Abstract
The most common neurodegenerative disorders, such as Alzheimer’s or Parkinson’s diseases, are characterized by synaptic dysfunction, neuronal loss and proteinaceous aggregates in central nervous system. The deposition of misfolded proteins constitutes neuropathological hallmarks of these diseases, grouped in the generic term of proteinopathies. Apart from these, other neurodegenerative diseases are characterized by genetic abnormalities like unstable repetitive simple sequence tracts (microsatellites) dispersed throughout the human genome. They are called repeat expansion disorders and include, for example, Huntington’s disease or frontotemporal dementia/amyotrophic lateral sclerosis phenotypes associated to an expansion in C9ORF72. The presence of the expanded DNA tract leads to molecular alterations at the DNA, RNA and protein levels associated to distinct mechanisms, such as loss-of-function (LOF), gain-of-function (GOF) including misfolding of physiological or mutant proteins, favoring their polymerization and aggregation. Therefore, specific proteinopathies also arise from these repeat expansion disorders. The molecular description of the nature and location of expanded tracts, highlighting the consequences onto clinical phenotypes will be first described. Specific focuses on the three pathomechanisms of the repeat expansions associated to proteinopathies will then be addressed. Lastly, we will show how progress in the understanding of these different mechanisms has led to recent advances in new/innovative therapeutic approaches and emergence of associated biomarkers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Neurodegenerative disorders are characterized by progressive loss of selectively vulnerable populations of neurons and can be classified according to clinical features and/or molecular abnormalities (Dugger and Dickson 2017). Indeed, the most prevalent neurodegenerative diseases are characterized by proteinaceous aggregates (hyper-phosphorylated tau and amyloid peptides in Alzheimer’s disease, alpha-synuclein in Parkinson’s disease, TDP-43 protein in frontotemporal dementia/amyotrophic lateral sclerosis…) (Noor et al. 2021). The detection of these aggregates in brain samples led to the protein-based molecular classification of neurodegenerative diseases (Kovacs 2017) which permits, in addition to specific clinical criteria, a definite diagnosis for these diseases (McKhann et al. 2011; Rascovsky et al. 2011; Watson et al. 2021). As a consequence, the concept of proteinopathies emerged and can be defined as diseases triggered by the aggregation of one or more physiological proteins becoming pathologically active after post-translational modifications and/or conformational changes, leading to an increased propensity to self-association and precipitation (Bayer 2015). The formation of protein aggregates follows a sequential distribution pattern in different brain regions, both protein nature and prototypic pattern being therefore specific to each neurodegenerative disease (Marsh 2019; Kovacs 2019). The better comprehension of these proteinopathies and an improved classification of patients’ cohorts paved the way to intensive research with the aim of developing effective disease-modifying treatments (Aisen et al. 2020; Cummings et al. 2019; Brys et al. 2019).
Among neurodegenerative disorders, a vast group of diseases is characterized by unstable repetitive simple sequence tracts (microsatellites) dispersed throughout the human genome. This group of conditions, called repeat expansion disorders (Depienne and Mandel 2021), includes most commonly inherited neurological disorders, such as Huntington’s disease, spinocerebellar ataxias, and most recently frontotemporal dementia/amyotrophic lateral sclerosis associated to an expansion in C9ORF72 (Loureiro et al. 2016; Rudnicki and Margolis 2003). In these diseases, the presence of an expanded tract in one specific gene is a mandatory condition to trigger various and intertwined molecular mechanisms at the DNA, RNA and protein levels, leading to neurodegeneration and disease onset (Malik et al. 2021; Schwartz et al. 2021). These molecular alterations are classically divided into loss-of-function (LOF) or gain-of-function (GOF) mechanisms (La Spada and Taylor 2010). Some of the latter may be responsible for the misfolding of physiological or mutant proteins, favoring their polymerization and aggregation. For this reason, specific proteinopathies also arise from repeat expansion disorders. Proteinopathies’ identification in clinical practice is less crucial than for Alzheimer’s disease and Parkinson’s disease, as laboratory diagnosis of repeat expansion disorders essentially relies on genetic testing (Paulson 2018). Nevertheless, the better understanding of the molecular mechanisms secondary to the presence of these unstable expansions allows for the conception of new disease-specific therapies and associated monitoring biomarkers (Benn et al. 2021; Hautbergue et al. 2021; Bakkar et al. 2015).
The first part of this review will focus on a molecular description of the nature and location of expanded tracts, highlighting the consequences of these genetic alterations onto clinical phenotypes. Then, three pathomechanisms of repeat expansion disorders leading to proteinopathies will be detailed in the second part of this article: the translation of mutant proteins containing repeated amino acids, the sequestration of RNA-binding proteins and the repeat associated non-AUG (RAN) translation of repeat peptides. Finally, the emergence of novel therapeutics and biomarkers associated to these particular molecular mechanisms will be discussed.
The molecular features of expanded DNA tracts in repeat expansion disorders
A first particular characteristic of repeat expansions is their dynamic behavior. Expanded alleles in repeat expansion disorders arise from polymorphic repeats in general population and often change size within or between tissues of affected individuals (Pearson et al. 2005). Moreover, expansions and longer normal alleles are more likely to increase than smaller normal alleles. Because of the size of these large expansions, genetic testing requires specific methodologies as high-throughput short-read sequencing remains unable to genotype long expansions (Chintalaphani et al. 2021). The mechanisms associated with such an instability are now better understood and would involve an alteration of DNA metabolic processes, such as replication, repair and/or recombination. Indeed, alternative DNA structures can be observed according to repeated sequences, including DNA triplexes, G-quadruplexes or hairpins, facilitating the emergence of large-scale expansions (Khristich and Mirkin 2020). In addition to genetic factors, the role of exogenous agents (as an example, different chemical compounds) was also explored (Gomes-Pereira and Monckton 2004). In parallel to somatic instability, germline instability is also described as an important mechanism of enlargement of expansions across generations (Bois and Jeffreys 1999). Paternal and maternal expansion or contraction biases exist, leading to an increased proportion of paternal or maternal disease-causing transmission to offspring in some repeat expansion diseases (Aziz et al. 2011).
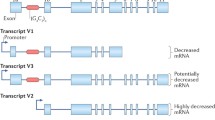
Different phenotypes can be observed according to the size of the repeat in a same gene. Fragile X-associated tremor ataxia syndrome (FXTAS) is a neurodegenerative disorder, with patients around 60 years old exhibiting a tremor and an ataxic gait, with possible neuropathy, parkinsonism and executive dysfunction. Patients harbor a pre-mutation in FMR1 with (CGG) repeats included between 55 and 200. On the contrary, an expansion with more than 200 (CGG) repeats in the same gene will lead to Fragile X syndrome, the most common cause of inherited intellectual disability and autism spectrum disorder (Cabal-Herrera et al. 2020). Such a genotype–phenotype correlation between repeat length and disease severity, named anticipation, is also observed for polyglutamine tract diseases (like Huntington’s disease) and DM1 (Fig. 1): the longer the repeat, the more severe the disease with an earlier onset (Fan et al. 2014; Kamsteeg et al. 2012).
