Abstract
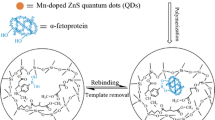
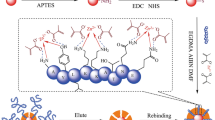
Molecularly imprinted polymer (MIP) nanofilms for alpha-fetoprotein (AFP) and the receptor binding domain (RBD) of the spike protein of SARS-CoV-2 using either a peptide (epitope-MIP) or the whole protein (protein-MIP) as the template were prepared by electropolymerization of scopoletin. Conducting atomic force microscopy revealed after template removal and electrochemical deposition of gold a larger surface density of imprinted cavities for the epitope-imprinted polymers than when using the whole protein as template. However, comparable affinities towards the respective target protein (AFP and RBD) were obtained for both types of MIPs as expressed by the KD values in the lower nanomolar range. On the other hand, while the cross reactivity of both protein-MIPs towards human serum albumin (HSA) amounts to around 50% in the saturation region, the nonspecific binding to the respective epitope-MIPs is as low as that for the non-imprinted polymer (NIP). This effect might be caused by the different sizes of the imprinted cavities. Thus, in addition to the lower costs the reduced nonspecific binding is an advantage of epitope-imprinted polymers for the recognition of proteins.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Templating is an ancient technology which was already applied by the Muiscas—the original population of South America—before the arrival of the Spanish conquistadores (1575) for the preparation of golden sculptures, e.g., the El Dorado Raft. Using the templating concept, the Russian scientist Polyakov proposed the first polymer with molecular memory in 1931 [1]. The template, functional monomers, and crosslinkers are polymerized together in these processes. Removal of the template forms template-complementary binding sites resembling the “lock and key principle.” However, a real breakthrough has been only achieved ~ 50 years later [2, 3]. Initially, imprinted polymers have been developed exclusively for low-molecular weight substances and these synthetic sorbents coined the name of molecular imprinted polymers (MIPs). Imprinting of large biological molecules and particles (e.g., viruses and cells) faces challenges due to their complex nature, fragility, solvent compatibility, high cost for pure templates, distribution of heterogeneous binding sites, and difficulties in highly reproducible mass production [4, 5].
Antibodies—the biological counterparts of MIPs—bind only to a small area of large antigens—the immunogenic determinant, so-called epitope. Mimicking this principle Rachkov and Minoura [6] applied the three amino acids on the N-terminal of the nonapeptide oxytocin as the template for a MIP which could subsequently bind the whole peptide. Later, this approach was successfully extended to proteins and cells and termed as epitope imprinting to highlight the similarity between epitope-MIPs and antibodies [6]. Such epitope-MIPs can recognize the template fragment as well as the respective whole protein [6,7,8,9,10,11,12]. A generally claimed advantage of the epitope imprinting concept is the lower cost of the template in the MIP synthesis resulting a 10- to 50-fold lower cost per sensing spot (refer to SI1 in the supporting information).
On the other hand, the differences in terms of target binding characteristics of both types of MIPs are still not well understood. Only a few reports compare the analytical performance of MIPs, which use an epitope or the whole protein as the template. Our group developed MIPs either using cytochrome c adsorbed on a mercaptoundecanoic acetic acid (MUA) layer or the cysteine extended C-terminal nonapeptide as the template [9, 13]. Sellergren’s and Piletsky’s groups developed MIPs for the recognition of immunoglobulin G (IgG) using the whole antibody molecule, the Fc domain and the C-terminal 10-mer peptide, respectively [10]. All three MIPs recognized both human and goat IgGs.
In this paper, we compare the analytical performance in terms of measuring range, KD, and interference by HSA of MIPs generated by using a peptide epitope or the whole protein as the template for the recognition of alpha-fetoprotein (AFP) and the receptor binding domain (RBD) of SARS-CoV-2. For mechanistic insights, the density of binding pockets for whole-protein MIPs and epitope-MIPs has been analyzed by conductive AFM (C-AFM).
