Abstract
Purpose
The aim of this study was to investigate roles of microRNA (miR)-183 in pediatric acute myeloid leukemia (AML).
Methods
miR-183 expression in bone marrow and patients’ sera of childhood AML was detected by real-time quantitative PCR. Functions of miR-183 in malignant phenotypes of two leukemia cell lines were then evaluated. Additionally, putative targets of miR-183 were predicted using three miRNA target prediction algorithms and validated by luciferase reporter assay. Clinical relevance of miR-183 and its target gene were further determined.
Results
miR-183 expression in bone marrow and patients’ sera of childhood AML was both significantly higher than those in the corresponding normal controls (both P < 0.001). Enforced expression of miR-183 dramatically enhanced cell proliferation and G1/S transition, but inhibited cell apoptosis of leukemia cells. Bioinformatics prediction and luciferase reporter assay identified programmed cell death 6 (PDCD6) as a direct target gene of miR-183. Moreover, high serum miR-183 combined with low serum PDCD6 mRNA was significantly associated with French–American–British classification subtype M7 (P = 0.01) and unfavorable karyotypes (P = 0.006). Further multivariate analysis identified the combination of serum miR-183 and PDCD6 levels as an independent prognostic factor for both relapse-free and overall survivals. Functionally, re-introduction of PDCD6 markedly reversed the effects of miR-183 in cell cycle, proliferation and apoptosis of two leukemia cell lines.
Conclusion
Combined serum miR-183 and PDCD6 mRNA may serve as a novel prognostic biomarker for pediatric AML. Interestingly, miR-183 may function as an oncogene and may enhance cell proliferation by targeting PDCD6, implying a potential therapeutic target for this malignancy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Acute myeloid leukemia (AML), a heterogeneous group of leukemias, is characterized by abnormal proliferation of myeloid progenitor cells, leading to the insufficient generation of normal mature blood cells (Rubnitz et al. 2008). This clonal disease is caused by malignant transformation of a bone-marrow-derived, self-renewing stem cell or myeloid progenitor, which usually has a reduced rate of apoptosis as well as aberrant and limited, differentiation capacity (Ziai et al. 2015). Statistically, AML contributes to 15–20 % of acute leukemia cases in childhood and adolescents and also represents approximately 30 % of cancer-related deaths in this age range (Dewan and Agarwal 2015). Growing evidence shows that the clinical outcome of patients with pediatric AML may often vary by host factors such as age at diagnosis, gender, race–ethnicity, body mass index, genetic polymorphism, socioeconomic status, phenotypes, cytogenetics, morphology, white blood cell counts and response to therapy (Forthun et al. 2016). During the past decades, accumulating great advances have been performed to enhance the diagnostic and therapeutic strategies and to improve the clinical outcome of patients with pediatric AML. However, the diagnosis of this disease remains according to the onset of symptoms, which may often progress to be apparent at advanced stages (Huang et al. 2015). Therefore, it is of great clinical significance to identify novel biomarkers in patients’ sera and to develop less invasive and cheaper diagnostic and prognostic approaches and to uncover efficient therapeutic targets of this malignancy.
MicroRNAs (miRNAs), a group of small (about 18–25 nucleotides in length), noncoding, endogenous, functional RNAs exert the functions via regulating the expression of the corresponding target mRNAs at a posttranscriptional level (Yan et al. 2015). MiRNAs are incorporated into the RNA-induced silencing complex and bind to the 3’-untranslated region (UTR) of mRNA sequences with imperfect complementarity (Danen-van Oorschot et al. 2012). Functionally, miRNAs play multiple roles in cell development, differentiation, growth, survival, apoptosis, migration and invasion (Ma et al. 2016). Accumulating studies have also revealed that the alterations of miRNAs often lead to various human diseases, including cancers. MiR-183, together with miR-96 and miR-182, belongs to the miR-183 family, which has high homogeneity and is clustered within 2–4 kb at chromosome 7q32 (Dambal et al. 2015). Recent studies have observed the aberrant expression of miRNAs from this locus in various cancer types, including leukemia, breast cancer, liver cancer and colorectal cancer (Liu et al. 2013; Li et al. 2014; Leung et al. 2015; Huangfu et al. 2016). Among them, miR-183 has been revealed to function as either an oncogene or a tumor suppressor in diverse human cancers by regulating different target genes (Zhang et al. 2013). However, its function and underlying mechanisms in this malignancy remain unclear.
