Abstract
This article describes a fluorescent molecularly imprinted polymer (MIP) capable of selective fluorescent turn-on recognition of the tumor biomarker α-fetoprotein. The technique is making use of amino-modified Mn-doped ZnS quantum dots (QDs) as solid supports, 4-vinylphenylboronic acid and methyl methacrylate as the functional monomers, γ-methacryloxypropyl trimethoxysilane as the grafting agent, and α-fetoprotein as a template. A graft imprint is created on the surface of the QDs. The functional monomers are shown to play an important role in the formation of the binding sites and in preventing nonspecific protein binding. The resulting MIP-QDs display a good linear response to α-fetoprotein in the 50 ng · L−1 to 10 μg · L−1 concentration range, and the limit of detection is 48 ng · L−1. In our perception, the method has a wide scope in that it may be adapted to various other glycoproteins.

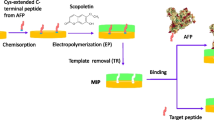
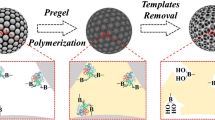
Schematic illustration of the synthesis of the MIP-QDs composites
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Glycosylation encompasses a diverse selection of sugar-moiety additions to proteins that with significant effects on protein folding, conformation, distribution, stability and activity [1]. The resultant glycoproteins play important roles in numerous biological events, and a large number of them have been served as disease biomarkers and therapeutic targets in clinical diagnostics [2, 3]. A thorough understanding of the molecular basis of disease requires the detailed analysis of the proteins being involved [4]. However, the dynamic nature of metabolism, the complexity of proteins and the low stoichiometry of protein post-translational modifications pose a serious technical challenge to specifically recognize target proteins from biological samples [5]. The enzyme-linked immuno-sorbent assay (ELISA) is the gold standard for clinical protein biomarker detection in complex biological samples. Unfortunately, antibodies are not always perfect owing to their high costs, time-consuming and instability when out of their native environment [6]. LC-MS-based proteomics have attracted considerable attention for biomarker discovery but they are currently too expensive and technically complex for routine clinical diagnostics [7]. Meanwhile, fluorescence methods show great promise for the detection of trace amounts of analytes owing to their sensitivity, simplicity, fast response, and cost-effective instrumentation [8]. Over the past few decades, there has become a remarkable growth in the use of fluorescence methods in the bioanalysis, cellular and molecular imaging. Among the various luminophores, quantum dots (QDs) have been widely employed due to their easy preparation, high luminescence efficiency, photo-stability and size dependent optical properties [9]. However, the major challenge to develop a QDs-based fluorescence probe is the development of a chemical or molecular recognition element [10, 11].
Molecular imprinting technology allows the creation of tailor-made recognition sites with a memory of the shape, size and functional groups of the template molecules in synthetic polymers. Molecularly imprinted polymers (MIPs) have shown considerable promise as substitutes for natural receptors and antibodies, and a wide range of application has been explored [12]. While molecular imprinting technology has seen use with small organic molecules, the imprinting of proteins has been and continues to be a sizable challenge. The difficulty is mainly due to their large size, structural complexity, conformations flexibility, and compatibility with solvents [13]. Hydrogen bonding plays an important role in molecular recognition of biomolecules in aqueous media. The use of water as solvent provides a biologically benign environment for proteins. However, water can reduce hydrogen bonding and electrostatic interactions between template molecules and functional monomers. Hydrogen bond-based molecular imprinting often shows non-specific recognition for small molecules and proteins in aqueous solutions. Covalent molecular imprinting with the template proteins conjugated with the functional monomers by covalent linkage may be able to provide more specific binding cavities. Boronic acids are ideal candidates for effective formation of glycoprotein recognition sites because they covalently and reversibly bind carbohydrates to form cyclic esters in an alkaline aqueous solution, and the cyclic esters dissociate when the medium is changed to acidic pH [14]. Recently, Liu and coworkers have been exploring the new field of boronate affinity-based MIP for the immunoassay of glycoproteins [15–17].