A second characteristic of repeat expansion diseases is the important variety of repetitive tracts and their location throughout genome. The first expansions discovered were trinucleotide repeats including different motifs: (CGG) and (CAG) repeats in FMR1 and in AR, respectively, both on chromosome X. Trinucleotide repeats represent the majority motif in these diseases; however, repetitions of tetra-nucleotides, penta-nucleotides, hexa-nucleotides and even dodeca-nucleotides can also be found (Malik et al. 2021; Loureiro et al. 2016). Expandable repeats are located in various regions of their resident genes: coding regions, 5′ and 3′ untranslated regions, introns and promoter regions (Mirkin 2007). The location of repeats (coding or non-coding DNA regions as an example) is directly linked to the molecular pathomechanisms involved in disease development. Only trinucleotide repeats were discovered so far in coding sequences, leading to the formation of polyglutamine (repeat of CAG codons) or polyalanine (repeat of GCN codons) tracts within proteins (Depienne and Mandel 2021). Trinucleotide repeats are also frequently present in non-coding DNA regions: (CGG) trinucleotide repeats within FMR1 5′UTR causing fragile X syndrome (Verkerk et al. 1991), (GAA) repeat expansions in intron 1 of FXN related to Friedreich’s ataxia (Campuzano et al. 1996) and myotonic dystrophy type 1 (CTG) repeats within DMPK 3’UTR (Mahadevan et al. 1992) are some examples of these triplet repeat disorders. Tetra-nucleotide, penta-nucleotide and hexa-nucleotide repetitions are present in intronic regions. Interestingly, most penta-nucleotide expansions consist in a different expanded pathogenic motif compared to the normal motif observed in general population. As an example, CANVAS, the acronym for cerebellar ataxia, neuropathy and vestibular areflexia syndrome, is associated to a recessive expansion of a mutated (AAGGG)n repeated unit in intron 2 of RFC1, whereas the most prevalent motif is (AAAAG)11 in healthy controls (Cortese et al. 2019).
Finally, the nature of the repeated patterns is not necessarily homogeneous in some expansion diseases. As an example, in spinocerebellar ataxia type 1 (SCA1), (CAT) trinucleotides, coding for histidine amino acid, interrupt the poly(CAG) tract both in normal and expanded alleles. In normal alleles, this phenomenon could increase its stability. In expanded alleles, such interruptions could delay age of onset (Kraus-Perrotta and Lagalwar 2016). Non-coding expansions can also include interruptions. In DM1, (CCG), (GGC), (CTC) and (CAG) interruptions within 5′ and 3′ends of DMPK expansions are found with a global frequency of 3–5% in DM1 patients and are considered as a potential genetic modifier of DM1 phenotype (Pesovic et al. 2018). Table 1 presents a description of repeat expansion diseases, including the genomic location and the nature of expanded alleles.
The translation of mutant proteins containing repeated amino acids leads to protein deposits
Among repeat expansion disorders with trinucleotide repeats in coding sequences, those with a repetition of a (CAG) triplet are the most prevalent and lead to an elongated polyglutamine (polyQ) tract in the corresponding protein. They are often referred to as polyglutamine tract diseases or (CAG)-polyglutamine repeat diseases and they share several characteristics. Nine diseases are related to this group, with Huntington’s disease being the most frequent (Stoyas and La Spada 2018). Moreover, spinal and bulbar muscular atrophy (SBMA) and numerous SCAs complete this family (Table 1). Their onset is typically occurring in the midlife after a long time of silent pre-symptomatic stage. Nevertheless, as previously described, anticipation phenomena occur between generations and a genotype–phenotype correlation with higher severity linked to the highest repeat length for this group of disease particularly. Larger expansions could occur in offspring, mainly when paternal transmission, leading to a decrease in the disease age of onset in successive generation. As a result, phenotypes could sometimes vary in the same family (Paulson 2018; Gatchel and Zoghbi 2005).
For all diseases, the normal polyQ repeat length is variable in the non-affected population but above a specific threshold of repeats, around 30–40 (CAG), an increasing size of the glutamine stretch leads to the production of a mutant and toxic isoform of the native protein. Studies exploring how the structure of the proteins are modified by the extended stretch in mutant polyQ proteins show that native proteins are mainly disordered but acquire β-strand rich conformations in the mutant isoforms (Wetzel 2012). This polyQ stretch in the mutant protein leads to conformational changes, abnormal folding of the protein, tends to form oligomers, then promoting aggregation into fibrils, finally forming inclusions (Cooper et al. 1998; Hoffner et al. 2005). Experiments showed that various monomers of different size can assembly into larger polymers called oligomers. These oligomers constitute an intermediate state before becoming insoluble and accumulating into larger amorphous aggregates (Hands and Wyttenbach 2010). The most toxic state of the mutant protein seems to be small or intermediate size oligomers but not aggregates, depending on the considered expansions (Miller et al. 2011; Takahashi et al. 2008). However, these inclusions constitute hallmarks of the diseases and can be detected in tissues, particularly in neurons where they are mainly but not exclusively localized into the nucleus (DiFiglia et al. 1997). In HD for example, neuropile inclusions of mutant huntingtin protein (mHtt) seem to be much frequent than nuclear aggregates in adult phenotypes (Gutekunst et al. 1999) and their size increases according to the diseases’ stages. A unique deposition pattern within the central nervous system seems to be linked to each disease, with a higher extensive number of inclusion associated to a highest severity of diseases (Davies et al. 1998).
Inclusions are polymorphic structures, containing different species of proteins. In HD aggregates, mHtt was the main component of inclusions, but other proteins have been detected such as ubiquitin and wild type Htt (Kazantsev et al. 1999). MHtt essentially truncated after proteolytic cleavage by caspase was also identified into inclusions, even early during the first stages of the disease progression in human (Goldberg et al. 1996; Wellington et al. 2002). Protein cleavage seems then an important feature for disease development. Indeed, in a mice model expressing mHtt and resistant to cleavage by caspase-6, mice maintain their neuronal functions without developing striatal neurodegeneration (Graham et al. 2006). Unfortunately, in these resistant mice models, alternative enzyme cleavage can restore the proteolytic event, leading finally to neurodegeneration (Wong et al. 2015). In another polyQ disease, spinocerebellar ataxia type 3 (SCA3), the inhibition of the calpain cleavage of the expanded Ataxin 3 protein appears to stop the formation of aggregates, highlighting once again the general importance of protein cleavage in the toxicity process (Haacke et al. 2007; Koch et al. 2011).
Apart from (CAG)-polyglutamine repeat diseases, polyalanine (polyA) tract expansions exist and the latter have been associated with different developmental diseases (Moumne et al. 2008). These polyA extensive tracts also give a propensity for the mutant proteins to self-assembly and form fibrils, leading to aggregation (Di Lascio et al. 2020).
RNA gain-of-toxicity leads to the sequestration of physiological proteins into foci
Many repeat expansion disorders are associated to a potential or proved RNA toxicity (Depienne and Mandel 2021). RNAs harboring repeat expansions adopt unusual secondary structures, varying according to the repeated motif from hairpins to stable G-quadruplexes (Bugaut et al. 2012). Other factors can also influence RNAs’ secondary structures as sequences flanking repeats, potential repeat interruptions and intermolecular associations (Ciesiolka et al. 2017). RNAs are then retained in the cell nucleus and sequester various RNA-binding proteins (RBPs), forming insoluble nuclear inclusions called foci (Chan 2014). Muscleblind-like proteins are RBPs sharing structural similarities, including four zinc-finger domains critical for recognizing a common consensus sequence in pre-mRNA and mRNA targets. As an example, Mbnl1 was detected in a variety of repeat-formed foci, including (CUG), (CCUG), (CAG) and (CGG) RNA inclusions. The sequestration of Mbnl1 directly impairs splicing of several key regulatory target pre-mRNAs (CLC2, IR2, cTNNT2…) in muscles and neural cells, explaining DM1 phenotypic features (Konieczny et al. 2014). Interestingly, the presence of Mbnl1 in RNA–RBP foci was reported to be not a consequence, but a necessary condition for the formation of foci with RNAs including (CUG) repeats (Querido et al. 2011).
Another example of RBPs’ sequestration is the interaction of nucleolin to (CAG) expanded RNAs. Nucleolin is a multifunctional protein involved in various steps of ribosome biogenesis. An alteration of these processes leads to nucleolar stress and apoptosis (Pfister 2019). This protein normally binds to an upstream control element of the rRNA promoter, thus protecting this region from CpG hypermethylation. Because of a competition with (CAG) repeats, nucleolin prevents no longer hypermethylation of rRNA promoter, leading to a reduction of rRNA expression (Marti 2016).
