Abstract
Although only a relatively small proportion of plant species form ectomycorrhizae with fungi, it is crucial for growth and survival for a number of widespread woody plant species. Few studies have attempted to investigate the fine scale spatial structure of entire root systems of adult ectomycorrhizal (EcM) plants. Here, we use the herbaceous perennial Bistorta vivipara to map the entire root system of an adult EcM plant and investigate the spatial structure of its root-associated fungi. All EcM root tips were sampled, mapped and identified using a direct PCR approach and Sanger sequencing of the internal transcribed spacer region. A total of 32.1% of all sampled root tips (739 of 2302) were successfully sequenced and clustered into 41 operational taxonomic units (OTUs). We observed a clear spatial structuring of the root-associated fungi within the root system. Clusters of individual OTUs were observed in the younger parts of the root system, consistent with observations of priority effects in previous studies, but were absent from the older parts of the root system. This may suggest a succession and fragmentation of the root-associated fungi even at a very fine scale, where competition likely comes into play at different successional stages within the root system.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Although a relatively small number of plant species form ectomycorrhizae compared to arbuscular mycorrhiza (Brundrett and Tedersoo 2018), ectomycorrhizal (EcM) fungi is crucial for growth and survival of a number of ecologically and economically important woody plant species in boreal, temperate and arctic regions (Smith and Read 2008). In these environments, EcM fungi have been shown to play a prominent role including, but not limited to; carbon sequestration (Clemmensen et al. 2013; Averill et al. 2014), nutrient cycling and decomposition of organic matter (Lindahl et al. 2007; Kyaschenko et al. 2017; Sterkenburg et al. 2018).
An increasing amount of studies have addressed the spatial variation in EcM fungal communities, ranging from local (Genney et al. 2006; Buée et al. 2007; Bahram et al. 2011, 2015; Anderson et al. 2014), regional (Jarvis et al. 2013; Miyamoto et al. 2015), continental (Talbot et al. 2014; Glassman et al. 2015; van der Linde et al. 2018) and even global scales (Tedersoo et al. 2010; Brundrett and Tedersoo 2018). At all scales, there is a high level of patchiness and a rapid turnover in species composition. Fungal communities in forest ecosystems are auto-correlated at relatively small spatial scales, typically less than 3 m (Lilleskov et al. 2004; Pickles et al. 2012), and forest microsite affects the EcM community composition (Tedersoo et al. 2008). At smaller scales, the abundance of EcM fungi in a mixed forest is highly variable at a 5 cm scale, and a complete turnover in species composition can be observed at a 50 cm scale (Tedersoo et al. 2003).
Although the community composition of EcM fungi has been widely studied at relatively local scales (cm–m), less is known about the spatial distribution at smaller scales (μm–cm) and within single root systems (Bogar and Peay 2017). Although individual EcM genets can cover relatively large belowground areas, (up to 18 m in some species) (Dahlberg and Stenlid 1994; Bergemann and Miller 2002), EcM root tips may be located less than 1 mm from each other in the soil, and individual hyphae interact at micrometre scale (Bogar and Peay 2017). The few studies conducted on a `single host plant scale` show that the EcM fungal community is very diverse even within an individual host plant. A single tree of Populus tremula hosted more than 200 root-associated fungal species, as well as dozens of individuals of the same species (Bahram et al. 2011). Even in seedlings and herbaceous plants, the number of species is surprisingly high. The number of root-associated operational taxonomic units (OTUs, a proxy for species) may range from 37 to 159 within single root systems of the EcM-forming plant B. vivipara (Yao et al. 2013), and Yoshida et al. (2014) found 40 species of EcM fungi associated with three Tsuga diversifolia seedlings.
The processes structuring EcM fungal communities and maintaining species richness at finer scales in root systems may be attributed to several selective processes, ecological drift and dispersal limitations (Bogar and Peay 2017). EcM fungi are phylogenetically diverse, and their morphology when colonising root tips varies between species, from contact exploration types (i.e. no emanating hyphae from the mantle covering the plant root tip), to rhizomorphic, long-distance exploration types (Agerer 2001). However, exploration type is often relatively conserved within genera or even family (Agerer 2001). The exploration types of EcM fungi associated with plant roots have been shown to correlate with both root density (Peay et al. 2011) and forest age (Clemmensen et al. 2015). Therefore, exploration types may potentially be important in structuring communities also at finer scales. Stochastic events, such as timing of arrival of a species, can have major effects on the outcome of the species distribution within a single root system (Kennedy and Bruns 2005; Kennedy et al. 2009), and may, at least in part, explain some variation in root-associated communities found across host plants at small scales (Yao et al. 2013). A stochastic event, such as the timing of arrival of a species (i.e. priority effect) is thought to play a major role in structuring EcM communities. There are, however, some contrasting results regarding the importance of priority effects in EcM fungal communities. Lilleskov & Bruns (2003) deemed competition more important than the priority effect, as a slower colonising species had a greater competitional advantage, in spite of a later colonisation. On the contrary, Kennedy & Bruns (2005) found competition to be an important factor, but a strong priority effect determined the outcome of the dominant species in their EcM root systems. They later confirmed that root systems were dominated by the earlier coloniser (Kennedy et al. 2009), and that priority effects determined the outcome. Yoshida et al. (2014) investigated the fine scale structure of three seedlings of T. diversifolia from a natural environment, and similarly observed that the priority effect was likely important in structuring the EcM fungal community, as clusters of root tips colonised by individual species were observed in the root system. However, in mature forest systems, competitive interactions seem to be the determining process in structuring EcM communities at finer scales (Pickles et al. 2012). There is strong evidence that priority effects play important roles in structuring EcM communities for seedlings, however, EcM competitive patterns may be life-stage dependent (Kennedy et al. 2011), and should thus be investigated further.
Most of the evidence for priority effects in EcM fungal communities comes from studies on seedlings. To the best of our knowledge, no exhaustive study looking at fine scale spatial structure of root-associated fungi have been conducted on adult EcM plants (i.e. investigating all EcM root tips within the root system). This may be because most ectomycorrhizae-forming plants are woody plant species with large root systems, and thus more challenging to investigate in detail. However, a few herbaceous plants also form ectomycorrhizae (Smith and Read 2008), including the Alpine bistort; a small, perennial herb (Bistorta vivipara (L.) Delarbre, synonym Persicaria vivipara (L). Ronse Decr.). Because of its small, condensed root system, it is possible to investigate the entire root systems, and several studies have shown that B. vivipara hosts phylogenetically diverse root-associated fungal communities (Brevik et al. 2010; Blaalid et al. 2012, 2014; Kauserud et al. 2012; Yao et al. 2013; Botnen et al. 2014; Mundra et al. 2015b).
In the present study, we take advantage of the small root system of B. vivipara to assess the diversity and spatial distribution of the root-associated fungal community within a single plant root system at a root-tip level. The root morphology of B. vivipara also allows us to investigate root system age, as the plant grows directionally with an unbranched rhizome, where the growth of the rhizome is restricted to the distal end (Diggle 1997). Numerous fine roots, where the EcM tips are formed, grows out from the rhizome. The exact turnover rates of B. vivipara fine roots are not known. However, in a study from the arctic focusing on temporal changes in the root-associated fungal communities of B. vivipara (Mundra et al. 2015a), fine roots were observed year-round, indicating they have a longer persistence time, not just being produced every growing season. Thus, we can expect that the fine roots in general are younger in the actively growing end of the rhizome and that there is an age gradient within the root system.
