Abstract
Primary bone tumors are rare cancers that cause significant morbidity and mortality. The recent identification of recurrent mutations in histone genes H3F3A and H3F3B within specific bone cancers, namely, chondroblastomas and giant cell tumors of bone (GCTB), has provided insights into the cellular and molecular origins of these neoplasms and enhanced understanding of how histone variants control chromatin function. Somatic mutations in H3F3A and H3F3B produce oncohistones, H3.3G34W and H3.3K36M, in more than nine of ten GCTB and chondroblastomas, respectively. Incorporation of the mutant histones into nucleosomes inhibits histone methyltransferases NSD2 and SETD2 to alter the chromatin landscape and change gene expression patterns that control cell proliferation, survival, and differentiation, as well as DNA repair and chromosome stability. The discovery of these histone mutations has facilitated more accurate diagnoses of these diseases and stratification of malignant tumors from benign tumors so that appropriate care can be delivered. The broad-scale epigenomic and transcriptomic changes that arise from incorporation of mutant histones into chromatin provide opportunities to develop new and disease-specific therapies. In this chapter, we review how mutant histones inhibit SETD2 and NSD2 function in bone tumors and discuss how this information could lead to better treatments for these cancers.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
4.1 Introduction
Primary bone tumors are rare cancers that produce significant morbidity and mortality in children and adults. Bone neoplasms cause debilitating pain, impede skeletal growth, and weaken bones. Benign bone tumors are treated surgically if they are painful or prominent, but many require no treatment. Malignant bone tumors will be treated surgically if possible and with antineoplastic regimens, such as adjunct chemotherapy and radiation. In some cases, amputation or major resection is required, which results in lifelong disability and medical care. Children treated with radiation and chemotherapy for bone cancers are at increased risk for other cancers and require continuous monitoring.
Bone cancers are categorized by the cell type from which they originate, by the type of tissue they produce (e.g., bone (osteo-) or cartilage (chondro-)), by appearance on radiographs or pathology sections, and increasingly by the presence of reoccuring genetic mutations. For example, recurrent chromosomal translocations characterize Ewing’s sarcoma and Ewing-related family of tumors [1]. Recently, somatic mutations in histone genes were found in more than 90% of chondroblastomas and giant cell tumors of the bone (GCTB) [2, 3]. This discovery launched experimentation to understand how mutated histones (oncohistones) contribute to tumorigenesis and enhanced basic understanding of chromatin organization. This chapter will review fundamentals of bone formation and homeostasis and then focus on how genetic mutations within histone genes cause changes to the epigenome and drive tumorigenesis in bone tissues. Ways in which this information can lead to new therapeutic strategies for debilitating bone tumors will be discussed.
4.2 Bone Development, Homeostasis, and Bone Tumors
Bones are highly dynamic organs not just during growth phases but throughout life. Bones provide structural support to skeletal muscles, protect our internal organs, store calcium and other minerals, and produce endocrine factors that affect overall health and function of other tissues, including the skeletal muscle, the gut, and the brain. Bones are formed by two processes: intramembranous ossification and endochondral ossification [4]. During intramembranous ossification, mesenchymal progenitor cells condense and differentiate into osteoblasts that produce an extracellular matrix rich in collagen 1 and mineralization promoting factors that stimulate hydroxyapatite incorporation into the matrix. This process forms the clavicle (collar bone) and many flat bones in the skull. Most bones form by endochondral ossification when mesenchymal progenitors differentiate into chondrocytes, which produce matrices rich in collagen 2, proteoglycans, and mineralization inhibitors. Vascular and neuronal invasion of these cartilaginous structures promotes recruitment of hematopoietic cells, including monocytes that differentiate into osteoclasts and carve out the marrow cavity. Mesenchymal progenitor cells that differentiate into osteoblasts are also recruited and gradually replace the chondrocytic matrix with the mineralized matrix.
Long bone growth continues until the cartilaginous epiphyseal growth plate closes. Chondrocytes in growth plates are organized in columnar structures and horizontal zones (Fig. 4.1). The most distal zone of growth plates on each end of a long bone contains proliferative cells that maintain the chondrocyte pool in developing bones. After proliferation ceases, chondrocytes undergo hypertrophy, and this enlargement in cell size drives longitudinal bone lengthening. Most hypertrophic chondrocytes will die by apoptosis, but some survive and take residence in the bone marrow where they are poised to contribute to fracture healing [5].
