Abstract
Background and aims
Crop yield and quality are generally limited by poor soils, which is a key limiting factor for sustainable development in modern agriculture. Wild soybean (Glycine soja) is an excellent wild resource, with tolerance to adverse environments, especially poor soil. This study aimed to reveal the key molecular mechanism of wild soybean to resist phosphorus deficiency in soil.
Methods
Differences in the types, amounts and metabolic pathways of small molecule metabolites and gene expression were compared and multi-omics integration analysis was performed between wild and cultivated soybean (Glycine max) seedling roots under sufficient and artificially simulated low-phosphorus in this study.
Results
Under low-phosphorus stress, wild soybean seedlings experienced less growth inhibition and root-specific growth compared with cultivated soybean. Genes encoding sulfoquinovosyl transferase (SQD2), catechol O-methyltransferase (COMT), glutathione S-transferase (GST) and peroxidase (POD) were up-regulated; levels of glutamic acid, glycine, putrescine, phenylalanine, tyrosine, catechol and neohesperidin were increased; and levels of glycerol-3-phosphate decreased. Integrated analysis showed that the above genes and metabolites were involved in glutathione metabolism, glycerolipid metabolism and phenylpropane biosynthesis.
Conclusions
These metabolic pathways are involved in phosphorus reuse, while membrane lipid remodelling and reactive oxygen species scavenging are carried out to maintain membrane stability and ensure plant survival under phosphorus deficiency. This study provides new ideas for the study of mechanism of tolerance to phosphorus deficiency in wild soybean and lays the theoretical foundation for developing varieties of cultivated soybean that tolerate poor soils.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Phosphorus is an important factor for maintaining the normal growth and development of crops and ensuring productivity in agricultural ecosystems (George et al. 2016; Kirkby and Johnston 2008). Phosphorus in the soil is often not absorbed by crops due to the adsorption of metal cations and the activities of microorganisms (Hinsinger, 2001; Schachtman et al. 1998). Excessive use of phosphate fertilizer not only causes economic loss to farmers, but also eutrophication of water bodies (Correll 1998; George et al. 2016). In addition, phosphate rock is a non-renewable resource (Abelson 1999; Vance et al. 2003). Exploring the molecular mechanisms of crop adaptation to phosphorus deficiency in soil is a hot issue in agricultural research from the perspective of the dynamic relationship between soil and crop.
Soybean is an important economic crop and oil crop, providing people with nutrients such as protein, isoflavones and phospholipids (Kofsky et al. 2018). Phosphorus deficiency can prevent the growth of root nodules, leading to a reduction in the nitrogen-fixation capacity of soybean, which in turn reduces yield and quality (Drevon and Hartwig 1997; Zhu et al. 2021). As the ancestor of cultivated soybeans, wild soybean is a natural plant resource with genetic diversity (Lam et al. 2010). It has been used many times for breeding improvement due to its high protein content and resistance to adversity (Li et al. 2017; Liu et al. 2020). Therefore, it is an important to study molecular mechanisms of tolerance to phosphorus deficiency in wild soybean to improve the quality of cultivated soybean.
Roots are organs that directly perceive a poor soil environment and respond quickly. Studies have shown that, under phosphorus deficiency, the phenotypic structure of roots changes (Ding et al. 2021; Lynch 2011), root secretions are altered (Wang et al. 2018), lipid metabolism and secondary metabolism are activated (Meena et al. 2021; Zhang et al. 2019), and genes related to phosphorus uptake and transport are induced (Qu et al. 2015; Wu et al. 2013). Thus, it is feasible to reveal the relationship between crops and soil from the perspective of roots.
Under phosphorus deficiency, electron transport and photosynthesis are restricted, leading to the accumulation of reactive oxygen species and membrane damage (Juszczuk et al. 2001). Studies have shown that in rice and rape, antioxidant enzyme activity and antioxidant levels are enhanced to scavenge oxygen free-radicals and protect membrane structure (Chen et al. 2015; Fu et al. 2014). Secondary metabolites are accumulated to improve plant tolerance in wheat, switchgrass and maize under phosphorus deficiency (Ding et al. 2021; Luo et al. 2019; Pontigo et al. 2018). Previous studies showed that phosphorus deficiency leads to the degradation of phospholipids to obtain phosphorus for plant use, while sulfolipids and galactolipids are synthesized to replace phospholipids and so maintain membrane stability (Mo et al. 2019; Byrne et al., 2011; Zhang et al. 2019). These findings provide a theoretical basis and new ideas for the study of the molecular mechanisms underlying the response of wild soybean roots to low phosphorus.
In this study, high-throughput sequencing technology and integrated transcriptomic and metabolomic analyses were used to compare differences in the physiology, types, amounts and metabolic pathways of small molecule metabolites and gene expression between wild and cultivated soybean seedling roots under phosphorus sufficiency and deficiency. The objectives were to (1) identify differentially expressed genes (DEGs) in wild soybean seedling roots under phosphorus deficiency, particularly up-regulated transcripts associated with membrane lipid phosphorus reusing and antioxidant protecting and (2) determine the changes in metabolites caused by changes in gene expression under phosphorus deficiency. The overall objective is to identify key metabolic pathways, key genes and key metabolites that play crucial roles in the resistance of wild soybeans to phosphorus deficiency, and to lay the theoretical foundation for the development of soybean tolerant to low phosphorus.
Materials and methods
Plant materials and growth
The experimental materials, cultivated soybean (Jinong24) (Glycine max) and wild soybean (Huinan06116) (Glycine soja), were kindly provided by the Jilin Academy of Agriculture Science, Changchun (China). The experiment was carried out on May 10, 2019. Seeds of both lines were planted in perforated pots of approximately 14 cm diameter, 23 cm high and 3.5 L volume filled with clean sand, with one seedling per pot. Soybeans were cultivated in the experimental site of Northeast Normal University, Changchun (China), where day/night temperature was 25 ± 3/17 ± 2 °C and relative humidity was about 60%.