Both targets are of high diagnostic relevance: AFP is the most frequently determined biomarker for the early detection of the neoplastic disease hepatocellular carcinoma (HCC) which is the 7th most common cancer [14, 15]. The 29.5-kDa receptor binding domain is a globular structure protein which anchors the SARS-CoV-2 to the angiotensin-converting enzyme (ACE2) of the host cell and it is frequently used as the target structure in SARS-CoV-2 immunoassays and lateral flow assays [16]. Mimicking the biological counterpart, we have chosen “the non-terminal epitope” GFNCYFP derived from the exposed binding site of the RBD to the ACE. The heptapeptide contains a central cysteine (C488) which can bind the peptide to the gold surface [17]. In case of the MIPs targeting AFP beside the whole AFP glycoprotein (MW 69 kDa), the two peptides AALGVC and SKTRAALGVC of the exposed C-terminus containing an additional C-terminal cysteine for immobilization were used as template.
Materials and methods
Chemicals and reagents
Sulfuric acid (H2SO4) (96%), absolute ethanol (99.9%), sodium chloride (NaCl), potassium chloride (KCl), disodium hydrogen phosphate dihydrate (Na2HPO4•2H2O), potassium dihydrogen phosphate (KH2PO4), potassium hydroxide (KOH), potassium hexacyanoferrate (II) trihydrate (K4[Fe(CN)6]•3H2O), and potassium hexacyanoferrate (III) (K3[Fe(CN)6]) were from ROTH (Karlsruhe, Germany). One molar hydrochloric acid (HCl), phosphate-buffered saline (PBS), scopoletin (7-hydroxy-6-methoxycoumarin), the receptor binding domain from SARS-CoV-2 (RBD, 25.9 kDa), and albumin from human serum (HSA, 66.5 kDa) were purchased from Sigma-Aldrich (Merck, Darmstadt, Germany). Human alpha-fetoprotein (AFP, 63 kDa) from Bio-Rad (Munich, Germany). HAuCl4•3H2O was obtained from Chinoin Zrt. (Budapest, Hungary). The solutions throughout this work were prepared using deionized and filtered water obtained from a water purification system Milli-Q from Sartorius (Göttingen, Germany).
All peptides listed in Table 1 were purchased from Biosyntan (Berlin, Germany) and possess an amide group at the C-terminus.
Apparatus and electrochemical experiments
All the electrochemical measurements were conducted at room temperature using a CHI 440 electrochemical workstation (CH Instruments, Austin, TX, USA) and a three-electrode system. Gold wires (diameter: 0.5 mm; active surface: 20 mm2) (Goodfellow, Germany), a platinum coil, and an Ag/AgCl system were utilized as the working, counter, and reference electrodes, respectively.
SWV measurements and electrochemical oxidation for template removal were carried out using a one-compartment polymethylmethacrylate (PMMA) cell with a volume of 1 mL with Ag/AgCl (1 M KCl) as the reference electrode. SWV experiments were carried out in 5 mM ferri/ferrocyanide in phosphate-buffered saline (PBS) (137 mM NaCl, 10 mM Na2HPO4, 1.8 mM KH2PO4, and 2.7 mM KCl, pH 7.4) by scanning the potentials from − 0.2 to + 0.5 V, with a frequency of 10 Hz, amplitude of 50 mV, and a step height of 3 mV.
Electropolymerization of scopoletin was conducted in a one-compartment polychlorotrifluoroethylene (PCTFE) cell with a volume of 2 mL with Ag/AgCl (3 M KCl) (CH Instruments) as the reference electrode.
Cleaning of working electrodes
Gold wires were used as working electrodes for the preparation of MIPs. They were initially treated in 2.5 M KOH for 4 h and then stored overnight in 96% H2SO4. The purity of the electrodes was verified by cyclic voltammetry in 0.5 M H2SO4. Finally, anodic oxidation was performed in PBS at 1.1 V for 30 s.
Electrosynthesis of MIP, template removal, and binding assays
Table 2 shows the preparation details of the different MIPs used in this study with either the whole protein or peptide templates. Generally (Scheme 1), to create a MIP, a gold wire electrode was initially incubated in template in PBS at room temperature. To follow the adsorption of the respective template, the SWV signal of ferri/ferrocyanide was measured. The adsorption step was finished when the suppression of the SWV signal reached around 50%. Then, the polymer was synthesized around the adsorbed template by electropolymerizing 0.5 mM scopoletin in 10 mM NaCl containing 5% ethanol using between 20 and 40 pulse cycles, each starting with 0 V for 5 s and followed by 0.7 V for 1 s. The synthesis of non-imprinted polymers (NIPs) was conducted following the same procedure, but without template. After polymerization, the electrodes were rinsed with ddH2O and dried using a gentle nitrogen stream. The adsorbed template was then removed from the polymer by anodic oxidation in PBS (applied potential and length of the pulses are summarized in Table 2). Subsequently, the electrodes were washed with deionized water and dried carefully with nitrogen. The electrodes were then immersed in 1 mL of stirred ferri/ferrocyanide solution. For the binding assays, 2 µL of the analyte stock solution at a specific concentration was introduced into the cell. Both template removal and binding were characterized by measuring the SWV signal of ferri/ferrocyanide every 2 min without stirring until steady-state was reached.
