Abstract
Mammalian target of rapamycin complex 1 (mTORC1) is activated by amino acids to promote cell growth via protein synthesis. Specifically, Ras-related guanosine triphosphatases (Rag GTPases) are activated by amino acids, and then translocate mTORC1 to the surface of late endosomes and lysosomes. Ras homolog enriched in brain (Rheb) resides on this surface and directly activates mTORC1. Apart from the presence of intracellular amino acids, Rag GTPases and Rheb, other mediators involved in intracellular amino acid signaling to mTORC1 activation include human vacuolar sorting protein-34 (hVps34) and mitogen-activating protein kinase kinase kinase kinase-3 (MAP4K3). Those molecular links between mTORC1 and its mediators form a complicate signaling network that controls cellular growth, proliferation, and metabolism. Moreover, it is speculated that amino acid signaling to mTORC1 may start from the lysosomal lumen. In this review, we discussed the function of these mediators in mTORC1 pathway and how these mediators are regulated by amino acids in details.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mammalian target of rapamycin complex 1 (mTORC1) promotes cell growth and changes cell size by stimulating protein synthesis. Signals such as nutrients, growth factors, and energy levels can influence mTORC1 activity (Deng et al. 2009, 2010; Tan et al. 2010). Once receiving and integrating these upstream signals, mTORC1, in turn, can regulate various growth-related cellular processes, such as transcription, translation, and autophagy (Wullschleger et al. 2006). Among these signals, amino acids especially leucine are specific nutrients for protein synthesis and cell growth (Kim et al. 2007; Wu et al. 2007; Wu et al. 2009; Kim 2009; Yin and Tan 2010; Li et al. 2011; Yao et al. 2012). Furthermore, the importance of amino acids is reflected not only in its potently stimulatory ability but also in its indispensability to mTORC1 activation by other stimuli, such as growth factors (Kim and Guan 2011). However, it has been demonstrated that amino acids act independently of insulin and the tuberous sclerosis complex (TSC). Thus, the insulin/phosphoinositide-3-OH kinase (PI3 K) pathway may not be involved in amino acids signaling to mTORC1 (Efeyan and Sabatini 2013). The molecular mechanisms of amino acids regulate mTORC1 signaling pathway remain largely unknown. However, several important advances have been found in the study of amino acid-induced mTORC1 activation. First, amino acids are sensed by amino acid sensors to modulate protein synthesis or degradation through mTORC1 signaling. (Long et al. 2005b; Efeyan et al. 2012). Several key mediators have been found to play a critical role in relaying amino acid signals to mTOR activation. These mediators include the Ras-related guanosine triphosphatases (Rag GTPases), Ras homolog enriched in brain (Rheb), human vacuolar sorting protein-34 (hVps34), and mitogen-activating protein kinase kinase kinase kinase-3 (MAP4K3). In the signal pathway, lysosome serves as a platform for mTORC1 activation. In brief, activation of mTORC1 requires at least two regulated steps: translocation of mTORC1 to the surface of lysosome where Rheb resides, and activation of mTORC1 by Rheb (Demetriades et al. 2014). This review discusses current proposed mediators of intracellular amino acids signaling to mTORC1 activation with special emphasis on biochemical mechanisms.
mTORC1 signaling in cell growth control
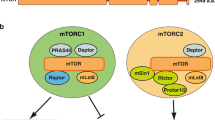
Cell growth determines the size of cells, organs and organisms (Li et al. 2004). mTOR signaling plays a key role in integrating a wide range of signals from growth factors, nutrients and energy status to regulate cell growth (Sarbassov et al. 2005). mTOR, a protein kinase, exists in two structurally and functionally distinct multi-protein complexes termed mTORC1 and mTORC2, respectively. mTORC1 positively mediates cell growth and is sensitive to rapamycin (Sarbassov et al. 2005; Betz and Hall 2013; Duan et al. 2015). Therefore, this review will focus on mTORC1 and the use of mTOR will refer solely to this complex. mTORC1 is composed of mTOR, regulatory-associated protein of mTOR (Raptor), mammalian LST8/G-protein β-subunit–like protein (mLST8/GβL), Deptor, proline-rich Akt/PKB substrate 40 kDa (PRAS40), and a member of the FK506-binding protein (FKBP) family, FKBP38 (Bai et al. 2007; Yang and Guan 2007; Betz and Hall 2013). Raptor is a scaffold protein. It plays a critical role in recruiting mTORC1 substrates, integrating various signals for mTOR modulation, and in stabilizing mTOR (Bai and Jiang 2010). Importantly, the raptor-mTOR interaction may be the target of upstream signals (De Virgilio and Loewith 2006). mLST8 may not be an indispensable component of mTORC1 function. It binds to mTOR kinase domain and activates the kinase activity independent of Raptor (Kim 2009). PRAS40 and FKBP38 act as negative regulators of mTORC1. FKBP38 suppresses mTORC1 activity through directly associating with mTORC1. Overexpression of PRAS40 and FKBP38 inhibits downstream targets of mTORC1 (Vander Haar et al. 2007; Bai et al. 2007). However, some other studies disagree with the role of FKBP38 as a negative regulator of mTORC1 (Wang et al. 2008; Maehama et al. 2008; Uhlenbrock et al. 2009).
Protein synthesis is thought to be one of the most highly regulated processes in cell growth (Gingras et al. 2001). Over the past years, modulation of mTORC1 on protein synthesis has been well-characterized and presented in a number of reviews (Duan et al. 2015). In brief, mTORC1 positively regulates protein synthesis and cell growth by phosphorylating the following two translational regulatory proteins: eukaryotic initiation factor 4E (eIF4E)-binding protein 1 (4E-BP1) and p70 ribosomal protein S6 kinase (S6K1) (Li et al. 2004; Inoki and Guan 2006; Arsham and Neufeld 2006; Yin et al. 2010).
Amino acids are required for activation of mTORC1. Other signals cannot activate mTORC1 in the case of lacking sufficient amino acids. And amino acids starvation can completely hinder mTORC1 activity, which cannot be compensated by other stimuli such as growth factors or energy (Gingras et al. 2001; Yang et al. 2013). The insulin/PI3 K pathway may not be involved in amino acids signaling to mTORC1 (Efeyan and Sabatini 2013). And the mechanism through which insulin activates mTOR is well-elucidated and has been shown in a number of reviews (Columbus et al. 2014). In particular, leucine, glutamine and arginine were identified as three important upstream mediators of the mTORC1 signaling pathway (Wang et al. 1998; Crespo et al. 2002; Yao et al. 2008; Tan et al. 2012; Kong et al. 2012; Yang et al. 2013). Therefore, this review will focus on amino acid-induced mTORC1 activation.
How intracellular amino acids are sensed
Before elaborating the function of these mediators (Rag GTPases, Rheb, hVps34, and MAP4K3) in mTORC1 activation and the regulation of these mediators by intracellular amino acids, we firstly find out how intracellular amino acids are sensed. Previous studies have presented evidence that intracellular amino acids are key regulators of mTORC1 (Christie et al. 2002; Beugnet et al. 2003; Avruch et al. 2009). Although the way that some amino acid (such as leucine) transporters transport amino acids from the intercellular space into the cell has been well-elucidated, the identity and location of the intracellular amino acid sensors implicated in the modulation of mTORC1 remains elusive (Wang and Proud 2009; Taylor 2009). Therefore, we will discuss proteins associated with intracellular amino acids sensing upstream of mTORC1 in detail.
