Abstract
The authors describe an aptasensor based on time-resolved fluorescence resonance energy transfer (TR-FRET) for the identification of Salmonella typhimurium (S. typhimurium). Aptamer-functionalized nanoparticles (NPs) is used as the energy donor, and a complementary oligonucleotide (cDNA) labeled with carboxyfluorescein (FAM) acts as the acceptor. The detection scheme is based on the hybridization between aptamer and cDNA, upon which photonic energy is transferred from NPs to FAM unless aptamer interacts with S. typhimurium. Due to the highly specific recognition ability of aptamer and the strong fluorescence intensity of NPs, the method is highly sensitive and selective for S. typhimurium. Under the optimal conditions, at excitation wavelength of 273 nm, a delay time of 100 μs and a gating time of 1 ms, the integrated time-resolved fluorescence intensity ratio (FAM520/Tb489) is linear in the 100 to 106 cfu·mL−1 range, and the limit of detection is as low as 25 cfu·mL−1. The assay was applied to the analysis of eggs and chicken meat for S. typhimurium, and the results were consistent with those of a plate-counting method.

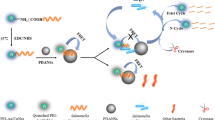
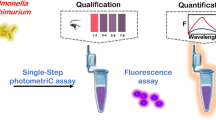
Schematic of a homogeneous time-resolved FRET assay for the determination of Salmonella typhimurium by FRET between NaYF4:Ce/Tb nanoparticle-tagged aptamer and FAM-cDNA, with a good linearity and the limit of detection of 25 cfu mL−1.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Salmonella typhimurium (S. typhimurium) is one of the leading causes of food-borne illness in human and animal hosts worldwide [1]. In United Stated almost million cases of salmonellosis with 19,000 hospitalizations and up to 300 deaths were caused by S. typhimurium each year [2]. In China, approximately 70 ~ 80% of food-borne bacteria outbreaks are thought to be caused by Salmonella and 90% of Salmonella was transmitted through the consumption of raw or uncooked vegetables, poultry, eggs, and fruits [3, 4]. Therefore, it is of great importance to develop new techniques with faster response time, better sensitivity and selectivity for S. typhimurium detection.
The existing methods for detecting S. typhimurium include conventional plate counting method, polymerase chain reaction (PCR) [5] and immunoassay methods [6]. Though the results of conventional method as well as PCR based method are accurate and reliable, the preparation work (such as pre-enrichment, selective enrichment, PCR amplification and/or cell culturing) is laborious, time consuming and unable to meet the rapid detection needs [7]. The immunoassay methods have become a powerful tool for biological research and clinical diagnostics, due to the convenient operation and the ability to test a large number of samples at the same time [8, 9]. However, these methods are heavily reliant on the quality of the antibodies, which are costly and susceptible to stability or modification issues.
To rival antibodies in these ways, aptamers with high affinity and selectivity are beginning to emerge and used for therapeutic and diagnostic applications. Aptamers are DNA or RNA molecules which can adopt specific three-dimensional conformations to combine with target analytes [10, 11]. Aptamers provide a variety of advantages over antibodies. For example, aptamers are stable, inexpensive, flexibly to chemically modify, and minimally immunogenic [12, 13]. In addition, aptamers can be modified with a variety of signal tags and designed as fluorescent [14], electrochemical [15], SERS [16] and colorimetric [17] sensors for a wide range of targets [18].
