Abstract
Main conclusion
The present study is the first to integrate physiological and proteomic data providing information on Fe, Mn and Zn deficiency-responsive mechanisms of potato plants in vitro.
Abstract
Micronutrient deficiency is an important limiting factor for potato production that causes substantial tuber yield and quality losses. To under the underlying molecular mechanisms of potato in response to Fe, Mn and Zn deficiency, a comparative proteomic approach was applied. Leaf proteome change of in vitro-propagated potato plantlets subjected to a range of Fe-deficiency treatments (20, 10 and 0 μM Na-Fe-EDTA), Mn-deficiency treatments (1 and 0 μM MnCl2·4H2O) and Zn-deficiency treatment (0 μM ZnCl2) using two-dimensional gel electrophoresis was analyzed. Quantitative image analysis showed a total of 146, 55 and 42 protein spots under Fe, Mn and Zn deficiency with their abundance significantly altered (P < 0.05) more than twofold, respectively. By MALDI-TOF/TOF MS analyses, the differentially abundant proteins were found mainly involved in bioenergy and metabolism, photosynthesis, defence, redox homeostasis and protein biosynthesis/degradation under the metal deficiencies. Signaling, transport, cellular structure and transcription-related proteins were also identified. The hierarchical clustering results revealed that these proteins were involved in a dynamic network in response to Fe, Mn and Zn deficiency. All these metal deficiencies caused cellular metabolic remodeling to improve metal acquisition and distribution in potato plants. The reduced photosynthetic efficiency occurred under each metal deficiency, yet Fe-deficient plants showed a more severe damage of photosynthesis. More defence mechanisms were induced by Fe deficiency than Mn and Zn deficiency, and the antioxidant systems showed different responses to each metal deficiency. Reprogramming of protein biosynthesis/degradation and assembly was more strongly required for acclimation to Fe deficiency. The signaling cascades involving auxin and NDPKs might also play roles in micronutrient stress signaling and pinpoint interesting candidates for future studies. Our results first provide an insight into the complex functional and regulatory networks in potato plants under Fe, Mn and Zn deficiency.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Micronutrients such as iron (Fe), manganese (Mn) and zinc (Zn) are essential for plant growth and development, which play crucial roles in various cellular functions (Pilon et al. 2009; Hänsch and Mendel 2009). Fe, Mn and Zn are required as a cofactor or activator for many enzymes directly involved in photosynthesis, respiration, redox reaction in chloroplast and mitochondrion, nitrogen fixation, DNA synthesis, transcription, and other biosynthetic and degradative reactions (Broadley et al. 2007; Andreini et al. 2008; Nouet et al. 2011). On the other hand, these three metals are of particular importance to photosynthesis in plants. Fe is an important component of photosynthetic apparatus (Balk and Schaedler 2014). It is found in all photosynthetic electron-transfer complexes including cytochrome (Cyt) b6f complex and ferredoxins (Briat et al. 2015). Some Fe-cofactor containing proteins also participate in chlorophyll biosynthesis (van Leeuwe and Stefels 2007). Mn is pivotal for photosystem II (PSII) as a component of oxygen-evolving complex, which catalyzes the water splitting reaction of PSII to produce oxygen and provides electrons for photosynthetic electron transport chain (Millaleo et al. 2010). Zn is a cofactor of carbonic anhydrases that catalyze the hydration of CO2 and promote carbon fixation in photosynthesis (Broadley et al. 2007).
Micronutrient deficiency often occurs during plant growth and development. It leads to a variety of morphological, physiological and biochemical alterations in plants, and causes metabolic remodeling for improving metals acquisition and distribution. Chlorosis is one of the most important symptoms of Fe deficiency (Abadía et al. 2011). It is attributed to the inhibition of chlorophyll biosynthesis, which requires the function of Fe-containing enzymes (van Leeuwe and Stefels 2007). Fe deficiency also causes alterations in thylakoid membrane structure and photosynthetic electron-transfer chain (Sharma 2007; Laganowsky et al. 2009), including the decline of light-harvesting complex (LHC) and Cyt b6f complex, which lead to the reduced photosynthetic efficiency. Previous study has revealed that the decreased LHCI subunits and PSI core subunits (PsaC and PsaD) can be as indicators of Fe-deficiency response in rice (Yadavalli et al. 2012). To cope with Fe deficiency, plants have devised various strategies for acquiring Fe. Two main strategies are known: Strategy I based on a reduction uptake system (dicots and non-graminaceous monocots) and Strategy II based on a chelation uptake system (graminaceous monocots) (Curie and Briat 2003; Abadía et al. 2011). It is known that potato belongs to Strategy I plant, which can induce Fe(III)-chelates reductase and Fe(II) transporter located at the plasma membrane of epidermal root cells, and release some low-molecular-weight compounds including carboxylates, flavins and phenolic compounds in response to Fe deficiency (Ivanov et al. 2012; Legay et al. 2012). Several phytohormones including auxin, jasmonic acid and cytokinins have also been reported to transmit Fe-deficiency signals for adaption change of root morphology (Bacaicoa et al. 2009; Maurer et al. 2011).
Mn is required for biosynthesis of carbohydrate, lipid and lignin in plants, and its deficiency can inhibit plant growth and decrease biomass (Marschner 2012). Mn deficiency often leads to inter-venal chlorosis of younger leaves, and it is most likely because the Mn complex is required for stabilizing D1 protein in PSII (Allen et al. 2007; Schmidt et al. 2016). The tissue necrosis of Mn-deficient plants is also common due to a decrease in MnSOD levels and an increase in oxygen free radicals (Allen et al. 2007). Most of the Mn transporters have been identified in plants, such as NRAMP (natural resistance-associated macrophage protein), ZIP (ZRT/IRT-like protein), YSL (yellow stripe-like), CAX (cation exchanger), CCX (calcium cation exchanger), CDF/MTP (cation diffusion facilitator/metal tolerance protein), P-type ATPases and VIT (vacuolar iron transporter) (Morris et al. 2008; Mills et al. 2008; Cailliatte et al. 2010; Milner et al. 2013).
Zn deficiency also has a negative effect on plants, which results in short internodes, a decrease in leaf size and delayed maturity (Assunção et al. 2010a). It can cause a decrease in Cu/Zn-SOD activity and accumulation of reactive oxygen species (ROS), resulting in oxidative damage of cell membrane (Cakmak 2000). The carbonic anhydrase activity is reduced under Zn deficiency, which inhibits the carbon fixation in photosynthesis (Cakmak et al. 2010). Several members of ZIP family transporters are induced in roots of Arabidopsis under Zn deficiency (Wintz et al. 2003). Two highly homologous transcription factors, AtbZIP19 and AtbZIP23, have been identified to regulate Zn-deficiency response in Arabidopsis (Assunção et al. 2010b).
Potato (Solanum tuberosum L.) is the fourth most important food crop worldwide and a model for vegetatively propagated storage organs. Micronutrients such as Fe, Mn and Zn play vital role in the primary growth and development of potato plants, and lateral tuber yield and quality formation (Vreugdenhil 2007). These metals deficiency can cause substantial tuber yield losses and decrease the nutritional quality of tubers (Chatterjee et al. 2006; Legay et al. 2012). Despite an advanced knowledge of Fe, Mn and Zn deficiency-responsive mechanisms in model plants, it remains largely unknown in potato. In addition, based on the potato genome sequencing data, a large number of functional genome researches would be carried out (PGSC 2011). It needs to establish the stable technology system for mutant phenotype analysis, genetic transformation and material acquisition. In vitro techniques can offer an optimal control of the experimental growth conditions, which is suitable for researching the Fe, Mn and Zn deficiency-responsive mechanisms in potato. Furthermore, recent reports have been published focusing on proteome changes in response to micronutrient deficiency in plants (Rellán-Álvarez et al. 2010; Lan et al. 2011; Hsieh et al. 2013). Our observations attempt to extend findings regarding the potential proteomic dynamics of in vitro-propagated plantlets of potato under Fe, Mn and Zn deficiency and to enrich the research content of Fe, Mn and Zn deficiency-responsive mechanisms in potato.
In the present study, a comparative proteomic approach was applied to investigate the molecular events of in vitro-propagated plantlets of potato in response to Fe, Mn and Zn deficiency, respectively. The differentially abundant proteins including well-known and novel Fe, Mn and Zn deficiency-responsive proteins were identified using two-dimensional gel electrophoresis (2-DE) coupled with MALDI-TOF/TOF MS and Mascot database searching. The findings will help drive further work to develop strategies for reducing Fe, Mn and Zn deficiency effects on potato, and to gain comprehensive knowledge of the underlying molecular mechanisms of potato in response to these metals deficiency.
Materials and methods
Plant materials, trace element treatments and sampling
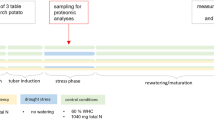
In vitro-propagated plantlets of potato (S. tuberosum L. cv. Atlantic) as the experimental materials (supplied by Gansu Key Laboratory of Crop Improvement and Germplasm Enhancement, Lanzhou, China) were propagated from single-node sections in MS medium (Murashige and Skoog 1962) containing 3% (w/v) sucrose and 0.8% (w/v) agar (pH 5.8). The MS medium contained the following macronutrients: 1.5 mM MgSO4·7H2O, 1.25 mM KH2PO4, 20.6 mM NH4NO3, 18.8 mM KNO3, 3 mM CaCl2·2H2O, 110 nM CoCl2·2H2O, 100 nM CuCl2·2H2O, 100 μM H3BO3, 5 μM KI, 100 μM MnCl2·4H2O, 1.03 μM Na2MoO4·2H2O, 30 μM ZnCl2 and 124 μM Na-Fe-EDTA. The single-node explants obtained from propagated plantlets were firstly transferred into liquid MS medium containing 3% (w/v) sucrose (pH 5.8) without Fe, Mn and Zn, respectively. The explants were inserted on the paper bridges in each culture bottles, and the liquid MS medium was under the paper bridges. Only the part below stem node of every explant was immersed in the liquid MS medium. They were cultured for 2 weeks at 25 ± 2 °C under 16-h photoperiod (100 μmol m−2 s−1 light intensity). For Fe-deficiency studies, the control (124 μM Na-Fe-EDTA) and Fe-deficiency treatments (20, 10 and 0 μM Na-Fe-EDTA) were prepared with liquid MS medium containing 3% (w/v) sucrose (pH 5.8). For Mn-deficiency studies, the control (100 μM MnCl2·4H2O) and Mn-deficiency treatments (1 and 0 μM MnCl2·4H2O) were prepared with liquid MS medium containing 3% (w/v) sucrose (pH 5.8). For Zn-deficiency studies, the control (30 μM ZnCl2) and Zn-deficiency treatment (0 μM ZnCl2) were prepared with liquid MS medium containing 3% (w/v) sucrose (pH 5.8). The detailed MS medium formulation for Fe, Mn and Zn deficiency is shown in Supplemental Table S1. Then, the cultures were transferred into these trace element treatment medium, respectively, and cultured at 25 ± 2 °C under 16-h photoperiod (100 μmol m−2 s−1 light intensity). There were about 300 explants used for each treatment (60 bottles for each treatment, and every five explants per bottle), and each treatment was replicated three times. After a 3-week culture, the fresh leaves were used for physiological analysis, and the leaves immediately frozen in liquid nitrogen were stored at − 80 °C for proteomic analysis.
