Abstract
Main conclusion
The keys to alkali-stress resistance of barren-tolerant wild soybean lay in enhanced reutilization of reserves in cotyledons as well as improved antioxidant protection and organic acid accumulation in young roots.
Abstract
Soil alkalization of farmlands is increasingly serious, adversely restricting crop growth and endangering food security. Here, based on integrated analysis of transcriptomics and metabolomics, we systematically investigated changes in cotyledon weight and young root growth in response to alkali stress in two ecotypes of wild soybean after germination to reveal alkali-resistance mechanisms in barren-tolerant wild soybean. Compared with barren-tolerant wild soybean, the dry weight of common wild soybean cotyledons under alkali stress decreased slowly and the length of young roots shortened. In barren-tolerant wild soybean, nitrogen-transport amino acids asparagine and glutamate decreased in cotyledons but increased in young roots, and nitrogen-compound transporter genes and genes involved in asparagine metabolism were significantly up-regulated in both cotyledons and young roots. Moreover, isocitric, succinic, and l-malic acids involved in the glyoxylate cycle significantly accumulated and the malate synthetase gene was up-regulated in barren-tolerant wild soybean cotyledons. In barren-tolerant wild soybean young roots, glutamate and glycine related to glutathione metabolism increased significantly and the glutathione reductase gene was up-regulated. Pyruvic acid and citric acid involved in pyruvate–citrate metabolism increased distinctly and genes encoding pyruvate decarboxylase and citrate synthetase were up-regulated. Integrated analysis showed that the keys to alkali-stress resistance of barren-tolerant wild soybean lay in enhanced protein decomposition, amino acid transport, and lipolysis in cotyledons as well as improved antioxidant protection and organic acid accumulation in young roots. This study provides new ideas for the exploitation and utilization of wild soybean resources.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Soil salinization and alkalization are widespread and important abiotic stresses worldwide that seriously affect the physiological and metabolic processes of plants, and limit their growth and yield (Zhang et al. 2022). The global saline-alkaline land area has exceeded 1 billion hectares; of this, 54% is sodic soil containing sodium bicarbonate (NaHCO3) and sodium carbonate (Na2CO3) that results in alkali stress to plants (Li et al. 2022b). Alkali stress is more damaging than salinity and greatly reduces agricultural productivity, thus impeding the development of global agriculture and leading to food security problems (Gao et al. 2022). Expansion of the sodic soil area caused by increased soil pH has become a new environmental problem (Dixit et al. 2020). Therefore, screening and identification of alkali-tolerant plant resources are imperative for the cultivation of high-quality crop varieties adapted to sodic soil and the sustainable development of future agriculture.
Wild plant resources are rich in good traits and genetic variation, and can provide candidate genes for crop improvement and breeding, helping to improve global agricultural production (Dempewolf et al. 2017; Bohra et al. 2022). Studies to enhance crop performance using wild relatives have been going on for decades and, for example, have resulted in successful breeding of wheat with leaf rust resistance, rye with multi-resistance, and cotton with high fiber (Laugerotte et al. 2022; Xu et al. 2022). Moreover, the contribution of wild relatives to the global economy is as much as 186.3 billion dollars annually (Tyack et al. 2020). Wild soybean (Glycine soja Siebold & Zucc.), a closely related wild species of cultivated soybean (Glycine max (L.) Merr.), has strong adaptability to alkali stress (Li et al. 2017). Thus, this is a natural gene pool for screening and identifying alkali-resistant genes for soybean breeding and is also ideal material for studying the mechanisms of alkali resistance in plants.
In addition to harm from osmotic effects and ion toxicity, alkali stress also has a high pH, which causes alkali burn to seeds, resulting in the loss of the selective absorption function of the cell membrane and catabolism disorder of stored substances in seeds (Han et al. 2022). In addition, growth of young roots can be more severely inhibited (Zaib-un-Nisa et al. 2022). The post-germination growth stage is the critical period for the establishment of soybean seedlings and is vulnerable to adverse environments (Du et al. 2021). Luckily, plant cotyledons have evolved a series of mechanisms to maintain the mobilization and utilization of their internal storages under saline-alkali stress, such as regulating the ratio and physiological activity of internal hormones and enhancing the activities of α-amylase, lipases, and proteases (Ashraf et al. 2009; Gogna and Bhatla 2020; Ma et al. 2022). The transport of the small molecular substances formed by the decomposition of storage compounds also improves the tolerance of hypocotyls and roots to saline-alkali stress (Chen et al. 2022c).
Young roots show various responses to alkali stress, for instance, changes in root morphology and microstructure, selective absorption and transportation of ions, accumulation and secretion of organic acids, and enhanced scavenging capacity for reactive oxygen species (ROS) (Li et al. 2019, 2022b; Gao et al. 2022; Zhao et al. 2022). Enhanced ROS scavenging capability is one of the most important mechanisms of plant’s response to alkali stress. Under alkali stress, the content of antioxidant enzymes increased significantly in the roots of Chinese cabbage, tomato, and broomcorn millet (Gong et al. 2014; Ma et al. 2021; Li et al. 2022c). Except for antioxidant enzymes, it was through metabolism of the non-enzymatic system that plants maintain the stability of intracellular ROS levels under saline-alkali stress, such as by ascorbic acid, reduced glutathione, flavonoids, and vitamins (Zhang et al. 2020; Ren et al. 2021). Previous studies showed that alfalfa and tomato removed ROS from their roots by enhancing the ascorbate–glutathione cycle under alkali stress; however, the roots of maize and Sorghum bicolor did this by flavonoid biosynthesis (Gong et al. 2014; Song et al. 2017; Ma et al. 2020; Li et al. 2022a). Thus, it is well worth investigating the mechanisms of the decomposition and utilization of cotyledon reserves and the adaptation of young roots in wild soybean under alkali stress.
With the development of “omics” in sequencing technologies and data analysis methods, the integrated analysis of multi-omics has been commonly used and has become an indispensable technique in scientific research, including genomics, epigenomics, transcriptomics, proteomics, metabolomics, and microbiomics (Pazhamala et al. 2021). Multi-omics integration analysis has provided great achievements concerning the response mechanisms of plants to abiotic stresses, such as drought, salinity, and cold. Using this method, researchers have successfully screened out candidate genes for enhancing salt-alkali tolerance and genes involved in cold stress response and recovery—these genes were targets for plant molecular improvement breeding in further studies (Raza et al. 2021; Lv et al. 2022). Multi-omics integration analysis is also a good platform for the study of complex quantitative traits controlled by multiple genes.
In this study, two different ecotypes of wild soybean were used as experimental materials, and alkali stress was simulated using a 1:1 molar ratio of Na2CO3:NaHCO3. After seed germination, cotyledon dry weight and young root length were measured for seven consecutive days. The quantities of small molecular substances and gene expression levels in cotyledons and young roots of the two wild soybeans were tested and metabolites and genes enriched in the same metabolic process were subjected to integrated analysis and comparison. This paper attempts to reveal the resistance mechanism of barren-tolerant wild soybean to alkali stress at post-germination growth stage and its gene expression regulatory mechanisms from the perspectives of mobilization and utilization of storage substances in cotyledons and the type and metabolic pathway of non-enzymatic antioxidants in young roots. The present study will provide a new standard for the evaluation and identification of barren-tolerant wild soybean germplasm resources, and provide candidate genes for the cultivation of new crop varieties with alkali tolerance.
