Abstract
The OsFBT4 belongs to a small sub-class of rice F-box proteins called TLPs (Tubby-like proteins) containing the conserved N-terminal F-box domain and a C-terminal Tubby domain. These proteins have largely been implicated in both abiotic and biotic stress responses, besides developmental roles in plants. Here, we investigated the role of OsFBT4 in abiotic stress signalling. The OsFBT4 transcript was strongly upregulated in response to different abiotic stresses in rice, including exogenous ABA. When ectopically expressed, in Arabidopsis, under a constitutive CaMV 35S promoter, the overexpression (OE) caused hypersensitivity to most abiotic stresses, including ABA, during seed germination and early seedling growth. At the 5-day-old seedling growth stage, the OE conferred tolerance to all abiotic stresses. The OE lines displayed significant tolerance to salinity and water deficit at the mature growth stage. The stomatal size and density were seen to be altered in the OE lines, accompanied by hypersensitivity to ABA and hydrogen peroxide (H2O2) and a reduced water loss rate. Overexpression of OsFBT4 caused upregulation of several ABA-regulated/independent stress-responsive genes at more advanced stages of growth, showing wide and intricate roles played by OsFBT4 in stress signalling. The OsFBT4 showed interaction with several OSKs (Oryza SKP1 proteins) and localized to the plasma membrane (PM). The protein translocates to the nucleus, in response to oxidative and osmotic stresses, but failed to show transactivation activity in the yeast system. The OE lines also displayed morphological deviations from the wild-type (WT) plants, suggesting a role of the gene also in plant development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Tubby protein was discovered in mouse and is a small group of 4–5 members, called TLPs (Tubby-like proteins) in mammals. The TLP family plays vast and indispensable roles in mammals, ranging from retinal maintenance, neuronal cell development, regulation of insulin pathway, spinal cord development during embryogenesis, skeletal development, stress signalling, vesicle trafficking, and so on (Boggon et al. 1999; Mukhopadhyay and Jackson 2011). These proteins, almost entirely constituted of a large C-terminal Tubby domain acts as a bi-partite transcription factor possessing both DNA binding and transcriptional activation properties. They localize to the inner side of the plasma membrane (PM) and translocate to the nucleus in response to extracellular signals, where they function as a transcription factor (Boggon et al. 1999; Santagata et al. 2001). Later, the TLPs were identified in diverse eukaryotic organisms, like humans, C. elegans, Drosophila, and chicken. Each having 4–5 members (Nishina et al. 1998; Heikenwälder et al. 2001).
The plant TLP family is more expanded and was first identified in Arabidopsis with 11 members (Lai et al. 2004), and later in several other plants, viz. fourteen in rice, eleven in poplar, nine in apple etc. (Yang et al. 2008; Kou et al. 2009; Xu et al. 2016). This is indicative of a more diversified and yet basic role of the plant TLPs. In plants, the TLPs are part of a highly expanded F-box family of proteins, as they possess an N-terminal F-box motif coupled to the C-terminal Tubby domain (Lai et al. 2004; Kou et al. 2009), which is not the case with the non-plant TLPs, having undefined N-terminal. The F-box proteins are part of the SCF (SKP1-Cullin1-F-box)-type E3 ligase that ubiquitinates the target proteins for their 26-S proteasome-mediated degradation and have been reported to be involved in all aspects of plant development (Vierstra. 2009). The plant TLPs might show functional divergence from the rest of the eukaryotes as, although they have been shown to possess dsDNA binding properties (Wardhan et al. 2012), but may not possess a strong transcriptional activation activity, unlike other eukaryotic TLPs (Lai et al. 2004; Wardhan et al. 2012). Moreover, they may be involved in the ubiquitin-mediated protein degradation pathway as a newly acquired plant-specific function (Bao et al. 2014). Plant TLPs have been shown to bind SKP1 (S-phase kinase-associated protein 1) subunit (also known as OSKs in rice and ASKs in Arabidopsis) of the SCF-E3 ligase complex and other target proteins with the help of F-box motif alone or in combination with the Tubby domain, but DNA binding is the sole property of Tubby domain (Du et al. 2014; Lai and Shaw 2012; Bao et al. 2014; Wardhan et al. 2016).
TLPs from several plants have been shown to display widespread expression pattern, both temporally and spatially, across all vegetative and reproductive stages of growth suggesting their basic as well as pleiotropic roles (Lai et al. 2004; Kou et al. 2009; Wardhan et al. 2012; Xu et al. 2016; Li et al. 2020; Zhang et al. 2020). Several studies on the functional characterization of plant TLPs have shown their diverse roles in plant development, abiotic stress signalling, hormone signalling, plant-pathogen interactions, seed coat mucilage biosynthesis, fruit ripening, and so on (Kou et al. 2009; Reitz et al. 2012; Bao et al. 2014; Wang et al. 2019; Xu et al. 2019; Zhang et al. 2020; Li et al. 2020, 2021). TLPs across a wide range of plants, such as Arabidopsis, apple, cassava, cotton, tomato, cucumber, and so on, have been shown to be extensively involved in abiotic stress signalling (Kou et al. 2009; Reitz et al. 2012; Wardhan et al. 2012; Xu et al. 2016; Dong et al. 2019; Li et al. 2020, 2021; Bano et al. 2021).
There are 14 TLP genes in rice (classified and designated as FBTs), viz. OsFBT1 to OsFBT14 (Jain et al. 2007; Yang et al. 2008). As true for other plant TLPs, the OsFBTs also exhibit differential expression in different tissues and at different stages of development (Liu 2008; Kou et al. 2009). The transcript level of OsFBT4 was found to be in abundance in all tissues and moreover, it expressed at a relatively higher basal level across tissues, compared to the other OsFBTs (Liu 2008; Kou et al. 2009; Yang et al. 2008), suggesting important roles. All OsFBTs, including OsFBT4, were shown to be upregulated in response to blight-causing bacteria, Xanthomonas oryzae, and also by wounding, showing that they are involved in plant–pathogen interaction (Kou et al. 2009). OsFBT12/ OsTLP2 was shown to interact with the pathogen-responsive element of the OsWRKY13 promoter and regulate its transcription, which plays an important role in defense response to pathogen attack (Cai et al. 2008).
Since several plant TLPs have been implicated in abiotic stress responses, including CaTLP1 and AtTLP2, the putative orthologs of OsFBT4 (showing 49.7% and 54% amino acid sequence similarities, respectively, at the protein level), we studied the possible role of OsFBT4 in abiotic stress signaling. Rice productivity is severely affected by water scarcity, high temperatures, and salinity, frequently encountered by this crop plant, and hence, it becomes essential to study stress pathways for future improvements. The stress response of the OsFBT4-OE lines was studied in Arabidopsis at the seed germination, 5-day-old seedling, and mature stages of growth. As was expected, OsFBT4 overexpression conferred tolerance to salt and dehydration stresses at a mature stage but caused hypersensitivity to abiotic stresses at the seed germination stage. At the seedling growth stage, tolerance was conferred by the overexpression of the gene to most stresses. The responses seemed to be both ABA dependent and independent. The OE lines showed altered behavior in response to exogenous ABA, and also exhibited differential expression of several ABA and stress-responsive genes.
Materials and methods
Plant materials and growth conditions
For all rice-related work, the PB1 variety of indica rice was used. Seeds were surface sterilized using 0.1% mercuric chloride and placed on ½ MS (Murashige and Skoog) salts supplemented with 1% sucrose and 0.2% phytagel (Sigma) for gelling, in 50-ml culture tubes. The rice plants were grown in the rice culture room conditions, i.e., 16 h light and 8 h dark at 28–30 °C, with 100–120 µmol m−2 s−1 light intensity. For hydroponics, a liquid rice growth medium (Yoshida et al. 1976) was used. The plant material used for transformation and stress analysis was Arabidopsis thaliana, ecotype Col-0. The seeds were surface sterilized using 2% solution of sodium hypochlorite. The seedlings/plants were grown in growth room conditions, i.e., 16 h light and 8 h dark at 22 °C, with 100–120 µmol m−2 s−1 light intensity. One-half MS salts supplemented with 1% sucrose and 0.8% agar (Qualigens) for gelling was used whenever MS basal medium is mentioned in this study. For mature plants, a mix of 50% soilrite and 50% coco peat, saturated with OS medium (Ogren and Somerville) was used. Transformation of Arabidopsis was performed by floral dip method (Clough and Bent 1998), using the GV3101 Agrobacterium strain.