To address this problem, we here firstly detected the expression levels of miR-183 in bone marrow and patients’ sera of childhood AML by real-time quantitative PCR. Then, we evaluated the regulatory effects of miR-183 in cell cycle, proliferation and apoptosis of two leukemia cell lines (HL60 and K562) following the transfection of miR-183 mimics and inhibitors. In addition, putative targets of miR-183 were predicted using three miRNA target prediction algorithms (miRanda, TargetScan and miRWalk) and validated by luciferase reporter assay. Furthermore, the clinical relevance and functions of miR-183 and its target gene were also determined.
Materials and methods
Cell lines and culture
Two human leukemia cell lines HL60 and K562 were purchased from American Type Culture Collection (ATCC, Manassas, VA, USA) and were cultured in RPMI-1640 medium (Invitrogen, Carlsbad, CA, USA) supplemented with 10 % fetal bovine serum. Cells were cultured at 37 °C with 5.0 % CO2 for further use.
Patients and tissue samples
In this study, we used the same patient cohort with our previous studies (Wang et al. 2013; Bai et al. 2012). Our study was approved by Huai’an First People’s Hospital Ethics Committee. Prior informed consent was obtained from the patients for the collection of specimens in accordance with the guidelines of Huai’an First People’s Hospital, China. All specimens were handled and made anonymous according to the ethical and legal standards. Please see detail information on patients and tissue samples in supplementary file S1. The clinical characteristics of 106 patients with AML are summarized in Table 1.
Cell transfection
The transfection of human leukemia cells with miR-183/control mimic (miR-183/NC_mimics, Applied Biosystems, CA, USA), miR-183/control inhibitor (miR-183/NC_inhibitor, Applied Biosystems, CA, USA) and pcDNA3.1-PDCD6 expression vector (en-PDCD6, Invitrogen, Carlsbad, CA, USA) by Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) according to the user’s instruction. At 48 h after transfection, cells were harvested for Western blot or real-time quantitative RT-PCR analyses.
Western blot analysis
Proteins were extracted from leukemia cells after the transfection using radio-immunoprecipitation assay buffer. The cell lysates were separated by 10 % SDS-PAGE and transferred onto polyvinylidene difluoride membranes (Qiagen China Co., Ltd.). Then, the membranes were incubated with the primary antibodies: anti-PDCD6 (dilution 1:500, Abcam Inc, MA, USA) and anti-GAPDH (dilution 1:1000, Abcam Inc, MA, USA), after blocking with 8 % milk in phosphate-buffered saline (PBS; pH 7.5). In addition, the membranes were incubated with the appropriate horseradish peroxidase-conjugated secondary antibodies (dilution 1:1000, Abcam Inc, MA, USA) after the incubation at 4 °C overnight. Finally, protein bands were detected by using an enhanced chemiluminescence system (Amersham Bioscience, Piscataway, NJ).
Real-time quantitative RT-PCR
Expression levels of miR-183 and PDCD6 mRNA in AML patients’ sera and cells were detected by real-time quantitative RT-PCR according to the protocols described in our previous studies (Wang et al. 2013; Bai et al. 2012). RNU6B and GAPDH were, respectively, used as internal reference genes for miR-183 and PDCD6. The primer sequences were as following: for miR-183: forward 5′-CTA TGG CAC TGG TAG AAT TCA CT-3′, reverse 5′-TCG TAT CCA GTG CAG GGT C-3′; for RNU6B: forward 5′-CGC TTC GGC AGC ACA TAT AC-3′, reverse 5′-TTC ACG AAT TTG CGT GTC AT-3′; for PDCD6 mRNA: forward 5′-GGA TGA TCG ATA AGA ACG AGC TGA A-3′, reverse 5′-ATG AGG ATG TCG TGG AAC TGG TC-3′; for GAPDH: forward 5′-ATG GAA ATC CCA TCA CCA TCT T-3′, reverse 5′-CGC CCC ACT TGA TTT TGG-3′. Relative quantifications of target miRNA and mRNA expression were evaluated using the comparative cycle threshold (CT) method (Schmittgen and Livak 2008). The relative expression level of miR-183 was obtained by the normalization to the values of U6. The relative expression level of PDCD6 mRNA was obtained by the normalization to the values of GAPDH. Each sample was examined in triplicate.