The integration of a signalling element into MIP has also attracted extensive research interest, because the resultant recognition materials are capable of transduction of binding events into readable signals [18]. Generally, there are two methods for the incorporation of fluorescent species in the imprinted polymer matrix. The first method involves the use of fluorophore-conjugated functional monomers [19, 20]. Unfortunately, syntheses of fluorescent functional monomers are time consuming, and the resultant fluorescent MIP often show high background noise due to fluorescent residues located outside the binding cavities [21]. The second method concerns the creation of a molecular imprinted layer over QDs [22–24]. However, the surface immobilization of functional groups on QDs remains complex and uncontrollable. Furthermore, the MIP-QD sensing materials often show low sensitivity because QDs were embedded into highly cross-linked MIP.
As a tumor biomarker, α-fetoprotein (AFP) is a glycoprotein with a molecular weight of approximately 70 kDa, which is secreted by fetal liver and yolk sac. The increased AFP concentration in adult plasma is usually considered as an early indication of hepatocellular or endodermal sinus tumor [25]. This paper demonstrates an alternative strategy for producing fluorescent MIP for selective fluorescence turn-on sensing of α-fetoprotein. We use amino modified Mn-doped ZnS QDs as the support, 4-vinylphenylboronic acid and methyl methacrylate as the functional monomers, γ-methacryloxypropyl trimethoxy silane as the grafting agent, and α-fetoprotein as a template to produce graft imprinting at the surface of the Mn-doped ZnS QDs. The MIP-QDs composites function as a fluorescent sensing material for specific recognition of α-fetoprotein, allowing for sensitive and selective detection of α-fetoprotein in biological samples.
Experimental section
Reagents and chemicals
All chemicals used were of analytical grade. ZnSO4 · 7H2O, Na2S · 9H2O, MnCl2 · 4H2O, were used as received from Alfa Aesar (http://www.alfachina.cn/, Tianjin, China). 3-Mercaptopropyltriethoxysilane (MPTS), γ-methacryloxypropyl trimethoxy silane (KH570), α-fetoprotein (AFP), lysozyme (Lyz), cytochrome c (Cyt), bovine serum albumin (BSA), bovine hemoglobin (BHb), horseradish peroxidase (HRP), myoglobin (Mb), ovalbumin (Ova), transferrin (TRF), and ribonuclease B (RNase B) were obtained from Sigma-Aldrich Co (http://www.sigma-aldrich.com/, St. Louis, MO). 4-Vinylphenylboronic acid (VPBA), methyl methacrylate (MMA), acrylamide (AAM), N, N, N’, N’-tetramethylethylethylenediamine (TEMED) and ammonium persulfate (APS) were purchased from Tianjian Chemical Reagents Co., Ltd (http://www.ectcr.com/, Tianjin, China). The mouse monoclonal to AFP antibody was from Jingtian Biotech Co. Ltd. (http://www.biococ.com.cn/, Shanghai, China). Other chemicals were purchased from Shanghai Chemical Reagent Company (http://www.scrri.com/, Shanghai, China). Ultrapure water from a laboratory water purification system (Sartorius, Germany) was used throughout this work.
Characterization
The morphology and microstructure of the QDs and MIP-QDs were characterized by high-resolution transmission electron microscopy (HRTEM) on a JEM-2100HR (JEOL; Japan). The X-ray diffraction (XRD) spectra were collected on a Bruker D8 Advance X-ray diffractometer (Bruker, Germany). UV–Vis absorption spectra were recorded with A Shimadzu UV–Vis 1700 Spectrophotometer. The fluorescence spectra were recorded on an F-2500 (Hitachi, Tokyo, Japan). The acidity was monitored with a pH211 microprocessor pH meter (Hanna instruments).
Synthesis of amino-capped Mn-doped ZnS QDs
The procedure for synthetically producing the amino-capped Mn-doped ZnS QDs used in this work is similar to the method previously described in literature [26]. 1.8 g of ZnSO4, 0.1 g of MnCl2, and 20 mL of ultrapure water were added to a 50 mL three-necked flask. After the mixture was stirred under nitrogen at room temperature for 10 min, 5 mL of aqueous solution containing 1.5 g of Na2S was added drop wise, and the mixture was kept stirring for 30 min. Then 5 mL of an ethanol solution containing 0.072 g of MPTS was added, and the mixture was kept stirring for 24 h. The resultant MPTS-capped Mn-doped ZnS QDs were centrifuged and washed with absolute ethanol three times before being dried under vacuum. In the second step, 10 mL of an absolute ethanol solution containing 200 μL of APTES, 0.35 mL of TEOS, and 250 mg of MPTS-capped Mn-doped ZnS QDs were added to a 25-mL flask. After the mixture was stirred for 10 min, 0.4 mL of concentrated ammonium hydroxide solution and 2.0 mL H2O were added, and the mixture was kept stirring for 18 h. The amino modified Mn-doped ZnS QDs were centrifuged and washed with ultrapure water three times then dried in vacuum at room temperature.