Materials and methods
Plant materials and alkali treatment
The seeds of common wild soybean, Glycine soja Siebold & Zucc., var. Huinan06116; (GS1) and barren-tolerant wild soybean var. Tongyu06311 (GS2) used in this study were provided by Jilin Crop Germplasm Introduction and Breeding Center, Changchun, China. Healthy, plump, and uniform size seeds were selected from the seeds of two ecotypes of wild soybean as experimental materials, and their seed coats were scored. These seeds were then evenly sown in pots containing washed sand, with 10 seeds per pot. The pots were about 15 cm in diameter and 17 cm in height, with holes about 1 cm in diameter in the bottom. Before putting the sand into pots, a layer of gauze was laid on the pot bottom. The experimental materials were cultivated on June 1, 2022, in an arched shelter covered with plastic sheeting at Northeast Normal University, Changchun, China, where the day and night temperatures were 25 ± 2 and 16 ± 3 °C, respectively, and relative humidity was about 60%.
Pots seeded with GS1 and GS2 were randomly divided into control and treatment groups. The control and treatment groups were irrigated adequately with distilled water and simulated alkali solution once a day, respectively, until the end of this experiment. In simulated alkali solution, the Na2CO3:NaHCO3 was 1:1 (v/v) and Na+ concentration was 50 mmol L−1. Seeds were considered germinated when the root tip and hypocotyl broke through the seed coat by 2 mm (Ducatti et al. 2022); the overwhelming majority of seeds had germinated 1 day after sowing. After 5 days of seed germination, three pots were randomly selected from each group. After removing seed coats and isolating cotyledons and young roots, RNA was extracted from cotyledons and young roots separately for transcriptome analysis. Four pots were randomly selected from each group for metabolome analysis of cotyledons and young roots at 6 days after seed germination, and the preliminary preparations were the same as above. For transcriptomic and metabolomic analyses, cotyledons or young roots from the same pot were considered as one biological duplicate. All harvested fresh samples were quickly frozen in liquid nitrogen and then stored at − 80 °C.
Measurements of cotyledon weight and young root length
The dry weight of cotyledons and the length of young roots of two wild soybeans under control and alkali stress conditions were measured for seven consecutive days from the day of germination. Three pots were randomly selected from each group for measurements every day, with each measurement repeated three times.
Metabolomics analysis
Of cotyledon and young root samples of the two wild soybeans under control and alkali stress, 50 mg was transferred into new Eppendorf tubes. Into each tube, 500 μL of extracting solution (Vmethanol/Vchloroform = 3:1, v/v) and 60 μL of ribitol were added. After blending, the tubes were put into ice water and sonicated for 5 min, and then centrifuged for 10 min at 12,000g and 4 °C. The supernatants were transferred into glass bottles specially designed for a gas chromatograph–mass spectrometer (GC–MS). The glass bottles were dried in a vacuum concentrator for 30 min, and then 120 μL of methoxyamine hydrochloride was added to them. After vortex mixing, they were derivatized in an oven at 37 °C for 120 min. Lastly, 100 μL of N,O-bis-(trimethylsilyl)-trifluoroacetamide reagent (containing 1% trimethylchlorosilane) was added to the dried samples and they were oscillated at 70 °C for 1 h. After dropping to room temperature, samples were tested and analyzed by Agilent 7890 GC–MS (Agilent Technologies, Santa Clara, CA, USA). Four biological replicates were randomly selected from each group for analysis each time, and the analysis was repeated three times.
ChromaTOF software (V 4.3x, LECO, St. Joseph, MI, USA), and EI-MS and FiehnLib databases were used to process data and identify metabolites (Chen et al. 2022a). Principal component analysis (PCA) and partial least-squares discriminant analysis (PLS-DA) were completed by SIMCA-P13.0 software (Umetrica, Umea, Sweden), and variable importance in projection (VIP) values were also calculated (Chen et al. 2022a). Student’s t test was performed using Excel 2013. The differential metabolites were defined and screened according to P < 0.05, VIP > 1, and similarity value > 500. The metabolic pathways of metabolites were queried using KEGG (https://www.genome.jp/kegg) and MetaboAnalyst 5.0 (https://www.metaboanalyst.ca).
Transcriptomics analysis
The total RNA from cotyledon and young root samples was extracted by TRIzol kit (Invitrogen, Carlsbad, CA, USA), and the concentration and integrity of extracted RNA were separately determined by NanoDrop 2000 (Thermo Scientific Inc., Wilmington, DE, USA) and Agilent Bioanalyzer 2100 system (Agilent Technologies). The NEBNext® Ultra™ RNA Library Prep Kit of Illumina® (NEB, USA) was used to generate cDNA sequencing libraries, and the quality of libraries was assessed by Agilent Bioanalyzer 2100 system. The library fragments were then sequenced on an Illumina Hiseq platform based on the principle of sequencing while synthesizing, and raw reads were generated. All the above steps were repeated three times, and three biological replicates were randomly selected from each group at a time.
Clean reads were obtained by removing adapters’ reads and reads with low-quality, and their Q30 percentage and GC content were further calculated. Then, the clean reads were mapped to the reference genome Williams 82 for analysis and annotation of exact matches. The differential expression analysis of cotyledons or young roots under control and alkali stress conditions was completed using the DESeq R package (1.10.1), and genes with P < 0.05 and |log2FC| ≥ 1 were defined as differentially expressed genes (DEGs) (Chen et al. 2022a), where FC (fold change) refers to the ratio of expression levels between alkali stress and the control. The expression levels of genes were expressed as FPKM values, which were equal to cDNA fragments/mapped fragments (millions) × transcript length (kb) (Florea et al. 2013). Finally, the GO enrichment and KEGG pathway enrichment of DEGs were determined using the GOseq R program package and KEGG website.
Real-time fluorescence quantitative PCR validation
Eight cotyledon genes and ten young root genes were randomly selected from the two wild soybeans, and their expression levels under alkali stress and control conditions were determined using real-time fluorescence quantitative PCR (qRT-PCR). Primers were designed by Primer Premier 6 software (Premier Biosoft International, PaloAlto, CA, USA), and their specificities were evaluated by TBtools software (https://www.tbtools.com). The designed primers sequences were listed in Tables S1 and S2. Total RNA was isolated from frozen samples by the TRUEscript RT MasterMix (DF Biotech, Chengdu, China) dedicated to qRT-PCR. The cDNA synthesis was performed according to the manufacturer’s instructions of TransScript First-Strand cDNA Synthesis kit (AiDLAB Biotech, Beijing, China). With 18S rRNA (GYMA06G315500WM82A2V1) as the internal reference gene, qRT-PCR was performed on an Analytik Jena-qTOWER2.2 fluorescence quantitative PCR instrument (Analytik Jena AG, Jena, Germany) (Chen et al. 2022a). All the above operations were repeated three times, and three loading sample points were set for each gene every time. Then, the obtained data were analyzed according to comparative CT method.
Integrated analysis of metabolomics and transcriptomics
Metaboanalyst 5.0 was used to calculate the correlation coefficients between metabolites and genes in the same metabolic process. Using these correlation coefficients, correlation networks were plotted using Cytoscape software (https://www.cytoscape.org). Visio 15.0 was used to visualize the mechanisms of resistance to alkali stress in cotyledons and young roots of barren-tolerant wild soybean.
Statistical analysis
All data were analyzed using Excel 2013. The differences between treatments were evaluated using Student’s t test, with P < 0.05 considered significant.