Amino acid sensors in the cytosol
Several putative cytosolic amino acid sensors linked to mTORC1 activation have been identified. To our knowledge, they are mainly leucyl-tRNA synthetase (LRS) (Han et al. 2012), inositol polyphosphate multikinase (IPMK) (Kim et al. 2011), glutamate dehydrogenase (GDH) (Durán et al. 2012), Ras-like protein A (RalA) (Maehama et al. 2008), and unbranched chain amino acid receptors 1 and 2 (UBR1-2) (Kume et al. 2010). Some of them (such as LRS, GDH, and UBR1-2) are able to directly bind amino acids (Taylor 2014).
LRS is an intracellular amino acid sensor which is sensitive to leucine concentration. LRS induces mTORC1 activation in a leucine concentration-dependent manner. (Han et al. 2012). In yeast, LRS positively mediates TORC1 pathway: when leucine is available, LRS associates with Gtr1 (the RagA/B homolog) and activates it by an unknown regulator (Efeyan et al. 2012). In mammalian cells, LRS directly binds to GTP-bound RagD in a leucine-dependent manner and acts as a GTPase-activating protein (GAP) for Rag GTPase to activate mTORC1 (Fig. 1) (Han et al. 2012). Interaction of LRS with RagD relys on the nucleotide-binding state of RagD. As GTP-bound RagD inhibits mTORC1 activation, LRS appears to interact with the inactive Rag heterodimer, thereby facilitating LRS conversion to the GDP-bound form, and then dissociate from the active Rag heterodimer to activate mTORC1. Of note, it is not the tRNA charging activity of LRS but the leucine recognition function that is implicated in mTORC1 activation (Efeyan et al. 2012; Han et al. 2012).
The role of leucyl-tRNA synthetase (LRS), an amino acid sensor in the cytosol, in amino acid-induced Mammalian target of rapamycin complex 1 (mTORC1) activation. When leucine is present, LRS senses its concentration and binds to it. Then, LRS directly interacts with GTP-bound RagD, facilitating its conversion to the GDP-bound form, and then dissociates from the active Rag heterodimer to activate mTORC1
IPMK was originally identified in yeast as an essential gene affecting responses to arginine and therefore was labeled Arg82, and is also known as yeast IPMK (Bechet et al. 1970; Saiardi et al. 1999; Odom et al. 2000). In mammalian cells, IPMK, possessing both inositol phosphate kinase and lipid kinase activities, regulates the effects of amino acids stimulation on mTOR pathway. IPMK depletion inhibits amino acid-stimulated mTOR signaling and markedly diminishes mTOR–raptor interaction (Fig. 2) (Guertin et al. 2006). Of note, IPMK modulation of mTOR is not via catalytic activity but by the unique amino terminus of IPMK. IPMK serves as an mTOR cofactor. IPMK is proposed to act noncatalytically to selectively stabilize the interaction between mTOR and raptor. Leucine depletion enhancing interaction between IPMK, mTOR, and raptor, which is reversed by leucine stimulation. The observations indicate that leucine mediates the affinity rather than the stoichiometry of the interaction complex (Kim et al. 2011). Collectively, IPMK functions as a physiologic cofactor between mTOR and Raptor in the presence of sufficient amino acid. It stabilizes mTOR–Raptor association in the mTORC1 complex through its amino-terminal sequence.
Glutamine is converted to α-ketoglutarate (α-KG) through a process termed glutaminolysis. In this process, the enzymes glutaminase (GLS) and GDH are the key regulators (Durán et al. 2012). GDH is allosterically mediated by several factors, including leucine as an activator and GTP as a negative mediator (Frigerio et al. 2008; Li et al. 2012). Leucine directly associates with GDH and activates it, leading to glutamate deamination and hence α-KG production (Fig. 3) (Durán et al. 2012). Moreover, glutamine flux facilitates leucine uptake, which in turn modulates mTORC1 (Nicklin et al. 2009). Thus, leucine and glutamine cooperate in activating glutaminolysis and mTORC1. In other words, glutaminolysis is responsible for an actual sensing mechanism, for at least leucine and glutamine. RagB in the GTP-bound form promotes lysosomal recruitment of mTORC1 (discussed in detail below), and α-KG enhances GTP loading of RagB to activate mTORC1 (Fig. 3). Therefore, suppression of glutaminolysis (such as GDH activity) hinders the translocation of mTOR to the lysosome, preventing the activation of mTORC1 by leucine and glutamine. On the contrary, up-regulation of glutaminolysis enhances the response of mTORC1 to amino acids (Durán et al. 2012). Thus, glutaminolysis, in particular the transition of glutamate to α-KG catalyzed by GDH, activates mTORC1 by enhancing its sensitivity to amino acids and its recruitment to the lysosome by stimulating GTP loading of RagB. Collectively, Rag, and thus mTORC1 senses glutamine and leucine through GDH stimulating glutaminolysis and production of α-KG (Taylor 2014).
The role of glutamate dehydrogenase (GDH), an amino acid sensor in the cytosol, in amino acid-induced mTORC1 activation. Leucine directly associates with GDH and activates it, leading to glutamate deamination and hence α-ketoglutarate (α-KG) production. α-KG enhances GTP loading of RagB to activate mTORC1
RalA, a member of the Ras small G-protein superfamily, is proposed to be involved in amino acid-induced mTORC1 activation (Maehama et al. 2008). Of note, RalA may also localize to lysosome. Amino acids can regulate RalA by increasing the levels of GTP-bound RalA, and hence activate mTORC1. Moreover, RalA knockdown suppresses mTORC1 pathway in cells overexpressing a hyperactive mutant of Rheb without influencing its nucleotide-bound status, placing RalA downstream of Rheb. However, it is unclear whether RalA directly interacts with mTORC1 or with FKBP38 (Maehama et al. 2008; Dodd and Tee 2012). Therefore, it raises the possibility that amino acids may activate mTORC1 downstream from Rheb through RalA.
UBR1 and UBR2, E3 ubiquitin ligases, might be cellular targets of leucine. They specifically recognize the identity of N-terminal residues, contributing to selective destabilization of target proteins according to the N-end rule (Kume et al. 2010). Leucine directly associates with the substrate-recognition domain of UBR2 and prevents degradation of N-end rule substrates in vitro, which promotes signaling via mTORC1. Moreover, overexpression of UBR1 and UBR2 leads to a reduction of S6K1 phosphorylation and hence inhibits mTOR signaling, which could be rescued with high concentrations of leucine. However, knockdown of UBR1 and UBR2 enhances S6K1 phosphorylation. Thus, UBR1 and UBR2 function as leucine binding proteins and negative mediators of mTORC1, and leucine activates mTOR signaling at least in part through inhibiting ubiquitin ligase activity of UBR1 and UBR2 (Kume et al. 2010; Dodd and Tee 2012).