Determination of chlorophyll content
Approximately, 1 g of fresh leaves was extracted in 10 mL of 80% chilled acetone. After centrifugation at 3000g for 2 min at 4 °C, the supernatant was used for the determination of chlorophyll content. The absorbance of supernatant was recorded at 663 nm and 645 nm, respectively. Chlorophyll content was calculated as described by Bhushan et al. (2007) and expressed as mg g FW−1.
Determination of sucrose, glucose and fructose content
Sugars were extracted as described by Matsuura-Endo et al. (2004) with some modifications. Approximately, 2 g of fresh leaves was homogenized in liquid nitrogen and incubated with 80% (v/v) ethanol at 70 °C for 3 h. The homogenate was centrifuged at 5000g for 20 min, and the supernatant was collected. The supernatant was then vacuum-dried, dissolved in distilled water, and passed though membrane filters (0.2-μm, Millipore, Bedford, MA, USA). The concentrations of sucrose, glucose and fructose in the filtrate were determined using HPLC (Agilent 1100 series, USA) with a ZORBAX Carbohydrate column (4.6 × 150 mm, USA).
Protein extraction
Total leaf proteins were extracted as described by Donnelly et al. (2005) with some modifications. Approximately, 2 g of leaves was homogenized in liquid nitrogen and precipitated overnight at − 20 °C by the addition of 25 mL of 10% (v/v) chilled trichloroacetic acid/acetone containing 0.07% (v/v) β-mercaptoethanol. After centrifugation at 20,000g for 20 min at 4 °C, the supernatant was removed. The pellet was re-suspended with 25 mL of chilled acetone containing 0.07% (v/v) β-mercaptoethanol and incubated at − 20 °C for 20 min. After centrifugation at 20,000g for 15 min at 4 °C, the pellet was collected. The steps were repeated until the pellet became pure white. The washed pellet was air-dried for 1 h and then solubilised in 300 μL of rehydration buffer [8 M urea, 2% (v/v) Triton X-100, 1% (w/v) DTT] for 2 h at room temperature. After centrifugation at 20,000g for 15 min at 4 °C, the supernatant was collected and stored at − 80 °C. The protein extraction was repeated for both control and treatments in triplicates, and the protein concentration was measured using Bio-Rad Protein Assay Reagent Kit (Bio-Rad, Hercules, CA, USA).
2-DE
The first dimension of the isoelectric focusing (IEF) was performed using 17 cm IPG strips (Bio-Rad) with pH 3–10 in PROTEAN IEF Cell System (Bio-Rad). The IPG strips were rehydrated overnight with 950 μg of total proteins diluted in rehydration buffer [7 M urea, 2 M thiourea, 2% (w/v) CHAPS, 0.3% (w/v) DTT, 0.5% (v/v) IPG buffer (pH 3–10) and 0.002% (w/v) bromophenol blue] to reach a final volume of 350 μL. After rehydration, the focusing was performed at 20 °C using the following settings: 50 V during 12 h, gradient to 250 V during 1 h, gradient to 1000 V in 1.5 h, gradient to 9000 V in 5 h, 9000 V until 62,000 Vh, 500 V during 24 h. Prior to second dimension electrophoresis, the IPG strips were equilibrated at room temperature for 15 min in 5 mL of equilibration buffer [6 M urea, 2% (w/v) SDS, 20% (v/v) glycerol, 0.375 M Tris-HCl (pH 8.8) and 0.2% (w/v) DTT], and subsequently for 15 min in the same buffer but 2.5% (w/v) iodoacetamine replacing DTT. The equilibrated strips were loaded and run on 12% SDS-PAGE gels using PROTEANII xi Cell System (Bio-Rad). The separated proteins were visualized by coomassie brilliant blue G-250 staining. For each protein sample, three replicates were run for each gel to ascertain reproducibility.
Image acquisition and data analysis
The CBB-stained 2-DE gels were scanned with a UMAX PowerLook 2100XL-USB scanner (Maxium Tech Inc., Taiwan, China) at a resolution of 300 dpi, and subsequently analyzed using PDQuest v8.0.1 (Bio-Rad). The gel image of control was selected as reference gel to align with gel image of Fe-deficiency treatments (20, 10 and 0 μΜ), Mn-deficiency treatments (1 and 0 μΜ) and Zn-deficiency treatment (0 μΜ), respectively. The abundance of one protein spot was expressed as the volume of that spot which was defined as the sum of the intensities of all the pixels that make up that spot. To minimize possible errors due to differences in the amount of protein loaded and the staining intensity, the spot abundance was normalized as a percentage of the total spot volume (relative V%) from three replicate gels, which were used to quantify and compare the spots. Only spots with statistical significance (P < 0.05) and reproducible changes were considered, and the spots with an abundance ratio at least twofold in relative abundance were selected as differentially abundant proteins.
Protein identification by MALDI-TOF/TOF MS
Protein pots with significantly differential abundance were excised, destained and in-gel digested with trypsin and peptides extracted according to Katayama et al. (2001). Peptide MS and MS/MS were performed on an ABI 5800 MALDI-TOF/TOF Plus mass spectrometer (AB SCIEX, Framingham, MA, USA). The MS spectra were recorded in the positive reflector mode in a mass range from 800 to 3500 Da with a focus mass of 2000. For one main MS spectrum, 25 subspectra with 125 shots per subspectrum were accumulated using a random search pattern. MS was used as a CalMix5 standard to calibrate the instrument (ABI 5800 Calibration Mixture). For MS calibration, autolysis peaks of trypsin (m/z 842.5100 and 2211.1046) were used as internal calibrates, and up to 10 of the most intense ion signals were selected as precursors for MS/MS acquisition, excluding the trypsin autolysis peaks and matrix ion signals. In MS/MS positive-ion mode, for one main MS spectrum, 50 subspectra with 50 shots per subspectrum were accumulated using a random search pattern. Collision energy was 1 kV, collision gas was air, and default calibration was set using the Glu1-Fibrino-peptide B (m/z 1570.6696) spotted onto Cal 7 positions of the MALDI target. Both the MS and MS/MS data were integrated and processed by using the GPS Explorer V3.6 software (AB SCIEX) with default parameters. Peptides were identified by searching the peak list against the NCBInr-S. tuberosum database 20160526 (43,985 sequences; 19,331,394 residues) using MASCOT V2.3 search engine (Matrix Science, London, UK). The parameters for searching were: taxonomy, S. tuberosum; trypsin as the digestion enzyme; one missed cleavage site; fixed modifications of Carbamidomethyl (C); partial modifications of Acetyl (Protein N-term), Deamidated (NQ), Dioxidation (W), Oxidation (M); 100 ppm for precursor ion tolerance and 0.3 Da for fragment ion tolerance. The significance threshold (P < 0.05) was set using the Mascot algorithm.
Functional classification and hierarchical clustering analysis
The functional classification of the identified proteins was conducted according to the putative functions assigned to each of the candidates using Geneontology database (http://amigo.geneontology.org/cgi-bin/amigo/blast.cgi). A hierarchical clustering analysis was performed by using the MultiExperiment Viewer (MeV) software. The data were taken in terms of fold expression with respect to the control expression value. Then, the data sets were log-transformed to the base 2 to level the scale of expression and reduce the noise. Only the clusters with n > 6 were taken to investigate the co-expression patterns for functionally similar proteins.
Statistical analysis
Statistical analysis was carried out with three biological replicates for proteomic and physiological analyses. The repeated measurement was given as mean ± standard error (SE). The results of spot abundance and physiological data were statistically evaluated by one-way analyses of variance (ANOVA) and the Duncan’s multiple range test to determine the significant difference among group means. In all cases, significance was defined as P < 0.05.
Results
Morphological and physiological changes under Fe, Mn and Zn deficiency
The morphological and physiological responses of potato seedlings in vitro were investigated under each metal-deficiency condition. The Fe-deficient plants displayed growth dwarfed, chlorosis in young leaves, and root growth inhibited (Fig. 1a). With the decrease of Fe concentration in medium, the chlorophyll content of leaves was significantly reduced by 40%, 51% and 87% at 20, 10 and 0 μM Na-Fe-EDTA treatments compared to control (124 μM Na-Fe-EDTA), respectively (Fig. 1b). There were no visible morphological changes in Mn-deficient plants (Fig. 1c), but the chlorophyll content of leaves was significantly declined by 20% at 0 μM MnCl2·4H2O treatments compared to control (100 μM MnCl2·4H2O), respectively (Fig. 1d). The Zn-deficient plants showed growth dwarfed, smaller leaves, and root growth inhibited (Fig. 1e). The chlorophyll content of leaves was significantly decreased by 29% at 0 μM ZnCl2 treatment compared to control (30 μM ZnCl2) (Fig. 1f). The glucose, fructose and sucrose content of leaves were also affected by these metals deficiency. For Fe-deficient plants, the glucose and fructose content of leaves was significantly increased by 57% and 200% at 0 μM Na-Fe-EDTA treatment compared to control, respectively (Fig. 2a, b), whereas the sucrose content was significantly decreased with the decrease of Fe concentration in medium (Fig. 2c). For Mn-deficient plants, the glucose content of leaves was significantly increased by 14% at 0 μM MnCl2·4H2O treatment compared to control (Fig. 2d), and the fructose content showed no obvious change at 0 μM MnCl2·4H2O treatment (Fig. 2e). Compared with the control, the sucrose content of leaves was significantly declined by 23% and 12% at 1 and 0 μM MnCl2·4H2O treatments, respectively (Fig. 2f). For Zn-deficient plants, the glucose and fructose contents of leaves were significantly decreased by 19% and 59% at 0 μM Na-Fe-EDTA treatment compared to control, respectively (Fig. 2g, h), whereas the sucrose content was significantly increased by 60% compared to control (Fig. 2i).