Results
Changes in cotyledon weight and young root growth
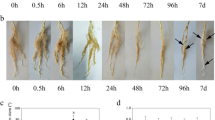
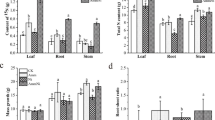
The cotyledon dry weight of both ecotypes of wild soybean showed a gradual decline during germination (Fig. 1a). However, the decreasing trend of cotyledon dry weight significantly differed between the two ecotypes and between control and alkali stress. Under alkali stress, the reduction in cotyledon dry weight was greater for barren-tolerant than common wild soybean, and there were also significant differences in cotyledon dry weight at 7 days post-germination. Without stress, the dry weight of GS1 and GS2 cotyledons at 7 days was reduced by 20.9% and 21.9% compared with that at 1 day, respectively, while under stress, they were reduced by 16.7% and 19.3% (P < 0.05). The alkali stress inhibited the growth of young roots, and the degree of inhibition increased gradually with longer stress time (Fig. 1b). At 7 days, compared with control, the young root length of GS1 was significantly decreased by 0.60-fold, while that of GS2 was only decreased by 0.47-fold (P < 0.01).
Multi-omics analysis of cotyledons
Metabolome analysis
The metabolites in cotyledons of two ecotypes of wild soybean under control and alkali stress conditions were identified and quantified by GC–MS. In the PCA, the first principal component (PC1) obviously distinguished GS1 from GS2 (Fig. 2a), explaining 44.5% of the variance—the second principal component (PC2) clearly separated control and treatment groups (Fig. 2a), explaining 26.2% of total variance. The PLS-DA indicated that metabolites with more contribution to PC1 included l-malic acid, d-arabitol, and sorbose, and the metabolites contributing to PC2 included citric acid, oxoproline, sucrose, and alanine (Fig. 2b, Table S3). In total, 21 kinds of differential metabolites were identified and quantified according to the screening criteria: nine amino acids, one non-amino acid nitrogenous compound, two lipids, five organic acids, and four sugars (Table 1).
Principal component analysis and loading plots of metabolites in two wild soybean cotyledons and young roots under control and alkali stress conditions. a Principal component analysis of metabolites in cotyledon. b Loading plots of metabolites in cotyledon. c Principal component analysis of metabolites in young root. d Loading plots of metabolites in young root
Under alkali stress, compared with control, the relative content of glutamine increased in the cotyledons of both wild soybeans but increased more in GS2, with a significant increase of 1.36-fold (P < 0.01). More interestingly, nitrogenous compounds threonine, tryptophan, phenylalanine, asparagine, glutamic acid, alanine and 4-aminobutyric acid significantly increased by 0.40-, 0.89-, 0.84-, 0.24-, 0.38-, 0.14-, and 0.63-fold, respectively, in GS1 cotyledons (P < 0.05), but decreased by 0.45-, 0.15-, 0.34-, 0.18-, 0.68-, 0.29-, and 0.15-fold in GS2.
In comparison with control, under alkali stress, linoleic acid, diglycerol, l-malic acid, succinic acid, and fumaric acid, related to the hydrolysis of stored lipids and glyoxylate cycle, increased in both wild soybean cotyledons but increased more in GS2, with increases of 0.93-, 0.31-, 1.58-, 0.72-, and 0.86-fold, respectively (P < 0.01). The relative contents of citric and isocitric acids, involved in the glyoxylate cycle, increased by 0.25- and 0.29-fold in GS2 cotyledons but decreased by 0.32- and 0.20-fold in GS1, respectively (P < 0.05).
Transcriptome analysis
The gene expression levels in cotyledons of two wild soybeans under control and alkali stress conditions were determined using RNA sequencing (RNA-seq). A total of 78.74 Gb clean data was obtained and the clean data of each sample reached 5.89 Gb. The percentages of Q30 and GC of the clean data were 95.51–96.29% and 44.8–45.36%, respectively. The sequence alignment efficiency between clean data and reference genome Williams 82 ranged within 92.91–95.74% (Table S4). The expression levels of eight cotyledon genes of both ecotypes of wild soybeans determined by qRT-PCR under control and alkali stress conditions were consistent with RNA-seq results, which verified the accuracy of the transcriptome data and confirmed that it could be further analyzed (Fig. S1).
Under alkali stress, 997 (548 up-regulated and 449 down-regulated) and 1122 DEGs (611 up-regulated and 511 down-regulated) were screened and identified in the GS1 and GS2 cotyledons, respectively (Fig. S2). Among them, 782 DEGs of GS1 cotyledons and 944 DEGs of GS2 cotyledons were annotated. The results showed that they were mainly enriched in biological processes such as response to salt stress (GO:0009651), protein metabolic process (GO:0019538), and lipid metabolic process (GO:0006629); cellular component such as intracellular organelle (GO:0043229); and molecular function such as hydrolase activity (GO:0016787) and transporter activity (GO:0005215) (Fig. S3a, b). Then, the top 20 KEGG pathways annotated by DEGs of the two wild soybean ecotypes were analyzed. The results indicated that nitrogen metabolism (ko00910) was the metabolic pathway co-enriched by DEGs of both wild soybeans, but glyoxylate and dicarboxylate metabolism (ko00630) was the metabolic pathway specifically enriched by DEGs of barren-tolerant wild soybean (Fig. S3c, d).
Compared with the control, under alkali stress, a serine protease gene involved in the decomposition of storage protein, was down-regulated by 1.15-fold in GS2 cotyledons. However, in GS1 cotyledons, this protease was encoded by another gene and significantly down-regulated by 2.39-fold. Three genes encoding serine protease inhibitors were up-regulated in cotyledons of both wild soybeans but the up-regulation was greater in GS1, with 2.00-, 2.60-, and 2.40-fold up-regulation, respectively. In addition, an asparagine synthase gene, involved in amino acid transport was up-regulated by 1.03-fold in GS2 cotyledons, under alkali stress compared with control, but there was no significant change in GS1. Moreover, genes responsible for transport of nitrogenous compounds were up-regulated by 1.27- and 1.09-fold in GS2 cotyledons. In GS1 cotyledons, another gene was responsible for this process but was significantly down-regulated by 1.54-fold (Table 2).
Under alkali treatment, compared with untreated samples, triacylglycerol lipase gene, phospholipase gene, diacylglycerol acyltransferase gene, and long-chain acyl-CoA synthetase gene involved in lipolysis, were significantly up-regulated by 1.48-, 1.31-, 1.08-, and 1.13-fold in GS2 cotyledons, respectively. However, in GS1 cotyledons, there were no significant changes in expression levels of genes other than one phospholipase gene. Moreover, the gene encoding malate synthase, the key enzyme of the glyoxylate cycle, was up-regulated by 1.15-fold in GS2 cotyledons but with no significant change in GS1 (Table 2).
Integrated analysis of transcriptome and metabolome
Integrated metabolome and transcriptome analysis showed that in GS2 cotyledons, the correlation coefficients of glutamine and asparagine with asparagine synthetase gene were 0.67 and − 0.93, respectively (P < 0.05). There were significant positive correlations of nitrogenous compounds alanine, serine, threonine, asparagine, oxoproline, 4-aminobutyric acid, glutamic acid, phenylalanine, glutamine and tryptophan with nitrogenous compound transporter genes in GS2 cotyledons, with correlation coefficients of 0.62–0.99 (P < 0.05). In addition, citric, isocitric, succinic, and l-malic acids in GS2 cotyledons were positively correlated with the gene encoding malate synthase of the glyoxylic acid cycle, with correlation coefficients of 0.95, 0.93, 0.96, and 0.93, respectively (P < 0.01) (Fig. 3, Table S5). However, in GS1 cotyledons, the correlations between genes and metabolites involved in the same metabolic process were weak and non-significant.
