Abstract
Molybdenum is found in the active site of enzymes usually coordinated by one or two pyranopterin molecules. Here, we mimic an enzyme with a mononuclear molybdenum-bis pyranopterin center by incorporating molybdenum in rubredoxin. In the molybdenum-substituted rubredoxin, the metal ion is coordinated by four sulfurs from conserved cysteine residues of the apo-rubredoxin and two other exogenous ligands, oxygen and thiol, forming a Mo(VI)-(S-Cys)4(O)(X) complex, where X represents –OH or –SR. The rubredoxin molybdenum center is stabilized in a Mo(VI) oxidation state, but can be reduced to Mo(IV) via Mo(V) by dithionite, being a suitable model for the spectroscopic properties of resting and reduced forms of molybdenum-bis pyranopterin-containing enzymes. Preliminary experiments indicate that the molybdenum site built in rubredoxin can promote oxo transfer reactions, as exemplified with the oxidation of arsenite to arsenate.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Metal substitutions in metalloproteins have been an attractive research field, contributing to the understanding of the structural/functional relationships. This strategy has been demonstrated to be a valuable tool for spectroscopic studies and for the mimic of active sites and the design of novel metalloproteins [1–3]. Rubredoxins (Rd) are the simplest iron–sulfur proteins, with a molecular mass of 5.6 kDa and containing a single iron atom tetrahedrally coordinated by the sulfur atoms of four cysteine residues (Fig. 1a) arranged in a –CX2C–X n –CX2C– motif [4, 5]. A variation of this binding mode is found in desulforedoxin, where the binding motif contains two adjacent cysteines, depicted as –CX2C–X n –CC– and has a ferredoxin-like fold [6–8]. Mutations on the desulforedoxin-binding motif by addition of two amino acid residues between the two adjacent cysteine residues converts the desulforedoxin iron site into a center with the spectroscopic characteristics of Rd [9].
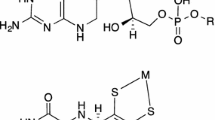
Schematic representation of the centers of a Fe-Rd and b Mo-bis PGD enzymes and of c pyranopterin cofactor. In (b), X and Y represent terminal O, S, Se atoms and/or Asp, Ser, Cys, and SeCys residues; for simplicity, only the dithiolate moiety of the pyranopterin cofactor is represented. In E. coli periplasmatic nitrate reductase, X and Y represent a terminal oxo group and cysteine residue; in Alcaligenes faecalis arsenite oxidase, X and Y represent terminal oxo and hydroxyl groups. The cofactor (c) is a pyranopterin-dithiolate moiety, which forms a five-membered ene-1,2-dithiolate chelate ring with the molybdenum atom; in Mo-bis PGD enzymes, the cofactor is found esterificated, as a pyranopterin guanosine dinucleotide (R represents guanosine monophosphate)
Rd provides an excellent system for obtaining a wide range of metal-substituted proteins, due to its small size and simple structure [10–13], in which the iron atom can be substituted by reconstitution of the apo-form (apo-Rd), prepared by acid precipitation under reducing conditions. Over the years, metal-substituted Rd-type proteins containing 57Fe(II), Co(II), Ni(II), Zn(II), Cd(II), Hg(II), Ga(III) and In(III) have been prepared and characterized [14–22]. Among these, nickel-substituted Rd was pointed out as a model compound of the nickel site of bacterial hydrogenase, displaying similar D/H exchange properties, as well as CO inhibition [16]. The preparation of copper-substituted Rd had remained elusive and was achieved only recently [23].
To date, a molybdenum-substituted rubredoxin (Mo-Rd) had not been reported. An obvious interest is the possibility of creating a sulfur-rich environment that mimics the active site of mononuclear molybdenum-bis pyranopterin-containing enzymes. The mononuclear molybdoenzymes harbor in their active site one molybdenum atom coordinated by the cis-dithiolene group of one or two pyranopterin cofactor molecules (Fig. 1c) and by oxygen, sulfur, or selenium atoms, in a diversity of arrangements that determines the classification of molybdoenzymes into three families: xanthine oxidase, sulfite oxidase, and dimethylsulfoxide reductase (DMSOR) [24–26]. The DMSOR family is the larger and more diverse family, comprising prokaryotic enzymes of different functions (e.g., enzymes involved in oxygen atom transfer, C–H bond cleavage, or hydration reactions) [27, 28]. The enzymes from this family (in the oxidized form) hold a trigonal prismatic L2MoXY core, where L stands for the pyranopterin cofactor and X and Y represent terminal =O, –OH, =S, and –SH groups and/or oxygen, sulfur, or selenium atoms from cysteine, selenocysteine, serine, or aspartate residue side chains (Fig. 1b). Because the molybdenum atom of these enzymes is coordinated by two pyranopterin guanosine dinucleotides, these enzymes are proposed to be grouped under a new and more meaningful denomination: molybdenum-bis pyranopterin guanosine dinucleotide-containing enzymes (Mo-bis PGD enzymes) [26, 29, 30].