Due to the likely influence of priority effects, we hypothesise that there will be a spatial clustering of fungal species in the root system, as observed earlier in other plant root systems (Kennedy and Bruns 2005; Kennedy et al. 2009; Yoshida et al. 2014). If competition and succession is a stronger driver of structuring the fungal community, we expect to observe a weaker signal of priority effects, and less spatial clustering of species. This may also be reflected as a change in exploration types along the age gradient. To investigate this, a single EcM root system of B. vivipara was dissected and all EcM root tips mapped and sampled. A direct PCR approach, where the entire internal transcribed spacer (ITS) region was amplified from crushed root tips without DNA extraction, followed by Sanger sequencing, was used to identify fungal OTUs within the root system of one Bistorta vivipara individual.
Materials and methods
Host plant description
The host plant, B. vivipara, is a long-lived perennial in the family Polygonaceae, first reported as ectomycorrhizae-forming by Hesselman (1900) and later confirmed in several studies (e.g. Massicotte et al. 1998; Brevik et al. 2010; Kauserud et al. 2012; Botnen et al. 2014). The plant is widely distributed in the Northern Hemisphere and is common in arctic and alpine ecosystems (Jonsell 2000). Reproductive structures include flowers, but more commonly, reproduction is through asexual bulbils. The plant produces an unbranched rhizome (see Fig. 1) that grows plagiotropically 3–4 cm below ground, from which fine roots are growing (Diggle 1997). The fungal community of B. vivipara has been found to be highly diverse, comprising a wide range of both endo- and ectomycorrhizal fungi as well as saprotrophs, parasites and endophytes. Basidiomycota and Ascomycota have been found to be the most dominant phyla, although Glomeromycota and Zygomycota are found as well (Massicotte et al. 1998; Carlsen 2002; Eriksen et al. 2002; Brevik et al. 2010; Blaalid et al. 2012, 2014; Yao et al. 2013; Botnen et al. 2014).
Schematic drawing of B. vivipara, with its unbranched rhizome with fine roots and EcM root tips. R1–R4 are the approximate parts the rhizome was dissected into, and subsequently used as age groups, where R1 is the youngest part of the root system, and R4 is the oldest. Figure credit: Erlend Y. Fines
Field description and sample preparation
Sampling was conducted August 1, 2012, at the glacier foreland in front of Blåisen, Finse, Norway (60° 33′ N, 7° 31′ E, 1388 m a.s.l.), which is an alpine ecosystem. The sampling site was sparsely vegetated, with Salix herbaceae L. and B. vivipara, as the dominating plant species. The site was chosen because of the very sparse vegetation, making it possible to sample and analyse a single root system without breaking off roots due to entanglement and intermixing with roots from other plants.
One randomly chosen B. vivipara plant and its entire root system was dug up and stored in a zip-lock bag in a cooler for transportation, and later, in a fridge at 4 °C until further processing. The root system was rinsed 1 day after sampling and transportation, first gently in tap water and then thoroughly in distilled water. A dissecting microscope was used to inspect the roots to ensure that all debris and soil attached to the plant’s rhizome or roots were removed. As a final rinsing step, the roots were washed three times in milli-Q H2O (mqH2O) for 30 s, before excess water was gently dried off with sterile tissue paper. The root system was stored in a petri dish at 4 °C until further processing.
Root tip sampling
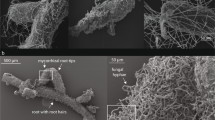
Within 2 days after sampling, the rhizome along with attached fine root was dissected into four approximately equal sized parts, which were named R1, R2, R3 and R4 (Fig. 1). R1 was the distal end (i.e. where the current year’s leaves and flowers grow) and thereby the youngest part of the rhizome and R4 the proximal and oldest part of the rhizome (Diggle 1997). The number of fine roots present at each rhizome part was highly variable, ranging from three (R2, R4) to 13 (R1), where newly formed roots were only observed in R1. The fine roots were numbered chronological from distal to proximal end and cut off from the rhizome, totalling 29 fine roots from the entire root system, and further, stored in individual petri dishes at − 18 °C until further processing. Root tips were sampled and numbered from top (closest to the rhizome) to bottom of the fine roots and simultaneously mapped onto an overview photograph to keep track of their relative position in the root system (see also Fig. S2 for a photograph of a fine root). If root tips were located on branches from the main fine root, this was noted as well. The number of root tips per fine root varied from three to 278 and were relatively even spaced on the fine roots. All root tips had either a fungal mantle or signs of fungal colonisation (i.e. hyphae were present when observed under the dissection microscope).
Dissection of the fine roots was conducted in a petri dish with mqH2O under a dissecting microscope. Each root tip was cut off using a scalpel, transferred to an Eppendorf tube with 100 μl mqH2O and stored at − 18 °C. The total number of root tips in the root system was 2302.
Molecular analyses
The ITS region was used to barcode the fungal component of the root tips. The fungal-specific primers ITS1F and ITS4, targeting the entire ITS region including ITS1, 5.8S and ITS2 (White et al. 1990; Gardes and Bruns 1993) were used for all PCR reactions. Before PCR, all samples were crushed using two Qiagen® Tungsten Carbide beads (3 mm) on a Rech Ball Mill MM301 (Verder Scientific, Haan, Germany) for 2 × 2 min at 20 rounds per sec. Several approaches were used to amplify as many root tips as possible. The Thermo Scientific Direct PCR Plant kit (Thermo Fisher Scientific Inc., Waltham, MA, USA) was used for DNA amplification of most samples. The kit allows for direct amplification without time-consuming DNA extraction and purifications steps. The kit was modified for fungi as follows: for each reaction, 2 μl crushed root tip in water (DNA template) was added to a master mix consisting of 10-μl 3× plant buffer, 2 μl and 5 μM ITS1F, 2 μl and 5 μM ITS4, 3.6 μl mqH2O and 0.4-μl Phire Hot Start Polymerase (Thermo Fisher Scientific Inc., Waltham, MA, USA). The PCR reaction was run with an initial step for 5 min at 98 °C, 40 cycles of denaturation at 98 °C for 20 s, primer binding at 53 °C for 20 s and extension at 72 °C for 2 min, and a final extension at 72 °C for 2 min, before cooling down to 10 °C. For samples with unsuccessful amplification after the first setup, a new PCR reaction was run as previously, but the number of PCR cycles was reduced to 36. For those that still did not amplify, a 50× dilution of the crushed root tips was used as template, using the above temperatures and 36 cycles. Further, as a last try, the REDExtract-N-Amp Plant PCR Kits (Sigma), slightly modifying the manufacturer’s instructions, was used to amplify the root tips that had not worked in the initial amplification attempts. In short, 2 μl of the crushed root tip and mqH2O solution was incubated in extraction solution for 10 min at 95 °C before adding 30 μl of dilution buffer. PCR was set up using illustra™ PuReTaq Ready-To-Go™ PCR Beads, according to the manufacturer’s instructions. The PCR products were cleaned using illustra™ ExoStar™ (VWR international Ldt., Dublin, Ireland) and submitted for Sanger sequencing at the ABI lab, University of Oslo or to the GATC biotech (Konstanz, Germany). The ITS amplicons were sequenced in one direction only. Initially, ITS4 was used as a sequencing primer for 63.9% of the amplicons, but due to poor quality of some of the sequences, ITS5 was used for the remaining samples.