After bone growth ceases, bones remain highly dynamic organs throughout life to keep bones strong and healthy. Every day, osteoblasts (bone forming cells) and osteoclasts (bone resorbing cells) are actively remodeling and refreshing the skeleton to ensure that the skeleton remains strong and that the body has a sufficient supply of essential minerals and growth factors. The tight communication between osteoblasts and osteoclasts is skewed during the menopause and with aging such that bone resorption exceeds bone formation and the skeleton weakens. This communication is also altered by primary and metastatic bone tumors that find bone to be a nurturing environment. Bone tumors have a lytic, blastic, or mixed bone phenotype and affect bone strength. Blastic tumors will produce a collagen 1-rich matrix, but the collagen is secreted in woven patterns that are mechanically weaker than the normal lamellar pattern. In contrast, osteolytic tumors activate osteoclasts that resorb bone matrix, lowering bone density and releasing growth factors to sustain their growth in a vicious cycle [6]. Primary bone tumors are relatively rare but frequently occur in children and young adults who are actively growing. Primary bone tumors that become metastatic typically colonize the lungs or other bones. Increased understanding of the molecular mechanisms that drive carcinogenesis, colonization, and metastasis of bone neoplasms will lead to better treatments that optimally treat the bone niche as well as the tumor.
4.3 Histones and Epigenetics
Histones are cellular proteins responsible for the storage, organization, and accessibility of DNA in the nucleus. Proteins from four histone families (H2A, H2B, H3, and H4) form an octamer consisting of two H3-H4 dimers and two H2A-H2B dimers upon which 166 nucleotides of DNA circle twice to form a nucleosome (Fig. 4.2). Proteins from a fifth histone family (H1) stabilize each nucleosome and link it to adjacent nucleosomes. Canonical histones are synthesized during S phase and incorporated into nucleosomes by chromatin assembly factor (CAF)-1-dependent mechanisms [7]. Histone variants may substitute for the core canonical histones in nucleosomes as genomic DNA is used during the cell cycle. Replication-independent histone variants are produced throughout the cell cycle and are inserted into nucleosomes by distinct molecular mechanisms involving histone regulator A (HIRA) [8]. Two canonical forms of H3 (H3.1 and H3.2) and the histone variant H3.3 have been linked to cancers [9]. Histone variant H3.3 varies from H3.1 and H3.2 by just five amino acids in the core domain and is present in open or active chromatin (i.e., euchromatin).
Nucleosome structures. The nucleosome is the basic unit of organization and consists of eight histone proteins consisting of two H2A, H2B, H3, and H4 with two full loops of DNA. Together with a short amount of linker DNA that extends toward the next nucleosome and another histone protein type (linker histone H1), each histone octamer organizes approximately 200 bp of DNA. N-termini of histones extend from the nucleosome core. One tail from a H3 molecule is shown here. The amino acids changed by genetic mutations in bone tumors are indicated
In addition to being scaffolds for genomic DNA, histones play an important communication role in cell nuclei as posttranslational modifications (PTM) generate codes that control gene transcription or other molecular events and allow cells to respond to their environment [10]. Canonical and variant histones within the nucleosome octamer are arranged such that their amino-termini can extend into the environment, away from the protein core and DNA (Fig. 4.2). This allows enzymes and other proteins to bind and posttranslationally modify select amino acids (e.g., lysines, arginines, and serines) in histones in response to environmental cues. Hundreds of PTMs exist [11], but methylation, acetylation, and phosphorylation are the best understood. In euchromatin (active or open chromatin), PTMs are added to histones by enzymes collectively referred to as “writers,” removed by enzymes called “erasers,” and functionally interpreted by proteins called “readers” [12, 13]. Writers and erasers typically have no DNA binding activity and are recruited to specific genomic regions by sequence-specific transcription factors. Readers are attracted by the presence or absence of the PTM and build a platform for the recruitment of other complexes that regulate gene expression and chromatin structure. The regulation of histone modifications allows for timely gene expression while maintaining nuclear structure.