In C9ORF72, amyotrophic lateral sclerosis and/or frontotemporal dementia (ALS/FTD), the biggest group among RBPs that bind and co-localize with RNA foci is the heterogeneous nuclear ribonucleoprotein group (hnRNPs) (Kumar et al. 2017). On another note, FUS is one of the proteins most consistently shown to bind to repeated (GGGGCC) expanded RNA, pinpointing convergent mechanisms between FUS ALS and C9ORF72 ALS. However, the involvement of RNA foci in disease pathogenesis remains debated. May be expanded RNAs’ toxicity could be primarily mediated before integration into foci, developing further interrogations and creating new research directions (Swinnen et al. 2020).
The products of repeat associated non-AUG (RAN) translation are prone to self-aggregation
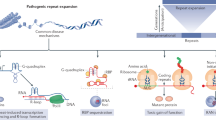
Repeat-associated non-AUG (RAN) translation is a non-canonical translational initiation process enabling elongation through a repeat strand in the absence of an (AUG) initiation codon and in multiple reading frames according to repetitive DNA tracts, producing multiple homo-polymeric proteins, dipeptide repeat- and more complex polypeptide-repeat proteins (Green et al. 2016). In 2011, Zu et al. first described this mechanism in SCA8 and DM1, harboring CAG and CTG repeats, respectively (Zu et al. 2011). Ten years later, RAN-translated proteins were described in Huntington’s disease and HDL2, DM1 and DM2, FXTAS, SCA2, SCA3, SCA8, SCA36 and ALS/FTD associated to C9ORF72. Interestingly, RAN translation can occur from coding and non-coding regions of both sense and antisense RNA transcripts carrying expansions (Fig. 2) (Castelli et al. 2021).
Different RAN products can be accumulated within a single cell, suggesting that this process can occur in multiple reading frames in parallel. However, not all theoretical repeat peptides are observed in disease. As an example, in fragile X-associated tremor/ataxia syndrome (FXTAS) associated to CGG repeats in FMR1, no products can be seen in (CGG-Arginine) reading frame even at larger repeat sizes (above 100 CGG repeats), whereas RAN translation occurs in (GGC-Glycine) reading frame within a normal range of repeats (Todd et al. 2013). This may suggest that RAN translation implies many interdependent factors, including the surrounding sequence of the repeat, the nature of amino acids that are produced and the length of the repeat (Kearse and Todd 2014). Many microsatellite expansion mutations are GC-rich sequences that form secondary structures, like G-quadruplex (G4) structures, similar to internal ribosome entry sites (IRES) (Hellen and Sarnow 2001). In this alternative initiation pathway, structured RNAs directly recruit the preinitiation complex (Komar and Hatzoglou 2011).
The toxicity of RAN peptides was particularly studied in ALS/FTD associated to C9ORF72. Five dipeptide repeat (DPR) proteins are generated according to (GGGGCC) repeats in both sense and antisense transcripts: poly-GA, poly-GP, poly-GR, poly-PR and poly-PA proteins. Marked differences exist between these five DPR, the most toxic species being poly-GR and poly-PR, due to their arginine-rich content (Nguyen et al. 2019). Their biophysical properties favor binding to other proteins such as hnRNPA1/2, leading to a disruption in pre-mRNA splicing and RNA biogenesis. Consequently, exposures of cell lines to these synthetic peptides led to a decrease of cell viability (Kwon et al. 2014). Following cellular uptake, the migration of poly-PR to nucleoli and the interaction to TIA-1 are associated to the formation of stress granules (Wen et al. 2014). Non-arginine-containing DRP proteins also appear important to neurodegeneration: as an example, the expression of poly-GA in primary mammalian neurons causes increased toxicity through impairment of the ubiquitin–proteasome system (Green et al. 2016) and enhanced the formation of toxic amyloid fibrils (Chang et al. 2016).
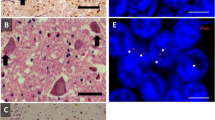
Neuropathological examinations of human tissues show aggregates of RAN proteins for different repeat expansion disorders (Castelli et al. 2021). In HD, RAN proteins (polyA, polyS, polyL, and polyC) accumulate most abundantly in brain regions with neuronal loss, microglial activation and apoptosis, including the striatum (caudate and putamen), a region severely affected in HD, but also white matter and cerebellum in juvenile-onset cases. Interestingly, polyQ aggregates are not detected in regions with the most intense RAN protein staining (Banez-Coronel et al. 2015). In DM2, anti-LPAC and anti-QAGR antibodies show positive staining in multiple brain regions. Cytoplasmic LPAC RAN proteins accumulate primarily in gray matter regions of the brain; on the contrary, immunohistochemistry shows that QAGR RAN proteins accumulate in the white matter regions (frontal cortex, basal ganglia and hippocampus) with punctate nuclear aggregates often located at the nuclear membrane (Zu et al. 2017). SCA8 human autopsy cases show polyS aggregates in the cerebellum, brainstem and cortex, increasing with age and disease progression (Ayhan et al. 2018). Finally, c9RAN proteins are a major component of TDP-43-negative, p62-positive inclusions in ALS/FTD associated to C9ORF72. Inclusions of poly-GP and poly-GA are abundant in cerebellum, hippocampus, and neocortical regions (Al-Sarraj et al. 2011; Gendron et al. 2015). Interestingly, cerebellar poly-GP concentrations seem to vary according to the clinical phenotype, with lower values for ALS (Gendron et al. 2015). Finally, the observation that DPR inclusions are rare in spinal cord (contrary to TDP43 inclusions) and absent from motor neurons in patients with C9ORF72 ALS (Gomez-Deza et al. 2015) led to the following question: to what extent does the production of DPR proteins confer neurodegeneration in vivo (Schmitz et al. 2021)?
The emergence of disease-modifying therapeutic opportunities for repeat expansions disorders
Current therapies for neurodegenerative disorders remain mostly aimed at symptomatic relief (Sudhakar and Richardson 2019). However, must research has been undertaken to develop disease-modifying treatments. As substantial progress has been made in our understanding of the pathogenesis of neurodegenerative disorders, different gene-based therapies are being evaluated today and can be divided into DNA-targeting and RNA-targeting approaches (Sun and Roy 2021), some of them already having a marketing authorization to treat spinal muscular atrophy (Lee et al. 2019). The possibility of using gene-delivery, gene-editing or knockdown techniques would vary for each repeat expansion disorder, according to the predominant pathological mechanism: the loss of-function of a physiological protein, the expression of a mutant protein, the presence of RNA foci and RAN translation.
The three main DNA-targeting approaches are ZFNs (zinc-finger nucleases), TALENs (transcription activator-like effector nucleases) and CRISPR/Cas (clustered regularly interspaced short palindromic repeats/CRISPR-associated protein). These approaches combine a specific DNA-binding element (small peptides for ZFN and TALEN or a single-guide RNA for CRISPR/Cas9) and a nuclease, ultimately leading to the knocking-out of the targeted gene (Gaj et al. 2013). These approaches could provide long-term treatment and eliminate inter-generational transmission of repeat expansion disorders (Tabrizi et al. 2020). As an example, the use of CRISPR-Cas9 reduced mutant Htt protein inclusions in a mouse model of HD (Ekman et al. 2019), reduced nuclear RNA foci in the muscle of DM1 mice (Lo Scrudato et al. 2019) and reduced RNA foci and DPR levels in human cell lines with (GGGGCC) repeats (Pinto et al. 2017). These approaches present several limitations, including viral delivery of these compounds; thus, immunogenicity, intrathecal administration, and potential off-targets could be irreversible (Tabrizi et al. 2020).
