Abstract
Abundant phosphorus (P) has been applied to paddy fields in the red soil region of subtropical China. Microbial communities play important roles in soil nutrient cycling; however, the effects of P surplus on soil microbial diversity and community composition are still unclear. Soils collected from paddy fields in subtropical China was incubated and subjected to four P treatments: 33 kg ha−1 (CK), 66 kg ha−1 (P1), 132 kg ha−1 (P2), and 264 kg ha−1 (P3). Changes in bacterial and fungal diversity and community composition were evaluated by high-throughput sequencing. The different P rates had no significant effect on bacterial diversity, whereas fungal richness and diversity indexes declined significantly by increasing P rates. Principle coordinate analysis (PCoA) also indicated a shift in fungal community composition when P rates were higher than 132 kg ha−1. Available P (AP) was the dominant factor affecting fungal community composition as evaluated by canonical correspondence analysis (CCA). Multivariate regression trees (MRT) revealed that the key threshold of 53.6 mg kg−1 of AP divided treatments into two distinct groups. Linear discriminant analysis effect size (LEfSe) showed that abundances of Pseudogymnoascus and Geomyces increased, but those of Penicillium and an unknown genus of Trichocomaceae decreased when AP was ≥ 53.6 mg kg−1.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
As one of the elements limiting crop production, P fertilizer is intensively applied to obtain high crop yield in conventional agriculture. It was predicted that by 2050, global P fertilizer input would increase by 64% compared to that in 2000 and reach 23 Tg year−1 (Bouwman et al. 2013). However, only 10–20% fertilizer P is recovered by crop in the short term and a large portion of input soluble P is fixed and accumulated in soil (Bouwman et al. 2009). It has been reported that soil P surplus could increase by 54% and reach 18 Tg year−1 by 2050 (Bouwman et al. 2013). The red soil region is the main grain-producing area in subtropical China and paddy fields are the major land use sites in this region. To obtain and maintain high rice yield, abundant P is added to paddy fields. Red soil is rich in ferric and aluminum ion; thus, applied soluble P is strongly fixed and its utilization efficiency is low (Zhang and Li 1998). This has led to P accumulation up to 300% in subtropical China (Lu et al. 2000). This P surplus may not only brings potential environmental problems, such as eutrophication, but also affects soil ecological processes. For example, N fixation and soil respiration often increased with P availability (Cleveland et al. 2011). Phosphorus also affected organic carbon, microbial biomass, microbial community composition, and metabolic activity of soil (Liu et al. 2013; Wakelin et al. 2017).
Microorganisms play essential roles in nutrient cycling, and microbial community composition is closely related to soil ecosystem processes and functions (Brussaard et al. 2007). Conversely, nutrient availability, particularly N and P, also has effects on microbial properties. However, contradictory effects of P on soil microbial diversity have often been observed. For instance, P addition was shown not only to increase (Tan et al. 2013; Zhong and Cai 2007) but also decrease the microbial Shannon diversity indexes in soil (Chhabra et al. 2013). Moreover, several studies showed that P amendments can affect biomass, activity, and composition of microbial communities (Beauregard et al. 2010; Chen et al. 2016; Huang et al. 2016; Liu et al. 2012; Wakelin et al. 2017), while others revealed no obvious changes in microbial community composition under P addition (Hamel et al. 2006; Huang et al. 2016; Shi et al. 2012).
These contradictory results depend on different P fertilization rates, crop rotation, or land use and also on the method used for microbial community analysis. For example, Beauregard et al. (2010) discovered that there was no significant effect of P input on the microbial community composition based on phospholipid fatty acid analysis, whereas the composition of bacterial and fungal communities was influenced by P addition and crop season as determined by denaturing gradient gel electrophoresis. In addition, different sensitivities of bacteria and fungi to P addition make estimation of the overall microbial response more complicated.