Fluorescence resonance energy transfer (FRET) is a typical and widely used homogeneous assay, due to its convenient experimental procedure [19, 20]. However, conventional FRET assays is severely compromised by autofluorescence interference, which limits practical application of the FRET technique. Time-resolved fluorescence resonance energy transfer (TR-FRET), which employing the long fluorescence emission lifetimes of lanthanide ion (Ln3+) compounds, can effectively reduce the background interference from ubiquitous endogenous fluorescent components [21, 22] and thus offers a signal with remarkably high signal-to-noise ratio in fluorescence biodetection as compared to conventional FRET [23, 24]. Ln3+-chelates as biolabels has been well developed and commercialized for decades [25]. However, most Ln3+-chelates is susceptible to photobleaching, have low labeling ratio and high cost, which strongly restricts the sensitivity of the assay and its widespread applications. As an alternative to Ln3+ chelates, Ln3+-doped inorganic nanoparticles possesses a series of advantages, such as high photochemical stability, excellent flexibility for bioconjugation, low cytotoxicity and low cost, thus can be used as a new fluorescence nano-bioprobes [26].
Herein, we introduce a TR-FRET method for rapid and sensitive detection of S. typhimurium using aptamer as a specific recognition element, time-resolved inorganic nanoparticles (NPs) as energy donor and fluorescent dye as acceptor. This method demonstrates the novelty use of long lifetime and strong fluorescence intensity of NPs and unique FRET between NPs and fluorescent dye for the quantitative analysis of pathogenic bacteria. In combination with the use of aptamer as recognition elements, this method shows high sensitivity, good selectivity and short detection time.
Materials and methods
Reagents
Tb(NO3)3·5H2O, Ce(NO3)3·6H2O and Y(NO3)3·6H2O were purchased from Aladdin Chemistry Co.Ltd. (China) (http://www.aladdin-e.com). 2-aminoethyl dihydrogen phosphate (AEP) was purchased from TCI (Shanghai) Development Co., Ltd. (http://www.tcichemicals.com). Ethylene glycol, ethanol, glutaraldehyde aqueous solution (25% in V/V), NaCl and the chemicals used to prepare the buffers and solutions were obtained from Sinopahrm chemical Reagent Co. Ltd. (China) (http://www.sinoreagent.com). Avidin was purchased from Sigma-aldrich Co. LLC (USA) (http://www.sigmaaldrich.com). The opaque 96-well microtitration microplates (300 mL/well) were purchased from Corning Inc. (USA) (http://www.corning.com).
The S. typhimurium aptamer was reported by Raghavendra Joshi et al. [27]. The DNA sequence of S. Typhimuriu aptamer is 5′-biotin-C6-TAT GGC GGC GTC ACC CGA CGG GGA CTT GAC ATT ATG ACA G-3′. Carboxyfluorescein (FAM) was used as fluorescent dye, and the complementary DNA (cDNA) sequence is 5′-GAC GCC GCC ATA-FAM-3′. The biotin labeled aptamer and FAM labeled cDNA were synthesized by the Shanghai Sangon Biological Science & Technology Company (Shanghai, China) (http://www.sangon.com).
Apparatus
The time-resolved fluorescence (TRFL) spectra was carried out on a multi-mode microplate reader with hybrid technology (Synergy H1, BioTek) upon excitation at 273 nm, where the delay time and gate time were set to be 100 μs and 1 ms, respectively. Power X-ray diffraction (XRD) pattern was collected using a D8-advance (Bruker AXS Ltd., Germany) with graphite-monochromatized Cu Kα radiation (λ = 0.15406 nm). TEM measurement was performed using a JEOL model 2100 HR instrument and operating at 200 kV accelerating voltage (TEM, JEOL Ltd., Japan). The infrared spectra of the bionanoparticles was collected using a Nicolet Nexus 470 Fourier transform infrared spectrophotometer (FTIR, Thermo Electron Co., U.S.A.) using the KBr method. Imaging of the aptamer-NPs (apt-NPs) connected with FAM-cDNA to construct NPs-FAM pair was performed with a ZEISS LSM 710 confocal microscope (Carl ZEISS, Germany). Ultraviolet-visible absorption spectra were recorded on a UV-1800 spectrophotometer (Shimadzu Co., Japan). The transient decay of the NPs was measured by employing an Edinburgh Instrument FLS920 spectrofluorometer equipped with both continuous (450 W) xenon and pulsed xenon lamp. The suspension of nanoparticles was prepared using an ultrasonic bath KJ-300 (Wuxi Kejie Electron Instruments Co. Ltd., China).