Amino acid sensors on endosomal membranes
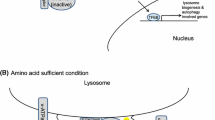
Once amino acid stimulation, mTORC1 is shuttled to the late endosomal and lysosomal compartments, where it associates with the Ragulator-Rag complex and is assembled into active mTORC1 (discussed below) (Goberdhan 2010). Therefore, amino acid transporters on endosomal such as lysosomal membranes may themselves function as intracellular amino acid sensors (Taylor 2014). The vacuolar H+-adenosine triphosphatase ATPase (v-ATPase) and solute carrier 36A1 (SLC36A1) H+-coupled amino acid transporter (aka PAT1) are almost exclusively anchored on lysosome. When sufficient amino acids are present, v-ATPase and PAT1 form the lysosomal-anchored “nutrisome” protein complex. They function as sensors of intralysosomal amino acid concentrations and physically bind to the Rag-Ragulator complex (Goberdhan 2010; Zoncu et al. 2011; ögmundsdottir et al. 2012). The v-ATPase is primarily responsible for pumping protons into the lysosome from the cytosol, along with hydrolyzing ATP to ADP. Therefore, v-ATPase helps to generate the acidic interior of the lysosomes compared with the slightly alkaline cytosol (pH 5–pH 7.4) (Taylor 2014). SLC36A1 regulates H+-dependent amino acid efflux from the lysosomal lumen into the cytosol. Also, SLC36A1 physically associates with the Rag GTPases and is essential to normal amino acid-dependent mTORC1 localization (Fig. 4) (Ögmundsdottir et al. 2012). When overexpressed in mammalian cell lines, SLC36A1 has a negative effect on lysosomal mTORC1 signaling (Zoncu et al. 2011). Taken together, the proton gradient generated by the v-ATPase is essential for amino acids to be transported into the subcellular compartment where the amino acid sensor SLC36A1 resides.
The molecular mechanism of the Rag guanosine triphosphatases (GTPases) in amino acid signaling to mTORC1 activation. The vacuolar H+-adenosine triphosphatase ATPase (v-ATPase) and solute carrier 36A1 (SLC36A1) H+-coupled amino acid transporter are almost exclusively anchored on lysosome. The v-ATPase is primarily responsible for pumping protons into the lysosome from the cytosol, generating a proton gradient across the lysosomal membrane. SLC36A1 physically associates with the Rag GTPases and regulates H+-dependent amino acid efflux from the lysosomal lumen into the cytosol. Then, these amino acids associates with Rag A/B, promoting Rag A/B GTP charging and activate it. Once Rag A/B activated, it is first localized to the lysosomal membranes through the v-ATPase-Ragulator complex. And then the Ragulator-Rag complex recruits mTORC1 by binding Raptor to the lysosomal membranes (where Ras homolog enriched in brain (Rheb) is localized) for activation
Key mediators involved in activating mTORC1 signaling by amino acids
Several mediators of amino acid signals have been identified to lie upstream of mTORC1, including the small GTPase Rag, Rheb, hVps34 and MAP4K3. Each protein activates mTORC1 through distinctive mechanisms.
Rag GTPases
Recent studies indicated that the Rag proteins, a family of four related small GTPases, play a crucial role in the modulation of mTORC1 activation in response to amino acids, especially in the spatial regulation of mTORC1 localization (Kim et al. 2008; Sancak et al. 2008; Suzuki and Inoki 2011). mTORC1 maybe translocated to the lysosomal membrane (which were previously termed Rab7-positive vesicles) in a fashion-dependent on Rag, whose activity is mediated by amino acid availability (Suzuki and Inoki 2011). Rag proteins consists of Rag A, B, C, and D. They function as heterodimers of Rag A/B combined with Rag C/D. Rag A/BGTP and Rag C/DGDP constitute the ‘active’ forms, which can associate with mTORC1 and potently activate mTORC1 signaling. Conversely, the ‘inactive’ forms (Rag A/BGDP − Rag C/DGTP) are unable to bind to mTORC1 and potently inhibit mTORC1 activity, even when amino acids are available (Sancak et al. 2010). Furthermore, the GTP-charged Rag B is regulated by amino acids. On amino acid starvation, the levels of GTP-charged Rag B are reduced (Kim et al. 2008; Sancak et al. 2008). Overall, the heterodimerization of Rag GTPases does not rely on amino acids, whereas the nucleotide loading status of them is mediated by amino acids through proteins as indicated (Yang et al. 2013).
How do Rag GTPases recruit mTORC1 to the lysosomal membrane? This process can be divided into at least two steps. In the first step, a trimeric protein complex named ‘Ragulator’ plays a key role. The Ragulator complex directly interacts with the Rag GTPases heterodimer and tethers it to the lysosomal surface. Moreover, the interaction between Rag GTPases and Ragulator, which is essential for proper intracellular localization and activation of mTORC1, is not influenced by amino acids (Sancak et al. 2010). The Ragulator complex consists of p18, p14, and MP1 (mitogen-activated protein kinase scaffold protein 1). p18 lies in the lysosomal membrane through N-terminal palmitoylation and myristoylation sites, and serves as a platform for associating p14 and MP1 to the lysosome (Nada et al. 2009). p14 and MP1 do not directly localize to the lysosomal membrane, but both proteins are required for the recruitment and activation of mTORC1 (Sancak et al. 2010). Lack of functions or one of the members of the MP1/p14/p18 complex failing to correctly localize Rag GTPases at the lysosomal membrane influences mTORC1 translocalization and its activity in response to amino acid stimulation (Sancak et al. 2010).
Once the Rag-Ragulator complex is assembled on the lysosomal membrane, a proton gradient generated by v-ATPase across the lysosomal membrane is required for activation of mTORC1 pathway (Zoncu et al. 2011; ögmundsdottir et al. 2012). It physically binds to Ragulator and indirectly links with Rag GTPases. Loss of v-ATPase components significantly inhibits amino acid-stimulated mTORC1 activation (Qi et al. 2007; Jefferies and Forgac 2008). Apart from Ragulator and v-ATPase, LRS, Adaptor protein p62, and SH3BP4 also bind and regulate Rag GTPases in response to amino acids, readers can find the details in elsewhere (Bonfils et al. 2012; Han et al. 2012; Yang et al. 2013). Collectively, Rag GTPases heterodimers are anchored at lysosomal membranes via the association with Ragulator complex (p18/MP1/p14) and v-ATPase (Yang et al. 2013).
In the second step, mTORC1 is recruited by activated Rag GTPases to the surface of lysosome, where mTORC1 is activated. More specifically, the active Rag heterodimers (Rag BGTP–Rag DGDP) associate with Raptor (an important component of mTORC1), and subsequently translocate mTORC1 to the surface of lysosome in response to amino acid stimulation (Sancak et al. 2008; Yang et al. 2013). The Rag–Raptor association is regulated by amino acids. The exact way of regulation is still unclear, but it is confirmed to exclude the modulation of Rag GTPase guanyl nucleotide charging (Oshiro et al. 2014). Rag heterodimers are constitutively expressed at LAMP2 (a late endosome or lysosome marker)—positive compartments. Interestingly, the amino acid-stimulated mTOR or Raptor localization at the LAMP2-positive compartments relies on the levels of GTP-bound form of Rag A/B (Sancak et al. 2010). Therefore, it raises the possibility that the interaction between the Rag heterodimer and Raptor relies on the levels of GTP-bound Rag A or Rag B in the heterodimer. Knockdown of Rag A/B or inhibiting Rag A/B expression markedly suppresses the mTORC1 signaling pathway. Furthermore, Amino acid stimulation induces co-localization of mTOR with LAMP2 (Sancak et al. 2008). Taken together, mTORC1 is finally recruited to lysosomal membranes in response to amino acids through the interaction between Rag GTPases and Raptor.
