Abstract
Main conclusion
Coffea karyotype organization and evolution has been uncovered by classical cytogenetics and cytogenomics. We revisit these discoveries and present new karyotype data.
Abstract
Coffea possesses ~ 124 species, including C. arabica and C. canephora responsible for commercial coffee production. We reviewed the Coffea cytogenetics, from the first chromosome counting, encompassing the karyotype characterization, chromosome DNA content, and mapping of chromosome portions and DNA sequences, until the integration with genomics. We also showed new data about Coffea karyotype. The 2n chromosome number evidenced the diploidy of almost all Coffea, and the C. arabica tetraploidy, as well as the polyploidy of other hybrids. Since then, other genomic similarities and divergences among the Coffea have been shown by karyotype morphology, nuclear and chromosomal C-value, AT and GC rich chromosome portions, and repetitive sequence and gene mapping. These cytogenomic data allowed us to know and understand the phylogenetic relations in Coffea, as well as their ploidy level and genomic origin, highlighting the relatively recent allopolyploidy. In addition to the euploidy, the role of the mobile elements in Coffea diversification is increasingly more evident, and the comparative analysis of their structure and distribution on the genome of different species is in the spotlight for future research. An integrative look at all these data is fundamental for a deeper understanding of Coffea karyotype evolution, including the key role of polyploidy in C. arabica origin. The ‘Híbrido de Timor’, a recent natural allotriploid, is also in the spotlight for its potential as a source of resistance genes and model for plant polyploidy research. Considering this, we also present some unprecedented results about the exciting evolutionary history of these polyploid Coffea.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Coffee is one of the most important beverages worldwide, with more than 2.25 billion cups consumed daily (Denoeud et al. 2014). More than 75% of traded coffee is produced by Coffea arabica L. and 35% by Coffea canephora Pierre ex A. Froehner. Coffea L. (Rubiaceae) genus has a recent origin (7.87 Mya, Tosh et al. 2013) and includes more than 124 species native to Africa, Australasia, Comoros, India, Madagascar, Mascarenes and Papua New Guinea. Almost all diploid (2n = 2x = 22) species are self-incompatible, with few exceptions, such as C. anthonyi Stoff. & F. Anthony. C. arabica is the only polyploid in the genus, with 2n = 4x = 44 chromosomes, and is self-compatible (Hamon et al. 2017).
Cytogenetics concerns genomic and epigenomic aspects of the chromosomes, including the number, structure, organization, function, evolution, and behavior during the cell cycle and meiosis (Singh 2016; Deakin et al. 2019). Coffea cytogenetics was initiated in the early twentieth century with the work of von Faber (1912). As in other plant species, limitations have hampered the cytogenetic investigations in Coffea. The main problem has been to obtain mitotic and/or meiotic chromosomes, i.e., the biological material to conduct cytogenetic studies. Even today, one of the main challenges is to seek alternatives to obtain Coffea chromosomes, mainly to cytogenomic experiments involving in situ hybridization applications. In addition, the small size of the chromosomes and low metaphasic index from root meristems also hampered Coffea cytogenetics. The further advance of protocols allowed the progress in Coffea chromosome complement research, such as the analysis of pachytene chromosomes (Pinto-Maglio and Cruz 1987), improved methods for mitotic chromosome preparation and the replacement of root meristems by cell aggregate suspensions (CAS, Clarindo and Carvalho 2006). From this, karyotype has been expanded and refined, contributing to a better understanding of the Coffea genome. So, Coffea cytogenetics is important for taxonomic and evolutionary studies, DNA sequence mapping, functional genome annotation (Hamon et al. 2009; Yuyama et al. 2012) and coffee breeding programs (Clarindo and Carvalho 2009). In this review, we revisited all cytogenetic Coffea studies to report the advances and contributions in this genomic era, as well as the main challenges and perspectives for further studies. In addition, we report new data about the 2n karyotype of the diploid C. eugenioides and the polyploid “Híbrido de Timor '' (HT), expanding the genomic data about Coffea and its diversification and evolution.
Initial steps—chromosome number determination
The first chromosome counting in a Coffea from micro- and megasporogenesis evidenced that C. arabica shows 2n = 16 chromosomes (von Faber 1912). Twenty years later, Homeyer (1932) counted 2n = 22 chromosomes for the same species. Based on chromosome numbers of other Rubiaceae genera, such as Sherardia L., Crucianella L., Asperula L. and Galium L., Homeyer suggested the basic chromosome number of x = 11 for Coffea. Most striking breakthroughs in Coffea cytogenetics were achieved in the 1930s, mainly at the Instituto Agronômico de Campinas, Brazil, and by French and Belgian researchers. Krug (1934) reported 2n = 44 chromosomes for five C. arabica varieties (‘Nacional’, ‘Bourbon’, ‘Laurina’, ‘Maragogipe’ and ‘Amarelo de Botucatu’) and 2n = 22 for C. canephora, Coffea liberica Hiern and Coffea congensis A. Froehner, confirming the basic chromosome number of x = 11. Therefore, from the 2n chromosome number, C. arabica was noticed as a tetraploid species of Coffea (Krug 1934) and started to stand out not only for its economic relevance but also for the genomic events involved in its karyotype evolution. Despite being a tetraploid, C. arabica demonstrates a diploid meiotic behavior, forming only bivalents from prophase I until metaphase I. Although Krug did not carry out a detailed morphological analysis, he described the Coffea chromosomes as small (approximately 1–2 µm) and homomorphic. Due to these karyotype features, some cytogenetic applications have been applied to characterize the chromosomes and to understand the evolution of the Coffea genome. In the following years, the chromosome number of many other species, varieties and hybrids of Coffea was determined as 2n = 22 or 2n = 44 (Krug 1937; Mendes 1938; Bouharmont 1959, 1963; Sybenga 1960; Conagin and Mendes 1961).
Karyotype morphology and the first karyograms
Coffea chromosomes have been considered a hindrance for karyogram assembly due to their small size and high similar morphology. Mendes (1938) was the first to report a morphological characterization of Coffea chromosomes, describing solely those of Coffea liberica ‘Dewevrei’ (De Wild. & T. Durand) Lebrun (syn. Coffea excelsa Chev.). Although C. liberica ‘Dewevrei’ chromosomes were larger compared to those of other Coffea species, they were also considered small and morphologically, ranging from 3.5 (chromosome 1) to 1.5 µm (chromosome 11). Only three classes of chromosomes were distinguishable based on their total length, being designated as A, B and C. Class A was composed of three chromosome pairs with 2–3.5 µm, class B of four pairs with around 2 µm, and class C of four pairs with 1–2 µm (Mendes 1938). Bouharmont (1959, 1963) determined the number and length of the mitotic chromosomes of sixteen Coffea species, characterizing them as small and homomorphic and highlighting the hypothesis that the Coffea basic chromosome number is x = 11. Owing to the high intra- and interspecific similarity among chromosomes, the author was only able to distinguish five (1, 2, 3, 4 and 11) of the eleven chromosomes.
Pachytene chromosomes allowed to overcome the hindrance associated with the small size of mitotic metaphase chromosomes. Pachytene chromosome characterization bearing the NOR provided insights on the C. arabica polyploid origin, by comparing the bivalent morphology with those of diploid species. One C. arabica NOR bivalent was similar to the Coffea eugenioides S. Moore and C. liberica ‘Dewevrei’ with regard to total and arm lengths. The other C. arabica NOR bivalent showed a simpler chromomeric pattern, most similar to the bivalents II of Coffea salvatrix Swynn. & Philipson and Coffea racemosa Lour (Pinto-Maglio and Cruz 1987).
C. arabica 22 pachytene chromosomes were after characterized, being four metacentric (1, 8, 11 and 18), fourteen submetacentric (2, 3, 4, 5, 6, 7, 9, 10, 12, 15, 16, 17, 19 and 20) and four acrocentric (13, 14, 21 and 22) chromosomes. Three bivalents presented the NOR (14, 20 and 21). In general, the chromomeres were located near the centromere, and similarities concerning the chromomeric pattern were shown among 54% of the 22 bivalents of C. arabica, evidencing some level of homoeology between them. Based on these cytogenetic markers and by gathering information from the bibliography, the authors suggested that such similarities among C. arabica bivalents are consistent with a segmental allopolyploid origin (Pinto-Maglio and Cruz 1998). So, the mitotic cytogenetics showed the ploidy level of the Coffea species, mainly the tetraploidy of C. arabica, and the meiotic cytogenetics indicated the genomic origin of this species. Unfortunately, new data has not been published about the Coffea meiotic chromosome.
