Abstract
A small group of tumors of breast and salivary glands contains squamous/epidermoid elements as a constitutive feature (e.g., squamous carcinoma, syringomatous tumors, and mucoepidermoid carcinoma). Other tumors (e.g., pleomorphic adenoma, adenomyoepithelial tumors, and adenoid cystic carcinoma) may show occasionally squamous differentiation. Furthermore, squamous metaplasia may be observed in non-neoplastic breast and salivary tissues. However, the histogenesis of these squamous differentiations is far from being understood. Based on our earlier in situ triple immunofluorescence and quantitative reverse transcription (RT)-PCR experiments for basal keratins K5/14 and p63 as well as for glandular keratins (K7/K8/18), squamous keratins (K10 and K13), and myoepithelial lineage markers (smooth muscle actin, SMA), we here traced the squamous/epidermoid differentiation lineage of 60 tumors of the breast and/or salivary glands, cultured tumor cells of 2 tumors, and of 7 squamous metaplasias of non-neoplastic breast and salivary tissues. Our results indicate that both the neoplastic lesions as well as the non-neoplastic squamous metaplasia contain p63/K5/14+ cells that differentiate toward K10/13+ squamous cells. Thus, cells with squamous/epidermoid differentiation undergo a transition from its original p63/K5/14+ precursor state to K10/13+ squamous lineage state, which can be pictured by triple-immunofluorescence experiments. Given the immunophenotypic similarity of p63/K5/14+ tumor cells to their physiological p63/K5/14+ counterparts in normal breast and salivary duct epithelium, we suggest that these cells provide an important histogenetic key to understanding the pathogenesis of squamous differentiation both in normal breast/salivary gland tissues and their corresponding tumors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In previous studies on adenosquamous carcinoma/syringomatous tumor of the breast and on tumors of breast/salivary glands with myoepithelial differentiation, we demonstrated that K5/K14-positive tumor cells coexpressing p63 are key constituent cells of these lesions [7, 8]. They give rise to glandular and/or myoepithelial cell lineages and, in some tumors, even to heterologous tissues. We now continued these studies with a focus on the phenomenon of epidermoid/squamous differentiation, including lesion categories which constitutively contain squamous differentiation (e.g., squamous carcinoma and adenosquamous/low/intermediate mucoepidermoid carcinoma) and lesions which occasionally show such differentiation (e.g., pleomorphic adenoma, papilloma, and adenomyoepithelial tumors) [6, 35, 36, 42, 51, 58, 86, 90, 94], and finally non-neoplastic tissues with such cellular changes [17, 37, 41, 43, 77, 91, 103]. The histogenesis of squamous differentiation in these lesions is far from fully understood. Based on the expression of p63 both in myoepithelial and in squamous cells, the most-held current view is that of a myoepithelial histogenesis of squamous differentiation in these lesions [27, 29, 33, 46, 88] [75, 89].
The aim of this study was to analyze the developmental potential of cells with special reference to squamous differentiation state in order to better understand the histogenesis of the squamous (epidermoid) lineage differentiation in these lesions. As in our previous studies [8], we used in situ triple immunofluorescence experiments to investigate the simultaneous expression of several keratin subtypes [39, 48, 61, 63], p63 (a p53 homologue) [4, 68, 83, 84], and a myoepithelial marker SMA [4, 30, 40, 102]. Corroborated by quantitative reverse transcription (RT)-PCR expression studies, our results imply that K5/K14 and p63 coexpressing cells are the key players from which squamous differentiation in all these tumors as well as in squamous metaplasia of non-neoplastic tissue develop.
Material and methods
Case selection and histological classification
Routinely processed formalin-fixed paraffin-embedded tissues from 8 squamous cell carcinomas of the breast and 2 of the salivary gland, 21 cases of low-grade adenosquamous carcinomas/syringomatous tumors of the breast, 14 mucoepidermoid carcinomas of the salivary glands, 4 pleomorphic adenomas, 3 epithelial-myoepithelial tumors, 2 fibroepithelial tumors, 2 papillary lesions, 4 no specific type carcinoma or medullary carcinomas, and 7 non-neoplastic tissues with squamous metaplasia were collected from the archives of the Department of Pathology of the University of Muenster (WB) and Albertinen Pathology, Hamburg/Salivary Gland Registry, Hamburg (ThL). Representative hematoxylin and eosin (H/E)-stained sections were reviewed in all cases (WB, ThL), using the criteria described in the WHO breast tumor classification [51, 86, 95] and the WHO classification of head and neck tumors [36]. Normal human breast tissues obtained from plastic surgery and normal salivary glands were used as controls.
Tissue probe processing and primary antibodies
Paraffin tissue sections (4 μm thick) were pretreated and stained as described elsewhere [13]. Antibodies were applied according to the manufacturers’ recommendations. Blocking the endogenous Fc receptors prior to incubation with primary antibodies was omitted, because we have recently reported that endogenous Fc receptors in routinely fixed cell and tissue probes do not retain their ability to bind Fc fragments of antibodies [12]. For immunostaining, we used primary antibodies raised against basal keratins K5 (rabbit monoclonal, MEDAC Diagnostica), K5/6 (mouse monoclonal, DAKO), K14 (mouse monoclonal, Jackson ImmunoRes), glandular keratins K7 (mouse monoclonal, DAKO), K8/18 (mouse monoclonal, Zytomed), K18 (mouse monoclonal, Sigma), squamous keratins K10 and K10/13 (mouse monoclonal, both from DAKO), SMA (rabbit polyclonal, AbCam), vimentin (mouse monoclonal, DAKO), Ki67 (rabbit monoclonal, Thermo Fisher Scientific Inc), and p63 (mouse monoclonal, DAKO). The exclusion of the primary antibody from the immunohistochemical reaction or substitution of primary antibodies with normal IgG (mouse or rabbit) at the same final concentration as that of primary antibodies resulted in lack of immunostaining.
Bright-field microscopy
After immunoreactions with primary antibodies, the sections were treated for 10 min with methanol containing 0.6 % H2O2 to quench endogenous peroxidase. The primary antibodies were then detected with AmpliStain™ horseradish peroxidase (HRP) conjugates (SDT GmbH, Baesweiler, Germany) according to the manufacturers’ instructions [11], and the HRP label was visualized using the DAB substrate kit (Vector Laboratories, Burlingame, CA, USA). When using secondary antibodies conjugated with alkaline phosphatase (AP), AP label was visualized with Dako LSAB REAL Detection System (naphthol phosphate/Fast Red, no. K5005, Dako Corporation, Hamburg, Germany). Finally, the sections were counterstained with hematoxylin. All steps were preceded by rinsing with phosphate-buffered saline (PBS) (pH 7.4).
