Abstract
Purpose
Pilocytic astrocytomas (PCAs) are characterized by two dominant molecular alterations of the BRAF gene, i.e., BRAFV600E mutation and KIAA1549-BRAF fusions which show a differential pattern of frequency across different age-groups.
Methods
Formalin-fixed paraffin-embedded tissues of 358 (pediatric 276 and adult 82) consecutive PCAs were evaluated for BRAFV600E mutation by Sanger sequencing and KIAA1549:BRAF fusion transcripts (KIAA1549:BRAF 16-9, KIAA1549:BRAF 15-9, and KIAA1549:BRAF 16-11) by reverse transcriptase polymerase chain reaction, which were correlated with different clinicopathological features.
Results
BRAFV600E mutation was detected in 8.9% pediatric and 9.75% adult PCAs, whereas 41.1% and 25.7% of pediatric and adult cases showed KIAA1549-BRAF fusions respectively. BRAFV600E did not show any statistically significant correlation with any of the clinical parameters (age, location, and gender). KIAA1549:BRAF fusions showed a significant statistical association with the pediatric age group and cerebellar location. KIAA1549-BRAF 16-9 was the commonest variant and was predominantly associated with cerebellar location than non-cerebellar whereas fusion variant 15-9 negatively correlated with cerebellar locations.
Conclusions
The present study showed overall frequency of 53.5% and 37.3% BRAF alterations in pediatric and adult PCA cases respectively. BRAF fusion in PCA cases showed a different distribution pattern across age groups and locations; while no such differential pattern was observed for BRAFV600E.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pilocytic astrocytoma (PCA) is the most frequent pediatric low-grade glioma and is also not uncommon in young adults, accounting for 15.3% among children and adolescents [1, 2]. It is WHO grade I circumscribed glioma classified under other astrocytic tumors in the current 2016 WHO classification [3]. It commonly occurs in the cerebellum and the optic pathway but can occur throughout the neuroaxis [4]. It is predominantly sporadic but is the most common familial syndrome associated glial tumor, especially in neurofibromatosis 1 (NF1). PCAs are of favorable prognosis with > 95% of 10-year overall survival [1].
PCAs are mediated by constitutive activation of mitogen-activated protein kinase (MAPK) pathway through the in-frame fusion between oncogene BRAF and KIAA1549 genes (resulting from breakpoint at multiple sites of KIAA1549 and BRAF generating different fusion variants), through oncogenic BRAFV600E mutation and uncommonly through alterations in fibroblast growth factor receptor 1 (FGFR1). KIAA1549 Exon 16-BRAF Exon 9 fusion is the commonest followed by KIAA1549 Exon 15-BRAF Exon 9 and KIAA1549 Exon 16-BRAF Exon 11 [5,6,7]. Other uncommon reported fusion variants are Exon 18-10, 19-9, 16-10, 15-11, 17-10 of KIAA1549 and BRAF [5, 6]. Rare non-KIAA gene fusion variants such as SRGAP3-BRAF [8], FAM131B:BRAF [9, 10], GIT2I-BRAF [11], TMEM106B-BRAF fusions [12], and BRAF gain of function mutation are also reported [13]. While BRAFV600E mutation results in constitutive activation of BRAF even in its monomeric form and also induces the same effect on MAPK pathway activation as in BRAF fusions [14]. FGFR1 alterations also induce MAPK/ERK pathway upregulation through intragenic duplications of the FGFR1 tyrosine kinase domain resulting in FGFR1 autophosphorylation. However, this is relatively frequent in grade II diffuse gliomas (24%) than PCAs and LGGs (7.4%) [15].
Although both the BRAF (fusions and V600E) alterations exert their effect of tumorigenesis through the MAPK pathway, but their frequency pattern varies according to age of diagnosis and location. KIAA1549-BRAF fusions are more common in younger age and in cerebellar location, whereas BRAFV600E is more common in non-cerebellar location [13, 16, 17]. KIAA1549-BRAF fusion was reported in around 60–70% of all PCAs, whereas BRAFV600E mutation in < 10% of both pediatric and adult cases [17,18,19].
This is a single institutional study aimed at evaluating the frequency of BRAFV600E mutation and KIAA1549-BRAF (16-9, 15-9, and 16-11) fusions in PCAs and to correlate it with clinicopathological features.
Materials and methods
Tumor samples
The study is approved by our institutional review board, comprised of 358 consecutive (2011–2018) histologically diagnosed cases of PCA (including pilomyxoid astrocytomas (PMA)). Cases with syndromic association (especially NF1) and inadequate tissue were excluded. Formalin-fixed paraffin-embedded (FFPE) tumor tissues were used for the study, which were essentially primary samples at diagnosis; 205 (pediatric 157/276, adult 48/82) were in-hospital and 153 (pediatric 119/276, adult 34/82) were referrals. Clinical and demographic details were collected from electronic medical records.
Immunohistochemistry
p53 (Dako, D07, 1:50 dilution)and ATRX protein (1:750; Polyclonal; Sigma; USA) expression was performed on 4 μm representative FFPE sections using the Ventana Benchmark XT autoimmunostainer by polymer detection kit.
p53 protein expression was graded as positive (> 50% tumor cells with strong intensity nuclear staining), focal positive (10–50% with strong nuclear staining/> 10% tumor cells with moderate intensity staining) and negative (absence of any staining or staining in < 10% tumor cells). ATRX expression was interpreted as retained (nuclear staining in tumor cells), loss (no staining in tumor cells with reactivity of native cells like endothelial and/or neurons as an internal control), and non-contributory (no staining in internal control).