Stress treatments and plant harvest
When the third triple compound leaf emerged, the seedling treatment began. Cultivated and wild soybean were treated with both control and low-phosphorus, with 12 pots in each group. Seedlings of the control group were irrigated with a complete Hoagland nutrient solution with a phosphorus concentration of 2.0 mM. Seedlings of the low-phosphorus stress group were irrigated with a Hoagland nutrient solution with a phosphorus concentration of 0.0025 mM, in which KCl of equal concentration was used instead of KH2PO4. After 6 days of treatment, three pots were randomly selected from each group as three replicates to use their roots for RNA extraction and analysis (Hosseini and Matthews 2014). After 14 days of treatment, five pots were randomly selected from each group to measure growth parameters and photograph plant status, and each parameter was determined for four replicates (Zhao et al. 2020). The roots of the remaining four pots were used as four replicates for metabolomics analysis (Zhang et al. 2016).
Growth parameter measurement and analysis
Root length, fresh weight and dry weight were determined according to Li et al. (Li et al. 2018). The longest tap root of each plant was selected for measurement. Data of growth parameter were presented as mean ± standard deviation of four biological replicates. The data were obtained and analyzed using SAS 9.2 (NCSU, N.C., USA).
Transcriptomics analysis
RNA extraction and sequencing
A NanoDrop 2000 (Thermo Scientific Inc., DE, USA) was used to measure the concentration of RNA, and the analysis kit in an Agilent Bioanalyzer 2100 system (Agilent Technologies, CA, USA) was used to assess the integrity of RNA. From each sample, 1 μg of RNA was used as input material to generate a sequencing library using the NEBNext® Ultra™ RNA Library Prep Kit from Illumina® (www.international.neb.com, USA). Library quality was assessed on the Agilent Bioanalyzer 2100 system after purification of the library fragments. The resulting libraries were sequenced on the Illumina Hiseq platform and reads were generated.
Data processing and analysis
Clean reads were processed by internal Perl scripts and their Q20, Q30 and GC content and sequence repeat levels were calculated. These pure reads were then positioned to the reference genomic sequence Williams 82 for analysis and annotation of exact matches. After quantifying the gene expression levels, the DESeq R software package (1.10.1) was used to perform differential expression analysis for the two conditions. Benjamini and Hochberg’s method was used to adjust P-values, and genes with P-values <0.05 were designated as DEGs (Liu et al. 2020). The GOseq R program package was used to perform Gene Ontology (GO) enrichment analysis on DEGs. The Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment of DEGs was carried out via the online KEGG Automated Annotation Server (http://www.genome.jp/kegg).
Quantitative real-time PCR analysis of DEGs
Eight genes were randomly selected in wild and cultivated soybean seedling roots, respectively. All 8 genes in wild soybean were differentially expressed genes. There were 8 genes in cultivated soybean, and 6 of them were differentially expressed genes. Among these randomly selected genes, 3 genes existed in both species and were differentially expressed under low-phosphorus stress. Primers were designed using Primer Premier 5.0 (Premier Biosoft International, PaloAlto, CA, USA). Gene IDs and primer sequences have been shown in Supplementary Table 1. Reverse transcription kit (Aidlab Biotechnologies Co., Ltd., BJ, CHN) was used to reverse transcribe 0.5 μg of RNA into cDNA. The qRT-PCR experiment was performed by the analytikjena-qTOWER2.2 fluorescence quantitative PCR instrument (Analytik Jena AG, Jena, GER). Each gene was set up 3 replicates in each sample. After initial denaturing at 95 °C for 3 min, the reaction was followed by 39 cycles at 94 °C for 10 s and 58 °C for 30 s. At the same time, 18 s rRNA (GYMA06G315500WM82A2V1) was selected as the reference gene, which was verified to be stably expressed in plants with good amplification efficiency (Ma et al. 2016; Xiao et al. 2016). The comparative CT method and SigmaPlot version 10.0 (Systat Software Inc., CA, USA) were used for data analysis (Schmittgen and Livak 2008).
Metabolomics analysis
Extraction and analysis of metabolites from seedling roots
Root samples of 50 mg from cultivated and wild soybean seedlings were mixed with 0.5 ml of methanol-chloroform (3:1) and 60 μl of ribitol. After centrifugation, the supernatant was placed into a 2-ml centrifuge tube. After drying, the samples were mixed with 80 μl of methoxamine reagent and placed in an oven for 2 h at 37 °C. Then, 100 μl of Bis(trimethylsilyl) trifluoroacetamide (containing 1% Trimethylchlorosilane, v/v) was added to each sample, mixed and stored for 1.5 h at 70 °C. After cooling to room temperature, the metabolites were detected using an Agilent 7890 gas chromatograph-mass spectrometer.
Data analysis
ChromaTOF software (V 4.3x, LECO, St. Joseph, MI, USA) and the EI-MS and FiehnLib databases were used for data collection and pre-processing as well as metabolite identification. The software SIMCA-P 13.0 (Umetrics, Umea, Sweden) was used to perform data principal component analysis (PCA), partial least squares discriminant analysis (PLS-DA), orthogonal partial least squares discriminant analysis (OPLS-DA) analysis and calculate variable importance values (VIP). The KEGG (www.genome.jP/kegg) and Metaboanalyst 3.0 (www.metaboanalyst.ca) databases were consulted to analyze metabolic pathways. Data were expressed as mean ± standard error of four biological replicates.
Integration analysis
Pearson correlation coefficients for transcriptomic and metabolomic data were calculated on Metaboanalyst 3.0 (www.metaboanalyst.ca). The integration network was drawn on Cytoscape (version 3.8.2) based on correlation coefficients between transcriptomic and metabolomic data. The key mechanism for counteracting phosphorus deficits was visualized using Visio software (Office Visio 2013, 15.0, Microsoft USA).
Results
Changes in root morphology
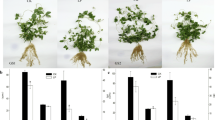
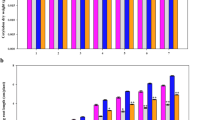
Root morphology of both wild and cultivated soybean seedlings was affected by low phosphorus, but with differences between them. Compared with control group, low phosphorus inhibited the number of lateral roots of wild and cultivated soybean seedlings, but promoted root length (Fig. 1). The length of the longest tap root of wild and cultivated soybean seedlings increased, by 0.15- and 0.04-fold, respectively (Table 1). After 14 days of low-phosphorus treatment, the fresh and dry weight of roots decreased by 0.57- and 1.10-fold in wild soybean and by 0.85- and 1.23-fold in cultivated soybean, respectively (Table 1). Thus, low phosphorus resulted in greater growth inhibition to the roots of cultivated soybean than for wild soybean seedlings.