The Fe, Mn and Zn deficiency-induced morphological responses in potato plants in vitro. The single-node explants obtained from in vitro-propagated plantlets of potato were transferred into Fe, Mn and Zn deficiency-treatment medium, respectively, and cultured for 3 weeks at 25 ± 2 °C under 16-h photoperiod (100 μmol m−2 s−1 light intensity). a, b The plant morphology and chlorophyll content of control (124 μM Na-Fe-EDTA) and Fe-deficiency treatments (20, 10 and 0 μM Na-Fe-EDTA). c, d The plant morphology and chlorophyll content of control (100 μM MnCl2·4H2O) and Mn-deficiency treatments (1 and 0 μM MnCl2·4H2O). e, f The plant morphology and chlorophyll content of control (30 μM ZnCl2) and Zn-deficiency treatment (0 μM ZnCl2). Each value is represented as mean ± SE for three independent experiments. Means followed by different small letters are significantly different at P < 0.05 according to Duncan’s multiple range test
The Fe, Mn and Zn deficiency-induced physiological responses in potato plants in vitro. The single-node explants obtained from in vitro-propagated plantlets of potato were transferred into Fe, Mn and Zn deficiency-treatment medium, respectively, and cultured for 3 weeks at 25 ± 2 °C under 16-h photoperiod (100 μmol m−2 s−1 light intensity). a–c The glucose, fructose and sucrose content of control (124 μM Na-Fe-EDTA) and Fe-deficiency treatments (20, 10 and 0 μM Na-Fe-EDTA). d, f The glucose, fructose and sucrose content of control (100 μM MnCl2·4H2O) and Mn-deficiency treatments (1 and 0 μM MnCl2·4H2O). g–i The glucose, fructose and sucrose content of control (30 μM ZnCl2) and Zn-deficiency treatment (0 μM ZnCl2). Each value is represented as mean ± SE for three independent experiments. Means followed by different small letters are significantly different at P < 0.05 according to Duncan’s multiple range test
Identification of Fe, Mn and Zn deficiency-responsive proteins by 2-DE and MS
Comparative proteomics analysis was used to investigate the changes of leaf protein profiles in potato seedlings in vitro under each deficiency condition. Total leaf proteins of control, Fe-deficiency treatments (20, 10 and 0 μM Na-Fe-EDTA), Mn-deficiency treatments (1 and 0 μM MnCl2·4H2O) and Zn-deficiency treatment (0 μM MnCl2·4H2O) were extracted and separated by 2-DE, and three replicate gels for control and each treatment were obtained (Supplemental Figs. S1–S3). Figure 3 showed the representative standard gel maps of each deficiency condition, respectively. More than 1100 protein spots were reproducibly detected and matched between all the gels. Quantitative image analyses showed a total of 146 protein spots from Fe-deficiency treatments, 55 protein spots from Mn-deficiency treatments and 42 protein spots from Zn-deficiency treatment with their abundance significantly altered (P < 0.05) by more than at least two-fold, respectively. Interestingly, the differentially abundant proteins in Fe-deficiency treatments were obviously more than Mn- and Zn-deficiency treatments, which indicated that the potato seedlings in vitro might be more sensitive to Fe deficiency. The expression profile of these proteins indicated that most of the proteins showed quantitative changes, and some proteins also showed qualitative changes (Fig. 3; Supplemental Tables S2–S4). Of the 146 differentially abundant proteins under Fe deficiency, 60 proteins were up-regulated, 29 proteins were down-regulated, one protein was disappeared, 30 proteins were induced at least one concentration under Fe deficiency, and the rest showed a mixed expression pattern. Of the 55 differentially abundant proteins under Mn deficiency, 28 proteins were up-regulated, 11 proteins were down-regulated, 10 or 6 proteins were induced or disappeared at least under one Mn-deficiency treatment. Of the 42 differentially abundant proteins under Zn deficiency, 31 proteins were up-regulated, 8 proteins were down-regulated, and 3 proteins were disappeared.
2-DE gel analysis of the leaf proteome of potato plants in vitro under Fe, Mn and Zn deficiency. Equal amounts (950 μg) of proteins were separated on pH 3–10 IPG strips (17 cm, linear) in the first dimension and by SDS-PAGE on 12% polyacrylamide gels in the second dimension. The gels were visualized by CBB staining. Three replicate CBB-stained gels for control and Fe-, Mn- and Zn-deficiency treatments were computationally combined using PDQuest v8.0.1 software, respectively. a 2-DE protein profile for control (124 μM Na-Fe-EDTA) and Fe-deficiency treatments (20, 10 and 0 μM Na-Fe-EDTA). b 2-DE protein profile for control (100 μM MnCl2·4H2O) and Mn-deficiency treatments (1 and 0 μM MnCl2·4H2O). c 2-DE protein profile for control (30 μM ZnCl2) and Zn-deficiency treatment (0 μM ZnCl2)
All the differentially abundant proteins were successfully identified by MALDI-TOF/TOF MS. The primary identification information of these differentially abundant proteins under Fe, Mn and Zn deficiency were presented in Supplemental Tables S5–S7, which are summarized in Supplemental Tables S2–S4, respectively. To generate a board survey of these identified proteins with altered abundance, a Venn diagram was conducted to show the dynamics of the number of differentially abundant proteins under Fe, Mn and Zn deficiency. Among these identified proteins, 17 proteins were identified under all deficiency condition (Fig. 4a), including 23-kDa oxygen-evolving protein of PSII (spot 2111), ATP synthase CF1 beta chain (spot 3703), photosynthetic NDH subunit of lumenal location 5 (spot 7003), ribulose-1,5-bisphosphate carboxylase/oxygenase large subunit (RuBisco large subunit, spot 8705), oxygen-evolving enhancer protein 1 (OEE 1, spot 7104), plastidic aldolase (spot 3511), enolase-like isoform X1 (spot 3710), glyceraldehyde 3-phosphate dehydrogenase (spot 5507), mitochondrial NAD-dependent malate dehydrogenase (spot 7410), dehydroascorbate reductase (DHAR, spots 4214 and 5106), chitinase (spots 6407, 7302 and 8315), peroxidase 12 (spot 8602), catalase (CAT, spot 9705) and elongation factor 1-alpha (spot 9602). However, most of these proteins showed differential responses to Fe, Mn and Zn deficiency. Only two proteins (enolase-like isoform X1 and DHAR) were up-regulated and two proteins (RuBisco large subunit and CAT) were down-regulated under all deficiency condition (Fig. 4b, c). Thirty-nine proteins were up-regulated only under Fe deficiency, 8 proteins were up-regulated only under Mn deficiency, and 12 proteins were up-regulated only under Zn deficiency (Fig. 4b). Meanwhile, 23 proteins were down-regulated only under Fe deficiency, 6 proteins were down-regulated only under Mn deficiency, and 3 proteins were down-regulated only under Zn deficiency (Fig. 4c). Otherwise, it was noted that the same protein migrated to different gel spots, and their function was common to different spots. Under Fe deficiency, 32 proteins were identified in two, three or five spots. For example, ferredoxin-NADP reductase was identified in two spots (spots 4402 and 4405), OEE 1 was identified in three spots (spots 1307, 1314 and 7104), and RuBisco large subunit was identified in five spots (spots 6306, 6713, 6722, 8401 and 8705) (Supplemental Table S2). Under Mn deficiency, two proteins were identified in two spots, that is, RuBisco large subunit (spots 7204 and 8705) and 23-kDa oxygen-evolving protein of PSII (spots 0205 and 2111) (Supplemental Table S3). Under Zn deficiency, three proteins were identified in two or three spots, that is, RuBisco large subunit (spots 6306 and 8705), glyceraldehyde 3-phosphate dehydrogenase (spots 5507, 6502 and 6514) and superoxide dismutase [Fe] (FeSOD, spots 3101 and 4102) (Supplemental Table S4).
Venn diagrams of the number of total proteins (a), up-regulated proteins (b) and down-regulated proteins (c) in potato leaves in vitro under Fe, Mn and Zn deficiency. Overlapping regions of the circles indicate the number of proteins shared by the two or three metals-deficiency treatments, whereas non-overlapping circles indicated the number of proteins only found under each metal-deficiency condition
Functional classification of the Fe, Mn and Zn deficiency-responsive proteins
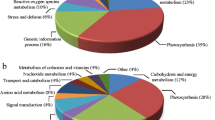
The identified proteins play a variety of functions during cellular adaptation to Fe, Mn and Zn deficiency, respectively. Under Fe deficiency, the 146 differentially abundant proteins were grouped into eleven functional classes (Fig. 5a; Supplemental Table S2). The largest percentages of these proteins were involved in defence (25%), bioenergy and metabolism (22%) and photosynthesis (19%), and the second classes corresponded functions were involved in protein biosynthesis/degradation (13%) and redox homeostasis (10%). Proteins were also found to play roles in transport (3%), miscellaneous (3%), signaling (2%), transcription (1%), cellular structure (1%) and unknown (1%). Under Mn deficiency, the 55 differentially abundant proteins were classified into nine functional categories (Fig. 5b; Supplemental Table S3), including photosynthesis, bioenergy and metabolism, defence, redox homeostasis, protein biosynthesis/degradation, transcription, transport, signaling, miscellaneous and unknown. The largest percentages of these proteins belong to photosynthesis (27%), bioenergy and metabolism (25%) and defence (20%). Redox homeostasis and protein biosynthesis/degradation-related proteins accounted 9% and 7%, respectively. Similarly, a wide range of cellular functions were also covered under Zn deficiency, which were grouped into nine functional classes (Fig. 5c; Supplemental Table S4). The major functional class corresponded proteins involved in bioenergy and metabolism (26%), photosynthesis (19%), redox homeostasis (19%), defence (15%) and protein biosynthesis/degradation (7%). Taken together, it appeared that the bioenergy and metabolism, photosynthesis, defence, redox homeostasis and protein biosynthesis/degradation-related proteins were significantly affected under each deficiency condition.