Molecular analysis: BRAF alterations
Selected FFPE blocks were subjected for detection of BRAF mutation by Sanger sequencing and KIAA1549-BRAF fusion transcript detection by Reverse transcriptase PCR (RT-PCR) for three different fusion transcripts; KIAA1549-BRAF 16-9 (124 bp), KIAA1549-BRAF 15-9 (159 bp) and KIAA1549-BRAF 16-11 (Table 1: primer sequences). See Appendix for the detailed methodology of BRAFV600 sequencing and BRAF fusion RT-PCR.
Validation of KIAA1549: BRAF fusion transcripts
Sanger sequencing was performed on 10% of the KIAA1549-BRAF fusion-positive samples to validate and confirm the presence of KIAA1549-BRAF fusion transcripts (Appendix). The sequences were aligned and compared with the reference sequences using the BLAST alignment tool.
Statistical analysis
The data was analyzed using SPSS 20.0 software. The BRAF mutation and KIAA1549-BRAF fusion results were correlated with the clinical and histological features using Fisher’s test and p value < 0.05 was considered statistically significant.
Results (Table 2 )
Sample size (n = 358)
214 were males and 144 were females (M:F ratio of 1.5:1). Age range was 1–50 years (interquartile range [IQR] 6–18 years; median10 years) with 276 pediatric cases (≤ 18 years) and 82 adults (19–39 years,77; >39 years, 5).
Pediatric group (n = 276)
Clinical
M:F was 1.4:1 and IQR was 5–13 years (0–3 years, 29; 4–14 years, 203; 15–18 years, 44) with median age of 8 years.The most common location was cerebellar (n 123; 44.6%) followed by 3rd ventricular (n 68; 24.6% (suprasellar, 53; 3rd ventricle,11; hypothalamic, 3; pineal, 1)), cerebrum (n 25; 9.1%), spinal (n 17; 6.2%), optic nerve and chiasma (n 15; 5.4%), thalamic (n 14; 5.1%), and brainstem (n 14; 5.1%). Histologically, 270 were PCA while 6 were PMAs.
BRAF alterations
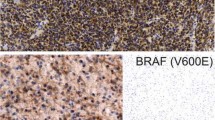
Interpretable BRAFV600E was in 246 cases, while the remaining 30 were uninterpretable (17 could not be amplified for beta-actin gene and 13 showed non-readable sequencing data). BRAFV600E was observed in 22 (8.9%) cases. All cases showed typical heterozygous BRAFV600E substitution replacing adenine in place of thymine nucleotide at codon 600 (Fig. 1).
Representative photomicrographs of pediatric PCA. a Cerebellar PCA (H&E; × 200) with electropherograms showing BRAFV600 and gel picture depicting KIAA1549-BRAF 16-9 fusion positive. b 3rdventricularPCA (H&E; × 200) with electropherograms showing BRAFV600E mutant (black arrow) and gel picture depicting KIAA1549-BRAF fusion negative. c Suprasellar PCA (H&E; × 200) with electropherograms showing BRAFV600 wild type and gel picture depicting KIAA1549-BRAF fusion negative
KIAA1549-BRAF fusion PCR could be performed in 239 pediatric cases (37 failed due to degraded RNA), 99 (41.1%) cases showed fusion transcripts. Sixty-six cases were positive for KIAA1549-BRAF 16-9 transcripts (Fig. 1) whereas KIAA1549-BRAF 15-9 and KIAA1549-BRAF 16-11 fusion transcripts were detected in 21 and 12 cases respectively. None of the cases showed the presence of more than one fusion transcript and were mutually exclusive with BRAFV600E.
Of the 276 cases, both BRAF mutation and KIAA1549-BRAF fusion transcript expression could be analyzed in 226 cases with 53.5% (121/226) of cases showing BRAF alterations. 43.8% (n 99) were KIAA1549-BRAF and only 9.7% cases demonstrated the presence of BRAFV600E. All PMA cases within the group were negative for BRAF alterations.
p53 (n = 190) and ATRX protein (n = 272) expression
p53 protein expression was not available in 86 cases, 17 (8.9%) were positive, 47 (23.9%) were focal positive and 126 (66.3%) were negative. All 272 cases showed retained ATRX expression (4 were non-contributory).
Adult group (n = 82)
Clinical
M:F was 1.9:1 and age range was 19–50 years with IQR of 21–28 years (19–39 years, 77; > 39 years, 5), and median age 24 years. Cerebellum was the commonest location (n 27; 32.9%), followed by cerebrum (n 16; 19.5%), 3rdventricular (n 13; 15.9% (suprasellar, 8; 3rd ventricle, 2; hypothalamic, 2; pineal, 1)), spinal (n = 12; 14.6%), thalamic (n 6; 7.3%), brainstem (n 4; 4.9%), and optic nerve/chaisma (n 4; 4.9%).
BRAF alterations
Ten were uninterpretable for BRAFV600E (7 had suboptimal DNA preservation and 3 showed noisy sequencing data); 9.7% (7/72 (19–39, 7/67; > 39, 0/5)) were BRAFV600E mutant (Fig. 2).
Representative photomicrographs of adult PCA. a Cerebellar PCA (H&E; × 100) with electropherograms showing BRAFV600 and gel picture depicting KIAA1549:BRAF 15-9 fusion positive. b Suprasellar PCA (H&E; × 100) with electropherograms showing BRAFV600E mutant (black arrow) and gel picture depicting KIAA1549:BRAF fusion negative. c spinal PCA (H&E; × 200) with electropherograms showing BRAFV600 wild type and gel picture depicting KIAA1549:BRAF fusion negative
KIAA1549-BRAF fusion was interpretable in 70 cases (12 had degraded RNA). Of these, 25.7% (18/70 (19–39, 16/65; > 39, 2/5)) were positive for KIAA1549-BRAF fusion (KIAA1549-BRAF 16-9, 11; KIAA1549-BRAF 15-9, 5; and KIAA1549-BRAF 16-11, 2) (Fig. 2).
