Abstract
Giant cell tumor of bone (GCTB) is a locally aggressive bone tumor that frequently shows local recurrence and occasionally shows malignant transformation to high-grade sarcoma. Histologically, conventional GCTB is composed mainly of three types of cells: mononuclear neoplastic cells with an osteoblastic precursor phenotype, mononuclear histiocytic cells, and osteoclast-like multinucleated giant cells. These cells interact with each other via the RANKL-RANK axis and other mechanisms for tumor formation. The vast majority of GCTBs were recently revealed to harbor H3F3A p.G34W mutation, and a minor subset have H3F3A p.G34L, p.G34M, p.G34R, or p.G34V mutation. H3.3 G34W mutant-specific immunohistochemistry is a highly sensitive and specific surrogate marker for H3F3A p.G34W mutation in GCTB and thus useful for differential diagnoses of histological mimics. H3.3 mutant-specific immunohistochemistry has also contributed to the understanding of the bone-forming ability of neoplastic cells of GCTB and the remarkable new bone formation after treatment with denosumab, an inhibitor of RANKL. In primary and secondary malignant GCTBs, the H3F3A gene allele can be preserved or lost with malignant transformation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
The clinical and histopathological features of GCTB
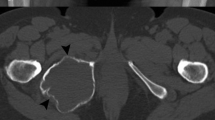
Giant cell tumor of bone (GCTB) is an osteolytic neoplasm that usually develops in the metaphysis to epiphysis of a long bone (such as the femur or tibia) or in the axial skeleton (such as the spine or sacrum) of young to middle-aged adults [1]. GCTB may occur in the metaphysis without involving the epiphysis, particularly in children whose growth plates are still open. Clinically, GCTB behaves as a locally aggressive tumor, sometimes with destruction of bone cortex and extraosseous invasion. GCTBs frequently show local recurrence. Some GCTBs have shown metastasis to the lung, in keeping with the histology of conventional GCTB. In rare instances, GCTB shows malignant transformation to high-grade sarcoma [2].
Histologically, conventional GCTB is composed mainly of three types of cells: mononuclear neoplastic cells, mononuclear histiocytic cells, and osteoclast-like multinucleated giant cells (Fig. 1a) [1]. The mononuclear neoplastic cells have round to spindle-shaped nuclei and various degrees of mitotic figures. Secondary aneurysmal bone cyst-like change is often associated with GCTB. The name "giant cell tumor" has been somewhat confusing regarding the nature of neoplastic cells. Recent studies have obtained evidence that mononuclear cells with an osteoblastic precursor phenotype are the true neoplastic element of GCTB, whereas mononuclear histiocytic cells and osteoclast-like multinucleated giant cells are considered non-neoplastic elements [3, 4]. However, mononuclear neoplastic cells and mononuclear histiocytic cells are indistinguishable only by cellular morphology on hematoxylin–eosin stained specimens. Mononuclear neoplastic cells can be identified by osteoblast-related markers such as RUNX2 and p63 [3]. In contrast, mononuclear histiocytic cells and osteoclast-like multinucleated giant cells are positive for CD68.
Histological appearance of a giant cell tumor of bone (GCTB) (a–c) and immunohistochemistry for H3.3 G34W mutant-specific antibody (d–f). Conventional GCTBs consist of mononuclear neoplastic cells, mononuclear histiocytic cells, and osteoclast-like multinucleated giant cells (a), and mononuclear neoplastic cells are positive for H3.3 G34W (d). Focal osteoid formation is seen in GCTB (b), and the osteoid-forming cells are positive for H3.3 G34W (e). Immature bone formation is remarkable in this denosumab-treated GCTB (c), and the bone-forming cells are positive for H3.3 G34W (f)
Despite the osteoblastic precursor phenotype of neoplastic cells, bone formation is usually absent and osteoid formation can be seen only focally in conventional GCTB (Fig. 1b) [5]. This matter is discussed later.
Several experimental data support that mononuclear neoplastic cells, mononuclear histiocytic cells, and osteoclastic giant cells interact with each other via growth factors and cytokines, and they are thought to play important roles in the development of bone-destructive lesions with unique histology [reviewed in Ref. 3].