Workflow of MIP synthesis and application shown for the preparation of AFP-binding MIP by using peptide epitope imprinting: (1) the template peptide is immobilized on the gold surface via terminal cysteine (blue dots), (2) the scopoletin monomer is electropolymerized, (3) the template is removed by anodic stripping, (4) binding of the whole AFP protein or the peptide by the liberated binding sites
Conductive atomic force microscopy measurements
To ensure planar MIP nanofilm specimens for C-AFM measurements gold slides were used instead of gold wires as working electrodes, which required slight changes in the surface pretreatment and MIP synthesis. Prior to use, the gold slides were rinsed successively with acetone and ethanol, dried under N2 stream, and then subjected to 10 min UV/ozone treatment (PSD Pro Series Digital UV Ozone System, Novascan, USA). For MIP preparation, a conical electrochemical cell made of methacrylate was used (CFLWCL-CONIC, Metrohm DropSens) connected to an Autolab PGSTAT12 Potentiostat/Galvanostat. A 3-electrode system was used with the gold slide, a platinum wire, and an Ag/AgCl (3 M KCl) (CH Instruments) as working, counter, and reference electrodes, respectively. For template immobilization, the conical cell was filled with 5 µM GFNCYFP peptide or 450 nM RBD in PBS and it was incubated at room temperature for 15 min by shaking at 350 rpm. After rinsing with deionized water, the electropolymerization was performed by cyclic voltammetry in a solution containing 0.5 mM scopoletin, 0.1 M NaCl, and 4 mM K4[Fe(CN)6]. Fifty cycles were made between 0.0 and 0.75 V at a scan rate of 50 mV s−1. Template removal was performed in PBS by chronoamperometry applying subsequently 0.9 V for 30 s, then 0 V for 5 s and finally again 0.9 V for 30 s. Gold deposition was performed in 0.2 mM HAuCl4 solution containing 0.1 M HCl and 2 mM sodium citrate by chronoamperometry using − 0.9 V for 30 s. C-AFM measurements were performed with a FlexAFM instrument (Nanosurf, Switzerland) at a setpoint of 20 nN and tip voltage of − 10 mV using an electrically conductive tip, PPP-ContPt (Nanosensors, Switzerland) with a tip radius of ca. 25 nm and a force constant of 0.2 N/m.
Results and discussion
MIPs for AFP using peptide epitope imprinting
Based on the prediction from modeling showing that already tripeptides could serve as templates for MIPs with µM measuring range [18], we applied the hexapeptide AALGVC (AC-MIP) as the template. However, the binding experiments for the AC-MIP revealed only negligible affinity towards the target AALGV (Figure S1). Already, the interaction of the AC-MIP with the unrelated peptide YFPLQS (Y-peptide from SARS-CoV-2 Spike protein) resulted in a considerably larger (non-specific) binding as revealed by the larger current suppression (Figure S1). Obviously, the template peptide (AALGVC) was too short for efficient rebinding of the imprinted peptide. To increase the binding strength of the MIP cavities, we applied a template which was extended at the N-terminus of the AC-peptide with four amino acids, resulting in SKTRAALGVC (SK-MIP). The elongation of the imprinted peptide template resulted in increased binding efficiency. The concentration dependence for the binding of the derived septa-peptide TRAALGV (TR-peptide) to the SK-MIP (Fig. 1) gave an estimated KD value of 14 µM. Imprinting with the longer SK-peptide also improved the discrimination of non-related C-terminal peptides, i.e., the pentapeptide VVSTQ from BSA (BSA-peptide), as shown by the significantly smaller relative current suppression as compared with the specific TR-peptide (Fig. 1b).