Tissue sampling and RT-qPCR analysis
For expression profiling of OsFBT4 under stress treatments in rice, the month-old plants grown hydroponically were transferred to stress agents in separate culture tubes as follows: salinity stress (200 mM NaCl solution), oxidative stress (1% H2O2 solution), ABA (50 µM in 0.1% ethanol); for drought stress gradual drying of the plants was performed on the Whatman sheet. For expression profiling of stress marker genes in Arabidopsis, the 3-day-old seedlings and 15-day-old plants, raised in MS basal petri plates, were harvested. The RT-qPCR analysis was performed twice (n = 2). The RNA isolation was done using Trizol reagent (Sigma), as per the manufacturer’s protocol. The cDNA synthesis was done with 1 µg of total RNA samples using the High-Capacity cDNA Reverse Transcription kit (Applied Biosystems). The RT-qPCR analysis was carried out using 2X Roche SYBR Green I master mix on a Roche Light Cycler 480 II instrument as per the manufacturer’s protocol.
Stress treatments and physiological assays
All stress assays were performed involving sub-lethal and gradual stress conditions to mimic environmental stress, allowing sufficient time to mount maximal responses.
Percentage germination assays
The seeds were plated directly on MS basal supplemented with stress agents; NaCl (150 mM), Mannitol (300 mM), ABA (3 µM), and 3 µM paraquat (Methyl viologen). The emergence of the radicle was scored as germinated, 24 h onwards till day 5. Cotyledon greening was scored on day 6. The assay was performed in three sets (n = 3), where each set consisted of three subsets of 30 seeds each.
Arabidopsis seedling growth assays on stress agents
The 5-day-old seedlings, raised on MS basal in a vertical position were transferred to MS basal supplemented with stress agents; 300 mM mannitol, 150 mM NaCl, 10 µM ABA, 1 µM paraquat (methyl viologen), and 40% polyethylene glycol 6000 (PEG) and allowed to grow in vertical position for 15 days before evaluation. For root length measurements, the ImageJ software was used. The assay was performed in triplicate (n = 3), where each set consisted of 15 seedlings each.
Stress assays with mature potted Arabidopsis plants
The Arabidopsis plants were grown in pots till the stage of bolting, before giving stress treatments. Salt stress of 150 mM NaCl was given for 3 weeks, every alternate day, followed by a recovery phase where excess salt was removed by passing excess water through the pots. Dehydration stress was also given gradually where a limited amount of water was given on a daily basis, insufficient for sustaining the potted plants, for introducing a gradual water deficit. For recovery, excess water was supplied to the pots for 1 week. All experiments were done thrice (n=3), where each set consisted of three pots each having eight plants. Response to exogenous ABA and oxidative stress was studied by transferring 15-day-old Arabidopsis plants, raised on MS basal, to stress media supplemented with 15 µM ABA and 2 µM paraquat for 14 days. Nearly 15 plants were taken per set, where n = 3. Membrane injury was studied by subjecting 15-day-old plants raised on a thin layer of soilrite mix in glass petri plates to different stress treatments for 72 h. The stress agents were administered as solutions by saturating the soilrite mix with their equal volumes, i.e., 35% PEG 6000, 250 mM NaCl, 25 µM ABA, and 5 µM paraquat. For membrane injury, each set consisted of three plants, where n = 3.
Water loss assay
The assay was performed in mature Arabidopsis plants, prior to bolting. The rosettes were detached, placed in trays covered on top with plastic wrap, and weighed every 30 min for 4 h. The assay was done in five sets (n = 5), each consisting of three mature rosettes.
Chlorophyll estimation
Chlorophyll estimation was done according to Porra (2002). One hundred-milligram leaf samples or three 15-day-old seedlings were harvested and added to 2 ml dimethyl sulphoxide (DMSO). The samples were incubated at 65 °C for 30 min. The absorbance was measured at 645 and 663 nm in 96-well micro-titer plates. The total chlorophyll was calculated using the equation: (Chl A + B) = (17.679 * A645) + (7.129 * A663). Chlorophyll values are represented in mg/g fresh weight or per 3 seedlings. The estimation was performed in a set of three (n = 3).
Proline estimation
Proline estimation was done according to the protocol given by Bates et al. (1973). A proline standard curve in the range of 0–100 µg was utilized in calculating the proline concentration in the samples. Readings from three plants per set were employed for estimating the proline values and 3 sets were used (n = 3).
MDA estimation
The malondialdehyde (MDA) estimation was done according to the protocol given by Heath and Packer (1968). The MDA concentration was calculated using the equation: C = A/155*L, where, C = MDA concentration, A = OD532 − OD600, and L = Pathlength. Readings from three plants per set were employed for estimating MDA and 3 sets were used (n = 3).
Fv/Fm measurements
The Fv/Fm values were measured using the Junior PAM instrument (PAM-210, H. Waltz). Before taking the measurements, the plants were darkly acclimatized for 2 h and the measurements were also performed in dark. Readings from ten plants per set were taken for calculating the Fv/Fm values, and three sets were employed (n = 3).
DAB staining
DAB (3,3′-diaminobenzidine) staining was done according to the protocol given by Kumar et al. (2014). The seedlings were incubated overnight in 10 ml DAB solution followed by de-staining. For staining, five plants were employed per set, where n = 3, and the best representation for each line is presented.
Ion leakage
The ion leakage was measured by quickly weighing 50-mg seedling tissues, washing them with Milli-Q water, and immersing them in 5-ml Milli-Q water for 60 min. One-milliliter sample (A) was removed after the incubation and the remainder volume was autoclaved (B). Both samples were measured for conductivity using a TDS meter (Eutech), and the ion leakage values were represented in percentage as (A/B *100). Readings from three plants per set were employed for calculating the ion leakage values, where n = 3.
Stomatal analysis
The stomata from the epidermal peels of the abaxial surface of mature leaves were isolated with the help of an adhesive tape and observed under a bright field microscope (Leica). The detached leaves were dipped in stomatal opening solution according to Eisele et al. (2016), for 2 h, followed by visualization of stomatal apertures, size, and density. Stomatal sensitivity to ABA and H2O2 was studied by immersing the leaves with fully open stomata in the stomatal opening solution supplemented with 10 µM ABA or 100 µM H2O2 for 90 min, before visualization. All experiments were done in a set of two (n = 2), where each set consisted of data from three leaves each divided into three parts, one each for control, ABA, and H2O2. The stomatal density was calculated from 5 images from random areas of each leaf. Stomatal size/aperture was calculated from 100 stomata from each set. The stomatal response was calculated from 10 random images for each treatment per set.
Protein cell localization
The coding sequence (CDS) of OsFBT4 was cloned in the pSITE-3CA vector where the YFP protein is fused to the N-terminal region of OsFBT4. Cell localization studies were performed in the onion peel cells using the PDS-1000 He system (Bio-Rad), according to the manufacturer’s instructions. The bombarded peels were visualized under a confocal microscope (Leica). For studying nuclear translocation in response to stress treatments, the bombarded peels were placed on MS basal supplemented with 1% H2O2, 20% PEG, and 10 µM ABA, for 4 h, before visualization. The experiment was done twice (n = 2) with similar observations.
Protein interaction studies in yeast and transactivation assay
For the OsFBT4-SKP1 interaction study, seven high-expressing rice SKP1 proteins (OSKs) were chosen, based on an earlier study by Kahloul et al. (2013), these were OSK1 (Os11g26910), OSK8 (Os11g48030), OSK11 (Os06g02350), OSK20 (Os09g36830), OSK23 (Os07g43270), OSK24 (Os12g40300), and OSK29 (Os08g28780). The CDS of OsFBT4 and OSKs were cloned in both pGBKT7 and pGADT7 vectors (Clontech) for studies in yeast. All yeast assays were confirmed at least twice and the best interpretation of the results has been presented.
Transactivation assay
The PGBKT7-OsFBT4 construct, where the gene of interest is cloned next to the GAL4 DNA binding domain was transformed into the AH109 strain of yeast and plated on SD-W medium supplemented with X-α-GAL, to check for the development of blue color.
Spot assay for yeast two-hybrid protein interaction studies
The reciprocal combinations of PGBKT7-OsFBT4 and PGADT7-OSKs; and PGBKT7-OSKs and PGADT7-OsFBT4 were co-transformed into the AH109 strain of yeast. These co-transformed cells were cultured on liquid SD-LW medium and were dropped on SD-HLW medium supplemented with 3 mM 3-AT (3-Amino-1,2,4-triazole). Growth of the cells was observed after 5–6 days.
ONPG assay and β-galactosidase colony-lift filter assay
These assays were performed following the protocols and instructions provided in the Yeast Protocols Handbook (Clontech).
Statistical analysis
The graphical representations have been done as averages with standard error. The statistical significance of the data has been analyzed using Student’s t-test in the Microsoft Excel program. The P-value less than 0.05 was considered to be significant.