From the above, we know that when amino acid level is low, the v-ATPase-Ragulator-Rag GTPases complex is in the inactive conformation and has no ability to associate with mTORC1, causing its cytoplasmic localization. When amino acids are sufficient, Rag GTPases are firstly localized to the lysosomal membranes through the v-ATPase-Ragulator complex. That means the Ragulator-Rag complex functions as an amino acid-mediated docking site for mTORC1 on the surface of lysosome. Amino acids promote Rag A/B GTP charging, and then the Ragulator-Rag complex recruits mTORC1 by binding Raptor to the lysosomal membranes (where Rheb is localized) for activation (Fig. 4). The function of Rheb is discussed in details below.
Rheb
Previous descriptions suggest that the Rag proteins are mainly responsible for translocating mTORC1 to the surface of lysosome without activating mTORC1 directly. The activator of mTORC1 was proved to be the Rheb (Inoki et al. 2003; Sancak et al. 2008). Knockdown of Rheb in mammalian cells, amino acids are ineffective in activating mTORC1 (Roccio et al. 2006).
Rheb is a Ras-related GTPases, which can be rapidly induced by synaptic activity in brain neurons (Yamagata et al. 1994; Aspuria and Tamanoi 2004). A body of evidence suggests that Rheb is also involved in communicating amino acid sufficiency to mTORC1 (Kwiatkowski and Manning 2005). The Rheb small GTPase is a direct positive upstream mediator for mTORC1 activation (Manning and Cantley 2003). It still remains elusive in regard to whether endogenous active Rheb stimulates mTOR in vivo (Long et al. 2005a). However, purified active sufficient Rheb induces mTORC1 kinase activity in vitro. These observations indicate that the role of Rheb in the activation of mTORC1 has been widely accepted (Sato et al. 2009). Importantly, the Rheb–mTOR interaction, which is able to activate mTORC1 activity in vitro, may be mediated by amino acids (Long et al. 2005b; Joseph Avruch 2009). As is the case of other small GTPases, active Rheb is the GTP-bound form while inactive Rheb is the GDP-bound form (Suzuki and Inoki 2011). mTORC1 can be directly activated by Rheb-GTP rather than Rheb GDP, and the intact mTORC1 complex is essential to in vitro activation of mTORC1 by Rheb (Sancak et al. 2007). GTP loading of Rheb can be regulated by amino acids via a distinct GAP or GTPase exchange factor (GEF) (Nobukuni et al. 2005). On amino acid deprivation, Rheb is no more GTP loaded even following stimulation from insulin. However, resupplementing amino aicds to cells that have been deprived for 2 h significantly increases Rheb-GTP levels (Roccio et al. 2006). Additionally, the increase of the Rheb-GTP level serves as a permissive signal for mTOR activation (Roccio et al. 2006). Rheb-GTP can also be promoted by PKB/Akt inhibiting TSC1/2. However, this has not been a consistent finding. TSC1/2 complex (a direct upstream regulator of Rheb) is not implicated in the effect of amino acids on mTORC1 (Roccio et al. 2006). Moreover, PKB/Akt can directly phosphorylate PRAS40, and relieve the inhibition of PRAS40 (a negative regulator) to mTORC1 (Sancak et al. 2007; Vander Haar et al. 2007). Interestingly, there are some similar aspects of Rheb and Rag-dependent modulation of mTORC1. More specifically, they both localize to the lysosomal surface, and are both GTPases that change their nucleotide state upon physiological fluctuations of their upstream signals (Efeyan and Sabatini 2013).
On amino acid stimulation, mTOR and Rheb are present in Rab7-positive structures. It raises the possibility that amino acids may mediate mTORC1 activity via the event of its movement to lysosomal membranes (Sancak et al. 2008). Rheb activates mTORC1 directly or indirectly through at least two mechanisms. One mechanism requires a direct Rheb-mTORC1 interaction. When associated with GTP, Rheb binds directly to the mTOR catalytic domain, sequesters FKBP38 which otherwise associates with and suppresses mTORC1, and then activates mTORC1 (Fig. 4) (Laplante and Sabatini 2009; Joseph Avruch 2009; Oshiro et al. 2014). The other mechanism occurs though production of phosphatidic acid (PA). The PA biosynthetic pathway is an important downstream of Rheb to activate mTORC1 signaling (Fang et al. 2001, 2003; Sun et al. 2008). PA is the product of phosphatidylcholine (PC) hydrolysis by phospholipase D (PLD). Down-regulation of PLD or suppressing PA production prevents Rheb-induced mTORC1 activation (Sun et al. 2008). PA associates with the FKBP12-rapamycin-binding domain of mTOR, competing with the FKBP12-rapamycin complex for mTOR binding. Therefore, increased cellular PA level makes cells less sensitive to rapamycin (Chen et al. 2003). Rheb may connect PLD and activating mTORC1 in amino acid signaling. However, supplementing PA is unable to directly activate mTORC1 in vitro (Chen et al. 2003; Bjorklund et al. 2006). The mechanism by which amino acids regulate PLD activity remains unknown, but Vps34 may be involved in regulating PLD function. Vps34 product, PtdIns(3)P (PI(3)P), accumulates on the endosomal membrane and binds PLD to recruit it into the signaling platform where Rheb resides (Kim and Guan 2011). Other groups have demonstrated that the class III PI 3-kinase hVps34 relays amino acid sufficiency to mTORC1 independently of Rheb (Byfield et al. 2005; Nobukuni et al. 2005). Therefore, PLD may communicate the signals from amino acids and Vps34 to mTORC1, and Rheb may mediate the signals via enhancing PLD activity. The role of Vps34 in amino acid signaling is discussed in detail below.
hVps34
hVps34, a class III phosphatidylinositol-3-kinase, has also been proposed as a critical factor implicated in the regulation of amino acids-induced mTORC1 activation. On Vps34 deprivation, amino acids cannot increase S6 K phosphorylation in mammalian cells, while overexpression of Vps34 enhances S6 K phosphorylation when amino acids are available (Nobukuni et al. 2005; Byfield et al. 2005). These observations suggest that Vps34 functions upstream of mTORC1 in amino acid sensing.
The potential mechanism by which hVps34 regulates amino acid signaling in mTORC1 activation is reported by Gulati et al. (Gulati et al. 2008). They show that amino acids do not directly mediate hVps34 activity. Instead, amino acids enhance cellular Ca2+ levels by facilitating Ca2+ influx, and then activate Vps34. It is worth noting that lack of Ca2+ results in mTORC1 inactivation. Once amino acid stimulation, Vps34 elevates PI(3)P on endosomes, which provide a docking site for proteins containing phox homology (PX) or Fab1/YOTB/-2K632.12/Vac1/EEA1 (FYVE) domains. Therefore, PI(3)P may recruit one or more unknown PX- or FYVE-domain containing proteins to the endosome to build an mTORC1 ‘signalsosome’ for activation (Fig. 5). Vps34 associates with the late endosomal GTPase Rab7 and forms a stable complex with mTORC1. Vps34 may indirectly regulate the kinase activity of mTORC1 by influencing intracellular vesicle trafficking in response to amino acids. Previously, it has been indicated that amino acids change mTORC1 conformation to allow mTOR access and phosphorylate S6 K (activation) (Kim et al. 2002). Furthermore, activation of S6 K was identified to be associated with a Ca2+-dependent shift of the kinase into a large protein complex (Hannan et al. 2003). Ectopic expression of hVps34 enhances S6K1 phosphorylation only in the presence of amino acids (Nobukuni et al. 2005). Collectively, Vps34 is activated by cellular increased Ca2+ levels stimulated by amino acids and associates with the late endosomal GTPase Rab7, forming a stable complex with mTORC1. Vps34 product, PI(3)P, accumulates on endosomes and recruits PX- or FYVE-domain containing proteins to the endosome to build an mTORC1 ‘signalsosome’ for mTORC1 activation.