The molybdoenzymes, and the Mo-bis PGD enzymes in particular, occupy a significant place in bioinorganic chemistry, as they carry out a wide range of metabolic reactions in the nitrogen, sulfur, and carbon biocycles [31, 32]. Therefore, structures of molybdenum coordinated with several sulfur atoms are interesting models for resting and/or reduced species of active site of Mo-bis PGD enzymes. Herein, we report the synthesis and characterization of a novel molybdenum center coordinated by the four cysteine residues that coordinate the iron site of Rd (Fe-(S-Cys)4) and we will discuss how this center can be regarded as a mimetic system for Mo-bis PGD enzymes.
Materials and methods
Unless otherwise stated, all reagents were of analytical grade or higher and purchased from Sigma-Aldrich, Merck, Fluka or Riedel-de-Häen.
Overexpression and purification of Desulfovibrio gigas rubredoxin
Desulfovibrio gigas Rd was heterologously expressed in E. coli BL21(DE3) as an iron-containing rubredoxin (Fe-Rd) [33], and the protein was purified as previously described [6, 34]. The purity of the Fe-Rd was checked by SDS-PAGE and UV–visible spectrum and presented an A 280nm/A 494nm purity ratio of 2.4.
Reconstitution of rubredoxin with molybdenum
Apo-Rd was prepared by precipitation of Fe-Rd with 80 % trichloroacetic acid (TCA), under reducing conditions, in the presence of dithiothreitol (DTT) [35]. The Mo-Rd was prepared as shown schematically in Fig. 2. The reconstitution protocol was initiated by adding H2MoO4 to β-mercaptoethanol (β-ME) (ratio 1:4) in lukewarm water (heat is required to dissolve the solid H2MoO4 in water). A red solution was obtained, with a maximum absorption band at 335 nm and a shoulder at 450 nm, which is consistent with the formation of MoO2(SR)4 (SR = aliphatic thiol) [36–40]. Therefore, in the reaction mixture, {MoO2(β-ME)4} is probably the complex that is formed. A molar excess (1.5- to 2-fold) of this complex was added to an apo-Rd solution (1.76 mM) in 50 mM Tris–HCl, pH 7.6, and incubated for 2 h on an ice bath. This solution was then dialyzed, at 4 °C, against 1000-fold volume of 25 mM acetate buffer, pH 5.0, for 24 h. The residual-free metal complex ({MoO2(β-ME)4}) was removed by passing the sample through a PD-10 desalting column (GE Healthcare), equilibrated with the same buffer at pH 5.0. No attempts of reconstitution were made below pH 4, because apo-Rd becomes unstable and precipitates, or above pH 6.5, since molybdenum, in the complex {MoO2(β-ME)4}, is reluctant to be coordinated by protein cysteine thiols at high pH values.
Preparation of oxidized oxo-Mo-Rd and sulfo-Mo-Rd and of reduced Mo-Rd. See “Materials and methods“ for details
Determination of the protein/metal ratio
Molybdenum content was determined by inductively coupled plasma (ICP) emission analysis using the multi-elements (Reagecon) as a standard solution in a concentration range of 0.01–1 ppm. The protein concentration was determined using the BCA kit (Sigma) with bovine serum albumin (Sigma) as standard. The Mo-Rd samples were analyzed after desalting using a PD-10 desalting column (GE Healthcare).
Spectroscopic methods
UV–visible absorption spectra were recorded in 1 cm path length cuvette, using a Shimadzu UV 1800 spectrophotometer, at room temperature.
Resonance Raman (RR) spectra were recorded using a Raman spectrometer (Jobin–Yvon U1000), equipped with a 1200 lines/mm grating and a liquid nitrogen-cooled CCD detector. A confocal microscope equipped with an Olympus 20× objective (working distance of 21 mm, numeric aperture of 0.35) was used for laser focusing onto the sample and light collection in the backscattering geometry. The 413 nm line from a Kr+ laser (Coherent Innova 302) was used as the excitation source. 2 μL of 400 μM Mo-Rd solution were placed onto a Linkam THMS 600 microscope stage and cooled with liquid N2. All spectra were recorded at 83 K. The laser power and accumulation time were set to 12 mW and 60 s, respectively; typically six to ten spectra were co-added to improve the signal/noise ratio.
X-band EPR (9.65 GHz) spectra of Mo-Rd frozen solution (400 µM), at 77 K, were recorded using a Bruker EMX 6/1 spectrometer and an ER4116DM rectangular cavity (Bruker); the samples were cooled with liquid helium in an Oxford Instruments ESR900 continuous-flow cryostat, fitted with a temperature controller. The acquisition conditions were a modulation frequency of 100 kHz, a modulation amplitude of 0.2 mT and a microwave power of 635 µW. The EPR spectra were simulated with WinEPR SimFonia (ver. 1.25) program from Bruker Analytische GmbH, considering the Hamiltonian,
where µ B (= 9.2740154(31) × 10−24JT−1) is the Bohr magneton, \( \hat{S} \) the electron spin angular moment operator and B the magnetic field.
19F NMR spectra were acquired on a Bruker ARX NMR spectrometer operating at 400 MHz and locked on the 2H resonance of the solvent (D2O). The NMR sample was a 400 µM Mo-Rd solution, which was incubated overnight at 4 °C with excess of 2,2,2-trifluoroethanethiol (TFET) in 25 mM acetate buffer, pH 5.0, with 20 % D2O.