Bioinformatics and statistical analyses
A total of 1891 ITS sequences were obtained and manually trimmed and edited using Geneious® 8.1.7 Biomatters (Biomatters Ltd., Auckland). After removing sequences with low quality chromatograms, i.e. sequences with more than ten ambiguities and read-length shorter than 350 base pairs (bp), 1080 sequences remained for further analyses.
Because several of the PCR negatives produced clear bands on the gel, additional quality control of the sequences was necessary to avoid including contaminants in the dataset. All 1080 sequences were aligned using custom settings in the MAFT package included in Geneious ® 8.1.7. A neighbour joining (NJ) tree was constructed from the alignment using custom settings in Genious® 8.1.7. Clusters were manually extracted from the cladogram. Each cluster of sequences was realigned using MAFT (as above), further manually inspected, edited and quality checked within the alignment, and one sequence from each alignment of identical sequences was BLASTED against UNITE (Kõljalg et al. 2013). Clusters of sequences that included negative controls were altogether removed from the dataset. Further, sequences that had top hits to known laboratory contaminants and sequences from several ubiquitous polyporid fungi were removed from the dataset (Table S2). Only sequences with top hit to EcM fungi and known root-associated fungi were included, with a few exceptions (see “Discussion”).
The remaining dataset of 739 sequences were clustered into OTUs using BlastClust (Alva et al. 2016) with 97.0% sequence similarity and minimum 70.0% query cover. All OTUs were manually inspected in Geneious® 8.1.7. Most of the OTUs contained only identical sequences.
All further analyses were conducted in R (R Core Team 2017). To examine if the sampling depth was sufficient, species accumulation curves were constructed (Oksanen et al. 2013), with fine roots as sites. Richness and Shannon diversity index (H′) was calculated for each part of the rhizome and each fine root using the vegan package (Oksanen et al. 2013) to assess species richness and diversity. OTUs that had taxomonic affinity to EcM fungi were assigned to exploration types by species using determination and characterisation of ectomycorrhizae (DEEMY) (Agerer & Rambold, 2004–2018). For instances where OTUs had poor match to a species, but exploration type was consistent throughout the genus, exploration type was also assigned. Eleven OTUs were assigned to mycorrhizal exploration type: contact, short distance smooth, medium distance smooth or medium distance fringe according to DEEMY.
Spatial structure
To investigate spatial structure in the root system, a matrix of the abundances of OTUs per fine root (site) was constructed. Fine roots with less than ten observations (successfully sequenced root tips) were excluded, leaving 18 fine roots and 701 root tips. Because ordination results strongly depend on the choice of ordination method and weighting function, a multiple parallel ordination (MPO) approach was used, as recommended by van Son & Halvorsen (2014). MPO was conducted using fine roots as plots, and OTUs as “species” observations. Three different weighting functions (data transformations) using power-function weighting (van der Maabel 1979; Clymo 1980) with raw data (w = 1), intermediate weighting (w = 0.5) and presence–absence (w = 0) were used, as described in van Son & Halvorsen (2014, for details see also Økland 1986). If ordination structure is similar between parametric and non-parametric ordination methods, there is an indication of a robust structuring of the dataset (Son and Halvorsen 2014). Therefore, two ordination methods were used: detrended correspondence analyses (DCA, Hill & Gauch 1980) and global non-metric multidimensional scaling (GNMDS, Kruskal 1964) ordinations were constructed using the vegan package v. 2.0–10 (Oksanen et al. 2013). For the DCA ordinations, default settings were used. For GNMDS, the following settings were used: two dimensions, dissimilarity measure used was Jaccard distance with an isometric geodesic step-across measure for Jaccard distance > 0.8 (Tenenbaum 2000); maximum number of iterations was set to 200 and 100 random initial starting configurations were used. The same settings were used for all three weighting functions. To see if axes of GNMDS and DCA correlated and which variables covaried with each other and the ordination axes, a correlation test with Kendall’s τ rank correlation coefficient (Kendall 1938; Kruskal 1958) was conducted. In addition, exploration types, diversity (Shannon’s H), richness per fine root, number of root tips (i.e. root density), length, branching and position of fine roots along the rhizome were fitted onto the ordination diagram using the envar function in the vegan package v. 2.0–10 (Oksanen et al. 2013), where only variables with a significant correlation to ordination axes were plotted.
To investigate whether a species tended to form clusters within the root system (i.e. indication of priority effect), a Jaccard distance matrix using the assigned OTU per root tip was constructed, where dissimilar OTUs would get a value of 1 and identical OTUs would get a value of 0. We used an approximation, because it is impossible to accurately estimate the actual position of root tips and how the fine roots are oriented in the soil before rinsing. Even though the spatial orientation in the soil and the actual physical distance between each root tip are unknown, root tips located on the same fine root or branch on fine roots were assumed to be in closer proximity than root tips located on other fine roots, and similarly, root tips that were sampled in sequence on the same branch were assumed to be located in close proximity in the soil (see also Fig. 1 for a schematic drawing of the B. vivipara root system and Fig. S2 for a photograph and description of our assumptions). Accordingly, three distance classes were defined: neighbours (strict neighbouring root tips, sampled in sequence on the same branch), adjacent (root-tip sample on the same branch on a fine root) and non-neighbours. These classes were used for constructing a distance matrix, such that strict neighbouring root tips were assigned a spatial distance of 1; adjacent root tips were scored as 2–23 (max number of root tips on branches), and non-neighbouring root tips were assigned a much higher number to emphasise non-adjacency (>> 300). Because initial tests showed very similar results for neighbouring and adjacent root tips, we only focus on neighbour (i.e. adjacent was merged into the neighbour class) and non-neighbour in the downstream results and discussion. Neighbours and non-neighbours were extracted from the matrix into data frames consisting of values of dissimilarities, so that pairs of root tips sharing an OTU obtained a value of 0, and pairs not sharing an OTU obtained a value of 1. The subsets were compared using a two-sided Welch T test, which takes into account the differences in sample size in the two classes. The root system was analysed as a whole, and subsequently, each rhizome part (R1–R4) was analysed separately, to see whether patterns were similar across the entire root system.
A Pearson’s χ2 test was conducted using the stats package (R Core Team 2017) to see whether the fungal community within parts of the root system was different from other parts than expected by chance. For each OTU, the actual distribution was tested against an expected random distribution in each part of the rhizome.
Results
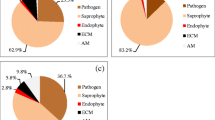
After discarding sequences of low quality and tentative contaminants, 739 sequences representing 32.1% of the total number of root tips in the root system remained for further analyses. The number of fine roots, root tips and OTUs varied considerably throughout the root system, as shown in Table 1. Three fine roots had the appearance of being newly formed (i.e. within the same year), and all three were found attached to R1 of the rhizome. A total of 41 fungal OTUs were observed in the root system. A large proportion (60.5%) of the successfully sequenced root tips was located in the youngest part (R1) of the root system, likely because this was also the part with the largest proportion of fine roots and total root tips. The most successfully sequenced part of the root system was R2, where 53.1% of all root tips gave an acceptable sequence. The frequency distribution of OTUs showed that the root system was dominated by a few abundant OTUs, and 36.6% of the OTUs was observed once (Fig. 2a). All non-singleton OTUs are listed in Table 2 (singleton OTUs in Table S2). Representative sequences of all OTUs are deposited to GenBank, under accession nos. MK614825–MK614865.