Functional classification of the differentially abundant proteins in potato leaves in vitro under Fe (a), Mn (b) and Zn (c) deficiency. The protein function classification was conducted according to the putative functions assigned to each of the candidate proteins using the protein functional database and displayed in the pie chart
Dynamics of the Fe, Mn and Zn deficiency-responsive protein networks
To summarize the identified proteins with similar expression profiles and achieve a comprehensive overview of the co-expression patterns for functionally similar proteins, a hierarchical clustering was applied to differentially abundant proteins identified under each deficiency condition. The clustering analysis yielded eight, six and two expression clusters under Fe, Mn and Zn deficiency (Figs. 6, 7, 8), respectively, and only the clusters with n > 6 were taken to study the co-expression patterns for functionally similar proteins. The detailed information on proteins within each cluster is presented in Supplemental Figs. S4–S6, respectively. Under Fe deficiency, the proteins involved in bioenergy and metabolism, defence, redox homeostasis and protein biosynthesis/degradation in Cluster 1 maintained almost steady state at 20 and 10 μM Na-Fe-EDTA treatments, and rapidly up-regulated at 0 μM Na-Fe-EDTA treatment (Fig. 6). Cluster 2 enriched in these proteins showed a gradual up-regulation with the decrease of Fe deficiency, and the co-clustering pattern was also found for unknown protein. However, the non-homogeneous expression patterns were also observed in proteins with these functions. The proteins involved in photosynthesis, protein biosynthesis/degradation, bioenergy and metabolism, redox homeostasis and signaling in Cluster 6 were observed to be down-regulated under Fe deficiency. The photosynthesis, protein biosynthesis/degradation, and transport-related proteins in Cluster 8 were only up-regulated at 20 μM Na-Fe-EDTA treatment, and then down-regulated. Under Mn deficiency, Cluster 3 as the most abundant group was found to maintain almost steady state at 1 μM MnCl2·4H2O treatment, and rapidly up-regulated at 0 μM MnCl2·4H2O treatment, which was enriched in proteins associated with bioenergy and metabolism, photosynthesis, redox homeostasis, defence, signaling and protein biosynthesis/degradation (Fig. 7). The proteins involved in defence, bioenergy and metabolism, redox homeostasis and transcription were down-regulated at 1 μM MnCl2·4H2O treatment and then up-regulated. Under Zn deficiency, the clustering analysis only yielded two expression clusters (Fig. 8). The most abundant group, Cluster 1 with 31 proteins, was found to be up-regulated, showing the maximum co-clustering for the proteins involved in bioenergy and metabolism and redox homeostasis. The proteins involved in photosynthesis and protein biosynthesis/degradation were found to be down-regulated. Due to heterogeneous composition, the miscellaneous category of proteins was represented in these two clusters and showed no clear clustering patterns.
Clustering analysis of the expression profiles of differentially abundant proteins in potato leaves in vitro under Fe deficiency. The hierarchical cluster tree is shown at the top, and the expression profiles are shown below. The four rows of hierarchical cluster tree represent control (124 μM Na-Fe-EDTA) and Fe-deficiency treatments (20, 10 and 0 μM Na-Fe-EDTA), respectively. Each individual protein is represented by a single column of color boxes. The up- and down-regulated proteins are indicated in red and green, respectively. The colors intensity is increased with the expression differences increasing, as shown in the bar. The expression profile of each individual protein in the cluster is depicted by gray lines, while the mean expression profile is marked in pink for each cluster. The number of proteins in each cluster is given in the left upper corner, and the cluster number is given in the right lower corner. Only the clusters with n > 6 were taken to investigate the co-expression patterns for functionally similar proteins. The detailed information on proteins within each cluster is presented in Supplemental Fig. S4
Clustering analysis of the expression profiles of differentially abundant proteins in potato leaves in vitro under Mn deficiency. The hierarchical cluster tree is shown at the top, and the expression profiles are shown below. The three rows of hierarchical cluster tree represent control (100 μM MnCl2·4H2O) and Mn-deficiency treatments (1 and 0 μM MnCl2·4H2O), respectively. Each individual protein is represented by a single column of color boxes. The up- and down-regulated proteins are indicated in red and green, respectively. The colors intensity is increased with the expression differences increasing, as shown in the bar. The expression profile of each individual protein in the cluster is depicted by gray lines, while the mean expression profile is marked in pink for each cluster. The number of proteins in each cluster is given in the left upper corner, and the cluster number is given in the right lower corner. Only the clusters with n > 6 were taken to investigate the co-expression patterns for functionally similar proteins. The detailed information on proteins within each cluster is presented in Supplemental Fig. S5
Clustering analysis of the expression profiles of differentially abundant proteins in potato leaves in vitro under Zn deficiency. The hierarchical cluster tree is shown at the top, and the expression profiles are shown below. The two rows of hierarchical cluster tree represent control (30 μM ZnCl2) and Zn-deficiency treatment (0 μM ZnCl2), respectively. Each individual protein is represented by a single column of color boxes. The up- and down-regulated proteins are indicated in red and green, respectively. The colors intensity is increased with the expression differences increasing, as shown in the bar. Due to only one treatment (0 μM ZnCl2) for Zn deficiency, there were only two expression clusters (39 up-regulated proteins and 8 down-regulated proteins), and no expression profile produced. Both the two clusters were taken to investigate the co-expression patterns for functionally similar proteins. The detailed information on proteins within each cluster is presented in Supplemental Fig. S6
Discussion
Micronutrient deficiency such as Fe, Mn and Zn is one of the important limiting factors for potato production that cause substantial tuber yield losses and decrease the nutritional quality of tubers (Chatterjee et al. 2006; Legay et al. 2012). Despite intensive studies on the mechanisms leading to Fe, Mn and Zn uptake, homeostasis and storage in model plants, little information is available regarding the strategies that improve their acquisition and cellular homeostasis in potato plants under conditions of low Fe, Mn and Zn availability. The application of proteomic approaches that allows the simultaneous and untargeted analysis of multiple proteins can provide insight into multiple cellular processes taking place under Fe, Mn and Zn deficiency. The present research describing the potential proteomic dynamics of in vitro-propagated plantlets of potato will help to gain comprehensive knowledge of the underlying molecular mechanisms of potato plants in response to Fe, Mn and Zn deficiency.
The leaf proteome of Fe-deficient plants showed more severe damage on photosynthesis than that of Mn- and Zn-deficient plants
It is well known that Fe, Mn and Zn are of particular importance to photosynthesis in plants (Broadley et al. 2007; Millaleo et al. 2010; Balk and Schaedler 2014). Each of these metals deficiency in the present study is linked to photosynthetic function. Some common responses were found, for example, the down-regulated or disappeared RuBisco large subunits (spots 6306, 6713, 6722, 8401 and 8705) suggested the reduced photosynthetic efficiency caused by Fe, Mn and Zn deficiency in potato plants. Also, some differential effects among these three metals deficiency were revealed by the proteomics data. On the one hand, Fe deficiency appeared to result in the inhibition of photosynthetic carbon flux. This is observed via the down-regulation of two aldolases including plastidic aldolase (spot 3511) and fructose-bisphosphate aldolase 1 (spot 2406), whereas they were up-regulated under Mn and Zn deficiency. Plastidic aldolase is a key enzyme involved in the carbon fixation of Calvin cycle which is an important metabolism pathway in photosynthesis (Patron et al. 2004). It has been reported that this enzyme participates in response to multiple abiotic stresses in plants (Yamada et al. 2000). The results suggested that the photosynthetic carbon partitioning might be inhibited by Fe deficiency, whereas Mn- and Zn-deficient plants might enhance NADPH synthesis for energy and maintenance of Calvin cycle. On the other hand, Fe deficiency strongly affected photosynthetic electron transport, but much less responses were showed in Mn- and Zn-deficient plants. Ferredoxin (Fd) is iron-sulfur protein that serves as electron carrier in the photosynthetic electron transport chain (Fukuyama 2004). Ferredoxin-NADP+ reductase (FNR) uses iron–sulfur proteins as electron donors to transfer electrons from PSI to NADPH (Aliverti et al. 2008). The down-regulation of Fd I (spot 0102) and FNR (spots 4402 and 4405) suggested that Fd and FNR might be the major targets of Fe deficiency as suggested by Hantzis et al. (2018). PSI-D subunit is a hydrophilic subunit of PSI, which has an important function in the docking of Fd to PSI (Sétif et al. 2002). The finding here that Fe deficiency caused the down-regulation of two putative PSI reaction center PSI-D subunit precursor (spots 9031 and 9032) was consistent with the observation in rice under Fe deficiency by Yadavalli et al. (2012). In contrast to Fe-deficient response, the PSI-D subunit was up-regulated under both Mn and Zn deficiency. The similar expression patterns of chloroplast NADH dehydrogenase complexes, mediating the Fd-dependent cyclic electron flow (Rumeau et al. 2007), were also found under these metals deficiency. OEEs as an auxiliary component of PSII manganese cluster can control O2 evolution and maintain the stability of PSII (Heide et al. 2004). Under both Fe and Mn deficiency, some OEEs were identified to be down-regulated or disappeared, including OEE 1 (spots 1307, 1314, 7104 and 9009) and 23 kDa OEE (spot 2106). It appeared that the oxygen-evolving activity of PSII might be decreased under Fe and Mn deficiency. Interestingly, there was a PSII repair protein PSB27-H1 (spot 8010) induced by Mn deficiency, which might have a certain function for repairing the damaged PSII. Additionally, the chloroplast manganese-stabilizing protein (spot 1312) was down-regulated in Fe-deficient plants, suggesting the inhibition of PSII core assembly/stability as suggested by Yi et al. (2005). All the results indicated that more severe and irreparable damage of photosynthesis might be occurred in potato plants under Fe deficiency, whereas Mn and Zn deficiency might trigger some positive responses to reduce damage.