The molecular mechanism of the human vacuolar sorting protein-34 (hVps34) in amino acid signaling to mTORC1 activation. Vps34 is activated by cellular increased Ca2+ levels stimulated by amino acids and associates with the late endosomal GTPase Rab7, forming a stable complex with mTORC1. Vps34 product, PI(3)P, accumulates on endosomes and recruits PX- or FYVE-domain containing proteins to the endosome to build an mTORC1 ‘signalsosome’ for mTORC1 activation
MAP4K3
MAP4K3 has been proposed as a factor involved in the regulation of amino acid-induced mTORC1 activation. First, Amino acids can regulate MAP4K3 activity. Previous studies have shown that MAP4K3 activity and its activation of mTORC1 signaling require phosphorylation of Ser170 in the activation segment of MAP4K3. And PR61, a PP2A-targeting subunit, plays a crucial role in regulating MAP4K3 phosphorylation in response to amino acids (Fig. 6) (Yan et al. 2010; Suzuki and Inoki 2011). On amino acid deprivation, PP2A/PR61 interacts with MAP4K3, promoting dephosphorylation of Ser170 and suppressing MAP4K3 activation; whereas amino acid stimulation also causes rapid MAP4K3 activation (Findlay et al. 2007; Yan et al. 2010). The MAP4K3 activation mediated by amino acids occurs before phosphorylation of S6 K and is not suppressed by rapamycin, suggesting that MAP4K3 functions upstream of mTORC1 (Kim and Guan 2011). Moreover, loss of MAP4K3 functions in mammalian cells inhibits the stimulating effect of amino acid on mTORC1 activity, while gain of MAP4K3 functions increases mTORC1 activity and makes the mTORC1 signaling pathway insensitive to amino acids (Resnik-Docampo and de Celis 2011). Interestingly, inhibiting Rag C and Rag D impaired MAP4K3 ability to activate mTORC1, indicating that Rag proteins are implicated in the MAP4K3-mTORC1 axis (Kim and Guan 2011). However, MAP4K3 may not be the major mediator as Rag proteins relaying the amino acid signaling to mTORC1.
The molecular mechanism of the Mitogen-activating protein kinase kinase kinase kinase-3 (MAP4K3) in amino acid signaling to mTORC1 activation. On amino acid deprivation, PP2A/PR61 interacts with MAP4K3, promoting dephosphorylation of Ser170 and suppressing MAP4K3 activation; on amino acid stimulation, amino acids phosphorylate MAP4K3 in Ser170 and activate MAP4K3 as well as mTORC1
The importance of cell organelle lysosome
Lysosome is traditionally viewed as a vesicular organelle rich in degradative enzymes (lipases, proteases, hydrolases, etc.), where macromolecules are broken down (Efeyan et al. 2012). In other words, the lysosome is the end point of autophagy, generating lots of amino acids during nutrient starvation through degrading proteins and organelles (Yu et al. 2010). Therefore, mammalian lysosomes may maintain a stable pool of luminal amino acids (Harms et al. 1981). Recently, such amino acids accumulated by autophagy are found to activate mTORC1 (Yu et al. 2010). Therefore, the lysosome may be not only the end point of autophagy but also the starting point of amino acid signaling to mTORC1 (Efeyan et al. 2012). Notably, the observation that mTORC1 is active in the lysosome has significantly promoted our understanding of mTORC1 modulation.
The localization of the Rag GTPases is in lysosomes rather than other Rheb-containing endomembranes, suggesting that this organelle plays a crucial role in amino acid signaling to mTORC1 (Flinn et al. 2010; Zoncu et al. 2011). Furthermore, preventing the formation of late endosomes or redirecting the Ragulator to other cellular compartments blocks the activation of mTORC1. The evidence indicates that the specialized subcellular environment of late endosomes and lysosomes is required for mTORC1 activation (Sancak et al. 2010). In particular, the lysosome contains all machinery that is required for sensing amino acids and activating the Rag GTPases (Zoncu et al. 2011). Accumulating evidence suggests that amino acids may be sensed inside the lysosomal lumen, for alcohol esters of amino acids freely cross membranes and then accumulate inside lysosomes. Moreover, amino acids inside lysosomes generate an ‘inside-out’ signal that causes Rag activation, and v-ATPase is required for regulate the inside-out signal (Zoncu et al. 2011; Efeyan et al. 2012).
Summary and perspectives
Our basic knowledge of amino acid signaling to mTORC1 activation has been greatly expanded over the past years. Growing evidence shows that Rag GTPases, Rheb, hVps34, and MAP4K3 are key mediators involved in amino acid signaling and lysosome serves as the platform for mTORC1 activation. Of note, understanding the regulation of mTORC1 activation may provide new strategies for many metabolic diseases, such as insulin resistance, diabetes. mTORC1 plays many significant physiological and pathological implications in vivo. Its deregulation underlies the pathogenesis of many diseases (Efeyan et al. 2012). On the other hand, lysosome, a key mediator of cellular catabolism, is at the core of mTORC1 regulation by amino acids. Key questions include the following: First, how do amino acids subtly regulate the activity of Rag proteins? How do amino acids regulate the interaction between Rag proteins and Raptor? Second, apart from the lysosomes, can mTORC1 be also activated at other organelles? Last but not the least, given the fact that lysosome is the place autophagy occurs, then what is the physiological meaning of mTORC1 activation on the surface of lysosome? Further studies are required for clearly addressing these questions.