In the present study, incubation experiments using different P fertilizer rates were carried out with a paddy soil sampled from the red soil region of subtropical China. High throughput sequencing of amplified bacterial 16S rDNA and fungal Internal Transcribed Spacer (ITS) region was done to get information on the diversity of the two microbial communities under P surplus. The specific objective of this study was to test the hypothesis that responses in composition of bacterial and fungal communities were different with different P fertilization rates.
Materials and methods
Site and soil collection
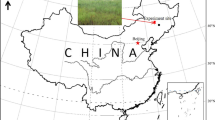
Paddy fields accounted for 30% China arable land and more than 90% of which were located in tropical or subtropical of China (Jia et al. 2010). Thus, a typical paddy filed in Yingtan, Jiangxi Province, subtropical China (28° 15′ 30′′ N, 116° 55′ 30′′ E) was selected as sampling site. Double rice was continuously cultivated before sampling. A red soil derived from Quaternary Red Clay was collected (0–15 cm) at the end of October. After sampling, soil was mixed thoroughly for the subsequent incubation experiment, air-dried at room temperature, and sieved (< 2 mm). The soil organic carbon (SOC), total N (TN), and total P (TP) contents were 40.7, 2.48, and 0.69 g kg−1, respectively, while the available N (AN) and P (AP) contents were 207 and 29.7 mg kg−1, respectively. The water holding capacity of soil was 61.0%.
Incubation experiment and soil measurement
Incubation experiment
The local P fertilization rate of the red soil region was 33 kg ha−1 P. We carried out the following four treatments: 33 kg ha−1 (CK), 66 kg ha−1 (P1), 132 kg ha−1 (P2), and 264 kg ha−1 (P3) of P; to get these rates, 36, 72, 144, and 288 mg calcium superphosphate (6% P) were mixed thoroughly with 190 g soil at a 40% water holding capacity gravimetrically. Additionally, 22 mg urea (46% N) and 19 mg potassium chloride (43% K), equivalent to 150 kg ha−1 N and 124 kg ha−1 K, respectively, were incorporated into soil. The mixture was then settled in a 600-ml plastic bottle sealed with a gas-permeable membrane. Each treatment was replicated three times. After incubation at 25 °C for 60 days, portions of the fresh samples were collected to be analyzed for microbial properties. The remaining soil samples were air-dried to determine the chemical characteristics.
Chemical property measurement
Soil pH was measured in 1:2.5 soil-to-water solutions (w/v) using the potentiometric method. After digesting with 0.8 M K2Cr2O4/concentrated H2SO4 (v/v, 1:1) SOC was measured using the Tyrin method (Pansu and Gautheyrou 2006). Kjeldahl and the alkali-hydrolysis diffusion methods were used to determine TN and AN (Pansu and Gautheyrou 2006). Soil was heated at 850 °C with anhydrous sodium carbonate, then TP was dissolved with 5 M sulfuric acid and detected by molybdenum-blue colorimetry (Pansu and Gautheyrou 2006). Soil AP was extracted by 0.5 M sodium bicarbonate and measured by molybdenum-blue colorimetry (Pansu and Gautheyrou 2006).
Microbial biomass P measurement
Microbial biomass P (MBP) was measured using the chloroform fumigation-extraction method (Brookes et al. 1982). Briefly, 5 g of non-fumigated and 5 g of chloroform fumigated soil samples were extracted by 0.5 M NaHCO3, as well as 5 g of spiked soil sample treated with KH2PO4 (equivalent to 25 mg P kg−1 soil). The extracted inorganic P was determined by molybdenum-blue colorimetry. MBP was calculated as MBP = E P / (K P × R P), where E P was the value of fumigated P minus non-fumigated P, K P was 0.4, representing 40% microbial P fractions extracted after fumigation, and R P was the P recovery efficiency estimated with the spiked soil.