Besides the small size and homogeneous morphology of Coffea chromosomes, the low seed germination rate and low metaphase index have also been pointed out as hindrances for cytogenetics (Conagin and Mendes 1961; Iacia and Pinto-Maglio 2013). To overcome this barrier, our research group (Clarindo and Carvalho 2006, 2008, 2009; Clarindo et al. 2012) replaced the root meristems by in vitro CAS as material source (Box 1, Fig. 1). Thereby, high metaphase indexes for C. canephora, C. congensis, C. eugenioides and C. arabica have been achieved, allowing to obtain chromosomes with total lengths of up to 5 µm (5.30 µm for chromosome 1 of C. arabica, for instance, Clarindo and Carvalho 2008). Such chromosomes were suitable for the assembly of the first karyograms at distinct levels of chromatin condensation, as well as the application of banding procedures. C. canephora and C. congensis possess 2n = 2x = 22 chromosomes, with two metacentric (4 and 9) and nine submetacentric pairs (1, 2, 3, 5, 6, 7, 8, 10 and 11). The submetacentric chromosome pair 6 of both species shows a secondary constriction (SC) in the short arm, which was confirmed for C. canephora by Ag-NOR and Hsc-FA. C. arabica has 2n = 4x = 44 chromosomes, being 5 pairs classified as metacentric (7, 8, 13, 14 and 20), 16 as submetacentric (1–6, 9–12, 15–19 and 21) and one pair as acrocentric (22).
“Híbrido de Timor '' (HT), a natural hybrid between C. canephora and C. arabica, had its karyotype characterized and karyogram assembled by our adapted protocol. The semi-fertile HT ‘CIFC 4106’, a vegetatively propagated accession derived from the original HT plant, shows 2n = 3x = 33 chromosomes. Therefore, it was considered an allotriploid (Clarindo et al. 2013). We classified the chromosomes, assembled the HT ‘CIFC 4106’ karyogram and showed it for the first time now. Due to its anorthoploid condition (odd number of chromosome complements), the karyotype exhibits not only chromosome pairs, but also individual ones and groups of three or four chromosomes (Fig. 2). Some chromosomes appear to be more similar to those of C. canephora or C. arabica, providing new cytogenetic information regarding the allopolyploid origin of HT. Such evolutionary aspects of the Coffea genome and interspecific hybridizations throughout the history of the genus will be discussed in the following topics.
HT ‘CIFC karyogram 4106’ evidencing individual and grouped chromosomes, which were defined according to the total length and classification. 19 chromosomes (1, 4–9, 12–14, 19, 22–24, 27, 30–33) showed at least one particular cytogenetic feature. The paired chromosomes were 2–3, 10–11, 20–21, 25–26, and 28–29; and the chromosomes grouped in four were 15–18. Bar = 5 μm
Chromosome banding and the onset of Coffea cytogenomics
C- and NOR-banding were initially applied in Coffea, mapping the constitutive heterochromatin and NOR, respectively. C-bands in Coffea occur preferentially at pericentromeric/centromeric regions. Corroborating to karyotype characterization, C-banding shows the karyotype asymmetry of the genus, varying from 62.3% (C. eugenioides) and 64.32% (C. liberica), with the prevalence of submetacentric chromosomes. On the other hand, the number of active NOR sites vary, with C. kapakata, C. congensis, C. eugenioides, C. liberica, C. canephora and C. liberica var. dewevrei displaying one pair with a positive NOR band (presumably number 3), and C. racemosa, C. salvatrix and C. stenophylla with two positive bands in chromosomes 1 and 3 (Pierozzi et al. 1999, 2012; Pierozzi 2013).
4′-6-diamidino-2-phenylindole (DAPI+) and Chromomycin A3 (CMA3+) differential staining have been carried out in Coffea species to map the AT- and GC-rich regions, respectively, contributing to identify specific karyotype signals in several species, even those with genetic or botanical status still under debate. DAPI+ bands were only found in C. stenophylla among 12 analyzed Coffea species (Pinto-Maglio et al. 2000, 2001; Barbosa et al. 2001; Lombello and Pinto-Maglio 2004a, b, c). After DNA denaturation/renaturation in a fluorescent in situ hybridization (FISH), specific interstitial DAPI+ bands in nine Coffea species were detected. CMA3+ bands were found in all Coffea species studied so far. In general, the strongest CMA3+ fluorescence signals are associated with the SC and co-localized with the ribosomal DNA (rDNA) loci detected by FISH. Additional, CMA3+ bands may also occur adjacent to, or interspersed with, the interstitial 5S rDNA (Hamon et al. 2009).
Karyotype knowledge obtained from classical cytogenetics has been the basis for queries, which have been investigated mainly by cytogenomics, allowing a more refined Coffea genome characterization. Cytogenomics made possible to: (1) discover the diploid progenitors of C. arabica (Raina et al. 1998; Lashermes et al. 1999): (2) obtain information for application in breeding programs (Herrera et al. 2007); and (3) obtain a larger amount of details on the organization and evolution of Coffea genomes (Hamon et al. 2009). The identity of C. arabica diploid progenitors has been in the spotlight of Coffea research since the discovery of its tetraploid condition by Krug (1934). Currently, wild C. arabica populations are found mainly in the Afromontane rainforests of southwest Ethiopia and the Boma Plateau of Sudan (Bawin et al. 2020). Although the geographic range of C. arabica does not overlap with that of any other Coffea species, the closeness with wild populations of C. eugenioides and of the Canephoroid species C. canephora, C. congensis and C. brevipes (Benth.) H.S. Irwin & Barneby, made these the most likely diploid progenitors. The first phylogenetic inferences based on chloroplast genes and rDNA confirmed the high genetic proximity among these species (Lashermes et al. 1995).
From genomic DNA probes (genomic in situ hybridization—GISH) of four diploid Coffea species, 22 C. arabica chromosomes hybridized preferentially with C. eugenioides probes, while the other 22 chromosomes hybridized more strongly with the C. congensis probes (Raina et al. 1998). Also using GISH, Lashermes et al. (1999) suggested that C. canephora and C. eugenioides, or their related ecotypes, are most likely the two diploid progenitors of the tetraploid C. arabica. In a phylogenomic approach using genotyping-by-sequencing, the genetic distances were estimated between C. arabica and 23 other species, including all those known as the most closely related to C. arabica, as C. eugenioides, C. canephora, C. congensis and C. brevipes. C. eugenioides and C. canephora were confirmed as the putative female and male progenitor species, respectively, that hybridized between 1.08 million and 543 thousand years ago (Bawin et al. 2020).
Integrating the DAPI+, CMA3+ and rDNA sites mapping, two to five chromosome pairs were discriminated from the karyotypes of 16 Coffea species, including the allotetraploid C. arabica. However, the results were not suitable to discriminate between C. canephora and C. congensis as putative progenitors. Differences of the number of 18S and 5S rDNA loci were revealed and related to the biogeographical region of the 16 analyzed species. Most of the East African species, as C. eugenioides, C. salvatrix and C. racemosa, possessed two chromosome pairs with the 18S rDNA locus and one pair with the 5S rDNA, while the majority of the West and Central African species exhibited one chromosome pair with the 18S and two with the 5S rDNA (Hamon et al. 2009).
Alien chromatin in interspecific hybrids between C. arabica and C. canephora, as well as in an introgressed line derived from C. arabica and C. liberica crossing, was identified from GISH and BAC-FISH (fluorescence in situ hybridization using bacterial artificial chromosomes). GISH results from the interspecific hybrids revealed close affinity between C. arabica and C. canephora genomes, evidencing that a low rate of structural modifications has occurred in both genomes since C. arabica speciation (Bawin et al. 2020). In addition, GISH/BAC-FISH in the introgressed line karyotype detected and physically located the C. liberica-introgressed DNA sequences carrying the SH3 factor for resistance against Hemileia vastatrix Berk. & Broome (Herrera et al. 2007).