Electrochemical studies
Cyclic voltammetry (CV) was performed, at 6 °C, on a potentiostat/galvanostat AUTOLAB system with an electrochemical cell consisting of a glassy carbon (GC) [or pyrolytic graphite (PGE)] working electrode, a platinum wire counter electrode, and a KCl saturated Ag/AgCl reference electrode. For protein film experiments, the protein solution (400 μM in 25 mM acetate, pH 5.0) with the co-adsorbent polymyxin-B (typically, 0.2 mg/mL) was used to coat the electrode. The buffer-electrolyte solution in the electrochemical cell consisted of 25 mM acetate buffer, pH 5.0. To stabilize the protein films, 0.2 mg/mL polymyxin-B was also added to the cell solution to promote the electrochemical response. Protein films were prepared by coating the freshly polished GC/PGE electrode with an ice-cold protein solution. The electrode was then transferred to the cell. Potentials were measured against internal ferrocene (Fc) and are reported relatively to normal hydrogen electrode (NHE).
Assay for arsenite oxidase activity
The Mo-Rd ability to oxidize arsenite was studied following the absorbance changes in the Mo-Rd spectrum under acidic conditions, in the presence of 0.1–5.0 mM arsenite, at room temperature. The confirmation that arsenate is the product of the Mo-Rd reaction with arsenite was performed with the molybdenum blue test [41, 42]. The same assay was used to quantify the amount of arsenate formed in the presence of Mo-Rd using different concentrations of arsenite. The molybdenum blue test is based on the reaction of ammonium molybdate with arsenate, under acidic and reducing conditions, to yield an arseno-molybdate blue complex that can be quantified at 840–890 nm [41, 42]. In short, the sample (1 mL) containing arsenate (formed after reaction of Mo-Rd with arsenite) was mixed with the colored reagent (500 µL) constituted by a mixture of 2:2:1:5 (v:v:v:v) of 10.8 % ascorbic acid: 3 % ammonium molybdate: 0.56 % potassium antimonyl tartrate: 13.98 % sulfuric acid. The absorbance at 880 nm was measured after 60 min. Arsenite and Mo-Rd were both assayed separately and no color formation was observed. A calibration curve was prepared with standards of known arsenate concentrations (1–20 μM) to determine the arsenate concentration formed by Mo-Rd in the presence of arsenite.
Results and discussion
Preparation of Mo-Rd
The reconstitution of apo-Rd with molybdenum was achieved at pH 5.0. The Mo-Rd obtained has a molybdenum/protein ratio of 1.2 and migrated as a single band in SDS-PAGE under reducing and non-reducing conditions (presence and absence of β-ME) (Fig. 3a), indicating that all the Rd cysteine residues are coordinating the metal ion prior to entering in the gel [23], and also that there is no apo-Rd after the reconstitution procedure. The UV–visible spectrum of the as-prepared Mo-Rd (Fig. 3b; black line) shows a maximum absorption band at 278 nm (ε = 17.77 mM−1 cm−1) with two unresolved shoulders at 314 and 450 nm (ε = 8.01 and 1.02 mM−1 cm−1, respectively). These shoulders can be assigned to Mo-thiolate charge transfer bands, which are similar to the ones observed in synthetic inorganic model complexes [36–40] and to the ones assigned for the protein YedY, though the molar extinction coefficient values are 10× for the 450 nm and both absorption bands are shifted to higher energy values [43]. It is important to mention that the choice of pH, under which the reconstitution procedure is carried out, is crucial for the successful formation of Mo-Rd. As mentioned in “Materials and methods“, pH values between 4.0 and 6.5 favor the incorporation of molybdenum and its coordination by the Rd cysteine thiols (Fig. 2). The drop of pH to 5.0 could be performed by either addition of acetic acid to the mixture containing apo-Rd and {MoO2(β-ME)4} or by dialysis of the mixture against acetate buffer at pH 5.0. However, in the first option, there was no incorporation of metal, due to the formation of polynuclear complexes of molybdenum [44]. In the second strategy, using dialysis to adjust the pH to 5.0, the reconstitution was accomplished. This can be explained by the fact that β-ME reacts with molybdenum and then it exchanges slowly with Rd cysteines thiols during the dialysis process. Therefore, we concluded that both pH and dialysis play a critical role in the insertion of molybdenum into apo-Rd, with the optimal conditions being achieved when {MoO2(β-ME)4} and apo-Rd were dialyzed against acetate buffer at pH 5.0, for 24 h, at 4 °C.
a SDS-PAGE gel under reducing and non-reducing conditions (presence and absence of β-ME, respectively) of Fe-Rd and Mo-Rd (after desalting); the migration of molecular weight markers (MWM) of 25, 15, and 10 kDa is indicated. b UV–visible spectrum of the as-prepared Mo-Rd (black line) in 25 mM acetate buffer, pH 5.0, and of the DTT-treated Mo-Rd (gray line)
Spectroscopic characterization of the coordination sphere of molybdenum in Mo-Rd
The Mo-Rd was probed by X-band electron paramagnetic resonance (EPR) spectroscopy. The as-prepared Mo-Rd is EPR silent, as expected for the oxidized Mo-Rd with the metal center in the Mo(VI) state. In the presence of DTT, the Mo-Rd EPR signal does not change, which suggests that DTT is not able to reduce Mo-Rd. Upon anaerobic reduction with sodium dithionite (20–25 min), an EPR-active species is detected at 77 K. This species has a rhombic spectrum (Fig. 4a) that can be simulated as a single signal, with g values of 2.02, 1.97 and 1.92, and is attributed to a Mo(V) species (d 1; S = ½) [45, 46]. Increasing the reduction time (to 2 h), the intensity of the Mo-Rd signal decreases suggesting that Mo(V) is further reduced to Mo(IV), which is EPR silent.