Synthesis of NaYF4: Ce/Tb nanoparticles
NaYF4: Ce/Tb was synthesized according to Tu’s and our previously reported procedure with minor alteration [28, 29]. Briefly, AEP (1 mmol) and NaCl (1 mmol) were dissolved in 30 mL ethylene glycol, then Y(NO3)3·6H2O (0.9 mmol), Ce(NO3)3·6H2O (0.05 mmol) and Tb(NO3)3·6H2O (0.05 mmol) were added under magnetic stirring to form a homogeneous solution. Subsequently, 10 mL ethylene glycol solution containing NH4F (4 mmol) was added dropwise. The mixed solution was stirred for 30 min and was transferred to a 50 mL Teflon-lined autoclave and sealed and subsequently treated solvothermally at 180 °C for 4 h. After cooling to room temperature (RT), the products were collected by centrifugation, and washed with ethanol and water three times.
Preparation of signal probes
The procedure for preparation of avidin-modified NPs was adapted from the classical glutaraldehyde method according to our previous work [29]. In brief, 1 mg of NaYF4: Ce/Tb NPs was first dissolved in 1 mL of phosphate buffered saline (PBS 137 mmol mL−1 NaCl, 2.7 mmol mL−1 KCl, 10 mmol mL−1 Na2HPO4, and 2 mmol mL−1 KH2PO4, pH 7.4) by ultrasonication for 15 min, and 250 μL of glutaradehyde solution was introduced to the mixture under gentle agitation for 3 h at 25 °C, the NPs was separated by centrifugation and were washed three times with PBS.
The 5′-biotin-labeled aptamer and avidin-functionalized NPs were bound via the high-affinity biotin-avidin system. Generally, the amount of the required 5′-biotin-labeled aptamer was introduced to the avidin-functionalized NPs solution and incubated for 6 h at 37 °C. The final products were washed with PBS three times, and redispersed in fresh PBS buffer.
Procedures for detection of S. Typhimurium
To each well, 100 μL of aptamer functionalized NPs (apt-NPs) dispersed in PBS solution was added, followed by the addition of the optimized FAM-cDNA. The plate was incubated for 40 min at 37 °C to form the NPs-FAM complex. Then various concentrations of S. typhimurium standard solutions were added to the complex (the final volume of each well is 200 μL) and future incubated at 37 °C for 50 min. Finally the plate was subjected to the measurement of TR-FRET spectra on a microplate reader (Synergy H1, BioTeK) upon excitation at 273 nm, where the delay time and gate time were set to be 100 μs and 1 ms, respectively.
Results and discussion
Detection principle
The principle of TR-FRET bioassay of S. typhimurium is illustrated in Scheme 1. In brief, avidin modified NPs is linked to biotin-labeled S. typhimurium aptamer, through the biotin-avidin affinity reaction. Complementary oligonucleotide (cDNA), which is labeled with FAM, is selected as energy acceptor, in view of its broad excitation peak centered at 495 nm matches well with the emission band of Tb3+ centered at 489 nm (Fig. 1). Through the hybridization between the aptamer and cDNA, the TR-FRET pair is constructed, where the excitation energy is transferred from the NPs to FAM. The energy transfer from the NPs donor would apparently lengthen the fluorescence lifetime of the acceptor, and the lengthened long-lived fluorescence can be temporally separated from their naturally short-lived fluorescence co-excited under UV excitation.