Cell proliferation assay
Cell proliferation of leukemia cells after the transfection were measured by using 3-(4,5-dimethylthiazol-2-yl)-2.5-diphenyl-2H-tetrazolium bromide (MTT) assays. In brief, cells were seeded at a density of 5.0 × 103 to 1.0 × 104 cells per well in 96-well plates. At 72 h after the transfection, fresh medium containing 10 % fetal bovine serum and 20 μl of MTT solution (5 μg/ml, Sigma-Aldrich, MA, USA) was added to each well, which was then incubated for an additional 4 h at 37 °C. Following the centrifugation for 10 min at 500×g, culture medium supernatant was removed from wells and the formazan was dissolved with dimethyl sulfoxide (DMSO). Finally, the amounts of MTT formazan generated were measured as absorbance using a microplate reader at 570 nm.
Cell cycle assay
Cell cycle phases of leukemia cells after the transfection were analyzed by measuring the DNA fragment staining with propidium iodide (PI, Sigma-Aldrich Co.) based on the instructions to users. In brief, at 48 h after the transfection, leukemia cells were cultured in serum-free medium for 22 h, and then, fetal bovine serum was supplied for additional 2 h. After that, the cells were harvested and washed twice with cold phosphate-buffered saline, followed by the incubation with PI dye solution containing RNaseA (30 mg/mL) for 30 min at room temperature in the dark. After PI staining, leukemia cells were analyzed for the DNA content immediately by flow cytometry with FACS calibers (BD Biosciences, San Jose, CA, USA) using CellQuest software (version 3.3, BD Biosciences, San Jose, CA, USA). PI-negative living cells were gated for the cell cycle assay. Each sample was examined in triplicate.
Cell apoptosis assay
Cell apoptosis of leukemia cells after the transfection was subsequently detected using an annexin V apoptosis detection kit (BD Biosciences, San Jose, CA, USA) based on the instructions to users. In brief, leukemia cells were resuspended in 100 mL binding buffer and stained with 5 mL each of annexin V and PI staining solution for 30 min. After that, flow cytometry with FACS calibers (BD Biosciences, San Jose, CA, USA) was used to analyze the cell apoptosis using the CellQuest software (version 3.3, BD Biosciences, San Jose, CA, USA).
MiRNA target prediction
Putative target mRNAs of miR-183 were predicted by three bioinformatic algorithms, including TargetScan (http://www.targetscan.org/vert_71/, Release 7.1 June 2016) (Garcia et al. 2011), microRNA.org-Targets and Expression (http://www.microrna.org/microrna/home.do, Last Update: 2010-11-01) (Betel et al. 2008) and miRWalk (version 2.0, http://zmf.umm.uni-heidelberg.de/apps/zmf/mirwalk2/) (Dweep and Gretz 2015), which are commonly used in the field for the miRNA target prediction. To overcome the limitations of each program, the putative target genes predicted by all three algorithms were chose as the candidate target genes for further validations.
Luciferase reporter assay
To confirm the binding efficiency of miR-183 on the 3′UTR of PDCD6 mRNA, leukemia cells were co-transfected with miR-183 mimic or the negative control and psiCHECK-2-PDCD6-3′-UTR-WT, psiCHECK-2-PDCD6-MUT. At 48 h after the transfection, cells were collected and detected with the Dual-Luciferase Reporter Assay System (Promega, CA, USA). Luciferase activity was measured by the GloMax fluorescence reader (Promega, CA, USA).
Statistical analysis
All statistical analyses were performed using the software of SPSS (version 11.0, SPSS Inc, IL, USA). Data were shown as mean ± standard deviation (SD). The differences between groups were analyzed using Student’s t test when there were only two groups, or assessed by one-way analysis of variance (ANOVA) when more than two groups were analyzed. The correlation between miR-183 expression and PDCD6 mRNA expression in AML patients’ sera was determined by Spearman’s correlation analysis. Associations of serum miR-183 combined with PDCD mRNA levels with clinicopathological features of pediatric AML were performed using the χ2 test for categorical variables. The Kaplan–Meier survival curves were used to determine any significant relationships between serum miR-183 combined with PDCD mRNA levels, and the status of the patients with respect to relapse-free survival (RFS) or overall survival (OS). P values less than 0.05 were considered to be statistically significant.