Histone H3.3 mutation and the diagnostic implications
As a basic element of chromatin, the nucleosome consists of a core of eight histone proteins (two each of H2A, H2B, H3, and H4) wrapped by DNA. The N-terminal tails of these histone proteins can be modified by histone modificatory enzymes, including acetyltransferases for acetylation and methyltransferases for methylation [6]. Histone modification plays an important role in the epigenetic regulation of gene expression. Histone H3.3 is encoded by two genes located in different loci: H3F3A on chromosome 1 and H3F3B on chromosome 17. Data are accumulating regarding histone H3.3 gene mutations in neoplasms, such as p.K27M and p.G34R/V in pediatric gliomas, p.G34W in GCTBs, and p.K36M in chondroblastomas (Fig. 2) [7,8,9].
Approximately 90% of GCTBs harbor H3F3A p.G34W mutation, and minor subsets (each < 2%) have H3F3A p.G34L, p.G34M, p.G34R, or p.G34V mutation [9,10,11,12]. Up to 90–95% of chondroblastomas have H3F3B p.K36M mutation [9, 11]. A monoclonal antibody against H3.3 G34W mutant protein (clone RM263) is now commercially available for immunohistochemistry. Several studies have revealed that H3.3 G34W mutant-specific immunohistochemistry is a highly sensitive and specific surrogate marker for the presence of H3F3A p.G34W mutation in GCTBs [12,13,14]. Immunoreactivity for H3.3 G34R and H3.3 G34V mutant-specific antibodies also well correlate with the presence of corresponding H3F3A mutation [12]. Likewise, an H3.3 K36M mutant-specific antibody is available for the diagnosis of chondroblastoma [15].
In GCTBs, mononuclear cells show nuclear expression of H3.3 G34W with a mosaic pattern (Fig. 1d) [12]. This phenomenon probably indicates that the H3.3 G34W-positive neoplastic cells and the H3.3 G34W-negative histiocytic cells are admixed within the mononuclear cell population.
GCTB should be distinguished from various types of bone lesions with osteoclastic giant cells, including chondroblastoma, non-ossifying fibroma, giant cell reparative granuloma, primary aneurysmal bone cyst, and giant cell-rich type osteosarcoma. These lesions (except for a minor subset of osteosarcomas) are negative for H3.3 G34W immunohistochemistry, indicating the utility of this immunohistochemistry for the differential diagnosis of morphologic mimics [12, 13].
RNNKL-RANK signal, denosumab therapy, and histopathological correlation
It is well known that receptor activator of nuclear factor kappa-B ligand (RANKL) and its receptor RANK axis play an important role in the development of GCTB [reviewed in Ref. 3]. RANKL is expressed by mononuclear neoplastic cells and binds to RANK on the surface of osteoclastic multinucleated giant cells, leading to the activation and proliferation of these giant cells (Fig. 3). The activated osteoclastic giant cells play an important role in bone absorption.
Hypothetical model of bone formation and absorption in GCTB. In conventional GCTBs, bone absorption by osteoclastic giant cells is activated, while bone formation by H3F3A-mutated neoplastic cell is very limited (a). In a post-denosumab GCTB, bone absorption by osteoclastic giant cells is suppressed due to the inhibition of the RANKL-RANK axis by denosumab (b). Instead, immature new bone formation by neoplastic cells is increased (b)
Denosumab, an inhibitor of RANKL, has shown a promising antitumor effect for GCTB [16, 17]. By targeting the RANKL-RANK axis, the osteoclastic activity of giant cell is considerably reduced. GCTBs after denosumab treatment have shown drastic histological changes such as giant cell depletion, various degrees of spindle-cell proliferation, and new bone formation (Fig. 1c) [12, 18,19,20]. According to these reports, the spindle cells and the cells in and around immature bone in denosumab-treated GCTBs are immunohistochemically positive for H3.3 G34W (Fig. 1f), and H3F3A mutations are consistently detected in the corresponding samples [12, 14, 19, 20]. These findings suggest that neoplastic cells may still be alive even after denosumab treatment. Indeed, a recurrent lesion after the discontinuation of denosumab treatment can show conventional GCTB morphology again.
The appearance of post-denosumab GCTBs is distinct from the histology of conventional GCTB and can be reminiscent of osteosarcoma or malignancy in GCTB, potentially leading to diagnostic difficulty [18]. In diagnostically challenging cases, it is therefore important to know the patient's complete clinical history and to perform immunohistochemistry for H3.3 G34W.