Sixty-seven cases were interpretable for both BRAF alterations, of which 25 (37.3%) BRAF alterations (25/67; BRAFV600E, 7; BRAF fusion, 18).
p53 (n = 59) and ATRX (n = 78) protein expression
p53 was positive in 13.6% (8/59) cases, focal positive in 20.3% (12/59) and 66.1% (39/59) negative cases. One case showed a mosaic pattern of ATRX expression (which was negative for BRAF alterations) and 77(98.7%) showed retained ATRX protein.
Validation of KIAA1549: BRAF fusion transcripts
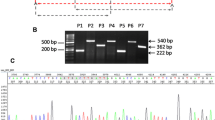
12/117 (10%) of KIAA1549-BRAF fusion-positive cases were confirmed by Sanger sequencing for all three fusion variants and their breakpoints (Fig. 3).
Correlation of BRAF alterations with clinicopathological parameters
Clinical
BRAFV00E was detected in the age range of 5–33 years (median age13). No significant difference (p value 0.82) was observed between BRAFV600E and age group (pediatric 8.9% and adult group 9.7%) and also for gender (p value 0.84). Of 117 KIAA1549-BRAF fusion positive cases, 78 were males and 39 were females with a median age of 8 years. There was no statistically significant difference observed for KIAA1549-BRAF fusion and gender (p value 0.15). KIAA1549-BRAF fusions were more common in the pediatric age group as compared with adults and this preponderance in ≤ 18 years (99/239; 41.4%) is statistically significant (p value 0.02) as compared with that in adults (> 18 years 18/70; 25.7%). Within the adult group, two cases were > 39 years (42 years of cerebellar location and 46 years of cervicomedullary location) showed presence of KIAA1549-BRAF 16-9 fusions.
Location
None of the tumor location showed significant statistical correlation for BRAFV600E mutation in pediatric and/or adult age group. None of the optic nerve/chiasma cases showed BRAF mutation. In the pediatric group, BRAF fusions were significantly associated with cerebellar as compared with non-cerebellar location (p = 0.01). It was the commonest location within fusion positive cases. 55/99 (55.6%) fusion positive pediatric cases were from cerebellar location. Among the three variants of fusion transcripts, KIAA1549-BRAF 16-9 correlated significantly with cerebellar location than other two fusion transcripts (p = 0.03) whereas fusion 15-9 transcript showed significant association with non-cerebellar location and negatively correlated with cerebellar location (p = 0.00).Adult cases also showed cerebellar (37.5%) as the predominant location for BRAF fusion but did not show a statistical significant association (p = 0.15).
p53 and ATRX protein expression
8.3% of BRAFV600E were in p53 positive and 8.9% were in focal positive while 11.5% were in negative cases. BRAF fusion transcripts were present in 33.3% p53 positive, 37.7% focal positive and 43.6% in p53 negative cases. No statistically significant correlation was observed. All except one case showed ATRX retained expression and thus no further comparative evaluation was done.
Discussion
PCAs are characterized by constitutive activation of the MAPK pathway through two major BRAF gene alterations—BRAF fusion and BRAFV600E mutation. The BRAF gene located at chromosome 7 (7q34) encodes for serine/threonine RAF kinase which acts as an intracellular transducer of the RAS/RAF/MEK/ERK pathway where activated RAS induces dimerization and activation of BRAF leading to MAPK activation, which causes tumorigenesis [20, 21]. The most common mechanism of alteration of MAPK pathway in PCAs is tandem duplication of approximately 2 MB 7q34 region with subsequent fusion resulting in KIAA1549-BRAF fusion variant [5]. The event of KIAA1549-BRAF fusion results in the replacement of the N-terminal regulatory region of BRAF with the N-terminal end of KIAA1549 resulting in constitutive activation of BRAF kinase domain and upregulation of the MAPK pathway. Another mechanism of constitutive MAPK activation is BRAFV600E mutation, the most common hotspot mutation observed across various cancers including glial tumors and is a missense substitution in exon 15 of BRAF gene at nucleotide 1799 replacing valine to glutamic acid at codon 600 within the activation segment coding region resulting in constitutive activation of BRAF in its monomeric form [20, 22]. BRAF fusion is the most common genetic event and characteristic of PCAs whereas BRAFV600E is uncommon. The latter is more frequent in non-PCA other astrocytic tumors like pleomorphic xanthoastrocytomas and gangliogliomas [17].