DNA extraction, PCR amplification, and Miseq Illumina sequencing
Microbial genomic DNA was extracted from 0.5 g fresh soil using a FastDNA™ SPIN Kit (MP Biomedicals, Santa Ana, CA, USA). Bacterial primers 515F (5′-GTGCCAGCMGCCGCGGTAA -3′) and 806R (5′-GGACTACHVGGGTWTCTAAT-3′) with a barcode marker were used to amplify the genes targeting the V4 region of the 16S rRNA genes (Caporaso et al. 2011). Fungal primers ITS5-1737F (5′-GGAAGTAAAAGTCGTAACAAGG-3′) and ITS2-2043R (5′-GCTGCGTTCTTCATCGATGC-3′) with a barcode marker were used to amplify the genes targeting the ITS1 region (Schmidt et al. 2013; White et al. 1990). Phusion® High-Fidelity PCR Master Mix with GC Buffer (New England Biolabs, Beverly, MA, USA) was used for the amplification reaction. The PCR cycling conditions were 98 °C for 1 min, 30 cycles of 98 °C for 10 s, 50 °C for 30 s, 72 °C for 30 s, and 72 °C for 5 min. Amplicons were purified using the GeneJET kit (Thermo Scientific, Waltham, MA, USA) and a library was constructed using a NEB Next® Ultra™ DNA Library Prep Kit for Illumina (New England Biolabs). High-throughput sequencing was performed on the Illumina MiSeq PE250 platform.
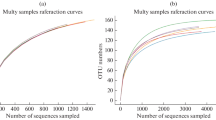
Sequencing raw data were processed using the Quantitative Insights Into Microbial Ecology (QIIME) software package (Caporaso et al. 2010). First, reads were filtered by QIIME quality filters to remove primers, barcodes, low-quality sequences (quality score Q < 25), or ambiguous nucleotides as evaluated by the database. Sequences with ≥ 97% similarity were then assigned to the same OTUs. A representative sequence for each OTU was then selected and taxonomic information was annotated by RDP classifier (Wang et al. 2007). Each sample was then rarefied to the same number of sequences for subsequent analyses. Finally, 10,947 high-quality sequences were grouped into 3859 OTUs for bacteria and 27,388 high-quality sequences were grouped into 547 OTUs for fungi for each sample. Sequences (raw data) of bacteria and fungi were submitted to Sequence Read Archive (SRA) of NCBI with the accession numbers SRP112637 and SRP112647.
Statistical analyses
Significant differences among treatments were determined by one way ANOVA with Duncan’s post hoc test at P < 0.05 (SPSS, Chicago, IL, USA). Microbial α and β diversity were analyzed by R (version 3.2.3; R Development Core Team). The Chao1 richness and Shannon diversity indexes were calculated by using the phyloseq package. Principal coordinate analysis (PCoA) and canonical correspondence analysis (CCA) was performed with the vegan package. The effects of P rates on the composition of microbial communities were determined by permutational multivariate analysis of variance (PERMANOVA) using adonis function within the vegan package. The vif.cca function was used to remove the high autocorrelation variables when CCA was conducted. Significant effects of total and specific edaphic properties on the microbial community were determined by the anova.cca and envfit functions within the vegan package. Multivariate regression trees (MRT) were generated with the mvpart package (De’Ath 2002). To identify indicator species in soils with different P availability, the linear discriminant analysis (LDA) effect size (LEfSe) method was used (http://huttenhower.sph.harvard.edu/lefse/) (Segata et al. 2011).
Results
Changes in edaphic properties and MBP under different P fertilization rates
Except for treatment P3, in which AN content was significantly higher than CK, there was no obvious difference in AN contents among the other P rates (Table 1). The total and available P contents were 0.69–0.81 and 41.4–75.0 mg kg−1, respectively, and these increased obviously with P fertilization rates (Table 1). The MBP was 14.0–21.0 mg kg−1, which was also significantly higher in treatments with higher than lower P rates (Table 1). The C/P and N/P ratios were 123–151 and 6.92–8.09, respectively, under different P fertilization rates (Table 1).
Changes in microbial diversity and community composition under different P fertilization rates
Bacterial Chao1 and Shannon indexes were not significantly affected by different P fertilization rates (Table 2). However, when input rates were > 132 kg ha−1, fungal Chao1 and Shannon indexes decreased markedly.