Triple-immunofluorescence staining experiments
Systematic triple-immunostaining experiments [8] were performed for basal (K5 and K14), glandular/luminal (K7, K8/18), and squamous keratins (K10, K10/13) and for the myoepithelial marker SMA. For these studies, we used secondary antibodies (purchased from Dianova and Molecular Probes) conjugated with Cy3, Alexa Fluor-488, Alexa Fluor-647, or with biotin [13, 14]. For simultaneous visualization of primary antibodies raised in the same species and even belonging to the same iso-species IgG isotype, such primary antibodies were non-covalently labeled in vitro with a reporter molecule employing monovalent IgG Fc-specific Fab fragments [10]. The reporter molecule was either fluorophore Cy3 or biotin. The latter was visualized using fluorophore-labeled streptavidin. Nuclei were counterstained with DAPI (5 μg/ml PBS, 15 s), and sections were mounted with VectaShield (Vector Laboratories, Burlingame, USA).
We also used archived slides with tissue sections immunostained for bright-field microscopy with antibodies bearing AP label visualized with Naphthol phosphate/Fast Red (Fig. 2b). This red chromogen routinely used for visualization of AP in histopathology is also highly fluorescent when using the excitation filter system in the red spectrum. After soaking in xylene for 2–3 days, cover slides were detached, and tissue sections were then rehydrated in graded ethanols to water. After rehydration and antigen retrieval, tissue sections were subjected to immunostaining with one or two additional primary antibodies using secondary antibodies conjugated with Alexa Fluor-488 (green channel) and/or Alexa Fluor-647 (deep-red channel). The Fast Red substrate label and hematoxylin counterstaining used for bright-field microscopy preparations did not interfere with the following immunostaining for fluorescent microscopy.
Image acquisition
Immunostained sections were examined using a Zeiss microscope (Axio Imager Z1). Images were captured and processed using an AxioCam digital microscope camera and the software AxioVision image processing (Carl Zeiss Vision, Germany). The images were imported as JPEG files into PhotoImpact 3.0 (Ulead Systems, Inc. Torrance, CA) and submitted with the final revision of the manuscript at 300 DPI.
AxioVision image processing program permits to present fluorophores in any artificial color when using the black and white digital microscope camera. Thus, in some figures, we can present the red fluorophore Cy3 in yellow color in cases when the other two labels are visualized with green and red fluorophores. Far-red fluorescence (Alexa 647) can also be presented in any artificial color—usually in pink (magenta) color.
Quantitative RT-PCR (qPCR)
Total RNA was extracted from manually microdissected formalin-fixed paraffin embedded (FFPE) tumor tissue sections using the RNeasy FFPE kit (Qiagen, Germany) as described [8]. Messenger RNAs (mRNAs) of basal keratins K5, K14, glandular K7, K8, K18, and epithelial specific squamous keratin 10 were reverse transcribed and amplified using the OneStep RT-PCR kit (Qiagen, Germany). RNA expressions were quantified by real-time fluorescence detection of amplified complementary DNA (cDNA) (ABI Life Technologies StepOne® system) using MGB probes. Gene expression levels were described as ratios between two absolute measurements (gene of interest/HPRT endogenous reference). All samples were run in triplicate for all genes. We compared the differential expression of six keratins (K5/K14/K7/K8/K18/K10) in tumors and in normal breast parenchyma and nipple squamous epithelium, respectively.
Cultured cells
The mucoepidermoid carcinoma cell lines UT-MUC1, derived from an intermediate grade tumor (kindly provided by Dr. Reidar Grenman, University of Turku, Finland), and NCI-H292, derived from a low-grade tumor (CRL-1848, ATCC), were cultured as previously described [5]. Cells were propagated in 25-cm2 flasks and maintained at 37 ° C in 5 % CO2. Tumor cells were seeded out on eight-well chamber slides (Nunc) and allowed to attach overnight. On the next day, the chambers were rinsed briefly with PBS, and the cells were fixed in methanol for 5 min at room temperature. Slides were air-dried for 60 min and subsequently subjected to immunostaining as described above.
CRTC1-MAML2 fusion gene screening
Total RNA was isolated from the two MEC cell lines UT-MUC1 and NCI-H292 and from paraffin tissue sections of 14 low/intermediate-grade MECs as previously described [5, 26]. DNase-treated (DNA-free™; Ambion, Austin, TX) total RNA was subsequently converted to cDNA using the SuperScript™ First-Strand Synthesis System according to the manufacturer’s manual (Invitrogen). The CRTC1-MAML2 fusion transcript was amplified by PCR using previously published primer sets [5, 26]. PCR products were gel-purified and sequenced using an ABI PRISM 310 Genetic Analyzer (Applied Biosystems, Foster City, CA).
Results
Tumors with constitutive squamous differentiation
Squamous carcinoma
Histologically, all ten tumors consisted of pure squamous carcinoma (eight breast, two salivary gland), eight invasive, and two pure squamous in situ lesions. Immunohistochemically, all tumors expressed p63 and K5/6 with a range from 70 to 90 % of tumor cells. The squamous keratins K10 and K10/13 were observed in all tumors with a range of 10–70 % of positive cells (Tables 1 and 2 and Fig. 1).
Invasive squamous carcinoma, grade 3. H/E staining and immunostainings for p63, K5, and K10. Note intensive staining for p63 and basal keratin K5. Patchy positive staining for keratin K10 indicates squamous differentiation. Nuclei counterstained with Ehrlich hematoxylin. Scale bar = 200 μm for the entire layout
In triple-immunofluorescence studies, three types of cells were observed (Fig. 2): (1) cells stained only with antibodies against p63 and basal keratins K5/14, (2) cells positively stained for p63/K5/14 and squamous keratins K10/13, (3) and cells which only stained for squamous keratins K10/13. In this cell lineage, a sequential expression pattern of basal keratins K5/14, p63 and squamous keratins K10/13 with a gradual decrease of p63 and K5/K14, and an increase of K10/13 could be demonstrated, which mirrors the characteristic keratin distribution patterns in normal non-neoplastic squamous epithelium [62, 63]. In well-differentiated squamous cell carcinoma, a zonation of p63, K514, and K10/13 could be observed with p63/K5/14+ cells in the periphery and a transition to K10/13+ squamous cells in central cells similar to normal squamous epithelium (Fig. 2). In contrast, grade 3 carcinomas usually displayed a much more chaotic expression of squamous keratins K10/13. Interestingly, one case coexpressed glandular keratins K7 and another case K8/18 with squamous keratins K10/13 in otherwise typical squamous carcinomas.