a X-band (9.65 GHz) EPR spectrum of dithionite-reduced Mo-Rd (400 μM solution, in 25 mM acetate buffer, pH 5.0, reduction time of 25 min). The experimental spectrum (exp) was acquired at 77 K, as described in “Materials and methods“, and the other experimental conditions were: modulation frequency, 100 kHz; modulation amplitude, 0.2 mT; microwave power, 635 μW. The spectrum was simulated (simul) as described in “Materials and methods“; the g value position is indicated by the arrows. b Cyclic voltammetry (100 mV s−1) and differential pulse voltammetry at a GC electrode of Mo-Rd (400 μM in 25 mM acetate buffer, pH 5.0, with 0.2 mg/mL polymyxin-B co-adsorbed; 4 °C)
It was anticipated that molybdenum coordination sphere would not be satisfied by only four cysteinyl ligands and thus it was expected that the metal would have a tendency to enlarge its coordination number. The coordination sphere of the molybdenum center in Mo-Rd was addressed by resonance Raman (RR) spectroscopy. Upon excitation into an electronic absorption band of a chromophore, RR spectra selectively reveal molecular details of the moiety that gives origin to electronic transition. In Mo-Rd, the resonance enhancement condition is fulfilled for 413 nm laser line which matches the assigned broad cysteinyl S-Mo(VI) charge transfer (CT) band centered at ~400 nm [47]. In the low-frequency region (~300–400 cm−1), RR spectra of Mo-Rd revealed Mo–S stretching and bending vibrational modes (Fig. 5a, upper trace) also observed in other molybdenum-containing proteins and model compounds [48–51]. Deconvolution of the spectra reveals four vibrational modes, centered at 347, 363, 385, and 401 cm−1 (Fig. 5a, inset), the most intense of which (363 cm−1) carries the highest stretching character. Well-resolved Fe–S modes are on average slightly downshifted (345, 361 and 376 cm−1) in the RR spectrum of native Rd (Fig. 5a, bottom trace) [52]. Note that strong features in the spectra marked with asterisks originate from lattice modes of the frozen buffer solution. The band centered at ~705 cm−1 (Fig. 5a, upper trace) reflects the first overtone or combination band of the Mo–S stretching modes. Another prominent feature in the RR spectrum of Mo-Rd, which is clearly absent from that of Rd, is a broad band at 942 cm−1. Similar bands were identified in RR spectra of other oxomolybdenum-containing enzymes, e.g., human sulfite oxidase (hSO) at 903 cm−1, DMSOR at 862 cm−1, and biotin sulfoxide reductase (BSO reductase) at 860 cm−1 and, based on isotopic labeling, attributed to Mo=O stretching mode [53]. Therefore, by analogy with the RR spectra of these enzymes and model compounds, we attributed the 942 cm−1 mode in the Mo-Rd RR spectra to Mo=O symmetric stretching, ν sym. It is ~40 cm−1 upshifted with respect to that of the hSO, and even more in comparison with DMSOR and BSO reductase, indicating a stronger bond; it is closer in energy to the Mo=O bond observed in cis dioxomolybdenum (VI) thiolate complexes [48–51]. Similarly to the BSO reductase [54], in Mo-Rd we observe an atypically large bandwidth of the 942 cm−1 mode (Δν ~ 52 cm−1). It suggests the presence of an unresolved asymmetric Mo=O stretching mode, which is, for example, well resolved in the RR spectra of hSO, in which the ν sym and ν asym are found ~20 cm−1 apart. Alternatively, it reflects heterogeneity in the active site structure or environment.
a Resonance Raman spectra of Mo-Rd (upper trace; 400 μM in 25 mM acetate buffer, pH 5.0) and of Fe-Rd (lower trace; 1 mM in 50 mM Tris–HCl buffer, pH 7.6). Inset shows deconvolution of Mo–S band. Bands marked with asterisks are from the buffer. b 19F NMR spectrum of Mo-Rd (400 μM in 25 mM acetate buffer, pH 5.0) in the presence of 2 mM TFET
Since it is well known that molybdenum ions in enzymes have a tendency to have higher coordination numbers, we can speculate that the metal center in Mo-Rd contains Mo=O(=O…H–protein) or Mo=O(–OH) (Fig. 2). This type of coordination is observed in the oxidized form of arsenite oxidase, where the molybdenum atom is coordinated by two pyranopterin molecules plus an oxo and a hydroxyl group [55, 56] (Fig. 1b) and also in tungsten-containing aldehyde:ferredoxin oxidoreductase [57].
