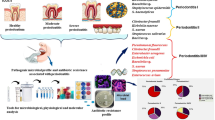
Abstract
Objectives
The aim of this study was to analyze the distribution of β-lactamase genes and the multidrug resistance profiles in β-lactam-resistant subgingival bacteria from patients with periodontitis.
Materials and methods
Subgingival samples were obtained from 130 Spanish patients with generalized periodontitis stage III or IV. Samples were grown on agar plates with amoxicillin or cefotaxime and incubated in anaerobic and microaerophilic conditions. Isolates were identified to the species level by the sequencing of their 16S rRNA gene. A screening for the following β-lactamase genes was performed by the polymerase chain reaction (PCR) technique: blaTEM, blaSHV, blaCTX-M, blaCfxA, blaCepA, blaCblA, and blaampC. Additionally, multidrug resistance to tetracycline, chloramphenicol, streptomycin, erythromycin, and kanamycin was assessed, growing the isolates on agar plates with breakpoint concentrations of each antimicrobial.
Results
β-lactam-resistant isolates were found in 83% of the patients. Seven hundred and thirty-seven isolates from 35 different genera were obtained, with Prevotella and Streptococcus being the most identified genera. blaCfxA was the gene most detected, being observed in 24.8% of the isolates, followed by blaTEM (12.9%). Most of the isolates (81.3%) were multidrug-resistant.
Conclusions
This study shows that β-lactam resistance is widespread among Spanish patients with periodontitis. Furthermore, it suggests that the subgingival commensal microbiota might be a reservoir of multidrug resistance and β-lactamase genes.
Clinical relevance
Most of the samples yielded β-lactam-resistant isolates, and 4 different groups of bla genes were detected among the isolates. Most of the isolates were also multidrug-resistant. The results show that, although β-lactams may still be effective, their future might be hindered by the presence of β-lactam-resistant bacteria and the presence of transferable bla genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Periodontitis is a complex infectious disease caused by a dysbiosis of the subgingival biofilm and a disproportionate response of the host’s immune system [1]. Supra- and subgingival debridement along with motivation and oral hygiene instructions are the standard treatment for periodontitis, and antimicrobials can be used as adjunctive therapy if patient’s conditions apply [2,3,4]. The microbial etiology of inflammatory periodontal diseases provides the rationale for the use of antimicrobial medication in periodontal therapy. Antibiotics may be specially indicated for periodontal patients 36 years of age or younger with periodontitis stage II or for patients with attachment or radiographic bone loss at more than two nonadjacent sites [5]. The most used antimicrobials in periodontitis are the β-lactams, particularly amoxicillin (AMX) which can be administered together with metronidazole [2]. It has been described that those bacteria resistant to β-lactams may also be resistant to other antibiotics such as tetracyclines, aminoglycosides, and chloramphenicol [6,7,8]. The most important resistance mechanism to β-lactam antibiotics are the β-lactamases, enzymes that break the β-lactam ring and inactivate the antimicrobial [9]. Extended spectrum β-lactamases (ESBLs) can hydrolyze a wide array of β-lactams such as penicillins, cephalosporins, and monobactams while inhibited by β-lactamase inhibitors such as clavulanic acid [8, 10, 11]. ESBLs were first detected in the 1980s in a Klebsiella pneumoniae isolate, and now these are found in both enteric and non-enteric microorganisms. More than 230 ESBLs have been described, including TEM-type, SHV-type, CTX-M-type, OXA-type, and KPC-type [11]. In the oral environment, blaTEM have been described as the most prevalent ESBLs, although there are only a few studies that have screened for these genes [12,13,14,15]. Other β-lactamases found in the oral biofilm include blaCfxA, blaCepA/CblA, and blaampC, suggesting that the mouth might be a reservoir for β-lactamase genes. In fact, the oral environment, being a transit place for bacteria entering the digestive tract and harboring more than 700 bacterial species that grow to form a biofilm, offers an excellent opportunity for horizontal gene transfer to occur [16, 17].
Spain is one of the European countries with the highest consumption of antibiotics, which has been suggested to be linked with the increase of antibiotic resistance [18]. Little is known about the distribution of β-lactamase genes in the subgingival microbiota of Spanish patients with periodontitis. For this reason, the aim of the current study was to screen for β-lactamase genes in the AMX- and cefotaxime-resistant subgingival microbiota isolated from 130 patients with periodontitis. Additionally, these isolates were tested for their multidrug resistance (MDR) to chloramphenicol, tetracycline, kanamycin, erythromycin, and streptomycin.
Materials and methods
Patients involved in the study
Subgingival samples were taken consecutively from 130 patients, recruited between 2016 and 2017, and diagnosed with generalized severe chronic periodontitis or generalized aggressive periodontitis according to the 1999 classification [19]. Nowadays, this diagnosis corresponds to generalized periodontitis stage III or IV, according to the new classification [20]. Stages were assessed according to the interproximal attachment loss. Grading was established considering the coefficient radiographic bone loss and patients age, adding the status of the systemic conditions such as smoking and diabetes. The samples were obtained from patients that attended at the Department of Periodontology of the Universitat Internacional de Catalunya (UIC) (Barcelona, Spain). All patients were supervised by the same clinician (CM), who also took the microbial samples. The study was previously approved by the Ethics Committee of the UIC (Study number: ODO-2014-01) and complied with the principles of the Declaration of Helsinki. Additionally, all the participants signed an Institutional Review Board approved informed consent form. None of the patients had taken antibiotics at least 3 months prior to the sampling.