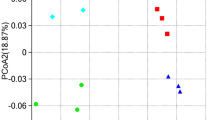
The first and second axes of PCoA explained 46.8 and 14.5% of the variation in the bacterial community composition, respectively. The bacterial communities were not clearly separated by the different P rates (Fig. 1). Axes 1 and 2 of PCoA interpreted 88.2 and 2.3% of changes in the composition of the fungal community. The fungal community of P2 and P3 treatments was separated from that of CK and P1 treatments along PCoA 1 (Fig. 1).
Principal coordinate analysis of changes in bacterial (a) and fungal (b) diversity on the basis of the Bray-Curtis distance of communities in a paddy soil treated with different P rates. CK, P fertilization rate of 33 kg ha−1; P1, P fertilization rate of 66 kg ha−1; P2, P fertilization rate of 132 kg ha−1; P3, P fertilization rate of 264 kg ha−1
Different P rates did not affect bacterial community composition as evaluated by PERMANOVA analysis (P > 0.05, Table 3). When P input rates were higher than 132 kg ha−1, the soil fungal community composition was significantly different from that of CK as evaluated by PERMANOVA analysis (P = 0.02, Table 3).
Impact factors for fungal community changes under different P fertilization rates
The first and second axes of CCA explained 77.6 and 4.7%, respectively, of the variation in the composition of fungal community by soil properties and MBP. Except for high autocorrelation variants, pH, TN, TP, AN, AP, C/N, and MBP as a whole contributed to changes in the composition of fungal community based on ANOVA (P = 0.01). Among above soil properties, only AP content (P = 0.001), TP content (P = 0.002), and MBP content (P = 0.005) were those most affecting fungal community composition changes (Fig. 2).
Canonical correspondence analysis of the effects of soil properties on the composition of fungal communities in a paddy soil treated with different P rates. MBP, microbial biomass P; AP, available P; TP, total P; AN, available N; TN, total N; CK, P fertilization rate of 33 kg ha−1; P1, P fertilization rate of 66 kg ha−1; P2, P fertilization rate of 132 kg ha−1; P3, P fertilization rate of 264 kg ha−1
Fungal communities of the different treatments were divided into two groups according to a threshold of 53.6 mg kg−1 of AP by MRT analysis (Fig. 3). The community of the CK and P1 treatments grouped together for AP content less than 53.6 mg kg−1, and those of P2 and P3 treatments grouped together over this value (Fig. 3). When the AP content was higher than 53.6 mg kg−1, abundances of Dothideomycetes, Leotiomycetes, and Sordariomycetes increased, but those of Eurotiomycetes, Un-s-Ascomycota sp, IS-s-Mortierella sp FSU10537, Agaricomycetes, Lecanoromycetes, IS-s-Mortierella sp FMR23_12, and Un-s-fungal sp. K6 decreased (Fig. 3).
Multivariate regression trees of fungal community in a paddy soil treated with different P rates, partitioned by available P. Bar plots indicated relative abundance of fungi classes. AP, available P; CK, P fertilization rate of 33 kg ha−1; P1, P fertilization rate of 66 kg ha−1; P2, P fertilization rate of 132 kg ha−1; P3, P fertilization rate of 264 kg ha−1
According to the results of MRT, we defined treatments CK and P1 as group 1 (soil P availability < 53.6 mg kg−1) and treatments P2 and P3 as group 2 (soil P availability ≥ 53.6 mg kg−1). When LDA effect size scores were < − 4.5, Penicillium and an unknown genus of Trichocomaceae were distinguished as an indicator fungi for group 1 by LEfSe analysis. Conversely, when LDA effect size scores were > 4.5, Pseudogymnoascus and Geomyces were considered indicator fungi for group 2 (Fig. 4).