Remodeling of cellular metabolic status was induced by all these metals deficiency
Micronutrient deficiency can cause cellular metabolic remodeling for improving metals acquisition and distribution in plants (López-Millán et al. 2013). Glycolysis and tricarboxylic acid (TCA) cycle is the most common metabolic pathway affected by micronutrient deficiency (Rellán-Álvarez et al. 2010; Khandakar et al. 2013). In the present study, most of the key enzymes involved in glycolysis and TCA cycle were identified to be up-regulated under Fe, Mn and Zn deficiency in potato plants, including glyceraldehyde 3-phosphate dehydrogenase, triose phosphate isomerase, enolase, phosphoglycerate kinase, and malate dehydrogenase. It reflected an increased energy requirement in response to these metals deficiency, and the enhancement of glycolysis and TCA cycle might be the conspicuous initial indicator of high energy production (Rellán-Álvarez et al. 2010; Khandakar et al. 2013). Additionally, several up-regulated ATP synthase subunits might further support the higher energy requirement, which could enhance ATP synthesis and provide more energy in response to Fe, Mn and Zn deficiency. The similar behavior was also described in Arabidopsis under abiotic stresses (Zhang et al. 2008). There was also a adenylate kinase 4 (spot 8326) induced by Fe deficiency, which could contribute to regulate multiple cellular energy dependent and nucleotide signaling processes through catalyzing phosphotransfer as suggested by Dzeja and Terzic (2009). Except for these common metabolic pathways, some specific changes were also found among these metals deficiency. The amino acid metabolism was affected by Fe and Mn deficiency. This is observed via the differential expression of key enzymes and proteins involved in amino acid biosynthesis, including S-adenosylmethionine synthetase, plastid glutamine synthetase, 5-methyltetrahydropteroyltriglutamate-homocysteine methyltransferase, cysteine synthase, glutamate dehydrogenase, leucine aminopeptidase and T-protein. There might have some shifts in the utilization of amino acids toward essential cellular function as suggested by Khandakar et al. (2013). Formate dehydrogenase (FDH) is a mitochondrial and NAD-dependent enzyme that catalyzes the oxidation of formate to CO2 in plants (Alekseeva et al. 2011). Three mitochondrial FDH precursors (spots 6509, 6112 and 7501) in Fe-deficient plants and one mitochondrial FDH precursor (spot 7501) in Zn-deficient plants were found to be up-regulated. With regard to the Fe deficiency-induced FDH, it can be interpreted that Fe deficiency might cause anoxia by the depletion of Fe from heme or the inhibition of heme biosynthesis; then the formate pathway would be induced either by the decrease of heme or by the reduced electron transport in the respiratory chain of mitochondria (Suzuki et al. 1998; López-Millán et al. 2000). Even through the similar change of FDH in Zn-deficient plants was not well known, it appeared to be a secondary effect caused by anoxia induced by Zn deficiency. Moreover, the up-regulation of perakine reductase (spot 3506), participating in the biosynthesis of monoterpenoid indole alkaloids (Sun et al. 2008), was found under Fe and Zn deficiency. It is suggested that the synthesis of secondary metabolite might be induced as an adaptation mechanism for Fe and Zn deficiency in potato plants.
Fe deficiency triggered more defense mechanisms than Mn and Zn deficiency, and the antioxidant systems showed different responses to these metals deficiency
To adapt these metals deficiency, various defense mechanisms were triggered in potato plants. With Fe deficiency, there were clear indications of a global defense against pathogens and pests. This is observed via the up-regulation or induction of pathogenesis-related protein (spots 1015, 2005 and 3002), chitinase (spots 8310, 8315, 6317, 7302, 4404, 6407, 7414 and 2203), beta-1,3-glucanase (spots 8414, 9303, 9409, 9417 and 9416), osmotin-like protein (spots 7211, 8206, 8103 and 8212), cyclophilin (spot 8112), patatin (spots 1207, 1305, 1515 and 2510) and proteinase inhibitor (spots 8013, 8014, 8123 and 8208). However, much less pathogenesis-related defense responses were found in Mn- and Zn-deficient plants, and there was no other defense response found in Zn-deficient plants. Alcohol dehydrogenase was up-regulated under Fe and Mn deficiency, which might avoid cells from being poisoned by acetic acid and promote the adaptability of potato plants (Strommer 2011). The up-regulation of lactoylglutathione lyase (spot 2403) also exhibited the detoxification of methylglyoxal in cells under Fe deficiency (Dixon et al. 1998). CBS domain-containing protein has been reported that its over-expression can improve salinity, oxidative and heavy metal tolerance in transgenic tobacco (Singh et al. 2012). The induction of this protein suggested its important role in response to Fe and Mn deficiency. Inhibitor II-chloramphenicol acetyltransferase (spot 7208), a wound-inducible protein (Keil et al. 1990), was also induced in potato plants under Fe deficiency. All these results indicated that more defense mechanisms were induced in Fe-deficient plants than in Mn- and Zn-deficient plants. On the other hand, these three metals deficiency appeared to activate different oxidative stress responses in potato plants, and various protective enzymes and antioxidants were induced to protect against oxidative damage. With Fe deficiency, the down-regulation of FeSOD (spots 3101 and 4102) and CAT (spots 7702 and 9705) might reflect the strong impact of Fe deficiency on redox homeostasis, not only because free Fe ions induce ROS formation via Fenton reactions, but also because many proteins involved in oxidative stress such as FeSOD and CAT are Fe-containing proteins (López-Millán et al. 2013). The down-regulation or induction of peroxidase 12 (spots 8602 and 8622) and 2-Cys peroxiredoxin BAS1 (spots 0208 and 0102) indicated the complex interplay between Fe and redox homeostasis mechanisms. The up-regulation of glutathione S-transferase L3-like (spots 1205 and 2202), probable l-ascorbate peroxidase 6 (spot 4304) and DHAR (spot 5106) suggested an induction of ascorbate–glutathione cycle in response to Fe deficiency (Zaharieva and Abadía 2003). For Mn- and Zn-deficient plants, the down-regulation of CAT also reflected the impact of Mn and Zn deficiency on redox homeostasis. However, compared with Mn deficiency, Zn deficiency induced more antioxidative systems to protect cells from oxidative damage, including the up-regulation of FeSOD (spots 3101 and 4102), peroxidase 12 (spot 8602), 2-Cys peroxiredoxin BAS1 (spot 0208) and DHAR (spots 4214 and 5106). All the results suggested that activation of different antioxidases was required for a detoxification mechanism in potato plants under Fe, Mn and Zn deficiency.
Reprogramming of protein biosynthesis, degradation and assembly was more strongly required for Fe-deficient plants than Mn- and Zn-deficient plants
A subset of elongation factors (spots 0307, 9602, 9710, 0305 and 5608), ribosomal proteins (spots 9208 and 0012), ribosome-recycling factor (spot 8216) and nascent polypeptide-associated complex subunit alpha-like protein (spot 0302) were up-regulated or induced under Fe deficiency, suggesting the enhancement of protein biosynthesis in potato plants. Elongation factor is a core translational protein that catalyzes the initiation and elongation of newly growing peptide chains (Singh et al. 2004). Ribosomal protein is one kind of highly conserved proteins that make up ribosomal subunits involved in the cellular process of translation (Rodnina and Wintermeyer 2011). Previous studies have reported that these proteins are highly dynamic and affected by environmental signals (Ma et al. 2010; Schippers and Mueller-Roeber 2010). It is, thus, speculated that differential expression of these proteins might control mRNA preference toward the translation of some genes that are crucial for the acclimation to Fe deficiency as suggested by Lan et al. (2011). Proteolysis is necessary for the removal of abnormal, modified and mistargeted proteins, and for altering the balance of proteins (Choudhary et al. 2009). Cysteine proteinase 3 (spot 1202) was induced, suggesting its role in maintaining strict protein quality control in response to Fe deficiency. The down-regulation of ATP-dependent Clp protease (spot 2809) indicated that ATP-dependent proteolysis in chloroplast might be inhibited when potato plants were subjected to Fe deficiency. Additionally, the induced 10-kDa chaperonin might protect potato plants against Fe deficiency by re-establishing normal protein conformation and cellular homeostasis (Wang et al. 2004). The up-regulation of peptidyl-prolyl cis–trans isomerase FKBP16-3 (spot 1115) and protein disulfide-isomerase-like (spot 0705) might also accelerate protein re-folding under Fe deficiency. For Mn- and Zn-deficient plants, only a few protein biosynthesis and degradation-related proteins were identified, such as three elongation factors (spots 5608, 9602 and 5926) and one 40S ribosomal protein (spot 9208) in Mn-deficient plants, and two elongation factors (spots 0307 and 9602) and one ATP-dependent Clp protease (spot 2809) in Zn-deficient plants. All the results suggested that reprogramming of protein biosynthesis/degradation and assembly might represent critical processes that contribute to the acclimation to Fe deficiency.