4.3.1 Lysine Methylation of Histones
Methylation of histones H3 and H4 is achieved by histone methyltransferases (HMTase) and requires S-adenosyl methionine as the methyl donor [14]. Histone methylation can involve the transfer of one to three methyl groups on a single lysine or arginine residue (Fig. 4.3), resulting in mono-, di-, or tri-methylated states (me1, me2, and me3, respectively). Methylation of specific lysines and arginines within histones controls transcriptional activation, repression, or elongation. For example, methylation of H3K4, H3K36, and H3K79 is typically associated with open chromatin and gene activation, while methylation of H3K9, H3K27, and H4K20 is associated with transcriptional repression of genes [15]. Beyond regulating gene expression, histone methylation may regulate other functions, such DNA repair and stability.
HMTases are recruited to specific regions of the genome by combinations of transcription factors, histone PTMs, DNA methylation, as well as noncoding RNAs [16,17,18]. HMTases are subdivided based on structure and function into three groups: (1) SET (Su(var)3-9, Enhancer of Zeste, Trithorax) domain-containing lysine methyltransferases, (2) non-SET-domain-containing lysine methyltransferases, and (3) protein arginine methyltransferases (PRMT). SET domain-containing HMTases demonstrate substrate specificity for lysines in histone tails, while HMTases lacking SET domains (e.g., DOT1) methylate lysines in the histone core sequence. The SET domain and flanking peptide sequences form flexible β-strand and β-sheet structures that can recognize many lysine substrates, including partially methylated lysines. The activity and/or expression of several HMTases are altered in many human cancers [19, 20]. In bone tumors, the functional activities of SET domain-containing HMTases, NSD2 and SETD2, are suppressed, and global histone H3K36 di- and tri-methylation are consequently reduced [21, 22].
4.3.2 Nuclear Receptor SET Domain-Containing (NSD) Methyltransferase 2
NSD2 is a SET domain-containing HMTase that is generally associated with open and active chromatin. It preferentially methylates the 36th amino acid (lysine) of H3 when it is incorporated in nucleosomes creating H3K36me2 and H3K36me3 (Fig. 4.3) but can also generate H3K4me3, H3K27me3, and H4K20me3 [23]. NSD2 is one of several genes on the short arm of chromosome 4 that is deleted in Wolf-Hirschhorn syndrome (WHSC1), a condition affecting many tissues, including the skeleton, and that is characterized by significant growth and cognitive delays. Also known as MMSET, NSD2 is also involved in a chromosomal translocation, t(4;14), present in 15% of patients with multiple myeloma, which is a hematopoietic malignancy of plasma cells that arises in the bone marrow. This chromosomal rearrangement places NSD2 downstream of an immunoglobulin promoter, which drives its transcription and overexpression in B-lineage cells [24]. Myeloma cells harboring this translocation also have increased global levels of H3K36me2 [23, 25]. Overexpression of catalytically active NSD causes aberrant enrichment of H3K36me2 at normally silent oncogenes (e.g., MET, PAK1, RRAS2, TGFA), triggering increased expression of these tumor-promoting pathways and transformation of primary B cells [26]. In musculoskeletal tumors that will be discussed below, NSD activity is inhibited by mutant histones. Thus, proper regulation of NSD2 activity and H3K36 methylation is necessary for maintaining cellular homeostasis and preventing cancer.
4.3.3 Su(Var)3-9, Enhancer of Zeste, Trithorax Domain-Containing (SETD) Methyltransferase 2
SETD2 is an HMTase that preferentially methylates H3K36me2 to create H3K36me3, open chromatin regions, and promote DNA repair and chromosomal stability. SETD2 is the primary H3K36 tri-methyltransferase responsible for the bulk of H3K36me3 in most cell types [27] (Fig. 4.3). SETD2 is highly conserved from Drosophila to humans and contains three functional regions: (1) triplicate AWS-SET-PostSET domains, (2) WW domain, and (3) Set2 Rpb1 interacting (SRI) domain. The AWS-SET-PostSET domains mediate histone H3K36-specific activities by transferring methyl groups from S-adenosyl-L-methionine to the amino group of lysine residues in histones [27, 28]. The WW domain contains two tryptophan residues that are 20 amino acids apart and mediates interactions of SETD2 with other proteins containing PPxYpSPpTP sequences, including the Huntington disease protein [29, 30]. Abnormal expression of the WW domain in SETD2 has been linked to various cancer types and Alzheimer’s disease [31,32,33]. The SRI domain facilitates interactions with hyperphosphorylated RNA polymerase II and couples histone H3K36me3 with transcriptional activation and elongation [34]. Deletion of the SRI domain of SETD2 eliminates the interaction between RNA polymerase II and reduces H3K36me3 and transcription elongation [35].