Preparation of MIP-QDs composites
Typically, NH2 modified Mn-doped ZnS QDs were first dispersed in 5 mL of carbonate buffer (0.01 mol · L−1, pH 9.0) by ultrasonic vibration. MMA, 4-vinylphenylboronic acid, AFP and KH570 were dissolved into the above solution. The mixing solution was purged with nitrogen for 30 min and stirred for 1 h before TEOS (0.3 mL of TEOS dissolved in 1 mL methanol) and 10 mg of APS were added into the solution. The mixing solution was purged with nitrogen and allowed to proceed under magnetic stirring for 5 h. The nonimprinted polymer (NIP) was synthesized in parallel but without the addition of the template. After the reaction, the MIP- and NIP-QDs composites were centrifuged and washed with ultrapure water to remove the unreacted monomers. Finally, the MIP-QDs composites were washed repeatedly with 0.2 mol · L−1 phosphoric acid solution containing 30 % acetonitrile (v/v) and shaken for 2 h to remove the template until no AFP in the supernatant was detected using a UV/vis spectrophotometer at 280 nm detection wavelength.
Evaluation of the adsorption characteristics of MIP-QDs
The capacity and kinetics of the rebinding molecules were investigated using UV–Vis absorption spectra. Ten milligrams of MIP-QDs composites were suspended in 5 mL of aqueous solution with different AFP concentrations. After incubation at room temperature for 12 h, the MIP-QDs composites were removed by centrifugation. The binding amount of template was calculated as the difference in concentration between the initial spike solution and the concentration of the supernatant after equilibration. The experimental data is presented as the adsorption capacity per unit mass (mg) of the nanoparticles, and the adsorption capacity (Q) calculated using \( Q=\frac{V\left({C}_0-{C}_{\mathrm{e}}\right)}{m} \), where Q (mg · g−1) is the mass of the template adsorbed by a unit mass of particles, C0 (mg mL−1) is the initial template concentration, Ce (mg · mL−1) is the template concentration of the supernatant, V (mL) is the volume of the initial solution, and m (g) is the mass of the particles.
Measurement procedure
The standard solutions of AFP for calibration were prepared by dissolved AFP with 0.01 mol · L−1 phosphate (PBS, pH 7.4) solution and stored at 4 °C. To a 5-mL calibrated test tube, 1 mL of 100 mg · L−1 MIP- or NIP-QDs solution, 1.0 mL of carbonate solution (CBS, pH 9.0, 0.01 mol · L−1), and different concentrations of AFP (1.0 mL) were sequentially added. The mixture was then diluted to volume with ultrapure water and mixed thoroughly to make it homogeneous. The response time of the MIP-QDs composites was set to be 15 min for subsequent experiments [27]. The experiments were conducted in the air conditioned room under the temperature of 25 °C.
Serum samples
The clinical serum samples were collected from volunteers of Guangzhou first people’s hospital. An appropriate dilution of serum (20-fold) was adopted before detection. The concentration of the AFP in serum samples were detected by the above method. The slit widths were 5 nm for both excitation and emission in the fluorescence mode. The excitation wavelength was set at 340 nm with a PMT voltage of 400 V.