RNA-based editing alters gene expression at the transcript level. Since RNA is transient, there is lesser risk of permanent deleterious effects but, on the contrary, the need for repeated administration is challenging (Sun and Roy 2021). The administration of small interfering RNAs (siRNAs) or microRNAs (miRNAs) leads to RNA interference (RNAi), a cellular process promoting the degradation of a target messenger RNA (mRNA) with a complementary sequence (Lam et al. 2015). Briefly, these small RNAs need to be loaded onto an argonaute protein to form the effector complex referred to as RNA‐induced silencing complex (RISC) (Nakanishi 2016; Valencia-Sanchez et al. 2006). Like DNA-targeting approaches, extensive research was performed to improve RNAi-inducing therapies delivery to brain cells (Sarisozen et al. 2015). RNAi approach improved motor coordination, restored cerebellar morphology and resolved characteristic ataxin-1 inclusions in Purkinje cells of SCA1 mice (Xia et al. 2004). Boudreau et al. reviewed the different proof-of-concept studies testing therapeutic RNAi for repeat expansion and other CNS disorders (Boudreau and Davidson 2010). The second RNA-targeting approach relies on the use of antisense oligonucleotides (ASOs). These synthetic, single-stranded, modified DNA molecules can bind to mRNAs or pre-mRNAs, forming an RNA–DNA hybrid that becomes a substrate for RNase H, which results in target mRNA degradation (Rinaldi and Wood 2018). Since the ASOs available today do not cross the blood–brain barrier, the application must be carried out by intrathecal injection in the case of CNS disorders and needs to be administered on a regular basis (Brenner et al. 2020). Single-dose ASOs reduced RNA foci, DPR proteins, and behavioral deficits in C9ORF72 mice (Jiang et al. 2016). In a DM1 mouse model, ASOs induced degradation of expanded DMPK transcripts, disrupted RNA foci and could release binding of Mbnl1 to the toxic RNA in skeletal muscle (Lee et al. 2012). Finally, in HD patients, the administration of ASO HTT led to a dose-dependent decrease in cerebrospinal fluid (CSF) concentrations of mHTT and clinical improvements in comparison to patients who received placebo (Tabrizi et al. 2020).
The emergence of disease-modifying strategies goes hand in hand with the development of biomarkers, which could permit to monitor the effectiveness of these treatments. As previously described, the measurement of CSF mHtt is important for the development of specific therapeutic strategies in HD (Wild et al. 2015). In a same way, poly-GP protein, one of C9RAN proteins associated to C9ORF72 expansions, was already detectable in CSF (Gendron et al. 2017). In parallel to these biomarkers linked to the pathomechanisms involved in a repeat expansion disorder, the use of surrogate markers could be of great value. Among these markers, neurofilament light chain protein (NfL), a marker of neuronal damage, seems particularly interesting, as it can be measured both in CSF and in serum (Khalil et al. 2020). As this biomarker reflects early neuronal injury, even before clinical expression, its iterative determination allows a longitudinal follow-up of patients (Byrne et al. 2017; Lambertsen et al. 2020). Moreover, NfL measurement could help monitoring treatment response (Yuan and Nixon 2021). Thus, biological markers appear as useful tools complementary to genetic testing, which remains essential for the diagnosis of repeat expansion disorders.
Conclusion
Repeat expansion disorders constitute a various group of diseases leading to neurological degeneration. The diversity of repetitive tracts and their location throughout genome cause multiple modifications on molecular and protein levels, leading to cell toxicity. Gain-of-function mechanisms lead to proteinopathies: mutant proteins, RNA-binding proteins in RNA foci or RAN products can form aggregates, thus leading to inclusions’ formation. Not all these mechanisms are necessarily fully exclusive, as combinations were proven in many diseases. The better understanding of these molecular and protein dysfunctions has contributed to the development of disease-modifying therapeutic strategies, numerous being under clinical evaluation. Moreover, it also permitted the identification of markers underlying specific pathomechanisms, whose measurements could present interest in the monitoring of drug efficacy.
References
Aisen PS, Cummings J, Doody R, Kramer L, Salloway S, Selkoe DJ, Sims J, Sperling RA, Vellas B (2020) The future of anti-amyloid trials. J Prev Alzheimer’s Dis 7(3):146–151. https://doi.org/10.14283/jpad.2020.24
Al-Sarraj S, King A, Troakes C, Smith B, Maekawa S, Bodi I, Rogelj B, Al-Chalabi A, Hortobagyi T, Shaw CE (2011) p62 positive, TDP-43 negative, neuronal cytoplasmic and intranuclear inclusions in the cerebellum and hippocampus define the pathology of C9orf72-linked FTLD and MND/ALS. Acta Neuropathol 122(6):691–702. https://doi.org/10.1007/s00401-011-0911-2
Ayhan F, Perez BA, Shorrock HK, Zu T, Banez-Coronel M, Reid T, Furuya H, Clark HB, Troncoso JC, Ross CA, Subramony SH, Ashizawa T, Wang ET, Yachnis AT, Ranum LP (2018) SCA8 RAN polySer protein preferentially accumulates in white matter regions and is regulated by eIFF. EMBO J https://doi.org/10.15252/embj.201899023
Aziz NA, van Belzen MJ, Coops ID, Belfroid RD, Roos RA (2011) Parent-of-origin differences of mutant HTT CAG repeat instability in Huntington’s disease. Eur J Med Genet 54(4):e413-418. https://doi.org/10.1016/j.ejmg.2011.04.002
Bakkar N, Boehringer A, Bowser R (2015) Use of biomarkers in ALS drug development and clinical trials. Brain Res 1607:94–107. https://doi.org/10.1016/j.brainres.2014.10.031
Banez-Coronel M, Ayhan F, Tarabochia AD, Zu T, Perez BA, Tusi SK, Pletnikova O, Borchelt DR, Ross CA, Margolis RL, Yachnis AT, Troncoso JC, Ranum LP (2015) RAN translation in Huntington disease. Neuron 88(4):667–677. https://doi.org/10.1016/j.neuron.2015.10.038
Bayer TA (2015) Proteinopathies, a core concept for understanding and ultimately treating degenerative disorders? Eur Neuropsychopharmacol 25(5):713–724. https://doi.org/10.1016/j.euroneuro.2013.03.007
Benn CL, Gibson KR, Reynolds DS (2021) Drugging DNA damage repair pathways for trinucleotide repeat expansion diseases. J Huntington’s Dis 10(1):203–220. https://doi.org/10.3233/JHD-200421
Bois P, Jeffreys AJ (1999) Minisatellite instability and germline mutation. Cell Mol Life Sci CMLS 55(12):1636–1648. https://doi.org/10.1007/s000180050402
Boudreau RL, Davidson BL (2010) RNAi therapeutics for CNS disorders. Brain Res 1338:112–121. https://doi.org/10.1016/j.brainres.2010.03.038