Abbreviations
- α-KG:
-
α-Ketoglutarate
- 4E-BP1:
-
4E-binding protein 1
- eIF4E:
-
Eukaryotic initiation factor 4E
- FKBP:
-
FK506-binding protein
- GAP:
-
GTPase-activating protein
- GDH:
-
Glutamate dehydrogenase
- GEF:
-
GTPase exchange factor
- hVps34:
-
Human vacuolar sorting protein-34
- IPMK:
-
Inositol polyphosphate multikinase
- LRS:
-
Leucyl-tRNA synthetase
- MAP4K3:
-
Mitogen-activating protein kinase kinase kinase kinase-3
- mLST8/GβL:
-
Mammalian LST8/G-protein β-subunit–like protein
- mTORC1:
-
Mammalian target of rapamycin complex 1
- PA:
-
Phosphatidic acid
- PC:
-
Phosphatidylcholine
- PI3K:
-
Phosphoinositide-3-OH kinase
- PI(3)P:
-
PtdIns(3)P
- PLD:
-
Phospholipase D
- PRAS40:
-
Proline-rich Akt/PKB substrate 40 kDa
- Rag GTPases:
-
Ras-related Guanosine triphosphatases
- RalA:
-
Ras-like protein A
- Raptor:
-
Regulatory-associated protein of mTOR
- Rheb:
-
Ras homolog enriched in brain
- S6K1:
-
p70 ribosomal protein S6 kinase
- SLC36A1:
-
Solute carrier 36A1
- TSC:
-
Tuberous sclerosis complex
- UBR1-2:
-
Unbranched chain amino acid receptors 1 and 2
- v-ATPase:
-
vacuolar H+-adenosine triphosphatase ATPase
References
Arsham AM, Neufeld TP (2006) Thinking globally and acting locally with TOR. Curr Opin Cell Biol 18(6):589–597
Aspuria PJ, Tamanoi F (2004) The Rheb family of GTP-binding proteins. Cell Signal 16(10):1105–1112
Avruch J, Long X, Ortiz-Vega S, Rapley J, Papageorgiou A, Dai N (2009) Amino acid regulation of TOR complex 1. Am J Physiol Endocrinol Metab 296(4):E592–E602
Bai X, Jiang Y (2010) Key factors in mTOR regulation. Cell Mol Life Sci 67(2):239–253
Bai XC, Ma DZ, Liu AL, Shen XY, Wang QJM, Liu YJ, Jiang Y (2007) Rheb activates mTOR by antagonizing its endogenous inhibitor, FKBP38. Science 318(5852):977–980
Bechet J, Greenson M, Wiame JM (1970) Mutations affecting the repressibility of arginine biosynthetic enzymes in Saccharomyces cerevisiae. Eur J Biochem 12(1):31–39
Betz C, Hall MN (2013) Where is mTOR and what is it doing there? J Cell Biol 203(4):563–574
Beugnet A, Tee AR, Taylor PM, Proud CG (2003) Regulation of targets of mTOR (mammalian target of rapamycin) signalling by intracellular amino acid availability. Biochem J 372(Pt 2):555–566
Bjorklund M, Taipale M, Varjosalo M, Saharinen J, Lahdenpera J, Taipale J (2006) Identification of pathways regulating cell size and cell-cycle progression by RNAi. Nature 439(7079):1009–1013
Bonfils G, Jaquenoud M, Bontron S, Ostrowicz C, Ungermann C, De Virgilio C (2012) Leucyl-tRNA synthetase controls TORC1 via the EGO complex. Mol Cell 46(1):105–110
Byfield MP, Murray JT, Backer JM (2005) hVps34 is a nutrient-regulated lipid kinase required for activation of p70 S6 kinase. J Biol Chem 280(38):33076–33082
Chen Y, Zheng Y, Foster DA (2003) Phospholipase D confers rapamycin resistance in human breast cancer cells. Oncogene 22(25):3937–3942
Christie GR, Hajduch E, Hundal HS, Proud CG, Taylor PM (2002) Intracellular sensing of amino acids in Xenopus laevis oocytes stimulates p70 S6 kinase in a target of rapamycin-dependent manner. J Biol Chem 277(12):9952–9957
Columbus DA, Fiorotto ML, Davis TA (2014) Leucine is a major regulator of muscle protein synthesis in neonates. Amino Acids
Crespo JL, Powers T, Fowler B, Hall MN (2002) The TOR-controlled transcription activators GLN3, RTG1, and RTG3 are regulated in response to intracellular levels of glutamine. Proc Natl Acad Sci 99(10):6784–6789
De Virgilio C, Loewith R (2006) The TOR signalling network from yeast to man. Int J Biochem Cell Biol 38(9):1476–1481
Demetriades C, Doumpas N, Teleman AA (2014) Regulation of TORC1 in Response to Amino Acid Starvation via Lysosomal Recruitment of TSC2. Cell 156(4):786–799
Deng D, Yao K, Chu WY, Li TJ, Huang RL, Yin YL, Liu HQ, Zhang JS, Wu GY (2009) Impaired translation initiation activation and reduced protein synthesis in weaned piglets fed a low-protein diet. J Nutr Biochem 20:544–552
Deng J, Wu X, Bin S, Li TJ, Huang R, Liu Z, Liu Y, Ruan Z, Deng Z, Hou Y, Yin YL (2010) Dietary amylose and amylopectin ratio and resistant starch content affects plasma glucose, lactic acid, hormone levels1and protein synthesis in splanchnic tissues. Zeitschrift für Tierphysiologie, Tierernährung und Futtermittelkunde 94:220–226
Dodd KM, Tee AR (2012) Leucine and mTORC1: a complex relationship. Am J Physiol Endocrinol Metab 302(11):E1329–E1342
Duan YH, Li FN, Liu HN, Li YH, Liu YY, Kong XF, Zhang YZ, Deng D, Tang YL, Feng ZM, Wu GY, Yin YL (2015) Nutritional and regulatory roles of leucine in muscle growth and fat reduction. Front Biosci (Landmark) 20:796–813
Durán RV, Oppliger W, Robitaille AM, Heiserich L, Skendaj R, Gottlieb E, Hall MN (2012) Glutaminolysis activates Rag-mTORC1 signaling. Mol Cell 47(3):349–358
Efeyan A, Sabatini DM (2013) Nutrients and growth factors in mTORC1 activation. Biochem Soc Trans 41(4):902–905
Efeyan A, Zoncu R, Sabatini DM (2012) Amino acids and mTORC1: from lysosomes to disease. Trends Mol Med 18(9):524–533
Fang Y, Vilella-Bach M, Bachmann R, Flanigan A, Chen J (2001) Phosphatidic acid-mediated mitogenic activation of mTOR signaling. Science 294(5548):1942–1945
Fang Y, Park IH, Wu AL, Du G, Huang P, Frohman MA, Walker SJ, Brown HA, Chen J (2003) PLD1 regulates mTOR signaling and mediates Cdc42 activation of S6K1. Curr Biol 13(23):2037–2044
Findlay GM, Yan L, Procter J, Mieulet V, Lamb RF (2007) A MAP4 kinase related to Ste20 is a nutrient-sensitive regulator of mTOR signalling. Biochem J 403:13–20
Flinn RJ, Yan Y, Goswami S, Parker PJ, Backer JM (2010) The Late Endosome is Essential for mTORC1 Signaling. Mol Biol Cell 21(5):833–841
Frigerio F, Casimir M, Carobbio S, Maechler P (2008) Tissue specificity of mitochondrial glutamate pathways and the control of metabolic homeostasis. Bba-Bioenergetics 1777(7–8):965–972
Gingras AC, Raught B, Sonenberg N (2001) Regulation of translation initiation by FRAP/mTOR. Genes Dev 15(7):807–826
Goberdhan DC (2010) Intracellular amino acid sensing and mTORC1-regulated growth. Curr Opin Investig Drugs 11(12):1360–1367