Loss of function from mutations in SETD2 and mutations altering its prime substrate H3K36 has been linked in numerous solid tumors and chemotherapy resistance. Various cancers (clear cell renal cell carcinomas, T-cell lymphoma, breast cancer, and leukemia) harbor inactivating mutations in SETD2 [36]. SETD2 was deemed as a tumor suppressor after a missense mutation that inactivated the protein that was identified in clear cell renal cell carcinoma patients [37]. SETD2 levels were also significantly reduced in breast cancers compared to adjacent noncancerous tissue [38].
4.4 Histone Mutations in Bone Tumors
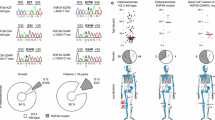
In a number of adult and pediatric cancers, somatic mutations in the H2F2A and H3F3B genes that encode H3.3 variants disrupt homeostatic control of histone PTMs, particularly histone methylation, by producing oncohistones that competitively inhibit the activity of HMTases, NSD2 and SETD2. The oncohistones cause chaos in the intricate processes controlling gene expression and consequently lead to activation of oncogenic pathways and/or inhibition of tumor suppressors. The amino acid changes in histone H3 proteins are highly specific to certain cancers. Recurrent H3K27M and H3G34V/R mutations have been discovered pediatric high-grade gliomas [39], and several years later, H3K36M and H3G34W/L are found in over 90% of chondroblastomas and giant cell tumors of bone, respectively [2, 3]. The effects of these mutations in bone cells are discussed below.
4.4.1 Chondroblastomas
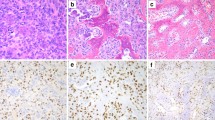
Chondroblastomas are aggressive rare bone tumors that present with pain in long bones. They are thought to arise from immature chondrocytes within secondary ossification centers in the epiphyses of long bones (Fig. 4.4). Chondroblastomas do not produce normal cartilage. Rather the matrix surrounding individual chondroblasts becomes calcified, producing a chicken-wire pattern on x-ray. Chondroblastomas are typically removed surgically by curettage. The rate of chondroblastoma recurrence ranges from 5% to 40%. The overall survival prognosis for a patient diagnosed with a chondroblastoma in early stages is 80–90%.
Approximately 95% of chondroblastomas contain a heterozygous mutation in H3F3B that replaces lysine 36 with methionine (K36M) in the histone variant H3.3 [3]. Mutant H3K36M molecules are integrated into the genome at sites of active transcription and produce global reductions in H3K36 di- and tri-methylation (HeK36me2 and H3K36me3) in human chondrocytes [21, 22]. Recurrent H3K36M mutations reprogram the transcriptome in chondroblastomas by binding with high affinity to HMTases, NSD2/MMSET and SETD2, and inhibiting their ability to methylate H3K36 (Fig. 4.3). H3K36M did not affect other HMTases, ASH1L and NSD1. Crystal structures of SETD2 bound to H3K36M or H3K36I peptides show that the mutant residues are positioned into the catalytic sites of SETD2 where they block enzymatic activity [40]. Elucidating the specific role of the H3K36me2 interaction in stabilizing NSD2 will be important in further understanding the function of NSD2 and the H3K36me2 modification under normal physiologic conditions and also in musculoskeletal cancers.
In addition to changing H3K36 methylation patterns, increases in H3K27me3 patterns were observed in cells expressing the mutant H3K36M, suggesting that the loss of H3K36 methylation provides a nucleosomal substrate for PRC2 [22]. The overall consequence is altered expression of genes that support chondrocytic proliferation, colony formation, cell survival, and DNA repair, along with suppression of genes (e.g., BMP2, RUNX2) controlling chondrocyte differentiation [21, 22]. It is difficult to precisely replicate this human disease in mice because of its natural anatomical location in developing skeletons, but subcutaneous injection of mesenchymal progenitor cells expressing H3K36M into mice generated undifferentiated sarcomas in mice. These data as a whole demonstrate that H3K36M expression is a driver of neoplasia in mesenchymal cells of the skeleton.
