications of the cancers. Therefore, the cut-off value for considering AI prediction as correct was set at 50%, and this method was utilized in all the validation process.
Schematic diagram for AI-based pathologic prediction in endoscopic videos. Initially, the gastric lesion is identified and outlined with red boundaries (cropped) with the lesion detection model. Subsequently, the cropped lesion is categorized as either cancer, adenoma, or non-neoplastic lesion by the lesion classification model. For lesions classified as cancer, the model computes confidence levels to predict differentiation status, invasion depth, lymphovascular invasion, and lymph node metastasis. Finally, the lesion is categorized into distinct pathologic classes utilizing a soft voting method. AI artificial intelligence, EGC early gastric cancer
Testing the performance of the AI model and endoscopists
A test set was designed to evaluate and compare the predictive performances of the AI model and endoscopists, using 260 endoscopic images and 10 videos distinct from the training set. Sixteen endoscopists, comprising nine experts and seven novices, participated in the test and predicted the differentiation status, invasion depth, LVI, and LNM of the target lesions in the test set. The test was performed as a questionnaire comprising 100 questions, each including two or three images of one target lesion, and additional 10 questions with video clips, each containing one target lesion, as shown in Supplementary Figure S2. Each novice endoscopist had 1 year of experience in upper endoscopy prior to this study, conducted at least 500 procedures independently, and diagnosed EGC in clinical practice. The experts comprised endoscopists from six tertiary hospitals in the Republic of Korea who had over 10 years of experience performing gastric ESD before this study and held positions of associate professor or higher.
Statistical analyses
The gold standard for prediction was derived from postoperative pathological reports of specimens obtained from ESD or surgery. Accuracy, sensitivity, specificity, and positive and negative predictive values of the predictions were calculated. The prediction metrics were presented as means with 95% confidence intervals and were compared using the Mann–Whitney U test. Receiver operating characteristic (ROC) curves and the corresponding area under the ROC curve (AUC) values for the AI model were calculated. The accuracies of the AI model and those of all the experts were compared using the McNemar’s test. The accuracies of the experts and novices were compared using the Mann–Whitney U test. All statistical tests were two-sided, and P < 0.05 were considered significant. Statistical analyses were performed using the R software 4.2.3. for Windows.
Results
Baseline characteristics of patients
Detailed clinical information of enrolled patients and pathologic characteristics of lesions in the datasets are presented in Supplementary Table S1. Baseline characteristics were comparable across the datasets, including mean age (64.5 years vs. 63.6 years vs. 63.5 years), male sex (68.7% vs. 59.1% vs. 69.0%), and mean tumor size (21.0 mm vs. 24.3 mm vs. 22.7 mm).
Internal validation of the AI model with images and videos
Table 2 presents the predictive metrics (per image or frame) of the AI model after training with images and videos. Based on images, the model achieved mean predictive accuracies of 91.9% (sensitivity, 85.3%; specificity, 94.1%) for undifferentiated histology, 88.4% (sensitivity, 82.4%; specificity: 90.2%) for submucosal invasion, 84.7% (sensitivity, 24.2%; specificity, 96.2%) for LVI, and 86.8% (sensitivity, 27.4%; specificity, 94.0%) for LNM.
For videos, the model demonstrated mean predictive accuracies of 89.7% (sensitivity, 83.4%; specificity, 92.1%) for undifferentiated histology, 88.0% (sensitivity, 80.4%; specificity, 91.1%) for submucosal invasion, 87.9% (sensitivity, 20.0%; specificity, 97.0%) for LVI, and 92.7% (sensitivity, 16.7%; specificity, 96.5%) for LNM.
There was no significant difference in the performance of the AI model between images and videos, except for the accuracy of predicting LNM, which was significantly higher for the videos (P = 0.008).