Cladogram of linear discriminant analysis effect size showing the fungal indicators with different abundances among treatments. Group 1 and group 2 included treatments with soil available P lower and higher than 53.6 mg kg−1, respectively. Circles from inside root to outside indicated six fungal taxonomical levels from kingdom to genus. Red and green represent different fungal indicators in corresponding groups. Prefixes of each fungal taxonomic name such as p, c, o, f, and g were the initial letters of phylum, class, order, family, and genus, respectively
Discussion
Effect of P fertilization rates on microbial diversity
The response of MBP to high P fertilization rates confirmed what was observed by Huang et al. (2016) in a forest soil. This may occur because microorganisms can accumulate P in their biomass over their physiological demand (Docampo et al. 2010; Khan and Joergensen 2012; Ryazanova et al. 2009). For instance, intracellular polyphosphate accumulated rapidly in arbuscular mycorrhizal fungus with P application (Ezawa et al. 2003). Usually, P is taken up by soil solution and transferred to the vacuoles of arbuscular mycorrhizal fungus and transformed to polyphosphate (Ezawa et al. 2002).
Long-term application of P increased bacterial diversity in pasture soil (Tan et al. 2013), whereas high P fertilization decreased bacterial Shannon diversity indexes in barley rhizosphere soil (Chhabra et al. 2013). Phosphorus addition also changed total and alkaline phosphomonoesterases-harboring bacterial population in the ryegrass rhizosphere (Lagos et al. 2016). However, effect of P fertilization on fungal diversity has been less studied than on bacterial diversity. For example, He et al. (2016) reported P addition to soil affected fungal diversity more than N addition. Lin et al. (2012) found that long-term P application decreased arbuscular mycorrhizal fungal diversity in an arable soil. On the contrary, phospholipid fatty acid analyses revealed no changes in soil microbial composition after different P fertilization regimes (Huang et al. 2016; Shi et al. 2012). Denaturing gradient gel electrophoresis has shown a significant impact of P addition on bacterial and fungal community composition (Beauregard et al. 2010). In the present study, bacterial diversity was not affected by P, whereas fungal diversity changed markedly when P application rates were higher than 132 kg ha−1. The reason for the different responses of bacteria and fungi might be due to the fact that fungi are more sensitive than bacteria to soil fertility and P addition (He et al. 2008; Li et al. 2015). Usually, fungal hyphae can extend into soil and increase the surface area for water and nutrient absorption. Therefore, easy accessibility to soil available and even immobile P might contribute the sensitivity of fungi to P addition.
The critical soil properties affecting fungal diversity
Microbial diversity can be affected by soil properties such as moisture, temperature, pH, and organic or inorganic nutrient supply (Hamel et al. 2006). In the present study, fungal diversity was significantly affected by soil P-related factors. For example, regression analysis showed that fungal richness and diversity indexes were correlated to the AP content (Supplementary Table S1). The AP content was also the most important factor influencing fungal community composition as evaluated by CCA analysis. Similarly, soil P content was the second most important driver of bacterial and fungal diversity in soil (Siciliano et al. 2014). Moreover, extractable P concentration affected changes in fungal diversity in soils across different land use types (Lauber et al. 2008; Siciliano et al. 2014). The effect of P availability on fungal diversity may depend on: (i) changes in soil pH or osmotic potential (Liu et al. 2013; Thirukkumaran and Parkinson 2000); however, soil pH did not change in the present study; (ii) increased organic C availability by the increased litter fall or root exudate, due to the increased plant growth promoted by the increased available P (Huang et al. 2016); however, this can be excluded since no plants were involved in the incubation; (iii) and an increase in the abundance of soil fungivore, which subsequently altered competitive patterns between different fungal group (He et al. 2016). Probably, the AP content and the increase in soil fungivore abundance affected fungal diversity in this study; the effect of the AP content probably occurred by changing the osmotic potential; the impact of fungivore might take place by exerting the selective pressures on fungal communities and by changing the competitive interactions (Crowther et al. 2011).