Integration network of differential metabolites and differential genes in barren-tolerant wild soybean under alkali stress. The dotted line represents a negative correlation and the solid line represents a positive correlation. The thicker the edge is, the stronger the correlation is. The size of a node is proportional to the correlation between nodes
Multi-omics analysis of young roots
Metabolome analysis
GC–MS was applied to identify and quantify the metabolites in young roots of two ecotypes of wild soybean under control and alkali stress. The PC1 of the PCA distinguished the treated groups from the control (Fig. 2c), explaining 42.9% of the variation, and PC2 separated the two wild soybeans (Fig. 2c), explaining 25.4% of the variation. The PLS-DA showed that l-malic acid, fructose, sorbose, and aspartic acid contributed more to PC1, and mannose, tagatose, and asparagine contributed more to PC2 (Fig. 2d, Table S6). On the basis of the screening standard, 25 differential metabolites were identified and quantified: seven amino acids, three non-amino acid nitrogenous compounds, two lipids, seven organic acids, and six sugars and their derivatives (Table 3).
Under alkali stress, compared with the control, nitrogenous compounds tryptophan, glycine, asparagine, glutamine, and 4-aminobutyric acid significantly decreased by 0.54-, 0.43-, 1.34-, 1.00-, and 0.47-fold in GS1 young roots, respectively (P < 0.05), but increased by 0.47-, 0.54-, 0.14-, 1.27-, and 0.18-fold in GS2. In addition, the relative contents of aspartic acid, glutamic acid, and oxoproline increased in young roots of both wild soybeans but increased more in GS2, with 5.21-, 3.52-, and 1.93-fold increases, respectively (P < 0.01).
Compared with the control, under alkali stress, the relative contents of sucrose, pyruvic acid, citric acid, l-malic acid and glutaric acid associated with pyruvate–citrate metabolism, increased in the young roots of both wild soybeans, but the increase was greater in GS2, with 4.77-, 1.61-, 2.88-, 1.99-fold, and 1.84-fold increases, respectively (P < 0.05).
Transcriptome analysis
After transcriptome analysis of young roots of two wild soybeans under control and alkali stress using RNA-seq, 92.89 Gb of clean data were obtained, with the clean data of each sample reaching 6.07 Gb. The Q30 and GC proportions of clean data were 95.47–96.06% and 45.25–45.86%, respectively. The sequence alignment efficiency between clean data and the reference genome was in the range of 91.37–94.01% (Table S7). The FPKM values of each data point ranged from 10−2 to 104, and the correlations between the expression levels of each biological replicate were relatively high. The expression levels of ten genes in young roots of two wild soybeans determined by qRT-PCR under control and alkali stress conditions were consistent with the transcriptome data (Fig. S4). So, it was feasible and credible to further analyze transcriptome data.
Under alkali stress, 1795 DEGs (1659 up-regulated and 136 down-regulated) and 2717 DEGs (2239 up-regulated and 478 down-regulated) were screened and identified in the young roots of GS1 and GS2, respectively (Fig. S5); correspondingly 1442 and 2197 DEGs were annotated. The GO annotation results of DEGs showed the enriched GO terms included primary metabolic process (GO:0044238), response to stimulus (GO:0050896) and response to oxidative stress (GO:0006979) in biological process, organelle (GO:0043226) in cellular component, and antioxidant activity (GO:0016209) in molecular function (Fig. S6a, b). The top 20 KEGG pathways annotated by DEGs of two wild soybeans were further analyzed. The results showed that the metabolic pathway co-enriched by DEGs of two wild soybeans was glutathione metabolism (ko00480), but more DEGs were enriched in this pathway in barren-tolerant wild soybean (Fig. S6c, d).
In comparison with the control, under alkali stress, nitrogen-compound transporter genes were up-regulated by 3.67- and 3.49-fold in GS2 young roots, but with no significant change for GS1. In addition, two asparaginase genes involved in asparagine catabolism were up-regulated in the young roots of both wild soybeans, with greater up-regulation in GS2, and up-regulated by 7.27- and 4.72-fold, respectively. Moreover, in GS2 young roots, the gene encoding glutathione reductase of glutathione metabolism was significantly up-regulated by 1.12-fold but there was no significant change for GS1 (Table 4).
Under alkali stress, compared with control, two pyruvate decarboxylase genes and genes encoding citrate synthase, 2-oxoglutarate dehydrogenase, and malate dehydrogenase of the citric acid cycle were significantly up-regulated by 5.42-, 2.85-, 3.50-, 1.90-, and 1.19-fold in GS2 young roots, respectively. However, in GS1 young roots, except for the gene encoding citrate synthase, there were no significant changes in other genes (Table 4).
Integrated analysis of transcriptome and metabolome
The integrated analysis of metabolomics and transcriptomics showed significant positive correlations of nitrogenous compounds tryptophan, glycine, asparagine, glutamine, aspartic acid, glutamic acid, oxoproline, ethanolamine, 4-aminobutyric acid and adipamide in GS2 young roots with two nitrogenous compound transporter genes, with correlation coefficients of 0.54–0.97 (P < 0.05). Asparagine and aspartic acid in GS2 young roots were positively correlated with two asparaginase genes, with correlation coefficients of 0.97 and 0.96, and 0.97 and 0.95, respectively (P < 0.01). Both glycine and glutamate had strong correlations with glutathione reductase gene of glutathione metabolism, with correlation coefficients of 0.95 and 0.97, respectively (P < 0.01). Moreover, pyruvic and citric acids in GS2 young roots were highly correlated with two pyruvate decarboxylase genes and citrate synthase gene, with correlation coefficients of 0.80, 0.82, and 0.93 for pyruvic acid and 0.83, 0.74, and 0.91 for citric acid, respectively (P < 0.05) (Fig. 3, Table S8). However, in GS1 young roots, correlations between genes and metabolites in the same metabolic process were weak and non-significant.
Discussion
Soil salinization and alkalization are some of the most serious abiotic stresses threatening agricultural development and affect the growth and development of plants throughout their whole life cycle, especially at early stages (Yi et al. 2022). Previous studies have shown that saline-alkali stress reduces the water absorption of seeds, restricts the mobilization and utilization of storage reserves, and inhibits the growth of young roots (Marques et al. 2013; Nimir et al. 2020; Zaib-un-Nisa et al. 2022). Consistent with these findings, this study showed that alkali stress inhibited the reduction in cotyledon dry weight and the growth of young roots of both wild soybeans compared with the control, and the inhibitions became serious with longer stress time. However, compared with GS2, the inhibitions suffered by GS1 were more severe. This suggested that under alkali stress, barren-tolerant wild soybean had a stronger ability to mobilize storage substances and maintain young root growth at the post-germination growth stage, compared with common wild soybean. This might be related to differences in physiology and gene regulation between the two wild soybean ecotypes.