4.4.2 Giant Cell Tumors of Bone (GCTB)
Giant cell tumors of bone are rare but locally aggressive cancers that cause pain and swelling and can destroy surrounding bones and joints. Though typically benign, some GCTBs produce lung metastases, and high-grade sarcomas can form near the benign GCTB [41]. GCTBs usually occur in the epiphyses of the long bones within the appendicular skeleton (Fig. 4.4) and are diagnosed by X-ray or other imaging techniques. These rare cancers (one per one million people) typically form near the knee of young adults (aged 20–40 years) but are also found in the hips, shoulders, wrists, and lower back. Giant cell tumors on average have a 16% mortality rate [42], and treatment options include surgical resection or curettage followed by bone grafting. Radiation and other treatments that can damage the affected joint are reserved for cases where surgery is not possible.
GCTBs are heterogeneous and consist of three cell types. Mesenchymal cells of the osteoblast lineage are the neoplastic component of the tumor and express mutant H3.3 proteins, as well as osteoblast products, osteocalcin, and alkaline phosphatase. The other two cell types (mononuclear histiocytic cells and multinucleated giant cells) originate from hematopoietic progenitors, express CD68 but not mutated histones, and serve to support the mutant osteoblastic cells. The giant cells that give the disease its pathological identity form as a result of fusion between several individual mononuclear cells into a single, larger cells. These large cells resemble osteoclasts and cause bone resorption and destruction.
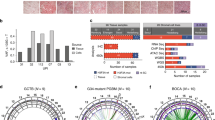
Somatic H3F3A mutations have been linked to over 92 percent of GCTBs [3] and are a molecular marker that separates GCTBs from other tumor types, including more malignant and metastatic bone tumors (e.g., osteosarcomas). The majority of mutations in H3F3A alter G34 in H3.3 to W, creating H3G34W, but other substitutions (G34V, G34R, and G34L) have also been discovered [43]. These mutations are created by single-base-pair changes in H3F3A that convert the W codon (GGG) to Arg/R (“AGG” and “CGG”), Trp/W (“TGG”), Val/V (“GTG”), Glu/E (“GAG”), or Ala/A (“GCG”) [21]. G34 itself is not posttranslationally modified, but it is a crucial residue for enzymatic processes that affect other nearby residues in histone tails, including H3K36 [44]. Reductions in H3K36me2 levels were noted in cells expressing G34W by chromatin immunoprecipitation assays, even though NSD2 was bound to the mutant chromatin [45]. Thus, G34 mutations in GCTBs likely inhibit lysine HMTases specific to H3K36 (NSD1, NSD2) to reduce global H3K36 methylation levels. RNA processing was also blocked by the G34W substitution in H3.3 [46]. Thus, the normal functions of H3.3 fail when there is an accumulation of H3.3G34W substitutions, which leads to hyper-proliferative activity and cancers in the epiphyses of long bones.
4.4.3 Other Bone Tumors
Since the discovery of mutant and oncogenic histones (oncohistones) in gliomas, chondroblastomas, and GCTBs, many other tumor genomes have been searched for histone gene variants. This effort identified additional skeletal tumors harboring such histone mutations but at very low relative frequencies. H3.1 K36M/I mutations were found in a case of pediatric undifferentiated mesenchymal sarcomas [22], and mutations in H3F3A and H3F3B that produce mutant H3.3G34R or H3.3G34W substitutions were found in less than one percent of osteosarcomas [43].
Osteosarcomas are malignant tumors of osteoblast origin that produce immature woven bone, which is mechanically weaker than normal lamellar bone. It is the most common type of cancer that arises in the skeleton and is usually found at the metaphyseal region of long bones. Most people diagnosed with osteosarcoma are under the age of 25 and are male. Patients with high-grade osteosarcoma in one location have a survival rate of about 68% [47]. Treatment usually includes a combination of surgery and chemotherapy. The presence of H3F3AG34W/R mutations in osteosarcomas is associated with epigenetic deregulation of oncogenic pathways such as PTEN [48]. The close relationship between H3F3A G34W/R mutant osteosarcomas and H3F3A G34W/L mutant GCBTs is consistent with a similar cellular origin from the osteoblastic lineage. More studies are necessary to determine how these H3F3AG34W/R mutations contribute to osteosarcoma pathogenesis and patient survival.