Further spectroscopic characterization of the molybdenum coordination in Mo-Rd was carried out exploiting the effect of the presence of exogenous thiol compounds. Upon addition of DTT to a Mo-Rd solution at pH 5.0, the UV–visible spectrum changes and bands arise at 336 nm (ε = 7.76 mM−1 cm−1) and 460 nm (ε = 1.91 mM−1 cm−1) (Fig. 3b; gray line), indicating that the molybdenum might be coordinated by a sulfur atom from the exogenous thiolate ligand. Therefore, DTT might serve as an exogenous thiol ligand to the Mo-Rd molybdenum center.
To further support the coordination of exogenous thiol ligands to the molybdenum center of Mo-Rd, the binding of 2,2,2-tri-fluoroethanethiol (TFET: CF3CH2SH) was studied by 19F-NMR spectroscopy. The TFET-reacted Mo-Rd 19F-NMR spectrum presents a triplet signal at −65.7, −66.0 and −66.3 ppm and a sharp peak at −76.9 ppm (Fig. 5b). While the −76.9 ppm peak is due to non-coordinated fluorothiol, the triplet signal is attributed to the molybdenum-coordinated fluorothiol [58–60]. The triplet signal suggests the formation of hydrogen bonds with fluorine atoms and demonstrates that also TFET can coordinate the Mo-Rd molybdenum center. Therefore, different thiol compounds can act as exogenous thiol ligands to the molybdenum center of Mo-Rd [61, 62].
We propose that the exogenous thiol compounds, either DTT or TFET, can replace the hydroxyl ligand in the molybdenum coordination sphere and the probable structures of the Mo-Rd center and thiol-treated Mo-Rd center are Rd-Mo(VI)=O(–OH) and Rd-Mo(VI)=O)(–SRexogenous), respectively (Fig. 2). This proposal is in agreement with the known molybdenum chemistry, which is dominated by the formation of oxides and sulfides and where the molybdenum centers show a strong tendency to easily lose an oxygen atom [63, 64]. Therefore, we suggest that the Mo-Rd metal site has a molybdenum atom coordinated by four cysteine sulfur atoms (Mo(S-Cys)4; the rubredoxin site), with two additional exogenous ligands, one oxo and either one thiol or one hydroxo group, to satisfy the higher coordination number found in Mo-bis PGD enzymes (Fig. 1b). The thiol-treated Mo-Rd, with the suggested structure of Rd-Mo(VI)(O)(SRexogenous), can be envisaged as a model for the E. coli or Rhodobacter sphaeroides periplasmatic nitrate reductase [65, 66], a molybdoenzyme that harbors the molybdenum atom coordinated by two pyranopterin molecules, one oxo group, and one cysteine sulfur atom. Conversely, the untreated Mo-Rd, with the suggested structure of Rd-Mo(VI)(O)(–OH), can constitute a model of arsenite oxidase with its molybdenum atom coordinated by two pyranopterin molecules, one oxo group, and one hydroxyl group (Fig. 1b) [55, 56, 67].
Electrochemical studies
To gain information on the redox properties of the Mo-Rd metal center, electrochemical techniques were used. Mo-Rd thin films displayed quasi-reversible CV peaks at E pc = −530 mV and E pa = −345 mV vs. NHE [ΔE = 185 mV, at 100 mV s−1 (Fig. 4b)]. The midpoint potential (E 1/2) determined was −493 mV vs. NHE and was assigned to the Mo(VI)/Mo(V) redox couple. This value is in reasonable agreement with the reduction potential of some molybdoenzymes. For example, the Desulfovibrio gigas aldehyde oxidoreductase (that harbors the molybdenum atom coordinated by one pyranopterin, two oxo, and one hydroxyl groups) displays reduction potential values of −415 and −539 mV for the Mo(VI)/Mo(V) and Mo(V)/Mo(IV) couples, respectively [68, 69]. Reduction peak potentials are shifted to more negative values and the separation between the two peaks is higher at higher scan rates, indicating a quasi-reversible behavior for the electron transfer of Mo-Rd through the electrode at high scan rates [E pc (reduction) = −785 mV, E pa (oxidation) = −529 mV and ΔE = 256 mV at 500 mV s−1; not shown] [70]. Furthermore, the asymmetric behavior indicates that the electron transfer rate for the oxidation is slower than that for the reduction process.