Detection of these BRAF alterations has a diagnostic value and has become an integral part of routine diagnostic practice in neuro-oncology. KIAA1549-BRAF fusion transcripts were detected using the most popular RT-PCR due to its ease to perform, cost-effectiveness, easy interpretation. However, RT-PCR requires good-quality RNA and a different set of primers for different variants with well optimization of each set [23]. The other method of detection is fluorescent in situ hybridization (FISH), this method is challenging especially in FFPE section. In view of the low cellularity of these tumors compounding with nuclear truncation usually result in variability in the signal pattern leading to difficulty in interpreting of fusion signals and is also endowed with inability to identify definite fusion variant [24]. However, a well-optimized FISH assay can also be used as an alternative method for BRAF fusion detection in diagnostic setting with skilled personnel for interpretation.Recent more accurate and sensitive techniques have also been developed based on droplet digital PCR, NanoString technology and Pyromark system but use of these techniques are restricted due to unavailability of these platforms in routine setups [25,26,27]. BRAF mutation can also be detected with high resolution melting curve analysis, allele-specific SNaP shot and by immunohistochemistry (IHC) with BRAF mutation-specific VE-1 antibody. The detection of BRAFV600E with VE-1 antibody is a robust, cheaper, and less time-consuming method and also showed high concordance with Sanger sequencing, but in some instances, it may yield non-specific staining necessitating the confirmation with sequencing [19, 28]. This study employed RT-PCR and Sanger sequencing for detection of limited BRAF-KIAA fusions (16-9, 15-9, and 16-11) and BRAFV600E in pediatric and adult groups.
The frequency of fusion transcripts within the PCA group varies from 50 to 80% across various studies (Table 3). The frequency of BRAF fusions in the present study was 41.1% in pediatric cases which is comparatively lower than reported, even in comparison with the three fusion variants evaluated. This possibly reflects the inherency of this population or probably due to the lesser sensitivity of the RT-PCR though only the cases with adequate representation of the tumor were selected [23, 24]. BRAF fusions were observed to be less frequent in adult PCAs and the frequency was observed to be 25.7% which is similar to previous studies [16, 29].
KIAA1549-BRAF fusion transcripts are predominantly seen in PCAs and PMAs, although the presence of these fusion transcripts are also reported in few non-PCA/PMA tumors like in 70–80% of low-grade glioneuronal tumors, diffuse leptomeningeal glioneuronal tumors [30,31,32], pediatric oligodendroglial tumors [33], and few reports of pediatric diffuse gliomas, NOS [17, 34,35,36,37]. However, none (although the number is small) of the PMA cases in the present study showed the presence of BRAF fusion.
KIAA1549-BRAF fusion variant of 16-9 (pediatric 66.7%, adults 61.1%) is the commonest followed by fusion 15-9 (pediatric 21.2%, adults 27.8%) and fusion 16-11 (pediatric 12.1%, adults11.1%), which is concordant with the predominant literature [5, 38]. However, one Egyptian study reported a higher incidence of 15-9 fusion variant [34]. Faulkner et al. and Kondo et al. have shown the presence of more than one type of fusion variant in PCA cases but none of the PCA cases of the present study showed the presence of more than one variant [23, 25]. KIAA1549-BRAF fusions were detected in both pediatric and adult cases ranging from 1 to 46 years of age but were significantly associated with the pediatric age group than adults, which is consistent with previous studies [16, 17, 35].
BRAFV600E was seen in 8.9% and 9.7% in pediatric and adult groups respectively which is consistent with the current reported literature [17, 19, 39]. A recent study has also showed the presence of novel somatic duplication mutation in BRAF gene (p.V504_R506dup) in five PCA cases [40]; however, the current study did not detect non-BRAFV600E mutations. Behling et al. and Schindler et al. have reported a high frequency of BRAF mutation in ≥ 30 years; however, in the present study, BRAFV600E was more common in < 30 years with a median age of 13 years [17, 19].
Schindler et al. [17] in their large cohort (n = 1320) showed an association of BRAF mutation with non-cerebellar origin of tumors whereas the present study did not show any significant difference of BRAF mutation between cerebellar (11/134, 8.2%) and non-cerebellar locations (18/186, 9.7%), while Horbinski et al. reported a higher frequency of BRAF mutation for cerebral hemispheric location [39]; but that was not the case in the present study; however, the number of cerebral hemispheric cases in the present study was very small (n = 25).
In the study under discussion, 55.6% of BRAF fusions in pediatric cases were cerebellar; among the three variants of fusion transcripts, fusion 16-9 was strongly associated with cerebellar location whereas fusion 15-9 showed a significant correlation with non-cerebellar location (16/21; 76.2%). The association of fusion 15-9 with non-cerebellar tumors was also demonstrated by Faulkner et al. in their study of 32 cases of PCAs [23]. Fusion 16-11 did not show any specific pattern for location.
BRAFV600E has been shown to be a negative prognostic marker in pediatric LGGs. Lassaletta A et al. have shown that BRAFV600E positive cases showed significantly poor progression-free survival (27%) as compared with BRAF wild type (60.2%) and showed a higher tumor recurrence rate after conventional chemotherapy [41]. However, BRAF fusion has been shown to be a positive prognostic marker and is dependent on the clinicopathological features such as age of patient and tumor location [42]. In addition to its diagnostic and prognostic significance, it also represents a good candidate for targeted therapy as both BRAF fusion and BRAFV600E constitutively activated the MAPK pathway. In a phase I trial of the MEK inhibitor selumetinib, Banerjee et al. demonstrated a promising effect showing 20% (5/23) of recurrent LGG children with sustained partial response. Among these 5 cases with partial response, authors could perform BRAF alterations in 4 and all were positive for BRAF alterations [43]. BRAFV600E has also been shown as an effective target in progressive optic pathway glioma (OPG) where biopsy and its molecular evaluation is not a routine clinical practice. Upadhyaya et al. (2018) in their case series demonstrated an excellent positive effect of vemurafenib on progressive NF1-negative OPG cases who did not respond to frontline chemotherapy and also highlighted a potential impact of BRAF alterations in OPG cases [44]. In the present study, 5.4% of cases were OPGs and 30.8% showed the presence of BRAF alterations; however, due to the small number of OPGs, this may lead to underrepresentation of the frequency. This biological implication does suggest the need for timely biopsy and the molecular evaluation in these refractory LGG cases to identify the subjects for targeted therapy. Drobysheva et al. and Miler et al. also demonstrated a positive response and improvement in cases with PCA when treated with MAPK inhibitors, thus providing supporting evidence for BRAF alterations as a candidate for MAPK targeted therapy [45,46,47]. The present study is majorly limited by the lack of clinical follow-up; thus, it does not address the issue of prognostic significance.