Guertin DA, Stevens DM, Thoreen CC, Burds AA, Kalaany NY, Moffat J, Brown M, Fitzgerald KJ, Sabatini DM (2006) Ablation in mice of the mTORC components raptor, rictor, or mLST8 reveals that mTORC2 is required for signaling to Akt-FOXO and PKC alpha but not S6K1. Dev Cell 11(6):859–871
Gulati P, Gaspers LD, Dann SG, Joaquin M, Nobukuni T, Natt F, Kozma SC, Thomas AP, Thomas G (2008) Amino acids activate mTOR complex 1 via Ca2 +/CaM signaling to hVps34. Cell Metab 7(5):456–465
Han JM, Jeong SJ, Park MC, Kim G, Kwon NH, Kim HK, Ha SH, Ryu SH, Kim S (2012) Leucyl-tRNA synthetase is an intracellular leucine sensor for the mTORC1-signaling pathway. Cell 149(2):410–424
Hannan KM, Thomas G, Pearson RB (2003) Activation of S6K1 (p70 ribosomal protein S6 kinase 1) requires an initial calcium-dependent priming event involving formation of a high-molecular-mass signalling complex. Biochem J 370(Pt 2):469–477
Harms E, Gochman N, Schneider JA (1981) Lysosomal pool of free-amino acids. Biochem Biophys Res Commun 99(3):830–836
Inoki K, Guan KL (2006) Complexity of the TOR signaling network. Trends Cell Biol 16(4):206–212
Inoki K, Li Y, Xu T, Guan KL (2003) Rheb GTPase is a direct target of TSC2 GAP activity and regulates mTOR signaling. Genes Dev 17(15):1829–1834
Jefferies KC, Forgac M (2008) Subunit H of the vacuolar (H +) ATPase inhibits ATP hydrolysis by the free V1 domain by interaction with the rotary subunit F. J Biol Chem 283(8):4512–4519
Joseph Avruch XL, Ortiz-Vega Sara, Rapley Joseph, Papageorgiou Angela, Dai Ning (2009) Amino acid regulation of TOR complex 1. Am J Physiol Endocrinol Metab 296:592–602
Kim E (2009) Mechanisms of amino acid sensing in mTOR signaling pathway. Nutr Res pract 3(1):64–71
Kim J, Guan KL (2011) Amino acid signaling in TOR activation. Annu Rev Biochem 80:1001–1032
Kim DH, Sarbassov DD, Ali SM, King JE, Latek RR, Erdjument-Bromage H, Tempst P, Sabatini DM (2002) mTOR interacts with Raptor to form a nutrient-sensitive complex that signals to the cell growth machinery. Cell 110(2):163–175
Kim SW, Mateo RD, Yin YL, Wu GY (2007) Functional amino acids and fatty acids for enhancing production performance of sows and piglets. Asian Austral J Anim 20(2):295–306
Kim E, Goraksha-Hicks P, Li L, Neufeld TP, Guan KL (2008) Regulation of TORC1 by Rag GTPases in nutrient response. Nat Cell Biol 10(8):935–945
Kim S, Kim SF, Maag D, Maxwell MJ, Resnick AC, Juluri KR, Chakraborty A, Koldobskiy MA, Cha SH, Barrow R, Snowman AM, Snyder SH (2011) Amino acid signaling to mTOR mediated by inositol polyphosphate multikinase. Cell Metab 13(2):215–221
Kong X, Tan B, Yin Y, Gao H, Li X, Jaeger LA, Bazer FW, Wu G (2012) l-Arginine stimulates the mTOR signaling pathway and protein synthesis in porcine trophectoderm cells. J Nutr Biochem 23(9):1178–1183
Kume K, Iizumi Y, Shimada M, Ito Y, Kishi T, Yamaguchi Y, Handa H (2010) Role of N-end rule ubiquitin ligases UBR1 and UBR2 in regulating the leucine-mTOR signaling pathway. Genes Cells 15(4):339–349
Kwiatkowski DJ, Manning BD (2005) Tuberous sclerosis: a GAP at the crossroads of multiple signaling pathways. Hum Mol Genet 14:R251–R258
Laplante M, Sabatini DM (2009) mTOR signaling at a glance. J Cell Sci 122(20):3589–3594
Li Y, Corradetti MN, Inoki K, Guan KL (2004) TSC2: filling the GAP in the mTOR signaling pathway. Trends Biochem Sci 29(1):32–38
Li F, Yin Y, Tan B, Kong X, Wu G (2011) Leucine nutrition in animals and humans: mTOR signaling and beyond. Amino Acids 41(5):1185–1193
Li M, Li CH, Allen A, Stanley CA, Smith TJ (2012) The structure and allosteric regulation of mammalian glutamate dehydrogenase. Arch Biochem Biophys 519(2):69–80
Long X, Lin Y, Ortiz-Vega S, Yonezawa K, Avruch J (2005a) Rheb binds and regulates the mTOR kinase. Curr Biol 15(8):702–713
Long X, Ortiz-Vega S, Lin Y, Avruch J (2005b) Rheb binding to mammalian target of rapamycin (mTOR) is regulated by amino acid sufficiency. J Biol Chem 280(25):23433–23436
Maehama T, Tanaka M, Nishina H, Murakami M, Kanaho Y, Hanada K (2008) RalA Functions as an Indispensable Signal Mediator for the Nutrient-sensing System. J Biol Chem 283(50):35053–35059
Manning BD, Cantley LC (2003) Rheb fills a GAP between TSC and TOR. Trends Biochem Sci 28(11):573–576
Nada S, Hondo A, Kasai A, Koike M, Saito K, Uchiyama Y, Okada M (2009) The novel lipid raft adaptor p18 controls endosome dynamics by anchoring the MEK-ERK pathway to late endosomes. EMBO J 28(5):477–489
Nicklin P, Bergman P, Zhang B, Triantafellow E, Wang H, Nyfeler B, Yang H, Hild M, Kung C, Wilson C, Myer VE, MacKeigan JP, Porter JA, Wang YK, Cantley LC, Finan PM, Murphy LO (2009) Bidirectional transport of amino acids regulates mTOR and autophagy. Cell 136(3):521–534
Nobukuni T, Joaquin M, Roccio M, Dann SG, Kim SY, Gulati P, Byfield MP, Backer JM, Natt F, Bos JL, Zwartkruis FJ, Thomas G (2005) Amino acids mediate mTOR/raptor signaling through activation of class 3 phosphatidylinositol 3OH-kinase. Proc Natl Acad Sci 102(40):14238–14243
Odom AR, Stahlberg A, Wente SR, York JD (2000) A role for nuclear inositol 1,4,5-trisphosphate kinase in transcriptional control. Science 287(5460):2026–2029
Ögmundsdottir MH, Heublein S, Kazi S, Reynolds B, Visvalingam SM, Shaw MK, Goberdhan DC (2012) Proton-assisted amino acid transporter PAT1 complexes with Rag GTPases and activates TORC1 on late endosomal and lysosomal membranes. PLoS ONE 7(5):e36616
Oshiro N, Rapley J, Avruch J (2014) Amino Acids Activate Mammalian Target of Rapamycin (mTOR) Complex 1 without Changing Rag GTPase Guanyl Nucleotide Charging. J Biol Chem 289(5):2658–2674
Qi J, Wang Y, Forgac M (2007) The vacuolar (H +)-ATPase: subunit arrangement and in vivo regulation. J Bioenerg Biomembr 39(5–6):423–426
Resnik-Docampo M, de Celis JF (2011) MAP4K3 Is a Component of the TORC1 Signalling Complex that Modulates Cell Growth and Viability in Drosophila melanogaster. PLoS ONE 6(1):e14528
Roccio M, Bos JL, Zwartkruis FJ (2006) Regulation of the small GTPase Rheb by amino acids. Oncogene 25(5):657–664