Since the nonradiative FRET relies heavily on the distance between the donor and the acceptor, in the absence of S. typhimurium, the distance between the donor (NPs) and the acceptor (FAM) is in close proximity, the excited state energy from the donor can be transferred, which leads to a reduction in the donor’s fluorescence intensity (here after referred to as Tb489) and an increase in the acceptor’s emission intensity (here after referred to as FAM520). In the presence of S. typhimurium, aptamer preferentially bound to S. typhimurium causes the dissociation of some cDNA, thereby liberating some FAM-cDNA, leading to a decreased fluorescence intensity of FAM520 and increased fluorescence intensity of Tb489. The concentration of S. typhimurium can be quantified by the ratio of the integrated TRFL intensities of FAM and Tb3+, as denoted by FAM520/Tb489. The TR-FRET detection based on Ln3+-doped inorganic NPs brings together the advantage of background-free signal of the TR technique and the separation-free convenience of homogeneous assay from FRET.
Characterizations of nanoparticles
In this work, amine-functionalized NaYF4: Ce/Tb NPs was synthesized by one-step solvothermal method with AEP as a surfactant and capping agent. Transmission electron microscopy (TEM) image of the NPs (Fig. 2) demonstrates that the NPs is monodisperse, uniform, roughly spherical with a diameter of approximately 30 nm. The cubic structure and phase purity of the NPs was characterized by powder XRD diffraction (Fig. S2). The elemental composition of the NaYF4: Ce/Tb NPs was determined by X-ray spectroscopy (EDS) (Fig. S3). To explore the potential application in TR-FRET detection, we measured the fluorescence lifetimes for Tb3+ doped NaYF4 NPs. The decay curves (Fig. S4) fit well to a single-exponential function, when monitoring the characteristic emissions at 544 nm. The fluorescence lifetime is determined to be 4.71 ms for the NPs samples.
Characterizations of nanoparticles conjugated to aptamer
UV-vis and FT-IR were used to characterize the conjugation of NPs with avidin and aptamer (data shown in Fig. S5, S6, S7 in the supporting information). The strong absorbance of avidin before conjugation to NPs can be seen at 280 nm (Fig. S5), and the absorbance of aptamer is at 260 nm (Fig. S7). After incubation, the absorbance of avidin and aptamer in supernatant became weaker at 280 nm (Fig. S5) and 260 nm (Fig. S7), which prove the successful preparation of aptamer bioprobes using an avidin-biotin system.
Control experiments
The TR-FRET relies heavily on the distance between NPs and FAM. In control experiments in which non aptamer labeled NPs was used in place of apt-NPs, under otherwise identical conditions, no FRET occurred (Fig. S8). The hybridization between aptamer and cDNA was directly conformed by confocal laser scanning microscopy. After apt-NPs incubated with FAM-cDNA for 40 min, the NPs-FAM pair was constructed. Apt-NPs shows bright green fluorescence after incubation with FAM-cDNA (Fig. S9).
Optimization of the method
The following parameters were optimized: (a) Concentration of FAM-cDNA; (b) Hybridization time between apt-NPs and FAM-cDNA; (c) Incubation time on FRET between apt-NPs and FAM after adding S. typhimurium. Respective data and figures are given in the Electronic Supplementary Material. The following experimental conditions were found to give best results: (a) FAM-cDNA concentration of 400 nM (Fig. S10); (b) Hybridization time of 40 min between apt-NPs and FAM-cDNA (Fig. S11); (c) Incubation time of 50 min after introducing S. typhimurium into the system (Fig. S12).
TR-FRET analysis of S. Typhimurium
We developed a TR-FRET biosensing platform for bacteria detection. Determination of S. typhimurium was performed in PBS buffer under optimal conditions. The integrated TRFL intensity ratio FAM520/Tb489 from the observed TRFL spectra (Fig. 3a) is used to quantify the concentration of S. typhimurium. As shown in Fig. 3a, FAM520/Tb489 is maximum without S. typhimurium, in the presence of S. typhimurium, the aptamer preferentially bound to bacteria and causes the dissociation of some FAM-cDNA from apt-NPs with a gradual decrease of the TR-FRET signal (FAM520/Tb489). Under optimal conditions, good linear relationship (y = 0.796–0.096×, R2 = 0.9932) between FAM520/Tb489 and the concentration of S. typhimurium is observed from 102 and 106 cfu mL−1 with a limit detection (LOD) of 25 cfu mL−1 (Fig. 3b).