Brenner D, Ludolph AC, Weishaupt JH (2020) Gene specific therapies - the next therapeutic milestone in neurology. Neurolog Res Pract 2:25. https://doi.org/10.1186/s42466-020-00075-z
Brys M, Fanning L, Hung S, Ellenbogen A, Penner N, Yang M, Welch M, Koenig E, David E, Fox T, Makh S, Aldred J, Goodman I, Pepinsky B, Liu Y, Graham D, Weihofen A, Cedarbaum JM (2019) Randomized phase I clinical trial of anti-alpha-synuclein antibody BIIB054. Movem Disord off J Movem Disord Soc 34(8):1154–1163. https://doi.org/10.1002/mds.27738
Bugaut A, Murat P, Balasubramanian S (2012) An RNA hairpin to G-quadruplex conformational transition. J Am Chem Soc 134(49):19953–19956. https://doi.org/10.1021/ja308665g
Byrne LM, Rodrigues FB, Blennow K, Durr A, Leavitt BR, Roos RAC, Scahill RI, Tabrizi SJ, Zetterberg H, Langbehn D, Wild EJ (2017) Neurofilament light protein in blood as a potential biomarker of neurodegeneration in Huntington’s disease: a retrospective cohort analysis. Lancet Neurol 16(8):601–609. https://doi.org/10.1016/S1474-4422(17)30124-2
Cabal-Herrera AM, Tassanakijpanich N, Salcedo-Arellano MJ, Hagerman RJ (2020) Fragile X-associated tremor/Ataxia syndrome (FXTAS): pathophysiology and clinical implications. Int J Mol Sci. https://doi.org/10.3390/ijms21124391
Campuzano V, Montermini L, Molto MD, Pianese L, Cossee M, Cavalcanti F, Monros E, Rodius F, Duclos F, Monticelli A, Zara F, Canizares J, Koutnikova H, Bidichandani SI, Gellera C, Brice A, Trouillas P, De Michele G, Filla A, De Frutos R, Palau F, Patel PI, Di Donato S, Mandel JL, Cocozza S, Koenig M, Pandolfo M (1996) Friedreich’s ataxia: autosomal recessive disease caused by an intronic GAA triplet repeat expansion. Science (new York, NY) 271(5254):1423–1427. https://doi.org/10.1126/science.271.5254.1423
Castelli LM, Huang WP, Lin YH, Chang KY, Hautbergue GM (2021) Mechanisms of repeat-associated non-AUG translation in neurological microsatellite expansion disorders. Biochem Soc Trans 49(2):775–792. https://doi.org/10.1042/BST20200690
Chan HY (2014) RNA-mediated pathogenic mechanisms in polyglutamine diseases and amyotrophic lateral sclerosis. Front Cell Neurosci 8:431. https://doi.org/10.3389/fncel.2014.00431
Chang YJ, Jeng US, Chiang YL, Hwang IS, Chen YR (2016) The glycine-alanine dipeptide repeat from C9orf72 Hexanucleotide expansions forms toxic amyloids possessing cell-to-cell transmission properties. J Biol Chem 291(10):4903–4911. https://doi.org/10.1074/jbc.M115.694273
Chintalaphani SR, Pineda SS, Deveson IW, Kumar KR (2021) An update on the neurological short tandem repeat expansion disorders and the emergence of long-read sequencing diagnostics. Acta Neuropathol Commun 9(1):98. https://doi.org/10.1186/s40478-021-01201-x
Ciesiolka A, Jazurek M, Drazkowska K, Krzyzosiak WJ (2017) Structural characteristics of simple RNA repeats associated with disease and their deleterious protein interactions. Front Cell Neurosci 11:97. https://doi.org/10.3389/fncel.2017.00097
Cooper JK, Schilling G, Peters MF, Herring WJ, Sharp AH, Kaminsky Z, Masone J, Khan FA, Delanoy M, Borchelt DR, Dawson VL, Dawson TM, Ross CA (1998) Truncated N-terminal fragments of huntingtin with expanded glutamine repeats form nuclear and cytoplasmic aggregates in cell culture. Hum Mol Genet 7(5):783–790. https://doi.org/10.1093/hmg/7.5.783
Cortese A, Simone R, Sullivan R, Vandrovcova J, Tariq H, Yau WY, Humphrey J, Jaunmuktane Z, Sivakumar P, Polke J, Ilyas M, Tribollet E, Tomaselli PJ, Devigili G, Callegari I, Versino M, Salpietro V, Efthymiou S, Kaski D, Wood NW, Andrade NS, Buglo E, Rebelo A, Rossor AM, Bronstein A, Fratta P, Marques WJ, Zuchner S, Reilly MM, Houlden H (2019) Biallelic expansion of an intronic repeat in RFC1 is a common cause of late-onset ataxia. Nat Genet 51(4):649–658. https://doi.org/10.1038/s41588-019-0372-4
Cummings J, Blennow K, Johnson K, Keeley M, Bateman RJ, Molinuevo JL, Touchon J, Aisen P, Vellas B (2019) Anti-tau trials for Alzheimer’s disease: a report from the EU/US/CTAD task force. J Prev Alzheimer’s Dis 6(3):157–163. https://doi.org/10.14283/jpad.2019.14
Davies SW, Beardsall K, Turmaine M, DiFiglia M, Aronin N, Bates GP (1998) Are neuronal intranuclear inclusions the common neuropathology of triplet-repeat disorders with polyglutamine-repeat expansions? Lancet 351(9096):131–133. https://doi.org/10.1016/S0140-6736(97)08360-8
Depienne C, Mandel JL (2021) 30 years of repeat expansion disorders: What have we learned and what are the remaining challenges? Am J Hum Genet 108(5):764–785. https://doi.org/10.1016/j.ajhg.2021.03.011
Di Lascio S, Benfante R, Cardani S, Fornasari D (2020) Research Advances on Therapeutic Approaches to Congenital Central Hypoventilation Syndrome (CCHS). Front Neurosci 14:615666. https://doi.org/10.3389/fnins.2020.615666
DiFiglia M, Sapp E, Chase KO, Davies SW, Bates GP, Vonsattel JP, Aronin N (1997) Aggregation of huntingtin in neuronal intranuclear inclusions and dystrophic neurites in brain. Science (new York, NY) 277(5334):1990–1993. https://doi.org/10.1126/science.277.5334.1990
Dugger BN, Dickson DW (2017) Pathology of Neurodegenerative Diseases. Cold Spring Harbor Perspect Biol. https://doi.org/10.1101/cshperspect.a028035
Ekman FK, Ojala DS, Adil MM, Lopez PA, Schaffer DV, Gaj T (2019) CRISPR-Cas9-mediated genome editing increases lifespan and improves motor deficits in a Huntington’s disease mouse model. Mol Therapy Nucleic Acids 17:829–839. https://doi.org/10.1016/j.omtn.2019.07.009
Fan HC, Ho LI, Chi CS, Chen SJ, Peng GS, Chan TM, Lin SZ, Harn HJ (2014) Polyglutamine (PolyQ) diseases: genetics to treatments. Cell Transplant 23(4–5):441–458. https://doi.org/10.3727/096368914X678454
Gaj T, Gersbach CA, Barbas CF 3rd (2013) ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol 31(7):397–405. https://doi.org/10.1016/j.tibtech.2013.04.004
Gatchel JR, Zoghbi HY (2005) Diseases of unstable repeat expansion: mechanisms and common principles. Nat Rev Genet 6(10):743–755. https://doi.org/10.1038/nrg1691
Gendron TF, van Blitterswijk M, Bieniek KF, Daughrity LM, Jiang J, Rush BK, Pedraza O, Lucas JA, Murray ME, Desaro P, Robertson A, Overstreet K, Thomas CS, Crook JE, Castanedes-Casey M, Rousseau L, Josephs KA, Parisi JE, Knopman DS, Petersen RC, Boeve BF, Graff-Radford NR, Rademakers R, Lagier-Tourenne C, Edbauer D, Cleveland DW, Dickson DW, Petrucelli L, Boylan KB (2015) Cerebellar c9RAN proteins associate with clinical and neuropathological characteristics of C9ORF72 repeat expansion carriers. Acta Neuropathol 130(4):559–573. https://doi.org/10.1007/s00401-015-1474-4