Results and discussion
Preparation and characterization of MIP- and NIP-QDs composites
Applying molecular imprinting techniques to the surface of QDs allows the preparation of MIP with accessible, surface-exposed binding sites and excellent optical properties. Among such QDs, Mn-doped ZnS QDs have been widely employed owing to their easy preparation, low cytotoxicity, wonderful biocompatibility and high photo-luminescence efficiency. As shown in Fig. 1, the surface of the QDs was modified with amino propyls that leave enough silanol groups for further surface-initiated poly-condensation to inhibit QDs leakage during the imprinting process [28]. Typically for the protein imprinting, functional monomers including acrylic acid derivatives, siloxane derivatives, and electropolymerizable monomers have been extensively used [29]. VPBA and MMA were chosen as the functional monomers based on the consideration that the boronic acid group covalently reacts with glycoproteins which containing cis-diols to form cyclic esters in an alkaline aqueous solution, and polymethyl methacrylate (PMMA) showed excellent hydrophobic ability to suppress nonspecific adsorption. We choose TEOS as the inorganic precursor, and KH570 as the coupling agent, which was used to form the covalent bonding between organic and inorganic phases. Subsequently, the MIP is fabricated from a sol–gel hydrolysis process and a pre-polymerization reaction through covalent bonds to yield organic–inorganic phase. After removing the template molecules, recognition cavities complementary to the template molecule in shape, size, and chemical functionality were formed in the cross-linked polymer matrix. Note that the controllability and hydrophilic the MIP relate to the specific binding properties. We designed and prepared several different MIP-QDs composites to find the optimal monomer composition. As shown in Table S1, MIP-QDs possesses a high adsorption capacity and selectivity with a molar ratio of 1:1, while the other molar ratios showed low adsorption capacity or selectivity. Our explanation for this result is that the low ratios of 4-vinylphenylboronic acid (VPBA) and methyl methacrylate (MMA) induce fewer non-specific binding sites in the polymer due to fewer complexes being formed between the functional monomer and the template, while the high ratios of VPBA to MMA result in high non-specific adsorption. Considering these two aspects, the molar ratio of 1:1 of VPBA to MMA was chosen to develop the MIP-QDs composites.
High-resolution transmission electron microscopy (HRTEM) image reveal that the QDs particles are spherical in shape and almost uniform in size, with a diameter of approximately 3 nm (n = 20, relative standard deviation: 1.8 %) (Fig. 2a). Fig. 2b and c show that the process aggregates and embeds the original QDs into larger particles after surface imprinting on the QDs, and there is no significant difference in the particle diameter between MIP-QDs and NIP-QDs. The TEM image of the MIP-QDs composites indicates that the QDs shown as black dots in the image are successfully embedded in the polymer matrix. The thickness of molecularly imprinted shell was difficult to characterize by TEM due to the aggregation of the MIP-QDs during drying on a Cu grid. The X-ray diffraction (XRD) pattern of the MIP-QDs composites (Fig. 2d) exhibits a cubic structure with peaks in (111), (220), and (311).
Next, the stability of the MIP-QDs composites was evaluated at room temperature by the repeated detections of the fluorescence intensity of the MIP-QDs. As shown in Fig. S1, the fluorescence intensity was stable as confirmed by the observed 2.3 % relative standard deviation for 9 replicate measurements over more than 2 h. The main reason for the stable fluorescence emission is that the inner Mn2+ is protected by the MIP shell.
Effect of pH on detection
Prior to using the MIP-QDs composites for fluorescence detection of α-fetoprotein, we investigated the effect of pH on the response of the sensing material. The enhanced fluorescence intensity (△F) normalized to the initial fluorescence intensity (F0) was used to evaluate the enhanced efficiency of the target protein toward the fluorescence intensity of the MIP-QDs. As shown in Fig. S2, a change in pH can influence the protonation of AFP and the boric acid groups in the polymers and thus can affect the interactions between AFP and the MIP-QDs. The enhanced fluorescence signals were stable in the subsequent 30 min. The maximum enhancement percentage of fluorescence intensity occurs at a pH of 9.0, so we used a 1.0 mL CBS buffer solution (0.01 mol · L−1, pH 9.0) for further experiments to obtain stable fluorescence intensity and high sensitivity. Note that this buffer solution is also a typical condition for boronate affinity interaction.
Analytical performance of MIP-QDs composites
Typical fluorescence enhancement of the MIP- and NIP-QDs composites by 5.0 × 10−8 to 1.0 × 10−5 g · L−1 AFP was showed in Fig. 3a and b. Both the MIP- and NIP-QDs composites exhibit a spectroscopic response to the target protein. However, the fluorescence enhancement is much more noticeable with the MIP-QDs composites than with the NIP-QDs composites. The larger enhancement efficiency of the MIP-QDs composites results from its specific binding affinity with AFP.