Activity of the Mo-Rd as arsenite oxidase
The Mo(O)(S-Cys)4 moiety of Mo-Rd depicts most of the features of the metal center functional unit (McFU) found in the active site of Mo-bis PGD enzymes of the Mo(O)(pyranopterin) 2 type [71]. Variations in the coordination chemistry and properties of the metal center occur with the binding of other ligands, which can include terminal =O, –OH, –OH2, =S, –SH group(s), and serine, aspartate, cysteine, or selenocysteine residue(s) from the polypeptide backbone of the protein. Having replicated the molybdenum first coordination sphere of a Mo-bis PGD enzyme, the next obvious step was to investigate the Mo-Rd hypothetical reactivity in oxygen atom transfer reactions [72, 73]. Given the similarity of the suggested molybdenum first coordination sphere of Mo-Rd with the one found in the oxidized arsenite oxidase catalytic center (molybdenum atom coordinated by two pyranopterin molecules plus an oxo and a hydroxyl group [55, 56] ), the oxidation of arsenite (As(III)O3 3−) to arsenate (As(V)O4 3−) by Mo-Rd (an oxygen atom insertion reaction) was studied. This reaction was monitored under acidic conditions, at room temperature, following the absorbance changes in the Mo-Rd spectrum, in the presence of 0.1–5.0 mM arsenite. Representative spectral changes of Mo-Rd in the presence of 1.0 mM arsenite are shown in Fig. 6, where the decrease in the absorption between 300 and 500 nm was observed. It is known that As(III) has a tendency to be coordinated by thiols, with the resulting CT electronic transition (broad) bands developing in the 250–320 nm range [74]. Because such bands did not emerge, the observed spectral changes were attributed to the Mo-Rd reduction by arsenite and not to arsenite-modified Mo-Rd. The dependence of the absorbance change at 360 nm on the arsenite concentration, for 0.1, 0.2, 0.5 and 1 mM, is shown in the inset of Fig. 6. No reaction took place in the presence of arsenate (no absorbance change was observed in the presence of arsenate; data not shown). The confirmation that arsenate is in fact the product of the Mo-Rd reaction with arsenite was achieved with the molybdenum blue test [41, 42], which was also used to quantify the amount of arsenate formed (Table 1). The arsenate formation was found to proceed slowly, with an initial rate of 0.58 ± 0.05 μM/min for the oxidation of 5 mM arsenite by 50 μM Mo-Rd, and the reaction extent was small, with <1 % of arsenate formed after 2 h (Table 1).
Spectral change of Mo-Rd (80 μM) reacted with 1 mM arsenite, in 25 mM acetate, pH 5, at room temperature. Spectra were recorded every 60 s, for 2 h; the decrease in absorbance (Abs) with time is indicated with the arrow. Inset absorbance change at 360 nm, as a function of time, observed in the presence of 0.1 (a), 0.2 (b), 0.5 (c) and 1 mM (d) arsenite
Even though Mo-Rd cannot be considered an efficient “arsenite oxidizer” (in comparison with the true arsenite oxidase enzyme, which has a k cat and a K m of 27 s−1 and 8 μM, respectively [67, 75]), our results demonstrate that Mo-Rd can promote arsenite oxidation. In line with the known mechanism of reaction of the molybdoenzymes, we suggest that the Mo-Rd {MoVIO2} moiety would be concomitantly reduced to {MoIVO}. The insertion of molybdenum into apo-Rd transforms a one-electron transfer protein (Fe-Rd) into an oxygen atom/two-electrons transfer “arsenite oxidase” (Mo-Rd). This alteration of function is noteworthy, in particular if one takes into account that the Mo-Rd does not hold a specific “substrate-binding pocket” to promote correct and efficient arsenite binding. Therefore, Mo-Rd can be considered as a functional model of arsenite oxidase.
Conclusions
In conclusion, Rd was successfully reconstituted with molybdenum and suggested harboring a Mo=O(OH)(–SCys)4 center or, after treatment with exogenous thiol compounds, a Mo=O(SR)(–SCys)4 center. The reconstituted site in Mo-Rd is, thus, suggested to mimic the active site of a Mo-bis PGD enzyme, in terms of (1) the nature of the ligands present in the first coordination sphere of molybdenum, (2) spectroscopic properties and (3) ability to insert an oxygen atom in a “substrate”. The Mo-Rd molybdenum center can thus be considered a structural and functional analog of some Mo-bis PGD enzymes. The rich sulfur coordination provided by the four apo-Rd cysteine residues can be extended by a wide range of ligands, including terminal oxo or sulfur-coordinating atoms in a versatile coordination sphere as seen in enzymes that include terminal =O, –OH, –OH2, =S, –SH group(s), and/or serine, aspartate, cysteine, or selenocysteine residue(s) from the polypeptide backbone of the protein. More detailed kinetic studies of the arsenite oxidase activity of Mo-Rd center are in progress, as well as of the reaction of this center with other relevant substrates.