4.5 Therapeutic Opportunities for Bone Tumors Harboring Oncohistones
The discoveries that most chondroblastomas and GCTBs harbor histone mutations that block HMTase activity, alter histone methylation patterns, and promote cell proliferation and survival have not only allowed for stratification of tumor types but also stimulated discussion on alternative therapeutic options specifically for tumors expressing oncohistones. At least three strategies to eliminate these cancers exist: (1) eliminate the mutant H3F3A or H3F3B allele through selective gene editing or RNA editing; (2) prevent biochemical interactions between oncohistones and HMTases or HIRA proteins; and (3) modify the activity of genes whose expression are differentially expressed as a result of alterations in the histone methylome. The first two strategies will require further structural information on the histone genes and protein complexes involved, as well as identification of active and specific agents (either biomolecules or chemicals) and effective drug delivery mechanisms. The third strategy involves indirect targeting of the oncogenic driver but could be more rapid if existing drugs can be repurposed. Two promising examples of this strategy are described next.
Ribonucleotide reductase subunit M2 (RRM2) is essential for regulating cellular dNTP levels [49]. RRM2 expression declines in SETD2-deficient cells. The WEE1 kinase inhibitor, AZD1775, also reduces RRM2 levels and synergizes with SETD2-deficiency to further deplete cells of dNTP, leading to S phase arrest and cancer cell death. Thus, WEE1 inhibition can selectively kill SETD2- and H3K36me3-deficient cancers by starving them of deoxyribonucleotides. It remains to be determined whether or not WEE1 inhibitors are efficacious on musculoskeletal tumors with H3K36me3 deficiency.
Understanding how oncohistones change gene expression profiles has led to new a therapeutic strategy for high-grade gliomas that harbor H3K27M mutations. In these tumors there is a global reduction in H3K27me2 and H3K27me3 levels but focal increases in H3K27me3 in genes associated with cancer progression [50]. Many gliomas with the H3K27M mutations overexpress the dopamine receptor D2 (DRD2). A DRD2/3 antagonist, ONC201, that blocks oncogenic AKT/ERK signaling pathways and anti-apoptotic in these tumors is currently in clinical trials for high-grade midline brain tumors [51]. If successful, this drug that can penetrate the blood-brain barrier will provide a much-needed option for inoperable brain tumors that have a poor prognosis. Moreover, this approach could be used to identify new therapies for bone tumors.
4.6 Conclusion and Perspectives
Advances in genome-wide sequencing technologies have facilitated the discovery of somatic histone mutations (oncohistones) in bone and brain cancers. These genetic mutations have further improved our understanding of epigenetics and mechanistic links between tumor epigenomes and cancer progression. Until recently, epigenomic and genetic alterations have been considered separate mechanisms contributing in tumorigenesis, but genomic sequencing has shown mutations in histone genes can drastically change the epigenome, leading to global chromatin chaos and activation of carcinogenic pathways within cells. Much remains to be learned about how H3K36 and H3G34 mutations drive the formation of chondroblastomas and GCTBs, as well as osteosarcomas and other musculoskeletal tumors. Furthering this knowledge could produce new therapies that eliminate these painful cancers and prevent fatal metastases.