The accumulation curve of OTUs did not level off, which was to be expected since only 32.1% of the root tips in the root system was analysed (Fig. 2b). The fungal diversity, assessed by Shannon’s H, varied consistently throughout the root system, with the older parts of the root system (R3 and R4) having higher scores than the younger parts (R1 and R2, Table 1).
The majority of OTUs of the sequenced root tips belonged to Basidiomycota (85.3%) whilst 13.9% belonged to Ascomycota. Most of the sequenced root tips (84.6%) that were included in the analyses had taxonomic affinity to known EcM fungi. More than half (58.0%) of the sequenced root tips belonged to the family Thelephoraceae, and the second most common family was Cortinariaceae (10.3%), followed by Russulaceae (6.2%). OTU1 (Pseudotomentella sp.) was the most abundant OTU throughout the root system, and more common in the younger part (R1) of the root system than would be expected by chance (Pearson’s χ2 test, χ2 = 40.825, df = 3, p < 0.001). OTU3 (Cortinarius sp. 1) was more abundant in R3 and R4 than expected by chance (Pearson’s χ2 test, χ2 = 25.244, df = 3, p < 0.001), and OTU4 (Russula subrubens) was more widespread in the oldest part (R4) than in an expected distribution (Pearson’s χ2 test, χ2 = 25.244, df = 3, p < 0.001). OTU12, although only represented by ten successfully sequenced root tips, had a top hit to the Geoglossaceae, Sarcoleotia globosa (99.9% similarity), which is not a known root associate, but still included in downstream analysis (see “Discussion”). This OTU was only found in the older parts of the root system (R3 and R4, Pearson’s χ2 test, χ2 = 14.665, df = 3, p = 0.002).
To investigate the spatial structure and compositional variation of fungi in the root system, ordination diagrams of OTU/fine root matrices were constructed, where each sample represents one fine root (as sites). DCA and GNMDS axes were strongly correlated with Kendall’s τ rank correlation test for all ordination methods, and in all ordination diagrams, point configurations were similar (Fig. S1), suggesting a robustly structured dataset (Son and Halvorsen 2014). Hence, we only focus on the GNMDS constructed using power-weighting (w = 0.5), which was the GNMDS with the lowest stress values (stress = 0.122). The ordination diagram showed a clear spatial structuring of the root-associated fungal community throughout the root system (Fig. 3 ), where fine roots from the different age classes clustered together. GNMDS1 represented to a large extent the length of the rhizome, as shown by the high correlation of position of fine roots along the rhizome with axis 1 (Fig. 3 ; Table S3). Fine roots from R1 were located at one end of the axis, and a shift from this part throughout to R4 was observed. Significant variables fitted onto the ordination diagrams showed that diversity was strongly positively correlated with the position of fine roots along the rhizome (proxy for age) and GNMDS1; short smooth exploration type (sh_smooth) was negatively correlated with GNMDS1. This indicate that this exploration type was more common in the younger part of the root system, whilst contact exploration type (contact) was positively correlated with position, indicating that this was more common in the older parts of the root system. The structure along GNMDS2 was mainly driven by richness.
GNMDS ordination diagram of power-weighted data (w = 0.5) for all analysed root tips, where fine roots are used as sites. The diagram is showing a gradient from the younger (R1) part of the root system through to the older (R4). Arrows depict significant variables fitted onto the ordination diagrams using envar function in the vegan package (Oksanen et al. 2013). Position = position of fine roots along the rhizome, diversity = Shannon’s H calculated per fine roots, richness = richness per fine root, length = relative length of fine roots, branching = number of branches per fine root, no_root_tip = number of root tips per fine root. Assigned exploration types are the following: contact, short smooth (sh_smooth), medium smooth (med_smooth), and additionally, hydrophilic for hydrophilic exploration types
To see whether root tips of the same OTU tended to cluster in the root system, a Bray–Curtis dissimilarity matrix of all successfully sequenced root tips and a distance matrix of their relative position in the root system were used to compare differences in mean dissimilarity between neighbouring and non-neighbouring root tips using Welch two-sided T test. The differences between dissimilarities in neighbouring and non-neighbouring root tips were significant (Welch two-sided T test, t = − 12.920, df = 515.620, p < 0.001) when analysing root system as a whole. The mean value of adjacent root tips was estimated to 0.53, meaning that almost half of the neighbouring root tips were identified to the same OTU, whilst for non-neighbours, the mean dissimilarity was estimated to 0.82, meaning that most pairs of root tips throughout the root system were dissimilar. The different parts of the root system were subjected to the same test, to see whether this pattern varied throughout the root system. The parts R1 and R3 had a significant different mean of dissimilarities in neighbouring and non-neighbouring (Welch two-sided T test, t = 11.569, df = 329.900, p < 0.001, and t = 4.759, df = 135.590, p < 0.001, respectively), whilst in R2 and R4, the difference was not significant (Welch two-sided T test, t = 1.727, df = 9.289, p = 0.117 and t = 1.213, df = 39.843, p = 0.232, respectively). There were probably not enough observations in R2 to infer any statistical significance (only 17 successfully sequenced root tips). From R1 through R4, there was an increase in estimated mean values for neighbouring and non-neighbouring root tips, indicating that it was more likely to be located next to root tips assigned to the same OTU in the younger of the root system than in the old (i.e. more clustering; Fig. 4 ). Clustering of root tips belonging to the same OTU is visualised schematically in Fig. 5 .
The proportion of pairs of root tips sharing the same OTU per age class (R1, R2, R3 and R4). Shared OTU between pairs of root tips is shown in dark grey, and pairs of root tips not sharing OTU is shown in light grey. The proportion of shared OTUs between pairs of root tips is larger in R1 and declines throughout the age classes
Schematic map of the entire root system the B. vivipara individual. R1–4 are the four parts the rhizome was dissected into and subsequently used as age classes in the analyses of the root system. Fine roots and branches are scaled in lengths relative to each other based on photographic measurements, but the spatial distance between fine roots and root tips are not realistically depicted in the map. Root tip colour indicate OTUs as described in the legend; light grey root tips were not identified
Discussion
Spatial structure within the root system
The ordination analyses showed a clear spatial structure within the root system for the community recovered from fine roots from different parts of the rhizome, with a shift in the fungal community from the youngest end of the rhizome to the older. This pattern was evident although only 32.1% of the total number of root tips was included in this analysis. Neither weighting functions nor ordination method (DCA or GNMDS) affected the overall structure; therefore, it is reasonable to believe that the structure found was not due to mathematical artefacts caused by weighting function or ordination method (Son and Halvorsen 2014). Previous studies have not found any clear structural patterns at finer scales (< 3 m) for the fungal communities of B. vivipara (Mühlmann et al. 2008; Blaalid et al. 2012, 2014; Yao et al. 2013; Botnen et al. 2014), except that Blaalid (2012) observed a spatial auto-correlation below 50-cm scale. In other studies, spatial heterogeneity and patchiness of fungal communities at finer scales have been observed (Taylor and Bruns 1999; Tedersoo et al. 2003; Lilleskov et al. 2004; Pickles et al. 2012; Blaalid et al. 2014). Several studies have found that some fungal species exhibit preference for certain microhabitats in the soil (Taylor and Bruns 1999; Tedersoo et al. 2003, 2008; Genney et al. 2006; Buée et al. 2007). However, for EcM species, the scale that species are occupying as root tips may be very different from the scale species are occupying as mycelium (Genney et al. 2006; Pickles et al. 2012). Yoshida et al. (2014) revealed a tendency towards a spatial clustering of individual EcM species within the root systems of T. diversifolia seedlings, but the present study is, to our knowledge, the first time, a clear spatial structuring at such a fine scale of the root-associated fungal community is shown for a full grown plant.