Clinical evaluation
The following clinical parameters were collected from a whole-mouth evaluation at baseline: probing pocket depth (PPD), clinical attachment level (CAL), full mouth plaque index (FMPI), full mouth bleeding on probing (FMBP), and mobility and furcation involvement. The PPD was measured at 6 sites per tooth (mesiobuccal, mid-buccal, distobuccal, mesiolingual, mid-lingual, and distolingual) as the distance in millimeters from the free gingival margin to the base of the probeable pocket using a handheld periodontal probe (PCP-UNC 15; Hu-Friedy Mfg. Co., Chicago, IL, USA). The CAL was measured at 6 sites per tooth from the cemento-enamel junction (CEJ) or from the base of the dental restoration or prosthesis to the bottom of the pocket. The plaque index (PI) was recorded according to the criteria described by Silness and Löe [21]. Bleeding on probing (BOP) was determined as being present or absent (±) within 30 s after probing of the aforementioned 6 sites per tooth. Only the parameters of PPD, CAL, PI, BOP, and mobility of the selected teeth, those with deepest PPD of each quadrant, were evaluated in the study.
Sample collection
Subgingival microbial samples were taken from the deepest periodontal pocket of each quadrant. Each area was isolated with cotton rolls, the supragingival plaque deposits were carefully removed with curettes, and subgingival microbial samples were obtained by inserting two sterile paper points in each subgingival pocket and keeping them in place for 20 s. Samples from each patient were pooled in a vial containing 1.5 ml of cold sterilized reduced transport medium without ethylenediaminetetraacetic acid (EDTA) [22] and sent to the microbiology laboratory at 4 °C for processing within the same day.
Microbial culture and bacterial selection
Subgingival biofilm samples were dispersed by vortex for 45 s. Serial tenfold dilutions were plated on blood agar (blood agar base no. 2; Oxoid Ltd., Basingstoke, UK) containing 5% of horse blood, hemin (5 mg/l), menadione (1 mg/l), and on the same media with 2 μg/ml of cefotaxime (CTX) or 8 μg/ml of AMX (Sigma Aldrich, St. Louis, MO, USA). Both concentrations were selected based on the breakpoint concentrations recommended by the European Committee on Antimicrobial Susceptibility Testing (EUCAST) and the Clinical and Laboratory Standards Institute (CLSI) [23, 24]. Given that the oral microbiota harbors many different bacterial genera and that many of these are not mentioned in either the EUCAST or the CLSI guidelines, these concentrations were chosen based on taxonomic relatedness to bacteria of the oral environment, using the higher concentration of antibiotics when in doubt. Therefore, bacteria that grew on plates with antibiotics were considered to be resistant. In order to obtain a wider array of the subgingival microbiota, plates were incubated at 37 °C under microaerophilic (5% CO2) and anaerobic (10% H2, 10% CO2, and 80% N2) conditions for 48–72 h. Resistant colonies were isolated according to their morphology (two of each morphology), replated to obtain pure cultures, and preserved at − 80 °C in a 30% sterilized glycerol solution.
DNA isolation and 16S rRNA gene sequencing
Genomic DNA extraction was performed on each isolate using the ATP™ Genomic DNA mini Kit (ATP Biotech Inc., Taipei City, Taiwan) following the manufacturer’s instructions. Once extracted, DNA was visualized in a 0.7% agarose gel with ethidium bromide and quantified using a Nanodrop 2000C UV-vis spectrophotometer (Nanodrop Technologies, Wilmington, DE, USA).
Isolates were identified to a species level, through 16S rRNA gene sequencing [25]. Zero point five (0.5) micromolar of primers 27F and 1544R (Table 1) were added to the polymerase chain reaction (PCR) mix with 30–100 ng of DNA, 1X PCR buffer, 1X dNTPs solution, 2.5 mM of MgCl2, and 1 unit of Taq polymerase (all reagents from Takara, Tokyo, Japan). Amplification was carried out using a T3000 Thermocycler (Biometra, Goettingen, Germany) under the following conditions: 5 min at 95 °C, followed by 35 cycles of 95 °C for 60 s, 57 °C for 60 s, and 72 °C for 60 s, and followed by 10 min at 72 °C. The PCR products were purified using the E.Z.N.A Gel Extraction Kit (Omega Bio-Tek, Norcross, GA, USA) and sent for sequencing to Macrogen Inc. (Amsterdam, Netherlands). The sequences obtained were aligned using Clustal Omega Software and analyzed using NCBI’s BLAST (available at http://www.ebi.acuk/ and https://blast.ncbi.nlm.nih.gov/Blast.cgi, respectively). Only sequences with ≥ 99% identity were accepted as proof to identify to the species level.