Squamous carcinoma of the breast. a Invasive squamous carcinoma, grade 2. H/E stain and triple immunofluorescence for p63 (yellow), K5 (green), and K10 (red) showing that p63/K5+ cells (tailed arrows) differentiate through intermediary cells (arrows) to squamous cells (asterisks). b Invasive squamous carcinoma, grade 3. Triple immunofluorescence for p63 (red), K5 (green), and squamous keratin K10 (pink). Nearly all tumor cells express high levels of K5 and p63. Squamous differentiation is observed showing increasing K10 staining through intermediary cells expressing p63. K5 and K10 (arrows) to K10+ squamous cells (asterisk)
The immunohistochemical findings on two cases of ductal carcinoma in situ (DCIS) showed the same p63, K5/14, and K10/13 pattern as described for invasive squamous carcinoma (Fig. 3). A compressed myoepithelial layer was confined to the outer layer similar to classical DCIS.
Ductal carcinoma in situ, squamous type, grade 3. H/E staining and immunostainings for p63, K5, SMA, and K10. The tumor cells display the same staining pattern as invasive squamous carcinoma with a chaotic K10 staining for differentiating squamous cells. The compressed myoepithelial cells express smooth muscle actin. Nuclei counterstained with Ehrlich hematoxylin
Low-grade adenosquamous carcinoma/syringomatous tumors
Twenty-one cases of low-grade adenosquamous carcinomas/syringomatous tumors of the breast were part of this and of another study, which has been published recently [9]. In addition to glandular differentiation, all these lesions contained areas of squamous differentiation (Table 3). The most noticeable finding was the strong immunoreaction for keratins K5/14 (70–90 %) and for p63 (30–90 %) in all tumors investigated. The antibodies to squamous keratins K10 and K10/13 displayed a range of positivity from 5 to 50 %.
The results of the triple-immunofluorescence experiments have been described in detail in a recent publication. Suffice it to say here that the squamous differentiation lineage was again characterized by a sequential expression of p63, K5/K14, and K10/K13. A small number of hybrid cells were observed which coexpressed glandular keratins K7 and/or K8/18 and squamous keratin K10 (not shown here). As shown previously, the involvement of myoepithelial cells in both glandular and squamous differentiations could be clearly ruled out [8]. However, the adjacent stroma of low-grade adenosquamous carcinoma/syringomatous tumor may contain SMA-positive myofibroblasts, which, in the vicinity of the tumor cells, may be misinterpreted as myoepithelial cells [23].
Mucoepidermoid carcinoma
We analyzed 14 cases of low/intermediate-grade mucoepidermoid carcinomas of the salivary gland origin All these cases were shown to express the mucoepidermoid carcinoma-specific CRTC1-MAML2 gene fusion (data not shown). All cases contained solid and cystic areas with smaller basaloid, intermediate/large cells with eosinophilic cytoplasm, clear, onkocytic, and even epidermoid cells. Mucinous cells were observed in all cases but usually in small numbers. Overtly squamous differentiation was observed only in one case.
Immunohistochemically, we observed a moderate to strong but variable reaction for p63, K5/6, and an intensive reaction for the glandular keratin K7 in most of the tumor cells (Table 4 and Fig. 4). A constant finding was the paucity of K10+, K10/13+ squamous cells accounting for less than 5 % of all tumor cells with either individual positive cells or positivity in small clusters of epidermoid cells. Interestingly, few cases did not stain with the K10 antibody but showed a clear reaction with the K10/13 antibody and vice versa. In two of the 14 cases, no staining was observed with both K10 and K10/13 antibodies.
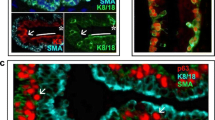
Mucoepidermoid carcinoma of the salivary gland. a H/E stain of typical mucoepidermoid carcinoma with solid and microcystic growth pattern with occasional mucinous cells and triple immunofluorescence for p63 (yellow), K5 (pink), and K7 (green) showing small clusters of p63/K5-positive cells (asterisk). Few cells express only K5 (arrow); some cells express either K5 and the glandular keratin K7 (crossed arrow) or only K7 (hollow arrow). Note that this staining pattern mimics normal glandular differentiation in excretory ducts of the salivary glands. b Another area of the same tumor. Triple immunofluorescence for p63 (yellow), K5 (pink), and K10 (green). Many cells in this picture express p63 and K5. Note that the few cells with K10+ squamous differentiation usually co-express K5 and p63 (arrows). c Cell culture of a mucoepidermoid carcinoma of the salivary gland. Triple immunofluorescence for K5 (pink), K7 (green), and K10 (red). A number of tumor cells express only K5 (arrows). K5-positive cells differentiate toward glandular cells expressing K7 (crossed arrows), and only few cells also differentiate to squamous cells (asterisks)
Triple-immunofluorescence experiments, performed on ten tumors, revealed that the bulk of tumor cells coexpressed p63, K5, and/or K7 (Fig. 5). This staining pattern was observed in both basaloid and intermediate cells. Larger “epidermoid cells” often expressed only K7, indicating glandular cell differentiation. The single cells or small clusters of cells positive for squamous keratins K10/13 usually coexpressed p63 and K5. Furthermore, p63/K5/14+ only cells are observed in all tumors.
Using triple-immunofluorescence experiments, we also analyzed cells in two CRTC1-MAML2 fusion-positive mucoepidermoid carcinoma cell lines. In both cell lines, we observed glandular and squamous differentiation (Fig. 5c). In contrast to the primary tumors, many tumor cells of both cell cultures, however, showed a progenitor cell phenotype expressing K5.
Tumors with occasional squamous differentiation
Areas of squamous differentiation in the form of relatively well-demarcated foci of cells with ample eosinophilic cytoplasm were occasionally seen in a number of benign or malignant neoplasms of breast and salivary glands. Here, we have analyzed two cases of papillomas, two cases of fibroadenoma/phyllodes tumor, four pleomorphic adenomas (one breast, three salivary gland), three epithelial-myoepithelial tumors (breast), and three carcinomas (breast) with such squamous differentiation. In all these lesions, we found histologically focal area(s) and in one case of phyllodes tumor, even widespread squamous differentiation. Immunohistochemically, the lesions displayed the well-known staining patterns in regard to p63, K5, K8/18, and SMA known from the literature. Thus, the two papillomas and the fibroadenoma showed P63, K5, and SMA positivity in the myoepithelial layer and K8/18 positivity in the luminal layer with occasional coexpression of K5 in the latter cells. Furthermore, pleomorphic adenomas and epithelial-myoepithelial tumors contained p63+/K5+ cells that differentiate toward glandular and/or myoepithelial cell lineages as has been published recently [7, 8]. The three carcinomas (two medullary and one NST, grade 3) showed positivity for p63, K5, and K8/18 but were negative for SMA. Although these tumors showed considerable heterogeneity in its cellular components in isTIF stainings, the common denominator of all these 14 lesions is the presence of p63+ and K5+ cells that lacked any of the lineage markers such as glandular (K7; K8/18) and myoepithelial markers (e.g., SMA and calponin). Furthermore, in regard to squamous differentiation transition from p63/K5+ cells to K10+ squamous cells with a zonation similar to normal squamous epithelium or with a more chaotic pattern of single or small groups of K10+, K10/13+ cells were observed (Fig. 6).