Several signal pathways and transport processes play roles in response to these metals deficiency
Nucleoside diphosphate kinases (NDPKs) are ubiquitous enzymes involved in equilibration of cellular nucleosides triphosphate pools (Hasunuma et al. 2003). Recent studies suggested that NDPKs also play roles in signal transduction pathways involved in oxidative stress (Haque et al. 2010; Kim et al. 2011). Under both Fe and Mn deficiency, the NDPKs (spots 8019 and 7106) were found to be induced or up-regulated. It is tempting to speculate that NDPKs might mediate Fe and Mn deficiency tolerance by signaling the transient expression of genes involved in antioxidant and protective processes, possibly through activation of the MAPK cascade (Yang et al. 2003). It is well known that the plant hormone auxin regulates many aspects of plant growth and development. Auxin signal is sensed and interpreted by distinct receptor systems that activate a broad range of cellular responses (Hagen 2015). Auxin-binding protein (ABP) functions as an auxin receptor in auxin signaling, which has an essential role in many developmental processes in plants (Paque et al. 2014). In the present study, the ABP19a-like (spot 6105) was down-regulated under Fe deficiency, whereas it was up-regulated under Zn deficiency. It suggested that ABP might be involved in the regulation of polar auxin transport in Fe- and Zn-deficient plants with a different manner, and thus affect local auxin concentration and early auxin gene regulation as suggested by Effendi et al. (2011). Indole-3-acetic acid-amido synthetase (GH3) is a kind of early auxin-responsive enzymes that widely exist in plants, which maintains auxin homeostasis by conjugating excess auxin to amino acids (Staswick et al. 2005). Previous studies have reported that GH3 modulated multiple developmental processes such as photomorphogenesis, light and auxin signaling, and auxin homeostasis in Arabidopsis (Tanaka et al. 2002; Staswick et al. 2005). The up-regulation of GH3.1 (spot 4513) was found in both Fe- and Mn-deficient plants, also suggesting an activation of auxin signaling pathway in response to Fe and Mn deficiency. In addition, some transport-related proteins were identified to be up-regulated or induced, indicating that there exists active transport of ions and metabolites for adaptation response to Fe, Mn and Zn deficiency. Mitochondrial outer membrane porin regulates metabolic and energetic flux across the outer mitochondrial membrane (De Pinto et al. 2010). The up-regulation or induction of 34-kDa outer mitochondrial membrane protein porin-like protein (spots 8306, 8308 and 7411) and 34-kDa porin (spot 9301) could enhance the exchange of ions and molecules between mitochondria and cytosol for maintaining intracellular homeostasis under Fe, Mn and Zn deficiency. Plant non-specific lipid-transfer proteins (nsLTPs) are responsible for the shuttling of phospholipids and other fatty acid groups between cell membranes (Carvalho and Gomes 2007). Recent evidences reported that nsLTPs modulate plant tolerance to multiple abiotic stresses and infection with bacterial and fungal pathogen (Sarowar et al. 2009; Gangadhar et al. 2016). The nsLTP 1-like (spots 9022 and 9002) were induced under both Fe and Mn deficiency, which suggested that nsLTPs might facilitate lipid transport between cell membranes in response to Fe and Mn deficiency.
Conclusions
This study first integrated physiological and proteomic data to provide useful information about the underlying molecular mechanisms of potato plants in response to Fe, Mn and Zn deficiency. Based on these proteomic data, the representative models for summarizing the functional and regulatory networks activated by these metals deficiency are illustrated in Fig. 9. Metal deficiency might be sensed inside the potato leaf cells via anterograde signals and/or organelle (chloroplast and mitochondrion) retrograde signals, and transduced for subsequent regulation of nuclear genes that are involved in various cellular processes in response to metal deficiency, via the activation of transcription factor cascades. The reduced photosynthetic efficiency occurred under each metal-deficiency condition, whereas Fe-deficient plants showed more severe and irreparable damage of photosynthesis including the inhibition of photosynthetic carbon flux and electron transport. The chloroplast metabolism might directly act as cellular metal sensors, which could produce retrograde signals for regulation of nuclear genes and thereby affect cellular metabolism. All these metals deficiency cause cellular metabolic remodeling for improving metals acquisition and distribution in potato plants, such as the enhancement of glycolysis, TCA cycle and ATP synthesis in Fe-, Mn- and Zn-deficient plants, the differential expression of several amino acid biosynthesis-related enzymes in Fe- and Mn-deficient plants, and the induction of secondary metabolism in Fe- and Zn-deficient plants. Fe deficiency triggered more defense mechanisms than Mn and Zn deficiency, especially defense against pathogens and pests. The antioxidant systems showed different responses to protect against oxidative damage under each metal-deficiency condition. Reprogramming of protein biosynthesis/degradation and assembly was more strongly required for Fe-deficient plants, which might control the translation of some genes for the acclimation to Fe deficiency, and maintain strict protein quality control for cellular homeostasis. An activation of auxin signaling pathway might regulate polar auxin transport in Fe-, Mn- and Zn-deficient plants. There also exists active transport of ions and metabolites for adaptation response to Fe, Mn and Zn deficiency. Future work should integrate transcriptomics, proteomics, and metabolomics approaches to gain a comprehensive knowledge of the sophisticated molecular networks of response and acclimation to Fe, Mn and Zn deficiency in potato plants.
The representative models for summarizing the functional and regulatory networks activated by Fe, Mn and Zn deficiency in potato plants in vitro. The identified proteins are displayed on the corresponding metabolic pathways. The small spheres are representative of Fe-, Mn- and Zn-deficiency treatment, respectively. The up or down arrow given on the right side of each small sphere represented that the protein was up- or down-regulated under corresponding metal deficiency. ABP auxin-binding protein, ADH alcohol dehydrogenase, AK adenylate kinase, APX ascorbate peroxidase, CAT catalase, DHAR dehydroascorbate reductase, EF elongation factor, FBA fructose-bisphosphate aldolase, Fd ferredoxin, FDH formate dehydrogenase, FeSOD superoxide dismutase [Fe], FNR ferredoxin NADP+ reductase, GADH glyceraldehyde 3-phosphate dehydrogenase, GH3 indole-3-acetic acid-amido synthetase, GRP glycine-rich RNA binding protein, GST glutathione S-transferase, hnRNP heterogeneous nuclear ribonucleoprotein, Inhibitor II-CAT inhibitor II-chloramphenicol acetyltransferase, MDH malate dehydrogenase, MSP manganese-stabilizing protein, NAC nascent polypeptide-associated complex, NDH photosynthetic NDH subunit of luminal, NDPK nucleoside diphosphate kinase, nsLTP non-specific lipid-transfer protein, OEE oxygen-evolving enhancer protein, PGK phosphoglycerate kinase, PPIase peptidyl-prolyl cis–trans isomerase, PR perakine reductase, PSI-D photosystem I reaction centre PSI-D subunit, RP ribosomal protein, RPF ribosome-recycling factor, RuBisco ribulose-1,5-bisphosphate carboxylase/oxygenase, TPI triose phosphate isomerase, 34 kDa VDAC 34 kDa outer mitochondrial membrane protein porin
Author contribution statement
FZ designed the study. LC, LY and SZ performed the experiments. LC, YW and BY analyzed the data. LC wrote the manuscript. All authors read and approved the final manuscript.
Abbreviations
- CAT:
-
Catalase
- 2-DE:
-
Two-dimensional gel electrophoresis
- DHAR:
-
Dehydroascorbate reductase
- Fd:
-
Ferredoxin
- FDH:
-
Formate dehydrogenase
- FeSOD:
-
Superoxide dismutase [Fe]
- FNR:
-
Ferredoxin NADP+ reductase
- NDPKs:
-
Nucleoside diphosphate kinases
- nsLTP:
-
Non-specific lipid-transfer protein
- OEE:
-
Oxygen-evolving enhancer protein
References
Abadía J, Vázquez S, Rellán-Álvarez R, El-Jendoubi H, Abadía A, Alvarez-Fernández A, López-Millán AF (2011) Towards a knowledge-based correction of iron chlorosis. Plant Physiol Biochem 49:471–482
Alekseeva AA, Savin SS, Tishkov VI (2011) NAD(+)-dependent formate dehydrogenase from plants. Acta Nat 3:38–54
Aliverti A, Pandini V, Pennati A, de Rosa M, Zanetti G (2008) Structural and functional diversity of ferredoxin-NADP+ reductases. Arch Biochem Biophys 474:283–291
Allen MD, Kropat J, Tottey S, Campo JAD, Merchant SS (2007) Manganese deficiency in Chlamydomonas results in loss of photosystem II and MnSOD function, sensitivity to peroxides, and secondary phosphorus and iron deficiency. Plant Physiol 143:263–277
Andreini C, Bertini I, Cavallaro G, Holliday G, Thornton J (2008) Metal ions in biological catalysis: from enzyme databases to general principles. J Biol Inorg Chem 13:1205–1218
Assunção AG, Schat H, Aarts MG (2010a) Regulation of the adaptation to zinc deficiency in plants. Plant Signal Behav 5:1553–1555
Assunção AG, Herrero E, Lin YF, Huettel B, Talukdar S, Smaczniak C, Immink RG, van Eldik M, Fiers M, Schat H, Aarts MG (2010b) Arabidopsis thaliana transcription factors bZIP19 and bZIP23 regulate the adaptation to zinc deficiency. Proc Natl Acad Sci USA 107:10296–10301
Bacaicoa E, Zamarreño ÁM, Leménager D, Baigorri R, García-Mina JM (2009) Relationship between the hormonal balance and the regulation of iron deficiency stress responses in cucumber. J Am Soc Hortic Sci 134:589–601