For comparative purposes, the linear ranges and LODs of several aptasensor for detecting S. typhimurium are summarized in Table 1. This method had a relatively low LOD compared with other methods. This benefited from the long fluorescence lifetime of NPs, which greatly improved the signal-to-background ratio. Moreover, comparing this novel method with our previous work [29], which needs magnetic separation, several steps of washing and reagent incubations, this homogeneous TR-FRET assay is more convenient and only required inexpensive equipment. It is therefore better than most of current existing methods.
Specificity assay
To assess the specificity of the TR-FRET-based aptasensor for S. typhimurium, the influences of other pathogetic bacteria including Staphylococcus aureus, E.coli, Bacillus cereus and Listeria monocytogenes were examined under the same conditions, and the concentrations of all bacteria were 105 cfu mL−1. As shown in Fig. 4, the result indicates that only S. typhimurium caused obvious change in the TR-FRET signal (FAM520/Tb489), however, the coexisting species do not exhibit any significant changes and are analogous to the TR-FRET signals of the blank. This designed aptamer biosensor using TR-FRET strategy is highly selective because the target bacteria only perfectly recognize and bind with specific NPs-labeled aptamer, whereas other coexisting species show weak binding with aptamer. The result clearly demonstrates that the designed TR-FRET based aptamer biosensor had good specificity for the detection of S. typhimurium.
Application of the new method in chicken meat and egg sample
To demonstrate the feasibility of the practical application of this method, we detected S. typhimurium in chicken meat and egg respectively. The pre-treated samples were spiked with between 102 and 104 cfu mL−1 S. typhimurium, and then analyzed. As shown in Table 2, the results obtained by this method are close to those obtained by the plate counting method, and no significant differences between the compared methods are observed. The application performance indicates that the TR-FRET assay with aptamer has the ability to efficiently detect and quantify bacteria in real samples.
Conclusions
We have developed an aptasensor via TR-FRET between NPs and fluorescent dye for rapid, sensitive and specific detection of S. typhimurium. The application of time-resolved fluorescence signal effectively removes the autofluorescence noise of biomolecules and debris (impurities) in samples. The use of the FRET system provides an efficient method for S. typhimurium detection in one single step. In addition, the aptamer exploited in our approach is easily available and more stable than commonly used antibodies, also the highly specific to the target bacteria demonstrated the potential to detect other bacteria by substituting suitable aptamers. We envision that the methods and principles presented here can be potentially used in pathogenic bacteria detection in food samples.
References
Olaimat AN, Holley RA (2012) Factors influencing the microbial safety of fresh produce: a review. Food Microbiol 32(1):1–19
Voetsch AC, Tauxe RV (2004) FoodNet estimate of the burden of illness caused by nontyphoidal Salmonella infections in the United States. Clin infect dis 38 (Supplement_3):127-134
Barnett C, Bell C, Vig K, Akpovo AC, Johnson L, Pillai S, Singh S (2011) Development of a LIBS assay for the detection of Salmonella enterica serovar Typhimurium from food. Anal Bioanal Chem 400(10):3323–3330