Detection of β-lactamase genes
Detection of β-lactamase genes was achieved using the primers described in Table 1. To ensure that most of the genetic variants would be detected in the screening, universal primers were used in the detection of blaCfxA, blaTEM, blaSHV, and blaCTX-M. PCR reactions were performed with 30–100 ng of DNA, 1X PCR buffer, 1X dNTPs solution, 2.5 mM MgCl2, 0.5 μM of each primer, and 1 unit Taq polymerase (Takara, Tokyo, Japan). Amplifications were carried out in a T3000 Thermocycler. The thermocycling conditions were as follows: (i) for the detection of blaCfxA, 1 min of denaturation at 95 °C, 1 min of annealing at 58 °C, and 1 min of extension at 72 °C for 26 cycles; (ii) for the detection of blaCepA/CblA, 30 s of denaturation at 95 °C, 30 s of annealing at 58 °C, and 30 s of extension at 72 °C for 29 cycles; (iii) for the detection of blaampC, 1 min of denaturation at 95 °C, 1 min of annealing at 50 °C, and 1 min of extension at 72 °C for 30 cycles; (iv) for the detection of blaOXA, 40 s of denaturation at 95 °C, 40 s of annealing at 55 °C, and 40 s of extension at 72 °C for 30 cycles; (v) for the detection of blaTEM, blaSHV, and blaCTX-M, a multiplex was performed as previously described [31]. In all reactions an initial, 5-min denaturation at 95 °C and a final 10 min extension at 72 °C were applied. DNA of the isolates carrying the genes screened was used as positive controls. To confirm the presence of the genes in the controls, PCR amplicons of the expected size were sequenced and annealed with reference sequences available at NCBI’s GenBank. Negative controls included water in place of DNA in the PCR mix.
PCR products were observed through electrophoresis using a 2% agarose gel with ethidium bromide. Gels were photographed using a UV light transilluminator GEL DOC™ XR+ system (Bio-Rad Laboratories Inc. Hercules, CA, USA).
In vitro antimicrobial resistance testing
MDR was determined for all isolates using blood agar plates containing 5% of horse blood, hemin (5 mg/l), and menadione (1 mg/l) and supplemented with (i) 1 μg/ml of erythromycin (ERY), (ii) 64 μg/ml of kanamycin (KAN), (iii) 8 μg/ml of chloramphenicol (CHL), (iv) 128 μg/ml of streptomycin (STR), (v) 8 μg/ml of tetracycline (TET), and (vi) 2 μg/ml of CTX or 8 μg/ml of AMX (all antimicrobials were obtained as pure powder from Sigma Aldrich, St. Louis, MO, USA). Incubation was performed at 37 °C in anaerobic and microaerophilic conditions for 48–72 h. The antibiotic concentrations used were set according to the recommendations provided by the EUCAST and CLSI and following the same criteria used for the initial selection for AMX and CTX.
Statistical analysis
The chi-square test was used as a paired statistical test for discrete variables to find statistical differences between the prevalence of the genes screened and phenotypical resistance to the antibiotics tested among the identified bacterial species of this study. p values of < 0.05 were accepted for statistical significance.
Results
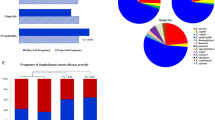
One hundred and thirty subgingival samples were obtained from patients with generalized periodontitis stage III or IV. Patients were between the ages of 24 and 82 years old (mean of 51.3) and showed a mean probing depth of 6.6 ± 1.8 mm, a mean clinical attachment loss of 7.6 ± 2.1 mm, and 92.1% of gingival sites with bleeding on probing. From the 130 samples, 63 (48.5%) and 33 (25.4%) samples had AMX-resistant isolates in anaerobic (ARIA) and in microaerophilic (ARIM) conditions, respectively, while 98 (75.4%) and 70 (53.6%) samples had CTX-resistant isolates in anaerobic (CRIA) and in microaerophilic (CRIM) conditions, respectively. Twenty-eight (21.5%) samples grown in anaerobiosis and 53 (40.8%) samples grown in microaerophilic conditions did not present any β-lactam-resistant isolates. Twenty-two samples (16.9%) did not present any β-lactam-resistant isolate in either microaerophilia or anaerobiosis. Samples grown in blood agar without antibiotics showed a mean bacterial load of 7.3 log10 colony-forming units per milliliter (cfu/ml) (± 1.3) in anaerobic conditions and 7.1 log10 cfu/ml (± 0.9) in microaerophilic conditions. The number of β-lactam-resistant bacteria averaged a 1 logarithmic reduction (Table 2).
The subgingival samples yielded 181 ARIAs, 84 ARIMs, 293 CRIAs, and 179 CRIMs, making a total of 737 isolates. Isolated bacteria were identified by the sequencing of their 16S rRNA gene, and β-lactam resistance genes were screened by PCR. Additionally, phenotypic resistance to CHL, STR, ERY, TET, and KAN was determined. Furthermore, AMX and CTX resistance was also tested depending on which antibiotic the isolates were selected for (Table 3). Data regarding the species detected, their phenotypic resistance, and their screened genes is shown in the Supplementary data (Table S1). Bacteria from the genus Prevotella were the most frequently isolated (n = 213), mainly the species Prevotella nigrescens (n = 70) and Prevotella intermedia (n = 65), followed by the genus Streptococcus (n = 153) with a wide variety of species within this genus (see Table S1). The genus Prevotella was significantly (p < 0.01) more prevalent in the ARIAs, while the genus Veillonella was more prevalent in the CRIAs (p < 0.01). Regarding bacteria grown in microaerophilic conditions, the genus Neisseria was significantly more prevalent in ARIMs (p < 0.05), while the genus Micrococcus was more prevalent in CRIMs (p < 0.01). More than 12 % (12.1%) of the isolates were identified at a genus level, and 8% could not be identified at all (Tables 3 and S1).