Performance of the AI model according to pathologic characteristics of EGC
Supplementary Table S2 shows the performance of the AI model for endoscopic images according to the differentiation status of the target lesion within the training set. For differentiated-type EGC, the model exhibited mean accuracies of 91.1% (sensitivity, 82.3%; specificity, 89.2%) for submucosal invasion, 83.5% (sensitivity, 27.3%; specificity, 95.4%) for LVI, and 90.1% (sensitivity, 29.7%; specificity, 96.3%) for LNM. For undifferentiated-type EGC, the model demonstrated mean accuracies of 87.5% (sensitivity, 79.3%; specificity, 93.3%) for submucosal invasion, 88.1% (sensitivity, 10.4%; specificity, 98.6%) for LVI, and 83.8% (sensitivity, 26.9%; specificity, 92.6%) for LNM. The model presented a significantly higher accuracy in differentiated-type EGC than in undifferentiated-type EGC for predicting submucosal invasion (P = 0.008) and LNM (P = 0.016).
Supplementary Table S3 shows the performance of the AI model for predicting LVI and LNM based on the invasion depth of EGC in the training set. For mucosal cancer, the model exhibited mean accuracies of 89.6% (sensitivity, 3.0%; specificity, 98.6%) for LVI and 96.1% (sensitivity, 9.4%; specificity, 97.4%) for LNM. For submucosal cancer, the model demonstrated mean accuracies of 63.0% (sensitivity, 35.9%; specificity, 83.3%) for LVI and 72.5% (sensitivity, 38.7%; specificity, 81.4%) for LNM. For submucosal cancer with SM2 invasion, the model presented mean accuracies of 64.7% (sensitivity, 41.4%; specificity, 81.5%) for LVI and 73.3% (sensitivity, 39.9%; specificity, 82.4%) for LNM.
Comparison of the predictive accuracies between AI model and endoscopists
Figure 4 shows the ROC curves demonstrating the performance of the AI model with the performance of the endoscopists, presented as dots (blue = expert, red = novice) in the test set. The AUC values of the model were 0.992 for undifferentiated histology, 0.902 for submucosal invasion, 0.706 for LVI, and 0.680 for LNM. All dots representing the performance of the endoscopists were positioned below the curves for predicting undifferentiated histology, submucosal invasion, and LNM.
Receiver operating characteristics curves for the AI model in the test set. a Prediction of undifferentiated histology (AUC = 0.992). b Prediction of submucosal invasion (AUC = 0.902). c Prediction of lymphovascular invasion (AUC = 0.706). d Prediction of lymph node metastasis (AUC = 0.680). Blue dots (n = 9) and red dots (n = 7) indicate the performance of the experts and novices, respectively. AI artificial intelligence, AUC area under the curve
Table 3 summarizes the performances of the AI model and endoscopists in the test. The model exhibited accuracies of 92.7% for undifferentiated histology, 87.3% for submucosal invasion, 76.4% for LVI, and 87.7% for LNM. The experts reported mean accuracies of 71.6% for undifferentiated histology, 72.6% for submucosal invasion, 69.7% for LVI, and 72.3% for LNM. The model showed significantly higher accuracy than the experts in predicting undifferentiated histology (P ≤ 0.001), submucosal invasion (P ≤ 0.012), and LNM (P ≤ 0.001). The experts showed significantly higher accuracy than the novices in identifying undifferentiated histology (P = 0.001) and submucosal invasion (P = 0.019). However, there was no significant difference between the experts and novices in detecting LVI (P = 0.525) and LNM (P = 0.790).
External validation of the AI model
Table 4 shows the performance of the AI model in the external validation. The model demonstrated predictive accuracies (per patient) of 75.6% (sensitivity, 81.7%; specificity, 72.5%) for undifferentiated histology, 71.9% (sensitivity, 53.3%; specificity, 80.6%) for submucosal invasion, 88.8% (sensitivity, 31.8%; specificity, 94.5%) for LVI, and 87.0% (sensitivity, 10.0%; specificity, 98.5%) for LNM.
Representative videos of the AI model in the test set and external validation are shown in Video 1 and 2, respectively. The resolution of images and videos in the external dataset (640 × 480) was lower than that in the internal dataset (1920 × 1080), owing to differences in the picture archiving and communication system between the two hospitals.