Results
miR-183 expression upregulates in patients with childhood AML
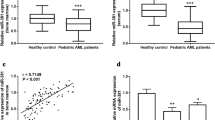
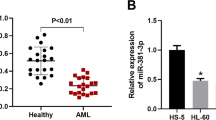
Expression levels of miR-183 (in bone marrow, AML vs normal: 2.95 ± 0.91 vs 1.57 ± 0.87, P < 0.001, Fig. 1a; in serum, AML vs normal: 2.82 ± 0.77 vs 1.46 ± 0.73, P < 0.001, Fig. 1b) in bone marrow and patients’ sera were both significantly higher than those in normal bone marrow and healthy sera. In addition, Spearman’s correlation analysis showed the closed correlations of miR-183 expression levels in the bone marrow of childhood AML patients with the corresponding expression in patients’ sera (Spearman’s correlation: r = 0.68, P = 0.01, Fig. 1c).
MiR-183 upregulation in bone marrow and patients’ sera of pediatric AML. a Expression levels of miR-183 in bone marrow of patients with pediatric AML and normal controls. b Expression levels of miR-183 in patients’ sera and healthy controls. c Correlations of miR-183 expression levels between the bone marrow and patients’ sera of pediatric AML
MiR-183 upregulation promotes cell proliferation of human leukemia cells
To elucidate the function of miR-183 in pathogenesis of pediatric AML, we here evaluated cell proliferation in two human leukemia cells HL60 and K562 in response to the transfection of miR-183 mimics and miR-183 inhibitor. As shown in Fig. 2a, b, the relative expression levels of miR-183 in leukemia cells transfected with miR-183 mimics were significantly increased compared with those transfected with control mimics (both P = 0.02), while its expression was dramatically downregulated by the transfection of miR-183 inhibitor (both P < 0.001).
MiR-183 upregulation promotes cell proliferation of human leukemia cells. a, b Expression levels of miR-183 in HL60 and K562 cells transfected with miR-183 mimics were significantly increased compared with those transfected with control mimics (both P = 0.02), while its expression in HL60 and K562 cells transfected with miR-183 inhibitor were significantly decreased (both P < 0.001); c, d MTT assay indicated that the transfection of miR-183 mimics significantly promoted cell proliferation in HL60 and K562 cells compared with normal controls (P < 0.01); in contrast, the loss of miR-183 by the specific inhibitor markedly suppressed cell proliferation of HL60 and K562 cells (P = 0.02 and 0.03, respectively)
Next, the results of MTT assay indicated that the transfection of miR-183 mimics significantly promoted cell proliferation in two leukemia cell lines (HL60 and K562) compared with normal controls (both P < 0.01, Fig. 2c, d). In contrast, the loss of miR-183 by the specific inhibitor markedly suppressed cell proliferation of two human leukemia cells HL60 and K562 (P = 0.02 and 0.03, respectively, Fig. 2c, d).
MiR-183 upregulation promotes G1/S transition and inhibits apoptosis of human leukemia cells
As shown in Fig. 3a, b, both HL60 and K562 cells that were transfected with miR-183 mimic displayed the significantly decreased ratio of G0-/G1-phase cells and significantly increased ratio of S-phase and G2-/M-phase cells, compared with the cells in the negative group (both P < 0.05). In contrast, the transfection of miR-183 inhibitor led to the increased cell population of G0-/G1-phase and the decreased cell population of the S-phase and G2-/M-phase of two leukemia cell lines (both P < 0.05).
MiR-183 upregulation promotes G1/S transition and inhibits apoptosis of human leukemia cells. a, b HL60 and K562 cells which were transfected with miR-183 mimic, respectively, display the significantly decreased ratio of G0-/G1-phase cells and significantly increased ratio of S-phase and G2-/M-phase cells, compared with the cells in the negative group (both P < 0.05). In contrast, the transfection of miR-183 inhibitor led to the increased cell population of G0-/G1-phase and the decreased cell population of the S-phase and G2-/M-phase of two leukemia cell lines (both P < 0.05). c, d Percentage of apoptotic HL60 and K562 cells, which were transfected with miR-183 mimic, were dramatically decreased (all P < 0.05), but those in miR-183 inhibitor transfection group were significantly increased, compared with the corresponding controls (all P < 0.05)
Moreover, the percentage of apoptotic HL60 and K562 cells that were transfected with miR-183 mimic were dramatically decreased (all P < 0.05, Fig. 3c, d), but those in miR-183 inhibitor transfection group were significantly increased, compared with the corresponding controls (all P < 0.05, Fig. 3c, d).
MiR-183 downregulates PDCD6 expression by directly targeting its 3′-UTR
We predicted the possible target genes of miR-183 by applying three miRNA target gene prediction systems and identified PDCD6 as one of the candidate genes, which were commonly predicted by three bioinformatics algorithms (Please see the list of candidate targets of miR-183 in Supplementary Table S1).