Abbreviations
- apo-Rd:
-
Apo-rubredoxin
- β-ME:
-
β-Mercaptoethanol
- BSO:
-
Biotin sulfoxide
- CT:
-
Charge transfer
- CV:
-
Cyclic voltammetry
- DMSOR:
-
Dimethylsulfoxide reductase
- DTT:
-
Dithiothreitol
- DPV:
-
Differential pulsed voltammetry
- Fe-Rd:
-
Iron-containing rubredoxin
- GC:
-
Glassy carbon
- hSO:
-
Human sulfite oxidase
- M:
-
Metal
- Mo-bis PGD:
-
Molybdenum-bis pyranopterin guanosine dinucleotide-containing enzyme
- Mo-Rd:
-
Molybdenum-substituted rubredoxin
- PGE:
-
Pyrolytic graphite electrode
- NHE:
-
Normal hydrogen electrode
- RR:
-
Resonance Raman
- Rd:
-
Rubredoxin
- TCA:
-
Trichloroacetic acid
- TFET:
-
2,2,2-Trifluoroethanethiol
References
Lu Y, Berry SM, Pfister TD (2001) Chem Rev 101:3047–3080
Liu J, Meier KK, Tian S, Zhang J, Guo H, Schulz CE, Robinson H, Nilges MJ, Münck E, Lu Y (2014) J Am Chem Soc 136:12337–12344
Yu F, Cangelosi VM, Zastrow ML, Tegoni M, Plegaria JS, Tebo AG, Mocny CS, Ruckthong L, Qayyum H, Pecoraro VL (2014) Chem Rev 114:3495–3578
Adman ET, Sieker LC, Jensen LH, Bruschi M, Le Gall J (1977) J Mol Biol 112:113–120
Chen CJ, Liu MY, Chen YT, LeGall J (2003) Biochem Biophys Res Commun 308:684–688
Moura I, Bruschi M, LeGall J, Moura JJG, Xavier AV (1977) Biochem Biophys Res Commun 75:1037–1044
Bruschi M, Moura I, Le Gall J, Xavier AV, Sieker LC, Couchoud P (1979) Biochem Biophys Res Commun 90:596–605
Czaja C, Litwiller R, Tomlinson AJ, Naylor S, Tavares P, LeGall J, Moura JJG, Moura I, Rusnak F (1995) J Biol Chem 270:20273–20277
Yu L, Kennedy M, Czaja C, Tavares P, Moura JJ, Moura I (1997) Rusnak FBiochem Biophys Res Commun 231:679–682
Dauter Z, Wilson KS, Sieker LC, Moulis JM, Meyer J (1996) Proc Natl Acad Sci USA 93:8836–8840
Archer M, Carvalho AL, Teixeira S, Moura I, Moura JJG, Rusnak F, Romao MJ (1999) Protein Sci 8:1536–1545
Ascenso C, Rusnak F, Cabrito I, Lima MJ, Naylor S, Moura I, Moura JJG (2000) J Bio Inorg Chem 5:720–729
Sun N, Dey A, Xiao Z, Wedd AG, Hodgson KO, Hedman B, Solomon EI (2010) J Am Chem Soc 132:12639–12647
May SW, Kuo JY (1978) Biochemistry 17:3333–3338
Kowal AT, Zambrano IC, Moura I, Moura JJG, LeGall J, Johnson MK (1988) Inorg Chem 27:1162–1166
Saint-Martin P, Lespinat PA, Fauque G, Berlier Y, Legall J, Moura I, Teixeira M, Xavier AV, Moura JJG (1988) Proc Natl Acad Sci USA 85:9378–9380
Moura I, Teixeira M, LeGall J, Moura JJG (1991) J Inorg Biochem 44:127–139
Henehan CJ, Pountney DL, Zerbe O, Vasak M (1993) Protein Sci 2:1756–1764
Ravi N, Prickril BC, Kurtz DM Jr, Huynh BH (1993) Biochemistry 32:8487–8491
Faller P, Ctortecka B, Tröger W, Butz T, Vasák M (2000) J Biol Inorg Chem 5:393–401
Maher M, Cross M, Wilce MCJ, Guss JM, Wedd AG (2004) Acta Cryst D 60:298–303
Mathies G, Almeida RM, Gast P, Moura JJG, Groenen EJJ (2012) J Phys Chem B 116:7122–7128
Thapper A, Rizzi AC, Brondino CD, Wedd AG, Pais RJ, Maiti BK, Moura I, Pauleta SR, Moura JJG (2013) J Inorg Biochem 127:232–237
Hille R (1996) Chem Rev 96:2757–2816
Brondino CD, Romao MJ, Moura I, Moura JJG (2006) Curr Opin Chem Biol 10:109–114
Hille R, Hall J, Basu P (2014) Chem Rev 114:3963–4038
Rothery RA, Workun GJ, Weiner JH (2008) Biochim Biophys Acta 1778:1897–1929
Grimaldi S, Schoepp-Cothenet B, Ceccaldi P, Guigliarelli B, Magalon A (2013) Biochem Biophys Acta 1827:1048–1085
Rajagopalan KV (1980) Sulfite Oxidase. In: Coughlan MP (ed) Molybdenum and molybdenum containing enzymes, chap 7. Pergamon Press, Oxford, pp 241–272
Pushie MJ, George GN (2011) Coord Chem Rev 255:1055–1084
Stiefel EL (2002) The biogeochemistry of molybdenum and tungsten. In: Sigel A, Sigel H (eds) Molybdenum and tungsten: their roles in biological processes. Metals ions in biological system, chap 1, vol 39. CRC Press, pp 1–29
Mendel RR (2005) Dalton Trans 3404–3409
Almeida RM, Pauleta SR, Moura I, Moura JJG (2009) J Inorg Biochem 103:1245–1253
Bruschi M, Hatchikian CE, Golovleva LA, LeGall J (1977) J Bacteriol 129:30–38
Lode ET, Coon MJ (1971) J Biol Chem 246:791–802