The decomposition and reuse of stored proteins are key events in seed germination and post-germination growth stages (Ehrhardt-Brocardo and Coelho 2022). Cysteine-, serine-, aspartic-, metallo-, and threonine-proteases play important roles in decomposition of storage proteins (Liu et al. 2018), but their catalytic activities can be inhibited under abiotic stress. The germinated seeds showed an increase in the abundance of protease inhibitors as well as a decrease in activities of proteases, under stress, which slowed the catabolism of stored proteins (Fercha et al. 2014; Ponte et al. 2014). A recent study found that a salt-tolerant sorghum variety could actively mobilize stored proteins in seeds by maintaining relatively higher protease activity under salt stress, thus alleviating the inhibition of salt stress on its germination and seedling growth (Punia et al. 2021). In the present study, compared with GS2, the serine protease gene was down-regulated more severely and up-regulation of serine protease inhibitor genes was greater in GS1 cotyledons. The total content of amino acids was decreased in GS2 cotyledons but increased in young roots, and the opposite was true for GS1. In addition, nitrogen-compound transporter genes were significantly up-regulated in both cotyledons and young roots of GS2, and there were significant positive correlations between amino acids and nitrogen-compound transport genes, in both cotyledons and young roots. Asparagine is the major nitrogen-transport amino acid due to its stability and high nitrogen/carbon ratio—the enzyme catalyzing its synthesis is asparagine synthase, which catalyzes the transfer of the amide group of glutamine to aspartate, yielding asparagine and participating in the mobilization of reserve nitrogen resources (García-Calderón et al. 2017). After asparagine is transported to sink tissues, it is mainly hydrolyzed to aspartic acid and ammonium by asparaginase, and the released ammonium can be re-assimilated by glutamate to generate other amino acids (Gaufichon et al. 2016). Under abiotic stress, asparagine metabolism in sensitive plants is significantly suppressed during germination, obstructing the mobilization and reutilization of stored nitrogen (Fercha et al. 2014). The enhancement of asparagine metabolism in salt-tolerant Bermuda grass facilitated the transport of nitrogen sources to the developing young roots, and improved its growth under salt stress (Hu et al. 2015). Our study showed that under alkali stress, glutamine increased more in GS2 cotyledons than in GS1 cotyledons, and the asparagine synthetase gene was significantly up-regulated. The integrated analysis showed that glutamine in GS2 cotyledon was positively correlated with asparagine synthetase gene. In addition, under alkali stress, compared with GS1, asparagine and aspartic acid increased significantly in GS2 young roots and up-regulation of asparaginase genes was greater. The integrated analysis showed positive correlations of asparagine and aspartic acid with asparaginase genes in GS2 young roots. The above results suggest that under alkali stress, barren-tolerant wild soybean had stronger ability to decompose protein and transport amino acids compared with common wild soybean, and the relative content of amino acids in cotyledons and young roots was due to amino acid transport rather than protein decomposition.
The decomposition of stored fat in cotyledons of leguminous plants is also a key event during the post-germination growth stage. After seed germination, fatty acids are released from stored fats by hydrolysis of lipases and, after activation, they are converted into acetyl-CoAs by β-oxidation and subsequently enter the glyoxylate cycle (Eastmond and Graham 2001). The glyoxylate cycle is a bridge linking fatty acid, energy, and amino acid metabolism, and acetyl-CoAs from stored fat decomposition are eventually converted into malate and succinate by isocitrate lyase and malate synthetase, providing energy for life activities and carbon skeletons for amino acid synthesis (Eastmond and Graham 2001). Previous studies have shown that under abiotic stresses such as drought and salt, the activities of key enzymes in the glyoxylate cycle significantly decreased (Sidari et al. 2008). Because of enhancement of the glyoxylate cycle, salt-tolerant castor and rice alleviated the inhibition of salt stress on fat decomposition and energy metabolism during seed germination (Asadi Aghbolaghi et al. 2022). Exogenous melatonin promoted the catabolic metabolism of storage lipids by enhancing the glyoxylic acid cycle, and ensured energy supply for cucumber seed germination and subsequent growth under stress (Zhang et al. 2017). Our study results showed that, under alkali stress, compared with GS1, glyoxylate cycle intermediates citric, isocitric, succinic, and malic acids increased more in GS2 cotyledons, and the malate synthase gene was significantly up-regulated. Moreover, the integrated analysis showed that glyoxylate cycle intermediates were positively correlated with the malate synthetase gene in GS2 cotyledons. This indicated that under alkali stress, compared with common wild soybean, barren-tolerant wild soybean could promote the catabolism of stored lipids by enhancing the glyoxylic acid cycle, which was conducive to providing energy security for subsequent life activities and maintaining the carbon and nitrogen balance.
Alkali stress induces overproduction of ROS and oxidative stress in plants. Glutathione metabolism is thought to play a central role in scavenging ROS, and its metabolic intensity is closely related to plant tolerance to saline-alkali stress, as well demonstrated in studies of alfalfa, maize, and wheat (Zhou et al. 2017; Lou et al. 2018). Glutathione is a small-molecule antioxidant synthesized from l-glutamic acid, l-cysteine, and glycine through two successive reactions and it can react directly with ROS to reduce cell damage caused by membrane lipid peroxidation (Zhou et al. 2017; Song et al. 2021). After the removal of ROS, reduced glutathione (GSH) is converted to oxidized glutathione (GSSG), which in turn can be reduced to GSH by glutathione reductase (GR); therefore, the ratio of GSH/GSSG is a key indicator to measure the intensity of glutathione metabolism (Ren et al. 2021). Under alkali stress, the glutathione metabolism of rice was disordered, leading to a distinct decrease in GSH/GSSG ratio and ROS scavenging capacity (Ren et al. 2021). Unlike the study of rice, our study showed under alkali stress, compared with GS1, glutamic acid and glycine increased significantly and the GR gene was up-regulated in GS2 young roots. In addition, glutamate and glycine in GS2 young roots were significantly positively correlated with the GR gene. This suggested that under alkali stress, compared with common wild soybean, barren-tolerant wild soybean could remove ROS from young roots by enhancing glutathione metabolism and improving the intracellular GSH/GSSG ratio.
Alkali stress causes high pH stress, hence, accumulation of organic acids is a key mechanism for plants to adapt to alkali stress (Li et al. 2022b; Song et al. 2021). However, the types of organic acids accumulated in plants vary among species and their regulatory mechanisms are also quite different. Under alkali stress, alkali-resistant gramineous plants mainly accumulated citric acid, while sea buckthorn mainly accumulated acetate (Yang et al. 2007; Chen et al. 2009). Oxalic acid was the main organic acid accumulated in grapevine and Kochia sieversiana roots under alkali stress, but phenolic acid was the main organic acid accumulated in roots of Suaeda salsa and Puccinellia tenuiflora (Yang et al. 2007; Guo et al. 2018; Chen et al. 2022b). The strategy of oats and barley to cope with alkali stress was enhancement of the TCA cycle, while the key of wheat resistance to alkali stress was malate metabolism (Han et al. 2019; Gao et al. 2022). Sea buckthorn enhanced its tolerance to alkali stress by increasing the rate of anaerobic respiration, but grapevine achieved this goal by promoting pyruvic acid and citric acid metabolisms, and Puccinellia tenuiflora by promoting fatty acid and phenolic acid metabolisms (Chen et al. 2009; Guo et al. 2018; Li et al. 2022b). In this study, compared with GS1, under alkali stress, pyruvic and citric acids were increased more and genes encoding pyruvate decarboxylase and citrate synthase were significantly up-regulated in GS2 young roots. The integrated analysis showed that pyruvic and citric acids in GS2 young roots were positively correlated with genes encoding pyruvate decarboxylase and citrate synthase. Thus, compared with common wild soybean, barren-tolerant wild soybean improved its tolerance to alkali stress by enhancing pyruvate–citrate metabolism.