In addition to a shift in the overall root-associated fungal community throughout the root system, richness and diversity of the fungal community also changed throughout the root system. For the EcM fungi, exploration type recovered from fine roots also changed from the younger to the older end of the rhizome. The higher abundance of the contact exploration type found in association with the older end of the rhizome was mainly because OTU4 (Russula pascua); the only OTU assigned to this exploration type was more abundant in this part of the root system.
Spatial distribution of OTUs within the root system
From the χ2 tests, the structuring of the fungal community was evident at a scale ranging from rhizome parts to individual OTUs. OTU1 (Pseudotomentella sp.), although common throughout the whole root system, was more common in the youngest part of the root system than expected by a random distribution. Although at very different spatial scales, Pseudotomentella has been reported to be observed in younger rather than older forest stands (Kyaschenko et al. 2017), suggesting that it may be a weak competitor. OTU3 (Cortinarius sp.) and OTU4 (Russula subrubens) were relatively more abundant in R3 and R4. There might be a succession with age of the root system of root-associated fungi within this root system, similar to what is observed at larger scales.
Priority effects?
Priority effects have been observed for several species (Alford and Wilbur 1985; Shorrocks and Bingley 1994), including fungi (Kennedy and Bruns 2005; Yoshida et al. 2014; Hiscox et al. 2015). For EcM fungi, the priority effect may be observed as clustering of species within a root system, and a dominance of one or a few species, when the timing of arrival of species is unknown (Yoshida et al. 2014). Although only relative distances are recorded in this study, root tips located in sequence on a branch from a fine root were assumed to be closer in proximity to each other than root tips not located on the same branch or on other fine roots. With these assumptions, it was possible to investigate whether or not the neighbourhood of a root tip included several root tips sharing the same OTU (i.e. clusters). There was a higher mean of neighbouring root tips sharing the same OTU than non-neighbouring root tips, indicating clustering of species, and this was particularly evident in the younger part of the rhizome where more than half of all neighbouring root tips shared OTUs (Fig. 3). The mean number of neighbouring root tips sharing OTUs was much higher in the younger parts (R1 and R2) than in the older parts (R3 and R4). In the oldest part of the root system, almost none of the investigated neighbouring root tips shared an OTU (estimated mean = 0.9; Fig. 4).
According to priority effects, colonisation in the early stages of root formation may result in clusters of species. Clusters similar to what Yoshida et al. (2014) found in their study were observed only in the youngest parts of this particular root system, but in the older parts, no evident clustering of OTUs was observed. As freshly formed fine roots were only observed in the youngest part of the rhizome, we deem it likely that these clusters are a result of priority effects rather than competition. This was seen from the T test on strict neighbours, but also for the neighbourhood (i.e. branches on fine roots, data not shown). Similarly, Pickles et al. (2012) found that in a mature forest system, competition, or processes generating similar patterns, was determining the fine scale distribution of EcM fungi. It is possible that succession and competition play a more prominent role in structuring the fungal community in more established root systems compared to younger root systems, where more newly formed roots are available colonisation.
Richness and community composition
A total of 41 root-associated fungal OTUs were recovered from the analysed root system. Although only 32.1% of all root tips were successfully sequenced in the present study, this accords well with previous studies on root-associated fungi in B. vivipara using high throughput sequencing of entire root systems (e.g. Blaalid et al. 2012, 2014; Yao et al. 2013; Botnen et al. 2014), where ranges of root-associated OTUs generally fall between (10–159). However, as indicated by the accumulation curve, the root system most likely possesses a somewhat richer community than we were able to recover. Few other studies have been conducted where the entire mycobiome of whole root systems has been profiled root tip by root tip. Bahram et al. (2011) recovered 122 EcM fungal species in 103 root samples from an extensively sampled root system of a P. tremula individually, whilst Yoshida et al. (2014) recovered 40 EcM species when mapping the entire EcM community of three T. diversifolia seedlings.
The OTUs followed a log-normal distribution, where a few OTUs were common and the majority was represented by ten or less root tips (Fig. 2a). Similar to these results, Yoshida et al. (2014) observed that the majority of fungal species recovered from three seedlings of T. diversifolia occupied less than ten root tips, with only one or a few species dominating the root system of a seedling. Bahram et al. (2011) observed the same pattern, where a single species was found in 59.2% of all samples. Dominance of one or a few species seems to be common in root-associated fungal communities.
In accordance with earlier studies on the fungal community of B. vivipara (Massicotte et al. 1998; Mühlmann et al. 2008; Brevik et al. 2010; Blaalid et al. 2012, 2014; Yao et al. 2013; Botnen et al. 2014), most of the detected OTUs were EcM fungi. The two most dominant orders, Thelephorales and Agaricales, are well established as EcM-forming linages (Smith & Read 2008; Tedersoo et al. 2010). In contrast to the abovementioned studies of B. vivipara, only typical EcM root tips or tips with signs of fungal colonisation were investigated, and a dominance of EcM fungi is therefore not surprising. The order Thelephorales accounted for more than half of all the successfully sequenced root tips. Thelephorales is found to be among the most common orders in several studies in arctic and alpine EcM communities (e.g. Clemmensen & Michelsen 2006; Mühlmann et al. 2008; Bjorbækmo et al. 2010; Timling et al. 2012; Blaalid et al. 2012; Yao et al. 2013; Botnen et al. 2014; Morgado et al. 2015; Mundra & Halvorsen 2015), suggesting that several species within this order tolerate harsh climatic and environmental conditions. All of the most common OTUs, Pseudotomentella sp. (OTU1), Thelephorales sp.1 (OTU2), Cortinarius sp. (OTU3, OTU15, OTU23) and Russula (OTU4) were ectomycorrhizal.
Due to problems with contaminants, a strict criterion for including OTUs in downstream analysis was used: OTUs with hits to all yeasts and/or fungi that had not previously been reported from plant roots were excluded with two exceptions: OTU12 (99.9% match to Sarcoleotia globosa) and OTU19 (99.0% match to Clitocybe subditopoda). Clitocybe is generally considered a saprotrophic genus, OTU19 clustered together with ITS sequences with affinity to Tricholoma, a known EcM genus, in the initial NJ tree, and Clitocybe subidtopoda has been found to be unrelated to the core Clitocybe group (Alvarado et al. 2015). Hence, it might be a root-associated fungus. S. globosa within Geoglossales forms small, stipitate fruit bodies, and due to its small size, most likely overlooked in field surveys. Its distribution has previously been described as arctic and sub-arctic, although stretching as far south as cool temperate (Jumpponen et al. 1997). It is often found in open vegetation and on disturbed ground (Schumacher and Sivertsen 1987; Jumpponen et al. 1997), which fits the sample site of the B. vivipara root system used in this study. It has been suggested to have saprotrophic lifestyle, however, in a survey of stable isotopes, a relatively high enrichment in δ15N was shown for species within Geoglossales (Griffith et al. 2002). Additionally, several collections have been made where S. globosa has been found in association with mosses or the ericaceous Empetrum nigrum (Schumacher and Sivertsen 1987). Extreme isotopic profiles for δ15N and δ13C and associations with plants may suggest a possibility of unknown biotrophic lifestyles (also discussed in Tedersoo et al. 2010).