Yang BW, Qu D, Zhang XL, Shen JL, Cui SH, Shi Y, Xi ML, Sheng M, Zhi SA, Meng JH (2010) Prevalence and characterization of Salmonella serovars in retail meats of marketplace in Shaanxi, China. Int J Food Microbiol 141(1–2):63–72
Jung SJ, Kim HJ, Kim HY (2005) Quantitative detection of Salmonella typhimurium contamination in milk, using real-time PCR. J Microbiol Biotechnol 15(6):1353–1358
Preechakasedkit P, Pinwattana K, Dungchai W, Siangproh W, Chaicumpa W, Tongtawe P, Chailapakul O (2012) Development of a one-step immunochromatographic strip test using gold nanoparticles for the rapid detection of Salmonella typhi in human serum. Biosens Bioelectron 31(1):562–566
Hadjinicolaou AV, Demetriou VL, Emmanuel MA, Kakoyiannis CK, Kostrikis GL (2009) Molecular beacon-based real-time PCR detection of primary isolates of Salmonella typhimurium and Salmonella Enteritidis in environmental and clinical samples. BMC Microbiol 9(1):1–14
Wiuff C, Jauho ES, Stryhn H, Andresen LO, Thaulov K, Boas U, Jakobsen MH, Heegaard PM (2000) Evaluation of a novel enzyme-linked immunosorbent assay for detection of antibodies against Salmonella, employing a stable coating of lipopolysaccharide-derived antigens covalently attached to polystyrene microwells. J Vet Diagn Investig 12(2):130–135
Lee HA, Wyatt GM, Bramham S, Morgan MR (1990) Enzyme-linked immunosorbent assay for Salmonella typhimurium in food: feasibility of 1-day Salmonella detection. Appl Environ Microbiol 56(6):1541–1546
Ellington AD, Szostak JW (1992) Selection in vitro of single-stranded DNA molecules that fold into specific ligand-binding structures. Nature 355:850–852
Tuerk C, Gold L (1990) Systematic evolution of Ligands by exponential enrichment: RNA Ligands to bacteriophage T4 DNA Polym. Sci 249(4968):505–510
Breaker RR (1997) DNA aptamers and DNA enzymes. Curr Opin Chem Biol 1(1):26–31
Yamamoto R, Baba T, Kumar PK (2000) Molecular beacon aptamer fluoresces in the presence of tat protein of HIV-1. Genes Cells 5(5):389–396
Duan N, Wu S, Chen X, Huang Y, Xia Y, Ma X, Wang Z (2013) Selection and characterization of aptamers against Salmonella typhimurium using whole-bacterium systemic evolution of Ligands by exponential enrichment (SELEX). J Agric Food Chem 61(13):3229–3234
Ma X, Jiang Y, Fei J, Ye Y, Jie C, Wang Z (2014) An aptamer-based electrochemical biosensor for the detection of Salmonella. Journal Microbiol Meth 98(1):94–98
Zhang H, Ma X, Liu Y, Duan N, Wu S, Wang Z, Xu B (2015) Gold nanoparticles enhanced SERS aptasensor for the simultaneous detection of Salmonella typhimurium and Staphylococcus aureus. Biosens Bioelectron 74:872–877
Yuan J, Tao Z, Yu Y, Ma X, Xia Y, Wang L, Wang Z (2014) A visual detection method for Salmonella typhimurium based on aptamer recognition and nanogold labeling. Food Control 37(1):188–192
Wu JJ, Zhu YY, Xue F, Mei ZL, Yao L, Wang X, Zheng L, Liu J, Liu GD, Peng CF, Chen W (2014) Recent trends in SELEX technique and its application to food safety monitoring. Microchim Acta 181(5–6):479–491
Zhou D, Piper JD, Abell C, Klenerman D, Kang DJ, Ying L (2005) Fluorescence resonance energy transfer between a quantum dot donor and a dye acceptor attached to DNA. Chem Commun 38(38):4807–4809
Chen Z, Li G, Zhang L, Jiang J, Li Z, Peng Z, Deng L (2008) A new method for the detection of ATP using a quantum-dot-tagged aptamer. Anal Bioanal Chem 392(6):1185–1188
Hanaoka K, Kikuchi K, Kobayashi S, Nagano T (2007) Time-resolved long-lived luminescence imaging method employing luminescent lanthanide probes with a new microscopy system. J Am Chem Soc 129(44):13502–13509