Coffea genomics and its integration with cytogenetics
During the 1990’s, the availability of DNA based molecular markers allowed a rapid progress in coffee genomics. As for other crops, early genomic studies in Coffea were mainly focused on assessing the genetic diversity and phylogenetic relationships, constructing genetic maps and identifying quantitative trait loci (QTLs) (reviewed by Lashermes et al. 2008; de Kochko et al. 2010). Genetic diversity was investigated from several molecular markers in wild Coffea species and, with a larger effort, in C. arabica and C. canephora. The considerably narrow genetic basis of both wild and cultivated C. arabica populations was the most relevant feature noticed by these studies (Vossen 1985; Lashermes et al. 1996; Scalabrin et al. 2020). This bottleneck is mainly explained by the autogamous reproduction system, with a rate of outcrossing around 10%, and the founder effect resulted from the small number of individuals introduced to America in the first commercial plantations (Carvalho and Krug 1949; Setotaw et al. 2013). In addition, C. arabica origin from a single allopolyploidization event also contributed to the low genetic diversity observed for the species (Lashermes et al. 2014). Such low genetic variability is a challenge for the identification and selection of superior genotypes. For this, molecular markers have played a fundamental role as to discriminate between genotypes (Sousa et al. 2017). Due to low genetic variability, breeding programs devoted efforts on the introgression of interesting genes from different species and hybrids, including C. canephora (Lashermes et al. 2011), C. liberica (Prakash et al. 2004) and HT (Setotaw et al. 2020). As the genomes of Coffea species exhibit considerable similarities, interspecific crossing (hybridization) is possible and often used for gene introgression (Charrier and Berthaud 1985; Anthony et al. 2011).
For C. canephora, the first genetic maps were constructed as soon as molecular markers became available for Coffea (reviewed by de Kochko et al. 2010). In 2014, the most complete genome sequence of C. canephora (accession number: PRJEB4211) was published along with a high-density genetic map constructed using several molecular markers and 3,230 loci distributed on 11 linkage groups, the same basic chromosome number of this species (x = 11). C. canephora genetic map was integrated with the sequenced genome, which covered 80% of the 710 Mbp total genome. There was a considerable variation in the physical:genetic map distances, with crossing overs occurring with higher frequencies in regions with lower density of repeats (Denoeud et al. 2014). The complexity of the C. arabica allotetraploid genome and its low genetic diversity were the main challenges to construct a high-density linkage map. Notwithstanding, a high-density linkage map was published for this species showing 22 linkage groups with 848 markers. This genetic map was also successfully used to identify QTLs associated with coffee yield, plant height, and bean size (Moncada et al. 2016).
As for the construction of genetic maps, the sequencing of a polyploid genome is also challenging, since the presence of homoeologous chromosomes hampers the assembly of each haploid complement. C. arabica draft genome (public accession number: PRJNA554647) was released in 2020, with the two component subgenomes independently assembled. In accordance with previous studies reported here, a low genetic diversity was confirmed, which was likely caused by a severe bottleneck resulting from a single event of polyploidization at the origin of the species C. arabica (Scalabrin et al. 2020). Moreover, two other genomic sequences have also been reported for C. arabica (NCBI: GCA_003713225.1 and Phytozome: genome ID 453). In addition to C. canephora and C. acabica, the C. eugenioides (accession number: PRJNA497891) and Coffea humblotiana Baill. (accession number: PRJNA665152, Raharimalala et al. 2021) genomes are also publicly available. The species C. humblotiana is the sole species from the Coffea genus endemic to the Comoros archipelago and that was probably cultivated and consumed in the past. The most noteworthy feature of C. humblotiana is the complete absence of caffeine in seeds and leaves, which is explained by the loss of one of the Caffeine Synthase (DXMT) genes, which converts theobromine into caffeine, most likely through illegitimate recombination (Raharimalala et al. 2021). Despite the significant difference in genome size, there is a high degree of synteny between the genomes of C. humblotiana (2C = 0.97 pg, Razafinarivo et al. 2012) and C. canephora (2C = 1.43 pg, Clarindo et al. 2012) with 32.16% more nuclear DNA content.
Recent advances in next-generation sequencing technologies and bioinformatic tools provided a growing amount of available genomic data, laying the background for the integration with cytogenetics and giving rise to the cytogenomics era (Talukdar and Sinjushin 2015). The integration of the linkage and cytogenetic maps and sequencing data is fundamental to define genome regions that are not yet well characterized (Kim et al. 2005). Because linkage map distances are based on recombination rates and not simply related to physical distances, physical mapping is also needed to confirm the locations of DNA sequences (Koo et al. 2008). Physical mapping might be obtained through genome sequencing or in situ localization using cytogenomics. The major challenge related to sequencing is the resolution of complex repetitive sequences. As the DNA must be fragmented into small contigs, usually around 100 bp, repeats create computational ambiguities during alignment and assembly, which might produce biases and errors when interpreting results (Schatz et al. 2012; Treangen and Salzberg 2012). Cytogenomics, on the other hand, is efficient to map repetitive sequences, revealing the physical localization in situ on the mitotic or meiotic chromosomes (Larracuente and Ferree 2015). Therefore, the linkage and cytogenetic maps and the sequencing are complementary, and integrating these data is fundamental for a more detailed and accurate Coffea genome knowdelege.
Completely Coffea sequenced genome released in 2014 for C. canephora showed that mobile elements represent more than 50% of its genome, among which ~ 85% belong to the LTR-retrotransposon class (Denoeud et al. 2014). Due to their high frequency (~ 15% to > 70%) and pivotal roles in plant genome organization, function and evolution, the analysis of mobile elements sequence (for example, the genes and repetitive sequences) and distribution along the genome has been performed in several plant species (Civáň et al. 2011; Wicker et al. 2018). Mobile elements comprise DNA sequences with the ability to insert themselves (transposons) or new copies of themselves (retrotransposons, RTEs) into new locations within a genome (Civáň et al. 2011). For the Coffea sequenced genomes analyzed so far, the proportions of LTR-retrotransposons, for instance, varied from 32% for C. humblotiana (1C = 0.49 pg, Razafinarivo et al. 2012) to 53% for Coffea heterocalyx Stoff. (1C = 0.87 pg, Noirot et al. 2003). The variation in the abundance and types of different mobile elements can reflect the divergence of botanical groups and also the evolution of species within these botanical groups (Guyot et al. 2016). From the genomic data provided by sequencing technologies, DNA sequence probes have been constructed for cytogenetic mapping of mobile elements. From this, the comparative cytogenomic analysis of the distribution patterns among different species can be performed. Therefore, this integration between genomics and cytogenetics provides valuable information to understand the karyotype evolution in Coffea.
Some mobile elements have been mapped in Coffea species, including DNA transposons (Lopes et al. 2013), Long Terminal Repeat (LTR) retrotransposons (Yuyama et al. 2012; Herrera et al. 2013; Lopes et al. 2013) and centromeric retrotransposons (Nunes et al. 2018). Nonetheless, the low longitudinal resolution of chromosomes did not allow the precise mapping, but an overview of distribution patterns along the genomes. DNA transposons MuDR and Tip100, for instance, exhibit a preferential clustering in terminal positions of C. canephora and C. eugenioides chromosomes, while C. arabica showed larger numbers of interstitial signals. This distribution indicates an increased transposition activity in the allotetraploid (Lopes et al. 2013), which is consistent with the well-known hypothesis that polyploidization can induce a burst in mobile element activity (Vicient and Casacuberta 2017).
The detailed analysis of the centromeric mobile elements composition of C. arabica, C. canephora and C. eugenioides revealed a considerable diversity in centromeric retrotransposons of Coffea (CRC) from the Ty3/Gypsy superfamily, which were divided in ten groups according to the sequence and similarity of the Reverse Transcriptase domain. Generalist probes for these CRCs exhibited a variable fluorescence signal pattern among species and among chromosomes of the same species. While in C. eugenioides the signals were identified only to centromeric regions, in C. canephora and C. arabica the signals appeared slightly scattered along interstitial regions and less specific to centromeres. In addition, C. arabica presented two pairs without bright signals, which might be homologous to those chromosomes without signals from the parental diploids C. canephora and C. eugenioides (one pair each) (Nunes et al. 2018).