Conclusion
For the first time, an attempt to map the entire root-associated fungal community of an adult EcM plant was conducted. Although the direct PCR approach was not suitable for recovering the entire fungal community associated with this plant, and only a third of the root-associated fungal community could be identified, a diverse and spatially structured fungal community was discovered. Indications of possible priority effects were restricted to the younger parts of the root system, whereas in the older parts, we did not observe these patterns, suggesting that competition may be more important in structuring mature root systems. This study provides a first glimpse into the fine scale structuring of the root-associated fungal community of an adult EcM plant.
References
Agerer R (2001) Exploration types of ectomycorrhizae: a proposal to classify ectomycorrhizal mycelial systems according to their patterns of differentiation and putative ecological importance. Mycorrhiza 11:107–114. https://doi.org/10.1007/s005720100108
Agerer & Rambold (2004-2018) DEEMY—an information system for characterization and determination of Ectomycorrhizae.www.deemy.de –München, Ger
Alford R, Wilbur H (1985) Priority effects in experimental pond communities: competition between Bufo and Rana. Ecology 66:1097–1105
Alva V, Nam S-Z, Söding J, Lupas AN (2016) The MPI bioinformatics toolkit as an integrative platform for advanced protein sequence and structure analysis. Nucleic Acids Res 44:W410–W415. https://doi.org/10.1093/nar/gkw348
Alvarado P, Moreno G, Vizzini A, Consiglio G, Manjón JL, Setti L (2015) Atractosporocybe, Leucocybe and Rhizocybe : three new clitocyboid genera in the Tricholomatoid clade (Agaricales) with notes on Clitocybe and Lepista. Mycologia 107:123–136. https://doi.org/10.3852/13-369
Anderson IC, Genney DR, Alexander IJ (2014) Fine-scale diversity and distribution of ectomycorrhizal fungal mycelium in a Scots pine forest. New Phytol 201:1423–1430. https://doi.org/10.1111/nph.12637
Averill C, Turner BL, Finzi AC (2014) Mycorrhiza-mediated competition between plants and decomposers drives soil carbon storage. Nature 505:543–545. https://doi.org/10.1038/nature12901
Bahram M, Peay KG, Tedersoo L (2015) Local-scale biogeography and spatiotemporal variability in communities of mycorrhizal fungi. New Phytol 205:1454–1463. https://doi.org/10.1111/nph.13206
Bahram M, Põlme S, Kõljalg U, Tedersoo L (2011) A single European aspen (Populus tremula) tree individual may potentially harbour dozens of Cenococcum geophilum ITS genotypes and hundreds of species of ectomycorrhizal fungi. FEMS Microbiol Ecol 75:313–320. https://doi.org/10.1111/j.1574-6941.2010.01000.x
Bergemann SE, Miller SL (2002) Size, distribution, and persistence of genets in local populations of the late-stage ectomycorrhizal basidiomycete, Russula brevipes. New Phytol 156:313–320. https://doi.org/10.1046/j.1469-8137.2002.00507.x
Bjorbækmo M, Carlsen T, Brysting A, Vrålstad T, Høiland K, Ugland K, Geml J, Schumacher T, Kauserud H (2010) High diversity of root associated fungi in both alpine and arctic Dryas octopetala. BMC Plant Biol 10:244. https://doi.org/10.1186/1471-2229-10-244
Blaalid R, Carlsen T, Kumar S et al (2012) Changes in the root-associated fungal communities along a primary succession gradient analysed by 454 pyrosequencing. Mol Ecol 21:1897–1908. https://doi.org/10.1111/j.1365-294X.2011.05214.x
Blaalid R, Davey ML, Kauserud H, Carlsen T, Halvorsen R, Høiland K, Eidesen PB (2014) Arctic root-associated fungal community composition reflects environmental filtering. Mol Ecol 23:649–659. https://doi.org/10.1111/mec.12622
Bogar LM, Peay KG (2017) Processes maintaining the coexistence of ectomycorrhizal fungi at a fine spatial scale. In: Tedersoo L (ed) Biogeography of mycorrhizal symbiosis. Springer International Publishing, Cham, pp 79–105
Botnen S, Vik U, Carlsen T, Eidesen PB, Davey ML, Kauserud H (2014) Low host specificity of root-associated fungi at an arctic site. Mol Ecol 23:975–985. https://doi.org/10.1111/mec.12646
Brevik A, Moreno-Garcia J, Wenelczyk J (2010) Diversity of fungi associated with Bistorta vivipara (L.) Delarbre root systems along a local chronosequence on Svalbard. Agarica 29:15–26
Brundrett MC, Tedersoo L (2018) Evolutionary history of mycorrhizal symbioses and global host plant diversity. New Phytol 220:1108–1115. https://doi.org/10.1111/nph.14976
Buée M, Courty PE, Mignot D, Garbaye J (2007) Soil niche effect on species diversity and catabolic activities in an ectomycorrhizal fungal community. Soil Biol Biochem 39:1947–1955. https://doi.org/10.1016/j.soilbio.2007.02.016
Carlsen TA (2002) Molecular diversity of root endophytes in an alpine Bistorta vivipara—Kobresia myosuroides tundra plant community (Cand. Scient. Thesis). University of Oslo
Clemmensen KE, Bahr A, Ovaskainen O, Dahlberg A, Ekblad A, Wallander H, Stenlid J, Finlay RD, Wardle DA, Lindahl BD (2013) Roots and associated fungi drive long-term carbon sequestration in boreal forest. Science 339:1615–1618. https://doi.org/10.1126/science.1231923
Clemmensen KE, Finlay RD, Dahlberg A, Stenlid J, Wardle DA, Lindahl BD (2015) Carbon sequestration is related to mycorrhizal fungal community shifts during long-term succession in boreal forests. New Phytol 205:1525–1536. https://doi.org/10.1111/nph.13208
Clemmensen KE, Michelsen A (2006) Integrated long-term responses of an arctic–alpine willow and associated ectomycorrhizal fungi to an altered environment. Can J Bot 84:831–843. https://doi.org/10.1139/b06-039
Clymo RS (1980) Preliminary survey of the peat-bog Hummel Knowe moss using various numerical methods. Vegetatio 42:129–148
Dahlberg A, Stenlid J (1994) Size, distribution and biomass of genets in populations of Suillus bovinus (L.: Fr.) Roussel revealed by somatic incompatibility. New Phytol 128:225–234. https://doi.org/10.1111/j.1469-8137.1994.tb04006.x
Diggle PK (1997) Extreme preformation in alpine Polygonum viviparum: an architectural and developmental analysis. Am J Bot 84:154–169. https://doi.org/10.2307/2446077
Eriksen M, Bjureke KE, Dhillion SS (2002) Mycorrhizal plants of traditionally managed boreal grasslands in Norway. Mycorrhiza 12:117–123. https://doi.org/10.1007/s00572-002-0165-x
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes—application to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118. https://doi.org/10.1111/j.1365-294X.1993.tb00005.x
Genney DR, Anderson IC, Alexander IJ (2006) Fine-scale distribution of pine ectomycorrhizas and their extramatrical mycelium. New Phytol 170:381–390. https://doi.org/10.1111/j.1469-8137.2006.01669.x