To demonstrate that the recognition specificity of the MIP-QDs composites, five other glycoproteins including horseradish peroxidase (HRP, PI 3.0–9.0, 4.0 × 6.7 × 11.7 nm3), myoglobin (Mb, pI 7.0, 2.5 × 3.5 × 4.5 nm3), ovalbumin (Ova, pI 4.6, 7.0 × 4.5 × 5.0 nm3), transferrin (TRF, PI, 5.2–6.2, 4.5 × 5.8 × 13.6 nm3), and ribonuclease B (RNase B, PI, 8.9, 10.1 × 3.3 × 7.4 nm3) were chosen for comparison. These proteins have large differences in isoelectric point and dimension. As shown in Fig. S3, the MIP-QDs showed a very sensitive response to the target protein AFP but were almost insensitive to the other proteins. The most relevant ions in biological fluids had no effect on the detection of AFP (Table S2). The fact that the delicate binding sites of AFP molecules in the MIP-QDs have the capability to recognize AFP and the hydrophobic ability of PMMA to suppress the non-specific adsorption can explain this result.
We next investigated the recyclability of MIP-QDs, since template removal during synthesis of the MIP-QDs was achieved by solvent extraction, and regeneration of the MIP-QDs composites after rebinding was performed the same way. As shown in Fig. S4, the enhancement efficiency of MIP-QDs to AFP was almost unchanged after six cycles of adsorption and desorption. We attribute this result to the effective regeneration of molecularly imprinted cavities, which provide special recognition sites for the target protein.
Using the MIP-QDs composites for selective detection of AFP in biological fluids
As shown in Fig. 4, the fluorescence intensity was proportionally increased by increasing AFP concentration in the MIP-QDs. The fluorescence enhancement in this system follows the Stern-Volmer equation:
where F0 and F are the fluorescence intensities in the absence and presence of the target protein respectively, C q is the concentration of the target protein, and K sv is the Stern-Volmer constant [23]. The calibration plot of F/F 0 − 1 versus C q has a good linear relationship in the range of 5.0 × 10−8 g · L−1 – 1.0 × 10−5 g · L−1 with a correlation coefficient of 0.998. The limit of detection (LOD) is calculated following the 3σ IUPAC criteria (3σ/S), where σ is the standard deviation of the blank signal and S is the slope of the linear calibration. The LOD for AFP was 4.8 × 10−8 g · L−1. The precision for 11 replicate detections of 1.0 × 10−6 g · L−1 AFP was 2.4 % (RSD).
The developed MIP-QDs composites-based fluorescence method was employed to detect AFP concentrations in clinical serum samples. The accuracy of AFP determination was examined by comparing the results with those from the ELISA analysis. As shown in Table 1, the analytical results for the detection of AFP in five serum samples were in good agreement with those obtained using an ELISA method. The results demonstrate that this sensing material has great potential for the determination of AFP in clinical samples.
Several methods have been described in literature for AFP detection and their analytical performances are summered in Table 2. Li et al. reported a sensitive photoelectrochemical immunosensing method for the determination of AFP but it needs a long incubation time and fabrication of the photoelectrochemical immunosensor is complicated [33]. It can be seen that without expensive instrumentation and extensive sample preparation, a level of analytical performance has been achieved by this approach. The LOD and linear range of this method is comparable or superior to those obtained by chemiluminescence and ELISA methods.
Possible increasement mechanism of AFP on the fluorescence of MIP-QDs composites
To explain the mechanism of AFP on the fluorescence of the MIP-QDs composites, it is important to understand both the fluorescence emission mechanism of the QDs and the recognition mechanism of the MIP. The well-known orange emission of Mn-doped ZnS QDs is excited either by direct recombination of a bound exciton with Mn2+ or via trapping of the hole by Mn2+. Subsequent recombinations with shallowly trapped electrons result in Mn2+ in an excited state and are followed by the well-known phosphorescence emission (Fig. S5). In aqueous systems, boronic acids exist in equilibrium between an undissociated neutral trigonal form and a dissociated anionic tetrahedral form. In the presence of 1, 2- or 1, 3-diols, cyclic boronate esters formed by reaction of the neutral boronic acid with a diol are generally considered hydrolytically unstable [35]. As shown in Fig. S6, the quenching mechanism can be described as an electron transfer process. In the absence of AFP, the fluorescence quenching occurs due to the electron transfer between the Mn-doped ZnS QDs and the boron moieties. On the other hand, when the boron moieties of the MIP-QDs becomes negatively charged after AFP binding, the electrostatic attraction between the QDs and the boron moieties is hindered, leading to a loss of quenching efficiency and an increase in fluorescence intensity.