As shown in Fig. 4a, one putative binding site for miR-183 was found in the 3′-UTR of PDCD6 at 227–233 bp, which are highly conserved across species. To confirm the predictions, the luciferase reporter assay was performed in HL60 and K562 cells. The relative luciferase activities were dramatically reduced in cells co-transfected with the psiCHECK-2-PDCD6-3′-UTR-WT luciferase reporter and miR-183 mimics than those in the negative control cells. However, both inhibitory and enhanced effects were abolished when 3′-UTRs that contained both mutant-binding sites were co-transfected with miR-183 (Fig. 4b).
MiR-183 downregulates PDCD6 expression by directly targeting its 3′-UTR. a The putative miR-183-binding sequence in the 3′-UTR of PDCD6 mRNA; b luciferase report assay showed that the relative luciferase activities were dramatically reduced in cells co-transfected with the psiCHECK-2-PDCD6-3′-UTR-WT luciferase reporter and miR-183 mimics than those in the negative control cells. However, both inhibitory and enhanced effects were abolished when 3′-UTRs that contained both mutant-binding sites were co-transfected with miR-183; c, d Enforced expression of miR-183 remarkably inhibited the endogenous expression of PDCD6 mRNA and protein in HL60 and K562 cells, respectively. e, f Loss of miR-183 increased the expression of endogenous PDCD6 at both the mRNA and protein levels in HL60 and K562 cells, respectively. The relative expression level of PDCD6 mRNA was obtained by the normalization to the values of GAPDH. Each sample was examined in triplicate. *P < 0.05, comparison with NC-mimics and NC-inhibitor groups
Next, the enforced expression of miR-183 remarkably inhibited the endogenous expression of PDCD6 mRNA and protein in HL60 and K562 cells (Fig. 4c, d). Consistently, the loss of miR-183 increased the expression of endogenous PDCD6 at both the mRNA and protein levels in HL60 and K562 cells (Fig. 4e, f). Our results indicate that both of the binding sites may be involved in miR-183 regulation, and miR-183 directly binds to the 3′-UTR of PDCD6 to regulate its expression.
These findings provide the evidence that PDCD6 may be a direct target of miR-183 in human leukemia cells.
Serum miR-183 in patients with pediatric AML negatively correlates with serum PDCD6 mRNA
The above data indicated that the serum levels of miR-183 (AML vs normal: 2.82 ± 0.77 vs 1.46 ± 0.73, P < 0.001) in patients with pediatric AML were significantly higher than those in healthy sera. In contrast, the expression levels of PDCD6 mRNA in pediatric AML patients’ sera were significantly lower than those in healthy sera (AML vs normal: 1.62 ± 0.88 vs 3.24 ± 1.03, P < 0.001, Fig. 5a). Then, Spearman’s correlation analysis showed the negative correlation between serum miR-183 and PDCD6 mRNA in patients with pediatric AML (Spearman’s correlation: r = −0.556, P < 0.001, Fig. 5b).
Serum miR-183 in patients with pediatric AML negatively correlates with serum PDCD6 mRNA. a Expression levels of PDCD6 mRNA in pediatric AML patients’ sera were significantly lower than those in healthy sera (AML vs normal: 1.62 ± 0.88 vs 3.24 ± 1.03, P < 0.001); the relative expression level of PDCD6 mRNA was obtained by the normalization to the values of GAPDH. Each sample was examined in triplicate. b Spearman’s correlation analysis showed the negative correlation between serum miR-183 and PDCD6 mRNA in patients with pediatric AML (Spearman’s correlation: r = −0.556, P < 0.001)
Associations of the combined serum miR-183 and PDCD6 mRNA levels with various clinical characteristics of patients with pediatric AML
The median values of serum miR-183 (2.67) and PDCD6 mRNA (1.71) levels were used as cutoff points to divide all 106 patients with pediatric AML into miR-183-low/miR-183-high (n = 52/54) and PDCD6-low/PDCD6-high (n = 55/51) expression groups. Among 106 patients, 13 (12.26 %) cases were both high expression of miR-183 and PDCD6, 14 (13.21 %) cases were both low expression of miR-183 and PDCD6, 41 (38.68 %) cases were miR-183-high and PDCD6-low expressions, and 38 (35.85 %) cases were miR-183-low and PDCD6-high expressions.