Discussion
In this study, we developed and evaluated an AI model that predicts postoperative pathologic results of EGC based on conventional white-light endoscopic images and videos. The performance of the model was compared with that of endoscopists in a test and externally validated using videos from another institution.
Categorizing the differentiation status of EGC is pivotal in deciding the indication for ESD, considering the significantly lower curative resection rate in undifferentiated-type EGC compared to differentiated-type EGC [22, 23]. Since approximately 18% of undifferentiated-type EGC can initially be misclassified as differentiated-type with forceps biopsy, endoscopic features of the lesions, including ME-NBI, must be combined for accurate diagnosis [24,25,26]. In a previous study, an AI model trained with ME-NBI showed an accuracy of 86.2% for classifying EGC differentiation status [27]. In our study, the AI model exhibited an accuracy of 89.7% in white-light endoscopic videos and outperformed the experts in identifying undifferentiated-type EGC. These results suggest that AI can assist endoscopists in predicting the differentiation status, with both white-light and ME-NBI endoscopic images.
Although EUS is commonly used to detect submucosal invasion in EGC, its advantages over conventional endoscopy are insignificant, with an accuracy of approximately 70% [10]. Notably, these findings are consistent with our results, where the experts showed a mean accuracy of 72.6% for predicting submucosal invasion of EGC in the test. In contrast, the AI model demonstrated significantly higher accuracy than the experts. Therefore, endoscopic findings indicative of submucosal invasion in EGC, such as clubbing, abrupt cutting or fusion of folds, uneven or nodular depression, and remarked redness of surface can be assessed without ultrasound [28,29,30,31], and AI enhances this process by learning an extensive dataset of conventional endoscopic images.
Several studies have investigated deep learning-based prediction of submucosal invasion in EGC using endoscopic images, reporting accuracies ranging from 84 to 94% [16, 32,33,34]. However, two studies revealed that undifferentiated-type EGC was associated with lower predictive accuracies compared with differentiated-type EGC [19, 35], a tendency also observed in our study. Furthermore, the significantly lower sensitivity for submucosal invasion was observed in undifferentiated EGC. Given that submucosal invasion with undifferentiated histology indicates non-curative ESD in EGC, these findings suggest that endoscopists still need to be more conservative when deciding to perform ESD for undifferentiated-type EGC than for differentiated-type EGC, even with the assistance of AI.
The lack of research on predicting LNM from endoscopic images using AI can be attributed to the low incidence of LNM in patients with EGC. The LNM rates have been reported to be < 9% for mucosal cancer and < 20% for submucosal cancer, according to large-scale studies based on surgical specimens of EGC [36, 37]. In addition, the LVI rate of EGC was approximately 13% in another study based on surgical specimens [38]. Although our study included as many surgical patients as possible, these inherently low rates of LVI and LNM in EGC induced an imbalance between positive and negative cases within the datasets. This is the reason our model exhibited lower sensitivity and positive prediction value, resulting in a low AUC value compared to its high accuracy in predicting LVI and LNM. However, excluding some patients with negative LVI or LNM to address this data imbalance could introduce significant selection bias. Therefore, despite the potential effect of data imbalance, we chose to include patients consecutively in the study.
Additionally, the absence of a significant difference in the mean accuracy between experts and novices suggests that the ability to detect LNM does not necessarily improve with clinical experience. However, the AI model in our study showed higher accuracy and sensitivity than the experts in predicting LNM. One possible explanation for this is that the AI may have adapted to associate a deeper invasion depth with an increased probability of LNM in EGC. This is supported by the fact that the model showed increased sensitivity for LNM in submucosal cancer compared to mucosal cancer, with the highest sensitivity in cancer with SM2 invasion. This trend was also observed in the prediction of LVI, a pathologic factor correlated with LNM in EGC [18, 39]. However, within the same categories of mucosal or submucosal cancer, there appeared to be no specific endoscopic features suggestive of LNM. Moreover, there are factors other than endoscopic findings associated with LNM in EGC, including age, sex, and tumor size, as reported in previous studies [40,41,42]. Therefore, future research could focus on integrating various clinical features indicative of LNM with endoscopic images to enhance AI-based detection of LNM in EGC.