Conclusions
We demonstrate an alternative strategy for producing fluorescent MIP for selective fluorescence turn-on sensing of α-fetoprotein. The resultant MIP exhibit excellent ability for specific binding of the target protein owing to a synergism between the boronate affinity binding sites and the shape exclusion of the imprinted cavity. Under optimal conditions, trace levels of α-fetoprotein are signalled with high sensitivity and selectivity by emission intensity changes of the Mn-doped ZnS QDs, which is embedded into the imprinted polymers.
References
Spiro R (2002) Protein glycosylation: nature, distribution, enzymatic formation, and disease implications of glycopeptide bonds. Glycobiology 12:43R–56R
Ohtsubo K, Marth JD (2006) Glycosylation in cellular mechanisms of health and disease. Cell 126:855–867
Rudd PM, Elliott T, Cresswell P, Wilson IA, Dwek RA (2001) Glycosylation and the immune system. Science 291:2370–2376
Mishra A, Verma M (2010) Cancer biomarkers: are we ready for the prime time? Cancers 2:190–208
Ye ML, Pan YB, Cheng K, Zou HF (2014) Protein digestion priority is independent of protein abundances. Nat Methods 11:220–222
Haupt K (2010) Biomaterials: plastic antibodies. Nat Mater 9:612–614
Zhang Y, Fonslow BR, Shan B, Baek MC, Yates JR (2013) Protein analysis by shotgun/bottom-up proteomics. Chem Rev 113:2343–2394
Yuan L, Lin WY, Zheng KB, He LW, Huang WM (2013) Far-red to near infrared analyte-responsive fluorescent probes based on organic fluorophore platforms for fluorescence imaging. Chem Soc Rev 42:622–661
Gill R, Zayats M, Willner I (2008) Semiconductor quantum dots for bioanalysis. Angew Chem Int Ed 47:7602–7625
You CC, Miranda OR, Gider B, Ghosh PS, Kim IB, Erdogan B, Krovi SA, Bunz UHF, Rotello VM (2007) Detection and identification of proteins using nanoparticle-fluorescent polymer ‘chemical nose’ sensors. Nat Nanotechnol 2:318–323
Zhang X, Du X, Huang X, Lv Z (2013) Creating protein-imprinted self-assembled monolayers with multiple binding sites and biocompatible imprinted cavities. J Am Chem Soc 135:9248–9251
Chen LX, Xu SF, Li JH (2011) Recent advances in molecular imprinting technology: current status, challenges and highlighted applications. Chem Soc Rev 40:2922–2942
Ge Y, Turner APF (2008) Too large to fit? Recent developments in macromolecular imprinting. Trends Biotechnol 26:218–224
Cambre JN, Sumerlin BS (2011) Biomedical applications of boronic acid polymers. Polymer 52:4631–4643
Bi XD, Liu Z (2014) Facile preparation of glycoprotein-imprinted 96-well microplates for enzyme-linked immunosorbent assay by boronate affinity-based oriented surface imprinting. Anal Chem 86:959–966
Li L, Lu Y, Bie ZJ, Chen HY, Liu Z (2013) Photolithographic boronate affinity molecular imprinting: a general and facile approach for glycoprotein imprinting. Angew Chem Int Ed 52:7451–7454
Ye J, Chen Y, Liu Z (2014) A boronate affinity sandwich assay: an appealing alternative to immunoassays for the determination of glycoproteins. Angew Chem Int Ed 53:10386–10389
Bompart M, Wilde YD, Haupt K (2010) Chemical nanosensors based on composite molecularly imprinted polymer particles and surface-enhanced raman scattering. Adv Mater 22:2343–2348
Sunayama H, Ooya T, Takeuchi T (2010) Fluorescent protein recognition polymer thin films capable of selective signal transduction of target binding events prepared by molecular imprinting with a post-imprinting treatment. Biosens Bioelectron 26:458–462
Wan W, Biyikal M, Wagner R, Sellergren B, Rurack K (2013) Fluorescent sensory microparticles that “Light-up” consisting of a silica core and a molecularly imprinted polymer (MIP) shell. Angew Chem Int Ed 52:7023–7027