Transcriptome analysis of roots under low-phosphorus stress
After transcriptome analysis of wild and cultivated soybean roots under low-phosphorus stress, a total of 89.42 Gb of clean data (Q30 > 93.96%) was obtained, and the clean data of each sample reached 6.52Gb. The clean reads of each sample showed alignment rates of 89.01%–93.87% with the designated reference genome, and the percentage of CG bases was 45.66%–46.23% (Table 2). After 6 days of low-phosphorus treatment, 340 and 306 transcripts were differentially expressed in wild and cultivated soybean seedling roots, respectively, including 6 common DEGs (Fig. S2a, Supplementary Table 5). There were 139 up-regulated and 201 down-regulated genes found in wild soybean roots, and corespondingly 200 and 106 in cultivated soybean (Fig. S2b). For wild and cultivated soybean seedling roots, 232 and 230 DEGs were annotated, respectively. The GO annotation analysis of DEGs showed that they were mainly distributed in biological processes such as metabolic process (GO:0008152) and response to stimulation (GO:0050896), cell components such as organelles (GO:0043226) and membranes (GO:0016020) and molecular functions such as catalytic activity (GO:0003824) and antioxidant activity (GO:0016209) (Fig. 2). In addition, the KEGG pathway enrichment analysis of DEGs showed that the metabolic pathways enriched in wild soybean roots included glutathione metabolism (ko00480), glycerolipid metabolism (ko00561), phenylpropanoid biosynthesis (ko00940) and starch and sucrose metabolism (ko00500). The metabolic pathways enriched in cultivated soybean roots included phenylpropanoid biosynthesis (ko00940) and glycerolipid metabolism (ko00561) (Fig. 2, Supplementary Table 2).
GO annotation and KEGG enrichment of DEGs in wild and cultivated soybean seedling roots. A, GO annotation of up-regulated genes in cultivated soybean seedling roots; B, GO annotation of down-regulated genes in cultivated soybean seedling roots; C, GO annotation of up-regulated genes in wild soybean seedling roots; D, GO annotation of down-regulated genes in wild soybean seedling roots; E, KEGG enrichment of up-regulated genes in cultivated soybean seedling roots; F, KEGG enrichment of down-regulated genes in cultivated soybean seedling roots; G, KEGG enrichment of up-regulated genes in wild soybean seedling roots; H, KEGG enrichment of down-regulated genes in wild soybean seedling roots
Glutathione metabolism and phenylpropane biosynthesis play important roles in antioxidant protection, and this experiment mapped annotated transcripts to these two pathways. Under low-phosphorus stress, 6 up-regulated genes in wild soybean seedling roots were annotated as genes encoding enzymes involved in phenylpropane biosynthesis. Gene encoding catechol O-methyltransferase (COMT) was up-regulated by 1.41-fold in wild soybean seedling roots. Genes encoding cationic perixidase (POD) were up-regulated by 1.25-, 1.39-, 1.62-, 1.04- and 1.02-fold, respectively, in wild soybean seedling roots (Table 3). Three genes encoding glutathione-s-transferases (GST) associated with scavenging reactive oxygen species in glutathione metabolism were up-regulated, by 1.86-, 1.12- and 1.24-fold, respectively, in wild soybean seedling roots. The GO analysis showed that they were involved in molecular functions (glutathione transferase activity, GO:0004364) and biological processes (cellular response to phosphate starvation, GO:0016036) (Table 3). In addition, under phosphorus deficiency, plants can reuse phosphorus through membrane lipid remodeling. The results showed that gene encoding a key enzyme sulfoquinovosyl transferase (SQD2) associated with membrane lipid remodeling, was highly up-regulated by 1.10-fold in wild soybean roots under low-phosphorus stress, and was annotated as cellular phosphate deficiency response (GO:0016036), plasma membrane (GO:0005886) and galactolipid biosynthesis process (GO:0019375) (Table 3).
Under low-phosphorus stress, gene encoding phospholipase D associated with phospholipid degradation was up-regulated by 1.54-fold in cultivated soybean seedling roots. Three genes encoding cationic peroxidase (POD) in phenylpropanoid biosynthesis were up-regulated, by 1.19-, 1.58- and 1.02-fold, respectively, in cultivated soybean seedling roots. However, three more genes encoding cationic peroxidase were significantly down-regulated, by 1.86-, 1.15- and 1.30-fold, respectively, in cultivated soybean seedling roots. In addition, TGA4 (TGACG motif-binding factor) gene associated with salicylic acid signaling in cultivated soybean was down-regulated by 1.69-fold under low-phosphorus stress, and was annotated as cellular response to stress (GO:0033554) in biological processes. The gene encoding S-adenosylmethionine decarboxylase was down-regulated by 1.57-fold in cultivated soybean seedling roots. The gene encoding asparagine synthase associated with nitrogen metabolism was down-regulated by 1.07-fold in cultivated soybean seedling roots, and was annotated as ammonium ion metabolic process (GO:0097164) in biological processes (Supplementary Table 6).
Transcription factors
Under low-phosphorus stress, differential expression of 30 and 23 transcription factors was detected in wild and cultivated soybean seedling roots, respectively. Among them, MYB308 expression was greatly down-regulated by 1.10-fold in wild soybean, but with no significant change in cultivated soybean (Fig. 3, Supplementary Table 3).
The validation of RNA-seq by qRT-PCR
To verify the reliability of RNA-seq results obtained by sequencing on the Illumina platform, we randomly selected 8 genes in wild and cultivated soybean seedling roots, respectively, and verified the accuracy of the RNA-seq data by qRT-PCR. The qRT-PCR results for these randomly selected genes were consistent with the trend of gene expression in the transcriptome data, verifying the accuracy of the transcriptome data (Fig. S1).