Results
OsFBT4 is differentially regulated at the transcript level by different abiotic stresses
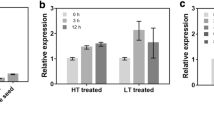
The RT-qPCR results showed that the gene was significantly up-regulated under drought stress and salt stress (Fig. 1). The transcript level increased slightly in response to oxidative stress caused by H2O2 treatment, although the response was not immediate. It indicates that the OsFBT4 transcript level is regulated by different abiotic stresses and the gene may play role in abiotic stress signalling. In response to exogenously supplied ABA, there was a sharp increase in the transcript level in the first hour of exposure followed by a sharp decline at 3 h, indicating the possibility of an early response to ABA and a possible feedback repression.
Expression profiling of OsFBT4 in rice, under different abiotic stress treatments, by RT-qPCR. Relative transcript level of OsFBT4 under different stress conditions have been graphically represented. Different controls (con) were used for different stress conditions; RO water was used for salt and H2O2 treatments, RGM (rice growth medium) was used for drought treatment and 0.1% EtOH was used for ABA treatment. The expression data was normalized with the geometric mean of UBQ5 and eEF-1α reference genes. The data is represented as mean ± SE. The asterisk marks (*) indicate significant difference (P-value < 0.05)
OsFBT4 interacts with several OSKs and might be part of larger SCF-E3 ligase complexes
F-box proteins play the role of a target-specific adaptor protein that functions to attach the target proteins to a larger multi-subunit complex called the SCF-type E3 ligase, for ubiquitination and ultimately degradation. SKP1 proteins (or OSKs in rice) are the component of the SCF complex that directly binds to the adapter F-box protein through the F-box motif. There are 31 OSKs in rice, showing expression in a wide range of tissues (Kahloul et al. 2013). We selected seven out of the nine OSKs that were showing widespread tissue expression, due to their higher chances of interaction with OsFBT4. The CDS of OsFBT4 and the chosen OSKs, viz. OSK1, 8, 11, 20, 23, 24, and 29 were cloned in both bait (with DNA binding domain of the GAL4 transcription factor) and prey (with activation domain of the GAL4 transcription factor) vectors, i.e., pGBKT7 and pGADT7, respectively. The OsFBT4 and OSK bait and prey constructs were co-transformed into the AH109 strain of yeast in reciprocal combinations and spot assays were performed on SD-HLW medium, supplemented with 3 mM 3-AT (Fig. 2A, B). The yeast-two-hybrid assays showed an interaction between OsFBT4 and OSK1, 20, 23, 24, and 29 when OsFBT4 was used in the bait construct and the OSKs in the prey construct, however, the interactions with OSK23 and 29 were weak. These interactions were re-confirmed when the reciprocal combination was used, i.e., OsFBT4 in the prey construct and the OSKs in the bait constructs. In the latter combination, the interaction between OsFBT4 and OSK8 was also seen which was not seen earlier. The results from the spot assays were also confirmed by β-GAL assay (Fig. 2A, B) and ONPG assay (Fig. 2C). The deeper blue color developed in the spots indicated stronger interaction in the β-GAL assay which was indicated by longer bars in the ONPG assay. Taken together, it was concluded that OsFBT4 interacts with OSK1, 8, 20, 23, 24, and 29, albeit with different strengths, and hence, possessed a functional F-box domain. Moreover, OsFBT4 did not show homo-dimerization in the yeast system, when the combination of OsFBT4 cloned in both bait and prey vectors was used.
Interaction study of OsFBT4 and OSKs by yeast two-hybrid. A Shows the spot assay and β-gal assay for all controls in all combinations, B Shows the spot assay and β-gal assay for interaction of OsFBT4 with the chosen OSKs in both reciprocal combinations. C Represents the ONPG assay performed to assess the strength of each interaction, where KT represents the pGBKT7 empty vector and AD represents pGADT7 empty vector. P53 + T7 served as positive control whereas LAM + T7 served as negative control. A 3-mM concentration of 3-AT was used for the spot assay. Intensity of the blue color developed in the β-gal assay is directly proportional to the strength of interaction. D Shows the transactivation assay, where the pGBKT7 empty vector served as negative control and the GAL4-AD fragment cloned in pGBKT7 served as positive control on SD-W + X-α-GAL medium
OsFBT4 may not possess transcriptional activation capability
Plant TLPs have been shown to possess a relatively weaker dsDNA binding property, but the transcriptional activation property has been shown for a few TLPs, and not all. To confirm the latter for OsFBT4, the pGBKT7-OsFBT4 bait construct, where the protein was fused with the DNA binding domain of the GAL4 transcription factor was transformed into the AH109 strain of yeast and plated on SD-W medium, supplemented with X-α-Gal. Activation of the Mel1 (α-galactosidase) reporter gene of the AH109 strain, due to the transcriptional activation property of the fused protein of interest, would have produced blue color due to the breakdown of X-α-Gal. No such activation was seen in the case of OsFBT4, although the positive control was able to activate the Mel1 gene (Fig. 2D). Hence, OsFBT4 did not show transactivation property in the yeast system.
OsFBT4 localizes to the PM and shows nuclear re-orientation in response to stress signals
In plants, TLPs have been shown to detach from the PM for nuclear translocation in response to stress signals (Reitz et al. 2012). The localization of OsFBT4 was studied in onion peel cells for which the CDS of the gene was cloned in the pSITE-3CA (YFP) vector, where YFP is fused N-terminal to OsFBT4. The construct was then used for the particle bombardment procedure. The empty vector when used as a control showed the distribution of YFP all throughout the cell, whereas the YFP-OsFBT4 localized to the PM, under control conditions (Fig. 3), which was expected. Nuclear translocation of YFP-OsFBT4 was observed when the bombarded onion peels were placed on a medium supplemented with 1% H2O2 and 20% PEG for 4 h, before visualization. No nuclear translocation of the protein was observed in response to 10 µM ABA. The results further strengthened the role of OsFBT4 in abiotic stress signalling.
Cell localization study of OsFBT4 by particle bombardment in onion peels. The first panel shows the sub-cellular localization of YFP protein as control, the second panel shows PM localization of OsFBT4, the third and fourth panels show the nuclear translocation of OsFBT4 from the PM after 4 h of 1% H2O2 and 20% PEG treatments. The fifth panel shows that 4 h of ABA treatment did not induce nuclear translocation of OsFBT4. The pSITE-3CA vector was used for the study where the YFP protein is fused to the N-terminus of the protein of interest. The nuclei in every image have been highlighted in red circles
Cloning of OsFBT4, generation of heterologous over-expression (OE) lines in Arabidopsis, and their phenotypic characterization
The ORF of the gene is 1287 bp in length and comprises of four exons that encode a protein of 428 amino acids (aa) comprising of an N-terminal F-box motif (56 aa) and a C-terminal Tubby domain (291 aa) (Fig. S1). The CDS of OsFBT4 was amplified from 12-day-old rice seedlings by RT-PCR and subsequently cloned into the binary vector, pBI121 (Clontech), which is a plant expression vector and the gene of interest is expressed constitutively under the CaMV 35S promoter. The construct was transformed into the Arabidopsis (Col-0) plants by the floral dip method. We obtained eight homozygous T3 OE lines which were confirmed for the presence of the transgene by PCR amplification of OsFBT4, using a combination of CaMV 35S promoter-specific forward primer and OsFBT4-specific reverse primer and also through PCR amplification of NPT-II (Neomycin Phosphotransferase-II) (Fig. S1). We also checked for the expression level of OsFBT4 in the OE lines (Fig. S1). Four OE lines, viz. OE 16, 18, 30, and 37 were chosen for further analysis as these lines displayed strongest phenotypes and showed the higher expression level of OsFBT4. Ectopic expression of OsFBT4 caused several developmental alterations in Arabidopsis. An increase in the length of lateral/secondary roots, while a significant reduction in their number was conspicuous at the seedling growth stage (Fig. 4A). Most of the OE lines showed deviation in leaf morphology, which included slightly broader leaves and increased waviness in their margins (Fig. 4B, C), causing an overall deviation from the normally oblong leaves to slightly obovate shape. There was a small reduction in the overall rosette size and plant height (Table S1), and there was a marked delay in the flowering time (bolt emergence) by 7–10 days across lines, under long-day condition (Fig. 4D, E). This delay was not attributed to the slow growth rate of the plants as there was a greater number of leaves at the time of bolting in the OE lines (Fig. 4F). From the above observations, it was clear that OsFBT4 plays pleiotropic roles and may regulate multiple developmental processes.