Regarding the bla genes, blaCfxA was the most prevalent gene (24.8%), followed by blaTEM (12.9%), blaCepA/CblA (1.1%), and blaSHV (0.8%). The blaCTX-M, blaOXA, and blaampC genes were not detected. A significantly higher (p < 0.01) percentage of blaCfxA was observed in those isolates grown in anaerobic conditions compared to those grown in microaerophilic conditions.
Phenotypic resistance to other antibiotics was observed in 599 isolates (81.3%). These isolates were resistant to, at least, one antimicrobial besides the β-lactam for which they were previously selected. Fifty-six point eighty-five percent (56.9%) of the isolates showed resistance to KAN, 54.6% to ERY, 29.4% to TET, 27.7% to STR, and 9.9% to CHL. Isolates selected for AMX resistance showed resistance to CTX more frequently (53.6%) than conversely (32.2%). Most of the MDR isolates were resistant to 1–3 antimicrobials (22.2%, 26.4%, and 25.7%, respectively), and 15 isolates (2.5%) were resistant to the 6 antimicrobials tested (Table 4). Statistical differences were observed between the percentage of ARIAs and ARIMs resistant to CHL and KAN and between the percentage of CRIAs and CRIMs resistant to AMX and KAN.
Discussion
This study analyzed the β-lactam-resistant subgingival microbiota isolated from patients with severe forms of periodontitis. Bacteria were isolated based on their resistance to AMX or to CTX. AMX was chosen for being the first-choice β-lactam for many bacterial infections including periodontitis [2, 32, 33]. The use of CTX, as a third-generation cephalosporin, is much more restricted, and therefore, resistance to this antibiotic should be scarcer. However, ESBLs such as CTX-M, which are active against it, are prevalent worldwide [34, 35], making them interesting targets to study when analyzing β-lactam resistance. An average of 7.8% of the culturable microbiota isolated in this study was resistant to AMX, slightly higher that what a previous study found in Spanish samples comparing subgingival microbiota from Spain and the Netherlands [36]. In our study population, CTX resistance turned out to be more prevalent, with 82.3% of the patients having at least one resistant isolate, than AMX resistance (55.4%). We detected a large amount of Veillonella isolates resistant to CTX (n = 76) and very few resistant to AMX (n = 5), which might be an explanation for the higher prevalence of CTX resistance. Previous studies have already shown that the genus Veillonella has a low susceptibility for β-lactams, probably due to the presence of penicillin-binding proteins with lower affinity for β-lactams [37,38,39,40]. Furthermore, other studies analyzed Veillonella isolates and found higher MICs of cefoxitin than AMX and ampicillin [38, 39], suggesting that Veillonella spp. might be more resistant to cephalosporins than aminopenicillins. P. nigrescens (n = 70), P. intermedia (n = 65), and Veillonella parvula (n = 60) were the species most often isolated. This agrees with previous reports of Prevotella being the main genus in the oral environment expressing β-lactam resistance [41,42,43]. Most of the studies analyzing β-lactam resistance have based their selection of isolates on the production of β-lactamases, and therefore, despite being β-lactam-resistant, the Veillonella genus has been ruled out because it does not produce β-lactamases [37, 39, 44].
In this study, MDR was analyzed in each isolate using breakpoint concentrations suggested by the EUCAST and the CLSI. All the isolates that grew on agar plates with antibiotics were considered resistant. Of the 737 β-lactam-resistant isolates, 81.3% were MDR. From these, 63.3% were selected for CTX resistance and 36.7% for AMX resistance, these values being very similar to the percentage of total CTX- and AMX-resistant isolates obtained (64% and 35%, respectively) with little difference regarding whether they were cultured in anaerobic or in microaerophilic conditions; this suggests that MDR does not depend on the initial AMX or CTX selection. Few differences were observed between the 4 groups of isolates, except for KAN, CHL, and AMX resistance. Kanamycin resistance showed a higher prevalence in anaerobic isolates due mainly to the inherent resistance of anaerobic bacteria to this antibiotic [45] and to the Prevotella isolates, which are known to present a higher tolerance to kanamycin than other oral bacteria [46]. Chloramphenicol resistance was observed in higher percentages in microaerophilic isolates due mainly to the Pseudomonas isolates, which are able to exhibit high resistance levels thanks to their multidrug efflux pumps [47, 48]. On the other hand, AMX resistance was less often observed in the CTX microaerophilic isolates, probably due to the lack of Prevotella isolates, which increased the prevalence of AMX resistance in the CTX-resistant anaerobic group.
Fifteen isolates were resistant to the 7 antibiotics tested in this study and were all obtained from different patients. The isolates identified were V. parvula (n = 3), Pseudomonas aeruginosa (n = 2), Acinetobacter guillouiae (n = 3), and Stenotrophomonas maltophilia (n = 1). Of the 15 isolates, 6 could not be identified through 16S rRNA gene sequencing. It has been reported that Pseudomonas, Acinetobacter, and Stenotrophomonas genera are rich in efflux pumps, which confer on them resistance to multiple antimicrobials [49]. Furthermore, tetracycline, kanamycin, and erythromycin resistance can be linked to the presence of transposons of the Tn916/1545 family, which are ubiquitous in the oral microbiota and carry genes that confer resistance to these antibiotics [50, 51]. These transposons have been described in Veillonella [52, 53], which might be responsible for the observed MDR.