In summary, this is the largest study from our subcontinent, essentially aimed at determining the frequency and distribution pattern of BRAF alterations in PCAs across all age groups. BRAF alterations were seen in 53.5% pediatric and 37.3% adult PCA cases. In keeping with the previous studies, BRAF fusion was found to be a key event in PCA than BRAF mutation and was strongly associated with PCA and cerebellar location. Interestingly, no significant difference was observed for BRAV600E for location and age group. This study will also serve as a baseline reference for other subsequent subcontinental studies.
References
Ostrom QT, Gittleman H, Truitt G, Boscia A, Kruchko C, Barnholtz-Sloan JS (2018) CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the United States in 2011–2015. Neuro-Oncology 20:iv1–iv86
Louis DN, Ohgaki H, Wiestler OD, Cavenee WK, Burger PC, Jouvet A, Scheithauer BW, Kleihues P (2007) The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol 114:97–109
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD, Kleihues P, Ellison DW (2016) The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131:803–820
Bornhorst M, Frappaz D, Packer RJ (2016) Pilocytic astrocytomas. In: Handbook of Clinical Neurology. Elsevier, Amsterdam, pp 329–344
Jones DTW, Kocialkowski S, Liu L, Pearson DM, Bäcklund LM, Ichimura K, Collins VP (2008) Tandem duplication producing a novel oncogenic BRAF fusion gene defines the majority of pilocytic astrocytomas. Cancer Res 68:8673–8677
Forshew T, Tatevossian RG, Lawson ARJ, Ma J, Neale G, Ogunkolade BW, Jones TA, Aarum J, Dalton J, Bailey S, Chaplin T, Carter RL, Gajjar A, Broniscer A, Young BD, Ellison DW, Sheer D (2009) Activation of the ERK/MAPK pathway: a signature genetic defect in posterior fossa pilocytic astrocytomas. J Pathol 218:172–181
Sievert AJ, Jackson EM, Gai X, Hakonarson H, Judkins AR, Resnick AC, Sutton LN, Storm PB, Shaikh TH, Biegel JA (2009) Duplication of 7q34 in pediatric low-grade astrocytomas detected by high-density single-nucleotide polymorphism-based genotype arrays results in a novel BRAF fusion gene. Brain Pathol 19:449–458
Jones DTW, Kocialkowski S, Liu L, Pearson DM, Ichimura K, Collins VP (2009) Oncogenic RAF1 rearrangement and a novel BRAF mutation as alternatives to KIAA1549:BRAF fusion in activating the MAPK pathway in pilocytic astrocytoma. Oncogene 28:2119–2123
Roth JJ, Santi M, Pollock AN, Harding BN, Rorke-Adams LB, Tooke LS, Biegel JA (2015) Chromosome band 7q34 deletions resulting in KIAA1549-BRAF and FAM131B-BRAF fusions in pediatric low-grade Gliomas. Brain Pathol 25:182–192
Cin H, Meyer C, Herr R, Janzarik WG, Lambert S, Jones DTW, Jacob K, Benner A, Witt H, Remke M, Bender S, Falkenstein F, van Anh TN, Olbrich H, von Deimling A, Pekrun A, Kulozik AE, Gnekow A, Scheurlen W, Witt O, Omran H, Jabado N, Collins VP, Brummer T, Marschalek R, Lichter P, Korshunov A, Pfister SM (2011) Oncogenic FAM131B-BRAF fusion resulting from 7q34 deletion comprises an alternative mechanism of MAPK pathway activation in pilocytic astrocytoma. Acta Neuropathol 121:763–774
Helgager J, Lidov HG, Mahadevan NR, Kieran MW, Ligon KL, Alexandrescu S (2017) A novel GIT2-BRAF fusion in pilocytic astrocytoma. Diagn Pathol 12:82
Hsiao SJ, Karajannis MA, Diolaiti D, Mansukhani MM, Bender JG, Kung AL, Garvin JH Jr (2017) A novel, potentially targetable TMEM106B-BRAF fusion in pleomorphic xanthoastrocytoma. Cold Spring Harb Mol Case Stud 3:a001396
Pathak P, Kumar A, Jha P, Purkait S, Faruq M, Suri A, Suri V, Sharma MC, Sarkar C (2017) Genetic alterations related to BRAF-FGFR genes and dysregulated MAPK/ERK/mTOR signaling in adult pilocytic astrocytoma. Brain Pathol 27:580–589
Wan PTC, Garnett MJ, Roe SM, Lee S, Niculescu-Duvaz D, Good VM, Project CG, Jones CM, Marshall CJ, Springer CJ, Barford D, Marais R (2004) Mechanism of activation of the RAF-ERK signaling pathway by oncogenic mutations of B-RAF. Cell 116:855–867
The St. Jude Children’s Research Hospital–Washington University Pediatric Cancer Genome Project (2013) Whole-genome sequencing identifies genetic alterations in pediatric low-grade gliomas. Nat Genet 45:602–612
Hasselblatt M, Riesmeier B, Lechtape B, Brentrup A, Stummer W, Albert FK, Sepehrnia A, Ebel H, Gerß J, Paulus W (2011) BRAF-KIAA1549 fusion transcripts are less frequent in pilocytic astrocytomas diagnosed in adults. Neuropathol Appl Neurobiol 37:803–806
Schindler G, Capper D, Meyer J, Janzarik W, Omran H, Herold-Mende C, Schmieder K, Wesseling P, Mawrin C, Hasselblatt M, Louis DN, Korshunov A, Pfister S, Hartmann C, Paulus W, Reifenberger G, von Deimling A (2011) Analysis of BRAF V600E mutation in 1,320 nervous system tumors reveals high mutation frequencies in pleomorphic xanthoastrocytoma, ganglioglioma and extra-cerebellar pilocytic astrocytoma. Acta Neuropathol 121:397–405