a SWVs curves characterizing the steps of MIP synthesis and the rebinding of the target TR-peptide (TRAALGV) to SK-MIP (SKTRAALGVC); b concentration dependent relative suppression of the SWV current signal for the SK-MIP upon binding the target protein AFP, the target TR-peptide (TRAALGV), and the non-related BSA-peptide (VVSTQ)
The parent protein AFP binding to the SK-MIP resulted in an even larger current suppression than the TR-peptide. Evaluation of the concentration dependence (Figs. 1b and 2) gave a KD value of 13.2 nM (Table 3). Since the non-imprinted polymer (NIP) is electrically insulating, this detection method based on the target-binding gated redox marker current is not applicable to measure the nonspecific AFP binding to the NIP. Still, it is a compulsory control experiment that indeed revealed insignificant signal change upon interacting the NIP with AFP as represented by a “formal” imprinting factor (IF) of 6.1 in the linear measuring range (Fig. 2). The selectivity of the SK-MIP was tested using HSA a protein of very similar size to AFP. An excellent discrimination discrimination was observed, i.e. the signal level was close to the background level measured on NIPs (Fig. 2).
MIPs for RBD using peptide epitope imprinting
We turned to confirm the conclusions of the AFP-MIP studies to another model system, i.e., RBD. Similar to the AFP case where, as shown in the “MIPs for AFP using peptide epitope imprinting” section (Figure S1), only a very weak binding of the pentapeptide AALGV to the AALGVC-based MIP was observed, the tetrapeptide EGFN showed no measurable binding to the G-MIP.
The MIP using the G-peptide as the template showed almost fourfold higher affinity towards the whole RBD (KD of 14.7 ± 0.9 nM) as compared with the G-peptide (58 nM) (Fig. 3b). The KD value for RBD is comparable with that determined by SPRi in a similar system, but with the epitope-imprinted nanofilms prepared on a planar chip by a combination of peptide microspotting and electrosynthesis, i.e., 2.2 ± 0.4 nM [19].
MIPs using the pre-adsorbed (whole) proteins as template
A sufficiently strong adsorption of the proteins to the gold substrate to withhold them during the electrosynthesis of the polymer is the prerequisite for the MIP synthesis around the adsorbed protein template. We found that both AFP and RBD bound spontaneously to the gold electrode (Figure S2), but could be effectively desorbed after a 30-s duration anodic pulse of 950 mV for AFP and 970 mV for RBD.
The MIPs prepared by using the whole protein as the template had affinities in the lower nM range for the binding of the protein templates, i.e., the KD values for AFP and RBD were 14.5 nM and 16.8 nM, respectively (Table 3). As shown in the “MIPs for AFP using peptide epitope imprinting” and “MIPs for RBD using peptide epitope imprinting” sections, the respective epitope-MIPs showed very similar affinities towards the target proteins. On the other hand, the KD values reported earlier for MIPs either using cytochrome c adsorbed on a MUA layer (10 µM) or the cysteine extended C-terminal nonapeptide as the template (8.5 µM) are almost three orders of magnitude larger [9, 13].
For both proteins, the magnitude of the relative suppression of the SWV signal at saturation was higher for the whole protein imprinted polymers than that of the epitope-MIPs (Figs. 2b and 3). In case of AFP, the difference was less significant than for RBD.
C-AFM investigation of the MIP nanofilms
To better understand and reveal contingent differences in the morphology of the MIP nanofilms prepared by using peptide or protein templates, we used AFM measurements. Molecular size cavities are difficult to be identified in polymer films [20]; therefore, we attempted to reveal them by electroplating gold in the imprinted cavities. Since during gold deposition gold nanoparticles may also form nonspecifically on the polymer film surface, we used C-AFM to discriminate between the gold grown through the imprinted cavities and nonspecific gold deposition. For this purpose, a small potential difference (− 10 mV) is applied between the gold substrate underneath the MIP and NIP nanofilm and the conducting AFM tip and as such the current map will reveal only the gold spots that are electrically connected to the gold electrode, i.e., the gold was grown through the imprinted cavities.