Remarkable immature bone formation in post-denosumab GCTBs may be related to the potential bone-forming ability of neoplastic cells and an interaction between the neoplastic cells and osteoclastic giant cells (Fig. 3). As mentioned above, focal or only limited osteoid formation is sometimes observed in denosumab-naïve GCTBs [1, 5]. Our recent study revealed that the cells within osteoid and immature bone was occasionally immunoreactive for H3.3 G34W in primary GCTB (Fig. 1e), suggesting that at least some population of osteoid-forming cells in GCTBs has a neoplastic nature, and these neoplastic cells have a latent bone-forming ability [12].
In experimental models, osteoclasts were likely to suppress osteoblast differentiation [3, 21]. The neoplastic cells of GCTB are able to differentiate into mature osteoblasts when separated from the osteoclasts, and a mineralized nodule or bone formation can be induced [22, 23]. In this context, the remarkable new bone formation in denosumab-treated GCTBs might be explained in part by the idea that the osteoblastic differentiation and bone-forming abilities of mononuclear neoplastic cells are activated once these cells are free from the influence of osteoclastic giant cells (Fig. 3) [12, 19]. Further research is needed to test this hypothesis.
Malignant giant cell tumor of bone
Malignant change in GCTB is a rare phenomenon, occurring in approx. 1% of GCTBs [1, 2]. Such a malignancy presents as either the primary tumor at presentation (primary malignant GCTB) or secondary malignant transformation in a recurrent GCTB, especially after radiation therapy (secondary malignant GCTB). The malignant component histologically corresponds to osteosarcoma, fibrosarcoma or, less frequently, undifferentiated high-grade pleomorphic sarcoma (formerly, malignant fibrous histiocytoma) [2]. At the molecular level, some population of malignant GCTBs have TP53 gene mutation in the malignant component [24].
Several recent studies demonstrated that both conventional GCTB and malignant components shared H3F3A p.G34W mutation and H3.3 G34W mutant protein expression, suggesting that this type of mutation is preserved even after the malignant transformation (Fig. 4a) [12, 25, 26]. In contrast, Yoshida et al. reported that, in some population of malignant GCTBs, H3F3A gene mutation and H3.3 G34W immunoreactivity are not present in the malignant component whereas the conventional GCTB component harbors this mutation (Fig. 4b) [27]. Those authors observed that one of the two alleles of H3F3A gene is deleted in the malignant cells in the malignant GCTB cases with loss of H3F3A mutation. Therefore, the loss of one allele of H3F3A (probably the mutant allele) in the malignant component may be responsible for the discordant H3F3A status between the two components. As the molecular mechanism of malignant transformation in GCTB is not fully elucidated, further investigation is expected.
Different H3.3 G34W expression status patterns in malignant GCTB. Concordant pattern (a) and discordant pattern (b) (each: hematoxylin–eosin stain with H3.3 G34W immunohistochemistry in inset). In the concordant pattern, the malignant component is positive for H3.3 G34W (a). In the discordant pattern, the malignant component is negative for H3.3 G34W (b). In both examples, the conventional GCTB component is positive for H3.3 G34W (not shown)
Conclusion
The discovery of the H3F3A gene mutation and the development of H3.3 mutant-specific immunohistochemistry have contributed to the improvement of the diagnosis of GCTB. In addition to the diagnostic aspect, the H3.3 mutant-specific immunohistochemistry has contributed to our understanding of the bone-forming ability of GCTBs and the remarkable new bone formation after the denosumab treatment. According to recent studies, H3F3A gene mutation status is presumably variable in the malignant transformation of GCTB. Further research is expected to clarify the molecular biology of GCTB and to establish better therapeutic strategies for this tumor.