Saiardi A, Erdjument-Bromage H, Snowman AM, Tempst P, Snyder SH (1999) Synthesis of diphosphoinositol pentakisphosphate by a newly identified family of higher inositol polyphosphate kinases. Curr Biol 9(22):1323–1326
Sancak Y, Thoreen CC, Peterson TR, Lindquist RA, Kang SA, Spooner E, Carr SA, Sabatini DM (2007) PRAS40 is an insulin-regulated inhibitor of the mTORC1 protein kinase. Mol Cell 25(6):903–915
Sancak Y, Peterson TR, Shaul YD, Lindquist RA, Thoreen CC, Bar-Peled L, Sabatini DM (2008) The Rag GTPases bind raptor and mediate amino acid signaling to mTORC1. Science 320(5882):1496–1501
Sancak Y, Bar-Peled L, Zoncu R, Markhard AL, Nada S, Sabatini DM (2010) Ragulator-Rag complex targets mTORC1 to the lysosomal surface and is necessary for its activation by amino acids. Cell 141(2):290–303
Sarbassov DD, Ali SM, Sabatini DM (2005) Growing roles for the mTOR pathway. Curr Opin Cell Biol 17(6):596–603
Sato T, Nakashima A, Guo L, Tamanoi F (2009) Specific activation of mTORC1 by Rheb G-protein in vitro involves enhanced recruitment of its substrate protein. J Biol Chem 284(19):12783–12791
Sun Y, Fang Y, Yoon MS, Zhang C, Roccio M, Zwartkruis FJ, Armstrong M, Brown HA, Chen J (2008) Phospholipase D1 is an effector of Rheb in the mTOR pathway. Proc Natl Acad Sci 105(24):8286–8291
Suzuki T, Inoki K (2011) Spatial regulation of the mTORC1 system in amino acids sensing pathway. Acta Biochim Biophys Sin 43(9):671–679
Tan BE, Yin YL, Kong XF, Li P, Li XL, Gao HJ, Li XG, Huang RL, Wu GY (2010) L-Arginine stimulates proliferation and prevents endotoxin-induced death of intestinal cells. Amino Acids 38:1227–1235
Tan BE, Li XG, Yin YL, Wu ZL, Liu C, Tekwe CD, Wu GY (2012) Regulatory roles for L-arginine in reducing white adipose tissue. Front Biosci-Landmrk 17:2237–2246
Taylor HSHaPM (2009) Amino acid transceptors: gate keepers of nutrient exchange and regulators of nutrient signaling. Am J Physiol Endocrinol Metab 296:E603–E613
Taylor PM (2014) Role of amino acid transporters in amino acid sensing. Am J Clin Nutr 99(1):223S–230S
Uhlenbrock K, Weiwad M, Wetzker R, Fischer G, Wittinghofer A, Rubio I (2009) Reassessment of the role of FKBP38 in the Rheb/mTORC1 pathway. FEBS Lett 583(6):965–970
Vander Haar E, Lee S, Bandhakavi S, Griffin TJ, Kim DH (2007) Insulin signalling to mTOR mediated by the Akt/PKB substrate PRAS40. Nat Cell Biol 9(3):316–323
Wang X, Proud CG (2009) Nutrient control of TORC1, a cell-cycle regulator. Trends Cell Biol 19(6):260–267
Wang XM, Campbell LE, Miller CM, Proud CG (1998) Amino acid availability regulates p70 S6 kinase and multiple translation factors. Biochem J 334:261–267
Wang XM, Fonseca BD, Tang H, Liu R, Elia A, Clemens MJ, Bommer UA, Proud CG (2008) Re-evaluating the Roles of Proposed Modulators of Mammalian Target of Rapamycin Complex 1 (mTORC1) Signaling. J Biol Chem 283(45):30482–30492
Wu GY, Bazer FW, Davis TA, Jaeger LA, Johnson GA, Kim SW, Knabe DA, Meininger CJ, Spencer TE, Yin YL (2007) Important roles for the arginine family of amino acids in swine nutrition and production. Livest Sci 112(1–2):8–22
Wu GY, Bazer FW, Davis TA, Kim SW, Li P, Rhoads JM, Satterfield MC, Smith SB, Spencer TE, Yin YL (2009) Arginine metabolism and nutrition in growth, health and disease. Amino Acids 37(1):153–168
Wullschleger S, Loewith R, Hall MN (2006) TOR signaling in growth and metabolism. Cell 124(3):471–484
Yamagata K, Sanders LK, Kaufmann WE, Yee W, Barnes CA, Nathans D, Worley PF (1994) rheb, a growth factor- and synaptic activity-regulated gene, encodes a novel Ras-related protein. J Biol Chem 269(23):16333–16339
Yan LJ, Mieulet V, Burgess D, Findlay GM, Sully K, Procter J, Goris J, Janssens V, Morrice NA, Lamb RF (2010) PP2A(T61 epsilon) Is an Inhibitor of MAP4K3 in Nutrient Signaling to mTOR. Mol Cell 37(5):633–642
Yang Q, Guan KL (2007) Expanding mTOR signaling. Cell Res 17(8):666–681
Yang H, Gong R, Xu Y (2013) Control of cell growth: Rag GTPases in activation of TORC1. Cell Mol Life Sci 70(16):2873–2885
Yao K, Yin YL, Chu WY, Li ZQ, Deng D, Li TJ, Huang RL, Zhang JS, Tan B, Wang W, Wu G (2008) Dietary arginine supplementation increases mTOR signaling activity in skeletal muscle of neonatal pigs. J Nutr 138(5):867–872
Yao K, Yin YL, Li XL, Xi PB, Wang JJ, Lei J, Hou YQ, Wu GY (2012) Alpha-ketoglutarate inhibits glutamine degradation and enhances protein synthesis in intestinal porcine epithelial cells. Amino Acids 42(6):2491–2500
Yin Y, Tan B (2010) Manipulation of dietary nitrogen, amino acids and phosphorus to reduce environmental impact of swine production and enhance animal health. J Food Agric Environ 8(3–4):447–462
Yin Y, Yao K, Liu Z, Gong M, Ruan Z, Deng D, Tan B, Liu Z, Wu G (2010) Supplementing l-leucine to a low-protein diet increases tissue protein synthesis in weanling pigs. Amino Acids 39(5):1477–1486
Yu L, McPhee CK, Zheng L, Mardones GA, Rong Y, Peng J, Mi N, Zhao Y, Liu Z, Wan F, Hailey DW, Oorschot V, Klumperman J, Baehrecke EH, Lenardo MJ (2010) Termination of autophagy and reformation of lysosomes regulated by mTOR. Nature 465(7300):942–946
Zoncu R, Bar-Peled L, Efeyan A, Wang S, Sancak Y, Sabatini DM (2011) mTORC1 senses lysosomal amino acids through an inside-out mechanism that requires the vacuolar H(+)-ATPase. Science 334(6056):678–683
Acknowledgments
The work in our laboratories was jointly supported by grants from the National Basic Research Program of China (2013CB127305, 2012CB124704), National Natural Science Foundation of China (31372325, 31110103909, and 31330075), The Chinese Academy of Science Visiting Professorships for Senior International Scientists (2013T2S0012), and Texas A&M AgriLife Research Hatch project (H-8200).
Conflict of interest
The authors declare that they have no competing financial interests.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Handling Editor: M. Hengstschlaeger.
Rights and permissions
About this article
Cite this article
Duan, Y., Li, F., Tan, K. et al. Key mediators of intracellular amino acids signaling to mTORC1 activation. Amino Acids 47, 857–867 (2015). https://doi.org/10.1007/s00726-015-1937-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-015-1937-x