As shown by the current map in Fig. 4, without imprinting the electrically insulating polyscopoletin film (NIP) proved to be largely pinhole-free after the gold electrodeposition. In turn, both the peptide and the protein imprinted films revealed locations with elevated current indicative of the electrodeposited gold through the imprinted cavities. While at this stage we cannot claim revealing all imprinted cavities, the results clearly suggest a higher surface density of the imprinted cavities in case of the peptide imprinting. However, apparently this higher density of binding cavities in case of peptide epitope imprinting is not beneficial for the much larger size proteins when using the target gated redox mediator current for detection as revealed by the larger Bmax values for whole protein imprinting shown in Table 3.
Since the purpose of these MIPs is ultimately the detection of the proteins, their analytical performance towards proteins is most relevant. Such KD values in the lower nanomolar range can be considered rather competitive given that the KD values for protein-MIPs are generally in the µM to nM range [13, 21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46] including AFP and RBD (Table S1).
On the other hand, the KD values for binding the template (or truncated) peptides to the epitope-MIPs for the G-peptide (58 nM) and TR-peptide (14 µM) are larger than for the respective parent protein. This suggests an apparently lower affinity for the peptide binding. However, considering only the binding to the imprinted cavity with 1:1 stoichiometry, the affinity of an epitope-MIP towards the parent protein should not exceed the affinity for the peptide. This relation was found earlier for the epitope-MIP templated with a cysteine extended C-terminal nonapeptide of Cyt c and direct readout by fluorescence, i.e., a lower affinity was found for the parent protein (KD of 8.54 µM) as compared with the target peptide (KD of 2.5 µM). On the other hand, when the “non-epitope” surface of the protein target nonspecifically interacts with the non-imprinted part of the MIP surface, an apparently stronger binding of the protein can be found. The MIP using the G-peptide as the template showed almost fourfold higher affinity towards the whole RBD as compared with the G-peptide and even higher towards the spike protein.
The higher (apparent) binding affinity of the MIPs determined by the indirect method of redox marker gating for both protein targets as compared with the respective template peptides might be caused by the lager “footprint”of the proteins. Thus, upon binding to the cavities of the epitope-MIP, the larger surface covered by the protein will disturb the non-linear diffusion of the redox marker to the pores (microelectrode behavior) [21] and can block multiple adjacent binding pockets while the peptide occupies only one pocket. This effect could explain the decrease of the KD by one to two orders of magnitude for the binding of the parent protein as compared with that of the peptide.
However, the benefit of using peptide imprinting is remarkable in terms of selectivity, i.e., both epitope-imprinted polymers (G-MIP and SK-MIP) show an excellent discrimination of the unrelated HSA (Figs. 2 and 3). It seems that the presumably smaller binding cavities generated by peptide imprinting allow the binding of the target protein via the exposed epitope but restrain the binding of unrelated proteins. In spite of the smaller size of RBD (MW 25.9 kD), the RBD-MIP binds the larger HSA. We earlier found this behavior for a MIP using the pre-adsorbed globular protein transferrin as the template [22] and concluded that this effect might be caused by the larger cavities in the protein-MIPs formed by partially unfolded template molecules at the gold surface [47]. Obviously, these lager cavities can accommodate protein molecules of comparable and even larger size than the template, i.e., facilitating the nonspecific binding of HSA.
Conclusion
The affinities as expressed by KD of the peptide and whole protein imprinted polymers are very similar for both model proteins, RBD and AFP. For a cautious extrapolation of this conclusion, it should be noted that we used at least 7 amino acid long peptides and the extremely thin polymer layers (ca. 10 nm) provided by the self-regulated electrosynthesis of insulating polymer films. For both AFP- and RBD-MIPs, the epitope-imprinted films had an apparently lower affinity for the peptide than for the parent protein that is most likely an artifact of the detection method due to the larger footprint of the protein on the MIP surface. The main advantage of the epitope imprinting in terms of analytical performance is the significantly better discrimination of nonrelated proteins as exemplified for HSA as compared to whole protein imprinted polymer films. Furthermore, for mass production the lower reagent cost is a clear advantage, to which adds the high purity of peptides stemming from a controlled synthetic process undergoing purification and quality control by HPLC and mass spectrometry.
