Abstract
Hepatozoon sp. are parasites that commonly infect frogs and arthropod vectors. This species has variability in the morphological and morphometric characteristics. Due to these variations, the naming of the species is thus impaired and only by visualizing the sporogonic cycle in vector and by molecular studies this problem can be solved. Recently, the use of molecular genetics has helped the species denomination. In this work, we collected 145 frogs (68 Leptodactylus chaquensis and 77 Leptodactylus podicipinus) in different sampling sites, where were found 18 (26.47 %) L. chaquensis and 24 (31.17 %) L. podicipinus parasitized; besides of gamonts, schizogonic forms were also seen in animals organs. The positivity difference between the collection sites for both frog species was not significant (p = 0.958). Comparing gamonts found in each species of anuran, we observed differences in morphology. The comparison in the molecular level for L. podicipinus was not possible due to small amount of blood obtained, just L. chaquensis had their parasites DNA sequenced. The amplified and sequenced samples, named HEP1 to HEP10, are presented in the phylogenetic tree as a different branch from other haemogregarines described on other hosts. Therefore, we have seen that, although the morphology and morphometry of the collected parasites at each site showed differences, the sequencing of these samples revealed identical species of Hepatozoon, and different compared to those from GenBank, thereby demonstrating that the species of Hepatozoon in L. chaquensis observed in this study probably represent a new species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The anuran amphibians have a complex life cycle that can be affected by many factors throughout its existence and can host ectoparasites and endoparasites. Among hemoparasites found, we can highlight the group of the haemogregarines, among them species of the Hepatozoon genus (Barta and Desser 1984).
This genus shows so diverse morphology in the blood of frogs that creates confusion among researchers when naming their found species, and different names were given to the same protozoan (Levine 1988; Desser et al. 1995; Smith 1996; Desser 2001). Costa et al. (1973) observed schizogonic forms in organs of Leptodactylus latrans and Leptodactylus pentadactylus and sporogonic forms in invertebrate (Haementeria lutzi). Desser et al. (1995) were the first to investigate the complete cycle of parasite development in vertebrates (Lithobates catesbeianus) and the sporogonic cycle in the vector (Culex territans). Posteriorly to this study, researchers such as Kim et al. (1998) and Harkness et al. (2010) used the experimental infections and reported morphological, morphometric, and molecular comparisons.
Regarding to the problem of identifying the haemogregarines found in anurans, several researchers have proposed the molecular analysis of 18S ribosomal RNA (rRNA) sequences (conserved regions) (Haag et al. 1998; Martin et al. 2002) and ITS (variable regions) to infer phylogenetic relationships among these protozoan (Schlegel 1991; Cai et al. 1992; Maia da Silva et al. 2004; Rodrigues et al. 2006; Ferreira et al. 2007).
Our goal was to determine the occurrence of Hepatozoon sp. in frogs Leptodactylus chaquensis and Leptodactylus podicipinus, from the region of the Pantanal (Brazil), and characterize the parasites by morphology, morphometry, and molecular analysis, in order to infer the phylogenetic relationships among the samples, as well as among the different species already described in the available databases.
Material and methods
The capture of specimens was performed under the approval of the Ethics Committee on Animal Experiments (CEEA) no. 1608, located at the Biosciences Institute, São Paulo State University (UNESP), and authorized by the Brazilian Institute of Environment and Renewable Natural Resources IBAMA, no. 16696-1.
We collected 145 adults individuals, 68 L. chaquensis and 77 L. podicipinus. The animals were captured in the Pantanal Study Base (BEP), located in Passo do Lontra, municipality of Corumbá – Mato Grosso Sul State, Brazil (19° 34′ 39″ S, 57° 0′ 44″ W) and in the EMBRAPA Research Center (18° 59′ 00″ S, 56° 39′ 00″ W), in Nhumirim farm, located in the subregion of Nhecolândia, also in the municipality of Corumbá. These collection sites are located at a distance of approximately 76 km apart one of other. The subregion of Passo do Lontra included in the Paraguay River basin, located at the lower portion of the Miranda River subbasin, under influence of the Abobral River, with rainy season from November to March (Polizer et al. 2000).