Squamous metaplasia in infarcted papilloma. a H/E staining. b Triple immunofluorescence for p63 (yellow), K5 (green), and smooth muscle actin (pink). Note that the cells with squamous differentiation lack any SMA staining. Insert: double immunofluorescence for p63 (red) and K10 (green). Note the focal expression of squamous keratin K10 (green)
Squamous metaplasia in non-neoplastic tissue
Squamous metaplasia is a rare event in normal breast and salivary gland tissues alone or in conjunction with inflammatory processes. Here, we analyzed one case with squamous metaplasia of lactiferous duct, two cases with displacement of breast epithelium after needle biopsy with squamous metaplasia, and four cases of sialometaplasia of the salivary glands. Three cases of sialometaplasia displayed only p63+ and K5/14+ cells. In one case of sialometaplasia and in all cases of squamous metaplasia in normal breast tissue p63/K5/14+, cells were observed which transformed from p63+, K5/14+, and K10/13+ cells to only K10/13+ squamous cells.
Identification of p63/K5/14 cells in normal breast and salivary duct epithelium
In order to understand the cellular relationship of these lesions to normal epithelium, we applied triple-immunofluorescence experiments on normal salivary and breast duct epithelium. p63/K5+ progenitor cells were observed in small clusters of a few cells basally located in salivary duct epithelium and at the interface between the myoepithelial and luminal layers of breast duct epithelium (Fig. 7).
Excretory duct of the salivary gland and normal nipple duct. a–c Excretory duct of the salivary gland. Triple immunofluorescence for p63 (red), K5 (yellow), and K7 (green) showing p63/K5+ cells at the base of the duct epithelium with formation of three more prominent clusters (asterisks). Some cells of these clusters are more tethered to the luminally located epithelium showing a decrease in p63 and finally leading to K5+ only cells (arrow). In K5+ cells, increasing staining of K7 is seen (crossed arrows). Finally, K7+ only glandular cells are observed which form the bulk of the duct epithelium. From these findings, we conclude that p63/K5+ cells function as glandular progenitors in excretory ducts. d–f Large segmental duct. Triple immunofluorescence for p63 (red), K5 (green), and SMA (pink). Note that p63 is expressed mainly in K5-positive basally located cells, most of which also co-express SMA (hollow arrows). Thus, these cells have to be interpreted as myoepithelial cells. Notice the occasional cells at the interface between myoepithelial and luminal layer which co-express p63 and K5 (asterisks). These cells do not express SMA and do not express K8/18 (not shown here). Interestingly, the more luminally situated cells express only K5 (arrows)
qPCR Analyses
The expressions of basal keratins K5 and K14, luminal keratins K8 and K18, and epidermal/squamous specific keratin K10 were studied by qPCR in 19 tumors with squamous differentiation (Fig. 8). As shown in this figure, we found similar levels of up-regulated mRNA of basal keratins in all tumors, whereas the K10 mRNA levels of mucoepidermoid carcinomas were found to be low corroborating the immunohistochemical observations of only single or few K10-positive cells in these lesions. Interestingly, the mRNA levels of luminal keratins K8 and K18 displayed a great variability with the highest levels observed in mucoepidermoid carcinomas. The mRNA expression data corroborated the immunohistochemical findings implying that these tumors represent basal-type tumors with squamous differentiation.
mRNA expression patterns for basal keratins K5 and K14, glandular keratins K8 and K18, and squamous keratin K10 in carcinomas with squamous differentiation. Note the high expression levels of basal keratins K5 and/or K14 in all carcinomas with squamous differentiation compared to normal breast tissue (zero line). Likewise, the K10 mRNA level of MECs is low to intermediate compared to squamous carcinoma. The glandular keratins K8 and K18 display variable levels in low-grade adenosquamous carcinomas/syringomatous tumors, but usually high mRNA levels in low-grade mucoepidermoid carcinoma. One case of squamous cell carcinoma showed down-regulation of both luminal keratins K8 and K18, whereas the other case, displayed up-regulated K18 mRNA level
Discussion
The histogenesis of squamous differentiation of breast and salivary gland tumors is far from being understood. In this study, we therefore analyzed the differential expression of different keratin subtypes [39, 62–65], p63 (a p53 homologue) [19, 68, 72], and SMA [16, 22, 25, 31, 38, 40, 53, 57, 66, 71, 75, 81, 84] to get a deeper insight into the cellular pathways of squamous differentiation in 60 tumors of breast and/or salivary glands and in 7 non-neoplastic squamous metaplasia of both glands. We used triple-immunofluorescence staining experiments for molecular mapping at the resolution of single cells and qPCR analyses of tissue sections for specifically identifying the phenotypic signature of the constituent cells. We demonstrated (1) that all non-neoplastic and neoplastic lesions with squamous/epidermoid differentiation contain progenitor cells expressing only the basal keratin K5 and p63 [4, 68, 84], but lacking lineage markers, and (2) that the squamous lineage revealed by triple-immunofluorescence staining can be arrayed in a succession of various cell types starting from p63/K5+ progenitor cells and, with increasing maturation, showing a transition to K10+ and/or K10/13+ state via an intermediary state of p63+, K5+, K10+, and/or K10/13+ cells. Therefore, non-neoplastic squamous metaplasia and squamous differentiation in these tumors recapitulate the tightly regulated differentiation-specific expression pattern of p63 and keratins in normal squamous epithelium [19, 63, 87, 97, 100] and in squamous carcinomas of other sites [62–64]. This interpretation contradicts the most held current viewpoint which, based on the p63 expression in both myoepithelial and squamous cells, explains the origin of squamous epithelium from myoepithelial cells [21, 28, 29]. In contrast, we regard the occurrence of myoepithelial and squamous and other elements in some of tumors with multilineage differentiation to represent the differentiative progeny capability state of p63/K5+ progenitor cells within this cell model. We furthermore hypothesize that the squamous and glandular differentiation potential [79, 98] may reflect the physiological differentiation potential of the ontogenetic ectodermal rudiment where breast and salivary glands are derived from. How these data modify the way, in which squamous metaplasia and tumors with squamous differentiation of breast and salivary glands can be thought or modeled, is shown in the schematic diagram in Fig. 9.