Morphological phenotypic analysis of OsFBT4-OE lines in Arabidopsis. (A) Shows longer but lesser number of lateral/secondary roots in the OE lines, at the 14-day-old stage. Morphological differences in leaf shape and rosette size are depicted in (B) and (C). (D) Shows late-flowering phenotype in the OE lines which is graphically represented in (E), and the associated increased leaf number phenotype at bolting time is represented in (F). All phenotypic observations were done several times. The flowering time data is represented as mean ± SE, where n = 3. The asterisk marks (*) indicate significant difference (P-value < 0.05)
OsFBT4 OE lines are sensitive to exogenous ABA and abiotic stress at the seed germination stage
We studied the germination and early seedling growth response of the OE lines to abiotic stresses. The results showed that the OE lines were significantly more sensitive to osmotic stress caused by mannitol, salt stress and exogenous ABA as the percentage of germination was significantly poor (Fig. 5A). A slightly lower germination rate was observed across the OE lines to oxidative stress caused by 3 µM paraquat, which was apparent only during the initial days. Growth characteristics of the seedlings were also observed, post germination, where the OE lines displayed a significantly lower percentage of cotyledon greening and expansion and also significantly reduced hypocotyl and radicle elongation on all abiotic stresses (Fig. 5B, C). Taken together, it could be concluded that ectopic expression of OsFBT4 caused enhanced sensitivity to ABA and abiotic stress, at the seed germination stage.
Response of the OsFBT4-OE lines to different abiotic stresses at the seed germination stage. Percentage seed germination from day 1 (D1) to day 5 (D5) for different stresses has been graphically presented in (A), where radicle emergence is scored as germinated. Cotyledon greening after 6 days of growth post germination has been graphically presented in (B). (C) depicts the growth characteristics of the OE lines in response to different stresses after 6 days of growth. Three micrometer ABA, 150 mM NaCl, 300 mM mannitol, and 3 µM paraquat were used for the assay. All results are represented as mean ± SE, where n = 3. The asterisk marks (*) indicate significant difference (P-value < 0.05)
Overexpression of OsFBT4 confers tolerance to 5-day-old Arabidopsis seedlings challenged with abiotic stresses
We checked for altered sensitivity in the OE lines towards different abiotic stresses at the 5-day-old seedling stage. After 14 days of stress treatment, the plants were evaluated for root growth and other physiological parameters. The primary and lateral roots were significantly longer in all the OE lines, in response to osmotic stress caused by mannitol and PEG, salt stress, oxidative stress, and exogenous ABA (Fig. 6A, D). An improved foliar growth was also seen in the case of salt stress, ABA, and oxidative stress, however, the osmotic stress caused by mannitol resulted in slightly reduced foliar growth (Fig. 6A). Higher root-to-shoot ratios are favored for tolerance to mild to medium drought conditions to conserve water (Verslues et al. 2006; Janiak et al. 2016). Foliar growth reduction was proportionate in the OE lines, in response to PEG, compared to the control condition. The biomass, as indicated by fresh weight per five seedlings, was significantly higher in the case of salt stress, exogenous ABA, and oxidative stress, but was not significant for mannitol and PEG (Fig. 6B). Finally, the percentage survival was calculated which was significantly higher in the OE lines for salt stress and oxidative stress (Fig. 6C). Survival rate was not affected by stress due to PEG, mannitol, and ABA, at the concentrations used in this study, but the growth characteristics were severely affected. Taken together, the results suggest tolerance in the OE lines towards osmotic stresses, salinity, exogenous ABA, and oxidative stress, at the seedling growth stage.
Response of the OsFBT4-OE lines to different abiotic stresses at the seedling growth stage. A Depicts the growth response of the OE lines to different abiotic stresses after 15 days on the stress media. Fresh weight or biomass per five seedlings has been graphically represented in (B). The percentage survival in response to stresses has been presented in (C). D Shows the total primary root length after 15 days of growth on stress media. 150 mM NaCl, 300 mM mannitol, 40% PEG, 10 µM ABA, and 1 µM paraquat were used for the assay. All results have been represented as mean ± SE, where n = 3. The asterisk marks (*) indicate significant difference (P-value < 0.05)
OsFBT4 confers tolerance to mature Arabidopsis plants subjected to salt and drought stress
At the mature stage, the plants were given gradual and sub-lethal exposure to salt and drought stresses, in order to allow them sufficient time to mount responses. The detrimental effects of salt were evident significantly earlier in the WT plants, relative to the OE lines (Fig. 7A), as also reflected in the higher chlorophyll and Fv/Fm values in the OE lines demonstrating lesser damage to the photosynthetic apparatus (Fig. 7C, D). At the end of the recovery phase, the survival rate was calculated which was significantly higher for the OE lines along with the onset of new growth which was far lesser in the WT plants (Fig. 7E). In response to dehydration, which was induced by a limited and insufficient daily supply of water, the OE lines showed a significantly lower and delayed wilting of the rosette, compared to WT (Fig. 7B). After 2 weeks the WT plants perished and showed poor revival during the recovery phase when excess water was supplied, whereas the OE lines showed much higher survival and recovery (Fig. 7F). In a separate experiment, the rate of loss of water was studied from the detached mature rosettes, just prior to bolting, to support the role of OsFBT4 in dehydration tolerance. As was expected, the OE lines showed a significantly lower rate of water loss due to transpiration, in comparison to WT, although there was no difference in the overall water content between the OE lines and the WT, as shown by the dry weights (Fig. 7G). The response of the OE lines to exogenous ABA and oxidative stress was also studied at 15-day-old plant stage (Fig. S2), which is closer to the mature stage. For this purpose, the Arabidopsis plants raised on MS medium were transferred to the stress media supplemented with 15 µM ABA or 2 µM paraquat. With the selected concentrations of the stress agents, there was no detrimental effect on survival, but the other physiological factors such as chlorophyll content, Fv/Fm value, and rosette size were affected. After 2 weeks on the stress media, the loss of chlorophyll content and drop in the Fv/Fm values were slightly lesser across the OE lines on both ABA and paraquat. Also, the reduction in the final rosette size was slightly lower in the OE lines in the case of ABA, but more or less proportionate in the case of oxidative stress. Taken together, it could be concluded that overexpression of OsFBT4 conferred tolerance to drought and salt stresses at the mature stage of growth. It also conferred slight tolerance to exogenous ABA and oxidative stress at this stage.
Response of the OsFBT4-OE lines to different abiotic stresses at the mature plant stage. A and B Depicts the response of the OE lines of OsFBT4 to salt stress and drought stress, respectively, at different stages of treatment, B represents the Fv/Fm values; and (D) shows the total chlorophyll content after exposure to salt stress. The percentage survival after the recovery phase has been presented in (E) and (F) for salt and drought stresses, respectively. The rate of water loss in the OE lines w.r.t. WT has been presented in (G). All results have been represented as mean ± SE, where n = 3, for water loss assay n = 5. The asterisk marks (*) indicate significant difference (P-value < 0.05)
Plants overexpressing OsFBT4 show lesser damage to membranes when challenged with stress
The stress of any form tends to jeopardize the normal functioning of cells and leads to increased production of reactive oxygen species (ROS) due to disruption of cell homeostasis. These ROS entities predominantly attack membrane structures and associated components, leading to cell damage. To support the positive role of OsFBT4 in stress signaling, we checked for membrane injury under stress conditions by exposing 15-day-old plants, raised on a thin layer of pot mix, to stress agents for 72 h (Fig. S3). ROS (H2O2) accumulation was studied by the DAB staining method which showed slightly lighter staining, caused by lower H2O2 accumulation, under most abiotic stresses, except oxidative stress, where similar staining was seen across the OE lines and WT. The chlorophyll level/3 seedlings and the Fv/Fm values were measured, where the loss of chlorophyll and decline in the Fv/Fm values were lower in the OE lines, in response to all abiotic stresses. The Fv/Fm ratio is the maximum quantum efficiency of photosystem II (PSII), indicative of its performance in plants. Ion leakage, as a function of membrane injury, was also found to be lower in the OE lines in response to all abiotic stresses. The MDA level was observed to be significantly lower in response to salt, PEG, and oxidative stress, while it was only slightly lower in the case of ABA. The proline level was higher in the OE lines in the case of salt, drought, and oxidative stresses, but was similar in the case of exogenous ABA. The results suggest that the OsFBT4-OE lines incur lesser damage to its membranes and the photosynthetic apparatus, in response to ABA and abiotic stress.
Overexpression of OsFBT4 causes altered stomatal response to exogenous ABA and H2O2
We checked for stomatal size, density, and response to exogenous ABA and H2O2 as the OE lines exhibited a reduced water loss rate. The stomata in the OE lines were significantly larger in size and lower in density (Figs. 8A, F, G). We found an altered stomatal response in the OE lines to both ABA and H2O2. A higher percentage of stomata was found to be completely closed across the OE lines in response to ABA and H2O2, although this response was stronger in the case of H2O2 treatment (Fig. 8A–D). The stronger stomatal response was also reflected in their smaller apertures across the OE lines, in response to both agents (Fig. 8E).