Nhecolândia is characterized by a mosaic of saline and freshwater lagoons, interspersing cordilleras with forest vegetation, connected by natural channels that receive water only during major floods, delimiting lanes of Brazilian savannah and grassland vegetation (Rodela 2006). The region has two distinct seasons, the rainy, from November to March, and the dry, from April to October (Soriano and Alves 2005).
L. chaquensis is a very abundant species in open areas in the Pantanal and surrounding plateau, both males and females measure 7.1 cm, whereas L. podicipinus, also abundant in Pantanal, males measure only 3.5 cm and females 3.9 cm. (Uetanabaro et al. 2008).
The capture was performed manually at night, in April, November, and December 2008, and January 2009, and all animals were packed in plastic bags and transferred to the laboratory. Subsequently, the animals were euthanized with a sodium thiopental injection; blood was collected from the heart to make the blood smears and the molecular analysis. The animals were necropsied and organs like the liver, spleen, and kidney were used to make the organs imprint, in order to visualize schizogonic stages of Hepatozoon sp.
All blood samples were frozen for molecular characterization by PCR. When it was not possible to collect blood, as in the case of very small specimens of L. podicipinus, the slides were scraped (Scopel et al. 2004; Sykes et al. 2008) and washed with lysis buffer (Qiagen) and the DNA extraction was performed normally. Blood smears and organs imprinting were prepared, fixed in methanol, and stained with Giemsa (10 %) for 30 min. The slides were examined under an optical microscope (×1000 magnification) for the diagnosis of the parasite.
The morphology and the morphometry of the parasite were obtained by using the image analyzer software Qwin lite 2.5 (Leica). The observed morphometric variables of the parasite were the area (BA), length (BL), and width (BW) of its body, as well the area (NA), length (NL), and width (NW) of its nucleus.
The extraction of total DNA was performed by using the commercial kit QIAmp DNA Blood Kit (Qiagen) according to the manufacturer’s instructions. The amplification of fragments of DNA samples was performed by means of HEMOI oligonucleotides 5′-TAT TGG GAA TAA TTT TTA CTA ATT TGA TTG-3′ and HEMOII 3′-CCT TTA CTT CTT CTC AGT AAG GTT GAT CAC-5′ (Perkins and Keller 2001) targeting the 18S rRNA region. The reactions were prepared for a final volume of 20 μl containing 10 μl of GoTaq Colorless Master Mix (Gotaq® DNA polymerase, GoTaq Colorless 2× Reaction buffer, pH 8.5, 400 μM dATP, 400 μM dGTP; 400 μM dCTP, 400 μM dTTP, and 3 mM MgCl2) (Promega), 10 pmol of each primer (forward and reverse), 1 μl of total DNA and H2O. All reactions were performed using a negative control.
Reactions were carried out using My Cycler (BioRad) thermocycler at the following conditions: initial cycle at 94 °C (hot start) for 3 min followed by 40 cycles of 94 °C for 45 s, 56 °C for 1 min and 72 °C for 1 min, with a final extension cycle at 72 °C for 7 min. The generated products were visualized on 1 % agarose gel stained with GelRed 0.5 μg/ml (Biotium). The amplified fragments were compared with the 1 kb DNA Ladder (Fermentas) used as standard and viewed in UVDI transluminator (Major Science). Positive samples showed approximately 900 bp in size, which were subjected to sequencing. Among the positive samples, ten were sequenced and compared to the sequences of the GenBank using the BLASTn software (http://www.ncbi.nlm.nih.gov/BLAST). The complete alignment of sequences was performed using the ClustalX (1.83) software. Methods of distance (neighbor-joining (NJ)) were used to construct the phylogenetic tree (Saitou and Nei 1987), and the bootstrap test was used to estimate the analysis consistency index of the branches distances of the NJ trees (Felsenstein 1985).