Allopolyploidy in Coffea
Coffea genus has a recent monophyletic origin around 5–25 Mya, and none event of whole genome duplication (WGD) occurred immediately prior to or after the irradiation of the Rubiaceae family (Orozco-Castillo et al. 1996; Wu et al. 2006; Mahé et al. 2007; Cenci et al. 2010). Therefore, the common ancestral shared by all Coffea species probably had the basic chromosome number of x = 11. In addition, the diversification and speciation of each Coffea species occurred by DNA sequence changes (mutations) and small chromosomal rearrangements, which have been progressively identified and characterized through classical cytogenetics and cytogenomics (Rijo 1974; Yu et al. 2011; Denoeud et al. 2014; Raharimalala et al. 2021). A micro-collinearity analysis between orthologous BACs of C. canephora and C. arabica, for instance, evidenced a high level of sequence similarity, but numerous small chromosomal rearrangements, including inversions, deletions and insertions (Yu et al. 2011; Cenci et al. 2012).
Allotetraploidy and diversification of C. arabica
Krug’s (1934) discovery the tetraploidy of C. arabica ignited a long debate on the origin (ancestry) and evolution (“omics” changes) of this species. As mentioned here, there was an agreement that C. arabica originated from hybridization between two diploid species with similar genomes, with the potential progenitors being C. eugenioides (Berthou 1983; Lopes et al. 1984; Orozco-Castillo et al. 1996; Raina et al. 1998; Lashermes et al. 1999; Ruas et al. 2003), C. canephora (Lashermes et al. 1997, 1999; Ruas et al. 2003; Clarindo and Carvalho 2009), C. congensis (Höfling and Oliveira 1981; Lashermes et al. 1997; Raina et al. 1998) and C. brevipes (Lashermes et al. 1997). Therefore, the focus has been the C. arabica ancestors. However, further studies should be accomplished to unravel the genomic and epigenomic outcomes of the C. arabica allopolyploid condition.
C. arabica polyploid origin from a crossing between two diploids with similar genomes is a consensus, but its genomic origin classification has been discussed. Initially, C. arabica was classified as a natural segmental allotetraploid (Orozco-Castillo et al. 1996; Pinto-Maglio and Cruz 1998). ‘Segmental allopolyploid’ in this context was based on the Stebbins (1949) definition as a type of allopolyploid that contain two partially differentiated genomes, and that was originated from hybridization between species close enough to allow the partial pairing between homoeologous chromosomes. Thus, segmental allopolyploids are intermediaries between autopolyploids and true (or genomic) allopolyploids, as the differentiation between the progenitor genomes is insufficient for complete allopolyploidy. Exceptionally, the allotetraploid C. arabica exhibits a stable diploid-like meiotic behavior (Mendelian segregation). This might be possible owing to the genetic system occurrence wherein the pairing between homoeologous chromosomes is avoided (Pinto-Maglio and Cruz 1998), such as the Ph gene (homoeologous pairing suppressor) found in Triticum L., Avena L., Festuca L., Gossypium L., Nicotiana L. and Lolium L. Further molecular evidence on this hypothesis suggested that the absence of homoeologous pairing in C. arabica is not a consequence of structural differentiation between the two parental genomes, but rather the effect of one or several pair-regulating genes, which could be similar to Ph (Lashermes et al. 2000).
C. arabica was also hypothesized as an amphidiploid species formed from the crossing between C. eugenioides as female progenitor and C. canephora, or its related ecotypes, as male progenitor. In addition, C. arabica origin is recent due to the low level of divergence between the two constitutive genomes of this species and the related parental genomes (Lashermes et al. 1999). Amphidiploid refers to segmental allotetraploids that have gone through an event of WGD after hybridization (Stebbins 1949). In the absence of homoeologous pairing, the WGD event may restore de fertility of the hybrid (homoploid), since the presence of two copies of each genome would enable the Mendelian pairing (bivalent) during meiosis I. The WGD event that gave rise to the fertile allotetraploid ancestors of C. arabica possibly involved either chromosome set doubling in a diploid interspecific hybrid or backcrossing of a spontaneous triploid (Lashermes et al. 1999).
The true allotetraploid nature of C. arabica was reinforced from classical cytogenetics and chromosomal image cytometry (Box 2) analyses (Clarindo and Carvalho 2008, 2009), corroborating with data based on molecular markers and GISH. The comparison with the karyotypes of two potential genitors, C. canephora and C. congensis, revealed the presence of identical chromosomes between C. arabica and both species concomitantly. In addition, C. arabica also exhibits a small acrocentric chromosome pair that is not present in either of these two species. Taken together these results also support the idea that only one of them participated in the origin of C. arabica (Clarindo and Carvalho 2009; Clarindo et al. 2012).
Cytogenetic evidence concerning the C. arabica origin was obtained from the SC/NOR chromosome. Three pachytene chromosomes of C. arabica (14, 20 and 21) have SC (Pinto-Maglio and Cruz 1998), which were confirmed by 18S rDNA signals in mitotic metaphases (Hamon et al. 2009). C. canephora and C. congensis exhibit a single SC in the short arm of the chromosome 6 (Clarindo et al. 2012), also confirmed by 18S rDNA (Hamon et al. 2009). Therefore, one of the SC/NOR chromosomes in C. arabica karyotype was probably derived from the canephoroid genitor (most likely C. canephora), while the other two were possibly inherited from C. eugenioides.
From orthologous coding sequence divergence analysis, C. arabica evolutionary history was also evidenced as an allotetraploid originated from a natural hybridization between C. canephora and C. eugenioides. Thus, the genome of C. arabica was represented by the genome formula CaCaEaEa (Lashermes et al. 1999; Yu et al. 2011). The most accurate estimation for the genomic origin time of the natural hybridization is in between 0.54 and 1.08 Mya (Bawin et al. 2020). Although the current geographical range of C. arabica do not overlap with any other Coffea species, including its possible progenitors, pollen records and lake sediment cores from the Congo basin and East Africa indicate that the Afromontane rainforests regularly expanded to lower altitudes during glacial periods between 0.60 and 1.05 Mya. Corroborating, C. canephora and C. eugenioides occurred in contact zone in this area. The changing environmental conditions might have also played a role on this speciation event, weakening interspecific reproductive barriers between C. arabica diploid parental species. In addition, the aridification of East Africa since 0.575 Mya is one of the possible explanations for the current non-overlapped distribution areas of C. arabica, C. eugenioides and C. canephora (Owen et al. 2018; Bawin et al. 2020).
C. canephora and C. arabica ‘Tall Mokka’ comparison from BAC clones (~ 140–160 kb), bearing the aforementioned orthologous coding sequences, aimed to show the outcomes of the C. arabica allopolyploidy from a cytogenomic perspective. Despite the high degree of sequence conservation in coding regions, genomic differences were found. Major chromosomal rearrangements were observed in the intergenic regions of these BACs, including a paracentric inversion between homoeologous regions within C. arabica (Ca and Ea). As C. eugenioides was not included in this study, it was not possible to distinguish if the chromosomal inversion occurred in C. arabica or was inherited from one of the two diploid progenitors. Therefore, the inversion might not be a consequence of the C. arabica allopolyploidy. Moreover, the specific insertion of a Ty1-copia retrotransposon in the Ca sub-genome of C. arabica was also reported, which might be related to the burst in TE activity (Yu et al. 2011).
Allotriploidy of the Timor hybrid
In addition to the polyploidization event that gave rise to the well-established allotetraploid species C. arabica, another recent allopolyploidy event in the Coffea genus originated the hybrid named HT, or the Timor hybrid. HT arose in a plantation of C. arabica ‘Typica’, established around 1917/18 on the Timor Island. All the accessions of this hybrid originated from this single plant, or from backcrosses between this plant and its progenitor C. arabica (Gonçalves et al. 1978). Besides the remarkable relevance of HT as a source of resistance genes in coffee breeding, this hybrid has also gained attention for its very recent allopolyploid origin of ~ 100 years (Gonçalves et al. 1978; Capucho et al. 2009). HT ‘CIFC 4106’, which has been vegetatively propagated, is a triploid hybrid with a chromosome number 2n = 3x = 33 and a nuclear DNA content of 1C = 2.10 pg. Therefore, we considered that this accession represents the first plant of HT found in the Timor Island.