References
Athanasou NA, Bansal M, Forsyth R, Reid RP, Sapi Z (2013) Giant cell tumour of bone. In: Fletcher CDM, Bridge JA, Hogendoorn PCW, Mertens F (eds) The WHO classification of tumours of soft tissue and bone. IARC, Lyon, pp 321–324
Gong L, Liu W, Sun X, Sajdik C, Tian X, Niu X, Huang X (2012) Histological and clinical characteristics of malignant giant cell tumor of bone. Virchows Arch 460:327–334
Cowan RW, Singh G (2013) Giant cell tumor of bone: a basic science perspective. Bone 52:238–246
Murata A, Fujita T, Kawahara N, Tsuchiya H, Tomita K (2005) Osteoblast lineage properties in giant cell tumors of bone. J Orthop Sci 10:581–588
Gupta R, Seethalakshmi V, Jambhekar NA, Prabhudesai S, Merchant N, Puri A, Agarwal M (2008) Clinicopathologic profile of 470 giant cell tumors of bone from a cancer hospital in western India. Ann Diagn Pathol 12:239–248
Tessarz P, Kouzarides T (2014) Histone core modifications regulating nucleosome structure and dynamics. Nat Rev Mol Cell Biol 15:703–708
Lowe BR, Maxham LA, Hamey JJ, Wilkins MR, Partridge JF (2019) Histone H3 mutations: an updated view of their role in chromatin deregulation and cancer. Cancers (Basel) 11:660
Schwartzentruber J, Korshunov A, Liu XY, Jones DT, Pfaff E, Jacob K, Sturm D, Fontebasso AM, Quang DA, Tönjes M, Hovestadt V, Albrecht S, Kool M, Nantel A, Konermann C, Lindroth A, Jäger N, Rausch T, Ryzhova M, Korbel JO, Hielscher T, Hauser P, Garami M, Klekner A, Bognar L, Ebinger M, Schuhmann MU, Scheurlen W, Pekrun A, Frühwald MC, Roggendorf W, Kramm C, Dürken M, Atkinson J, Lepage P, Montpetit A, Zakrzewska M, Zakrzewski K, Liberski PP, Dong Z, Siegel P, Kulozik AE, Zapatka M, Guha A, Malkin D, Felsberg J, Reifenberger G, von Deimling A, Ichimura K, Collins VP, Witt H, Milde T, Witt O, Zhang C, Castelo-Branco P, Lichter P, Faury D, Tabori U, Plass C, Majewski J, Pfister SM, Jabado N (2012) Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 482:226–231
Behjati S, Tarpey PS, Presneau N, Scheipl S, Pillay N, Van Loo P, Wedge DC, Cooke SL, Gundem G, Davies H, Nik-Zainal S, Martin S, McLaren S, Goody V, Robinson B, Butler A, Teague JW, Halai D, Khatri B, Myklebost O, Baumhoer D, Jundt G, Hamoudi R, Tirabosco R, Amary MF, Futreal PA, Stratton MR, Campbell PJ, Flanagan AM (2013) Distinct H3F3A and H3F3B driver mutations define chondroblastoma and giant cell tumor of bone. Nat Genet 45:1479–1482
Presneau N, Baumhoer D, Behjati S, Pillay N, Tarpey P, Campbell PJ, Jundt G, Hamoudi R, Wedge DC, Loo PV, Hassan AB, Khatri B, Ye H, Tirabosco R, Amary MF, Flanagan AM (2015) Diagnostic value of H3F3A mutations in giant cell tumour of bone compared to osteoclast-rich mimics. J Pathol Clin Res 1:113–123
Kervarrec T, Collin C, Larousserie F, Bouvier C, Aubert S, Gomez-Brouchet A, Marie B, Miquelestorena-Standley E, Le Nail LR, Avril P, Christophe Pagès J, de Pinieux G (2017) H3F3 mutation status of giant cell tumors of the bone, chondroblastomas and their mimics: a combined high resolution melting and pyrosequencing approach. Mod Pathol 30:393–406
Yamamoto H, Iwasaki T, Yamada Y, Matsumoto Y, Otsuka H, Yoshimoto M, Kohashi K, Taguchi K, Yokoyama R, Nakashima Y, Oda Y (2018) Diagnostic utility of histone H3.3 G34W, G34R, and G34V mutant-specific antibodies for giant cell tumors of bone. Hum Pathol 73:41–50
Amary F, Berisha F, Ye H, Gupta M, Gutteridge A, Baumhoer D, Gibbons R, Tirabosco R, O'Donnell P, Flanagan AM (2017) H3F3A (Histone 3.3) G34W immunohistochemistry: a reliable marker defining benign and malignant giant cell tumor of bone. Am J Surg Pathol 41:1059–1068