Models of progenitor relationships. In normal breast duct epithelium (and in salivary gland duct, not shown here), p63+/K5+ progenitors give rise to early glandular cells (K5+), then mature to intermediary glandular cells (K5+/K7+/K8/18+), and finally to glandular cells (K7+/K8/18+). During glandular differentiation, expression of p63 is down-regulated in normal duct epithelium. Likewise, in normal nipple squamous epithelium, immunophenotypically identical p63+/K5+ bipotent progenitors give rise to smooth muscle actin-positive myoepithelial cells and further to p63+/K5+/K10+ cells, which then mature to K7+ and/orK8/18+ glandular cells. The similarity in immunophenotypical appearance of the progenitors in these epithelia and the fact that they differentiate in either glandular or myoepithelial cell fates suggest robust lineage commitments in each site. All three tumors (squamous carcinoma, low-grade adenosquamous carcinoma/syringomatous tumor, and mucoepidermoid carcinoma) and squamous metaplasia contain p63+/K5+ progenitors, which differentiate to squamous (K10+) and glandular (K7+/K8/18+) cells through intermediary stages. It should be noted that the glandular differentiation in most tumors starts with down-regulation of p63 similar to the normal duct epithelium. The squamous lineage differentiation of all tumor types is a mirror of normal squamous epithelium of the ectoderm in which the organs arise. It is characterized by a sequential expression of p63 and K5, on one hand, and K10, on the other hand. Both p63 and basal keratin K5 may finally be down-regulated, whereas K10 is up-regulated. We assume that different cell lineages may arise via reprogramming of the target physiological progenitors during malignant transformation. Thickness of arrows here corresponds to the amount of differentiation found in these tumors
Each classification system of tumors draws its life from analogies to normal tissues. The data shown here provided several important insights and raise critical questions relevant to understanding and exploiting progenitor cell function of p63/K5+ cells. Recognizing that in situ triple-immunofluorescence experiments provide a simple method for specifically identifying and tracing such cells and their progeny in tissue sections, the authors took advantage to determine the anatomical location of cells of identical phenotype in normal breast and salivary gland tissues. We have found physiological progenitors with an identical p63/K5+ immunophenotype, similar to non-neoplastic squamous metaplasia and in tumors with squamous differentiations, residing at the interface of myoepithelial and luminal cells of breast ducts [8], and furthermore, identical cells were observed in a basal position of excretory and striated ducts of the salivary glands [20, 44].
However, the question arises, do p63+ and K5+ tumor progenitor cells originate from normal cells of the same phenotype or can tumor cells acquire this phenotype due to the mutagenic events? Our results provided by this observational study are clearly not sufficient to prove a p63/K5+ cell of origin for these tumors. However, our findings raise the interesting possibility that these physiological counterparts or a subpopulation of these cells may play a role in the development of squamous metaplasia and tumors with squamous differentiation. This interpretation is in agreement with data that p63 positivity is found in reserve cells of the human endometrium and cervical epithelium and its metaplastic (squamous or mucinous) epithelia, either alone or in conjunction with hyperplasias or carcinomas [60, 69, 70, 76, 78]. And finally, in the authors’ experience, squamous differentiations in adnexal tumors of the skin [47, 54] may be further examples of the same theme. These findings are also in line with observations on squamous epithelium of the esophagus, squamous metaplasia in the tracheobronchial tree, and low-grade squamous carcinoma showing K5 and p63 colocalization in less-differentiated cells, whereas more differentiated regions do not express K5 and p63 [19, 68, 85].
However, despite the idea of phenotypically defined p63/K5+ progenitors with their specific differentiation capacities in both normal tissues and their matching tumors, several questions arise. Specifically, all tumors analyzed are assumed to be characterized by a proportion of p63/K5/14+ precursors. However, some lesions (low-grade mucoepidermoid carcinoma, adenoid cystic carcinoma) contained only few progenitors or small groups of cells expressing only p63 and K5/14, whereas p63/K5/14+ progenitors were easily found in other lesions (e.g., squamous carcinoma, low-grade adenosquamous carcinoma/syringomatous tumor pleomorphic adenoma, and non-neoplastic squamous metaplasia). Furthermore, K10/13+ squamous differentiation varies tremendously, and K10/13+ squamous differentiation is only a minor component or even lacking in some tumors (for example in mucoepidermoid carcinoma). Vice versa, glandular differentiation is the main component in low-grade mucoepidermoid carcinoma but usually a minor component in low-grade adenosquamous carcinoma/syringomatous tumor. Interestingly, studies of two CRTC1-MAML2 fusion-positive low/intermediate-grade mucoepidermoid carcinoma cell lines showed the majority of cultured tumor cells displaying the classical p63+/K5/14+ progenitor phenotype. Thus, we conclude from our findings as indicating that low-grade mucoepidermoid carcinomas contain tumor cells with a progenitor cell phenotype as evidenced by the expression of p63 both in primary tumors [33] and in mucoepidermoid carcinoma-derived cell lines. Further molecular studies of these tumors are expected to give insights into the complex processes of cellular differentiation and the molecular pathogenesis of these interesting lesions.
Recent microarray-based gene expression profiling and/or immunohistochemical studies [18, 39, 49, 50, 56, 64, 67, 73, 74, 92, 93, 101] have led to a paradigm shift in understanding breast and salivary gland tumors, with a particular focus on basal-like tumors [3, 45, 80, 82, 96, 99] According to this concept, look-alike tumors with overt myoepithelial differentiation, tumors such as secretory carcinomas and carcinomas with squamous differentiation, preferentially pertain to the basal-like subgroup and have been shown to express basal keratins (K5, K14, and K17) and p63 in most, if not all, cases [1, 2, 15, 24, 32–34, 50, 52, 59, 82] Of interest, in benign lesions with squamous differentiation and also of non-neoplastic squamous metaplasias, the cellular components regarding squamous differentiation are identical to the carcinomas tested. More important, many tumors of this group are characterized by the presence of p63/K5+ progenitors which show the ability to enter into different phenotypic states associated with alternate lineage differentiations. This characterizes the p63/K5+ progenitor in regard to differentiation capacity as a highly versatile cell. These data are in contrast to the recent findings of basal-like breast cancers arising in women carrying mutations in the BRCA1 gene. Thus, Lim et al. (2009) observed that these tumors were more similar to normal luminal progenitor cells (p63-negative; but K5- and K8/18-positive) [55]. Identification of these different subtypes of basal-like tumors is thought to be relevant from a biological point of view and may provide a conceptual framework to understand the complexity and heterogeneity of these lesions.
In summary, our findings demonstrate that cells of non-neoplastic squamous metaplasia and of carcinomas with constitutively or occasionally found squamous/epidermoid differentiation undergo a transition from its original p63/K5/14+ precursor state to a new K10/13+ squamous lineage state, which can be demonstrated by in situ triple immunofluorescence labeling and mRNA expression studies. Given the phenotypic similarity of p63/K5/14+ tumor cells to their physiological counterparts in the normal breast and salivary duct epithelium and considering the fact that these cells in a number of epithelial tissues give rise to different progeny with specific differentiation lineages, we suggest that the p63/K5+ progenitor cell type and its differentiation potential provide an important ontogenetic key to a better understanding of the pathogenesis of these lesions which are proposed to constitute a special type of basal-like tumors.