Table 1 summarizes the associations of the combined serum miR-183 and PDCD6 mRNA levels with various clinical characteristics of patients with pediatric AML. High serum miR-183 combined with low serum PDCD6 mRNA was significantly associated with French–American–British classification subtype M7 (P = 0.01, Table 1) and unfavorable karyotypes (P = 0.006, Table 1). There were no significant associations between their serum levels and patients’ gender and age, leukocyte count, extramedullary disease and day 7 response to treatment (all P > 0.05, Table 1).
Prognostic values of the combined serum miR-183 and PDCD6 mRNA levels in patients with pediatric AML
Kaplan–Meier curve analyses revealed that pediatric AML patients with high serum miR-183 combined with low serum PDCD6 mRNA levels had the shortest RFS and OS in all four groups according to the combination expression of miR-183/PDCD6 (P = 0.001, Fig. 6). Moreover, univariate analyses found that RFS and OS of patients with pediatric AML were both significantly associated with the French–American–British classification subtype M7 (both P = 0.01, Table 2), unfavorable cytogenetic abnormalities (both P < 0.001, Table 2) and miR-183-high/PDCD6-low level (both P = 0.01, Table 2). Furthermore, Cox proportional hazards multivariate analyses of the univariate predictors identified the cytogenetic abnormalities (P = 0.01 and 0.009, respectively, Table 3) and miR-183-high/PDCD6-low level (both P = 0.03, Table 3) as independent prognostic factors for RFS and OS.
MiR-183 promotes cell proliferation in human leukemia cells via regulating PDCD6
To determine whether miR-183 promoted cell proliferation of human leukemia cells via regulating its target gene PDCD6, pcDNA3.1-PDCD6 expression vector was transfected into HL60 and K562 cells. As shown in Fig. 7a, b, the endogenous expression levels of PDCD6 protein were detected by Western blot in HL60 and K562 cells transfected with the miR-183 mimic in the presence of PDCD6 or vector control for 24 h. Next, re-introduction of PDCD6 markedly reversed the enhancing effects of cell proliferation in the miR-183-expressing cells (both P < 0.05, Fig. 7c for HL60 and Fig. 7d for K562), which was similar to the findings in cell cycle and cell apoptosis assays (all P < 0.05, Fig. 8).
MiR-183 promotes cell proliferation in human leukemia cells via regulating PDCD6. a, b Endogenous expression levels of PDCD6 protein were detected by western blot in HL60 and K562 cells transfected with the miR-183 mimic in the presence of PDCD6 or vector control for 24 h. c, d Re-introduction of PDCD6 markedly reversed the enhancing effects of cell proliferation in the miR-183-expressing HL60 and K562 cells
MiR-183 promotes G1/S transition and inhibits cell apoptosis in human leukemia cells via regulating PDCD6. a, b Re-introduction of PDCD6 markedly reversed the effects of miR-183 overexpression on cell cycle of HL60 and K562 cells, respectively. c, d Re-introduction of PDCD6 markedly reversed the effects of miR-183 overexpression on cell apoptosis of HL60 and K562 cells, respectively
Discussion
The identification of reliable and stable diagnostic and prognostic markers and efficient therapeutic targets for pediatric AML, as a complex and heterogeneous malignancy, remains a significant clinical challenge. In the current study, we demonstrated that miR-183 was upregulated in sera of patients with pediatric AML, and the re-expression and the knockdown of miR-183, respectively, promoted and suppressed AML cell proliferation in vitro. Moreover, PDCD6, a direct and functional target of miR-183, was also identified. Its expression was frequently decreased in sera of patients with pediatric AML, and the combination of high miR-183 expression and low PDCD6 expression was remarkedly associated with aggressive clinicopathological features and unfavorable prognosis in patients with pediatric AML. Furthermore, we found that miR-183 functionally promoted cell proliferation by silencing PDCD6. Taken together, these findings suggest that the dysregulation of miR-183-PDCD6 axis may play a crucial role during the progression of pediatric AML and may be a novel potential therapeutic target for the treatment of this malignancy.