Gendron TF, Chew J, Stankowski JN, Hayes LR, Zhang YJ, Prudencio M, Carlomagno Y, Daughrity LM, Jansen-West K, Perkerson EA, O’Raw A, Cook C, Pregent L, Belzil V, van Blitterswijk M, Tabassian LJ, Lee CW, Yue M, Tong J, Song Y, Castanedes-Casey M, Rousseau L, Phillips V, Dickson DW, Rademakers R, Fryer JD, Rush BK, Pedraza O, Caputo AM, Desaro P, Palmucci C, Robertson A, Heckman MG, Diehl NN, Wiggs E, Tierney M, Braun L, Farren J, Lacomis D, Ladha S, Fournier CN, McCluskey LF, Elman LB, Toledo JB, McBride JD, Tiloca C, Morelli C, Poletti B, Solca F, Prelle A, Wuu J, Jockel-Balsarotti J, Rigo F, Ambrose C, Datta A, Yang W, Raitcheva D, Antognetti G, McCampbell A, Van Swieten JC, Miller BL, Boxer AL, Brown RH, Bowser R, Miller TM, Trojanowski JQ, Grossman M, Berry JD, Hu WT, Ratti A, Traynor BJ, Disney MD, Benatar M, Silani V, Glass JD, Floeter MK, Rothstein JD, Boylan KB, Petrucelli L (2017) Poly(GP) proteins are a useful pharmacodynamic marker for C9ORF72-associated amyotrophic lateral sclerosis. Sci Transl Med. https://doi.org/10.1126/scitranslmed.aai7866
Goldberg YP, Nicholson DW, Rasper DM, Kalchman MA, Koide HB, Graham RK, Bromm M, Kazemi-Esfarjani P, Thornberry NA, Vaillancourt JP, Hayden MR (1996) Cleavage of huntingtin by apopain, a proapoptotic cysteine protease, is modulated by the polyglutamine tract. Nat Genet 13(4):442–449. https://doi.org/10.1038/ng0896-442
Gomes-Pereira M, Monckton DG (2004) Chemically induced increases and decreases in the rate of expansion of a CAG*CTG triplet repeat. Nucleic Acids Res 32(9):2865–2872. https://doi.org/10.1093/nar/gkh612
Gomez-Deza J, Lee YB, Troakes C, Nolan M, Al-Sarraj S, Gallo JM, Shaw CE (2015) Dipeptide repeat protein inclusions are rare in the spinal cord and almost absent from motor neurons in C9ORF72 mutant amyotrophic lateral sclerosis and are unlikely to cause their degeneration. Acta Neuropathol Commun 3:38. https://doi.org/10.1186/s40478-015-0218-y
Graham RK, Deng Y, Slow EJ, Haigh B, Bissada N, Lu G, Pearson J, Shehadeh J, Bertram L, Murphy Z, Warby SC, Doty CN, Roy S, Wellington CL, Leavitt BR, Raymond LA, Nicholson DW, Hayden MR (2006) Cleavage at the caspase-6 site is required for neuronal dysfunction and degeneration due to mutant huntingtin. Cell 125(6):1179–1191. https://doi.org/10.1016/j.cell.2006.04.026
Green KM, Linsalata AE, Todd PK (2016) RAN translation-What makes it run? Brain Res 1647:30–42. https://doi.org/10.1016/j.brainres.2016.04.003
Gutekunst C-A, Li S-H, Yi H, Mulroy JS, Kuemmerle S, Jones R, Rye D, Ferrante RJ, Hersch SM, Li X-J (1999) Nuclear and neuropil aggregates in Huntington’s disease: relationship to neuropathology. J Neurosci 19(7):2522. https://doi.org/10.1523/JNEUROSCI.19-07-02522.1999
Haacke A, Hartl FU, Breuer P (2007) Calpain inhibition is sufficient to suppress aggregation of polyglutamine-expanded ataxin-3. J Biol Chem 282(26):18851–18856. https://doi.org/10.1074/jbc.M611914200
Hands SL, Wyttenbach A (2010) Neurotoxic protein oligomerisation associated with polyglutamine diseases. Acta Neuropathol 120(4):419–437. https://doi.org/10.1007/s00401-010-0703-0
Hautbergue GM, Cleary JD, Guo S, Ranum LPW (2021) Therapeutic strategies for C9orf72 amyotrophic lateral sclerosis and frontotemporal dementia. Curr Opin Neurol. https://doi.org/10.1097/WCO.0000000000000984
Hellen CU, Sarnow P (2001) Internal ribosome entry sites in eukaryotic mRNA molecules. Genes Dev 15(13):1593–1612. https://doi.org/10.1101/gad.891101
Hoffner G, Island ML, Djian P (2005) Purification of neuronal inclusions of patients with Huntington’s disease reveals a broad range of N-terminal fragments of expanded huntingtin and insoluble polymers. J Neurochem 95(1):125–136. https://doi.org/10.1111/j.1471-4159.2005.03348.x
Jiang J, Zhu Q, Gendron TF, Saberi S, McAlonis-Downes M, Seelman A, Stauffer JE, Jafar-Nejad P, Drenner K, Schulte D, Chun S, Sun S, Ling SC, Myers B, Engelhardt J, Katz M, Baughn M, Platoshyn O, Marsala M, Watt A, Heyser CJ, Ard MC, De Muynck L, Daughrity LM, Swing DA, Tessarollo L, Jung CJ, Delpoux A, Utzschneider DT, Hedrick SM, de Jong PJ, Edbauer D, Van Damme P, Petrucelli L, Shaw CE, Bennett CF, Da Cruz S, Ravits J, Rigo F, Cleveland DW, Lagier-Tourenne C (2016) Gain of Toxicity from ALS/FTD-linked repeat expansions in C9ORF72 is alleviated by antisense oligonucleotides targeting GGGGCC-containing RNAs. Neuron 90(3):535–550. https://doi.org/10.1016/j.neuron.2016.04.006
Kamsteeg EJ, Kress W, Catalli C, Hertz JM, Witsch-Baumgartner M, Buckley MF, van Engelen BG, Schwartz M, Scheffer H (2012) Best practice guidelines and recommendations on the molecular diagnosis of myotonic dystrophy types 1 and 2. Eur J Hum Genet EJHG 20(12):1203–1208. https://doi.org/10.1038/ejhg.2012.108
Kazantsev A, Preisinger E, Dranovsky A, Goldgaber D, Housman D (1999) Insoluble detergent-resistant aggregates form between pathological and nonpathological lengths of polyglutamine in mammalian cells. Proc Natl Acad Sci 96(20):11404. https://doi.org/10.1073/pnas.96.20.11404
Kearse MG, Todd PK (2014) Repeat-associated non-AUG translation and its impact in neurodegenerative disease. Neurotherapeutics 11(4):721–731. https://doi.org/10.1007/s13311-014-0292-z
Khalil M, Pirpamer L, Hofer E, Voortman MM, Barro C, Leppert D, Benkert P, Ropele S, Enzinger C, Fazekas F, Schmidt R, Kuhle J (2020) Serum neurofilament light levels in normal aging and their association with morphologic brain changes. Nat Commun 11(1):812. https://doi.org/10.1038/s41467-020-14612-6
Khristich AN, Mirkin SM (2020) On the wrong DNA track: Molecular mechanisms of repeat-mediated genome instability. J Biol Chem 295(13):4134–4170. https://doi.org/10.1074/jbc.REV119.007678
Koch P, Breuer P, Peitz M, Jungverdorben J, Kesavan J, Poppe D, Doerr J, Ladewig J, Mertens J, Tüting T, Hoffmann P, Klockgether T, Evert BO, Wüllner U, Brüstle O (2011) Excitation-induced ataxin-3 aggregation in neurons from patients with Machado-Joseph disease. Nature 480(7378):543–546. https://doi.org/10.1038/nature10671
Komar AA, Hatzoglou M (2011) Cellular IRES-mediated translation: the war of ITAFs in pathophysiological states. Cell Cycle 10(2):229–240. https://doi.org/10.4161/cc.10.2.14472
Konieczny P, Stepniak-Konieczna E, Sobczak K (2014) MBNL proteins and their target RNAs, interaction and splicing regulation. Nucleic Acids Res 42(17):10873–10887. https://doi.org/10.1093/nar/gku767
Kovacs GG (2017) Concepts and classification of neurodegenerative diseases. Handb Clin Neurol 145:301–307. https://doi.org/10.1016/B978-0-12-802395-2.00021-3
Kovacs GG (2019) Molecular pathology of neurodegenerative diseases: principles and practice. J Clin Pathol 72(11):725–735. https://doi.org/10.1136/jclinpath-2019-205952