References
Abd El-Rehim DM, Pinder SE, Paish CE et al (2004) Expression of luminal and basal cytokeratins in human breast carcinoma. J Pathol 203:661–671
Azoulay S, Lae M, Freneaux P et al (2005) KIT is highly expressed in adenoid cystic carcinoma of the breast, a basal-like carcinoma associated with a favorable outcome. Mod Pathol 18:1623–1631
Badve S, Dabbs DJ, Schnitt SJ et al (2011) Basal-like and triple-negative breast cancers: a critical review with an emphasis on the implications for pathologists and oncologists. Mod Pathol 24:157–167
Barbareschi M, Pecciarini L, Cangi MG et al (2001) p63, a p53 homologue, is a selective nuclear marker of myoepithelial cells of the human breast. Am J Surg Pathol 25:1054–1060
Behboudi A, Enlund F, Winnes M et al (2006) Molecular classification of mucoepidermoid carcinomas-prognostic significance of the MECT1-MAML2 fusion oncogene. Gene Chromosome Cancer 45:470–481
Bennett AK, Mills SE, Wick MR (2003) Salivary-type neoplasms of the breast and lung. Semin Diagn Pathol 20:279–304
Boecker W, Junkers T, Reusch M et al (2012) Origin and differentiation of breast nipple syringoma. Sci Rep 2:226
Boecker W, Stenman G, Loening T et al (2013) K5/K14-positive cells contribute to salivary gland-like breast tumors with myoepithelial differentiation. Mod Pathol 26:1086–1100
Boecker W, Stenman G, Loening T et al (2014) Differentiation and histogenesis of syringomatous tumour of the nipple and low-grade adenosquamous carcinoma: evidence for a common origin. Histopathology 65:9–23
Brown JK, Pemberton AD, Wright SH et al (2004) Primary antibody-Fab fragment complexes: a flexible alternative to traditional direct and indirect immunolabeling techniques. J Histochem Cytochem 52:1219–1230
Buchwalow I, Boecker W, Wolf E et al (2013) Signal amplification in immunohistochemistry: loose-jointed deformable heteropolymeric HRP conjugates vs. linear polymer backbone HRP conjugates. Acta Histochem 115:587–594
Buchwalow I, Samoilova V, Boecker W et al (2011) Non-specific binding of antibodies in immunohistochemistry: fallacies and facts. Sci Rep 1:28
Buchwalow IB, Boecker W (2010) Immunohistochemistry: basics and methods. Springer Berlin Heidelberg
Buchwalow IB, Minin EA, Boecker W (2005) A multicolor fluorescence immunostaining technique for simultaneous antigen targeting. Acta Histochem 107:143–148
Calkins LA, Pearce EW (1958) Labor in the American Negro. Am J Obstet Gynecol 75:575–585, Discussion 585–589
Clarke C, Sandle J, Lakhani SR (2005) Myoepithelial cells: pathology, cell separation and markers of myoepithelial differentiation. J Mammary Gland Biol Neoplasia 10:273–280
Crile G Jr, Chatty EM (1971) Squamous metaplasia of lactiferous ducts. Arch Surg 102:533–534
Dairkee SH, Ljung BM, Smith H et al (1987) Immunolocalization of a human basal epithelium specific keratin in benign and malignant breast disease. Breast Cancer Res Treat 10:11–20
Daniely Y, Liao G, Dixon D et al (2004) Critical role of p63 in the development of a normal esophageal and tracheobronchial epithelium. Am J Physiol Cell Physiol 287:C171–C181
Dardick I, Byard RW, Carnegie JA (1990) A review of the proliferative capacity of major salivary glands and the relationship to current concepts of neoplasia in salivary glands. Oral Surg Oral Med Oral Pathol 69:53–67
Dardick I, Gliniecki MR, Heathcote JG et al (1990) Comparative histogenesis and morphogenesis of mucoepidermoid carcinoma and pleomorphic adenoma. An ultrastructural study. Virchows Arch A Pathol Anat Histopathol 417:405–417
Deugnier MA, Teuliere J, Faraldo MM et al (2002) The importance of being a myoepithelial cell. Breast Cancer Res 4:224–230
Di Tommaso L, Pasquinelli G, Damiani S (2003) Smooth muscle cell differentiation in mammary stromo-epithelial lesions with evidence of a dual origin: stromal myofibroblasts and myoepithelial cells. Histopathology 42:448–456
Diallo R, Schaefer KL, Bankfalvi A et al (2003) Secretory carcinoma of the breast: a distinct variant of invasive ductal carcinoma assessed by comparative genomic hybridization and immunohistochemistry. Hum Pathol 34:1299–1305
Eddinger TJ, Murphy RA (1991) Developmental changes in actin and myosin heavy chain isoform expression in smooth muscle. Arch Biochem Biophys 284:232–237
Fehr A, Meyer A, Heidorn K et al (2009) A link between the expression of the stem cell marker HMGA2, grading, and the fusion CRTC1-MAML2 in mucoepidermoid carcinoma. Gene Chromosome Cancer 48:777–785
Foschini MP, Eusebi V (1998) Carcinomas of the breast showing myoepithelial cell differentiation. A review of the literature. Virchows Arch 432:303–310
Foschini MP, Krausz T (2010) Salivary gland-type tumors of the breast: a spectrum of benign and malignant tumors including “triple negative carcinomas” of low malignant potential. Semin Diagn Pathol 27:77–90
Foschini MP, Pizzicannella G, Peterse JL et al (1995) Adenomyoepithelioma of the breast associated with low-grade adenosquamous and sarcomatoid carcinomas. Virchows Arch 427:243–250
Foschini MP, Scarpellini F, Gown AM et al (2000) Differential expression of myoepithelial markers in salivary, sweat and mammary glands. Int J Surg Pathol 8:29–37
Foschini MP, Simpson JF, O’Malley FO (2012) Ductal adenoma. In: Lakhani SR et al (eds) WHO classification of tumors of the breast. IARC, Lyon, pp 117–118
Geyer FC, Lacroix-Triki M, Savage K et al (2011) beta-Catenin pathway activation in breast cancer is associated with triple-negative phenotype but not with CTNNB1 mutation. Mod Pathol 24:209–231
Geyer FC, Lambros MB, Natrajan R et al (2010) Genomic and immunohistochemical analysis of adenosquamous carcinoma of the breast. Mod Pathol 23:951–960
Gill MS, Paish EC, Ronan J, et al (2012) Comparison of the PharmDx immunohistochemical system with standard methods for assessing estrogen and progesterone receptors in invasive carcinoma of the breast. Appl Immunohistochem Mol Morphol 21(1):90–93. doi:10.1097/PAI.0b013e3182609202
Goode AW, El-Naggar AK (2005) Mucoepidermoid carcinoma. In: Barnes L et al (eds) WHO: head and neck tumours. IARS Press, Lyon, pp 219–220