The blaCfxA gene was detected in 24.8% of the isolates, of which 71.0% (n = 136) were bacteria from the Prevotella genus. This genus has been previously associated with the blaCfxA gene, acquiring resistance to a variety of penicillins and cephalosporins [54]. As previously reported by other studies [12, 13, 55], we observed a high prevalence of blaCfxA in isolates of the Capnocytophaga genus (83.3%). To our knowledge, this is the first report of blaCfxA in the Staphylococcus, Alloprevotella, Streptococcus, and Veillonella genera. Since oral biofilm is a favorable environment for horizontal gene transfer [56, 57], it might be possible for blaCfxA to have been transferred from the Prevotella or Capnocytophaga genera, which are usual carriers of the gene [13]. The blaTEM gene was found in 23.1% of the samples and in 12.9% of the isolates, mainly in Veillonella spp. and Prevotella spp. The prevalence of this gene in our samples was low when compared to the study conducted by Ioannidis et al. which observed a prevalence of between 46.2 and 72.7% of blaTEM in subgingival and tongue samples from Greek subjects [58]. These differences might be related to the higher consumption of β-lactams by the Greek population according to the European Centre for Disease Prevention and Control [59], or methodologically related, since the detection of blaTEM was done from a pool of subgingival bacteria in each sample, regardless of their resistance patterns or ability to grow in isolation. The detection of the genes blaTEM, blaSHV, and blaCfxA in streptococci is noteworthy, given that there is some controversy about the presence of bla genes in this genus. Although it has been suggested that streptococci are unable to acquire foreign bla genes [60], at least two studies have reported the presence of these genes in Streptococcus pneumoniae [61, 62]. The detection of blaTEM by Ding et al. was questioned due to possible contamination of the Taq polymerase used in the reaction [63]; however, in our study, the negative controls did not suggest the presence of any contamination. Therefore, it would be interesting to conduct further studies to analyze the presence of bla genes among oral streptococci. Both blaCepA/CblA and blaSHV were found in low numbers as observed by the previous studies [13, 43, 64], suggesting that they may not play a critical role in β-lactam resistance in the oral environment.
With the data obtained in this study, we conclude that β-lactam resistance is widespread among the subgingival bacteria of Spanish patients with periodontitis. Prevotella, Veillonella, and Streptococcus were the genera with the highest number of β-lactam-resistant isolates, suggesting that in this population, oral commensal microbiota might be a reservoir of β-lactam resistance. Of special importance is the presence of β-lactamases that are coded in transferrable genes such as blaCfxA and blaTEM, which could transfer to other oral or transient bacteria. Moreover, a high prevalence of MDR was observed, constraining the number of antibiotics available against bacterial infections, where these to be needed. Despite the reports warning about increasing antimicrobial resistance, antimicrobials are usually prescribed in the clinical practice without studying microbial profiles and without performing antibiograms, a practice that favors the spread of antimicrobial resistance. The above, together with the high percentages of β-lactam resistance observed in this study, underlines the risk of a currently successful antibiotic treatment becoming ineffective.
References
Meyle J (2000) Chapple I (2015) molecular aspects of the pathogenesis of periodontitis. Periodontol 69:7–17. https://doi.org/10.1111/prd.12104
Herrera D, Sanz M, Jepsen S, Needleman I, Roldán S (2002) A systematic review on the effect of systemic antimicrobials as an adjunct to scaling and root planing in periodontitis patients. J Clin Periodontol 29:136–159. https://doi.org/10.1034/j.1600-051X.29.s3.8.x
Matesanz-Pérez P, García-Gargallo M, Figuero E, Bascones-Martínez A, Sanz M, Herrera D (2013) A systematic review on the effects of local antimicrobials as adjuncts to subgingival debridement, compared with subgingival debridement alone, in the treatment of chronic periodontitis. J Clin Periodontol 40:227–241. https://doi.org/10.1111/jcpe.12026
Rabelo CC, Feres M, Gonçalves C, Faver M, Tu Y-K, Chambrone L (2015) Systemic antibiotics in the treatment of aggressive periodontitis. A systematic review and a Bayesian Network meta-analysis. J Clin Periodontol 42:647–657. https://doi.org/10.1111/jcpe.12427