Abbreviations
- CAF-1:
-
Chromatin assembly factor 1
- DRD2:
-
Dopamine receptor D2
- GCTB:
-
Giant cell tumor of bone
- H3:
-
Histone 3
- HIRA:
-
Histone regulator A
- HMTase:
-
Histone methyltransferase
- K:
-
Lysine
- KDM:
-
Lysine demethylase
- KMT:
-
Lysine methylation transferase
- Me:
-
Methyl group
- MMSET:
-
Multiple myeloma SET domain
- NSD:
-
Nuclear receptor SET domain-containing
- PRC2:
-
Polycomb-repressive complex 2
- PRMT:
-
Protein arginine methyltransferases
- PTM:
-
Posttranslational modification
- RRM2:
-
Ribonucleotide reductase regulatory subunit M2
- SETD:
-
Su(var)3-9, Enhancer of Zeste, Trithorax domain-containing
- SRI:
-
Set2 Rpb1 interacting domain
- WHSC1:
-
Wolf-Hirschhorn syndrome gene (aka, NSD2)
References
Sankar S, Lessnick SL (2011) Promiscuous partnerships in Ewing’s sarcoma. Cancer Genet 204(7):351–365
Franchi A et al (2016) Histone 3.3 mutations in giant cell tumor and giant cell-rich sarcomas of bone. Lab Invest 96:17a–18a
Behjati S et al (2013) Distinct H3F3A and H3F3B driver mutations define chondroblastoma and giant cell tumor of bone. Nat Genet 45(12):1479–U105
Karsenty G, Kronenberg HM, Settembre C (2009) Genetic control of bone formation. Annu Rev Cell Dev Biol 25:629–648
Aghajanian P, Mohan S (2018) The art of building bone: emerging role of chondrocyte-to-osteoblast transdifferentiation in endochondral ossification. Bone Res 6:19
Siclari VA, Guise TA, Chirgwin JM (2006) Molecular interactions between breast cancer cells and the bone microenvironment drive skeletal metastases. Cancer Metastasis Rev 25(4):621–633
Kadyrova LY, Blanko ER, Kadyrov FA (2013) Human CAF-1-dependent nucleosome assembly in a defined system. Cell Cycle 12(20):3286–3297
Burgess RJ, Zhang Z (2013) Histone chaperones in nucleosome assembly and human disease. Nat Struct Mol Biol 20(1):14–22
Mohammad F, Helin K (2017) Oncohistones: drivers of pediatric cancers. Genes Dev 31(23-24):2313–2324
Jenuwein T, Allis CD (2001) Translating the histone code. Science 293(5532):1074–1080
Khoury GA, Baliban RC, Floudas CA (2011) Proteome-wide post-translational modification statistics: frequency analysis and curation of the swiss-prot database. Sci Rep 1:90
Rothbart SB, Strahl BD (2014) Interpreting the language of histone and DNA modifications. Biochim Biophys Acta 1839(8):627–643
Xu GG et al (2018) Signaling specificity in the c-di-GMP-dependent network regulating antibiotic synthesis in Lysobacter. Nucleic Acids Res 46(18):9276–9288
Serefidou M, Venkatasubramani AV, Imhof A (2019) The impact of one carbon metabolism on histone methylation. Front Genet 10:764
Greer EL, Shi Y (2012) Histone methylation: a dynamic mark in health, disease and inheritance. Nat Rev Genet 13(5):343–357
Joh RI et al (2014) Regulation of histone methylation by noncoding RNAs. Biochim Biophys Acta-Gene Regul Mech 1839(12):1385–1394
Benveniste D et al (2014) Transcription factor binding predicts histone modifications in human cell lines. Proc Natl Acad Sci U S A 111(37):13367–13372
Bannister AJ, Kouzarides T (2011) Regulation of chromatin by histone modifications. Cell Res 21(3):381–395
Bennett RL et al (2017) The role of nuclear receptor-binding SET domain family histone lysine methyltransferases in cancer. Cold Spring Harb Perspect Med 7(6):a026708
Wagner EJ, Carpenter PB (2012) Understanding the language of Lys36 methylation at histone H3. Nat Rev Mol Cell Biol 13(2):115–126
Fang D et al (2016) The histone H3.3K36M mutation reprograms the epigenome of chondroblastomas. Science 352(6291):1344–1348
Lu C et al (2016) Histone H3K36 mutations promote sarcomagenesis through altered histone methylation landscape. Science 352(6287):844–849
Kuo AJ et al (2011) NSD2 links Dimethylation of histone H3 at lysine 36 to oncogenic programming. Mol Cell 44(4):609–620
Chesi M et al (1998) The t(4;14) translocation in myeloma dysregulates both FGFR3 and a novel gene, MMSET, resulting in IgH/MMSET hybrid transcripts. Blood 92(9):3025–3034
Martinez-Garcia E et al (2011) The MMSET histone methyl transferase switches global histone methylation and alters gene expression in t(4;14) multiple myeloma cells. Blood 117(1):211–220