Sunayama H, Ooya T, Takeuchi T (2014) Fluorescent protein-imprinted polymers capable of signal transduction of specific binding events prepared by a site-directed two-step postimprinting modification. Chem Commun 50:1347–1349
Wang YZ, Li DY, He XW, Li WY (2015) Epitope imprinted polymer nanoparticles containing fluorescent quantum dots for specific recognition of human serum albumin. Microchim Acta 182:1465–1472
Wei X, Zhou Z, Hao T, Xu Y, Li H, Lu K, Dai J, Zheng X, Gao L, Wang J, Yan Y, Zhu Y (2015) Specific recognition and fluorescent determination of aspirin by using core-shell CdTe quantum dot-imprinted polymers. Microchim Acta 182:1527–1534
Ren X, Liu H, Chen L (2015) Fluorescent detection of chlorpyrifos using Mn (II)-doped ZnS quantum dots coated with a molecularly imprinted polymer. Microchim Acta 182:193–200
Rebischung C, Pautier P, Morice P, Lhomme C, Duvillard P (2000) Alpha-fetoprotein production by a malignant mixed Müllerian tumor of the ovary. Gynecol Oncol 77:203–205
Tan L, Huang C, Peng R, Li W, Tang Y (2014) Development of hybrid organic–inorganic surface imprinted Mn-doped ZnS QDs and their application as a sensing material for target proteins. Biosens Bioelectron 61:506–511
Tan L, Kang C, Xu S, Tang Y (2013) Selective room temperature phosphorescence sensing of target protein using Mn-doped ZnS QDs-embedded molecularly imprinted polymer. Biosens Bioelectron 48:216–223
Cumbo A, Lorber B, Corvini PF, Meier W, Shahgaldian P (2013) A synthetic nanomaterial for virus recognition produced by surface imprinting. Nat Commun 4:1503–1509
Dechtrirat D, Gajovic-Eichelmann N, Bier FF, Scheller FW (2014) Hybrid material for protein sensing based on electrosynthesized MIP on a mannose terminated self-assembled monolayer. Adv Funct Mater 24:2233–2239
Lin JH, He CY, Zhang LJ, Zhang SS (2009) Sensitive amperometric immunosensor for a-fetoprotein based on carbon nanotube/ gold nanoparticle doped chitosan film. Anal Biochem 384:130–135
Chen ZG, Lei YL, Chen X (2012) Immunoassay for serum alpha fetoprotein using silver nanoparticles and detection via resonance light scattering. Microchim Acta 179:241–248
Wu BY, Wang HF, Chen JT, Yan XP (2011) Fluorescence resonance energy transfer inhibition assay for α-fetoprotein excreted during cancer cell growth using functionalized persistent luminescence nanoparticles. J Am Chem Soc 133:686–688
Li YJ, Ma MJ, Zhu JJ (2012) Dual-signal amplification strategy for ultrasensitive photoelectrochemical immunosensing of alpha-fetoprotein. Anal Chem 84:10492–10499
Chen H, Guan Y, Wang S, Ji Y, Gong M, Wang L (2014) Turn-on detection of a cancer marker based on near-infrared luminescence energy transfer from NaYF4:Yb, Tm/NaGdF4 core-shell upconverting nanoparticles to gold nanorods. Langmuir 30:13085–13091
Gamsey S, Suri J, Wessling R, Singaram B (2006) Continuous glucose detection using boronic acid-substituted viologens in fluorescent hydrogels: linker effects and extension to fiber optics. Langmuir 22:9067–9074
Acknowledgments
We gratefully acknowledge financial support from the National Natural Science Foundation of China (No: 21505026) and Medical Science and Technology Project of Guangzhou (No: 20141A010055).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 397 kb)
Rights and permissions
About this article
Cite this article
Tan, L., Chen, K., Huang, C. et al. A fluorescent turn-on detection scheme for α-fetoprotein using quantum dots placed in a boronate-modified molecularly imprinted polymer with high affinity for glycoproteins. Microchim Acta 182, 2615–2622 (2015). https://doi.org/10.1007/s00604-015-1642-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00604-015-1642-1