Stomatal response from the mature OsFBT4-OE plants to ABA and H2O2. (A) Pictorially depicts the stomatal response of the OE lines and WT to stomatal opening solution (control), ABA (10 µM), and H2O2 (100 µM). Also note the larger stomatal size and lower stomatal density in the OE lines. (B, C, and D) graphically represents percentages of stomata in open/partially open/closed states when leaves were treated with ABA and H2O2 for 90 min from the fully open state (control). (E) Represents the reduction in stomatal aperture in response to ABA and H2O2 after 90-min exposure. (F) Represents the stomatal density; and (G) shows the stomatal size comparisons between the OE lines and WT. The results are represented as mean ± SE, where n = 2. The asterisk marks (*) indicate significant difference (P-value < 0.05)
Overexpression of OsFBT4 causes differential regulation of several stress-responsive genes, at different stages of growth
Overexpression of OsFBT4 confers tolerance to most abiotic stresses, including ABA, at advanced stages of growth. To find out the plausible mechanism of conferring the general tolerance to abiotic stress, we checked the expression level of several known stress-responsive genes in the 15-day-old MS medium-grown plants under unstressed condition (Fig. 9). Most of the genes quantified are known to be involved in several layers of stress signalling, both ABA-dependent and independent, and were found to be significantly upregulated in the OsFBT4-OE lines, such as SOD1, CAT1, DREB2A, ERD6, RD20, RD22, RD26, RD29A, ATAF1, RAB18, and MYB2. Since the OsFBT4-OE lines showed increased sensitivity to abiotic stresses at the germination and seedling establishment stages, we checked the transcript level of these marker genes at the 3-day-old seedling stage also. Most of these marker genes were slightly downregulated at this stage, except RAB18, which did not show significant deviation from the WT plants. The differential regulation of these diverse stress marker genes in the OE lines, in an unstressed condition, at different stages of growth shows that OsFBT4 may regulate both ABA-dependent and independent stress signalling pathways.
Transcript quantification of stress-marker genes in the OsFBT4-OE lines in Arabidopsis at 3-day and 15-day-old plant stages, by RT-qPCR. The expression data was normalized with Actin-II. The results are represented as mean ± SE. The asterisk marks (*) indicate significant difference (P-value < 0.05)
Discussion
In the present study, we showed that OsFBT4 plays pleiotropic roles in plant development, most notably in leaf and root architecture and vegetative to floral transition. The OsFBT4-OE lines also exhibit slight dwarfism, which is contrary to the CaTLP1-OE lines, the ortholog of OsFBT4, where an enhanced growth rate was seen (Wardhan et al. 2012, 2016). TLPs may be involved in plant development as they are shown to be regulated by major plant hormones (Lai et al. 2004). The OsFBT4-OE lines were moderately to mildly insensitive to ABA at the seedling and mature stages, respectively. ABA insensitivity has been associated with dwarfism (Gonzalez-Guzman et al. 2012; Zhao et al. 2018), which may partly explain the dwarfism observed in the OsFBT4-OE lines. Moreover, the late flowering phenotype observed in the OsFBT4-OE lines may also be attributed to ABA insensitivity. ABA promotes flowering by upregulating the florigen gene (FT) through the Gigantea and Constans signalling pathway, as several ABA-deficient and insensitive mutants have been shown to be late flowering (Riboni et al. 2013; Hwang et al. 2019). We showed that OsFBT4 interacts with OSK1, 8, 20, 23, 24, and 29 and hence possessed a functional F-box domain, and may participate in protein turnover. OsFBT4 failed to show transcriptional activation property, in yeast, localized to the PM and exhibited nuclear translocation in response to oxidative and osmotic stresses. In plants, the TLPs may be playing dual roles; as weak transcription factors and as F-box proteins. Several previous studies support the latter, as TLPs from several plants have been shown to bind SKP1 proteins, which may be mediated jointly through both F-box and Tubby domains (Lai et al. 2004; Lai and Shaw 2012; Bao et al. 2014; Li et al. 2021). It is essential for the F-box proteins to bind to the SKP1 subunit of the SCF-E3-ligase complex for participating in the targeted protein turnover. Several SKP1 proteins interact with several sub-classes of F-box proteins, while some of them are quite specific in their interactions (Risseeuw et al. 2003). Kuroda et al. (2012) revealed that AtTLPs interact with highly specific SKP1 proteins, ASK1 and ASK2, in Arabidopsis, compared to other sub-classes of the F-box proteins. The rice SKP1 proteins, OSK1 and OSK20, are orthologous to ASK1 and ASK2, and similarly showed interactions with several sub-classes of rice F-box proteins (Kahloul et al. 2013). Both interacted with OsFBT4 in the present study. Several previous studies have shown nuclear/partially cytosolic translocation of the plant TLPs from their usual PM-bound cell localization, in response to abiotic stress (Reitz et al. 2012; Wardhan et al. 2012; Bao et al. 2014; Li et al. 2021). However, ABA and other phytohormones failed to induce nuclear translocation of AtTLP3 (Reitz et al. 2012), similar to what was observed for OsFBT4 in the present study. The PM-binding function of the TLPs is mediated exclusively by the Tubby domain and may even be essential for the protein function, as seen for MdTLP7 (Bao et al. 2014; Xu et al. 2019). The nuclear translocation of plant TLPs may be facilitated by interaction with other nuclear-bound proteins, such as the Nuclear Factor Y subunit C3 (NF-YC3), as seen in the case of AtTLP2 (Wang et al. 2019). CaTLP1, AtTLP9, and SlTLFP8 were shown to be unable to activate transcription from the GAL4 promoter when fused to the GAL4 DNA-binding domain in the yeast two-hybrid system (Lai et al. 2004; Wardhan et al. 2012; Li et al. 2020, 2021). On the contrary, in rice, OsTLP2 was shown to transcriptionally regulate OsWRKY13 by binding to its promoter and AtTLP2 activated the expression of UGE1 (UDP-glucose epimerase) in Arabidopsis mesophyll protoplasts (Cai et al. 2008; Wang et al. 2019). Hence, plant TLPs may function as weaker transcription factors and may require specific post-translational modifications or additional interacting mediator proteins for regulating transcription.
In the present study, we found OsFBT4 to be upregulated by abiotic stress, including ABA. Several TLPs have been shown to be under direct transcriptional regulation by ABA and abiotic stress (Lai et al. 2004; Bao et al. 2014; Xu et al. 2016; Dong et al. 2019; Bano et al. 2021; Li et al. 2021). The OE lines of OsFBT4, in Arabidopsis, were significantly sensitive to exogenous ABA and abiotic stress at the seed germination and seedling establishment stages, but exhibited tolerance to ABA, salt, osmotic, and oxidative stresses at the later stages of growth. Several plant TLPs have been demonstrated to play roles in abiotic stress signalling that may involve both ABA-regulated and independent pathways. Overexpression of CaTLP1, an ortholog of OsFBT4, has also been shown to confer tolerance to ABA, salt, osmotic, and oxidative stresses at multiple stages of plant growth, including during seed germination, in Arabidopsis and tobacco (Wardhan et al. 2012, 2016). Overexpression of SlTLFP8, in tomato, conferred tolerance to drought stress, while its loss-of-function CRISPER lines were more sensitive (Li et al. 2020). Overexpression of CsTLP8 inhibited seed germination in Arabidopsis and cell growth in yeast, in response to ABA, salt, and osmotic stresses (Li et al. 2021). MdTLP7, when expressed in Arabidopsis and E.coli, conferred tolerance to salt, dehydration, heat, and cold stresses (Du et al. 2014; Xu et al. 2019).
Loss of function mutants, attlp3 and attlp9, were observed to be insensitive to exogenous ABA and osmotic stress during seed germination, while overexpression of AtTLP9 caused hypersensitivity to ABA during the same stage. The attlp3-attlp9 double mutant is further insensitive to ABA, suggesting their involvement in redundant as well as overlapping ABA-signalling pathways (Lai et al. 2004; Bao et al. 2014). Moreover, AtTLP9 interacts with a RING-H2 zinc-finger protein, XERICO, in Arabidopsis, the upregulation of which causes a dramatic increase in the endogenous ABA levels, caused by transcriptional upregulation of ABA-biosynthesis gene, NCED3, resulting in increased sensitivity to exogenous ABA and abiotic stress during seed germination, while conferring drought tolerance at the mature stage due to greater percentage of ABA-led stomatal closure (Ko et al. 2006). We made similar observations in the case of OsFBT4, where overexpression of the gene caused increased sensitivity to ABA and stress during seed germination and seedling establishment, but conferred tolerance to both at the later stages. AtTLP7 and AtTLP11 interact with an ABA and stress-inducible protein, NHL6 (NDR1/HIN1-like 6), that regulates seed germination and early seedling growth in response to ABA-mediated stress signalling (Bao et al. 2016; Song et al. 2019). Hence, it is possible that OsFBT4 may upregulate endogenous ABA level during seed germination, causing delayed germination and seedling establishment, or may interact with other proteins to regulate stress responses.