This study has several limitations. First, the AI model was trained using retrospective images after the selection process, potentially introducing bias into our study. To compensate for this, we also included videos from patients enrolled prospectively under the same indications for ESD and found consistent performance of the model between images and videos. Furthermore, we tested the performance of the model by comparing it with experts from various hospitals across the nation. Second, the predictive performance of the AI model was lower in the external validation than in the internal tests. This discrepancy can be partially explained by the inferior resolution of images and videos in the external dataset compared to those in the internal dataset. Additionally, previous studies reported the “overfitting effect” in AI, where the learning process becomes excessively adapted to the training data [43, 44]. Several studies on deep learning-based prediction of invasion depth in gastric neoplasms have also reported significant differences in accuracies between internal and external tests [19, 45, 46]. Nevertheless, the external validation of our model showed predictive accuracy above 70% for invasion depth, which was higher than reported predictive accuracy of EUS. The performance could be further improved by training the model with images from various institutions in the future. Third, this study did not evaluate ME-NBI images and videos of EGC. Training this model with NBI data can improve the histological diagnosis of EGC, and it is essential to train the model with NBI images and videos in further studies. Fourth, incorporating both ESD and surgical cases into the dataset may have affected the model’s performance due to heterogeneity among the data. The longer section intervals in surgical specimens compared to ESD specimens could potentially lead to underestimation of submucosal invasion and LVI in surgical specimens [47]. Finally, the findings of this study should be confirmed in randomized controlled trials, and we are planning to conduct prospective studies to apply this AI model in clinical practice.
In conclusion, this study suggests that AI has the potential to assist endoscopists in determining the optimal treatment strategy for EGC, showing high performance in predicting the differentiation status and invasion depth based on conventional endoscopic images and videos. However, the detection of LVI and LNM using deep learning-based methods requires further research.
Change history
04 September 2024
A Correction to this paper has been published: https://doi.org/10.1007/s10120-024-01549-8
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71:209–49.
Vasconcelos AC, Dinis-Ribeiro M, Libânio D. Endoscopic resection of early gastric cancer and pre-malignant gastric lesions. Cancers (Basel). 2023;15:3084.
Ahn HS, Lee HJ, Yoo MW, Kim SG, Im JP, Kim SH, et al. Diagnostic accuracy of T and N stages with endoscopy, stomach protocol CT, and endoscopic ultrasonography in early gastric cancer. J Surg Oncol. 2009;99:20–7.
Wani AH, Parry AH, Feroz I, Choh NA. Preoperative staging of gastric cancer using computed tomography and its correlation with histopathology with emphasis on multi-planar reformations and virtual gastroscopy. J Gastrointest Cancer. 2021;52:606–15.
Kim TH, Kim IH, Kang SJ, Choi M, Kim BH, Eom BW, et al. Korean practice guidelines for gastric cancer 2022: An evidence-based, multidisciplinary approach. J Gastric Cancer. 2023;23:3–106.
Association JGC. Japanese Gastric Cancer Treatment Guidelines 2021. Gastric Cancer. 2023;26:1–25.
Shim CN, Kim H, Kim DW, Chung HS, Park JC, Lee H, et al. Clinicopathologic factors and outcomes of histologic discrepancy between differentiated and undifferentiated types after endoscopic resection of early gastric cancer. Surg Endosc. 2014;28:2097–105.
Aliaga Ramos J, Pedrosa MS, Yoshida N, Abdul Rani R, Arantes VN. Histopathologic diagnosis discrepancies between preoperative endoscopic forceps biopsies and specimens resected by endoscopic submucosal dissection in superficial gastric neoplasms. J Clin Gastroenterol. 2023;57:74–81.