In summary, the hybridization/polyploidization events involving C. canephora, C. eugenioides, HT and C. arabica might be explained as follows. The allotetraploid C. arabica originated around 0.543 and 1.08 Mya (Bawin et al. 2020) from the fusion between a reduced reproductive cell from C. canephora (n = x = 11) and another from C. eugenioides (n = x = 11, Lashermes et al. 1999). After a polyploidization event in the sterile homoploid, fertility would be restored, resulting in the fertile allotetraploid with 2x = 4x = 44 chromosomes, being 22 from C. canephora and 22 from C. eugenioides (Ca and Ea subgenomes, respectively). A natural backcross dating from ~ 100 years ago occurred between C. arabica and its progenitor C. canephora, involving the fusion of reduced reproductive cells from both species and resulting in the HT with 2x = 3x = 33 chromosomes. Therefore, our research group recently launched the hypothesis that the genome of HT ‘CIFC 4106’ is represented by the formula CCaEa. To provide more information regarding the karyotypic evolution of C. eugenioides, C. canephora, C. arabica and HT ‘CIFC 4106’, we have been combining classical and molecular cytogenetics, as well as flow cytometry for nuclear DNA content measurements (Box 2).
The six chromosome pairs presented by HT 'CIFC 4106' (2–3, 10–11, 12–13, 20–21, 25–26, 28–29) might represent the CCa subgenomes, being C from C. canephora and Ca from C. arabica (Figs. 2, 3). A look at the HT ‘CIFC 4106’ karyogram shows that the possible pairs 2–3, 10–11, 12–13, 20–21, 25–26 and 28–29 may represent the chromosomes 1, 4, 5, 7, 9 and 11 of C. canephora, respectively (Fig. 3). HT ‘CIFC 4106’ group 15–18 is similar to pair 6 and 7 of C. canephora (Fig. 3). Another evidence that supports the CCaEa genome hypothesis, specifically regarding the E genome, is that chromosomes 1, 4 and 9 of HT ‘CIFC 4106’ are similar to 1, 2 and 3 of C. eugenioides, respectively, considering morphometry. In addition, chromosomes 7 and 11 of C. eugenioides are similar to 19 and 33 of HT ‘CIFC 4106’, respectively (Fig. 3).
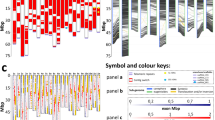
Karyograms obtained from metaphase chromosomes of: a C. eugenioides exhibiting 2 metacentric chromosome pairs (7, 10), 9 submetacentric (1–6, 8, 9 and 11) and 2 chromosomes pairs with SC (3 and 5); b C. canephora with 2 metacentric chromosome pairs (4 and 9), 9 submetacentric (1–3, 5–8, 10 and 11) and 1 with SC (chromosome 6); c C. arabica exhibits 5 metacentric chromosome pairs (7, 8, 13, 14 and 20), 16 submetacentric (1–6, 9–12, 15–19 and 21) and 1 acrocentric pair (22); and d HT 'CIFC 4106' possess 6 metacentric (10, 11, 19, 25, 26, 30) and 27 submetacentric chromosomes (1–9, 12–18, 20–24, 27–29, 31–33). Bar = 5 μm
Our new data about 5S rDNA mapping performed in C. eugenioides and C. canephora showed that the two diploids exhibit one 5S rDNA loci, in the interstitial region of chromosome 4 long arm for C. eugenioides, and in the pericentromeric portion of the long arm of chromosome 8 in C. canephora. The allotriploid HT ‘CIFC 4106’ shows two loci located interstitially in the long arm of chromosomes 9 and 13 (Fig. 4). The association of these data with chromosome class provide additional evidence for HT origin, as C. eugenioides chromosome 4 and HT 'CIFC 4106' chromosome 9 both exhibit one 5S rDNA loci in the same position. Therefore, the HT 'CIFC 4106' chromosome 9 was probably inherited from a reduced cell of C. arabica (CaEa), precisely from the Ea subgenome. In addition, the nuclear DNA content of HT 'CIFC 4106' (1C = 2.10 pg, Clarindo et al. 2013) is equivalent to the sum of the mean 2C nuclear genome size of C. canephora (2C = 1.41 pg, 1C = 0.705, Clarindo and Carvalho 2009) and the 1C nuclear value of C. eugenioides (2C = 1.38 pg, 1C = 0.690 pg, Sanglard et al. 2019) and HT 'CIFC 4106' chromosome number (2n = 3x = 33) corresponds to the fusion of one reproductive cell of C. arabica (CaEa; n = 2x = 22) and one of C. canephora (C; n = x = 11). These data also support the CCaEa hypothesis. A summary of the main cytogenetic features and evolutionary relations among the two allopolyploid Coffea and its parental diploids is depicted in Fig. 5.
FISH mapping of 5S rDNA genes (red) on Coffea metaphase chromosomes using probes labeled with tetramethyl-rhodamine 5-dUTP. Chromosomes were counterstained with 4’,6-diamidino-2-phenylindole dihydrochloride (DAPI) (blue). a C. eugenioides with the 5S rDNA signal at the interstitial region of chromosome 4 long arm. b C. canephora exhibited a pericentromeric signal in the long arm of chromosome 8. c HT ‘CIFC 4106’ has two positive 5S rDNA signals, one in chromosome 9 and the other in chromosome 13, both interstitial. Bars = 5 μm
Summary of the main cytogenetic features with representative idiograms of two Coffea allopolyploids, C. arabica and HT ‘CIFC 4106’, and the diploids C. canephora and C. eugenioides, as well as their evolutionary relations. The data presented for these Coffea were obtained from Pinto-Maglio and Cruz (1998), Noirot et al. (2003), Clarindo and Carvalho (2006, 2009), Hamon et al. (2009), Bawin et al. (2020) and from the present work. For these Coffea were reported: 1C nuclear DNA content measured by flow cytometry; 2n chromosome number; ploidy level; class of each chromosome of the karyotype (M metacentric, SM submetacentric, A acrocentric); number and chromosomes with secondary constrictions (SC); number of 18S (green) and 5S (red) rDNA sites
Natural neoallopolyploids are valuable materials intensively used as evolutionary models in plant polyploidy research, such as Spartina anglica C. E. Hubb. (Ainouche et al. 2004), Senecio cambrensis Rosser, Senecio eboracensis Abbott & Lowe (Abbott and Lowe 2004), Tragopogon mirus Ownbey, Tragopogon miscellus Ownbey (Soltis et al. 2004), Cardamine × schulzii Urbanska-Worytkiewicz (Urbanska et al. 1997) and Mimulus peregrinus Vallejo-Marín (Vallejo-Marín 2012). As HT is a recent allotriploid hybrid originated only ~ 100 years ago, the refined study of its karyotype, in comparison to its progenitor C. arabica, would certainly provide substantial information on the genomic rearrangements associated with interspecific hybridization and polyploidy. Although not an established species due to the allotriploid condition and meiosis irregularities, HT may also be considered an interesting asset for evolutionary research regarding plant polyploidy in the context of natural populations.
Concluding remarks
Advances in Coffea cytogenetics over the years have been remarkable. Thus far, the development of improved protocols for classical and molecular cytogenetics, alongside with genomics, allowed a deeper understanding on the structure, organization and evolution of Coffea genomes. The relevance of these data also reaches applied research, with flow cytometry and karyotyping being used, for instance, to distinguish species, varieties and hybrids, including HT cytotypes. The potential role of cytogenetics and flow/image cytometries on aiding Coffea genomes assembly is also worthy of notice (Box 2). The aim of any genome sequencing project is to achieve an assembly to the chromosome level, with each scaffold assigned and oriented onto a chromosome. Therefore, the refinement of sequence contigs assignment to chromosomes often requires the integration between the complementary data obtained from sequencing technologies and molecular cytogenetic mapping. The basis of another relevant information, sequencing coverage, relies on reliably measuring the total genome size of the target species, for which flow cytometry has been considered the ‘gold standard technique’. In addition to nuclear genome size, both flow and image cytometries also allow measuring the size of each chromosome of the karyotype, aiding to estimate the coverage at the chromosome level.