Metabolome analysis of roots under low-phosphorus stress
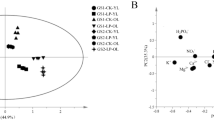
In this experimental study, metabolomic tests and analyses were further carried out on wild and cultivated soybean seedling roots under control and low-phosphorus stress. The first principal component, separating the two research objects of wild and cultivated soybean, scored 54.30% (Fig. 4a), with myo-inositol and ethanolamine playing crucial roles in the separation (Fig. 4b, Supplementary Table 4). The second principal component, separating the two treatments of low phosphorus stress and control, scored 19.40% (Fig. 4a), with sorbose and L-Malic acid playing key roles in the separation (Fig. 4b, Supplementary Table 4).
There were 36 differential metabolites screened according to VIP (variable importance projection values) > 1, similarity >700 and P ≤ 0.05 (Gao et al. 2019; Idborg et al. 2004), including the following types: amino acids, fatty acids, organic acids, sugars and sugar alcohols, and secondary metabolites. Compared with control, the relative contents of glycine, glutamic acid and putrescine in wild soybean seedling roots increased, by 1.56-, 0.43- and 0.85-fold, respectively, under low-phosphorus stress. Glutamic acid and putrescine decreased by 0.85-fold, respectively, and glycine increased by 0.35-fold in cultivated soybean roots, but the change was not significant. The relative level of glycerol-3-phosphate decreased by 1.19- and 0.09-fold in wild and cultivated soybean roots, respectively. The relative levels of phenylalanine and tyrosine increased by 1.91- and 1.67-fold in wild soybean roots and decreased by 0.52- and 0.30-fold in cultivated soybean roots, respectively. The 4-vinylphenol increased by 6.13- and 0.01-fold in wild and cultivated soybean roots, respectively. After low-phosphorus treatment, the relative content of neohesperidin increased by 0.42-fold in wild soybean roots, but decreased 0.07-fold in cultivated soybean. The relative content of catechol increased by 0.21- and 0.09-fold in wild and cultivated soybean roots, respectively (Table 4).
Discussion
In response to low phosphorus, dry and fresh weight revealed more inhibition of overall root growth in cultivated soybean, compared to wild soybean, while root length was not inhibited in either species. Under low-phosphorus stress, the root lengths of wild and cultivated soybean both increased, indicating that both absorbed phosphorus by increasing root length.
In addition to changes in root morphology, wild soybean also adjusted some metabolic pathways by changing gene expression to better adapt to low phosphorus. Metabolomic and transcriptomic integration analyses were performed to obtain a network of gene and metabolite interactions in wild soybean roots under low phosphorus stress (Fig. 5). Under low-phosphorus stress, lipid remodeling could also change the distribution of phosphorus in cells and so maintain the level of inorganic phosphorus. When plants are in a phosphorus-deficient state, phospholipids are degraded to generate free inorganic phosphorus and so alleviate the harm of low-phosphorus stress, and the consumed phospholipids could be replaced by newly synthesized sulfolipids and glycolipids (Müller et al. 2015; Zhang et al. 2019). Phospholipases are key enzymes that catalyse the hydrolysis of phospholipids in membranes and play a very important role in regulating plant development under phosphorus deficiency (Deepika and Singh 2021). Previous studies have shown that SQD2 was involved in the reaction of the UDP sulfoquinose with diacylglycerol to form sulfolipids and maintained the membrane stability of plants under phosphorus deficiency in Arabidopsis (Yu et al. 2002); SQD2 has also been identified as the responsive gene to phosphorus deficiency (Okazaki et al. 2013; Wang et al. 2018). In this study, the expression of the gene encoding SQD2 was significantly increased, and the relative content of glycerol-3-phosphate significantly decreased in wild soybean roots. Gene encoding phospholipase D was up-regulated, and level of glycerol-3-phosphate was decreased in cultivated soybean seedling roots, but to a lesser extent than in wild soybean. This showed that cultivated soybean had the ability to reuse phosphorus internally, but less than that of wild soybean. Although the phospholipids that make up biofilms could be damaged by redistribution of phosphorus in the cell, the synthesis of sulfolipids would be increased to maintain biofilm stability in wild soybean. However, no significantly genetic changes associated with sulfolipid synthesis were detected in cultivated soybean seedling roots. The integrity and stability of the membrane is essential for plants to maintain normal metabolism and withstand adverse external environments.
Integration network of metabolites and genes in wild soybean seedling roots under low-phosphorus stress. Genes and metabolites are represented by numbers and letters, respectively. 1, Glyma.07G048800.Wm82.a2.v1; 2, Glyma.03G208200.Wm82.a2.v1; 3, Glyma.09G277900.Wm82.a2.v1; 4, Glyma.10G022500.Wm82.a2.v1; 5, Glyma.18G055400.Wm82.a2.v1; 6, Glyma.18G055500.Wm82.a2.v1; 7, Glyma.07G032700.Wm82.a2.v1; 8, Glyma.03G078300.Wm82.a2.v1; 9, Glyma.03G018900.Wm82.a2.v1; 10, Glyma.20G238000.Wm82.a2.v1; 11, Glyma.01G040000.Wm82.a2.v1; 12, Glyma.11G198500.Wm82.a2.v1; 13, Glyma.11G024000.Wm82.a2.v1; 14, Glyma.08G175200.Wm82.a2.v1; A, 4-vinylphenol; B, phenylalanine; C, tyrosine; D, neohesperidin; E, 4-hydroxycinnamic acid; F, catechol; G, caffeic acid; H, D-glyceric acid; I, glycerol-3-phosphate; J, ethanolamine; K, oxoproline; L, putrescine; M, glutamic acid; N, dehydroascorbic acid; O, glycine. The thicker the edge is, the stronger the correlation is. The size of a node is proportional to the correlation between nodes
Under low-phosphorus stress, the electron transfer system is slowed, leading to excessive accumulation of oxygen free-radicals and increased membrane lipid peroxidation injury in plants (Juszczuk et al. 2001). Previous studies have shown that GST acts as an antioxidant enzyme that catalyzes the conversion of glutathione to glutathione disulfide (Nianiou-Obeidat et al., 2017; Rahantaniaina et al. 2013). This process can eliminate the toxicity of reactive oxygen and catalyze the reversible S-glutathionization of protein thiol residues, thereby ensuring that membrane structure and protein are protected from oxidative damage (Srivalli and Khanna-Chopra 2008). The integrated analysis of transcriptomics and metabolomics showed that in the glutathione metabolism pathway, the gene encoding GST was significantly up-regulated, and the relative contents of glutamate, glycine and putrescine were significantly increased in wild soybean roots. Glutamic acid and glycine can be used as precursors for the glutathione synthesis (Hasanuzzaman et al. 2017). Due to its polycationic nature, putrescine can interact directly with oxygen radicals to scavenge reactive oxygen species and can also transmit stress signals (Gupta et al. 2013). This suggests that at low-phosphorus levels, wild soybean could enhance the activity of the antioxidant enzyme GST and promote the synthesis of antioxidants to scavenge the excess reactive oxygen species generated by phosphorus deficiency.