Kim Y, Yoon HJ, Kim JH, Chun J, Youn YH, Park H, et al. Effect of histologic differences between biopsy and final resection on treatment outcomes in early gastric cancer. Surg Endosc. 2020;34:5046–54.
Choi J, Kim SG, Im JP, Kim JS, Jung HC, Song IS. Comparison of endoscopic ultrasonography and conventional endoscopy for prediction of depth of tumor invasion in early gastric cancer. Endoscopy. 2010;42:705–13.
Shi D, Xi XX. Factors affecting the accuracy of endoscopic ultrasonography in the diagnosis of early gastric cancer invasion depth: A meta-analysis. Gastroenterol Res Pract. 2019;2019:8241381.
Takamaru H, Yoshinaga S, Takisawa H, Oda I, Katai H, Sekine S, et al. Endoscopic ultrasonography miniature probe performance for depth diagnosis of early gastric cancer with suspected submucosal invasion. Gut Liver. 2020;14:581–8.
Lee J, Lee H, Chung JW. The role of artificial intelligence in gastric cancer: Surgical and therapeutic perspectives: A comprehensive review. J Gastric Cancer. 2023;23:375–87.
Chang YH, Shin CM, Lee HD, Park JB, Jeon J, Kang S, et al. Clinical evaluation of an artificial intelligence system for upper gastrointestinal endoscopy on detecting and diagnosing gastric lesions pathologically pre-diagnosed with atypia and gastric dysplasia: A pilot study. Gastrointest Endosc. 2023;97:764.
Bang CS. Artificial intelligence in the analysis of upper gastrointestinal disorders. Korean J Helicobacter Up Gastrointest Res. 2021;21:300–10.
Kim JH, Oh SI, Han SY, Keum JS, Kim KN, Chun JY, et al. An optimal artificial intelligence system for real-time endoscopic prediction of invasion depth in early gastric cancer. Cancers (Basel). 2022;14:6000.
Park CH, Yang DH, Kim JW, Kim JH, Kim JH, Min YW, et al. Clinical practice guideline for endoscopic resection of early gastrointestinal cancer. Korean J Helicobacter Up Gastrointest Res. 2020;20:117–45.
Lee S, Kim SG, Cho SJ. ision to perform additional surgery after non-curative endoscopic submucosal dissection for gastric cancer based on the risk of lymph node metastasis: A long-term follow-up study. Surg Endosc. 2023;37:7738–48.
Goto A, Kubota N, Nishikawa J, Ogawa R, Hamabe K, Hashimoto S, et al. Cooperation between artificial intelligence and endoscopists for diagnosing invasion depth of early gastric cancer. Gastric Cancer. 2023;26:116–22.
Tan M, Le Q 2019 EfficientNet Rethinking model scaling for convolutional neural networks. In Kamalika C, Ruslan S, (eds). Proceedings of the 36th international conference on machine learning Proceedings of the machine learning research, PMLR, London
Karras T, Laine S, Aittala M, Hellsten J, Lehtinen J, Aila T. Analyzing and improving the image quality of StyleGAN. IEEE Publications. 2019;2020:8107–16.
Ahn JY, Jung HY, Choi KD, Choi JY, Kim MY, Lee JH, et al. Endoscopic and oncologic outcomes after endoscopic resection for early gastric cancer: 1370 cases of absolute and extended indications. Gastrointest Endosc. 2011;74:485–93.
Bang CS, Park JM, Baik GH, Park JJ, Joo MK, Jang JY, et al. Therapeutic outcomes of endoscopic resection of early gastric cancer with undifferentiated-type histology: A Korean ESD registry database analysis. Clin Endosc. 2017;50:569–77.
Lee JH, Kim JH, Rhee K, Huh CW, Lee YC, Yoon SO, et al. Undifferentiated early gastric cancer diagnosed as differentiated histology based on forceps biopsy. Pathol Res Pract. 2013;209:314–8.