For the statistical analysis, we used the EPI-INFO software to perform the chi-squared test. The level of significance was set at p < 0.05. In this test, we verified whether there was a significant difference between the positivity of the species of frogs and their capture sites and also if there were significant differences of positivity between the species of frogs. For determination of variation of standard deviation in the morphology parameters of the parasitic species found in the frogs, we used the following formula (Costa Neto 1994): cv = 100σ / X, in which cv is the coefficient of variation, σ is the standard deviation, and X is the mean.
Results
Collection sites
In the first collection site (BEP), 73 frogs were captured (38 L. chaquensis and 35 L. podicipinus), while at Nhumirim farm, 72 frogs were captured (30 L. chaquensis and 42 L. podicipinus), totaling 145 animals captured, 68 L. chaquensis and 77 L. podicipinus.
Positivity for Hepatozoon spp.
The positivity of the animals in relation to Leptodactylus species and collection site can be seen in Table 1.
Of the 68 captured L. chaquensis, 8 (11.76 %) were positive for smear microscopy of blood or organs, 4 from BEP and 4 from Nhumirim, while for PCR technique, 18 animals (26.47 %) were positive, 13 from BEP and 5 from Nhumirim. Among the Hepatozoon sp. samples positive for blood smears technique, three were false-negative, i.e., were negative by PCR even having gamonts and schizonts typically seen in Hepatozoon sp.
Among the 77 L. podicipinus captured, 2 (2.60 %) were positive for smear microscopy of blood or organs, both collected at Nhumirim, while for PCR, 24 (31.17 %) were positive, 8 from BEP and 16 from Nhumirim. Among the positive samples on slides, only one was false-negative.
Collection sites and positive animals for PCR
In BEP 73, animals were collected, of which 21 (28.77 %) were positive for Hepatozoon sp., 13 (34.21 %) L. chaquensis, and 8 (22.86 %) L. podicipinus.
In Nhumirim, 72 animals were collected, of which 21 (29.16 %) were positive for Hepatozoon sp., 5 (16.67 %) L. chaquensis, and 16 (38 %) L. podicipinus (Table 1).
For PCR, statistical analysis showed that significant differences were not found when considering both frogs’ collection site (χ 2 = 0.003, p = 0.958), considering only L. chaquensis collection site (χ 2 = 2.651, p = 0.104), and considering only L. podicipinus collection site (χ 2 = 2.066, p = 0.151).
Considering the factor “used analysis techniques,” for both anurans species, statistically significant difference was found in the positivity between PCR and microscopy (χ 2 = 23.995, p < 0.001).
Morphological characterization of Hepatozoon sp.
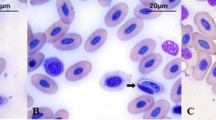
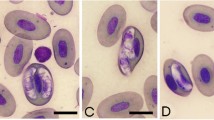
The gamonts presented intraerythrocytic and extraerythrocytic characteristics. There were variable forms, some more rounded in with the nucleus taking the entire width of the gamont, and others with rounded ends, and sometimes the nucleus was dislocated towards the periphery, with nuclear chromatin ranging between fragmented and condensate. The gamonts showed format varying from “sausage” to the globular, with the coloration of basophilic cytoplasm and nucleus varying from blue-gray to pink. In most of the erythrocytes, the nucleus has been displaced to the periphery in the presence of the parasite (Figs. 1 and 2). Gamonts enlarging the host erythrocytes were not found. Means and standard deviations of morphometric parameters of gamonts are shown in Table 2. The morphometric analysis of gamonts showed that the coefficient of variation was relatively higher in L. chaquensis than in L. podicipinus.
For 145 captured animals, a total of 133 organ imprints (spleen, liver, and kidney) were performed, 51 of L. chaquensis and 82 of L. podicipinus. Schizonts were found in only four animals (3 %), three L. chaquensis (5.88 %), which presented three schizonts in the liver and ten in the spleen, and one in L. podicipinus (1.22 %), with one schizont in the liver. Four L. chaquensis (7.84 %) presented gamonts in the organs, six in the liver, one in the spleen, and one in the kidney, whereas in L. podicipinus, gamonts were observed in two animals (2.44 %), one in the spleen and one in the kidney.