Pretzl B, Sälzer S, Ehmke B, Schlagenhauf U, Dannewitz B, Dommisch H, Eickholz P, Jockel-Schneider Y (2019) Administration of systemic antibiotics during non-surgical periodontal therapy—a consensus report. Clin Oral Investig 23:3073–3085. https://doi.org/10.1007/s00784-018-2727-0
Handal T, Olsen I, Walker CB, Caugant DA (2004) β-Lactamase production and antimicrobial susceptibility of subgingival bacteria from refractory periodontitis. Oral Microbiol Immunol 19:303–308. https://doi.org/10.1111/j.1399-302x.2004.00159.x
Jacoby GA (2009) AmpC β-Lactamases. Clin Microbiol Rev 22:161–182. https://doi.org/10.1128/CMR.00036-08
Perez F, Endimiani A, Hujer K, Bonomo R (2007) The continuing challenge of ESBLs. Curr Opin Pharmacol 7:459–469. https://doi.org/10.1016/j.coph.2007.08.003
Danziger LH, Pendland SL (1995) Bacterial resistance to β-lactam antibiotics. Am J Heal Pharm 52:S3–S8. https://doi.org/10.1093/ajhp/52.6_Suppl_2.S3
Lee JH, Bae IK, Hee Lee S (2012) New definitions of extended-spectrum β-lactamase conferring worldwide emerging antibiotic resistance. Med Res Rev 32:216–232. https://doi.org/10.1002/med.20210
Turner PJ (2005) Extended-spectrum β-lactamases. Clin Infect Dis 41:S273–S275. https://doi.org/10.1086/430789
Handal T, Giraud-Morin C, Caugant DA, Madinier I, Olsen I, Foose T (2005) Chromosome- and plasmid-encoded β-lactamases in Capnocytophaga spp. Antimicrob Agents Chemother 49:3940–3943. https://doi.org/10.1128/AAC.49.9.3940-3943.2005
Handal T, Olsen I, Walker CB, Caugant DA (2005) Detection and characterization of β-lactamase genes in subgingival bacteria from patients with refractory periodontitis. FEMS Microbiol Lett 242:319–324. https://doi.org/10.1016/j.femsle.2004.11.023
Kim S-M, Kim HC, Lee S-WS (2011) Characterization of antibiotic resistance determinants in oral biofilms. J Microbiol 49:595–602. https://doi.org/10.1007/s12275-011-0519-1
Søraas A, Olsen I, Sundsfjord A, Handal T, Bjørang O, Jenum PA (2014) Extended-spectrum beta-lactamase-producing bacteria are not detected in supragingival plaque samples from human fecal carriers of ESBL-producing Enterobacteriaceae. J Oral Microbiol 6:24026. https://doi.org/10.3402/jom.v6.24026
Olsen I, Tribble GD, Fiehn NE, Wang BY (2013) Bacterial sex in dental plaque. J Oral Microbiol 5:1. https://doi.org/10.3402/jom.v5i0.20736
Roberts AP, Mullany P (2010) Oral biofilms: a reservoir of transferable, bacterial, antimicrobial resistance. Expert Rev Anti-Infect Ther 8:1441–1450. https://doi.org/10.1586/eri.10.106
ECDC (European Centre for Disease Prevention and Control), EFSA (European Food Safety Authority), and EMA (European Medicines Agency) (2017) ECDC/EFSA/EMA second joint report on the integrated analysis of the consumption of antimicrobial agents and occurrence of antimicrobial resistance in bacteria from humans and food-producing animals - joint interagency antimicrobial consumption and resistance analysis (JIACRA) report. EFSA J 15:4872–5007. https://doi.org/10.2903/j.efsa.2017.4872
Armitage GC (1999) Development of a classification system for periodontal diseases and conditions. Ann Periodontol 4:1–6. https://doi.org/10.1902/annals.1999.4.1.1
Caton JG, Armitage G, Berglundh T, Chapple ILC, Jepsen S, Kornman KS, Mealey BL, Papanou PN, Sanz M, Tonetti MS (2018) A new classification scheme for periodontal and peri-implant diseases and conditions - introduction and key changes from the 1999 classification. J Clin Periodontol 45:S1–S8. https://doi.org/10.1111/jcpe.12935
Silness J, Löe H (1964) Periodontal disease in pregnancy II. Correlation between oral hygiene and periodontal condition. Acta Odontol Scand 22:121–135. https://doi.org/10.3109/00016356408993968
Loesche WJ, Syed SA, Stoll J (1987) Trypsin-like activity in subgingival plaque. J Periodontol 58:266–273. https://doi.org/10.1902/jop.1987.58.4.266
CLSI (2017) Performance standards for antimicrobial susceptibility testing (27th ed). Wayne, PA
The European Committee on Antimicrobial Susceptibility Testing (EUCAST) (2018) Breakpoint tables for interpretation of MICs and zone diameters, version 8.0. http://www.eucast.org/clinical_breakpoints
Shigematsu T, Hayashi M, Kikuchi I, Ueno S, Masaki H, Fujii T (2009) A culture-dependent bacterial community structure analysis based on liquid cultivation and its application to a marine environment. FEMS Microbiol Lett 293:240–247. https://doi.org/10.1111/j.1574-6968.2009.01536.x
Fosse T, Madinier I, Hannoun L, Giraud-Morin C, Hitzig C, Charbit Y, Ourang S (2002) High prevalence of cfxA β-lactamase in aminopenicillin-resistant Prevotella strains isolated from periodontal pockets. Oral Microbiol Immunol 17:85–88. https://doi.org/10.1046/j.0902-0055.2001.00096.x