Shibagaki K, Amano Y, Ishimura N, Taniguchi H, Fujita H, Adachi S, et al. Diagnostic accuracy of magnification endoscopy with acetic acid enhancement and narrow-band imaging in gastric mucosal neoplasms. Endoscopy. 2016;48(1):16–25.
Muto M, Yao K, Kaise M, Kato M, Uedo N, Yagi K, et al. Magnifying endoscopy simple diagnostic algorithm for early gastric cancer (MESDA-G). Dig Endosc. 2016;28(4):379–93.
Ling T, Wu L, Fu Y, Xu Q, An P, Zhang J, et al. A deep learning-based system for identifying differentiation status and delineating the margins of early gastric cancer in magnifying narrow-band imaging endoscopy. Endoscopy. 2021;53:469–77.
Kim GH. Systematic endoscopic approach to early gastric cancer in clinical practice. Gut Liver. 2021;15(6):811–7.
Choi J, Kim SG, Im JP, Kim JS, Jung HC, Song IS. Endoscopic prediction of tumor invasion depth in early gastric cancer. Gastrointest Endosc. 2011;73(5):917–27.
Tsujii Y, Kato M, Inoue T, Yoshii S, Nagai K, Fujinaga T, et al. Integrated diagnostic strategy for the invasion depth of early gastric cancer by conventional endoscopy and EUS. Gastrointest Endosc. 2015;82(3):452–9.
Abe S, Oda I, Shimazu T, Kinjo T, Tada K, Sakamoto T, et al. Depth-predicting score for differentiated early gastric cancer. Gastric Cancer. 2011;14(1):35–40.
Zhu Y, Wang QC, Xu MD, Zhang Z, Cheng J, Zhong YS, et al. Application of convolutional neural network in the diagnosis of the invasion depth of gastric cancer based on conventional endoscopy. Gastrointest Endosc. 2019;89:806-815.e1.
Hamada K, Kawahara Y, Tanimoto T, Ohto A, Toda A, Aida T, et al. Application of convolutional neural networks for evaluating the depth of invasion of early gastric cancer based on endoscopic images. J Gastroenterol Hepatol. 2022;37:352–7.
Nagao S, Tsuji Y, Sakaguchi Y, Takahashi Y, Minatsuki C, Niimi K, et al. Highly accurate artificial intelligence systems to predict the invasion depth of gastric cancer: Efficacy of conventional white-light imaging, nonmagnifying narrow-band imaging, and indigo-carmine dye contrast imaging. Gastrointest Endosc. 2020;92:866-873.e1.
Yoon HJ, Kim S, Kim JH, Keum JS, Oh SI, Jo J, et al. A lesion-based convolutional neural network improves endoscopic detection and depth prediction of early gastric cancer. J Clin Med. 2019;8:1310.
Hirasawa T, Gotoda T, Miyata S, Kato Y, Shimoda T, Taniguchi H, et al. Incidence of lymph node metastasis and the feasibility of endoscopic resection for undifferentiated-type early gastric cancer. Gastric Cancer. 2009;12:148–52.
Abdelfatah MM, Barakat M, Lee H, Kim JJ, Uedo N, Grimm I, et al. The incidence of lymph node metastasis in early gastric cancer according to the expanded criteria in comparison with the absolute criteria of the Japanese Gastric Cancer Association: A systematic review of the literature and meta-analysis. Gastrointest Endosc. 2018;87:338–47.
Choi S, Song JH, Lee S, Cho M, Kim YM, Kim HI, et al. Lymphovascular invasion: traditional but vital and sensible prognostic factor in early gastric cancer. Ann Surg Oncol. 2021;28(13):8928–35.
Hatta W, Gotoda T, Oyama T, Kawata N, Takahashi A, Yoshifuku Y, et al. A Scoring System to Stratify Curability after Endoscopic submucosal Dissection for Early Gastric Cancer: “eCura system.” Am J Gastroenterol. 2017;112:874–81.
Zhang M, Ding C, Xu L, Feng S, Ling Y, Guo J, et al. A nomogram to predict risk of lymph node metastasis in early gastric cancer. Sci Rep. 2021;11:22873.