The schizonts were found in immature and mature phases of the cycle. Mature schizonts and merozoites were found in small number in these organs (Fig. 3a, b), where we could see mature schizonts with formed merozoites (Fig. 3a), and schizonts with merozoites radially arranged around a residual body (Fig. 3b). The immature schizonts showed a “double line” around and contained coarse-looking material with no evident nucleus (Fig. 3c, d). As schizonts found were in different developmental stages, their measures were quite variable. In L. chaquensis, they measure 297.23 ± 169.1 μm2 (121.2–574.7) of area, whereas the unique schizont found in L. podicipinus has 168.8 μm2 of area.
Molecular characterization
The use of HepF and HepR primers, in this study, was not able to amplify the sequences of Hepatozoon sp. for the size of the desired fragment of 625 bp, despite of many attempts changing the temperature and concentration of the PCR reaction products. Only the HEMOI and HEMOII oligonucleotides were efficient in amplification. The sequencing of all ten samples was conducted only for L. chaquensis. Due to lack of sufficient DNA in sample, the sequencing of L. podicipinus samples was not possible.
After their appropriate patterning, the use of oligonucleotides HEMOI and HEMOII (Perkins and Keller 2001) gave us products with expected size of approximately 900 bp.
Phylogenetic analysis
All Hepatozoon sp. samples were amplified and sequenced (HEP1 to HEP10) generated sequences with 99 % of bootstrap. All samples (HEP1 to HEP10, GenBank sequence accession JX987775) were presented in the phylogenetic tree as a different branch from other haemogregarines described on other hosts. Samples identified in frogs showed more similarity to Hepatozoon sp. Boiga (unpublished) and are on a separate branch of other sequences described as Hepatozoon ayorgbor n. sp., Hepatozoon sp. DG1, Hepatozoon sp. BV1, and in a more distant branch of Hepatozoon cf. catesbianae (Fig. 4). In Table 3, there are the species of Hepatozoon with their intermediate hosts, used in the construction of the phylogenetic tree.
Discussion
The intraerythrocytic gamonts of L. chaquensis and L. podicipinus showed cytological and morphological changes that agree to the results of other studies performed worldwide (Fantham et al. 1942; Mohammed and Mansour 1966; Costa et al. 1973; Kim et al. 1998; Smith et al. 2000; Desser 2001; Leal et al. 2009). The gamonts of L. chaquensis are smaller than those reported by Costa et al. (1973), who obtained longer and wider gamonts. However, in relation to L. podicipinus, the size of gamonts was closest. We emphasize that the frogs studied by Costa et al. (1973) were Leptodactylus latrans and Leptodactylus pentadactylus.
The standard deviation between gamonts in two frogs species of our study showed that L. chaquensis gamonts were more varied in size and had lower size than those of L. podicipinus. A similar result was also observed in a previous study (Leal et al. 2009) with the same frog species. This variation may be attributable to the different developmental stages of the gamonts simultaneously present in the blood; this fact has been reported by Hull and Camin (1960) and Clark and Bradford (1969). This variability of gamont size was also observed by Smith et al. (1994) in Nerodia sipedon sipedon, parasitized with Hepatozoon sipedon, whose reptiles were exhibiting in your bloodstream many gamonts in the immature stage.
Leptodactylus chaquensis from BEP had more homogeneous forms of gamonts with predominance of medium size (Fig. 1a, b), and those caught in Nhumirim farm showed more variation, with small, medium, and large forms (Fig. 1c, d). It was not possible to affirm that they represent the same Hepatozoon species, despite phylogenetic analysis.
Using microscopic analysis, it was not possible to observe gamonts in L. podicipinus collected in BEP. The gamonts observed in L. podicipinus caught at Nhumirim farm were homogeneous in their morphology and presented basophilic nucleus and cytoplasm.