Kim SH, Gunther JR, Katzenellenbogen JA (2010) Monitoring a coordinated exchange process in a four-component biological interaction system: development of a time-resolved terbium-based one-donor/three-acceptor multicolor FRET system. J Am Chem Soc 132(13):4685–4692
Cohen N, Zahavy E, Zichel R, Fisher M (2016) An internal standard approach for homogeneous TR-FRET immunoassays facilitates the detection of bacteria, biomarkers, and toxins in complex matrices. Anal Bioanal Chem 408(19):5179–5188
Ju Q, Liu Y, Tu D, Zhu H, Li R, Chen X (2011) Lanthanide-doped multicolor GdF3 nanocrystals for time-resolved Photoluminescent Biodetection. Chem 17(31):8549–8554
Bünzli JCG, Eliseeva SV (2013) Intriguing aspects of lanthanide luminescence. Chem Sci 4(5):1939–1949
Zhang H, Xu Y, Yang W, Li Q (2007) Dual-lanthanide-chelated silica nanoparticles as labels for highly sensitive time-resolved Fluorometry. Chem Mater 19(24):5875–5881
Joshi R, Janagama H, Dwivedi HP, Kumar TS, Jaykus L-A, Schefers J, Sreevatsan S (2009) Selection, characterization, and application of DNA aptamers for the capture and detection of Salmonella enterica serovars. Mol Cell Probes 23(1):20–28
Tu D, Liu L, Ju Q, Liu Y, Zhu H, Li R, Chen X (2011) Time-resolved FRET biosensor based on amine-functionalized lanthanide-doped NaYF4 nanocrystals. Angew Chem Int Ed 50(28):6306–6310
Wang X, Huang Y, Wu S, Duan N, Xu B, Wang Z (2016) Simultaneous detection of Staphylococcus aureus and Salmonella typhimurium using multicolor time-resolved fluorescence nanoparticles as labels. Int J Food Microbiol 237:172
Duan YF, Ning Y, Song Y, Deng L (2014) Fluorescent aptasensor for the determination of Salmonella typhimurium based on a graphene oxide platform. Microchim Acta 181(5):647–653
Duan N, Wu S, Dai S, Miao T, Chen J, Wang Z (2015) Simultaneous detection of pathogenic bacteria using an aptamer based biosensor and dual fluorescence resonance energy transfer from quantum dots to carbon nanoparticles. Microchim Acta 182(5):917–923
Liu K, Yan X, Mao B, Wang S, Deng L (2016) Aptamer-based detection of Salmonella enteritidis using double signal amplification by Klenow fragment and dual fluorescence. Microchim Acta 183(2):643–649
Lei P, Tang H, Ding S, Ding X, Zhu D, Shen B, Cheng Q, Yan Y (2015) Determination of the invA gene of Salmonella using surface plasmon resonance along with streptavidin aptamer amplification. Microchim Acta 182(1):289–296
Jia F, Duan N, Wu S, Dai R, Wang Z, Li X (2016) Impedimetric Salmonella aptasensor using a glassy carbon electrode modified with an electrodeposited composite consisting of reduced graphene oxide and carbon nanotubes. Microchim Acta 183(1):337–344
Acknowledgements
This work was partly supported by National Natural Science Foundation of China (21375049, 31401575, and 31401576), Key Research and Development Program of Jiangsu Province BE2016306, China Postdoctoral Science Foundation (2016 T90430) and Synergetic Innovation Center of Food Safety and quality control of Jiangsu Province.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no competing interests.
Electronic supplementary material
ESM 1
(DOCX 899 kb)
Rights and permissions
About this article
Cite this article
Wang, X., Niazi, S., Yukun, H. et al. Homogeneous time-resolved FRET assay for the detection of Salmonella typhimurium using aptamer-modified NaYF4:Ce/Tb nanoparticles and a fluorescent DNA label. Microchim Acta 184, 4021–4027 (2017). https://doi.org/10.1007/s00604-017-2399-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00604-017-2399-5