Conclusion
Alkali stress severely inhibited the growth of plants after germination, mainly due to restriction of decomposition and transport of storage compounds in seeds. Meanwhile, nutrient deficiency and ROS damage in young roots were also the causes of growth inhibition under alkali stress. Alkali-tolerant plants or varieties can deal with alkali stress by adjusting physiological metabolism during seed germination. Through the measurement of growth parameters after germination and the comprehensive analysis and comparison of metabolomics and transcriptomics, this study revealed that under alkali stress, barren-tolerant wild soybean guaranteed the mobilization and reutilization of reserves in cotyledons by promoting asparagine metabolism, nitrogen compound transport, and the glyoxylic acid cycle. In addition, enhanced glutathione and pyruvate-citrate metabolism were key for barren-tolerant wild soybean to protect its young roots from damage caused by ROS and high pH stress induced by alkali stress (Fig. 4). This study will provide new insight for the cultivation of new varieties of alkali-tolerant crops and lay a theoretical foundation for the rational development and protection of wild germplasm resources.
Key mechanisms of resistance to alkali stress in two ecotypes of wild soybean. GS1 common wild soybean, GS2 barren-tolerant wild soybean. The darker the fill color of the ellipse, the greater the fold change value of metabolite. TGL triacylglycerol lipase, DGAT diacylglycerol acyltransferase, LACS long chain acyl-CoA synthetase, CS citrate synthase, ACO aconitate hydratase, aceA isocitrate lyase, aceB malate synthase, MDH malate dehydrogenase, ASN asparagine synthetase, NSE asparaginase, GLUL glutamine synthetase, GCLC glutamate–cysteine ligase catalytic subunit, GSS glutathione synthase, GSR glutathione reductase, GSH glutathione, GSSG glutathione disulfide, PDC pyruvate decarboxylase, IDH isocitrate dehydrogenase, OGDH 2-oxoglutarate dehydrogenase
Author contribution statement
XNW, YNH, TZ, and LXS conceived and designed research. XNW, YNH, YMW, and SJG conducted experiments. XNW and YNH analyzed data. XNW, JXG and LXS wrote the manuscript. XNW and YDW revised and proofread the manuscript. All authors read and approved the manuscript.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- DEGs:
-
Differentially expressed genes
- GS1:
-
Wild soybean var. Huinan06116
- GS2:
-
Wild soybean var. Tongyu06311
- ROS:
-
Reactive oxygen species
References
Asadi Aghbolaghi M, Sedghi M, Sharifi RS, Dedicova B (2022) Germination and the biochemical response of pumpkin seeds to different concentrations of humic acid under cadmium stress. Agriculture 12(3):374. https://doi.org/10.3390/agriculture12030374
Ashraf MY, Ashraf M, Naveed NH, Akram NA, Arshad M (2009) Salt-induced biochemical changes in germinating seeds of three rice cultivars differing in salt tolerance. Agrochimica LIII 53(5):308–321
Bohra A, Kilian B, Sivasankar S, Caccamo M, Mba C, McCouch SR, Varshney RK (2022) Reap the crop wild relatives for breeding future crops. Trends Biotechnol 40(4):412–431. https://doi.org/10.1016/j.tibtech.2021.08.009
Chen WC, Cui PJ, Sun HY, Guo WQ, Yang CW, Jin H, Fang B, Shi DC (2009) Comparative effects of salt and alkali stresses on organic acid accumulation and ionic balance of seabuckthorn (Hippophae rhamnoides L). Ind Crops Prod 30(3):351–358. https://doi.org/10.1016/j.indcrop.2009.06.007
Chen J, Zhou J, Li MX, Li M, Hu YN, Zhang T, Shi LX (2022a) Membrane lipid phosphorus reusing and antioxidant protecting played key roles in wild soybean resistance to phosphorus deficiency compared with cultivated soybean. Plant Soil 474(1–2):99–113. https://doi.org/10.1007/s11104-022-05316-5
Chen Q, Xie HS, Wei GY, Guo XR, Zhang J, Lu XY, Tang ZH (2022b) Metabolic differences of two constructive species in saline-alkali grassland in China. BMC Plant Biol 22(1):53. https://doi.org/10.1186/s12870-021-03401-y
Chen XF, Zhang RD, Li B, Cui T, Liu C, Liu CJ, Chen BR, Zhou YF (2022c) Alleviation of oxidative damage induced by CaCl2 priming is related to osmotic and ion stress reduction rather than enhanced antioxidant capacity during germination under salt stress in sorghum. Front Plant Sci 13:881039. https://doi.org/10.3389/fpls.2022.881039
Dempewolf H, Baute G, Anderson J, Kilian B, Smith C, Guarino L (2017) Past and future use of wild relatives in crop breeding. Crop Sci 57(3):1070–1082. https://doi.org/10.2135/cropsci2016.10.0885
Dixit VK, Misra S, Mishra SK, Tewari SK, Joshi N, Chauhan PS (2020) Characterization of plant growth-promoting alkalotolerant Alcaligenes and Bacillus strains for mitigating the alkaline stress in Zea mays. Anton Leeuw Int J G 113(7):889–905. https://doi.org/10.1007/s10482-020-01399-1
Du CX, Li H, Liu C, Fan HF (2021) Understanding of the postgerminative development response to salinity and drought stresses in cucumber seeds by integrated proteomics and transcriptomics analysis. J Proteomics 232:104062. https://doi.org/10.1016/j.jprot.2020.104062
Ducatti KR, Batista TB, Hirai WY, Luccas DA, Moreno LA, Guimaraes CC, Bassel GW, Silva EAA (2022) Transcripts expressed during germination Sensu Stricto are associated with vigor in soybean seeds. Plants 11(10):1310. https://doi.org/10.3390/plants11101310
Eastmond PJ, Graham IA (2001) Re-examining the role of the glyoxylate cycle in oilseeds. Trends Plant Sci 6(2):72–78. https://doi.org/10.1016/S1360-1385(00)01835-5
Ehrhardt-Brocardo NCM, Coelho CMM (2022) Mobilization of seed storage proteins is crucial to high vigor in common bean seeds. Cienc Rural 52:e20200894. https://doi.org/10.1590/0103-8478cr20200894
Fercha A, Capriotti AL, Caruso G, Cavaliere C, Samperi R, Stampachiacchiere S, Laganà A (2014) Comparative analysis of metabolic proteome variation in ascorbate-primed and unprimed wheat seeds during germination under salt stress. J Proteom 108:238–257. https://doi.org/10.1016/j.jprot.2014.04.040
Florea L, Song L, Salzberg SL (2013) Thousands of exon skipping events differentiate among splicing patterns in sixteen human tissues. F1000esearch 2:188.v2. https://doi.org/10.12688/f1000research.2-188.v2
Gao YG, Jin YL, Guo W, Xue YW, Yu LH (2022) Metabolic and physiological changes in the roots of two oat cultivars in response to complex saline-alkali stress. Front Plant Sci 13:835414. https://doi.org/10.3389/fpls.2022.835414