Goode RK, El-Naggar A (2005) Muccoepidermoid carcinoma. In: Barnes L et al. (eds) Pathology and genetics of head and neck tumours. IARC Press, Lyon, pp 219–220
Gottfried MR (1986) Extensive squamous metaplasia in gynecomastia. Arch Pathol Lab Med 110:971–973
Gudjonsson T, Ronnov-Jessen L, Villadsen R et al (2002) Normal and tumor-derived myoepithelial cells differ in their ability to interact with luminal breast epithelial cells for polarity and basement membrane deposition. J Cell Sci 115:39–50
Gusterson BA, Ross DT, Heath VJ et al (2005) Basal cytokeratins and their relationship to the cellular origin and functional classification of breast cancer. Breast Cancer Res 7:143–148
Gusterson BA, Warburton MJ, Mitchell D et al (1982) Distribution of myoepithelial cells and basement membrane proteins in the normal breast and in benign and malignant breast diseases. Cancer Res 42:4763–4770
Habif DV, Perzin KH, Lipton R et al (1970) Subareolar abscess associated with squamous metaplasia of lactiferous ducts. Am J Surg 119:523–526
Horii R, Akiyama F, Ikenaga M et al (2006) Muco-epidermoid carcinoma of the breast. Pathol Int 56:549–553
Hurt MA, Az-Arias AA, Rosenholtz MJ et al (1988) Posttraumatic lobular squamous metaplasia of breast. An unusual pseudocarcinomatous metaplasia resembling squamous (necrotizing) sialometaplasia of the salivary gland. Mod Pathol 1:385–390
Ihrler S, Blasenbreu-Vogt S, Sendelhofert A et al (2004) Regeneration in chronic sialadenitis: an analysis of proliferation and apoptosis based on double immunohistochemical labelling. Virchows Arch 444:356–361
Jacquemier J, Padovani L, Rabayrol L et al (2005) Typical medullary breast carcinomas have a basal/myoepithelial phenotype. J Pathol 207:260–268
Jones MW, Norris HJ, Snyder RC (1989) Infiltrating syringomatous adenoma of the nipple. A clinical and pathological study of 11 cases. Am J Surg Pathol 13:197–201
Kazakov DV, Michal M, Kazerovska D et al (2012) Neoplasms with multilineage differentiation. Cutaneous adnexal tumors. Wolters Kluwer, Lippincott Williams and Wilkins, Philadelphia, pp 417–442
Korsching E, Packeisen J, Agelopoulos K et al (2002) Cytogenetic alterations and cytokeratin expression patterns in breast cancer: integrating a new model of breast differentiation into cytogenetic pathways of breast carcinogenesis. Lab Investig 82:1525–1533
Laakso M, Tanner M, Nilsson J et al (2006) Basoluminal carcinoma: a new biologically and prognostically distinct entity between basal and luminal breast cancer. Clin Cancer Res 12:4185–4191
Lae M, Freneaux P, Sastre-Garau X et al (2009) Secretory breast carcinomas with ETV6-NTRK3 fusion gene belong to the basal-like carcinoma spectrum. Mod Pathol 22:291–298
Lakhani SR, Ellis IO, Schnitt SJ et al (2012) WHO classification of tumors of the breast. IARC, Lyon
Lambros MB, Tan DS, Jones RL et al (2009) Genomic profile of a secretory breast cancer with an ETV6-NTRK3 duplication. J Clin Pathol 62:604–612
Lazard D, Sastre X, Frid MG et al (1993) Expression of smooth muscle-specific proteins in myoepithelium and stromal myofibroblasts of normal and malignant human breast tissue. Proc Natl Acad Sci U S A 90:999–1003
LeBoit PA, Burg G, Weedon D, Sarasin A (2006) Pathology and genetics of skin tumours. IARC Press, Lyon
Lim E, Vaillant F, Wu D et al (2009) Aberrant luminal progenitors as the candidate target population for basal tumor development in BRCA1 mutation carriers. Nat Med 15:907–913
Livasy CA, Karaca G, Nanda R et al (2006) Phenotypic evaluation of the basal-like subtype of invasive breast carcinoma. Mod Pathol 19:264–271
Longtine JA, Pinkus GS, Fujiwara K et al (1985) Immunohistochemical localization of smooth muscle myosin in normal human tissues. J Histochem Cytochem 33:179–184
Luchtrath H, Moll R (1989) Mucoepidermoid mammary carcinoma. Immunohistochemical and biochemical analyses of intermediate filaments. Virchows Arch A Pathol Anat Histopathol 416:105–113
Marchio C, Weigelt B, Reis-Filho JS (2010) Adenoid cystic carcinomas of the breast and salivary glands (or ‘The strange case of Dr Jekyll and Mr Hyde’ of exocrine gland carcinomas). J Clin Pathol 63:220–228
Martens JE, Smedts F, van Muyden RC et al (2007) Reserve cells in human uterine cervical epithelium are derived from mullerian epithelium at midgestational age. Int J Gynecol Pathol 26:463–468
McCarthy KP, Slack DN, Sloane JP (1994) The polymerase chain reaction in diagnosing lymphoid disorders. Mol Biol Rep 19:69–77
Moll R, Divo M, Langbein L (2008) The human keratins: biology and pathology. Histochem Cell Biol 129:705–733
Moll R, Franke WW, Schiller DL et al (1982) The catalog of human cytokeratins: patterns of expression in normal epithelia, tumors and cultured cells. Cell 31:11–24
Moll R, Krepler R, Franke WW (1983) Complex cytokeratin polypeptide patterns observed in certain human carcinomas. Differentiation 23:256–269
Moll R, Zimbelmann R, Goldschmidt MD et al (1993) The human gene encoding cytokeratin 20 and its expression during fetal development and in gastrointestinal carcinomas. Differentiation 53:75–93
Moritani S, Kushima R, Sugihara H et al (2002) Availability of CD10 immunohistochemistry as a marker of breast myoepithelial cells on paraffin sections. Mod Pathol 15:397–405
Nagle RB, Bocker W, Davis JR et al (1986) Characterization of breast carcinomas by two monoclonal antibodies distinguishing myoepithelial from luminal epithelial cells. J Histochem Cytochem 34:869–881
Nylander K, Vojtesek B, Nenutil R et al (2002) Differential expression of p63 isoforms in normal tissues and neoplastic cells. J Pathol 198:417–427
O’Connell JT, Mutter GL, Cviko A et al (2001) Identification of a basal/reserve cell immunophenotype in benign and neoplastic endometrium: a study with the p53 homologue p63. Gynecol Oncol 80:30–36
Park JJ, Sun D, Quade BJ et al (2000) Stratified mucin-producing intraepithelial lesions of the cervix: adenosquamous or columnar cell neoplasia? Am J Surg Pathol 24:1414–1419
Pavlakis K, Zoubouli C, Liakakos T et al (2006) Myoepithelial cell cocktail (p63 + SMA) for the evaluation of sclerosing breast lesions. Breast 15:705–712