The increase of the sequencing information will also allow a deeper analysis of Coffea evolution, and the integration with the molecular cytogenetics is fundamental to associate the ex situ with the in situ localization of DNA sequences. Nonetheless, mitotic metaphases and pachytene chromosomes from different species of Coffea, especially wild species of strict geographical distribution, will still be one of the main challenges for cytogenomics. Partial genome sequence analysis already revealed interesting patterns regarding the association of mobile elements with Coffea diversification and the recent publication of a new genome sequence from the wild C. humblotiana will open new fields for comparative analysis. We also believe that the allotriploid HT is an interesting target for cytogenomics, especially to study the early genomic dynamics after an allopolyploidization event, such as the TE burst, sequence losses or gains and global effects on epigenetic regulation. Therefore, we hope that the continuous improvement in techniques and a closer integration of different research areas will lay the ground for future ground-breaking discoveries about Coffea genome.
Author contribution statement
All authors equally contributed to this work and approved the final manuscript version for submission.
Data availability statement
All data generated or analyzed during this study are included in this published article.
References
Abbott RJ, Lowe AJ (2004) Origins, establishment and evolution of new polyploid species: Senecio cambrensis and S. eboracensis in the British Isles. Biol J Linn Soc 82:467–474. https://doi.org/10.1111/j.1095-8312.2004.00333.x
Ainouche ML, Baumel A, Salmon A (2004) Spartina anglica C. E. Hubbard: a natural model system for analysing early evolutionary changes that affect allopolyploid genomes. Biol J Linn Soc 82:475–484. https://doi.org/10.1111/j.1095-8312.2004.00334.x
Anthony F, Bertrand B, Etienne H, Lashermes P (2011) Coffea and Psilanthus. Wild crop relatives: genomic and breeding resources. Springer, Berlin, pp 41–61
Barbosa R, Pierozzi N, Silvarolla M, Pinto-Maglio C (2001) Caracterização citomolecular de três espécies de Coffea L.: C. eugenioides, C. canephora, e C. dewevrei. In: II Simpósio De Pesquisa Dos Cafés Do Brasil. Vitória, Brazil, pp 293–298
Bawin Y, Ruttink T, Staelens A et al (2020) Phylogenomic analysis clarifies the evolutionary origin of Coffea arabica. J Syst Evol. https://doi.org/10.1111/jse.12694
Berthou F (1983) Chloroplast and mitochondrial DNA variation as indicator of phylogenetic relationships in the genus Coffea L. Theor Appl Genet 65:77–84. https://doi.org/10.1007/BF00276268
Bouharmont J (1963) Somatic chromosomes of Coffea species. Euphytica 12:254–257
Bouharmont J (1959) Recherches sur les affinités chromosomiques dans le genre Coffea. Publications de l’Institut National pour l’Etude Agronomique du Congo Belge
Capucho AS, Caixeta ET, Zambolim EM, Zambolim L (2009) Herança da resistência do Híbrido de Timor UFV 443–03 à ferrugem-do-cafeeiro. Pesq Agropec Bras 44:276–282. https://doi.org/10.1590/S0100-204X2009000300009
Carvalho A, Krug C (1949) Agentes de polinização da flor do cafeeiro (Coffea arabica L.). Bragantia 9:11–24. https://doi.org/10.1590/S0006-87051949000100002
Cenci A, Combes MC, Lashermes P (2010) Comparative sequence analyses indicate that Coffea (Asterids) and Vitis (Rosids) derive from the same paleo-hexaploid ancestral genome. Mol Genet Genom 283:493–501. https://doi.org/10.1007/s00438-010-0534-7
Cenci A, Combes MC, Lashermes P (2012) Genome evolution in diploid and tetraploid Coffea species as revealed by comparative analysis of orthologous genome segments. Plant Mol Biol 78:135–145. https://doi.org/10.1007/s11103-011-9852-3
Charrier A, Berthaud J (1985) Botanical classification of coffee. In: Coffee. Springer, pp 13–47
Civáň P, Švec M, Hauptvogel P (2011) On the coevolution of transposable elements and plant genomes. J Bot 2011:1–9. https://doi.org/10.1155/2011/893546
Clarindo W, Carvalho C (2006) A high quality chromosome preparation from cell suspension aggregates culture of Coffea canephora. Cytologia (tokyo) 71:243–249. https://doi.org/10.1508/cytologia.71.243
Clarindo WR, Carvalho CR (2008) First Coffea arabica karyogram showing that this species is a true allotetraploid. Plant Syst Evol 274:237–241. https://doi.org/10.1007/s00606-008-0050-y
Clarindo WR, Carvalho CR (2009) Comparison of the Coffea canephora and C. arabica karyotype based on chromosomal DNA content. Plant Cell Rep 28:73–81. https://doi.org/10.1007/s00299-008-0621-y
Clarindo WR, Carvalho CR, Mendonça MAC (2012) Cytogenetic and flow cytometry data expand knowledge of genome evolution in three Coffea species. Plant Syst Evol 298:835–844. https://doi.org/10.1007/s00606-012-0595-7
Clarindo WR, Carvalho CR, Caixeta ET, Koehler AD (2013) Following the track of “Híbrido de Timor” origin by cytogenetic and flow cytometry approaches. Genet Resour Crop Evol 60:2253–2259. https://doi.org/10.1007/s10722-013-9990-3
Conagin C, Mendes A (1961) Pesquisas citológicas e genéticas em três espécies de Coffea. Auto-compatibilidade em Coffea canephora Pierre ex Froehner. Bragantia 20:1–74
Cros J, Gavalda M-C, Chabrillange N et al (1994) Variations in the total nuclear DNA content in African Coffea species (Rubiaceae). Café Cacao Thé 38:3–3
Cros J, Combes M-C, Chabrillange N et al (1995) Nuclear DNA content in the subgenus Coffea (Rubiaceae): inter-and intra-specific variation in African species. Can J Bot 73:14–20. https://doi.org/10.1139/b95-002
Deakin J, Potter S, O’Neill R et al (2019) Chromosomics: bridging the gap between genomes and chromosomes. Genes 10:627. https://doi.org/10.3390/genes10080627
Denoeud F, Carretero-Paulet L, Dereeper A et al (2014) The coffee genome provides insight into the convergent evolution of caffeine biosynthesis. Science 345:1181–1184. https://doi.org/10.1126/science.1255274
Fehér A, Pasternak TP, Dudits D (2003) Transition of somatic plant cells to an embryogenic state. Plant Cell Tiss Organ Cult 74:201–228. https://doi.org/10.1023/A:1024033216561
Fontes BPD (2003) Citogenética, citometria de fluxo e citometria de imagem em Coffea spp. Universidade Federal de Viçosa
Fowler M (1984) Large-scale cultures of cells in suspension. In: Laboratory procedures and their applications. Academic Press, pp 167–174
Gonçalves M, Rodrigues M, Mexia J, Daehnhardt E (1978) Melhoramento da cafeicultura em Timor face à Hemileia vastatrix B. & Br. Garcia De Orta-Sér Est Agr 5:3–10
Guyot R, Darré T, Dupeyron M et al (2016) Partial sequencing reveals the transposable element composition of Coffea genomes and provides evidence for distinct evolutionary stories. Mol Genet Genomics 291:1979–1990. https://doi.org/10.1007/s00438-016-1235-7
Hamon P, Siljak-Yakovlev S, Srisuwan S et al (2009) Physical mapping of rDNA and heterochromatin in chromosomes of 16 Coffea species: a revised view of species differentiation. Chromosome Res 17:291–304. https://doi.org/10.1007/s10577-009-9033-2
Hamon P, Grover CE, Davis AP et al (2017) Genotyping-by-sequencing provides the first well-resolved phylogeny for coffee (Coffea) and insights into the evolution of caffeine content in its species. Mol Phylogenet Evol 109:351–361. https://doi.org/10.1016/j.ympev.2017.02.009