MiR-183 behaves as an oncomiRNA, and its elevated level has been observed in various human cancers. Miao et al. (2016) revealed that miR-183 enhanced the proliferation, invasion and metastasis of human pancreatic adenocarcinoma cells; Lowery et al. (2010) found that miR-183 dysregulation was correlated with estrogen receptor and HER2/neu receptor expression, and the induced overexpression of miR-183 inhibited migration of breast cancer cells; Gu et al. (2014) indicated that miR-183 suppressed apoptosis and promoted proliferation and invasion of gastric cancer cells by targeting PDCD4; Qiu et al. (2014) showed that miR-183 efficiently increased cell proliferation, migration and invasion via targeting protein phosphatase 2A in renal cancer cells. In contrast, a recent study of Fan et al. (2016) reported that miR-183 also exerted the tumor suppressive roles in cervical cancer and inhibited cell invasion and metastasis through targeting MMP-9. In pediatric AML, the current study confirmed the increased expression levels of miR-183 in bone marrow and patients’ sera compared to the corresponding normal controls. We also determined the diagnostic and prognostic potentials of miR-183 in this malignancy. To further explore its functions, we in the present study demonstrated that the overexpressed miR-183 promoted cell proliferation and G1/S transition and inhibited apoptosis of human leukemia cell lines, expanding our understanding on the pathophysiology of pediatric AML.
Since miRNAs can act as oncogenes or tumor suppressors depending on the function of the target genes, we identified the downstream target genes for miR-183 in order to elucidate its underlying molecular mechanisms involved into malignant progression of pediatric AML. Combined with bioinformatics algorithms of miRNA target prediction and luciferase reporter assay, PDCD6 was identified as a candidate target of miR-183. As a calcium-binding modulator protein, PDCD6 functions as a regulator of cell proliferation and death (Hashemi et al. 2015; Zhou et al. 2015). Its aberrant expression has also been observed in a variety of human cancers, and it may exert either an oncogenic role or a tumor suppressive role with a cancer-dependent manner. For example, PDCD6 has been found to be highly expressed in metastatic ovarian cancer cells and positively regulated cell migration and invasion (Su et al. 2012). In contrast, the decreased copy numbers and mRNA expression of PDCD6 were both found in gastric cancer specimens and also were identified as significant prognostic biomarkers for advanced patients (Yoon et al. 2012). In the present study, our clinical data showed that PDCD6 mRNA expression was dramatically downregulated in patients’ sera of pediatric AML. Its downregulation combined with miR-183 upregulation was found to be associated with French–American–British classification subtype M7 and unfavorable karyotypes. More interestingly, pediatric AML patients with miR-183-high/PDCD6-low expression had the shortest RFS and OS of four groups (miR-183-low/PDCD6-high, miR-183-low/PDCD6-low, miR-183-high/PDCD6-high and miR-183-high/PDCD6-low). Of note, the multivariate survival analysis showed that the combined aberrant expression of miR-183 and PDCD6 was an independent prognostic factor for both RFS and OS in pediatric AML. Furthermore, we speculated a novel mechanism in which the re-expression of miR-183 may directly downregulate PDCD6, resulting in the promotion of AML cell proliferation and G1/S transition and the inhibition of cell apoptosis. This is the first study to provide evidence that as a critical cell proliferation inducer, miR-183 may directly regulated PDCD6 at the posttranscriptional level and contributes to its oncogenic function of AML progression.
In conclusion, the dysregulation of miR-183-PDCD6 axis may be involved into the aggressive progression of pediatric AML. Combined serum miR-183 and PDCD6 mRNA may serve as a novel prognostic biomarker for this malignancy. Interestingly, miR-183 may function as an oncogene that may enhance AML cell proliferation by targeting PDCD6, implying a potential therapeutic target for pediatric AML.