Balk J, Schaedler TA (2014) Iron cofactor assembly in plants. Annu Rev Plant Biol 65:125–153
Bhushan D, Pandey A, Choudhary MK, Datta A, Chakraborty S, Chakraborty N (2007) Comparative proteomics analysis of differentially expressed proteins in chickpea extracellular matrix during dehydration stress. Mol Cell Proteomics 6:1868–1884
Briat JF, Dubos C, Gaymard F (2015) Iron nutrition, biomass production, and plant product quality. Trends Plant Sci 20:33–40
Broadley MR, White PJ, Hammond JP, Zelko I, Lux A (2007) Zinc in plants. New Phytol 173:677–702
Cailliatte R, Schikora A, Briat JF, Mari S, Curie C (2010) High affinity manganese uptake by the metal transporter NRAMP1 is essential for Arabidopsis growth in low manganese conditions. Plant Cell 22:904–917
Cakmak I (2000) Possible roles of zinc in protecting plant cells from damage by reactive oxygen species. New Phytol 146:185–205
Cakmak I, Pfeiffer WH, Mc Clafferty B (2010) Biofortification of durum wheat with zinc and iron. Cereal Chem 87:10–20
Carvalho AO, Gomes VM (2007) Role of plant lipid transfer proteins in plant cell physiology-a concise review. Peptides 28:1144–1153
Chatterjee C, Gopal R, Dube BK (2006) Impact of iron stress on biomass, yield, metabolism and quality of potato (Solanum tuberosum L.). Sci Hortic 108:1–6
Choudhary MK, Basu D, Datta A, Chakraborty N, Chakraborty S (2009) Dehydration responsive nuclear proteome of rice (Oryza sativa L.) illustrates protein network, novel regulators of cellular adaptation, and evolutionary perspective. Mol Cell Proteomics 8:1579–1598
Curie C, Briat JF (2003) Iron transport and signaling in plants. Annu Rev Plant Biol 54:183–206
De Pinto V, Messina A, Lane DJR, Lawen A (2010) Voltage-dependent anion-selective channel (VDAC) in the plasma membrane. FEBS Lett 584:1793–1799
Dixon DP, Cummins L, Cole DJ, Edwards R (1998) Glutathione-mediated detoxification systems in plants. Curr Opin Plant Biol 1:258–266
Donnelly BE, Madden RD, Ayoubi P, Porter DR, Dillwith JW (2005) The wheat (Triticum aestivum L.) leaf proteome. Proteomics 5:1624–1633
Dzeja PP, Terzic A (2009) Adenylate kinase and AMP signaling networks: metabolic monitoring, signal communication and body energy sensing. Int J Mol Sci 10:1729–1772
Effendi Y, Rietz S, Fischer U, Scherer GF (2011) The heterozygous abp1/ABP1 insertional mutant has defects in functions requiring polar auxin transport and in regulation of early auxin-regulated genes. Plant J 65:282–294
Fukuyama K (2004) Structure and function of plant-type ferredoxins. Photosynth Res 81:289–301
Gangadhar BH, Sajeesh K, Venkatesh J, Baskar V, Abhinandan K, Yu JW, Prasad R, Mishra RK (2016) Enhanced tolerance of transgenic potato plants over-expressing non-specific lipid transfer protein-1 (StnsLTP1) against multiple abiotic stresses. Front Plant Sci 7:1228
Hagen G (2015) Auxin signal transduction. Essays Biochem 58:1–12
Hänsch R, Mendel RR (2009) Physiological functions of mineral micronutrients (Cu, Zn, Mn, Fe, Ni, Mo, B, Cl). Curr Opin Plant Biol 12:259–266
Hantzis LJ, Kroh GE, Jahn CE, Cantrell M, Peers G, Pilon M, Ravet K (2018) A program for iron economy during deficiency targets specific Fe proteins. Plant Physiol 176:596–610
Haque ME, Yoshida Y, Hasunuma K (2010) ROS resistance in Pisum sativum cv. Alaska: the involvement of nucleoside diphosphate kinase in oxidative stress responses via the regulation of antioxidants. Planta 232:367–382
Hasunuma K, Yabe N, Yoshida Y, Ogura Y, Hamada T (2003) Putative functions of nucleoside diphosphate kinase in plants and fungi. J Bioenerg Biomembr 35:57–65
Heide H, Kalisz HM, Follmann H (2004) The oxygen evolving enhancer protein 1 (OEE) of photosystem II in green algae exhibits thioredoxin activity. J Plant Physiol 161:139–149
Hsieh SI, Castruita M, Malasarn D, Urzica E, Erde J, Page MD, Yamasaki H, Casero D, Pellegrini M, Merchant SS, Loo JA (2013) The proteome of copper, iron, zinc, and manganese micronutrient deficiency in Chlamydomonas reinhardtii. Mol Cell Proteomics 12:65–86
Ivanov R, Brumbarova T, Bauer P (2012) Fitting into the harsh reality: regulation of iron-deficiency responses in dicotyledonous plants. Mol Plant 5:27–42
Katayama H, Nagasu T, Oda Y (2001) Improvement of in-gel digestion protocol for peptide mass fingerprinting by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Rapid Commun Mass Spectrom 15:1416–1421
Keil M, Sánchezserrano J, Schell J, Willmitzer L (1990) Localization of elements important for the wound-inducible expression of a chimeric potato proteinase inhibitor II-CAT gene in transgenic tobacco plants. Plant Cell 2:61–70
Khandakar J, Haraguchi I, Yamaguchi K, Kitamura Y (2013) A small-scale proteomic approach reveals a survival strategy, including a reduction in alkaloid biosynthesis, in Hyoscyamus albus roots subjected to iron deficiency. Front Plant Sci 4:331
Kim YH, Kim MD, Choi YI, Park SC, Yun DJ, Noh EW, Lee HS, Kwak SS (2011) Transgenic poplar expressing Arabidopsis NDPK2 enhances growth as well as oxidative stress tolerance. Plant Biotechnol J 9:334–347
Laganowsky A, Gómez SM, Whitelegge JP, Nishio JN (2009) Hydroponics on a chip: analysis of the Fe deficient Arabidopsis thylakoid membrane proteome. J Proteomics 72:397–415
Lan P, Li W, Wen TN, Shiau JY, Wu YC, Lin W, Schmidt W (2011) iTRAQ protein profile analysis of Arabidopsis roots reveals new aspects critical for iron homeostasis. Plant Physiol 155:821–834
Legay S, Guignard C, Ziebel J, Evers D (2012) Iron uptake and homeostasis related genes in potato cultivated in vitro under iron deficiency and overload. Plant Physiol Biochem 60:180–189
López-Millán AF, Morales F, Andaluz S, Gogorcena Y, Abadía A, De Las Rivas J, Abadía J (2000) Responses of sugar beet roots to iron deficiency. Changes in carbon assimilation and oxygen use. Plant Physiol 124:885–898
López-Millán AF, Grusak MA, Abadía A, Abadía J (2013) Iron deficiency in plants: an insight from proteomic approaches. Front Plant Sci 4:254
Ma F, Liu Z, Wang TW, Hopkins MT, Peterson CA, Thompson JE (2010) Arabidopsis eIF5A3 influences growth and the response to osmotic and nutrient stress. Plant Cell Environ 33:1682–1696
Marschner P (2012) Marschner’s mineral nutrition of higher plants. Academic Press, Boston
Matsuura-Endo C, Kobayashi A, Noda T, Takigawa S, Yamauchi H, Mori M (2004) Changes in sugar content and activity of vacuolar acid invertase during low-temperature storage of potato tubers from six Japanese cultivars. J Plant Res 117:131–137
Maurer F, Müller S, Bauer P (2011) Suppression of Fe deficiency gene expression by jasmonate. Plant Physiol Biochem 49:530–536
Millaleo R, Reyes-Díaz M, Ivanov AG, Mora ML, Alberdi M (2010) Manganese as essential and toxic element for plants: transport, accumulation and resistance mechanisms. J Soil Sci Plant Nutr 10:476–494
Mills RF, Doherty ML, López-Marqués RL, Weimar T, Dupree P, Palmgren MG, Pittman JK, Williams LE (2008) ECA3, a Golgi-localized P2A-type ATPase, plays a crucial role in manganese nutrition in Arabidopsis. Plant Physiol 146:116–128
Milner MJ, Seamon J, Craft E, Kochian LV (2013) Transport properties of members of the ZIP family in plants and their role in Zn and Mn homeostasis. J Exp Bot 64:369–381
Morris J, Tian H, Park S, Sreevidya CS, Ward JM, Hirschi KD (2008) AtCCX3 is an Arabidopsis endomembrane H+-dependent K+ transporter. Plant Physiol 148:1474–1486
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497
Nouet C, Motte P, Hanikenne M (2011) Chloroplastic and mitochondrial metal homeostasis. Trends Plant Sci 16:395–404
Paque S, Mouille G, Grandont L, Alabadí D, Gaertner C, Goyallon A, Muller P, Primard-Brisset C, Sormani R, Blázquez MA, Perrot-Rechenmann C (2014) AUXIN BINDING PROTEIN1 links cell wall remodeling, auxin signaling, and cell expansion in Arabidopsis. Plant Cell 26:280–295
Patron NJ, Rogers MB, Keeling PJ (2004) Gene replacement of fructose-1,6-bisphosphate aldolase supports the hypothesis of a single photosynthetic ancestor of chromalveolates. Eukaryot Cell 3:1169–1175
Pilon M, Cohu CM, Ravet K, Abdel-Ghany SE, Gaymard F (2009) Essential transition metal homeostasis in plants. Curr Opin Plant Biol 12:347–357
Potato Genome Sequencing Consortium (PGSC) (2011) Genome sequence and analysis of the tuber crop potato. Nature 475:189–195
Rellán-Álvarez R, Andaluz S, Rodríguez-Celma J, Wohlgemuth G, Zocchi G, Álvarez-Fernández A, Fiehn O, López-Millán AF, Abadía J (2010) Changes in the proteomic and metabolic profiles of Beta vulgaris root tips in response to iron deficiency and resupply. BMC Plant Biol 10:120
Rodnina M, Wintermeyer W (2011) The ribosome as a molecular machine: the mechanism of tRNA-mRNA movement in translocation. Biochem Soc Trans 39:658–662
Rumeau D, Peltier G, Cournac L (2007) Chlororespiration and cyclic electron flow around PSI during photosynthesis and plant stress response. Plant Cell Environ 30:1041–1051
Sarowar S, Kim YJ, Kim KD, Hwang BK, Ok SH, Shin JS (2009) Overexpression of lipid transfer protein (LTP) genes enhances resistance to plant pathogens and LTP functions in long-distance systemic signaling in tobacco. Plant Cell Rep 28:419–427
Schippers JHM, Mueller-Roeber B (2010) Ribosomal composition and control of leaf development. Plant Sci 179:307–315
Schmidt SB, Jensen PE, Husted S (2016) Manganese deficiency in plants: the impact on photosystem II. Trends Plant Sci 21:622–632
Sétif P, Fischer N, Lagoutte B, Bottin H, Rochaix JD (2002) The ferredoxin docking site of photosystem I. BBA-Bioenergetics 1555:204–209
Sharma S (2007) Adaptation of photosynthesis under iron deficiency in maize. Plant Physiol 164:1261–1267