Phenylalanine and tyrosine are used as precursors to synthesize a variety of secondary metabolites in the phenylpropane biosynthetic pathway (Dong and Lin 2021). COMT is used to catalyze the conversion of caffeic acid to ferulic acid and is involved in synthesis of syringyl units. Studies have shown that COMT expression increases the syringyl/guaiacyl ratio, which in turn is beneficial for improving plant stress tolerance (Vanholme et al. 2008; Yang et al., 2020). Under low-phosphorus stress, COMT overexpression in tomato plants can enhance low-phosphorus tolerance by increasing the absorption, transportation and utilization of phosphorus (Yang et al., 2020). Transcriptomics analysis showed that under low-phosphorus stress, the genes encoding COMT and peroxidase were up-regulated, resulting in the activation of phenylpropane biosynthesis in wild soybean roots. This result was further validated by metabolomic analysis, which showed that the relative levels of phenylalanine, tyrosine, neohesperidin and catechol all obviously increased in wild soybean roots. Thus, precursors would accumulate and expression of key genes would be enhanced in phenylalanine biosynthesis of wild soybean, promoting phenylalanine metabolism to produce secondary metabolites such as neohesperidin and catechol. These metabolites can help plants scavenge reactive oxygen species and improve their tolerance to phosphorus deficiency (Liu et al. 2020; Pontigo et al. 2018) (Fig. 6). The expression of gene encoding peroxidase was disordered, and the levels of phenylalanine, tyrosine and 4-hydroxycinnamic acid were significantly decreased in cultivated soybean seedling roots. Under low-phosphorus stress, genes and metabolites of phenylpropanoid biosynthesis changed irregularly in cultivated soybean seedling roots.
Key mechanisms of resistance to low phosphorus stress in wild soybean. PAL, phenylalanine ammonia lyase; C4H, cinnamate-4-hydroxylase; TAL, tyrosine ammonia lyase; C3H, p-coumaric acid 3-hydroxylase; F5H, Ferulate-5-hydroxylase; GR, Glutathione reductase; GSPSA, glutathionylspermidine amidase; GSPS, glutathionylspermidine synthetase; SPDS, spermidine synthase; GSHS, glutathione synthetase; GSH, glutathione; GSSG, glutathione disulfide
There are many transcriptional repressors in the MYB transcription factor family whose down-regulation has a facilitative effect on metabolic processes (Jiao et al. 2019). Studies have shown that MYB308 overexpression in tobacco and snapdragon inhibits the activity of 4-coumaric acid: CoA ligase, cinnamic acid-4-hydroxylase and cinnamyl alcohol dehydrogenase in phenylpropane metabolism and leads to a reduction in lignin monomers and phenolic acid content, leaving plants vulnerable to adverse environmental stresses (Omer et al. 2013; Tamagnone et al. 1998). In this study, the MYB308 gene in wild soybean seedling roots was significantly down-regulated. Under low-phosphorus stress, the down-regulation of MYB308 provides further evidence for enhanced phenylpropane metabolism in wild soybean. It is clear from these changes that phenylpropane metabolism is of great help to wild soybean in resisting low-phosphorus stress.
Under low-phosphorus stress, genes associated with salicylic acid signaling, S-adenosylmethionine decarboxylase and asparagine synthase were significantly down-regulated in cultivated soybean seedling roots. The levels of salicylic acid, putrescine and some important amino acids such as aspartic acid, asparagine and glutamic acid also decreased obviously. Network obtained by metabolomic and transcriptomic integration analyses revealed that salicylic acid signaling and important amino acid metabolism were severely inhibited in cultivated soybean seedling roots under low-phosphorus stress (Fig. S3). This led to weakened signal transduction and active oxygen scavenging in cultivated soybeans, and cultivated soybeans were susceptible to membrane lipid damage caused by low phosphorus (Gupta et al. 2017; Liu et al. 2015; Sun et al. 2018; Zhang et al. 2019). More seriously, the negative impact of low-phosphorus stress on basic metabolism such as nitrogen metabolism in cultivated soybeans made plants unable to maintain normal life activities (Laure et al. 2010, 2016). The weaker plant-soil adaptation of cultivated soybeans than wild soybeans may be due to the low phosphorus effectiveness to crops in intensive agricultural and the loss of plant resistance in the pursuit of high-quality traits during artificial domestication of cultivated soybeans.
Conclusion
The damage from an adverse environment to plants depends on the type, intensity and time of stress and the adaptation of plants to an adverse environment depends on morphological structure, physiology, biochemistry and intracellular molecular metabolism. In this experiment, integrated transcriptomic and metabolomic analyses confirmed that wild soybean degrades phospholipids to release phosphorus for plant reuse. In addition, wild soybean synthesizes sulpholipids to replace phospholipids and enhances phenylpropane biosynthesis and glutathione metabolism to scavenge reactive oxygen species to maintain the stability of the membrane system, helping plants to survive phosphorus deficiency. Under low-phosphorus stress, salicylic acid signaling and important amino acid metabolism are more sensitive in cultivated soybean seedling roots. This may explain the vulnerability of cultivated soybean to phosphorus deficiency in soil. This study provides a theoretical basis for developing cultivated soybean varieties that are tolerant of nutrient-poor soils.