For the collection site factor, the statistical analysis showed no significant difference in positivity of L. chaquensis and L. podicipinus infections with Hepatozoon sp., despite the ecological differences between the two sites, but there are other factors that were not taken into account in this study that may have influenced this positivity, such as the density of vectors or sexual difference of host (Bardsley and Harmsen 1973).
The scraping of blood smears was already used by some researchers, as Scopel et al. (2004) and Sykes et al. (2008), and demonstrated a lower sensitivity in comparison to whole-blood DNA extraction.
Statistical analysis of the false-positive results found in this work showed that there is no significant evidence that slide scraping is a factor that leads to the occurrence of false-negatives results (χ 2 = 0.325, p = 0.569). In this case, the corrected value of χ 2 was used due to the presence of classes with frequencies lower than 5.
Anurans can be infected with many Hepatozoon species and can act also as parathenic hosts for others species (Smith 1996). Viana et al. (2012) proved that L. chaquensis is a parathenic host for Hepatozoon caimani. Cystozoites of H. caimani were identified in fresh liver impression smears of L. chaquensis after the ingestion with sporulated oocysts from laboratory-bred Culex (Melanoconion) mosquitoes. In our study, no cystozoites were found in organs of L. chaquensis or L. podicipinus, so we cannot incriminate the studied specimens as parathenic hosts.
Developmental stages of Hepatozoon sp. were found in anurans tissues. Immature and mature schizonts were observed in the spleen and the liver of L. chaquensis and in the liver of L. podicipinus. Mansour and Mohammed (1966), Levine and Nye (1977), and Desser et al. (1995) also observed schizonts; however, these forms were only found in the liver. The mature forms of schizonts observed in this study are similar to those reported by Mohammed and Mansour (1966) and Levine and Nye (1977); however, the sizes reported by them are larger than we have found. Costa et al. (1973) reported mature schizonts relatively smaller than those of the present study.
The morphology and morphometry of our samples of L. chaquensis showed differences in gamonts and schizonts when compared to the literature; however, when analyzing the phylogenetic tree, we found a high degree of similarity among our samples, forming a separate branch. Phylogenetic analysis showed that the Hepatozoon species parasitizing L. chaquensis were identical, although these animals have been collected at points about 76 km apart one of other.
The use of HepF and HepR primers, in this study, was not able to amplify the desired sequences of Hepatozoon sp., agreeing to Moço et al. (2012), who used two pairs of primers (HEPF/HEPR and PIRO A1/PIRO B) to unsuccessfully detect Hepatozoon spp. in snakes. The success obtained with HEMOI and HEMOII oligonucleotides for amplification of fragments, agrees to O’Dwyer et al. (2013), who used added to HEMOI/HEMOII, the HepF300/Hep900 primers in combination, getting better results for amplifications of fragments of different isolated of Hepatozoon spp. in snakes.
It was also observed that the fragments (HEP1 to HEP10) are very close to those isolated of Boiga irregulares, an animal from Australia, whose fragments generated of haemogregarine, when compared with our samples, revealed to have little substitution between them. The study of Jakes et al. (2003), with H. boigae of these same reptile species, showed intraerythrocytic gamonts that did not distended the cell of host, but causing displacement of the erythrocyte nucleus, and the infected erythrocytes that stained more clearly than the nonparasitized cells. The gamonts observed in our study were not preferentially erythrocytic, and there was no whitening of the cell in the presence of the parasite. Hepatozoon ayorgbor n. sp., another species that is found in a branch near to our samples in the phylogenetic tree, presents some differences. In the study reported by Sloboda et al. (2007), the gamonts of H. ayorgbor n. sp. found in Phyton regius showed the body and the nucleus longer and narrower than those measured in our study. Sloboda et al. (2007) reported liver schizonts which were much smaller than those we found.
In comparison with Hepatozoon cf. catesbianae, a present species in Rana catesbeiana, our samples were positioned on a most distant branch and were quite different morphometrically. This morphological difference was also observed when comparing our results with those reported by Desser et al. (1995). The gamonts found by them were bigger, showed a more elongated shape, basophilic weakly stained cytoplasm, compact and central nucleus. The gamonts we observed were not as elongated, and the nucleus were quite different, which were not only compact and central. Only the cytoplasm of gamonts was stained in a similar manner to that reported by the authors.