References
Polyakov MV (1931) Adsorption properties and structure of silica gel. Zhur Fiz Khim 2:799–895
Wulff G, Sarhan A (1972) Über die Anwendung von enzymanalog gebauten Polymeren zur Racemattrennung. Angew Chemie 84:364–364. https://doi.org/10.1002/ange.19720840838
Arshady R, Mosbach K (1981) Synthesis of substrate-selective polymers by host-guest polymerization. Die Makromol Chemie 182:687–692. https://doi.org/10.1002/MACP.1981.021820240
Birnbaumer GM, Lieberzeit PA, Richter L et al (2009) Detection of viruses with molecularly imprinted polymers integrated on a microfluidic biochip using contact-less dielectric microsensors. Lab Chip 9:3549–3556. https://doi.org/10.1039/B914738A
Amorim MS, Sales MGF, Frasco MF (2022) Recent advances in virus imprinted polymers. Biosens Bioelectron X 10:100131. https://doi.org/10.1016/J.BIOSX.2022.100131
Rachkov A, Minoura N (2000) Recognition of oxytocin and oxytocin-related peptides in aqueous media using a molecularly imprinted polymer synthesized by the epitope approach. J Chromatogr A 889:111–118
Rachkov A, Minoura N (2001) Towards molecularly imprinted polymers selective to peptides and proteins. The epitope approach. Biochim Biophys Acta - Protein Struct Mol Enzymol 1544:255–266. https://doi.org/10.1016/S0167-4838(00)00226-0
Yoshimatsu K, Yamazaki T, Hoshino Y et al (2014) Epitope discovery for a synthetic polymer nanoparticle: a new strategy for developing a peptide tag. J Am Chem Soc 136:1194–1197. https://doi.org/10.1021/ja410817p
Dechtrirat D, Jetzschmann KJ, Stöcklein WFM et al (2012) Protein rebinding to a surface-confined imprint. Adv Funct Mater 22:5231–5237. https://doi.org/10.1002/adfm.201201328
Moczko E, Guerreiro A, Cáceres C et al (2019) Epitope approach in molecular imprinting of antibodies. J Chromatogr B 1124:1–6. https://doi.org/10.1016/j.jchromb.2019.05.024
Nishino H, Huang CS, Shea KJ (2006) Selective protein capture by epitope imprinting. Angew Chemie - Int Ed 45:2392–2396. https://doi.org/10.1002/anie.200503760
Singh M, Gupta N, Raghuwanshi R (2017) Epitope imprinting approach to monitor diseases. J Mol Genet Med 11:1–6. https://doi.org/10.4172/1747-0862.1000270
Bosserdt M, Gajovic-Eichelman N, Scheller FW (2013) Modulation of direct electron transfer of cytochrome c by use of a molecularly imprinted thin film. Anal Bioanal Chem 405:6437–6444. https://doi.org/10.1007/s00216-013-7009-8
Wright LM, Kreikemeier JT, Fimmel CJ (2007) A concise review of serum markers for hepatocellular cancer. Cancer Detect Prev 31:35–44. https://doi.org/10.1016/J.CDP.2006.11.003
Yuen MF, Lai CL (2005) Serological markers of liver cancer. Best Pract Res Clin Gastroenterol 19:91–99. https://doi.org/10.1016/J.BPG.2004.10.003
Liu Z, Xiao X, Wei X et al (2020) Composition and divergence of coronavirus spike proteins and host ACE2 receptors predict potential intermediate hosts of SARS-CoV-2. J Med Virol 92:595–601. https://doi.org/10.1002/JMV.25726
Zhang X, Waffo AT, Yarman A et al (2022) How an ACE2 mimicking epitope-MIP nanofilm recognizes template-related peptides and the receptor binding domain of SARS-CoV-2. Nanoscale 14:18106–18114. https://doi.org/10.1039/D2NR03898F
Settipani J, Karim K, Chauvin A et al (2018) Theoretical aspects of peptide imprinting: screening of MIP (virtual) binding sites for their interactions with amino acids, di- and tripeptides. J Chinese Adv Mater Soc 6:301–310. https://doi.org/10.1080/22243682.2018.1467279
Bognár Z, Supala E, Yarman A et al (2022) Peptide epitope-imprinted polymer microarrays for selective protein recognition. Application for SARS-CoV-2 RBD protein. Chem Sci 13:1263–1269. https://doi.org/10.1039/D1SC04502D