References
Abelson PH (1999) A Potential Phosphate Crisis. Science, 283(5410):2015–2015. https://doi.org/10.1126/science.283.5410.201
Byrne SL, Foito A, Hedley PE, Morris J, Stewart D, Barth S (2011) Early response mechanisms of perennial ryegrass (Lolium perenne) to phosphorus deficiency. Ann Bot-London 107(2):243–254. https://doi.org/10.1093/aob/mcq234
Correll DL (1998) The role of phosphorus in the eutrophication of receiving waters: a review. J Environ Qual 27:261–266. https://doi.org/10.2134/jeq1998.00472425002700020004x
Chen S, Zhao H, Ding G et al (2015) Genotypic differences in antioxidant response to phosphorus deficiency in Brassica napus. Plant Soil 391:19–32. https://doi.org/10.1007/s11104-015-2395-7
Drevon JJ, Hartwig U (1997) Phosphorus deficiency increases the argon-induced decline of nodule nitrogenase activity in soybean and alfalfa. Planta 201:463–469. https://doi.org/10.1007/s004250050090
Ding N, Huertas R, Torres-Jerez I, Liu W, Watson B, Scheible WR, Udvardi M (2021) Transcriptional, metabolic, physiological and developmental responses of switchgrass to phosphorus limitation. Plant Cell Environ 44(1):186–202. https://doi.org/10.1111/pce.13872
Dong NQ, Lin HX (2021) Contribution of phenylpropanoid metabolism to plant development and plant-environment interactions. J Integr Plant Biol 63(1):180–209. https://doi.org/10.1111/jipb.13054
Deepika D, Singh A (2021) Plant phospholipase D: novel structure, regulatory mechanism, and multifaceted functions with biotechnological application. Crit rev Biotechnol, 1–19. Advance online publication. https://doi.org/10.1080/07388551.2021.1924113
Fu YQ, Yang XJ, Shen H (2014) The physiological mechanism of enhanced oxidizing capacity of rice (Oryza sativa L.) roots induced by phosphorus deficiency. Acta Physiol Plant 36:179–190. https://doi.org/10.1007/s11738-013-1398-3
Gupta K, Dey A, Gupta B (2013) Plant polyamines in abiotic stress responses. Acta Physiol Plant 35:2015–2036. https://doi.org/10.1007/s11738-013-1239-4
George TS, Hinsinger P, Turner BL (2016) Phosphorus in soils and plants – facing phosphorus scarcity. Plant Soil 401:1–6. https://doi.org/10.1007/s11104-016-2846-9
Gao Y, Jiang Z, Du M et al (2019) Photosynthetic and metabolic responses of eelgrass Zostera marina L. to short-term high-temperature exposure. J Ocean Limnol 37:199–209. https://doi.org/10.1007/s00343-019-7319-6
Gupta, S., Yadav, B. S., Raj, U., Freilich, S., & Varadwaj, P. K (2017) Transcriptomic Analysis of Soil Grown T aestivum cv Root to Reveal the Changes in Expression of Genes in Response to Multiple Nutrients Deficiency Front Plant Sci, 8, 1025. https://doi.org/10.3389/fpls.2017.01025
Hinsinger P (2001) Bioavailability of soil inorganic P in the rhizosphere as affected by root-induced chemical changes: a review. Plant Soil 237:173–195. https://doi.org/10.1023/A:1013351617532
Hosseini P, Matthews BF (2014) Regulatory interplay between soybean root and soybean cyst nematode during a resistant and susceptible reaction. BMC Plant Biol 14:300. https://doi.org/10.1186/s12870-014-0300-9
Hasanuzzaman M, Nahar K, Anee TI, Fujita M (2017) Glutathione in plants: biosynthesis and physiological role in environmental stress tolerance. Physiol Mol Biol Pla 23(2):249–268. https://doi.org/10.1007/s12298-017-0422-2
Idborg H, Edlund PO, Jacobsson SP (2004) Multivariate approaches for efficient detection of potential metabolites from liquid chromatography/mass spectrometry data. Rapid Commun Mass Sp, 18(9):944–954. https://doi.org/10.1002/rcm.1432
Juszczuk IM, Malusa M, Rychter AM (2001) Phosphate deficiency induced oxidative stress in bean (Phaseolus vulgaris L.). in: Horst W.J. et al. (eds) plant nutrition. Developments in plant and soil sciences, vol 92. Springer, Dordrecht. https://doi.org/10.1007/0-306-47624-X_71
Jinglong Z et al (2019) Transcriptome analysis reveals candidate genes related to phosphorus starvation tolerance in sorghum. BMC Plant Biol 19:1. https://doi.org/10.1186/s12870-019-1914-8
Jiao B, Zhao X, Lu W, Guo L, Luo K (2019) The R2R3 MYB transcription factor MYB189 negatively regulates secondary cell wall biosynthesis in Populus. Tree Physiol 39(7):1187–1200. https://doi.org/10.1093/treephys/tpz040
Kirkby EA, Johnston AE (2008) Soil and fertilizer phosphorus in relation to crop nutrition. In: white PJ, Hammond JP (eds) the ecophysiology of plant-phosphorus interactions. Plant ecophysiology, vol 7. Springer, Dordrecht. https://doi.org/10.1007/978-1-4020-8435-5_9
Kofsky J, Zhang H, Song BH (2018) The untapped genetic reservoir: the past, current, and future applications of the wild soybean (Glycine soja). Front Plant Sci 9:949. https://doi.org/10.3389/fpls.2018.00949
Lynch JP (2011) Root phenes for enhanced soil exploration and phosphorus acquisition: tools for future crops. Plant Physiol 156(3):1041–1049. https://doi.org/10.1104/pp.111.175414
Li MX, Guo R,Jiao Y, Jin XF, Zhang HY, Shi LX (2017) Comparison of salt tolerance in Soja based on metabolomics of seedling roots. Front plant Sci, 2017,8: https://doi.org/10.3389/fpls.2017.01101
Liu DP, Li MX, Liu Y, Shi LX (2020) Integration of the metabolome and transcriptome reveals the resistance mechanism to low nitrogen in wild soybean seedling roots. Environ Exp bot, 175. 104043.https://doi.org/10.1016/j.envexpbot.2020.104043