Singh BN, Mishra RN, Agarwal PK, Goswami M, Nair S, Sopory SK, Reddy MK (2004) A pea chloroplast translation elongation factor that is regulated by abiotic factors. Biochem Biophys Res Commun 320:523–530
Singh AK, Kumar R, Pareek A, Sopory SK, Singla-Pareek SL (2012) Overexpression of rice CBS domain containing protein improves salinity, oxidative, and heavy metal tolerance in transgenic tobacco. Mol Biotechnol 52:205–216
Staswick PE, Serban B, Rowe M, Tiryaki I, Maldonado MT, Maldonado MC, Suza W (2005) Characterization of an Arabidopsis enzyme family that conjugates amino acids to indole-3-acetic acid. Plant Cell 17:616–627
Strommer J (2011) The plant ADH gene family. Plant J 66:128–142
Sun L, Ruppert M, Sheludko Y, Warzecha H, Zhao Y, Stöckigt J (2008) Purification, cloning, functional expression and characterization of perakine reductase: the first example from the AKR enzyme family, extending the alkaloidal network of the plant Rauvolfia. J Plant Mol Biol 67:455–467
Suzuki K, Itai R, Suzuki K, Nakanishi H, Nishizawa NK, Yoshimura E, Mori S (1998) Formatede hydrogenase, an enzyme of anaerobic metabolism, is induced by iron deficiency in barley roots. Plant Physiol 116:725–736
Tanaka S, Mochizuki N, Nagatani A (2002) Expression of the AtGH3a gene, an Arabidopsis homologue of the soybean GH3 gene, is regulated by phytochrome B. Plant Cell Physiol 43:281–289
van Leeuwe MA, Stefels J (2007) Photosynthetic responses in Phaeocystis antarctica towards varying light and iron conditions. Biogeochemistry 83:61–70
Vreugdenhil D (2007) Potato biology and biotechnology. Elsevier, Wageningen
Wang W, Vinocur B, Shoseyov O, Altman A (2004) Role of plant heat shock proteins and molecular chaperones in the abiotic stress response. Trends Plant Sci 9:244–252
Wintz H, Fox T, Wu YY, Feng V, Chen W, Chang HS, Zhu T, Vulpe C (2003) Expression profiles of Arabidopsis thaliana in mineral deficiencies reveal novel transporters involved in metal homeostasis. J Biol Chem 278:47644–47653
Yadavalli V, Neelam S, Rao ASVC, Reddy AR, Subramanyam R (2012) Differential degradation of photosystem I subunits under iron deficiency in rice. J Plant Physiol 169:753–759
Yamada S, Komori T, Hashimoto A, Kuwata S, Imaseki H, Kubo T (2000) Differential expression of plastidic aldolase genes in Nicotiana plants under salt stress. Plant Sci 154:61–69
Yang KA, Moon HJ, Kim GT, Lim CJ, Hong JC, Lim CO, Yun DJ (2003) NDP kinase 2 regulates expression of antioxidant genes in Arabidopsis. Proc Jpn Acad Ser Biol 79:86–91
Yi X, McChargue M, Laborde S, Frankel LK, Bricker TM (2005) The manganese-stabilizing protein is required for photosystem II assembly/stability and photoautotrophy in higher plants. J Biol Chem 280:16170–16174
Zaharieva BT, Abadía J (2003) Iron deficiency enhances the levels of ascorbate, glutathione, and related enzymes in sugar beet roots. Protoplasma 221:269–275
Zhang XX, Liu SK, Takano T (2008) Overexpression of a mitochondrial ATP synthase small subunit gene (AtMtATP6) confers tolerance to several abiotic stresses in Saccharomyces cerevisiae and Arabidopsis thaliana. Biotechnol Lett 30:1289–1294
Acknowledgements
This work was supported by programs for National Key R&D Program of China (2017YFD0101905), National Natural Science Foundation of China (31471433, 31171477), Gansu High Educational Scientific Special Project (2018C-17), Gansu Provincial Key Laboratory of Aridland Crop Science (GSCS-2016-02).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
425_2019_3163_MOESM1_ESM.tif
Supplemental Fig. S1 The primary 2-DE gel maps at least three biological replicates for the potato leaf proteome of control (124 μM Na-Fe-EDTA) and Fe-deficiency treatments (20, 10 and 0 μM Na-Fe-EDTA) in vitro under Fe deficiency (TIFF 47071 kb)
425_2019_3163_MOESM2_ESM.tif
Supplemental Fig. S2 The primary 2-DE gel maps at least three biological replicates for the potato leaf proteome of control (100 μM MnCl2·4H2O) and Mn-deficiency treatments (1 and 0 μM MnCl2·4H2O) in vitro under Mn deficiency (TIFF 34022 kb)
425_2019_3163_MOESM3_ESM.tif
Supplemental Fig. S3 The primary 2-DE gel maps at least three biological replicates for the potato leaf proteome of control (30 μM ZnCl2) and Zn-deficiency treatment (0 μM ZnCl2) in vitro under Zn deficiency (TIFF 21350 kb)
425_2019_3163_MOESM4_ESM.tif
Supplemental Fig. S4 The detailed information on differentially abundant proteins within each cluster in the clustering analysis under Fe deficiency. The four columns of hierarchical cluster tree represent control (124 μM Na-Fe-EDTA) and Fe-deficiency treatments (20, 10 and 0 μM Na-Fe-EDTA), respectively. Each rows represent individual proteins. The up- and down-regulation of proteins are indicated in red and green, respectively. The intensity of colors is increased when the expression differences increased, as shown in the bar at the top. The differentially abundant proteins were grouped into eight clusters under Fe deficiency. The detailed information on these proteins within each cluster is presented, including the protein identification number and protein name (TIFF 889 kb)
425_2019_3163_MOESM5_ESM.tif
Supplemental Fig. S5 The detailed information on differentially abundant proteins within each cluster in the clustering analysis under Mn deficiency. The three columns of hierarchical cluster tree represent control (100 μM MnCl2·4H2O) and Mn-deficiency treatments (1 and 0 μM MnCl2·4H2O), respectively. Each rows represent individual proteins. The up- and down-regulation of proteins are indicated in red and green, respectively. The intensity of colors is increased when the expression differences increased, as shown in the bar at the top. The differentially abundant proteins were grouped into six clusters under Mn deficiency. The detailed information on these proteins within each cluster is presented, including the protein identification number and protein name (TIFF 496 kb)
425_2019_3163_MOESM6_ESM.tif
Supplemental Fig. S6 The detailed information on differentially abundant proteins within each cluster in the clustering analysis under Zn deficiency. The two columns of hierarchical cluster tree represent control (30 μM ZnCl2) and Zn-deficiency treatment (0 μM ZnCl2), respectively. Each rows represent individual proteins. The up- and down-regulation of proteins are indicated in red and green, respectively. The intensity of colors is increased when the expression differences increased, as shown in the bar at the top. The differentially abundant proteins were grouped into two clusters under Zn deficiency. The detailed information on these proteins within each cluster is presented, including the protein identification number and protein name (TIFF 308 kb)
425_2019_3163_MOESM8_ESM.docx
Supplemental Table S2 Identification of differentially abundant proteins of potato leaves in vitro under Fe deficiency (DOCX 14218 kb)
425_2019_3163_MOESM9_ESM.docx
Supplemental Table S3 Identification of differentially abundant proteins of potato leaves in vitro under Mn deficiency (DOCX 198 kb)
425_2019_3163_MOESM10_ESM.docx
Supplemental Table S4 Identification of differentially abundant proteins of potato leaves in vitro under Zn deficiency (DOCX 141 kb)
425_2019_3163_MOESM11_ESM.xlsx
Supplemental Table S5 The primary identification information of differentially abundant proteins of potato leaves in vitro under Fe deficiency using MALDI-TOF/TOF MS. Protein identifications were performed by searching for the NCBInr-S. tuberosum database using peptide mass fingerprinting (PMF) and MS/MS data from a MALDI-TOF/TOF mass spectrometry analysis. a The spot number of identified proteins. b Gene identification number as in GenBank. c The number of matched peptides. d MASCOT score reported after searching against the NCBInr database. e The sequence coverage percentage (%). f Theoretical pI and mass (Da) values of identified proteins estimated with MASCOT (XLSX 194 kb)
425_2019_3163_MOESM12_ESM.xlsx
Supplemental Table S6 The primary identification information of differentially abundant proteins of potato leaves in vitro under Mn deficiency using MALDI-TOF/TOF MS. Protein identifications were performed by searching for the NCBInr-S. tuberosum database using peptide mass fingerprinting (PMF) and MS/MS data from a MALDI-TOF/TOF mass spectrometry analysis. a The spot number of identified proteins. b Gene identification number as in GenBank. c The number of matched peptides. d MASCOT score reported after searching against the NCBInr database. e The sequence coverage percentage (%). f Theoretical pI and mass (Da) values of identified proteins estimated with MASCOT (XLSX 88 kb)
425_2019_3163_MOESM13_ESM.xlsx
Supplemental Table S7 The primary identification information of differentially abundant proteins of potato leaves in vitro under Zn deficiency using MALDI-TOF/TOF MS. Protein identifications were performed by searching for the NCBInr-S. tuberosum database using peptide mass fingerprinting (PMF) and MS/MS data from a MALDI-TOF/TOF mass spectrometry analysis. a The spot number of identified proteins. b Gene identification number as in GenBank. c The number of matched peptides. d MASCOT score reported after searching against the NCBInr database. e The sequence coverage percentage (%). f Theoretical pI and mass (Da) values of identified proteins estimated with MASCOT (XLSX 70 kb)
Rights and permissions
About this article
Cite this article
Cheng, L., Zhang, S., Yang, L. et al. Comparative proteomics illustrates the complexity of Fe, Mn and Zn deficiency-responsive mechanisms of potato (Solanum tuberosum L.) plants in vitro. Planta 250, 199–217 (2019). https://doi.org/10.1007/s00425-019-03163-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-019-03163-w