Horbinski C (2013) To BRAF or not to BRAF: is that even a question anymore? J Neuropathol Exp Neurol 72:2–7
Behling F, Barrantes-Freer A, Skardelly M, Nieser M, Christians A, Stockhammer F, Rohde V, Tatagiba M, Hartmann C, Stadelmann C, Schittenhelm J (2016) Frequency of BRAF V600E mutations in 969 central nervous system neoplasms. Diagn Pathol 11:55
Matallanas D, Birtwistle M, Romano D, Zebisch A, Rauch J, von Kriegsheim A, Kolch W (2011) Raf family kinases: old dogs have learned new tricks. Genes Cancer 2:232–260
Pakneshan S, Salajegheh A, Smith RA, Lam AK-Y (2013) Clinicopathological relevance of BRAF mutations in human cancer. Pathology 45:346–356
Davies H, Bignell GR, Cox C, Stephens P, Edkins S, Clegg S, Teague J, Woffendin H, Garnett MJ, Bottomley W, Davis N, Dicks E, Ewing R, Floyd Y, Gray K, Hall S, Hawes R, Hughes J, Kosmidou V, Menzies A, Mould C, Parker A, Stevens C, Watt S, Hooper S, Wilson R, Jayatilake H, Gusterson BA, Cooper C, Shipley J, Hargrave D, Pritchard-Jones K, Maitland N, Chenevix-Trench G, Riggins GJ, Bigner DD, Palmieri G, Cossu A, Flanagan A, Nicholson A, Ho JWC, Leung SY, Yuen ST, Weber BL, Seigler HF, Darrow TL, Paterson H, Marais R, Marshall CJ, Wooster R, Stratton MR, Futreal PA (2002) Mutations of the BRAF gene in human cancer. Nature 417:949–954
Faulkner C, Ellis HP, Shaw A, Penman C, Palmer A, Wragg C, Greenslade M, Haynes HR, Williams H, Lowis S, White P, Williams M, Capper D, Kurian KM (2015) BRAF fusion analysis in pilocytic astrocytomas: KIAA1549-BRAF 15-9 fusions are more frequent in the midline than within the cerebellum. J Neuropathol Exp Neurol 74:867–872
Tian Y, Rich BE, Vena N, Craig JM, MacConaill LE, Rajaram V, Goldman S, Taha H, Mahmoud M, Ozek M, Sav A, Longtine JA, Lindeman NI, Garraway LA, Ligon AH, Stiles CD, Santagata S, Chan JA, Kieran MW, Ligon KL (2011) Detection of KIAA1549-BRAF fusion transcripts in formalin-fixed paraffin-embedded pediatric low-grade gliomas. J Mol Diagn 13:669–677
Kondo A, Shimizu Y, Adachi S et al (2018) A comprehensive method for detecting fusion genes in paediatric brain tumours. Cancer Genomics - Proteomics 15:343–348
Appay R, Fina F, Macagno N, Padovani L, Colin C, Barets D, Ordioni J, Scavarda D, Giangaspero F, Badiali M, Korshunov A, M. Pfister S, T.W. Jones D, Figarella-Branger D (2018) Duplications of KIAA1549 and BRAF screening by droplet digital PCR from formalin-fixed paraffin-embedded DNA is an accurate alternative for KIAA1549-BRAF fusion detection in pilocytic astrocytomas. Mod Pathol 31:1490–1501
Ryall S, Arnoldo A, Krishnatry R, Mistry M, Khor K, Sheth J, Ling C, Leung S, Zapotocky M, Guerreiro Stucklin A, Lassaletta A, Shago M, Tabori U, Hawkins CE (2017) Multiplex detection of pediatric low-grade glioma signature fusion transcripts and duplications using the NanoString nCounter system. J Neuropathol Exp Neurol 76:562–570
Ida CM, Vrana JA, Rodriguez FJ, Jentoft ME, Caron AA, Jenkins SM, Giannini C (2013) Immunohistochemistry is highly sensitive and specific for detection of BRAF V600E mutation in pleomorphic xanthoastrocytoma. Acta Neuropathol Commun 1:20
Theeler BJ, Ellezam B, Sadighi ZS, Mehta V, Tran MD, Adesina AM, Bruner JM, Puduvalli VK (2014) Adult pilocytic astrocytomas: clinical features and molecular analysis. Neuro-Oncology 16:841–847
Qaddoumi I, Orisme W, Wen J, Santiago T, Gupta K, Dalton JD, Tang B, Haupfear K, Punchihewa C, Easton J, Mulder H, Boggs K, Shao Y, Rusch M, Becksfort J, Gupta P, Wang S, Lee RP, Brat D, Peter Collins V, Dahiya S, George D, Konomos W, Kurian KM, McFadden K, Serafini LN, Nickols H, Perry A, Shurtleff S, Gajjar A, Boop FA, Klimo PD, Mardis ER, Wilson RK, Baker SJ, Zhang J, Wu G, Downing JR, Tatevossian RG, Ellison DW (2016) Genetic alterations in uncommon low-grade neuroepithelial tumors: BRAF, FGFR1, and MYB mutations occur at high frequency and align with morphology. Acta Neuropathol 131:833–845
Deng MY, Sill M, Chiang J, Schittenhelm J, Ebinger M, Schuhmann MU, Monoranu CM, Milde T, Wittmann A, Hartmann C, Sommer C, Paulus W, Gärtner J, Brück W, Rüdiger T, Leipold A, Jaunmuktane Z, Brandner S, Giangaspero F, Nozza P, Mora J, Morales la Madrid A, Cruz Martinez O, Hansford JR, Pietsch T, Tietze A, Hernáiz-Driever P, Stoler I, Capper D, Korshunov A, Ellison DW, von Deimling A, Pfister SM, Sahm F, Jones DTW (2018) Molecularly defined diffuse leptomeningeal glioneuronal tumor (DLGNT) comprises two subgroups with distinct clinical and genetic features. Acta Neuropathol 136:239–253