Luo B, Ma P, Nie Z, Zhang X, He X, Ding X, Feng X, Lu Q, Ren Z, Lin H, Wu Y, Shen Y, Zhang S, Wu L, Liu D, Pan G, Rong T, Gao S (2019) Metabolite profiling and genome-wide association studies reveal response mechanisms of phosphorus deficiency in maize seedling. Plant J 97(5):947–969. https://doi.org/10.1111/tpj.14160
Laure G, Reisdorf-Cren M, Rothstein SJ, Chardon F, Suzuki A (2010) Biological functions of asparagine synthetase in plants. Plant Sci 179(3):141–153, ISSN 0168-9452. https://doi.org/10.1016/j.plantsci.2010.04.010
Laure G, Rothstein SJ, Suzuki A (2016) Asparagine metabolic pathways in arabidopsis. Plant Cell Physiol 57(4):675–689. https://doi.org/10.1093/pcp/pcv184
Liu JH, Wang W, Wu H, Gong X, Moriguchi T (2015) Polyamines function in stress tolerance: from synthesis to regulation. Front Plant Sci 6:827. https://doi.org/10.3389/fpls.2015.00827
Lam HM, Xu X, Liu X, Chen W, Yang G, Wong FL, Li MW, He W, Qin N, Wang B, Li J, Jian M, Wang J, Shao G, Wang J, Sun SS, Zhang G (2010) Resequencing of 31 wild and cultivated soybean genomes identifies patterns of genetic diversity and selection. Nat Genet 42(12):1053–1059. https://doi.org/10.1038/ng.715
Li M, Xu J, Wang X, Fu H, Zhao M, Wang H, Shi L (2018) Photosynthetic characteristics and metabolic analyses of two soybean genotypes revealed adaptive strategies to low-nitrogen stress. J Plant Physiol 229:132–141. https://doi.org/10.1016/j.jplph.2018.07.009
Ma QP, Hao S, Chen X et al (2016) Validation of reliability for reference genes under various abiotic stresses in tea plant. Russ J Plant Physiol 63:423–432. https://doi.org/10.1134/S1021443716030080
Meena SK, Pandey R, Sharma S et al (2021) Cross tolerance to phosphorus deficiency and drought stress in mungbean is regulated by improved antioxidant capacity, biological N2-fixation, and differential transcript accumulation. Plant Soil 466:337–356. https://doi.org/10.1007/s11104-021-05062-0
Mo X, Zhang M, Liang C, Cai L, Tian J (2019) Integration of metabolome and transcriptome analyses highlights soybean roots responding to phosphorus deficiency by modulating phosphorylated metabolite processes. Plant Physiol Bioch 139:697–706. https://doi.org/10.1016/j.plaphy.2019.04.033
Müller J, Gödde V, Niehaus K, Zörb C (2015) Metabolic adaptations of white Lupin roots and shoots under phosphorus deficiency. Front Plant Sci 6:1014. https://doi.org/10.3389/fpls.2015.01014
Nianiou-Obeidat I, Madesis P, Kissoudis C, Voulgari G, Chronopoulou E, Tsaftaris A, Labrou NE (2017) Plant glutathione transferase-mediated stress tolerance: functions and biotechnological applications. Plant Cell Rep 36(6):791–805. https://doi.org/10.1007/s00299-017-2139-7
Omer S, Kumar S, Khan BM (2013) Over-expression of a subgroup 4 R2R3 type MYB transcription factor gene from Leucaena leucocephala reduces lignin content in transgenic tobacco. Plant Cell Rep 32(1):161–171. https://doi.org/10.1007/s00299-012-1350-9
Okazaki Y, Otsuki H, Narisawa T, Kobayashi M, Sawai S, Kamide Y, Kusano M, Aoki T, Hirai MY, Saito K (2013) A new class of plant lipid is essential for protection against phosphorus depletion. Nat Commun 4:1510. https://doi.org/10.1038/ncomms2512
Pontigo S, Ulloa M, Godoy K, Nikolic N, Nikolic M, Mora ML, Cartes P (2018) Phosphorus efficiency modulates phenol metabolism in wheat genotypes. J Plant Nutr Soil Sc 18:904–920. https://doi.org/10.4067/S0718-95162018005002603
Qu B, He X, Wang J, Zhao Y, Teng W, Shao A, Zhao X, Ma W, Wang J, Li B, Li Z, Tong Y (2015) A wheat CCAAT box-binding transcription factor increases the grain yield of wheat with less fertilizer input. Plant Physiol 167(2):411–423. https://doi.org/10.1104/pp.114.246959
Rahantaniaina MS, Tuzet A, Mhamdi A, Noctor G (2013) Missing links in understanding redox signaling via thiol/disulfide modulation: how is glutathione oxidized in plants? Front Plant Sci 4:477. https://doi.org/10.3389/fpls.2013.00477
Sun T, Busta L, Zhang Q, Ding P, Jetter R, Zhang Y (2018) TGACG-BINDING FACTOR 1 (TGA1) and TGA4 regulate salicylic acid and pipecolic acid biosynthesis by modulating the expression of SYSTEMIC ACQUIRED RESISTANCE DEFICIENT 1 (SARD1) and CALMODULIN-BINDING PROTEIN 60g (CBP60g). New Phytol 217(1):344–354. https://doi.org/10.1111/nph.14780
Srivalli S, Khanna-Chopra R (2008) Role of glutathione in abiotic stress tolerance. In: Khan NA, Singh S, Umar S (eds) Sulfur assimilation and abiotic stress in plants. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-540-76326-0_10
Schmittgen T, Livak K (2008) Analyzing real-time PCR data by the comparative CT method. Nat Protoc 3:1101–1108. https://doi.org/10.1038/nprot.2008.73
Schachtman DP, Reid RJ, Ayling SM (1998) Phosphorus uptake by plants: from soil to cell. Plant Physiol 116(2):447–453. https://doi.org/10.1104/pp.116.2.447
Tamagnone L, Merida A, Parr A, Mackay S, Culianez-Macia FA, Roberts K, Martin C (1998) The AmMYB308 and AmMYB330 transcription factors from antirrhinum regulate phenylpropanoid and lignin biosynthesis in transgenic tobacco. Plant Cell 10(2):135–154. https://doi.org/10.1105/tpc.10.2.135
Vanholme R, Morreel K, Ralph J, Boerjan W (2008) Lignin engineering. Curr Opin Plant Biol 11(3):278–285. https://doi.org/10.1016/j.pbi.2008.03.005
Vance CP, Uhde-Stone C, Allan DL (2003) Phosphorus acquisition and use: critical adaptations by plants for securing a nonrenewable resource. New Phytol 157(3):423–447. https://doi.org/10.1046/j.1469-8137.2003.00695.x