In this study, both analyzed species of anurans had a high positivity of infection with Hepatozoon sp. under the PCR analysis technique and there was no significant difference between the positivity for the two collection sites.
All comparisons done among the species of Hepatozoon of this study and those reported in the literature showed that our samples are different morphologically, morphometrically, and genetically, thereby demonstrating that the species of Hepatozoon in L. chaquensis observed in this study can represent a new species.
References
Bardsley JE, Harmsen R (1973) The trypanosomes of anura. In: Dawes B (ed) Advance parasitology, vol 7. Academic Press, London, pp 1–73
Barta JR, Desser SS (1984) Blood parasites of amphibians from Algonquin Park, Ontario. J Wildl Dis 20(3):180–189. doi:10.7589/0090-3558-20.3.180
Cai JMD, Collins MAC, Donald V, Thompson DE (1992) PCR cloning and nucleotide sequence determination of the 18S rRNA genes and internal transcribed spacer 1 of the protozoan parasites Cryptosporidium parvum and Cryptosporidium muris. Biochim Biophys Acta 1131:317–320. doi:10.1016/0167-4781(92)90032-U
Costa SCG, Pessoa SB, Pereira NM, Colombo T (1973) The life history of Hepatozoon leptodactyli (Lesage, 1908) Pessoa, 1970—a parasite of the common laboratory animal—the frog of the genus Leptodactylus. Mem Inst Oswaldo Cruz 71(1/2):1–8. doi:10.1590/S0074-02761973000100001
Costa Neto PLO (1994) Estatística. Edgard Blucher LTDA, São paulo
Clark GW, Bradford J (1969) Blood parasites of some reptiles of the Pacific northwest. J Protozool 16:576–581. doi:10.1111/j.1550-7408.1969.tb.02316.x
Desser SS, Hong H, Martin DS (1995) The life history, ultrastructure, and experimental transmission of Hepatozoon catesbianae n. comb., an apicomplexan parasite on the bullfrog, Rana catesbeiana and the mosquito, Culex territans in Algoquin Park, Ontário. J Parasitol 81:212–222. doi:10.2307/3283922
Desser SS (2001) The blood parasites of anurans from Costa Rica with reflections on the taxonomy of their trypanosomes. J Parasitol 87(1):152–160. doi:10.1645/0022-3395(2001)087[0152:TBPOAF]2.0.CO;2
Fantham HB, Porter A, Richardson LR (1942) Some Haematozoa observed in vertebrates in earsten Canada. Parasitol 34:199–226. doi:10.1017/S0031182000016176
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evol 39(4):783–791. doi:10.2307/2408678
Ferreira RC, Campaner M, Viola LB, Takata CSA, Takeda GF, Teixeira MMG (2007) Morphological and molecular diversity and phylogenetic relationships among trypanosomes from the Amazonia, Atlantic Forest and Pantanal biomes in Brazil. Parasitol 19:1–16. doi:10.1017/S0331182007003058
Haag J, O’Huigin C, Overath P (1998) The molecular phylogeny of trypanosomes: evidence for an early divergence of the salivaria. Mol Biochem Parasitol 91:37–49. doi:10.1016/S0166-6851(97)00185-0
Harkness LM, Drohan AE, Dickson CM, Smith TG (2010) Experimental transmission of Hepatozoon clamatae (Apicomplexa: Adeleida) to the wood frog, Rana sylvatica, and to the mosquito Culex pipiens. J Parasitol 96(2):434–436. doi:10.1645/GE-2317.1
Hull RW, Camin JH (1960) Haemogregarines in snakes: The incidence of the erythrocytic stages. J Parasitol 46(4):515–523. doi:10.2307/3275151