Neusser G, Eppler S, Bowen J et al (2017) FIB and MIP: understanding nanoscale porosity in molecularly imprinted polymers via 3D FIB/SEM tomography. Nanoscale 9:14327–14334. https://doi.org/10.1039/C7NR05725C
Kondo T, Udagawa I, Aikawa T et al (2016) Enhanced sensitivity for electrochemical detection using screen-printed diamond electrodes via the random microelectrode array effect. Anal Chem 88:1753–1759. https://doi.org/10.1021/ACS.ANALCHEM.5B03986/ASSET/IMAGES/LARGE/AC-2015-03986V_0008.JPEG
Zhang X, Yarman A, Erdossy J et al (2018) Electrosynthesized MIPs for transferrin: plastibodies or nano-filters? Biosens Bioelectron 105:29–35. https://doi.org/10.1016/J.BIOS.2018.01.011
Taheri N, Khoshsafar H, Ghanei M et al (2022) Dual-template rectangular nanotube molecularly imprinted polypyrrole for label-free impedimetric sensing of AFP and CEA as lung cancer biomarkers. Talanta 239:123146. https://doi.org/10.1016/J.TALANTA.2021.123146
Cieplak M, Szwabinska K, Sosnowska M et al (2015) Selective electrochemical sensing of human serum albumin by semi-covalent molecular imprinting. Biosens Bioelectron 74:960–966. https://doi.org/10.1016/j.bios.2015.07.061
Bosserdt M, Erdőssy J, Lautner G et al (2015) Microelectrospotting as a new method for electrosynthesis of surface-imprinted polymer microarrays for protein recognition. Biosens Bioelectron 73:123–129. https://doi.org/10.1016/j.bios.2015.05.049
Cai D, Ren L, Zhao H et al (2010) A molecular-imprint nanosensor for ultrasensitive detection of proteins. Nat Nanotechnol 5:597–601. https://doi.org/10.1038/nnano.2010.114
Karimian N, Vagin M, Zavar MHA et al (2013) An ultrasensitive molecularly-imprinted human cardiac troponin sensor. Biosens Bioelectron 50:492–498. https://doi.org/10.1016/j.bios.2013.07.013
Yarman A (2018) Electrosynthesized molecularly imprinted polymer for laccase using the inactivated enzyme as the target. Bull Korean Chem Soc 39:483–488. https://doi.org/10.1002/bkcs.11413
Drobysh M, Ramanaviciene A, Viter R et al (2022) Biosensors for the determination of SARS-CoV-2 virus and diagnosis of COVID-19 infection. Int J Mol Sci 23:666. https://doi.org/10.3390/IJMS23020666
Ayankojo AG, Boroznjak R, Reut J et al (2022) Molecularly imprinted polymer based electrochemical sensor for quantitative detection of SARS-CoV-2 spike protein. Sensors Actuators B Chem 353:131160. https://doi.org/10.1016/J.SNB.2021.131160
Palladino P, Minunni M, Scarano S (2018) Cardiac troponin T capture and detection in real-time via epitope-imprinted polymer and optical biosensing. Biosens Bioelectron 106:93–98. https://doi.org/10.1016/J.BIOS.2018.01.068
Zhao C, Ma X, Li J (2017) An insulin molecularly imprinted electrochemical sensor based on epitope imprinting. Chinese J Anal Chem 45:1360–1366. https://doi.org/10.1016/S1872-2040(17)61039-9
Cheng Z, Zhang L, Li Y (2004) Synthesis of an enzyme-like imprinted polymer with the substrate as the template, and its catalytic properties under aqueous conditions. Chem – A Eur J 10:3555–3561. https://doi.org/10.1002/CHEM.200305370
Li S, Yang K, Liu J et al (2015) Surface-imprinted nanoparticles prepared with a His-tag-anchored epitope as the template. Anal Chem 87:4617–4620. https://doi.org/10.1021/ac5047246
Li S, Yang K, Zhao B et al (2016) Epitope imprinting enhanced IMAC (EI-IMAC) for highly selective purification of His-tagged protein. J Mater Chem B 4:1960–1967. https://doi.org/10.1039/C5TB02505B
Bognár Z, Kozma J, Kovács N, Gyurcsányi RE (2023) Novel functional monomer for the electrochemical synthesis of highly affine epitope-imprinted polymers. Electroanalysis e202300025. https://doi.org/10.1002/ELAN.202300025
Yang K, Li S, Liu J et al (2016) Multiepitope templates imprinted particles for the simultaneous capture of various target proteins. Anal Chem 88:5621–5625. https://doi.org/10.1021/acs.analchem.6b01247