Pavlov K, Maley CC (2010) New models of neoplastic progression in Barrett’s oesophagus. Biochem Soc Trans 38:331–336
Perou CM, Jeffrey SS, van de Rijn M et al (1999) Distinctive gene expression patterns in human mammary epithelial cells and breast cancers. Proc Natl Acad Sci U S A 96:9212–9217
Perou CM, Sorlie T, Eisen MB et al (2000) Molecular portraits of human breast tumours. Nature 406:747–752
Prasad AR, Savera AT, Gown AM et al (1999) The myoepithelial immunophenotype in 135 benign and malignant salivary gland tumors other than pleomorphic adenoma. Arch Pathol Lab Med 123:801–806
Quade BJ, Yang A, Wang Y et al (2001) Expression of the p53 homologue p63 in early cervical neoplasia. Gynecol Oncol 80:24–29
Raju GC, Wee A (1990) Spindle cell carcinoma of the breast. Histopathology 16:497–499
Regauer S, Reich O (2007) CK17 and p16 expression patterns distinguish (atypical) immature squamous metaplasia from high-grade cervical intraepithelial neoplasia (CIN III). Histopathology 50:629–635
Reis-Filho JS, Lakhani SR, Gpbbi H et al (2012) Metaplastic carcinoma. In: Sunil Lakhani, et.al (eds) WHO Classification of Tumours of the Breast, 4th edn. IARC, Lyon, p 48–52
Reis-Filho JS, Milanezi F, Carvalho S et al (2005) Metaplastic breast carcinomas exhibit EGFR, but not HER2, gene amplification and overexpression: immunohistochemical and chromogenic in situ hybridization analysis. Breast Cancer Res 7:R1028–R1035
Reis-Filho JS, Milanezi F, Paredes J et al (2003) Novel and classic myoepithelial/stem cell markers in metaplastic carcinomas of the breast. Appl Immunohistochem Mol Morphol 11:1–8
Reis-Filho JS, Milanezi F, Steele D et al (2006) Metaplastic breast carcinomas are basal-like tumours. Histopathology 49:10–21
Reis-Filho JS, Preto A, Soares P et al (2003) p63 expression in solid cell nests of the thyroid: further evidence for a stem cell origin. Mod Pathol 16:43–48
Reis-Filho JS, Schmitt FC (2002) Taking advantage of basic research: p63 is a reliable myoepithelial and stem cell marker. Adv Anat Pathol 9:280–289
Reis-Filho JS, Simpson PT, Martins A et al (2003) Distribution of p63, cytokeratins 5/6 and cytokeratin 14 in 51 normal and 400 neoplastic human tissue samples using TARP-4 multi-tumor tissue microarray. Virchows Arch 443:122–132
Reis Filho J, Lakhani A, Gobbi H et al (2012) Metaplastic carcinoma. In: Lakhani SR et al (eds) WHO classification of tumors of the breast. IARC, Lyon, pp 48–52
Roop DR (1987) Regulation of keratin gene expression during differentiation of epidermal and vaginal epithelial cells. Curr Top Dev Biol 22:195–207
Rosen PP (2009) Rosen’s breast pathology. Wolters Kluwer, Lippincott Wiliams & Wilkins, Philadelphia
Rothnagel JA, Mehrel T, Idler WW et al (1987) The gene for mouse epidermal filaggrin precursor. Its partial characterization, expression, and sequence of a repeating filaggrin unit. J Biol Chem 262:15643–15648
Sapino A, Sneige N, Euseby V (2012) Adenoid cystic carcinoma. In: Lakhani SR et al (eds) WHO classification of tumors of the breast. IARC, Lyon, pp 56–57
Shousha S (1989) An unusual breast cyst. Histopathology 14:423–425
Sorlie T, Perou CM, Tibshirani R et al (2001) Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci U S A 98:10869–10874
Sotiriou C, Neo SY, McShane LM et al (2003) Breast cancer classification and prognosis based on gene expression profiles from a population-based study. Proc Natl Acad Sci U S A 100:10393–10398
Tavassoli FA (1999) Pathology of the breast. Appleton-Lange, Stanford
Tavassoli FA, Devilee, P (2003) WHO classification of tumours: tumours of the breast and female genital organs. In: Tavassoli FA, Devilee P (ed). IARC Press, Lyon
Tot T (2000) The cytokeratin profile of medullary carcinoma of the breast. Histopathology 37:175–181
Tseng SC, Jarvinen MJ, Nelson WG et al (1982) Correlation of specific keratins with different types of epithelial differentiation: monoclonal antibody studies. Cell 30:361–372
Weidner N, Dabbs DJ (2012) Metaplastic breast carcinoma. In: Dabbs DJ (ed) Breast pathology. Elsevier Saunders, Philadelphia, pp 479–501
Weigelt B, Kreike B, Reis-Filho JS (2009) Metaplastic breast carcinomas are basal-like breast cancers: a genomic profiling analysis. Breast Cancer Res Treat 117:273–280
Weiss RA, Eichner R, Sun TT (1984) Monoclonal antibody analysis of keratin expression in epidermal diseases: a 48- and 56-kdalton keratin as molecular markers for hyperproliferative keratinocytes. J Cell Biol 98:1397–1406
Wetzels RH, Kuijpers HJ, Lane EB et al (1991) Basal cell-specific and hyperproliferation-related keratins in human breast cancer. Am J Pathol 138:751–763
Yaziji H, Gown AM, Sneige N (2000) Detection of stromal invasion in breast cancer: the myoepithelial markers. Adv Anat Pathol 7:100–109
Zuska JJ, Crile G Jr, Ayres WW (1951) Fistulas of lactifierous ducts. Am J Surg 81:312–317
Acknowledgments
Part of this study was supported by grants from the Swedish Cancer Society and BioCARE—a National Strategic Research Program at the University of Gothenburg.
The abstract with a title: “A discrete population of p63+/K5/14+ cells implicated in the pathogenesis of salivary gland-like tumors of the breast” has been selected for presentation at the 2013 San Antonio Breast Cancer Symposium, December 10–14, 2013 in San Antonio, Texas.
Conflict of interests
The authors declare no competing financial interests.
Author information
Authors and Affiliations
Corresponding author
Additional information
The authors dedicate this work to Prof. Dr. Dr. mult. Ekkehard Grundman, the predecessor of W. Boecker on the chair of Pathology at the University of Münster.
Rights and permissions
About this article
Cite this article
Boecker, W., Stenman, G., Loening, T. et al. Squamous/epidermoid differentiation in normal breast and salivary gland tissues and their corresponding tumors originate from p63/K5/14-positive progenitor cells. Virchows Arch 466, 21–36 (2015). https://doi.org/10.1007/s00428-014-1671-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-014-1671-x