Jakes K, O’Donoghue PJ, Cameron SL (2003) Phylogenetic relationship of Hepatozoon (Haemogregarina) boigae, Hepatozoon sp., Haemogregarina clelandi and Haemoproteus chelodina form Australian reptilies to other Apicomplexa based on cladistic analyses of ultrastructural and life-cycle characters. Parasitol 126:555–559. doi:10.1017/S0031182003003111
Kim B, Smith TG, Desser SS (1998) The life history and host specificity of Hepatozoon clamatae (Apicomplexa: Adeleorina) and ITS-1 nucleotide sequence variation of Hepatozoon species of frogs and mosquitoes from Ontario. J Parasitol 84:789–797. doi:10.2307/3284589
Leal DDM, O’Dwyer LH, Ribeiro VC, Silva RJ, Ferreira VL, Rodrigues RB (2009) Hemoparasites of the genus Trypanosoma (Kinetoplastida: Trypanosomatidae) and hemogregarines in anurans of the São Paulo and Mato Grosso do Sul States - Brazil. An Acad Bras Cienc 81(2):199–206. doi:10.1590/S0001-37652009000200006
Levine ND, Nye RR (1977) A survey of blood tissue parasites of leopard frogs Rana pipiens in the United States. J Wildl Dis 13:17–23. doi:10.7589/0090-3558-13.1.17
Levine ND (1988) The protozoan phylum Apicomplexa. CRC, Boca Raton, FL
Maia da Silva F, Rodrigues AC, Campaner M, Takata CS, Brigido MC, Junqueira AC, Coura JR, Takeda GF, Shaw JJ, Teixeira MM (2004) Randomly amplified polimorphic DNA analysis of Trypanosoma rangeli and allied species from human, monkeys and other sylvatic mammals of the Brazilian Amazon disclosed a new group and a species-specific marker. Parasitol 128:283–294. doi:10.1017/S003118203004554
Mansour NS, Mohammed AH (1966) Development of Haemogregarina pestanae in the toad Bufo regularis. J Protozool 13(2):265–269. doi:10.1111/j.1550-7408.1966.tb.01905.x
Martin DS, Wright ADG, Barta JR, Desser SS (2002) Phylogenetic position of the giant trypanosomes Trypanosoma chanttoni, Trypanosoma fallisi, Trypanosoma mega, Trypanosoma neveulemairei, and Trypanosoma ranarum inferred from 18S rRNA gene sequences. J Parasitol 88(3):566–571. doi:10.1645/0022-3395(2002)088[0566:PPOTGA]2.0.CO:2
Moço TC, Silva RJ, Madeira NG, Paduan KS, Rubini AS, Leal DDM, O’Dwyer LH (2012) Morphological, morphometric, and molecular characterization of Hepatozoon spp. (Apicomplexa, Hepatozoidae) from naturally infected Caudisona durissa terrifica (Serpentes, Viperidae). Parasitol Res 110:1393–1401. doi:10.1007/s00436-011-2639-2
Mohammed AH, Mansour NS (1966) Development of Haemogregarina boueti in the toad Bufo regularis. J Protozool 13(2):259–264. doi:10.1111/j1550-7408.1966.tb01904.x
O’Dwyer LH, Moço TC, Paduan KS, Spenassatto C, Silva RJ, Ribolla PEM (2013) Description of three new species of Hepatozoon (Apicomplexa, Hepatozoidae) from Rattlesnakes (Crotalus durissus terrificus) based on molecular, morphometric and morphologic characters. Exp Parasitol 135:200–207. doi:10.1016/j.exppara.2013.06.019
Perkins SL, Keller AK (2001) Phylogeny of Nuclear subunit rRNA genes of hemogregarines amplified with specific primers. J Parasitol 87(4):870–876. doi:10.1645/0022-3395(2001)087[0870:PONSSR]2.0.CO:2
Polizer M, Lastória G, Rondon MAC (2000) Características físicas da região do Passo do Lontra. In: I Simpósio Brasileiro de Recursos hídricos do Centro Oeste, 2000, Brasília: Distrito Federal
Rodela LG (2006) Unidades de Vegetação e pastagens nativas do Pantanal da Nhecolândia, Mato Grosso do Sul. 252 p. Tese de Doutorado. Universidade de São Paulo