Disruption or overexpression of several components of ABA signalling has been shown to produce altered sensitivity to ABA and stress, specifically only during seed germination, similar to what was observed in the OsFBT4-OE lines. The ABA-hypersensitive germination mutants, ahg1, ahg2, and ahg3, are defective in the protein phosphatase 2C (PP2C) encoding genes and serve as negative regulators of ABA signalling. These mutants exhibit pronounced ABA and stress hypersensitivity during the seed germination stage by transiently enhancing the endogenous ABA levels, but behave normally at advanced stages of growth (Nishimura et al. 2005, 2007). ABI5 is a positive regulator of ABA signalling in mature seeds and its loss of function mutant is insensitive to ABA and stress only during the seed germination stage, but not during other stages (Gonzalez-Guzman et al. 2012; Shu et al. 2016; Zhao et al. 2020). Overexpression of a positive regulator of ABA-signalling, SnRK2.6, caused hypersensitivity to ABA and stress during seed germination and early seedling growth but conferred improved growth at advanced stages (Zheng et al. 2010).
ABA and ROS (H2O2) regulate opening and closing of stomata (Li et al. 2017; He et al. 2018). Reduced stomatal density and increased stomatal cell size contribute to reduced transpiration and increased water use efficiency leading to drought tolerance (Yoo et al. 2010; Wang et al. 2016; Guo et al. 2019). Several TLPs, such as CaTLP1, AtTLP2, and CsTLP8, which conferred drought tolerance also caused increased stomatal sensitivity to ABA (Wardhan et al. 2016; Li et al. 2021). Drought-tolerant SlTLFP8-OE lines exhibited lower stomatal density, but larger stomatal size (Li et al. 2020). Similar results were observed for the OsFBT4-OE lines, in the present study, where they displayed larger stomatal size, lower stomatal density, and increased responsiveness to both ABA and H2O2, leading to a lower water loss rate and improved drought tolerance.
Several stress marker genes, such as SOD1, CAT1, DREB2A, ERD6, RD20, RD22, RD26, RD29A, ATAF1, RAB18, and MYB2 were found to be upregulated in the OsFBT4-OE lines in un-stressed condition, at a mature stage of development. Most of these genes were, however, down-regulated or unaltered in expression at the 3-day-old seedling stage. The differential expression pattern of these stress-responsive genes at different developmental stages may partly explain the different stage-specific responses of the OsFBT4-OE lines. SOD1 and CAT1 are known to scavenge ROS, generated in response to stress thereby protecting the cells from oxidative damages (Du et al. 2008; Lin et al. 2019). CsTLP8 OE lines showed reduced activities of ROS scavenging enzymes which led to more ROS accumulation in stress condition causing inhibition of seed germination and reduced early seedling growth (Li et al. 2021). RD20, RD22, RD26, and RD29A are ABA-dependent stress-regulated genes that play several roles in ABA signalling and ABA-mediated stress responses (Fujita et al. 2004; Aubert et al. 2010; Msanne et al. 2011; Harshavardhan et al. 2014; Kamranfar et al. 2021). ATAF1 induces ABA biosynthesis by binding to the promoter of ABA-biosynthesis gene, NCED3, and regulates stress signalling (Wu et al. 2009; Jensen et al. 2013; Wang et al. 2018). Expression of DREB2A, RAB18, MYB2, and ERD6 is regulated by ABA and stress and are involved in dehydration and salt stress responses (Lång and Palva 1992; Yoo et al. 2005; Kim et al. 2011; Slawinski et al. 2021). Several of these ABA/stress-responsive genes are known to be downregulated in the ABA-insensitive mutants (Yoshida et al. 2010; Gonzalez-Guzman et al. 2012; Zhao et al. 2018). Moreover, several genes, such as MYB2, ATAF1, RD20, and RD22 may also be upregulated in an ABA-independent manner (Fujita et al. 2004; Yoshida et al. 2010; Zhao et al. 2018). In the present study, these ABA/stress-responsive genes were downregulated/unaltered at the ABA-hypersensitive germination/seedling establishment stage, while they were upregulated at the less ABA-sensitive advanced stages of growth, suggesting the involvement of both ABA-dependent and independent pathways in stress responses.
Conclusion
In this study, we characterized a rice Tubby-like protein (TLP) encoding gene, OsFBT4, by analyzing the OE lines in Arabidopsis. OsFBT4 exhibits properties common to plant TLPs and plays extensive roles in abiotic stress signalling. Over-expression of the gene conferred tolerance to ABA and abiotic stress at most stages of plant growth, except the seed germination stage, where hypersensitivity to abiotic stress, including ABA, was seen. Higher endogenous ABA level at the seed germination stage is a possibility which could result in the increased sensitivity to stress at this stage, although we have not checked the ABA levels. At more advance stages of growth, the OsFBT4 overexpression seemed to activate more intricate and wide stress response pathways, both ABA-dependent and independent, conferring an overall tolerance to abiotic stress. The future studies should be aimed at deciphering the molecular mechanisms of functioning of the plant TLPs, as little is known in this direction and they differ substantially from the other eukaryotic TLPs in their domain/structural organization.
Data Availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Aubert Y, Vile D, Pervent M et al (2010) RD20, a stress-inducible caleosin, participates in stomatal control, transpiration and drought tolerance in Arabidopsis thaliana. Plant Cell Physiol 51:1975–1987. https://doi.org/10.1093/pcp/pcq155
Bano N, Fakhrah S, Mohanty CS, Bag SK (2021) Genome-wide identification and evolutionary analysis of gossypium Tubby-like protein (TLP) gene family and expression analyses during salt and drought stress. Front Plant Sci 12:1–24. https://doi.org/10.3389/fpls.2021.667929
Bao Y, Song WM, Jin YL et al (2014) Characterization of Arabidopsis Tubby-like proteins and redundant function of AtTLP3 and AtTLP9 in plant response to ABA and osmotic stress. Plant Mol Biol 86:471–483. https://doi.org/10.1007/s11103-014-0241-6
Bao Y, Song WM, Pan J et al (2016) Overexpression of the NDR1/HIN1-like gene NHL6 modifies seed germination in response to abscisic acid and abiotic stresses in Arabidopsis. PLoS ONE 11:1–16. https://doi.org/10.1371/journal.pone.0148572
Bates LS, Waldren RP, Teare ID (1973) Rapid determination of free proline for water-stress studies. Plant Soil 39:205–207. https://doi.org/10.1007/BF00018060
Boggon TJ, Shan WS, Santagata S et al (1999) Implication of tubby proteins as transcription factors by structure- based functional analysis. Science 286:2119–2125. https://doi.org/10.1126/science.286.5447.2119
Cai M, Qiu D, Yuan T et al (2008) Identification of novel pathogen-responsive cis-elements and their binding proteins in the promoter of OsWRKY13, a gene regulating rice disease resistance. Plant Cell Environ 31:86–96. https://doi.org/10.1111/j.1365-3040.2007.01739.x
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743. https://doi.org/10.1046/j.1365-313X.1998.00343.x
Dong MY, Fan XW, Pang XY, Li YZ (2019) Decrypting tubby-like protein gene family of multiple functions in starch root crop cassava. AoB Plants 11:1–13. https://doi.org/10.1093/aobpla/plz075
Du F, Xu JN, Zhan CY et al (2014) An obesity-like gene MdTLP7 from apple (Malus × domestica) enhances abiotic stress tolerance. Biochem Biophys Res Commun 445:394–397. https://doi.org/10.1016/j.bbrc.2014.02.005
Du YY, Wang PC, Chen J, Song CP (2008) Comprehensive functional analysis of the catalase gene family in Arabidopsis thaliana. J Integr Plant Biol 50:1318–1326. https://doi.org/10.1111/j.1744-7909.2008.00741.x
Eisele JF, Fäßler F, Bürgel PF, Chaban C (2016) A rapid and simple method for microscopy-based stomata analyses. PLoS ONE 11:1–13. https://doi.org/10.1371/journal.pone.0164576