Kraus-Perrotta C, Lagalwar S (2016) Expansion, mosaicism and interruption: mechanisms of the CAG repeat mutation in spinocerebellar ataxia type 1. Cerebell Ataxias 3:20. https://doi.org/10.1186/s40673-016-0058-y
Kumar V, Hasan GM, Hassan MI (2017) Unraveling the role of RNA mediated toxicity of C9orf72 repeats in C9-FTD/ALS. Front Neurosci 11:711. https://doi.org/10.3389/fnins.2017.00711
Kwon I, Xiang S, Kato M, Wu L, Theodoropoulos P, Wang T, Kim J, Yun J, Xie Y, McKnight SL (2014) Poly-dipeptides encoded by the C9orf72 repeats bind nucleoli, impede RNA biogenesis, and kill cells. Science (new York, NY) 345(6201):1139–1145. https://doi.org/10.1126/science.1254917
La Spada AR, Taylor JP (2010) Repeat expansion disease: progress and puzzles in disease pathogenesis. Nat Rev Genet 11(4):247–258. https://doi.org/10.1038/nrg2748
Lam JK, Chow MY, Zhang Y, Leung SW (2015) siRNA versus miRNA as therapeutics for gene silencing. Mol Therapy Nucleic Acids 4:e252. https://doi.org/10.1038/mtna.2015.23
Lambertsen KL, Soares CB, Gaist D, Nielsen HH (2020) Neurofilaments: The C-reactive protein of neurology. Brain Sci. https://doi.org/10.3390/brainsci10010056
Lee JE, Bennett CF, Cooper TA (2012) RNase H-mediated degradation of toxic RNA in myotonic dystrophy type 1. Proc Natl Acad Sci USA 109(11):4221–4226. https://doi.org/10.1073/pnas.1117019109
Lee BH, Collins E, Lewis L, Guntrum D, Eichinger K, Voter K, Abdel-Hamid HZ, Ciafaloni E (2019) Combination therapy with nusinersen and AVXS-101 in SMA type 1. Neurology 93(14):640–641. https://doi.org/10.1212/WNL.0000000000008207
Lo Scrudato M, Poulard K, Sourd C, Tome S, Klein AF, Corre G, Huguet A, Furling D, Gourdon G, Buj-Bello A (2019) Genome editing of expanded CTG repeats within the human DMPK gene reduces nuclear RNA foci in the muscle of DM1 mice. Mol Therapy 27(8):1372–1388. https://doi.org/10.1016/j.ymthe.2019.05.021
Loureiro JR, Oliveira CL, Silveira I (2016) Unstable repeat expansions in neurodegenerative diseases: nucleocytoplasmic transport emerges on the scene. Neurobiol Aging 39:174–183. https://doi.org/10.1016/j.neurobiolaging.2015.12.007
Mahadevan M, Tsilfidis C, Sabourin L, Shutler G, Amemiya C, Jansen G, Neville C, Narang M, Barcelo J, O’Hoy K et al (1992) Myotonic dystrophy mutation: an unstable CTG repeat in the 3’ untranslated region of the gene. Science (new York, NY) 255(5049):1253–1255. https://doi.org/10.1126/science.1546325
Malik I, Kelley CP, Wang ET, Todd PK (2021) Molecular mechanisms underlying nucleotide repeat expansion disorders. Nat Rev Mol Cell Biol 22(9):589–607. https://doi.org/10.1038/s41580-021-00382-6
Marsh AP (2019) Molecular mechanisms of proteinopathies across neurodegenerative disease: a review. Neurolog Res Practice 1:35. https://doi.org/10.1186/s42466-019-0039-8
Marti E (2016) RNA toxicity induced by expanded CAG repeats in Huntington’s disease. Brain Pathol 26(6):779–786. https://doi.org/10.1111/bpa.12427
McKhann GM, Knopman DS, Chertkow H, Hyman BT, Jack CR Jr, Kawas CH, Klunk WE, Koroshetz WJ, Manly JJ, Mayeux R, Mohs RC, Morris JC, Rossor MN, Scheltens P, Carrillo MC, Thies B, Weintraub S, Phelps CH (2011) The diagnosis of dementia due to Alzheimer’s disease: recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimer’s Dementia 7(3):263–269. https://doi.org/10.1016/j.jalz.2011.03.005
Miller J, Arrasate M, Brooks E, Libeu CP, Legleiter J, Hatters D, Curtis J, Cheung K, Krishnan P, Mitra S, Widjaja K, Shaby BA, Lotz GP, Newhouse Y, Mitchell EJ, Osmand A, Gray M, Thulasiramin V, Saudou F, Segal M, Yang XW, Masliah E, Thompson LM, Muchowski PJ, Weisgraber KH, Finkbeiner S (2011) Identifying polyglutamine protein species in situ that best predict neurodegeneration. Nat Chem Biol 7(12):925–934. https://doi.org/10.1038/nchembio.694
Mirkin SM (2007) Expandable DNA repeats and human disease. Nature 447(7147):932–940. https://doi.org/10.1038/nature05977
Moumne L, Dipietromaria A, Batista F, Kocer A, Fellous M, Pailhoux E, Veitia RA (2008) Differential aggregation and functional impairment induced by polyalanine expansions in FOXL2, a transcription factor involved in cranio-facial and ovarian development. Hum Mol Genet 17(7):1010–1019. https://doi.org/10.1093/hmg/ddm373
Nakanishi K (2016) Anatomy of RISC: how do small RNAs and chaperones activate Argonaute proteins? Wiley Interdisciplin Rev RNA 7(5):637–660. https://doi.org/10.1002/wrna.1356
Nguyen L, Cleary JD, Ranum LPW (2019) Repeat-Associated Non-ATG Translation: Molecular Mechanisms and Contribution to Neurological Disease. Annu Rev Neurosci 42:227–247. https://doi.org/10.1146/annurev-neuro-070918-050405
Noor A, Zafar S, Zerr I (2021) Neurodegenerative Proteinopathies in the Proteoform Spectrum-Tools and Challenges. Int J Mol Sci. https://doi.org/10.3390/ijms22031085
Paulson H (2018) Repeat expansion diseases. Handb Clin Neurol 147:105–123. https://doi.org/10.1016/B978-0-444-63233-3.00009-9
Pearson CE, Nichol Edamura K, Cleary JD (2005) Repeat instability: mechanisms of dynamic mutations. Nat Rev Genet 6(10):729–742. https://doi.org/10.1038/nrg1689
Pesovic J, Peric S, Brkusanin M, Brajuskovic G, Rakocevic-Stojanovic V, Savic-Pavicevic D (2018) Repeat interruptions modify age at onset in myotonic dystrophy type 1 by stabilizing DMPK expansions in somatic cells. Front Genet 9:601. https://doi.org/10.3389/fgene.2018.00601
Pfister AS (2019) Emerging role of the nucleolar stress response in autophagy. Front Cell Neurosci 13:156. https://doi.org/10.3389/fncel.2019.00156
Pinto BS, Saxena T, Oliveira R, Mendez-Gomez HR, Cleary JD, Denes LT, McConnell O, Arboleda J, Xia G, Swanson MS, Wang ET (2017) Impeding transcription of expanded microsatellite repeats by deactivated Cas9. Mol Cell 68(3):479-490 e475. https://doi.org/10.1016/j.molcel.2017.09.033
Querido E, Gallardo F, Beaudoin M, Menard C, Chartrand P (2011) Stochastic and reversible aggregation of mRNA with expanded CUG-triplet repeats. J Cell Sci 124(Pt 10):1703–1714. https://doi.org/10.1242/jcs.073270
Rascovsky K, Hodges JR, Knopman D, Mendez MF, Kramer JH, Neuhaus J, van Swieten JC, Seelaar H, Dopper EG, Onyike CU, Hillis AE, Josephs KA, Boeve BF, Kertesz A, Seeley WW, Rankin KP, Johnson JK, Gorno-Tempini ML, Rosen H, Prioleau-Latham CE, Lee A, Kipps CM, Lillo P, Piguet O, Rohrer JD, Rossor MN, Warren JD, Fox NC, Galasko D, Salmon DP, Black SE, Mesulam M, Weintraub S, Dickerson BC, Diehl-Schmid J, Pasquier F, Deramecourt V, Lebert F, Pijnenburg Y, Chow TW, Manes F, Grafman J, Cappa SF, Freedman M, Grossman M, Miller BL (2011) Sensitivity of revised diagnostic criteria for the behavioural variant of frontotemporal dementia. Brain 134(Pt 9):2456–2477. https://doi.org/10.1093/brain/awr179