Glassman SI, Peay KG, Talbot JM, Smith DP, Chung JA, Taylor JW, Vilgalys R, Bruns TD (2015) A continental view of pine-associated ectomycorrhizal fungal spore banks: a quiescent functional guild with a strong biogeographic pattern. New Phytol 205:1619–1631. https://doi.org/10.1111/nph.13240
Griffith GW, Easton GL, Jones AW (2002) Ecology and diversity of waxcap (Hygrocybe spp.) fungi. Bot J Scotl 54:7–22. https://doi.org/10.1080/03746600208685025
Hesselman H (1900) Om mykorrhizabildningar hos arktiske väkster. Bihang till Svenska Vetenskaps- Akademiens handlinger 26:1–46
Hill MO, Gauch HG (1980) Detrended correspondence analysis: an improved ordination technique. Vegetatio 42:47–58. https://doi.org/10.1007/BF00048870
Hiscox J, Savoury M, Müller CT, Lindahl BD, Rogers HJ, Boddy L (2015) Priority effects during fungal community establishment in beech wood. ISME J 9:2246–2260. https://doi.org/10.1038/ismej.2015.38
Jarvis S, Woodward S, Alexander IJJ, Taylor A, FSFS (2013) Regional scale gradients of climate and nitrogen deposition drive variation in ectomycorrhizal fungal communities associated with native Scots pine. Glob Chang Biol 19:1688–1696. https://doi.org/10.1111/gcb.12178
Jonsell B (ed) (2000) Flora Nordica 1. Ber, Stockholm
Jumpponen A, Weber NS, Trappe JM, Cázares E (1997) Distribution and ecology of the ascomycete Sarcoleotia globosa in the United States. Can J Bot 75:2228–2231. https://doi.org/10.1139/b97-933#.W78K7PmYOM8
Kauserud H, Kumar S, Brysting AK, Nordén J, Carlsen T (2012) High consistency between replicate 454 pyrosequencing analyses of ectomycorrhizal plant root samples. Mycorrhiza 22:309–315. https://doi.org/10.1007/s00572-011-0403-1
Kendall MG (1938) A new measure of rank correlation. Biometrika 30:81–93. https://doi.org/10.1093/biomet/30.1-2.81
Kennedy PG, Bruns TD (2005) Priority effects determine the outcome of ectomycorrhizal competition between two Rhizopogon species colonizing Pinus muricata seedlings. New Phytol 166:631–638. https://doi.org/10.1111/j.1469-8137.2005.01355.x
Kennedy PG, Higgins LM, Rogers RH, Weber MG (2011) Colonization-competition tradeoffs as a mechanism driving successional dynamics in ectomycorrhizal fungal vommunities. PLoS One 6:e25126. https://doi.org/10.1371/journal.pone.0025126
Kennedy PG, Peay KG, Bruns TD (2009) Root tip competition among ectomycorrhizal fungi: are priority effects a rule or an exception? Ecology 90:2098–2107. https://doi.org/10.1890/08-1291.1
Kõljalg U, Nilsson RH, Abarenkov K, Tedersoo L, Taylor AFS, Bahram M, Bates ST, Bruns TD, Bengtsson-Palme J, Callaghan TM, Douglas B, Drenkhan T, Eberhardt U, Dueñas M, Grebenc T, Griffith GW, Hartmann M, Kirk PM, Kohout P, Larsson E, Lindahl BD, Lücking R, Martín MP, Matheny PB, Nguyen NH, Niskanen T, Oja J, Peay KG, Peintner U, Peterson M, Põldmaa K, Saag L, Saar I, Schüßler A, Scott JA, Senés C, Smith ME, Suija A, Taylor DL, Telleria MT, Weiss M, Larsson KH (2013) Towards a unified paradigm for sequence-based identification of fungi. Mol Ecol 22:5271–5277. https://doi.org/10.1111/mec.12481
Kruskal JB (1964) Nonmetric multidimensional scaling: a numerical method. Psychometrika 29:115–129. https://doi.org/10.1007/BF02289694
Kruskal WH (1958) Ordinal measures of association. J Am Stat Assoc 53:814. https://doi.org/10.2307/2281954
Kyaschenko J, Clemmensen KE, Hagenbo A, Karltun E, Lindahl BD (2017) Shift in fungal communities and associated enzyme activities along an age gradient of managed Pinus sylvestris stands. ISME J 11:863–874. https://doi.org/10.1038/ismej.2016.184
Lilleskov EA, Bruns TD, Horton TR et al (2004) Detection of forest stand-level spatial structure in ectomycorrhizal fungal communities. FEMS Microbiol Ecol 49:319–332. https://doi.org/10.1016/j.femsec.2004.04.004
Lilleskov EA, Bruns TD (2003) Root colonization dynamics of two ectomycorrhizal fungi of contrasting life history strategies nutrient patches. New Phytol 159:141–151. https://doi.org/10.1046/j.0028-646x.2003.00794.x
Lindahl BD, Ihrmark K, Boberg J, Trumbore SE, Högberg P, Stenlid J, Finlay RD (2007) Spatial separation of litter decomposition and mycorrhizal nitrogen uptake in a boreal forest. New Phytol 173:611–620. https://doi.org/10.1111/j.1469-8137.2006.01936.x
Massicotte HB, Melville LH, Peterson RL, Luoma DL (1998) Anatomical aspects of field ectomycorrhizas on Polygonum viviparum (Polygonaceae) and Kobresia bellardii (Cyperaceae). Mycorrhiza 7:287–292. https://doi.org/10.1007/s005720050194
Miyamoto Y, Sakai A, Hattori M, Nara K (2015) Strong effect of climate on ectomycorrhizal fungal composition: evidence from range overlap between two mountains. ISME J 9:1870–1879. https://doi.org/10.1038/ismej.2015.8
Morgado LN, Semenova TA, Welker JM, Walker MD, Smets E, Geml J (2015) Summer temperature increase has distinct effects on the ectomycorrhizal fungal communities of moist tussock and dry tundra in Arctic Alaska. Glob Chang Biol 21:959–972. https://doi.org/10.1111/gcb.12716
Mühlmann O, Bacher M, Peintner U (2008) Polygonum viviparum mycobionts on an alpine primary successional glacier forefront. Mycorrhiza 18:87–95. https://doi.org/10.1007/s00572-007-0156-z
Mundra S, Bahram M, Tedersoo L, Kauserud H, Halvorsen R, Eidesen PB (2015a) Temporal variation of Bistorta vivipara-associated ectomycorrhizal fungal communities in the high arctic. Mol Ecol 24:6289–6302. https://doi.org/10.1111/mec.13458
Mundra S, Halvorsen R, Kauserud H, Müller E, Vik U, Eidesen PB (2015b) Arctic fungal communities associated with roots of Bistorta vivipara do not respond to the same fine-scale edaphic gradients as the aboveground vegetation. New Phytol 205:1587–1597. https://doi.org/10.1111/nph.13216
Oksanen J, Blanchet F, Kindt R, et al (2013) Vegan: community ecology package
Peay KG, Kennedy PG, Bruns TD (2011) Rethinking ectomycorrhizal succession: are root density and hyphal exploration types drivers of spatial and temporal zonation? Fungal Ecol 4:233–240. https://doi.org/10.1016/j.funeco.2010.09.010
Pickles BJ, Genney DR, Anderson IC, Alexander IJ (2012) Spatial analysis of ectomycorrhizal fungi reveals that root tip communities are structured by competitive interactions. Mol Ecol 21:5110–5123. https://doi.org/10.1111/j.1365-294X.2012.05739.x
Core Team R (2017) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Schumacher T, Sivertsen S (1987) Sarcoleotia globosa (Sommerf.: Fr.) Korf, taxonomy, ecology and distribution. In: Laursen GA, Ammirati JF, Redhead SA (eds) Arctic and alpine mycology II. Springer US, Boston, MA, pp 163–176