Fujita M, Fujita Y, Maruyama K et al (2004) A dehydration-induced NAC protein, RD26, is involved in a novel ABA-dependent stress-signaling pathway. Plant J 39:863–876. https://doi.org/10.1111/j.1365-313X.2004.02171.x
Gonzalez-Guzman M, Pizzio GA, Antoni R et al (2012) Arabidopsis PYR/PYL/RCAR receptors play a major role in quantitative regulation of stomatal aperture and transcriptional response to abscisic acid. Plant Cell 24:2483–2496. https://doi.org/10.1105/tpc.112.098574
Guo XY, Wang Y, Zhao PX et al (2019) AtEDT1/HDG11 regulates stomatal density and water-use efficiency via ERECTA and E2Fa. New Phytol 223:1478–1488. https://doi.org/10.1111/nph.15861
Harshavardhan VT, Van Son L, Seiler C et al (2014) AtRD22 and AtUSPL1, members of the plant-specific BURP domain family involved in Arabidopsis thaliana drought tolerance. PLoS ONE 9:1–14. https://doi.org/10.1371/journal.pone.0110065
He F, Wang HL, Li HG et al (2018) PeCHYR1, a ubiquitin E3 ligase from Populus euphratica, enhances drought tolerance via ABA-induced stomatal closure by ROS production in Populus. Plant Biotechnol J 16:1514–1528. https://doi.org/10.1111/pbi.12893
Heath RL, Packer L (1968) Photoperoxidation in isolated chloroplasts. I. Kinetics and stoichiometry of fatty acid peroxidation. Arch Biochem Biophys 125:189–198. https://doi.org/10.1016/0003-9861(68)90654-1
Heikenwälder MF, Koritschoner NP, Pajer P et al (2001) Molecular cloning, expression and regulation of the avian tubby-like protein 1 (tulp1) gene. Gene 273:131–139. https://doi.org/10.1016/S0378-1119(01)00578-9
Hwang K, Susila H, Nasim Z et al (2019) Arabidopsis ABF3 and ABF4 transcription factors act with the NF-YC complex to regulate SOC1 expression and mediate drought-accelerated flowering. Mol Plant 12:489–505. https://doi.org/10.1016/j.molp.2019.01.002
Jain M, Nijhawan A, Arora R et al (2007) F-Box proteins in rice. Genome-wide analysis, classification, temporal and spatial gene expression during panicle and seed development, and regulation by light and abiotic stress. Plant Physiol 143:1467–1483. https://doi.org/10.1104/pp.106.091900
Janiak A, Kwaśniewski M, Szarejko I (2016) Gene expression regulation in roots under drought. J Exp Bot 67:1003–1014. https://doi.org/10.1093/jxb/erv512
Jensen MK, Lindemose S, de Masi F et al (2013) ATAF1 transcription factor directly regulates abscisic acid biosynthetic gene NCED3 in Arabidopsis thaliana. FEBS Open Bio 3:321–327. https://doi.org/10.1016/j.fob.2013.07.006
Kahloul S, Hajsalah El Beji I, Boulaflous A et al (2013) Structural, expression and interaction analysis of rice SKP1-like genes. DNA Res 20:67–78. https://doi.org/10.1093/dnares/dss034
Kamranfar I, Balazadeh S, Mueller-Roeber B (2021) NAC transcription factor RD26 is a regulator of root hair morphogenic plasticity. bioRxiv. https://doi.org/10.1101/2021.04.21.440803
Kim JS, Mizoi J, Yoshida T et al (2011) An ABRE promoter sequence is involved in osmotic stress-responsive expression of the DREB2A gene, which encodes a transcription factor regulating drought-inducible genes in Arabidopsis. Plant Cell Physiol 52:2136–2146. https://doi.org/10.1093/pcp/pcr143
Ko JH, Yang SH, Han KH (2006) Upregulation of an Arabidopsis RING-H2 gene, XERICO, confers drought tolerance through increased abscisic acid biosynthesis. Plant J 47:343–355. https://doi.org/10.1111/j.1365-313X.2006.02782.x
Kou Y, Qiu D, Wang L et al (2009) Molecular analyses of the rice tubby-like protein gene family and their response to bacterial infection. Plant Cell Rep 28:113–121. https://doi.org/10.1007/s00299-008-0620-z
Kumar D, Yusuf MA, Singh P, Sardar M, Sarin NB (2014) Histochemical detection of superoxide and H2O2 accumulation in Brassica juncea. Bio-protocol 4:1–4. https://doi.org/10.21769/BioProtoc.1108
Kuroda H, Yanagawa Y, Takahashi N et al (2012) A comprehensive analysis of interaction and localization of Arabidopsis SKP1-LIKE (ASK) and F-Box (FBX) proteins. PLoS One 7. https://doi.org/10.1371/journal.pone.0050009
Lai CP, Lee CL, Chen PH et al (2004) Molecular analyses of the Arabidopsis Tubby-like protein gene family. Plant Physiol 134:1586–1597. https://doi.org/10.1104/pp.103.037820
Lai CP, Shaw JF (2012) Interaction analyses of Arabidopsis tubby-like proteins with ASK proteins. Bot Stud 53:447–458
Lång V, Palva ET (1992) The expression of a RAB-related gene, RAB18, is induced by abscisic acid during the cold acclimation process of Arabidopsis thaliana (L.) Heynh. Plant Mol Biol 20:951–962. https://doi.org/10.1007/BF00027165
Li J, Li Y, Yin Z et al (2017) OsASR5 enhances drought tolerance through a stomatal closure pathway associated with ABA and H2O2 signalling in rice. Plant Biotechnol J 15:183–196. https://doi.org/10.1111/pbi.12601
Li S, Wang Z, Wang F et al (2021) A tubby-like protein CsTLP8 acts in the ABA signaling pathway and negatively regulates osmotic stresses tolerance during seed germination. BMC Plant Biol 21:1–14. https://doi.org/10.1186/s12870-021-03126-y
Li S, Zhang J, Liu L et al (2020) SlTLFP8 reduces water loss to improve water-use efficiency by modulating cell size and stomatal density via endoreduplication. Plant Cell Environ 43:2666–2679. https://doi.org/10.1111/pce.13867
Lin KH, Sei SC, Su YH, Chiang CM (2019) Overexpression of the Arabidopsis and winter squash superoxide dismutase genes enhances chilling tolerance via ABA-sensitive transcriptional regulation in transgenic Arabidopsis. Plant Signal Behav 14:1–12. https://doi.org/10.1080/15592324.2019.1685728
Liu Q (2008) Identification of rice Tubby-like genes and their evolution. FEBS J 275:163–171. https://doi.org/10.1111/j.1742-4658.2007.06186.x
Msanne J, Lin J, Stone JM, Awada T (2011) Characterization of abiotic stress-responsive Arabidopsis thaliana RD29A and RD29B genes and evaluation of transgenes. Planta 234:97–107. https://doi.org/10.1007/s00425-011-1387-y
Mukhopadhyay S, Jackson PK (2011) The tubby family proteins. Genome Biol 12:1–9. https://doi.org/10.1186/gb-2011-12-6-225
Nishimura N, Kitahata N, Seki M et al (2005) Analysis of ABA Hypersensitive Germination2 revealed the pivotal functions of PARN in stress response in Arabidopsis. Plant J 44:972–984. https://doi.org/10.1111/j.1365-313X.2005.02589.x
Nishimura N, Yoshida T, Kitahata N et al (2007) ABA-Hypersensitive Germination1 encodes a protein phosphatase 2C, an essential component of abscisic acid signaling in Arabidopsis seed. Plant J 50:935–949. https://doi.org/10.1111/j.1365-313X.2007.03107.x
Nishina PM, North MA, Ikeda A et al (1998) Molecular characterization of a novel tubby gene family member, TULP3, in mouse and humans. Genomics 54:215–220. https://doi.org/10.1006/geno.1998.5567
Porra RJ (2002) The chequered history of the development and use of simultaneous equations for the accurate determination of chlorophylls a and b. Photosynth Res 73:149–156
Reitz MU, Bissue JK, Zocher K et al (2012) The subcellular localization of tubby-like proteins and participation in stress signaling and root colonization by the mutualist Piriformospora indica. Plant Physiol 160:349–364. https://doi.org/10.1104/pp.112.201319
Riboni M, Galbiati M, Tonelli C, Conti L (2013) GIGANTEA enables drought escape response via abscisic acid-dependent activation of the florigens and suppressor of overexpression of CONSTANS11[c][w]. Plant Physiol 162:1706–1719. https://doi.org/10.1104/pp.113.217729
Risseeuw EP, Daskalchuk TE, Banks TW, Liu E, Cotelesage J, Hellmann H, Estelle M, Somers DE, Crosby WL (2003) Protein interaction analysis of SCF ubiquitin E3 ligase subunits from Arabidopsis. Plant J 34:753–767
Santagata S, Boggon TJ, Baird CL et al (2001) G-protein signaling through tubby proteins. Science 292:2041–2050. https://doi.org/10.1126/science.1061233
Shu K, Liu XD, Xie Q, He ZH (2016) Two faces of one seed: hormonal regulation of dormancy and germination. Mol Plant 9:34–45. https://doi.org/10.1016/j.molp.2015.08.010
Slawinski L, Israel A, Artault C et al (2021) Responsiveness of early response to dehydration six-like transporter genes to water deficit in Arabidopsis thaliana leaves. Front Plant Sci 12:1–21. https://doi.org/10.3389/fpls.2021.708876