References
Bai J, Guo A, Hong Z et al (2012) Upregulation of microRNA-100 predicts poor prognosis in patients with pediatric acute myeloid leukemia. Onco Targets Ther 5:213–219
Betel D, Wilson M, Gabow A et al (2008) The microRNA.org resource: targets and expression. Nucleic Acids Res 36(Database issue):D149–D153
Dambal S, Shah M, Mihelich B, Nonn L (2015) The microRNA-183 cluster: the family that plays together stays together. Nucleic Acids Res 43:7173–7188
Danen-van Oorschot AA, Kuipers JE, Arentsen-Peters S et al (2012) Differentially expressed miRNAs in cytogenetic and molecular subtypes of pediatric acute myeloid leukemia. Pediatr Blood Cancer 58:715–721
Dewan K, Agarwal K (2015) Acute lymphoblastic leukemia with normal platelet count. Cancer Transl Med 1:181–182
Dweep H, Gretz N (2015) miRWalk2.0: a comprehensive atlas of microRNA-target interactions. Nat Methods 12:697
Fan D, Wang Y, Qi P et al (2016) MicroRNA-183 functions as the tumor suppressor via inhibiting cellular invasion and metastasis by targeting MMP-9 in cervical cancer. Gynecol Oncol 141:166–174
Forthun RB, Hinrichs C, Dowling TH et al (2016) The past, present and future subclassification of patients with acute myeloid leukemia. Curr Pharm Biotechnol 17:6–19
Garcia DM, Baek D, Shin C et al (2011) Weak seed-pairing stability and high target-site abundance decrease the proficiency of lsy-6 and other microRNAs. Nat Struct Mol Biol 18:1139–1146
Gu W, Gao T, Shen J et al (2014) MicroRNA-183 inhibits apoptosis and promotes proliferation and invasion of gastric cancer cells by targeting PDCD4. Int J Clin Exp Med 7:2519–2529
Hashemi M, Yousefi J, Hashemi SM et al (2015) Association between programmed cell death 6 interacting protein insertion/deletion polymorphism and the risk of breast cancer in a sample of Iranian Population. Dis Markers 2015:854621
Huang H, Gu J, Yao S et al (2015) MicroRNAs are related to rituximab in combination with cyclophosphamide, doxorubicin, vincristine, and prednisone resistance in patients with diffuse large b-cell lymphoma. Cancer Transl Med 1:6–10
Huangfu L, Liang H, Wang G et al (2016) miR-183 regulates autophagy and apoptosis in colorectal cancer through targeting of UVRAG. Oncotarget 7:4735–4745
Leung WK, He M, Chan AW et al (2015) Wnt/β-Catenin activates MiR-183/96/182 expression in hepatocellular carcinoma that promotes cell invasion. Cancer Lett 362:97–105
Li P, Sheng C, Huang L et al (2014) MiR-183/-96/-182 cluster is up-regulated in most breast cancers and increases cell proliferation and migration. Breast Cancer Res 16:473
Liu YL, Lensing SY, Yan Y et al (2013) Deficiency of CREB and over expression of miR-183 in juvenile myelomonocytic leukemia. Leukemia 27:1585–1588
Lowery AJ, Miller N, Dwyer RM et al (2010) Dysregulated miR-183 inhibits migration in breast cancer cells. BMC Cancer 10:502
Ma Y, Liang AJ, Fan YP et al (2016) Dysregulation and functional roles of miR-183-96-182 cluster in cancer cell proliferation, invasion and metastasis. Oncotarget. In press
Miao F, Zhu J, Chen Y et al (2016) MicroRNA-183-5p promotes the proliferation, invasion and metastasis of human pancreatic adenocarcinoma cells. Oncol Lett 11:134–140
Qiu M, Liu L, Chen L et al (2014) microRNA-183 plays as oncogenes by increasing cell proliferation, migration and invasion via targeting protein phosphatase 2A in renal cancer cells. Biochem Biophys Res Commun 452:163–169
Rubnitz JE, Gibson B, Smith FO (2008) Acute myeloid leukemia. Pediatr Clin North Am 55:21–51
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 3:1101–1108
Su D, Xu H, Feng J et al (2012) PDCD6 is an independent predictor of progression free survival in epithelial ovarian cancer. J Transl Med 10:31
Wang Z, Hong Z, Gao F et al (2013) Upregulation of microRNA-375 is associated with poor prognosis in pediatric acute myeloid leukemia. Mol Cell Biochem 383:59–65
Yan W, Xu L, Sun Z et al (2015) MicroRNA biomarker identification for pediatric acute myeloid leukemia based on a novel bioinformatics model. Oncotarget 6:26424–26436
Yoon JH, Choi YJ, Kim SG et al (2012) Programmed cell death 6 (PDCD6) as a prognostic marker for gastric cancers. Tumour Biol 33:485–494
Zhang QH, Sun HM, Zheng RZ et al (2013) Meta-analysis of microRNA-183 family expression in human cancer studies comparing cancer tissues with noncancerous tissues. Gene 527:26–32
Zhou B, Bai P, Xue H et al (2015) Single nucleotide polymorphisms in PDCD6 gene are associated with the development of cervical squamous cell carcinoma. Fam Cancer 14:1–8
Ziai JM, Siddon AJ, Education Committee of the Academy of Clinical Laboratory Physicians and Scientists (2015) Pathology consultation on gene mutations in acute myeloid leukemia. Am J Clin Pathol 144:539–554
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, X., Zuo, D., Yuan, Y. et al. MicroRNA-183 promotes cell proliferation via regulating programmed cell death 6 in pediatric acute myeloid leukemia. J Cancer Res Clin Oncol 143, 169–180 (2017). https://doi.org/10.1007/s00432-016-2277-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-016-2277-2