Herrera JC, D’Hont A, Lashermes P (2007) Use of fluorescence in situ hybridization as a tool for introgression analysis and chromosome identification in coffee ( Coffea arabica L.). Genome 50:619–626. https://doi.org/10.1139/G07-034
Herrera J-C, Camayo G, De-La-Torre G, et al (2013) Identification and chromosomal distribution of copia-like retrotransposon sequences in the coffee (Coffea L.) genome. 31:268–278
Höfling J, Oliveira A (1981) A serological investigation of some Coffea species with emphasis on the origin of C. arabica L. Ciência e Cultura 33:66–72
Homeyer H (1932) Zur zytologie der Rubiaceen. Planta 18:640
Iacia AAS, Pinto-Maglio CAF (2013) Mapping pachytene chromosomes of coffee using a modified protocol for fluorescence in situ hybridization. AoB PLANTS 5:40. https://doi.org/10.1093/aobpla/plt040
Kim J-S, Islam-Faridi M, Klein PE et al (2005) Comprehensive molecular cytogenetic analysis of sorghum genome architecture: distribution of euchromatin, heterochromatin, genes and recombination in comparison to rice. Genetics 171:1963–1976. https://doi.org/10.1534/genetics.105.048215
De Kochko A, Akaffou S, Andrade AC, et al (2010) Advances in Coffea genomics. In: Advances in botanical research. Elsevier, pp 23–63
Koo D-H, Jo S-H, Bang J-W et al (2008) Integration of cytogenetic and genetic linkage maps unveils the physical architecture of tomato chromosome 2. Genetics 179:1211–1220. https://doi.org/10.1534/genetics.108.089532
Krug C (1937) Observações citológicas em Coffea. III Instituto Agronômico De Campinas 37:1–19
Krug C (1934) Contribuição para o estudo da cytologia do gênero Coffea. Instituto Agronômico de Campinas
Larracuente AM, Ferree PM (2015) Simple method for fluorescence DNA in situ hybridization to squashed chromosomes. J vis Exp JoVE 95:52288. https://doi.org/10.3791/52288
Lashermes P, Trouslot P, Anthony F et al (1996) Genetic diversity for RAPD markers between cultivated and wild accessions of Coffea arabica. Euphytica 87:59–64. https://doi.org/10.1007/BF00022965
Lashermes P, Combes MC, Trouslot P, Charrier A (1997) Phylogenetic relationships of coffee-tree species (Coffea L.) as inferred from ITS sequences of nuclear ribosomal DNA. Theor Appl Genet 94:947–955. https://doi.org/10.1007/s001220050500
Lashermes P, Combes M-C, Robert J et al (1999) Molecular characterisation and origin of the Coffea arabica L. genome. Mol Gen Genet MGG 261:259–266. https://doi.org/10.1007/s004380050965
Lashermes P, Paczek V, Trouslot P et al (2000) Single-locus inheritance in the allotetraploid Coffea arabica L. and interspecific Hybrid C. arabica X C. canephora. J Hered 91:81–85. https://doi.org/10.1093/jhered/91.1.81
Lashermes P, Combes M-C, Ansaldi C et al (2011) Analysis of alien introgression in coffee tree (Coffea arabica L.). Mol Breed 27:223–232. https://doi.org/10.1007/s11032-010-9424-2
Lashermes P, Combes MC, Hueber Y, Severac D, Dereeper A (2014) Genome rearrangements derived from homoeologous recombination following allopolyploidy speciation in coffee. Plant J 78(4):674–685. https://doi.org/10.1111/tpj.12505
Lashermes P, Combes M-C, Cros J, et al (1995) Origin and genetic diversity of Coffea arabica L. based on DNA molecular markers. ASIC 16th:10
Lashermes P, Andrade AC, Etienne H (2008) Genomics of coffee one of the world’s largest traded commodities. In: Genomics of tropical crop plants. Springer, pp 203–226
Lombello RA, Pinto-Maglio CAF (2004a) Cytogenetic studies in Coffea L. and Psilanthus Hook.f. using CMA/DAPI and FISH. Cytologia 69:85–91. https://doi.org/10.1508/cytologia.69.85
Lombello RA, Pinto-Maglio CAF (2004b) Heterochromatin and rDNA sites in Coffea L. chromosomes revealed by FISH and CMA/DAPI II: C canephora cv. Apoatã C Salvatrix and c Sessiliflora. Caryologia 57:138–143. https://doi.org/10.1080/00087114.2004.10589383
Lombello RA, Pinto-Maglio CAF (2004c) Heterochromatin and rDNA sites in Coffea L. chromosomes revealed by FISH and CMA/DAPI. I: C. humilis, C. kapakata, C. sp Moloundou and c Stenophylla. Caryologia 57:11–17. https://doi.org/10.1080/00087114.2004.10589366
Lopes C, Cunha R, Blotta L (1984) Estudo taxonômico do gênero Coffea com emprego de métodos numéricos. Turrialba 34:421–430
Lopes FR, Jjingo D, da Silva CRM et al (2013) Transcriptional activity, chromosomal distribution and expression effects of transposable elements in Coffea genomes. PLoS ONE 8:e78931. https://doi.org/10.1371/journal.pone.0078931
Mahé L, Combes M-C, Lashermes P (2007) Comparison between a coffee single copy chromosomal region and Arabidopsis duplicated counterparts evidenced high level synteny between the coffee genome and the ancestral Arabidopsis genome. Plant Mol Biol 64:699–711. https://doi.org/10.1007/s11103-007-9191-6
Mendes A (1938) Morfologia dos cromossomos de Coffea excelsa Chev. Instituto Agronômico De Campinas 56:1–8
Moncada MDP, Tovar E, Montoya JC et al (2016) A genetic linkage map of coffee (Coffea arabica L.) and QTL for yield, plant height, and bean size. Tree Genet Genomes 12:5. https://doi.org/10.1007/s11295-015-0927-1
Noirot M, Poncet V, Barre P et al (2003) Genome size variations in diploid African Coffea species. Ann Bot 92:709–714. https://doi.org/10.1093/aob/mcg183
Nunes RC, Orozco-Arias S, Crouzillat D et al (2018) Structure and distribution of centromeric retrotransposons at diploid and allotetraploid Coffea centromeric and pericentromeric regions. Front Plant Sci 9:175. https://doi.org/10.3389/fpls.2018.00175
Orozco-Castillo C, Chalmers KJ, PoweH W, Waugh R (1996) RAPD and organelle specific PCR re-affirms taxonomic relationships within the genus Coffea. Plant Cell Rep 15:337–341. https://doi.org/10.1007/BF00232367
Ortega-Ortega J, Ramírez-Ortega FA, Ruiz-Medrano R, Xoconostle-Cázares B (2019) Analysis of genome size of sixteen Coffea arabica cultivars using flow cytometry. HortScience 54:998–1004. https://doi.org/10.21273/HORTSCI13916-19
Owen RB, Muiruri VM, Lowenstein TK et al (2018) Progressive aridification in East Africa over the last half million years and implications for human evolution. Proc Natl Acad Sci 115:11174–11179. https://doi.org/10.1073/pnas.1801357115
Pflug JM, Holmes VR, Burrus C, et al (2020) Measuring genome sizes using read-depth, k-mers, and flow cytometry: methodological comparisons in beetles (Coleoptera). G3 10:3047–3060. https://doi.org/10.1534/g3.120.401028
Pierozzi NI (2013) Orcein, C- and NOR-banding on mitotic chromosomes of some wild diploid coffee species (Coffea L.). Caryologia 66:76–83. https://doi.org/10.1080/00087114.2013.787206
Pierozzi NI, Pinto-Maglio CAF, Cruz ND (1999) Characterization of somatic chromosomes of two diploid species of Coffea L. with acetic orcein and C-band techniques. Caryologia 52:1–8. https://doi.org/10.1080/00087114.1998.10589147
Pierozzi NI, Borghi TC, Silvarolla MB (2012) A karyologicals study in some species of Coffea L. and in the closest relative Psilanthus travancorensis (Wight & Arn.) J.-F Leroy. Not Bot Horti Agrobot Cluj-Napoca 40:39–45. https://doi.org/10.15835/nbha4028195
Pinto-Maglio C, Cruz N (1987) Pachytene chromosome morphology in Coffea L. I. Nucleolar chromosomes. Caryologia 40:7–23. https://doi.org/10.1080/00087114.1987.10797806
Pinto-Maglio C, Cruz N (1998) Pachytene chromosome morphology in Coffea L II c Arabica l Complement. Caryologia 51:19–35. https://doi.org/10.1080/00087114.1998.10589117