Erdőssy J, Horváth V, Yarman A et al (2016) Electrosynthesized molecularly imprinted polymers for protein recognition. TrAC Trends Anal Chem 79:179–190. https://doi.org/10.1016/J.TRAC.2015.12.018
Zhang X, Caserta G, Yarman A et al (2021) “Out of pocket” protein binding—a dilemma of epitope imprinted polymers revealed for human hemoglobin. Chemosensors 9:128. https://doi.org/10.3390/CHEMOSENSORS9060128
Amouzadeh Tabrizi M, Fernández-Blázquez JP, Medina DM, Acedo P (2022) An ultrasensitive molecularly imprinted polymer-based electrochemical sensor for the determination of SARS-CoV-2-RBD by using macroporous gold screen-printed electrode. Biosens Bioelectron 196:113729. https://doi.org/10.1016/J.BIOS.2021.113729
Parisi OI, Francomano F, Dattilo M et al (2022) The evolution of molecular recognition: from antibodies to molecularly imprinted polymers (MIPs) as artificial counterpart. J Funct Biomater 13:12. https://doi.org/10.3390/JFB13010012
Raziq A, Kidakova A, Boroznjak R et al (2021) Development of a portable MIP-based electrochemical sensor for detection of SARS-CoV-2 antigen. Biosens Bioelectron 178:113029. https://doi.org/10.1016/j.bios.2021.113029
Zang Y, Zhang Y, Wei R et al (2023) Difunctional molecularly imprinted polymers and heterostructured CdS nanoparticle-sensitized ZnO nanorod arrays for antibody-free photoelectrochemical alpha-fetoprotein sensor. J Electroanal Chem 944:117631. https://doi.org/10.1016/J.JELECHEM.2023.117631
Shen X, Ma Y, Zeng Q et al (2016) Molecularly imprinted electrochemical sensor for advanced diagnosis of alpha-fetoprotein. Anal Methods 8:7361–7368. https://doi.org/10.1039/c6ay01922f
Wu Y, Wang Y, Wang X et al (2019) Electrochemical sensing of α-fetoprotein based on molecularly imprinted polymerized ionic liquid film on a gold nanoparticle modified electrode surface. Sensors (Switzerland) 19:3218. https://doi.org/10.3390/s19143218
Lai Y, Zhang C, Deng Y et al (2019) A novel α-fetoprotein-MIP immunosensor based on AuNPs/PTh modified glass carbon electrode. Chinese Chem Lett 30:160–162. https://doi.org/10.1016/J.CCLET.2018.07.011
Scheller F, Renneberg R (1982) Electrochemical studies of proteins. In: Dryhurst G, Kadish KM, Scheller F, Renneberg R (eds) Biological electrochemistry, 1st edn. Academic Press, London, UK, pp 398–521
Acknowledgements
The authors would like to thank the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) under Germany’s Excellence Strategy—EXC 2008–390540038 (Unifying Systems in Catalysis—UniSysCat) and the German Ministry of Education and Research (BMBF, 01DH20018). This work was partially supported by the Ministry of Innovation and Technology of Hungary from the National Research, Development and Innovation Fund under the TKP2021 funding scheme, project no. BME-EGA-02.
Author information
Authors and Affiliations
Contributions
Conceptualization, R.E.G. and F.W.S.; methodology and formal analysis, X.Z., A.Y., N.K., Z.B., and R.E.G.; investigation, X.Z., N.K., and Z.B.; writing—original draft preparation, X.Z., R.E.G., and F.W.S.; writing—review and editing, R.E.G. and F.W.S.; visualization, X.Z. Z.B., and N.K.; supervision, A.Y., R.E.G., and F.W.S.; project administration, F.W.S.; funding acquisition, R.E.G., F.F.B., and F.W.S. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
The paper is dedicated to both pioneers of MIP development Günther Wulff and Klaus Mosbach.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, X., Yarman, A., Kovács, N. et al. Specific features of epitope-MIPs and whole-protein MIPs as illustrated for AFP and RBD of SARS-CoV-2. Microchim Acta 191, 242 (2024). https://doi.org/10.1007/s00604-024-06325-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-024-06325-0