Bou G, Martinez-Beltran J (2000) Cloning, nucleotide sequencing, and analysis of the gene encoding an AmpC β-lactamase in Acinetobacter baumannii. Antimicrob Agents Chemother 44:428–432. https://doi.org/10.1128/AAC.44.2.428-432.2000
Danel F, Hall LM, Gur D, Livermore DM (1995) OXA-14, another extended-spectrum variant of OXA-10 (PSE-2) β-lactamase from Pseudomonas aeruginosa. Antimicrob Agents Chemother 39:1881–1884. https://doi.org/10.1128/AAC.39.8.1881
Paterson DL, Hujer KM, Hujer AM, Yeiser B, Bonomo MD, Rice LB, Bonomo RA, International Klebsiella Study Group (2003) Extended-spectrum β-lactamases in Klebsiella pneumoniae bloodstream isolates from seven countries: dominance and widespread prevalence of SHV- and CTX-M-type β-lactamases. Antimicrob Agents Chemother 47:3554–3560. https://doi.org/10.1128/AAC.47.11.3554-3560.2003
Boyd DA, Tyler S, Christianson S, McGeer A, Muller MP, Willey BM, Bryce E, Gardam M, Nordmann P, Mulvey MR, Canadian Nosocomial Infection Surveillance Program (2004) Complete nucleotide sequence of a 92-Kilobase plasmid harboring the CTX-M-15 extended-spectrum Beta-lactamase involved in an outbreak in long-term-care facilities in Toronto, Canada. Antimicrob Agents Chemother 48:3758–3764. https://doi.org/10.1128/AAC.48.10.3758-3764.2004
Monstein H-J, Östholm-Balkhed Å, Nilsson MV, Nilsson M, Dornbusch K, Nilsson LE (2007) Multiplex PCR amplification assay for the detection of blaSHV, blaTEM and blaCTX-M genes in Enterobacteriaceae. APMIS 115:1400–1408. https://doi.org/10.1111/j.1600-0463.2007.00722.x
Slots J (2012) Low-cost periodontal therapy. Periodontol 2000(60):110–137. https://doi.org/10.1111/j.1600-0757.2011.00429.x
Segura-Egea JJ, Velasco-Ortega E, Torres-Lagares D, Velasco-Ponferrada MC, Monsalve-Guil L, Llamas-Carreras JM (2010) Pattern of antibiotic prescription in the management of endodontic infections amongst Spanish oral surgeons. Int Endod J 43:342–350. https://doi.org/10.1111/j.1365-2591.2010.01691.x
Paterson DL, Bonomo RA (2005) Extended-spectrum β-lactamases: a clinical update. Clin Microbiol Rev 18:657–686. https://doi.org/10.1128/CMR.18.4.657-686.2005
D’Andrea MM, Arena F, Pallecchi L, Rossolini GM (2013) CTX-M-type β-lactamases: a successful story of antibiotic resistance. Int J Med Microbiol 303:305–317. https://doi.org/10.1016/j.ijmm.2013.02.008
Van Winkelhoff AJ, Herrera Gonzales D, Winkel EG, Dellemijn-Kippuw N, Vanden-broucke-Grauls CMJE, Sanz M (2000) Antimicrobial resistance in the subgingival microflora in patients with adult periodontitis: a comparison between the Netherlands and Spain. J Clin Periodontol 27:79–86. https://doi.org/10.1034/j.1600-051x.2000.027002079.x
Ready D, Bedi R, Mullany P, Wilson M (2012) Penicillin and amoxicillin resistance in oral Veillonella spp. Int J Antimicrob Agents 40:188–189. https://doi.org/10.1016/j.ijantimicag.2012.04.007
Nyfors S, Könönen E, Bryk A, Syrjänen R, Jousimies-Somer H (2003) Age-related frequency of penicillin resistance of oral Veillonella. Diagn Microbiol Infect Dis 46:279–283. https://doi.org/10.1016/S0732-8893(03)00082-8
Reig M, Mir N, Baquero F (1997) Penicillin resistance in Veillonella. Antimicrob Agents Chemother 41:1210. https://doi.org/10.1128/aac.41.5.1210
Theron MM (2003) Penicillin-binding proteins involved in high-level piperacillin resistance in Veillonella spp. J Antimicrob Chemother 52:120–122. https://doi.org/10.1093/jac/dkg297
Kuriyama T, Williams DW, Yanagisawa M, Iwahara K, Shimizu C, Nakagawa K, Yamamoto E, Karasawa T (2007) Antimicrobial susceptibility of 800 anaerobic isolates from patients with dentoalveolar infection to 13 oral antibiotics. Oral Microbiol Immunol 22:285–288. https://doi.org/10.1111/j.1399-302X.2007.00365.x
van Winkelhoff AJ, Herrera D, Oteo A, Sanz M (2005) Antimicrobial profiles of periodontal pathogens isolated from periodontitis patients in the Netherlands and Spain. J Clin Periodontol 32:893–898. https://doi.org/10.1111/j.1600-051X.2005.00782.x
Fernández-Canigia L, Cejas D, Gutkind G, Radice M (2015) Detection and genetic characterization of β-lactamases in Prevotella intermedia and Prevotella nigrescens isolated from oral cavity infections and peritonsillar abscesses. Anaerobe 33:8–13. https://doi.org/10.1016/j.anaerobe.2015.01.007
Rams TE, Degener JE, van Winkelhoff AJ (2013) Prevalence of β-lactamase-producing bacteria in human periodontitis. J Periodontal Res 48:493–499. https://doi.org/10.1111/jre.12031
Pancoast SJ (1988) Aminoglycoside antibiotics in clinical use. Med Clin North Am 72:581–612
Holbrook WP, Ogston SA, Ross PW (1978) A method for the isolation of Bacteroides melaninogenicus from the human mouth. J Med Microbiol 11:203–207. https://doi.org/10.1099/00222615-11-2-203