Roy DM, Walsh LA, Chan TA (2014) Driver mutations of cancer epigenomes. Protein Cell 5(4):265–296
Sun XJ et al (2005) Identification and characterization of a novel human histone H3 lysine 36-specific methyltransferase. J Biol Chem 280(42):35261–35271
Mohammad F et al (2017) EZH2 is a potential therapeutic target for H3K27M-mutant pediatric gliomas. Nat Med 23(4):483
Macias MJ et al (1996) Structure of the WW domain of a kinase-associated protein complexed with a proline-rich peptide. Nature 382(6592):646–649
Lu PJ et al (1999) Function of WW domains as phosphoserine- or phosphothreonine-binding modules. Science 283(5406):1325–1328
Sze CI et al (2004) Down-regulation of WW domain-containing oxidoreductase induces Tau phosphorylation in vitro – a potential role in Alzheimer’s disease. J Biol Chem 279(29):30498–30506
Yendamuri S et al (2003) WW domain containing oxidoreductase gene expression is altered in non-small cell lung cancer. Cancer Res 63(4):878–881
Li J et al (2016) SETD2: an epigenetic modifier with tumor suppressor functionality. Oncotarget 7(31):50719–50734
Kizer KO et al (2005) A novel domain in Set2 mediates RNA polymerase II interaction and couples histone H3 K36 methylation with transcript elongation. Mol Cell Biol 25(8):3305–3316
McDaniel SL, Strahl BD (2017) Shaping the cellular landscape with Set2/SETD2 methylation. Cell Mol Life Sci 74(18):3317–3334
Newbold RF, Mokbel K (2010) Evidence for a tumour suppressor function of SETD2 in human breast cancer: a new hypothesis. Anticancer Res 30(9):3309–3311
Duns G et al (2010) Histone methyltransferase gene SETD2 is a novel tumor suppressor gene in clear cell renal cell carcinoma. Cancer Res 70(11):4287–4291
Linne H et al (2017) Functional role of SETD2, BAP1, PARP-3 and PBRM1 candidate genes on the regulation of hTERT gene expression. Oncotarget 8(37):61890–61900
Wan YCE, Liu J, Chan KM (2018) Histone H3 mutations in cancer. Curr Pharmacol Rep 4(4):292–300
Yang S et al (2016) Molecular basis for oncohistone H3 recognition by SETD2 methyltransferase. Genes Dev 30(14):1611–1616
Palmerini E et al (2019) Malignancy in giant cell tumor of bone: a review of the literature. Technol Cancer Res Treat 18:1533033819840000
Domovitov SV, Healey JH (2010) Primary malignant Giant-cell tumor of bone has high survival rate. Ann Surg Oncol 17(3):694–701
Behjati S et al (2014) Distinct H3F3A and H3F3B driver mutations define chondroblastoma and giant cell tumor of bone (vol 45, pg 1479, 2013). Nat Genet 46(3):316–316
Wojcik J, Cooper K (2017) Epigenetic alterations in bone and soft tissue tumors. Adv Anat Pathol 24(6):362–371
Shi L et al (2018) Histone H3.3 G34 mutations Alter histone H3K36 and H3K27 methylation in cis. J Mol Biol 430(11):1562–1565
Lim J et al (2017) The histone variant H3.3 G34W substitution in giant cell tumor of the bone link chromatin and RNA processing. Sci Rep 7(1):13459
Ottaviani G, Jaffe N (2009) The epidemiology of osteosarcoma. Cancer Treat Res 152:3–13
Koelsche C et al (2017) Histone 3.3 hotspot mutations in conventional osteosarcomas: a comprehensive clinical and molecular characterization of six H3F3A mutated cases. Clin Sarcoma Res 7:9
Pfister SX et al (2015) Inhibiting WEE1 selectively kills histone H3K36me3-deficient cancers by dNTP starvation. Cancer Cell 28(5):557–568
Chan KM et al (2013) Neuro-Oncology 15:176–177
Ralff MD et al (2017) ONC201: a new treatment option being tested clinically for recurrent glioblastoma. Transl Cancer Res 6:S239–S243
Disclosures
The authors have no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Taylor, E.L., Westendorf, J.J. (2021). Histone Mutations and Bone Cancers. In: Fang, D., Han, J. (eds) Histone Mutations and Cancer. Advances in Experimental Medicine and Biology, vol 1283. Springer, Singapore. https://doi.org/10.1007/978-981-15-8104-5_4
Download citation
DOI: https://doi.org/10.1007/978-981-15-8104-5_4
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-8103-8
Online ISBN: 978-981-15-8104-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)