Spence JT, Chang HYY (1963) Inorg Chem 2:319–323
Buchanan I, Minelli M, Ashby MT, King TJ, Enemark JH, Garner CD (1984) Inorg Chem 23:495–500
Bishop PT, Dilworth JR, Hutchinson JP, Zubieta JA (1990) Trans Met Chem 15:177–182
Huang TJ, Haight GP Jr (1970) J Am Chem Soc 92:2336–2342
Ueyama N, Okamura T, Nakamura A (1992) J Am Chem Soc 114:8129–8137
Murphy J, Riley JP (1962) Anal Chim Acta 27:31–36
Tsang S, Phu F, Baum MM, Poskrebyshev GA (2007) Talanta 71:1560–1568
Yang J, Rothery R, Sempombe J, Weiner JH, Kirk ML (2009) J Am Chem Soc 131(43):15612–15614
Liu S, Sun X, Zubieta JA (1988) J Am Chem Soc 110:3324–3326
Brondino CD, Rivas MG, Romao MJ, Moura JJG, Moura I (2006) Acc Chem Res 29:7887–7896
Klein EL, Astashkin AV, Raitsimring AM, Enemark JH (2013) Coord Chem Rev 257:110–118
Izumi Y, Glaser T, Rose K, McMaster J, Basu P, Enemark JH, Hedman B, Hodgson KO, Solomon EI (1999) J Am Chem Soc 121:10035–10046
Subramanian P, Burgmayer S, Richards S, Szalai V, Spiro TG (1990) Inorg Chem 29:3849–3853
Johnson MK (2004) Prog Inorg Chem 52:213–266
Ueyama N, Nakata M, Araki T, Nakamura A, Yamashita S, Yamashita T (1981) Inorg Chem 20:1934–1937
Garrett RM, Rajagopalan KV (1996) J Biol Chem 271:7387–7391
Czernuszewicz RS, LeGall J, Moura I, Spiro TG (1986) Inorg Chem 25:696–700
Garton SD, Garrett RM, Rajagopalan KV, Johnson MK (1997) J Am Chem Soc 119:2590–2591
Garton SD, Temple CA, Dhawan IK, Barber MJ, Rajagopalan KV, Johnson MK (2000) J Biol Chem 275:6798–6805
Ellis PJ, Conrads T, Hille R, Kuhn P (2001) Structure 9:125–132
Conrads T, Hemann C, George GN, Pickering IJ, Prince RC, Hille R (2002) J Am Chem Soc 124:11276–11277
Roy R, Adams MW (2002) Met Ions Biol Syst 39:673–697
Caradonna JP, Harlan EW, Holm RH (1986) J Am Chem Soc 108:7856–7858
Tierney DL, Gassner GT, Luchinat C, Bertini I, Ballou DP, Penner-Hahn JE (1999) Biochemistry 38:11051–11061
Bonaccio M, Ghaderi N, Borchardt D, Dunn MF (2005) Biochemistry 44:7656–7668
Bray RC, Adams B, Smith AT, Bennett B, Bailey S (2000) Biochemistry 39:11258–11269
Klein EL, Raitsimring AM, Astashkin AV, Rajapakshe A, Johnson-Winters K, Arnold AR, Potapov A, Goldfarb D, Enemark JH (2012) Inorg Chem 51:1408–1418
Burgrnayer SJN, Stlefel EI (1985) J Chem Edu 62:943–953
Harlan EE, Berg JM, Holm RH (1986) J Am Chem Soc 108:6992–7000
Arnoux P, Sabaty M, Alric J, Frangioni B, Guigliarelli B, Adriano JM, Pignol D (2003) Nat Struct Biol 10:928–934
Jepson BJN, Mohan S, Clarke TA, Gates AJ, Cole JA, Butler CS, Butt JN, Hemmings AM, Richardson DJ (2007) J Biol Chem 282:6425–6437
Andersong GL, Williamsll J, Hille R (1992) J Biol Chem 267:23674–23682
Cammack R, Barber MJ, Bray RC (1976) Biochem J 157:469–478
Moura JJ, Xavier AV, Cammack R, Hall DO, Bruschi M, Le Gall J (1978) Biochem J 173:419–425
Weber K, Creager SE (1994) Anal Chem 66:3164–3172
Enemark JH, Garner CD (1997) J Biol Inorg Chem 2:817–822
Schröder I, Rech S, Krafft T, Macy JM (1997) J Biol Chem 272:23765–23768
Watts CA, Ridley H, Dridge EJ, Leaver JT, Reilly AJ, Richardson DJ, Butler CS (2005) Biochem Soc Trans 33:173–175
Spuches AM, Kruszyna HG, Rich AM, Wilcox DE (2005) Inorg Chem 44:2964–2972
Hoke KR, Cobb N, Armstrong FA, Hille R (2004) Biochemistry 43:1667–1674
Acknowledgments
We acknowledge Fundação para a Ciência e Tecnologia (FCT) for the financial support Granted to REQUIMTE (PEst-C/EQB/LA0006/2013) and UCIBIO-REQUIMTE (UID/Multi/04378/2013), BKM (SFRH/BPD/63066/2009) and SRP (FCT-ANR/BBB-MET0023/2012). NMR data were collected on 400-MHz spectrometer that is part of the National NMR Network, also supported by FCT (RECI/BBB-BQB/0230/2012).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Maiti, B.K., Maia, L.B., Silveira, C.M. et al. Incorporation of molybdenum in rubredoxin: models for mononuclear molybdenum enzymes. J Biol Inorg Chem 20, 821–829 (2015). https://doi.org/10.1007/s00775-015-1268-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-015-1268-0