Song W-m, Cheng Z-h, Guo X-t et al (2019) Overexpression of NHL6 affects seed production in transgenic Arabidopsis plants. Plant Growth Regul 88:41–47. https://doi.org/10.1007/s10725-019-00486-2
Verslues PE, Agarwal M, Katiyar-Agarwal S et al (2006) Methods and concepts in quantifying resistance to drought, salt and freezing, abiotic stresses that affect plant water status. Plant J 45:523–539. https://doi.org/10.1111/j.1365-313X.2005.02593.x
Vierstra RD (2009) The ubiquitin-26S proteasome system at the nexus of plant biology. Nat Rev Mol Cell Biol 10:385–397. https://doi.org/10.1038/nrm2688
Wang C, Liu S, Dong Y et al (2016) PdEPF1 regulates water-use efficiency and drought tolerance by modulating stomatal density in poplar. Plant Biotechnol J 14:849–860. https://doi.org/10.1111/pbi.12434
Wang J, Zhang L, Cao Y et al (2018) CsATAF1 positively regulates drought stress tolerance by an ABA-dependent pathway and by promoting ROS scavenging in cucumber. Plant Cell Physiol 59:930–945. https://doi.org/10.1093/pcp/pcy030
Wang M, Xu Z, Ahmed RI et al (2019) Tubby-like Protein 2 regulates homogalacturonan biosynthesis in Arabidopsis seed coat mucilage. Plant Mol Biol 99:421–436. https://doi.org/10.1007/s11103-019-00827-9
Wardhan V, Jahan K, Gupta S et al (2012) Overexpression of CaTLP1, a putative transcription factor in chickpea (Cicer arietinum L.), promotes stress tolerance. Plant Mol Biol 79:479–493. https://doi.org/10.1007/s11103-012-9925-y
Wardhan V, Pandey A, Chakraborty S, Chakraborty N (2016) Chickpea transcription factor CaTLP1 interacts with protein kinases, modulates ROS accumulation and promotes ABA-mediated stomatal closure. Sci Rep 6:1–16. https://doi.org/10.1038/srep38121
Wu Y, Deng Z, Lai J et al (2009) Dual function of Arabidopsis ATAF1 in abiotic and biotic stress responses. Cell Res 19:1279–1290. https://doi.org/10.1038/cr.2009.108
Xu J, Xing S, Sun Q et al (2019) The expression of a tubby-like protein from Malus domestica (MdTLP7) enhances abiotic stress tolerance in Arabidopsis. BMC Plant Biol 19:1–8. https://doi.org/10.1186/s12870-019-1662-9
Xu JN, Xing SS, Zhang ZR et al (2016) Genome-wide identification and expression analysis of the tubby-like protein family in the Malus domestica genome. Front Plant Sci 7:1–12. https://doi.org/10.3389/fpls.2016.01693
Yang Z, Zhou Y, Wang X et al (2008) Genomewide comparative phylogenetic and molecular evolutionary analysis of tubby-like protein family in Arabidopsis, rice, and poplar. Genomics 92:246–253. https://doi.org/10.1016/j.ygeno.2008.06.001
Yoo CY, Pence HE, Jin JB et al (2010) The Arabidopsis GTL1 transcription factor regulates water use efficiency and drought tolerance by modulating stomatal density via transrepression of SDD1. Plant Cell 22:4128–4141. https://doi.org/10.1105/tpc.110.078691
Yoo JH, Park CY, Kim JC et al (2005) Direct interaction of a divergent CaM isoform and the transcription factor, MYB2, enhances salt tolerance in Arabidopsis. J Biol Chem 280:3697–3706. https://doi.org/10.1074/jbc.M408237200
Yoshida S, Forno DA, Cock JH, Gomez KA (1976) Laboratory manual for physiological studies of rice, 3rd edn. International Rice Research Institute, Manila
Yoshida T, Fujita Y, Sayama H et al (2010) AREB1, AREB2, and ABF3 are master transcription factors that cooperatively regulate ABRE-dependent ABA signaling involved in drought stress tolerance and require ABA for full activation. Plant J 61:672–685. https://doi.org/10.1111/j.1365-313X.2009.04092.x
Zhang Y, He X, Su D et al (2020) Comprehensive profiling of tubby-like protein expression uncovers ripening-related TLP genes in tomato (Solanum lycopersicum). Int J Mol Sci 21:1–14. https://doi.org/10.3390/ijms21031000
Zhao H, Nie K, Zhou H et al (2020) ABI5 modulates seed germination via feedback regulation of the expression of the PYR/PYL/RCAR ABA receptor genes. New Phytol 228:596–608. https://doi.org/10.1111/nph.16713
Zhao Y, Zhang Z, Gao J et al (2018) Arabidopsis duodecuple mutant of PYL ABA receptors reveals PYL repression of ABA-independent SnRK2 activity. Cell Rep 23:3340-3351.e5. https://doi.org/10.1016/j.celrep.2018.05.044
Zheng Z, Xu X, Crosley RA et al (2010) The protein kinase SnRK2.6 mediates the regulation of sucrose metabolism and plant growth in Arabidopsis. Plant Physiol 153:99–113. https://doi.org/10.1104/pp.109.150789
Funding
This research was funded by the Department of Science and Technology (DST), Government of India and J.C. Bose National Fellowship to JPK (SB/SR/JCB-13/2013) by the Science and Engineering Research Board (SERB), Government of India. Authors also acknowledge infrastructural support by the Department of Science and Technology (FIST and PURSE programs), Government of India, and the University Grants Commission (UGC-SAP), New Delhi. NJ thanks the Council of Scientific and Industrial Research (CSIR), New Delhi, for providing financial assistance in the form of RA Fellowship.
Author information
Authors and Affiliations
Contributions
NJ and JPK designed the research plan. NJ performed the experiments, analyzed the data and wrote the manuscript. JPK and PK critically supervised the progress of the research, gave vital inputs in the improvement of the research work, and edited the manuscript. JPK funded the cost of the entire research through his grants. All authors have read and approve the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Handling Editor: Bhumi Nath Tripathi
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Keynote
The rice Tubby-like protein (TLP) encoding gene, OsFBT4, plays role in both ABA-dependent and independent stress signalling and its overexpression confers stage-specific tolerance to salt, drought, and other abiotic stresses in Arabidopsis.
Supplementary Information
Below is the link to the electronic supplementary material.
709_2022_1831_MOESM1_ESM.pdf
Supplementary file1 (PDF 455 KB) Fig S1. Exon organization of OsFBT4 and its translated product and validation of the OsFBT4-OE lines in Arabidopsis. (A) and (B) represents the organization of the OsFBT4- cDNA and its translated product, respectively. (C) and (D) represents the genomic DNA PCR- based confirmation of the OE lines, using a combination of CaMV 35S promoter-F and OsFBT4-R primers (1.3 kb), and NPT-II gene specific primers, respectively. The transcript level of OsFBT4 in the OE lines quantified by RT-qPCR using Actin-II as internal control is presented in (E). The asterisk marks (*) indicate significant difference (P-value<0.05). Fig S2. Response of the OsFBT4-OE lines at 15-d old plant stage to ABA and paraquat. (A) depicts growth response of the OE lines to exogenous ABA (15 µM) and paraquat (2 µM) after 15 days on the stress media. Physiological measurements such as Fv/Fm values (B); total chlorophyll content (C); and final rosette diameter (D) after completion of 15 days on the stress media are graphically presented. The results are represented as mean ± SE, where n=3. The asterisk marks (*) indicate significant difference (P-value<0.05). Fig S3. Estimation of membrane injury parameters in 15-d old OsFBT4-OE plants in response to different abiotic stresses. (A) depicts overall ROS (H2O2) accumulation in 15-d old plants as seen by DAB staining; (B) graphically represents the Fv/Fm values; (C) shows the total chlorophyll content (represented in mg/g fresh wt.); the ion leakage percentages are presented graphically in (D). (E) and (F) shows the MDA and proline content of the plants, respectively, in response to different stresses. The results are represented as mean ± SE, where n=3. The asterisk marks (*) indicate significant difference (P-value<0.05).
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jain, N., Khurana, P. & Khurana, J.P. Overexpression of a rice Tubby–like protein-encoding gene, OsFBT4, confers tolerance to abiotic stresses. Protoplasma 260, 1063–1079 (2023). https://doi.org/10.1007/s00709-022-01831-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-022-01831-5