Sui W, Chen Z, Li C, Chen P, Song K, Wei Z, et al. Nomograms for predicting the lymph node metastasis in early gastric cancer by gender: A retrospective multicentric study. Front Oncol. 2021;11: 616951.
Kim SM, Min BH, Ahn JH, Jung SH, An JY, Choi MG, et al. Nomogram to predict lymph node metastasis in patients with early gastric cancer: A useful clinical tool to reduce gastrectomy after endoscopic resection. Endoscopy. 2020;52:435–43.
Cho BJ, Bang CS. Artificial intelligence for the determination of a management strategy for diminutive colorectal polyps: Hype, hope, or help. Am J Gastroenterol. 2020;115:70–2.
Yang YJ, Bang CS. Application of artificial intelligence in gastroenterology. World J Gastroenterol. 2019;25:1666–83.
Cho BJ, Bang CS, Lee JJ, Seo CW, Kim JH. Prediction of submucosal invasion for gastric neoplasms in endoscopic images using deep-learning. J Clin Med. 2020;9:1858.
Gong EJ, Bang CS, Lee JJ, Baik GH, Lim H, Jeong JH, et al. Deep learning-based clinical decision support system for gastric neoplasms in real-time endoscopy: Development and validation study. Endoscopy. 2023;55:701–8.
Kim YI, Kook MC, Choi JE, Lee JY, Kim CG, Eom BW, et al. Evaluation of submucosal or lymphovascular invasion detection rates in early gastric cancer based on pathology section interval. J Gastric Cancer. 2020;20(2):165–75.
Acknowledgements
This research was supported by grants from the National Research Foundation of Korea (#NRF-2022R1A2B5B01001430), the Korean Society of Gastrointestinal Endoscopy (2021), and Ainex Corporation (2021). The funding source had no role in the study design, analysis and interpretation of the data, drafting of the article, critical revision of the article, and final approval for submission of the article.
Funding
Open Access funding enabled and organized by Seoul National University. National Research Foundation of Korea, #NRF-2022R1A2B5B01001430, Soo-Jeong Cho, Korean Society of Gastrointestinal Endoscopy, Ainex Corporation.
Author information
Authors and Affiliations
Contributions
S.L. contributed to study concept and design, data collection, statistical analysis, interpretation of the data, and drafting of the manuscript; J.J. and J.P. contributed to data processing, statistical analysis, and development of the artificial intelligence model; Y.H.C. and C.M.S. contributed to material support and patient enrollment; M.J.O. and S.H.K. contributed to data collection and material support; S.K. and S.H.P contributed to data collection; S.G.K. contributed to critical revision of the manuscript for important intellectual content; H.J.L., and H.K.Y. contributed to material support; H.S.L. contributed to pathologic analysis of the data; S.J.C. contributed to study concept and design, critical revision of the manuscript for important intellectual content, obtaining funding, patient enrollment, and study supervision. All authors reviewed the results and revised the manuscript. The corresponding author had full access to all the data in the study and had final responsibility for the decision to submit the manuscript for publication. Special thanks to Ji Yong Ahn (Asan Medical Center, Republic of Korea), Jong Yeul Lee (National Cancer Center, Republic of Korea), Sung Kwan Shin (Yonsei Severance Hospital, Republic of Korea), Byung-Hoon Min (Samsung Medical Center, Republic of Korea), Bokyung Kim (Seoul National University Hospital, Republic of Korea), and Hyunsoo Chung (Seoul National University Hospital, Republic of Korea) for participating as expert endoscopists in the test.
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised to update affiliation 2.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file6 (MPG 157,906 KB)
Supplementary file7 (MPG 36,368 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, S., Jeon, J., Park, J. et al. An artificial intelligence system for comprehensive pathologic outcome prediction in early gastric cancer through endoscopic image analysis (with video). Gastric Cancer 27, 1088–1099 (2024). https://doi.org/10.1007/s10120-024-01524-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10120-024-01524-3