Pinto-Maglio C, Cuéllar T, Barbosa R (2000) Aplicação de técnicas de citogenética molecular na caracterização dos cromossomos da espécie Coffea arabica L. In: Anais do I Simpósio da Pesquisa dos Cafés do Brasil. Poço das Caldas, Brazil, pp 444–446
Pinto-Maglio C, Barbosa R, Cuéllar T, et al (2001) Chromosome characterization in Coffea arabica L. using cytomolecular techniques. In: 14th International Chromosome Conference. Würzburg, Germany, p 100
Praça-Fontes MM, Carvalho CR, Clarindo WR, Cruz CD (2011) Revisiting the DNA C-values of the genome size-standards used in plant flow cytometry to choose the “best primary standards.” Plant Cell Rep 30:1183–1191. https://doi.org/10.1007/s00299-011-1026-x
Prakash N, Marques D, Varzea V et al (2004) Introgression molecular analysis of a leaf rust resistance gene from Coffea liberica into C. arabica L. Theor Appl Genet 109:1311–1317. https://doi.org/10.1007/s00122-004-1748-z
Raharimalala N, Rombauts S, McCarthy A et al (2021) The absence of the caffeine synthase gene is involved in the naturally decaffeinated status of Coffea humblotiana, a wild species from Comoro archipelago. Sci Rep 11:8119. https://doi.org/10.1038/s41598-021-87419-0
Raina SN, Mukai Y, Yamamoto M (1998) In situ hybridization identifies the diploid progenitor species of Coffea arabica (Rubiaceae). Theor Appl Genet 97:1204–1209. https://doi.org/10.1007/s001220051011
Razafinarivo NJ, Rakotomalala J-J, Brown SC et al (2012) Geographical gradients in the genome size variation of wild coffee trees (Coffea) native to Africa and Indian Ocean islands. Tree Genet Genomes 8:1345–1358. https://doi.org/10.1007/s11295-012-0520-9
Rijo L (1974) Observações cariológicas no cafeeiro “Híbrido de Timor.” Port Acta Biol 8:157–168
Ruas PM, Ruas CF, Rampim L et al (2003) Genetic relationship in Coffea species and parentage determination of interspecific hybrids using ISSR (Inter-Simple Sequence Repeat) markers. Genet Mol Biol 26:319–327. https://doi.org/10.1590/S1415-47572003000300017
Sanglard NA, Amaral-Silva PM, Sattler MC et al (2019) Indirect somatic embryogenesis in Coffea with different ploidy levels: a revisiting and updating study. Plant Cell Tissue Organ Cult PCTOC 136:255–267. https://doi.org/10.1007/s11240-018-1511-9
Scalabrin S, Toniutti L, Di Gaspero G et al (2020) A single polyploidization event at the origin of the tetraploid genome of Coffea arabica is responsible for the extremely low genetic variation in wild and cultivated germplasm. Sci Rep 10:1–13. https://doi.org/10.1038/s41598-020-61216-7
Schatz MC, Witkowski J, McCombie WR (2012) Current challenges in de novo plant genome sequencing and assembly. Genome Biol 13:1–7. https://doi.org/10.1186/gb-2012-13-4-243
Setotaw TA, Caixeta ET, Pereira AA et al (2013) Coefficient of parentage in Coffea arabica L. cultivars grown in Brazil. Crop Sci 53:1237–1247. https://doi.org/10.2135/cropsci2012.09.0541
Setotaw TA, Caixeta ET, Zambolim EM et al (2020) Genome introgression of Híbrido de Timor and its potential to develop high cup quality C. arabica cultivars. J Agric Sci 12:64–76. https://doi.org/10.5539/jas.v12n4p64
Singh R (2016) Plant cytogenetics. CRC Press
Soltis DE, Soltis PS, Pires JC et al (2004) Recent and recurrent polyploidy in Tragopogon (Asteraceae): cytogenetic, genomic and genetic comparisons. Biol J Linn Soc 82:485–501. https://doi.org/10.1111/j.1095-8312.2004.00335.x
Sousa TV, Caixeta ET, Alkimim ER et al (2017) Population structure and genetic diversity of coffee progenies derived from Catuaí and Híbrido de Timor revealed by genome-wide SNP marker. Tree Genet Genomes 13:1–16. https://doi.org/10.1007/s11295-017-1208-y
Stebbins GL (1949) Variation and evolution in plants. Columbia University Press, New York
Sybenga J (1960) Genetics and cytology of coffee: a literature review. Bibliogr Genet 10:83–137
Talukdar D, Sinjushin A (2015) Cytogenomics and mutagenomics in plant functional biology and breeding. PlantOmics: the omics of plant science. Springer, New Delhi, pp 113–156
Tosh J, Dessein S, Buerki S et al (2013) Evolutionary history of the Afro-Madagascan Ixora species (Rubiaceae): species diversification and distribution of key morphological traits inferred from dated molecular phylogenetic trees. Ann Bot 112:1723–1742. https://doi.org/10.1093/aob/mct222
Treangen TJ, Salzberg SL (2012) Repetitive DNA and next-generation sequencing: computational challenges and solutions. Nat Rev Genet 13:36–46. https://doi.org/10.1038/nrg3117
Urbanska KM, Hurka H, Landolt E et al (1997) Hybridization and evolution in Cardamine (Brassicaceae) at Urnerboden, Central Switzerland: biosystematic and molecular evidence. Plant Syst Evol 204:233–256. https://doi.org/10.1007/BF00989208
Vallejo-Marín M (2012) Mimulus peregrinus (Phrymaceae): a new British allopolyploid species. PhytoKeys 14:1–14. https://doi.org/10.3897/phytokeys.14.3305
van Boxtel J, Berthouly M (1996) High frequency somatic embryogenesis from coffee leaves: factors influencing embryogenesis, and subsequent proliferation and regeneration in liquid medium. Plant Cell Tissue Organ Cult 44:7–17. https://doi.org/10.1007/BF00045907
van der Vossen HA (1985) Coffee selection and breeding. Coffee. Springer, Boston, pp 48–96
Vicient CM, Casacuberta JM (2017) Impact of transposable elements on polyploid plant genomes. Ann Bot 120:195–207. https://doi.org/10.1093/aob/mcx078
von Faber FC (1912) Morphologisch-physiologische untersuchungen an blüten von Coffea-Arten. Ann Jard Buitenzorg 25:59–160
Wicker T, Gundlach H, Spannagl M et al (2018) Impact of transposable elements on genome structure and evolution in bread wheat. Genome Biol 19:103. https://doi.org/10.1186/s13059-018-1479-0
Wu F, Mueller LA, Crouzillat D et al (2006) Combining bioinformatics and phylogenetics to identify large sets of single-copy orthologous genes (COSII) for comparative, evolutionary and systematic studies: a test case in the Euasterid plant clade. Genetics 174:1407–1420. https://doi.org/10.1534/genetics.106.062455
Yotoko KSC, Dornelas MC, Togni PD et al (2011) Does variation in genome sizes reflect adaptive or neutral processes? New Clues from Passiflora. PLoS ONE 6:e18212. https://doi.org/10.1371/journal.pone.0018212
Yu Q, Guyot R, de Kochko A et al (2011) Micro-collinearity and genome evolution in the vicinity of an ethylene receptor gene of cultivated diploid and allotetraploid coffee species (Coffea): recent speciation event of Coffea arabica. Plant J 67:305–317. https://doi.org/10.1111/j.1365-313X.2011.04590.x
Yuyama PM, Pereira LFP, dos Santos TB et al (2012) FISH using a gag -like fragment probe reveals a common Ty 3- gypsy -like retrotransposon in genome of Coffea species. Genome 55:825–833. https://doi.org/10.1139/gen-2012-0081
Acknowledgements
We would like to thank the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq, Brasília—DF, Brazil), Fundação de Amparo à Pesquisa do Espírito Santo (FAPES, Vitória-ES, Brazil; grants: 65942604/2014 and 82/2017), and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES, Brasília—DF, Brazil) for financial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Communicated by Gerhard Leubner.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sattler, M.C., de Oliveira, S.C., Mendonça, M.A.C. et al. Coffea cytogenetics: from the first karyotypes to the meeting with genomics. Planta 255, 112 (2022). https://doi.org/10.1007/s00425-022-03898-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-022-03898-z