McCarthy K (2015) Pseudomonas aeruginosa: evolution of antimicrobial resistance and implications for therapy. Semin Respir Crit Care Med 36:044–055. https://doi.org/10.1055/s-0034-1396907
Puzari M, Chetia P (2017) RND efflux pump mediated antibiotic resistance in gram-negative bacteria Escherichia coli and Pseudomonas aeruginosa: a major issue worldwide. World J Microbiol Biotechnol 33:24. https://doi.org/10.1007/s11274-016-2190-5
Dougherty TJ, Pucci MJ (2012) Antibiotic discovery and development. Springer US, Boston
Roberts AP, Mullany P (2011) Tn916-like genetic elements: a diverse group of modular mobile elements conferring antibiotic resistance. FEMS Microbiol Rev 35:856–871. https://doi.org/10.1111/j.1574-6976.2011.00283.x
Santoro F, Vianna ME, Roberts AP (2014) Variation on a theme; an overview of the Tn916/Tn1545 family of mobile genetic elements in the oral and nasopharyngeal streptococci. Front Microbiol 5:1. https://doi.org/10.3389/fmicb.2014.00535
Ready D, Pratten J, Roberts AP, Bedi R, Mullany P, Wilson M (2006) Potential role of Veillonella spp. as a reservoir of transferable tetracycline resistance in the oral cavity. Antimicrob Agents Chemother 50:2866–2868. https://doi.org/10.1128/AAC.00217-06
Clewell DB, Flannagan SE, Jaworski DD, Clewell DB (1995) Unconstrained bacterial promiscuity: the Tn916-Tn1545 family of conjugative transposons. Trends Microbiol 3:229–236. https://doi.org/10.1016/S0966-842X(00)88930-1
Edwards R, Beringer R, Greenwood D (1996) Characterization of β-lactamases of Prevotella species. Anaerobe 2:217–221. https://doi.org/10.1006/anae.1996.0030
Ehrmann E, Handal T, Tamanai-Shacoori Z, Bonnaure-Mallet M, Fosse T (2014) High prevalence of β-lactam and macrolide resistance genes in human oral Capnocytophaga species. J Antimicrob Chemother 69:381–384. https://doi.org/10.1093/jac/dkt350
Kolenbrander PE (2000) Oral microbial communities: biofilms, interactions, and genetic systems. Annu Rev Microbiol 54:413–437. https://doi.org/10.1146/annurev.micro.54.1.413
Roberts AP, Kreth J (2014) The impact of horizontal gene transfer on the adaptive ability of the human oral microbiome. Front Cell Infect Microbiol 4:1. https://doi.org/10.3389/fcimb.2014.00124
Ioannidis I, Sakellari D, Spala A, Arsenakis M, Konstantinidis A (2009) Prevalence of tetM, tetQ, nim and blaTEM genes in the oral cavities of Greek subjects: a pilot study. J Clin Periodontol 36:569–574. https://doi.org/10.1111/j.1600-051X.2009.01425.x
European Centre for Disease Prevention and Control (ECDC) (2014) Surveillance of antimicrobial consumption in Europe 2012. https://www.ecdc.europa.eu/en/publications-data/surveillance-antimicrobial-consumption-europe-2012
Haenni M, Lupo A, Madec J-Y (2018) Antimicrobial resistance in Streptococcus spp. Microbiol Spectr 6:1. https://doi.org/10.1128/microbiolspec.ARBA-0008-2017
Ding Y, Zhang J, Mi Z, Qin L, Tao YZ, Qi X (2004) Study on the molecular epidemiology of beta-lactamase TEM gene in isolated Streptococcus pneumoniae. Zhonghua Liu Xing Bing Xue Za Zhi 25:970–972
Chang C-Y, Lin H-J, Li B-R, Li Y-K (2016) A novel metallo-β-lactamase involved in the ampicillin resistance of Streptococcus pneumoniae ATCC 49136 strain. PLoS One 11:e0155905. https://doi.org/10.1371/journal.pone.0155905
Koncan R, Valverde A, Morosini M-I, García-Castillo M, Cantón R, Cornaglia G, Baquero F, del Campo R (2007) Learning from mistakes: Taq polymerase contaminated with β-lactamase sequences results in false emergence of Streptococcus pneumoniae containing TEM. J Antimicrob Chemother 60:702–703. https://doi.org/10.1093/jac/dkm239
Naito M, Sato K, Shoji M, Yukitake H, Ogura Y, Hayashi T, Nakayama K (2011) Characterization of the Porphyromonas gingivalis conjugative transposon CTnPg1: determination of the integration site and the genes essential for conjugal transfer. Microbiology 157:2022–2032. https://doi.org/10.1099/mic.0.047803-0
Funding
No external funding, apart from the support of the authors’ institution, was available for this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the Universitat Internacional de Catalunya research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 69.7 kb).
Rights and permissions
About this article
Cite this article
Arredondo, A., Blanc, V., Mor, C. et al. Resistance to β-lactams and distribution of β-lactam resistance genes in subgingival microbiota from Spanish patients with periodontitis. Clin Oral Invest 24, 4639–4648 (2020). https://doi.org/10.1007/s00784-020-03333-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00784-020-03333-1