García-Calderón M, Pérez-Delgado CM, Credali A, Vega JM, Betti M, Márquez AJ (2017) Genes for asparagine metabolism in Lotus japonicus: differential expression and interconnection with photorespiration. BMC Genom 18(1):781. https://doi.org/10.1186/s12864-017-4200-x
Gaufichon L, Rothstein SJ, Suzuki A (2016) Asparagine metabolic pathways in Arabidopsis. Plant Cell Physiol 57(4):675–689. https://doi.org/10.1093/pcp/pcv184
Gogna M, Bhatla SC (2020) Salt-tolerant and -sensitive seedlings exhibit noteworthy differences in lipolytic events in response to salt stress. Plant Signal Behav 15(4):1737451. https://doi.org/10.1080/15592324.2020.1737451
Gong B, Wen D, Bloszies S, Li X, Wei M, Yang FJ, Shi QH, Wang XF (2014) Comparative effects of NaCl and NaHCO3 stresses on respiratory metabolism, antioxidant system, nutritional status, and organic acid metabolism in tomato roots. Acta Physiol Plant 36(8):2167–2181. https://doi.org/10.1007/s11738-014-1593-x
Guo SH, Niu YJ, Zhai H, Han N, Du YP (2018) Effects of alkaline stress on organic acid metabolism in roots of grape hybrid rootstocks. Sci Hortic 227:255–260. https://doi.org/10.1016/j.scienta.2017.09.051
Han L, Xiao CX, Xiao BB, Wang M, Liu JT, Bhanbhro N, Khan A, Wang H, Wang H, Yang CW (2019) Proteomic profiling sheds light on alkali tolerance of common wheat (Triticum aestivum L.). Plant Physiol Biochem 138:58–64. https://doi.org/10.1016/j.plaphy.2019.02.024
Han PL, Li SX, Yao KS, Geng HY, Liu JY, Wang YN, Lin JX (2022) Integrated metabolomic and transcriptomic strategies to reveal adaptive mechanisms in castor plant during germination stage under alkali stress. Environ Exp Bot 203:105031. https://doi.org/10.1016/j.envexpbot.2022.105031
Hu LX, Chen L, Liu L, Lou YH, Amombo E, Fu JM (2015) Metabolic acclimation of source and sink tissues to salinity stress in bermudagrass (Cynodon dactylon). Physiol Plant 155(2):166–179. https://doi.org/10.1111/ppl.12312
Laugerotte J, Baumann U, Sourdille P (2022) Genetic control of compatibility in crosses between wheat and its wild or cultivated relatives. Plant Biotechnol J 20(5):812–832. https://doi.org/10.1111/pbi.13784
Li MX, Guo R, Jiao Y, Jin XF, Zhang HY, Shi LX (2017) Comparison of salt tolerance in soja based on metabolomics of seedling roots. Front Plant Sci 8:1101. https://doi.org/10.3389/fpls.2017.01101
Li JS, Hussain T, Feng XH, Guo K, Chen HY, Yang C, Liu XJ (2019) Comparative study on the resistance of Suaeda glauca and Suaeda salsa to drought, salt, and alkali stresses. Ecol Eng 140:105593. https://doi.org/10.1016/j.ecoleng.2019.105593
Li CX, Jia Y, Zhou RY, Liu LW, Cao MN, Zhou Y, Wang ZH, Di H (2022a) GWAS and RNA-seq analysis uncover candidate genes associated with alkaline stress tolerance in maize (Zea mays L.) seedlings. Front Plant Sci 13:963874. https://doi.org/10.3389/fpls.2022.963874
Li H, Xu CY, Han L, Li CY, Xiao BB, Wang H, Yang CW (2022b) Extensive secretion of phenolic acids and fatty acids facilitates rhizosphere pH regulation in halophyte Puccinellia tenuiflora under alkali stress. Physiol Plant 174(2):e13678. https://doi.org/10.1111/ppl.13678
Li N, Cao BL, Chen ZJ, Xu K (2022c) Root morphology ion absorption and antioxidative defense system of two Chinese cabbage cultivars (Brassica rapa L.) reveal the different adaptation mechanisms to salt and alkali stress. Protoplasma 259(2):385–398. https://doi.org/10.1007/s00709-021-01675-5
Liu HJ, Hu MH, Wang Q, Cheng L, Zhang ZB (2018) Role of papain-like cysteine proteases in plant development. Front Plant Sci 871:1717. https://doi.org/10.3389/fpls.2018.01717
Lou YH, Guan R, Sun MJ, Han F, He W, Wang H, Song FP, Cui XM, Zhuge YP (2018) Spermidine application alleviates salinity damage to antioxidant enzyme activity and gene expression in alfalfa. Ecotoxicology 27(10):1323–1330. https://doi.org/10.1007/s10646-018-1984-7
Lv Y, Ma J, Wei H, Xiao F, Wang YY, Jahan N, Hazman M, Qian Q, Shang LG, Guo LB (2022) Genome-wide domestication and a transcriptomic analysis reveals the loci and natural alleles of salt tolerance in rice (Oryza sativa L.). Front Plant Sci 13:912637. https://doi.org/10.3389/fpls.2022.912637
Ma SQ, Lv L, Meng C, Zhang CS, Li YQ (2020) Integrative analysis of the metabolome and transcriptome of Sorghum bicolor reveals dynamic changes in flavonoids accumulation under saline-alkali stress. J Agric Food Chem 68(50):14781–14789. https://doi.org/10.1021/acs.jafc.0c06249
Ma Q, Wu CY, Liang SH, Yuan YH, Liu CJ, Liu JJ, Feng BL (2021) The alkali tolerance of broomcorn millet (Panicum miliaceum L.) at the germination and seedling stage: the case of 296 broomcorn millet genotypes. Front Plant Sci 12:711429. https://doi.org/10.3389/fpls.2021.711429
Ma Q, Yuan YH, Wu EG, Wang HL, Dang K, Feng Y, Ivanistau A, Feng BL (2022) Endogenous bioactive gibberellin/abscisic acids and enzyme activity synergistically promote the phytoremediation of alkaline soil by broomcorn millet (Panicum miliaceum L.). J Environ Manag 305:114362. https://doi.org/10.1016/j.jenvman.2021.114362
Marques EC, de Freitas PAF, Alencar NLM, Prisco JT, Gomes-Filho E (2013) Increased Na+ and Cl− accumulation induced by NaCl salinity inhibits cotyledonary reserve mobilization and alters the source-sink relationship in establishing dwarf cashew seedlings. Acta Physiol Plant 35(7):2171–2182. https://doi.org/10.1007/s11738-013-1254-5
Nimir NEA, Zhou GS, Zhu GL, Ibrahim ME (2020) Response of some sorghum varieties to GA3 concentrations under different salt compositions. Chilean J Agric Res 80(4):478–486. https://doi.org/10.4067/S0718-58392020000400478
Pazhamala LT, Kudapa H, Weckwerth W, Millar AH, Varshney RK (2021) Systems biology for crop improvement. Plant Genome 14(2):e20098. https://doi.org/10.4067/10.1002/tpg2.20098
Ponte LFA, Silva ALCD, Carvalho FEL, Maia JM, Voigt EL, Silveira JAG (2014) Salt-induced delay in cotyledonary globulin mobilization is abolished by induction of proteases and leaf growth sink strength at late seedling establishment in cashew. J Plant Physiol 171(15):1362–1371. https://doi.org/10.1016/j.jplph.2014.06.001
Punia H, Tokas J, Mor VS, Bhuker A, Malik A, Singh N, Satpal AAA, Hefft DI (2021) Deciphering reserve mobilization, antioxidant potential, and expression analysis of starch synthesis in sorghum seedlings under salt stress. Plants 10(11):2463. https://doi.org/10.3390/plants10112463