Shorrocks B, Bingley M (1994) Priority effects and species coexistence: experiments with fungal-breeding Drosophila. J Anim Ecol 63:799. https://doi.org/10.2307/5257
Smith SE, Read D (2008) Mycorrhizal symbiosis, 3rd edn. Elsevier, London
van Son TC, Halvorsen R (2014) Multiple parallel ordinations: the importance of choice of ordination method and weighting of species abundance data. Sommerfeltia 37:1–37. https://doi.org/10.2478/som-2014-0001
Sterkenburg E, Clemmensen KE, Ekblad A, Finlay RD, Lindahl BD (2018) Contrasting effects of ectomycorrhizal fungi on early and late stage decomposition in a boreal forest. ISME J 12:2187–2197. https://doi.org/10.1038/s41396-018-0181-2
Talbot JM, Bruns TD, Taylor JW, Smith DP, Branco S, Glassman SI, Erlandson S, Vilgalys R, Liao HL, Smith ME, Peay KG (2014) Endemism and functional convergence across the North American soil mycobiome. Proc Natl Acad Sci 111:6341–6346. https://doi.org/10.1073/pnas.1402584111
Taylor DL, Bruns TD (1999) Community structure of ectomycorrhizal fungi in a Pinus muricata forest: minimal overlap between the mature forest and resistant propagule communities. Mol Ecol 8:1837–1850. https://doi.org/10.1046/j.1365-294x.1999.00773.x
Tedersoo L, Hallenberg N, Larsson K (2003) Fine scale distribution of ectomycorrhizal fungi and roots across substrate layers including coarse woody debris in a mixed forest. New Phytol 159:153–165. https://doi.org/10.1046/j.0028-646x.2003.00792.x
Tedersoo L, May TW, Smith ME (2010) Ectomycorrhizal lifestyle in fungi: global diversity, distribution, and evolution of phylogenetic lineages. Mycorrhiza 20:217–263. https://doi.org/10.1007/s00572-009-0274-x
Tedersoo L, Suvi T, Jairus T, Kõljalg U (2008) Forest microsite effects on community composition of ectomycorrhizal fungi on seedlings of Picea abies and Betula pendula. Environ Microbiol 10:1189–1201. https://doi.org/10.1111/j.1462-2920.2007.01535.x
Tenenbaum JB (2000) A global geometric framework for nonlinear dimensionality reduction. Science 290:2319–2323. https://doi.org/10.1126/science.290.5500.2319
Timling I, Dahlberg A, Walker D et al (2012) Distribution and drivers of ectomycorrhizal fungal communities across the North American Arctic. Ecosphere 3(11):1–25
van der Linde S, Suz LM, Orme CDL, Cox F, Andreae H, Asi E, Atkinson B, Benham S, Carroll C, Cools N, de Vos B, Dietrich HP, Eichhorn J, Gehrmann J, Grebenc T, Gweon HS, Hansen K, Jacob F, Kristöfel F, Lech P, Manninger M, Martin J, Meesenburg H, Merilä P, Nicolas M, Pavlenda P, Rautio P, Schaub M, Schröck HW, Seidling W, Šrámek V, Thimonier A, Thomsen IM, Titeux H, Vanguelova E, Verstraeten A, Vesterdal L, Waldner P, Wijk S, Zhang Y, Žlindra D, Bidartondo MI (2018) Environment and host as large-scale controls of ectomycorrhizal fungi. Nature 558:243–248. https://doi.org/10.1038/s41586-018-0189-9
van der Maabel E (1979) Transformation of cover-abundance values in phytosociology and its effects on community similarity. Vegetatio 39:97–114. https://doi.org/10.1007/BF00052021
White TJ, Bruns TD, Lee SB, Taylor JW (1990) Analysis of phylogenetic relationships by amplification and direct sequencing of ribosomal RNA genes. In: Innis M, Gelfand D, Sninsky J, White T (eds) PCR protocols: a guide to methods and applications. Academic Press, New York, pp 315–322
Yao F, Vik U, Brysting AK, Carlsen T, Halvorsen R, Kauserud H (2013) Substantial compositional turnover of fungal communities in an alpine ridge-to-snowbed gradient. Mol Ecol 22:5040–5052. https://doi.org/10.1111/mec.12437
Yoshida N, Son JA, Matsushita N et al (2014) Fine-scale distribution of ectomycorrhizal fungi colonizing Tsuga diversifolia seedlings growing on rocks in a subalpine Abies veitchii forest. Mycorrhiza 24:247–257. https://doi.org/10.1007/s00572-013-0535-6
Acknowledgments
We would like to acknowledge Cecilie Mathiesen for an excellent effort in preparation of samples for sequencing, Rune Halvorsen for valuable input on ordination methods and Erlend Y. Fines for assistance in graphical design and excellent drawing.
Funding
This project was funded by University of Oslo.
Author information
Authors and Affiliations
Contributions
All authors have contributed to the completion of the article. The main ideas and design of the study were done by TC, HK and ET. Sampling and laboratory work were conducted by mainly ET, but also ABA and UV. ABA contributed substantially in ideas on analyses of the data. Drafting and writing of the manuscript was mainly performed by ET, but all authors (ABA, UV, AKB, IS, TC, HK) contributed substantially to supervision, ideas and discussion of the results in the writing process.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 58 kb)
Fig. S1
Multiple parallel ordinations (MPO) using DCA (A, B, C) and GNMDS (D, E, F) with raw (A, D), power-weighted (w=0.5, B, E) and presence/absence (C, F) data using fine roots as sites. All ordination methods and weighting functions show similar point configuration, supporting a strong gradient in the dataset. Arrows depicts significant variables fitted onto the ordination diagrams using envar function in the vegan package (Oksanen et al. 2013). position=postion of fine root along the rhizome, diversity=Shannon’s H calculated per fine root, richnes=richness per fine root, length=relative length of fine root, branch=number of branches per fine root, no_root_tip=number of root tips per fine root, no_exp_types=number of exploration type per fine root. Assigned exploration types are: contact, short smooth (sh_smooth), medium smooth (med_smooth) and additionally, hydrophilic for hydrophilic exploration types (PNG 323 kb)
Fig. S2
Photograph of one of the fine roots from the B. vivipara root system, showing how root tips are growing from fine roots and branches from the main fine roots. Because the real orientation of the main fine roots and branches in the soil is unknown, we assigned the root tips to three distance classes; neighbour, adjacent and non-neighbour. We assumed that, for instance, A and B were located in very close proximity in the soil (i.e. strict neighbours), and that C, on the same branch, is closer to A and B (i.e. adjacent) than it was to D, which was most likely located further away from A, B and C (i.e. non-neighbour) (PNG 1808 kb)
Rights and permissions
About this article
Cite this article
Thoen, E., Aas, A.B., Vik, U. et al. A single ectomycorrhizal plant root system includes a diverse and spatially structured fungal community. Mycorrhiza 29, 167–180 (2019). https://doi.org/10.1007/s00572-019-00889-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00572-019-00889-z