Chiang JCH, Harreld JH, Orr BA, Sharma S, Ismail A, Segura AD, Ellison DW (2017) Low-grade spinal glioneuronal tumors with BRAF gene fusion and 1p deletion but without leptomeningeal dissemination. Acta Neuropathol 134:159–162
Kumar A, Pathak P, Purkait S, Faruq M, Jha P, Mallick S, Suri V, Sharma MC, Suri A, Sarkar C (2015) Oncogenic KIAA1549-BRAF fusion with activation of the MAPK/ERK pathway in pediatric oligodendrogliomas. Cancer Genet 208:91–95
Taha H, Yehia M, Mahmoud M, el-Beltagy M, Ghabriel M, el-Naggar S (2015) Incidence of KIAA1549-BRAF fusion gene in Egyptian pediatric low grade glioma. Clin Transl Med 4:10
Gierke M, Sperveslage J, Schwab D, Beschorner R, Ebinger M, Schuhmann MU, Schittenhelm J (2016) Analysis of IDH1-R132 mutation, BRAF V600 mutation and KIAA1549–BRAF fusion transcript status in central nervous system tumors supports pediatric tumor classification. J Cancer Res Clin Oncol 142:89–100
Antonelli M, Badiali M, Moi L, Buttarelli FR, Baldi C, Massimino M, Sanson M, Giangaspero F (2015) KIAA1549:BRAF fusion gene in pediatric brain tumors of various histogenesis: KIAA1549:BRAF fusion gene in pediatric brain tumors. Pediatr Blood Cancer 62:724–727
Myung JK, Cho H, Park C-K et al (2012) Analysis of the BRAF(V600E) mutation in central nervous system tumors. Transl Oncol 5:430–436
Dahiya S, Yu J, Kaul A, Leonard JR, Gutmann DH (2012) Novel BRAF alteration in a sporadic pilocytic astrocytoma. Case Rep Med 2012:1–4
Horbinski C, Nikiforova MN, Hagenkord JM, Hamilton RL, Pollack IF (2012) Interplay among BRAF, p16, p53, and MIB1 in pediatric low-grade gliomas. Neuro-oncology 14:777–789
Khater F, Langlois S, Cassart P et al (2018) Recurrent somatic BRAF insertion (p.V504_R506dup): a tumor marker and a potential therapeutic target in pilocytic astrocytoma. Oncogene 38:2994–3002
Lassaletta A, Zapotocky M, Mistry M, Ramaswamy V, Honnorat M, Krishnatry R, Guerreiro Stucklin A, Zhukova N, Arnoldo A, Ryall S, Ling C, McKeown T, Loukides J, Cruz O, de Torres C, Ho CY, Packer RJ, Tatevossian R, Qaddoumi I, Harreld JH, Dalton JD, Mulcahy-Levy J, Foreman N, Karajannis MA, Wang S, Snuderl M, Nageswara Rao A, Giannini C, Kieran M, Ligon KL, Garre ML, Nozza P, Mascelli S, Raso A, Mueller S, Nicolaides T, Silva K, Perbet R, Vasiljevic A, Faure Conter C, Frappaz D, Leary S, Crane C, Chan A, Ng HK, Shi ZF, Mao Y, Finch E, Eisenstat D, Wilson B, Carret AS, Hauser P, Sumerauer D, Krskova L, Larouche V, Fleming A, Zelcer S, Jabado N, Rutka JT, Dirks P, Taylor MD, Chen S, Bartels U, Huang A, Ellison DW, Bouffet E, Hawkins C, Tabori U (2017) Therapeutic and prognostic implications of BRAF V600E in pediatric low-grade gliomas. J Clin Oncol 35:2934–2941
Brandner S, von Deimling A (2015) Diagnostic, prognostic and predictive relevance of molecular markers in gliomas: molecular markers in gliomas. Neuropathol Appl Neurobiol 41:694–720
Banerjee A, Jakacki RI, Onar-Thomas A, Wu S, Nicolaides T, Young Poussaint T, Fangusaro J, Phillips J, Perry A, Turner D, Prados M, Packer RJ, Qaddoumi I, Gururangan S, Pollack IF, Goldman S, Doyle LA, Stewart CF, Boyett JM, Kun LE, Fouladi M (2017) A phase I trial of the MEK inhibitor selumetinib (AZD6244) in pediatric patients with recurrent or refractory low-grade glioma: a pediatric brain tumor consortium (PBTC) study. Neuro-Oncology 19:1135–1144
Upadhyaya SA, Robinson GW, Harreld JH, Klimo PD, Hoehn ME, Orr BA, Qaddoumi IA (2018) Marked functional recovery and imaging response of refractory optic pathway glioma to BRAFV600E inhibitor therapy: a report of two cases. Childs Nerv Syst 34:605–610
Ross JS, Wang K, Chmielecki J, Gay L, Johnson A, Chudnovsky J, Yelensky R, Lipson D, Ali SM, Elvin JA, Vergilio JA, Roels S, Miller VA, Nakamura BN, Gray A, Wong MK, Stephens PJ (2016) The distribution of BRAF gene fusions in solid tumors and response to targeted therapy: BRAF fusions in solid tumors. Int J Cancer 138:881–890
Miller C, Guillaume D, Dusenbery K, Clark HB, Moertel C (2017) Report of effective trametinib therapy in 2 children with progressive hypothalamic optic pathway pilocytic astrocytoma: documentation of volumetric response. J Neurosurg Pediatr 19:319–324
Drobysheva A, Klesse LJ, Bowers DC, Rajaram V, Rakheja D, Timmons CF, Wang J, Koral K, Gargan L, Ramos E, Park JY (2017) Targeted MAPK pathway inhibitors in patients with disseminated pilocytic astrocytomas. J Natl Compr Cancer Netw 15:978–982
Acknowledgments
The authors wish to acknowledge Mrs. Prachi Gogte, Mr.Vinayak Kadam, Mr. Sandeep Dhanavade, and Mrs.Dipika Dhanavade for their technical assistance.