Rinaldi C, Wood MJA (2018) Antisense oligonucleotides: the next frontier for treatment of neurological disorders. Nat Rev Neurol 14(1):9–21. https://doi.org/10.1038/nrneurol.2017.148
Rudnicki DD, Margolis RL (2003) Repeat expansion and autosomal dominant neurodegenerative disorders: consensus and controversy. Expert Rev Mol Med 5(21):1–24. https://doi.org/10.1017/S1462399403006598
Sarisozen C, Salzano G, Torchilin VP (2015) Recent advances in siRNA delivery. Biomol Concepts 6(5–6):321–341. https://doi.org/10.1515/bmc-2015-0019
Schmitz A, Pinheiro Marques J, Oertig I, Maharjan N, Saxena S (2021) Emerging Perspectives on Dipeptide Repeat Proteins in C9ORF72 ALS/FTD. Front Cell Neurosci 15:637548. https://doi.org/10.3389/fncel.2021.637548
Schwartz JL, Jones KL, Yeo GW (2021) Repeat RNA expansion disorders of the nervous system: post-transcriptional mechanisms and therapeutic strategies. Crit Rev Biochem Mol Biol 56(1):31–53. https://doi.org/10.1080/10409238.2020.1841726
Stoyas CA, La Spada AR (2018) The CAG-polyglutamine repeat diseases: a clinical, molecular, genetic, and pathophysiologic nosology. Handb Clin Neurol 147:143–170. https://doi.org/10.1016/b978-0-444-63233-3.00011-7
Sudhakar V, Richardson RM (2019) Gene Therapy for Neurodegenerative Diseases. Neurotherapeutics 16(1):166–175. https://doi.org/10.1007/s13311-018-00694-0
Sun J, Roy S (2021) Gene-based therapies for neurodegenerative diseases. Nat Neurosci 24(3):297–311. https://doi.org/10.1038/s41593-020-00778-1
Swinnen B, Robberecht W, Van Den Bosch L (2020) RNA toxicity in non-coding repeat expansion disorders. EMBO J 39(1):e101112. https://doi.org/10.15252/embj.2018101112
Tabrizi SJ, Flower MD, Ross CA, Wild EJ (2020) Huntington disease: new insights into molecular pathogenesis and therapeutic opportunities. Nat Rev Neurol 16(10):529–546. https://doi.org/10.1038/s41582-020-0389-4
Takahashi T, Kikuchi S, Katada S, Nagai Y, Nishizawa M, Onodera O (2008) Soluble polyglutamine oligomers formed prior to inclusion body formation are cytotoxic. Hum Mol Genet 17(3):345–356. https://doi.org/10.1093/hmg/ddm311
Todd PK, Oh SY, Krans A, He F, Sellier C, Frazer M, Renoux AJ, Chen KC, Scaglione KM, Basrur V, Elenitoba-Johnson K, Vonsattel JP, Louis ED, Sutton MA, Taylor JP, Mills RE, Charlet-Berguerand N, Paulson HL (2013) CGG repeat-associated translation mediates neurodegeneration in fragile X tremor ataxia syndrome. Neuron 78(3):440–455. https://doi.org/10.1016/j.neuron.2013.03.026
Valencia-Sanchez MA, Liu J, Hannon GJ, Parker R (2006) Control of translation and mRNA degradation by miRNAs and siRNAs. Genes Dev 20(5):515–524. https://doi.org/10.1101/gad.1399806
Verkerk AJ, Pieretti M, Sutcliffe JS, Fu YH, Kuhl DP, Pizzuti A, Reiner O, Richards S, Victoria MF, Zhang FP et al (1991) Identification of a gene (FMR-1) containing a CGG repeat coincident with a breakpoint cluster region exhibiting length variation in fragile X syndrome. Cell 65(5):905–914. https://doi.org/10.1016/0092-8674(91)90397-h
Watson N, Brandel JP, Green A, Hermann P, Ladogana A, Lindsay T, Mackenzie J, Pocchiari M, Smith C, Zerr I, Pal S (2021) The importance of ongoing international surveillance for Creutzfeldt-Jakob disease. Nat Rev Neurol 17(6):362–379. https://doi.org/10.1038/s41582-021-00488-7
Wellington CL, Ellerby LM, Gutekunst CA, Rogers D, Warby S, Graham RK, Loubser O, van Raamsdonk J, Singaraja R, Yang YZ, Gafni J, Bredesen D, Hersch SM, Leavitt BR, Roy S, Nicholson DW, Hayden MR (2002) Caspase cleavage of mutant huntingtin precedes neurodegeneration in Huntington’s disease. J Neurosci 22(18):7862–7872. https://doi.org/10.1523/jneurosci.22-18-07862.2002
Wen X, Tan W, Westergard T, Krishnamurthy K, Markandaiah SS, Shi Y, Lin S, Shneider NA, Monaghan J, Pandey UB, Pasinelli P, Ichida JK, Trotti D (2014) Antisense proline-arginine RAN dipeptides linked to C9ORF72-ALS/FTD form toxic nuclear aggregates that initiate in vitro and in vivo neuronal death. Neuron 84(6):1213–1225. https://doi.org/10.1016/j.neuron.2014.12.010
Wetzel R (2012) Physical Chemistry of Polyglutamine: Intriguing Tales of a Monotonous Sequence. J Mol Biol 421(4):466–490. https://doi.org/10.1016/j.jmb.2012.01.030
Wild EJ, Boggio R, Langbehn D, Robertson N, Haider S, Miller JR, Zetterberg H, Leavitt BR, Kuhn R, Tabrizi SJ, Macdonald D, Weiss A (2015) Quantification of mutant huntingtin protein in cerebrospinal fluid from Huntington’s disease patients. J Clin Investig 125(5):1979–1986. https://doi.org/10.1172/JCI80743
Wong BKY, Ehrnhoefer DE, Graham RK, Martin DDO, Ladha S, Uribe V, Stanek LM, Franciosi S, Qiu X, Deng Y, Kovalik V, Zhang W, Pouladi MA, Shihabuddin LS, Hayden MR (2015) Partial rescue of some features of Huntington Disease in the genetic absence of caspase-6 in YAC128 mice. Neurobiol Dis 76:24–36. https://doi.org/10.1016/j.nbd.2014.12.030
Xia H, Mao Q, Eliason SL, Harper SQ, Martins IH, Orr HT, Paulson HL, Yang L, Kotin RM, Davidson BL (2004) RNAi suppresses polyglutamine-induced neurodegeneration in a model of spinocerebellar ataxia. Nat Med 10(8):816–820. https://doi.org/10.1038/nm1076
Yuan A, Nixon RA (2021) Neurofilament Proteins as Biomarkers to Monitor Neurological Diseases and the Efficacy of Therapies. Front Neurosci 15:689938. https://doi.org/10.3389/fnins.2021.689938
Zu T, Gibbens B, Doty NS, Gomes-Pereira M, Huguet A, Stone MD, Margolis J, Peterson M, Markowski TW, Ingram MA, Nan Z, Forster C, Low WC, Schoser B, Somia NV, Clark HB, Schmechel S, Bitterman PB, Gourdon G, Swanson MS, Moseley M, Ranum LP (2011) Non-ATG-initiated translation directed by microsatellite expansions. Proc Natl Acad Sci USA 108(1):260–265. https://doi.org/10.1073/pnas.1013343108
Zu T, Cleary JD, Liu Y, Banez-Coronel M, Bubenik JL, Ayhan F, Ashizawa T, Xia G, Clark HB, Yachnis AT, Swanson MS, Ranum LPW (2017) RAN translation regulated by muscleblind proteins in myotonic dystrophy type 2. Neuron 95(6):1292–1305. https://doi.org/10.1016/j.neuron.2017.08.039
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
AF and IQ authors drafted the work or revised it critically for important intellectual content and approved the submitted version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest:
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Fourier, A., Quadrio, I. Proteinopathies associated to repeat expansion disorders. J Neural Transm 129, 173–185 (2022). https://doi.org/10.1007/s00702-021-02454-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00702-021-02454-5