Wang Y, Lysøe E, Armarego-Marriott T, Erban A, Paruch L, van Eerde A, Bock R, Liu-Clarke J (2018) Transcriptome and metabolome analyses provide insights into root and root-released organic anion responses to phosphorus deficiency in oat. J Exp Bot 69(15):3759–3771. https://doi.org/10.1093/jxb/ery176
Wu P, Shou H, Xu G, Lian X (2013) Improvement of phosphorus efficiency in rice on the basis of understanding phosphate signaling and homeostasis. Curr Opin Plant Biol 16(2):205–212. https://doi.org/10.1016/j.pbi.2013.03.002
Xiao Z, Sun X, Liu X, Li C, He L, Chen S, Su J (2016) Selection of reliable reference genes for gene expression studies on Rhododendron molle G. Don Front Plant Sci 7:1547. https://doi.org/10.3389/fpls.2016.01547
Yang Xu, Shi Q, Gong B (2020) Characterization of COMT1-mediated low phosphorus resistance mechanism by metabolomics in tomato plants. Environ Exp Bot 179:104187. https://doi.org/10.1016/j.envexpbot.2020.104187
Yu B, Xu C, Benning C (2002) Arabidopsis disrupted in SQD2 encoding sulfolipid synthase is impaired in phosphate-limited growth. P Natl Acad Sci USA 99(8):5732–5737. https://doi.org/10.1073/pnas.082696499
Zhu SN, Chen ZJ, Xie BX, Guo Q, Chen MH, Liang CY, Bai ZL, Wang XR, Wang HC, Liao H, Tian J (2021) A phosphate starvation responsive malate dehydrogenase, GmMDH12 mediates malate synthesis and nodule size in soybean (Glycine max). Environ Exp Bot, Volume 189:104560, ISSN0098-8472. https://doi.org/10.1016/j.envexpbot.2021.104560
Zhao M, Guo R, Li M, Liu Y, Wang X, Fu H, Wang S, Liu X, Shi L (2020) Physiological characteristics and metabolomics reveal the tolerance mechanism to low nitrogen in Glycine soja leaves. Physiol Plantarum 168:819–834. https://doi.org/10.1111/ppl.13022
Zhang S, Tang D, Korpelainen H, Li C (2019) Metabolic and physiological analyses reveal that Populus cathayana males adopt an energy-saving strategy to cope with phosphorus deficiency. Tree Physiol 39(9):1630–1645. https://doi.org/10.1093/treephys/tpz074
Zhang J, Yang D, Li M, Shi L (2016) Metabolic profiles reveal changes in wild and cultivated soybean seedling leaves under salt stress. PLoS One 11(7):e0159622. https://doi.org/10.1371/journal.pone.0159622
Acknowledgments
We thank International Science Editing (http://www.internationalscienceediting.com) for editing this manuscript.
Funding
This work was supported by the National Natural Science Foundation of China (No. 32072012) and Natural Science Foundation of Jilin Province, China (No. 20200201134JC).
Author information
Authors and Affiliations
Contributions
Software: Jing Chen and Ji Zhou; Project administration, Methodology, Writing-review & editing, Supervision: Lianxuan Shi and Tao Zhang; Data curation: Jing Chen, Ji Zhou and Mu Li; Formal analysis: Jing Chen, Yunan Hu, and Mingxia Li.
Corresponding authors
Ethics declarations
Competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Responsible Editor: Jiayin Pang.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information

Figure S3
Changes in Additional Outcomes. Line diagrams showing changes in scores on a) the Word List Total Recall, b) the NIH Toolbox Fluid Cognition Composite, c) PROMIS anxiety, d) PROMIS depression, and e) PROMIS Satisfaction with Social Roles in the active and sham tDCS groups across baseline and follow-up (x-axis). Bars represent 95% confidence intervals (PNG 27 009 kbIntegration network of metabolites and genes in cultivated soybean seedling roots under low-phosphorus stress. Genes and metabolites are represented by numbers and letters, respectively. 1, Glyma.17G070500.Wm82.a2.v1; 2, Glyma.11G171400.Wm82.a2.v1; 3, Glyma.06G158700.Wm82.a2.v1; 4, Glyma.10G194800.Wm82.a2.v1; A, glutamic acid; B, proline; C, putrescine; D, pyruvic acid; E, 4-aminobutyric acid; F, alpha-ketoglutaric acid; G, asparagine; H, succinic acid; I, fumaric acid; J, aspartic acid; K, citric acid; L, salicylic acid. The thicker the edge is, the stronger the correlation is. The size of a node is proportional to the correlation between nodes (PNG 359 kb)
Table S1
Primers for qRT-PCR of genes in wild and cultivated soybean seedling roots (DOCX 14 kb)
Table S2
KEGG pathway of DEGs in wild and cultivated soybean seedling roots (DOCX 21 kb)
Table S3
Changes of transcription factors in wild and cultivated soybean seedling roots under low-phosphorus stress (DOCX 18 kb)
Table S4
Contribution rate of wild and cultivated soybean seedling root metabolites to the first and second principal components (PC1 and PC2, respectively) (DOCX 25 kb)
Table S5
DEGs co-expressed in wild and cultivated soybean seedling roots under low-phosphorus stress (DOCX 15 kb)
Table S6
KEGG annotation, GO annotation and log2(LP/CK) for some DEGs in cultivated soybean seedling roots under low-phosphorus stress (DOCX 14 kb)

Supplementary Figure 1.
qRT-PCR of genes in the roots of wild and cultivated soybean seedlings (PNG 57.1 KB)

Supplementary Figure 2.
DEGs of wild and cultivated soybean seedling roots under low-phosphorus stress and control: (a)Venn and (b) volcano diagrams (PNG 132 KB)
Rights and permissions
About this article
Cite this article
Chen, J., Zhou, J., Li, M. et al. Membrane lipid phosphorus reusing and antioxidant protecting played key roles in wild soybean resistance to phosphorus deficiency compared with cultivated soybean. Plant Soil 474, 99–113 (2022). https://doi.org/10.1007/s11104-022-05316-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-022-05316-5