Funding
This study was financially supported by the Terry Fox Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interests in the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix. Detailed methodology of nucleic acid extraction, PCR with subsequent sequencing for BRAFV600E and RT-PCR for BRAF fusions
Appendix. Detailed methodology of nucleic acid extraction, PCR with subsequent sequencing for BRAFV600E and RT-PCR for BRAF fusions
DNA and RNA extraction
Selected FFPE blocks were subjected to genomic DNA and total RNA extraction from four sections each of 10 μm thickness. Sections were deparaffinized with limonene (Sigma Aldrich, USA) followed by overnight digestion. DNA was extracted using QIAamp DNA mini kit (Qiagen) as per manufacturer’s instructions. Extracted DNA was checked for quality (260:280 ratio) and quantity by Nanodrop (Thermo Scientific, USA). The integrity of the DNA was assessed by PCR for beta-actin (ACTB-208 bp) housekeeping gene (Table 1) and the positive samples were then subjected to PCR for BRAF Exon 15. Total RNA extraction was performed using RecoverAll total nucleic acid isolation kit (Ambion, Thermo Scientific, USA). RNA was quantitated using Nanodrop (Thermo Scientific, USA) and complementary DNA (cDNA) was synthesized from 100 ng RNA using RevertAid™ H Minus First Strand cDNA Synthesis Kit (Thermo Scientific, USA). The quality of cDNA was determined using ACTB housekeeping gene PCR and cDNA amplifiable for ACTB gene were subjected to RT-PCR for KIAA1549-BRAF fusion.
BRAF polymerase chain reaction and sequencing
Amplification was carried out in a total volume of 20 μl reaction containing 10 μl of 2× master mix (Thermo Scientific, USA), 1 μl each of 10 pmol forward and reverse primer (Table 1), and 100 ng of template DNA. PCR conditions were initial denaturation at 94 °C for 3 min, 35 cycles of 94 °C for 30 s, annealing at 54 °C for 45 s, 72 °C for 45 s, and a final extension at 72 °C for 10 min. PCR products were electrophoresed on 1.5% agarose gel and purified with EXOSAP-IT (USB, Affymetrix). Direct DNA sequencing was performed on the purified PCR products using BigDye v3.1 cycle sequencing kit (Thermo Scientific, USA) followed by purification with BigDye × terminator kit. The purified products were sequenced on ABI3500 Genetic Analyzer (Thermo Scientific, USA) and sequences analyzed using Chromas Lite software and compared with the reference sequence for BRAFV600E mutation and other adjoining mutations.
KIAA1549-BRAF fusions: reverse transcriptase–polymerase chain reaction
RT-PCR was performed for 3 different fusion transcripts; KIAA1549-BRAF 16-9 (124 bp), KIAA1549-BRAF 15-9 (159 bp) and KIAA1549-BRAF 16-11 (182 bp). PCR reaction contained 2 μl of template cDNA, 5 μl of 2X PCR master mix (Thermo Scientific, USA), and 0.5 μl of forward and reverse primers specific for each fusion transcript (Table 1) in a total volume of 10 μl. PCR program was as follows; 94 °C for 3 min followed by 40 cycles of 94 °C for 45 s, 57–61 °C for 45 s, and 72 °C for 45 s, and completed with an extension step at 72 °C for 10 min. Annealing temperature for KIAA1549-BRAF 16-9 (124 bp), KIAA1549-BRAF 15-9 (159 bp) andKIAA1549-BRAF 16-11 (182 bp) fusion transcripts were 57 °C, 59 °C, and 61 °C respectively. PCR products were separated on 10% polyacrylamide gel, stained with ethidium bromide, and visualized under UV illumination (Alpha Imager, Bioscreen, USA).
Rights and permissions
About this article
Cite this article
Kurani, H., Gurav, M., Shetty, O. et al. Pilocytic astrocytomas: BRAFV600E and BRAF fusion expression patterns in pediatric and adult age groups. Childs Nerv Syst 35, 1525–1536 (2019). https://doi.org/10.1007/s00381-019-04282-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00381-019-04282-1