Rodrigues AC, Paiva F, Campaner M, Stevens JR, Noyes HA, Teixeira MMG (2006) Phylogeny of Trypanosoma (Megatrypanum) theileri and related trypanosomes reveals lineages of isolates associated with artiodactyl hosts diverging on SSU and ITS ribossomal sequences. Parasitol 132:215–224. doi:10.1017/S0031182005008929
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4(4):406–425
Scopel KKG, Fontes CJF, Nunes ÁC, Horta MF, Braga ÉM (2004) Low sensitivity of nested PCR using Plasmodium DNA extracted from stained thick blood smears: an epidemiological retrospective study among subjects with low parasitaemia in an endemic area of the Brazilian Amazon region. Malar J 3:1–6. doi:10.1186/1475-2875-3-8
Schlegel M (1991) Protist evolution and phylogeny as discerned from small subunit ribossomal RNA sequence comparisons. Eur J Protistol 27:207–219. doi:10.1016/S0932-4739(11)80059-3
Sloboda M, Kamler M, Bulantová J, Votýpka J, Modrý DA (2007) A new species of Hepatozoon (Apicomplexa: Adeleorina) from Phyton regius (serpentes: Phytonidae) and its experimental transmission by a mosquito vector. J Parasitol 93(5):1189–1198. doi:10.1645/GE-1200R.1
Smith TG, Desser SS, Martim DD (1994) The development of Hepatozoon sipedon n. sp. (Apicomplexa: Adeleina: Hepatozoidae) in its natural host, the Northern water snake (Nerodia sipedon sipedon), culicine vectors, Culex pipiens and Culex territans, and an intermediate host, the Northern leopard frog (Rana pipiens). Parasitol Res 80(7):559–568. doi:10.1007/BF00933003
Smith TG (1996) The genus Hepatozoon (Apicomplexa: Adeleina). J Parasitol 82(4):565–585. doi:10.2307/3283781
Smith TG, Kim B, Hong H, Desser SS (2000) Intraerytrocytic development of species of Hepatozoon infecting ranid frogs: evidence for convergence of life cycle characteristics among apicomplexans. J Parasitol 86(3):451–458. doi:10.2307/3284856
Soriano BMA, Alves MJM (2005) Boletim agrometeorológico ano 2002 para a sub-região da Nhecolãndia, Pantanal, Mato Grosso do Sul. EMBRAPA Pantanal, Corumbá, Brasil
Sykes JE, Owens SD, Terry JC, Lindsay LL, Pusterla N (2008) Use of dried blood smears for detection of feline hemoplasmas using real-time polymerase chain reaction. J Vet Diagn Investig 20(5):616–620. doi:10.1177/1040638708020000513
Uetanabaro M, Almeida Prado CP, Rodrigues DJ, Gordo M, Campos Z (2008) Field guide to the anurans of the Pantanal and surrounding Cerrados. Universidade do Mato Grosso do Sul, Brasil
Viana LA, Soares P, Silva JE, Paiva F, Coutinho ME (2012) Anurans as paratenic host in the transmission of Hepatozoon caimani to caimans Caiman yacare and Caiman latirostris. Parasitol Res 110(2):883–886. doi:10.1007/s00436-011-2570-6
Acknowledgments
This study received financial support from Coordination for the improvement of Higher Education Personnel (CAPES) and Foundation for UNESP Development, Process No. 00237/10-DFP (FUNDUNESP). We gratefully acknowledge Dr. Vanda Lucia Ferreira, Associate Professor of the Biology Department, Biological Sciences and Health Center of Mato Grosso do Sul Federal University, for valuable technical assistance.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Menezes Leal, D.D., Dreyer, C.S., da Silva, R.J. et al. Characterization of Hepatozoon spp. in Leptodactylus chaquensis and Leptodactylus podicipinus from two regions of the Pantanal, state of Mato Grosso do Sul, Brazil. Parasitol Res 114, 1541–1549 (2015). https://doi.org/10.1007/s00436-015-4338-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-015-4338-x