Raza A, Su W, Hussain MA, Mehmood SS, Zhang XK, Cheng Y, Zou XL, Lv Y (2021) Integrated analysis of metabolome and transcriptome reveals insights for cold tolerance in rapeseed (Brassica napus L.). Front Plant Sci 12:721681. https://doi.org/10.3389/fpls.2021.721681
Ren XN, Shan Y, Li X, Wang LL, Li YY, Ma LJ, Li XM (2021) Endophytic infection programs the ascorbate-glutathione cycle in rice (Oryza sativa L.) under Na2CO3 stress. Appl Ecol Environ Res 19(3):1895–1907. https://doi.org/10.15666/aeer/1903_18951907
Sidari M, Mallamaci C, Muscolo A (2008) Drought, salinity and heat differently affect seed germination of Pinus pinea. J for Res 13(5):326–330. https://doi.org/10.1007/s10310-008-0086-4
Song TT, Xu HH, Sun N, Jiang L, Tian P, Yong YY, Yang WW, Cai H, Cui GW (2017) Metabolomic analysis of alfalfa (Medicago sativa L.) root-symbiotic rhizobia responses under alkali stress. Front Plant Sci 8:1208. https://doi.org/10.3389/fpls.2017.01208
Song TT, Sun N, Dong L, Cai H (2021) Enhanced alkali tolerance of rhizobia-inoculated alfalfa correlates with altered proteins and metabolic processes as well as decreased oxidative damage. Plant Physiol Biochem 159:301–311. https://doi.org/10.1016/j.plaphy.2020.12.021
Tyack N, Dempewolf H, Khoury CK (2020) The potential of payment for ecosystem services for crop wild relative conservation. Plants 9(10):1–14. https://doi.org/10.3390/plants9101305
Xu ZZ, Chen JD, Meng S, Xu P, Zhai CJ, Huang F, Guo Q, Zhao L, Quan YG, Shangguan YX, Meng Z, Wen T (2022) Genome sequence of Gossypium anomalum facilitates interspecific introgression breeding. Plant Commun 3(5):100350. https://doi.org/10.1016/j.xplc.2022.100350
Yang CW, Chong JN, Li CY, Kim C, Shi DC, Wang DL (2007) Osmotic adjustment and ion balance traits of an alkali resistant halophyte Kochia sieversiana during adaptation to salt and alkali conditions. Plant Soil 294(1–2):263–276. https://doi.org/10.1007/s11104-007-9251-3
Yi YK, Peng YQ, Song T, Lu SQ, Teng ZN, Zheng Q, Zhao FK, Meng S, Liu BH, Peng Y, Chen GH, Zhang JH (2022) NLP2-NR module associated NO is involved in regulating seed germination in rice under salt stress. Plants 11(6):795. https://doi.org/10.3390/plants11060795
Zaib-un-Nisa MX, Anwar S, Chen C, Jin XX, Yu LJ, Ali N, Chen C (2022) Strigolactone enhances alkaline tolerance in soybean seeds germination by altering expression profiles of ABA biosynthetic and signaling genes. J Plant Biol. https://doi.org/10.1007/s12374-022-09357-2
Zhang N, Zhang HJ, Sun QQ, Cao YY, Li XS, Zhao B, Wu P, Guo YD (2017) Proteomic analysis reveals a role of melatonin in promoting cucumber seed germination under high salinity by regulating energy production. Sci Rep 7(1):503. https://doi.org/10.1038/s41598-017-00566-1
Zhang HH, Li X, Guan YP, Li MB, Wang Y, An MJ, Zhang YH, Liu GJ, Xu N, Sun GY (2020) Physiological and proteomic responses of reactive oxygen species metabolism and antioxidant machinery in mulberry (Morus alba L.) seedling leaves to NaCl and NaHCO3 stress. Ecotoxicol Environ Saf 193:110259. https://doi.org/10.1016/j.ecoenv.2020.110259
Zhang X, Yang F, Ma HY, Li JP (2022) Evaluation of the saline-alkaline tolerance of rice (Oryza sativa L.) mutants induced by heavy-ion beam mutagenesis. Biology 11(1):126. https://doi.org/10.3390/biology11010126
Zhao HX, Du XD, Chen SQ, Yang LM, Peng F, Huang XQ, Ma WD, Zheng HY, Cai YS, Pan GJ (2022) Rice adapts to alkali stress by microstructural changes. Pak J Bot 54(4):1181–1190. https://doi.org/10.30848/PJB2022-4(13)
Zhou Y, Wen ZL, Zhang JW, Chen XJ, Cui JX, Xu W, Liu HY (2017) Exogenous glutathione alleviates salt-induced oxidative stress in tomato seedlings by regulating glutathione metabolism, redox status, and the antioxidant system. Sci Hortic 220:90–101. https://doi.org/10.1016/j.scienta.2017.02.021
Acknowledgements
We thank Jilin Crop Germplasm Introduction and Breeding Center for providing the experimental materials for this study and thank International Science Editing for polishing this manuscript. This work was supported by the National Natural Science Foundation of China (No. 32072012) and the Natural Science Foundation of Jilin Province, China (No. 20200201134JC).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Communicated by Dorothea Bartels.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
425_2023_4129_MOESM1_ESM.tif
Fig. S1 qRT-PCR validation of eight cotyledon genes of two ecotypes of wild soybean. a Common wild soybean. b Barren-tolerant wild soybean (TIF 129 KB)
425_2023_4129_MOESM2_ESM.tif
Fig. S2 The number of up-regulated and down-regulated DEGs in cotyledons of two ecotypes of wild soybean. a Common wild soybean. b Barren-tolerant wild soybean (TIF 172 KB)
425_2023_4129_MOESM3_ESM.tif
Fig. S3 GO annotation and KEGG enrichment of DEGs in cotyledons of two ecotypes of wild soybean. a GO annotation of DEGs in common wild soybean cotyledon. b GO annotation of DEGs in barren-tolerant wild soybean cotyledon. c KEGG enrichment of DEGs in common wild soybean cotyledon. d KEGG enrichment of DEGs in barren-tolerant wild soybean cotyledon (TIF 1089 KB)
425_2023_4129_MOESM4_ESM.tif
Fig. S4 qRT-PCR validation of ten young root genes of two ecotypes of wild soybean. a Common wild soybean. b Barren-tolerant wild soybean (TIF 147 KB)
425_2023_4129_MOESM5_ESM.tif
Fig. S5 The number of up-regulated and down-regulated DEGs in young roots of two ecotypes of wild soybean. a Common wild soybean. b Barren-tolerant wild soybean (TIF 219 KB)
425_2023_4129_MOESM6_ESM.tif
Fig. S6 GO annotation and KEGG enrichment of DEGs in young roots of two ecotypes of wild soybean. a GO annotation of DEGs in common wild soybean young root. b GO annotation of DEGs in barren-tolerant wild soybean young root. c KEGG enrichment of DEGs in common wild soybean young root. d KEGG enrichment of DEGs in barren-tolerant wild soybean young root (TIF 940 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, X., Hu, Y., Wang, Y. et al. Integrated metabolomic and transcriptomic strategies to reveal alkali-resistance mechanisms in wild soybean during post-germination growth stage. Planta 257, 95 (2023). https://doi.org/10.1007/s00425-023-04129-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-023-04129-9