Lüke J, von Baer A, Schreiber J, Lübbehüsen C, Breining T, Mellert K, Marienfeld R, Schultheiss M, Möller P, Barth TFE (2017) H3F3A mutation in giant cell tumour of the bone is detected by immunohistochemistry using a monoclonal antibody against the G34W mutated site of the histone H3.3 variant. Histopathology 71:125–133
Amary MF, Berisha F, Mozela R, Gibbons R, Guttridge A, O'Donnell P, Baumhoer D, Tirabosco R, Flanagan AM (2016) The H3F3 K36M mutant antibody is a sensitive and specific marker for the diagnosis of chondroblastoma. Histopathology 69:121–127
Chawla S, Henshaw R, Seeger L, Choy E, Blay JY, Ferrari S, Kroep J, Grimer R, Reichardt P, Rutkowski P, Schuetze S, Skubitz K, Staddon A, Thomas D, Qian Y, Jacobs I (2013) Safety and efficacy of denosumab for adults and skeletally mature adolescents with giant cell tumour of bone: interim analysis of an open-label, parallel-group, phase 2 study. Lancet Oncol 14:901–908
Ueda T, Morioka H, Nishida Y, Kakunaga S, Tsuchiya H, Matsumoto Y, Asami Y, Inoue T, Yoneda T (2015) Objective tumor response to denosumab in patients with giant cell tumor of bone: a multicenter phase II trial. Ann Oncol 26:2149–2154
Wojcik J, Rosenberg AE, Bredella MA, Choy E, Hornicek FJ, Nielsen GP, Deshpande V (2016) Denosumab-treated giant cell tumor of bone exhibits morphologic overlap with malignant giant cell tumor of bone. Am J Surg Pathol 40:72–80
Kato I, Furuya M, Matsuo K, Kawabata Y, Tanaka R, Ohashi K (2018) Giant cell tumours of bone treated with denosumab: histological, immunohistochemical and H3F3A mutation analyses. Histopathology 72:914–922
Girolami I, Mancini I, Simoni A, Baldi GG, Simi L, Campanacci D, Beltrami G, Scoccianti G, D'Arienzo A, Capanna R, Franchi A (2016) Denosumab treated giant cell tumour of bone: a morphological, immunohistochemical and molecular analysis of a series. J Clin Pathol 69:240–247
Kubota K, Sakikawa C, Katsumata M, Nakamura T, Wakabayashi K (2002) Platelet-derived growth factor BB secreted from osteoclasts acts as an osteoblastogenesis inhibitory factor. J Bone Miner Res 17:257–265
Huang L, Teng XY, Cheng YY, Lee KM, Kumta SM (2004) Expression of preosteoblast markers and Cbfa-1 and Osterix gene transcripts in stromal tumour cells of giant cell tumour of bone. Bone 34:393–401
James IE, Dodds RA, Olivera DL, Nuttall ME, Gowen M (1996) Human osteoclastoma-derived stromal cells: correlation of the ability to form mineralized nodules in vitro with formation of bone in vivo. J Bone Miner Res 11:1453–1460
Oda Y, Sakamoto A, Saito T, Matsuda S, Tanaka K, Iwamoto Y, Tsuneyoshi M (2001) Secondary malignant giant-cell tumour of bone: molecular abnormalities of p53 and H-ras gene correlated with malignant transformation. Histopathology 39:629–637
Tsukamoto Y, Futani H, Kihara T, Watanabe T, Kumanishi S, Matsuo S, Hirota S, Ueda T, Yamamoto H, Yoshiya S (2018) An extremely rare case of primary malignancy in giant cell tumor of bone, arising in the right femur and harboring H3F3A mutation. Pathol Res Pract 214:1504–1509
Righi A, Mancini I, Gambarotti M, Picci P, Gamberi G, Marraccini C, Dei Tos AP, Simi L, Pinzani P, Franchi A (2017) Histone 33 mutations in giant cell tumor and giant cell-rich sarcomas of bone. Hum Pathol 68:128–135
Yoshida KI, Nakano Y, Honda-Kitahara M, Wakai S, Motoi T, Ogura K, Sano N, Shibata T, Okuma T, Iwata S, Kawai A, Ichimura K, Yoshida A (2019) Absence of H3F3A mutation in a subset of malignant giant cell tumor of bone. Mod Pathol 8:89. https://doi.org/10.1038/s41379-019-0318-5
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yamamoto, H., Ishihara, S., Toda, Y. et al. Histone H3.3 mutation in giant cell tumor of bone: an update in pathology. Med Mol Morphol 53, 1–6 (2020). https://doi.org/10.1007/s00795-019-00238-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00795-019-00238-1