Recent studies indicated that soil P accumulation can influence biological diversity of terrestrial ecosystems (Ceulemans et al. 2014; Kuramae et al. 2011). For example, microbial communities of grassland soil with neutral pH and soil under different plant succession were separated by the soil AP content (Kuramae et al. 2011). When AP content was higher than 100 mg kg−1, specific bacterial members became dominant (Kuramae et al. 2011). However, this AP value was not reached in our study and the red soil has different properties than the neutral pH soil used by Kuramae et al. (2011). We found the threshold of 53.6 mg kg−1 to be the AP value causing a significant shift in the fungal diversity.
Fungal indicators of different P availability
Phosphorus availability might influence microbial diversity by changing the presence of specific microorganisms in soil. For example, when soil AP content was greater than 100 mg kg−1, soil-borne copiotrophic bacteria were dominant in arable field soils (Kuramae et al. 2011). Increasing AP also stimulated the growth of r-strategy microorganisms in soils under long-term fertilization (Malý et al. 2009). The results of LEfSe indicated that Penicillium (belonging to class Eurotiomycetes) and unknown Trichocomaceae (belonging to class Eurotiomycetes) were fungal indicator of soils with AP < 53.6 mg kg−1, whereas Pseudogymnoascus (belong to Dothideomycetes) and Geomyces (belong to Leotiomycetes) were indicators for soils with AP ≥ 53.6 mg kg−1. Many strains of Penicillium are considered phosphate-solubilizing fungi contributing to the solubilization phosphate rocks (Gómez-Muñoz et al. 2017; Illmer et al. 1995; Rice et al. 1994). Some isolates of Trichocomaceae, including Aspergillus and Penicillium, can also use complex P sources, such as phytic acid and DNA (Daynes et al. 2008). Some members of Pseudogymnoascus and Geomyces were previously reported to be involved in decomposition of organic matter. For instance, Pseudogymnoascus roseus was related to the degradation of wood and its abundance increased in the rhizosphere of tree stumps (Kwasna 2001, 2004; Sigler et al. 2000). Moreover, some strains of Geomyces were reported to be involved in the decomposition and nutrient cycling in Antarctica (Arenza et al. 2006).
In soil ecosystem, fungi contribute to the degradation of lignocellulose in leaf litter and wood (Hammel 1997); mycorrhizal fungi help plant growth by P uptake (Schachtman et al. 1998). Hence, fungal diversity changes may play important roles in ecological processes under P surplus. However, this study did not give the evidence that fungi can affect C and P transformation. Lagos et al. (2016) detected the phosphomonoesterase encoding genes in bacteria as corresponding enzymes can mineralized organic P and increase soil available P. Some fungi such as Trichocomaceae also have the phosphomonoesterase activities and can grow on organic P (Daynes et al. 2008). Reduction of certain fungi may lead to the decrease of these enzyme activities with the subsequent effect on soil P.
Conclusion
The AP content was the most important soil property influencing fungal α and β diversity. When the AP content was ≥ 53.6 mg kg−1, Penicillium and Trichocomaceae decreased but Pseudogymnoascus and Geomyces increased. Fungi might play important roles in organic matter decomposition and P cycling under P surplus.
References
Arenza BE, Helda BW, Jurgensa JA, Farrellb RL, Blanchette RA (2006) Fungal diversity in soils and historic wood from the Ross Sea Region of Antarctica. Soil Biol Biochem 38(10):3057–3064. https://doi.org/10.1016/j.soilbio.2006.01.016
Beauregard MS, Atul-Nayyar CH, St-Arnaud M (2010) Long-term phosphorus fertilization impacts soil fungal and bacterial diversity but not AM fungal community in alfalfa. Microb Ecol 59(2):379–389. https://doi.org/10.1007/s00248-009-9583-z
Bouwman AF, Beusen AHW, Billen G (2009) Human alteration of the global nitrogen and phosphorus soil balances for the period 1970–2050. Global Biogeochem Cy 23:GB0A04
Bouwman L, Goldewijk KK, Hoek KWVD, Beusen AHW, Vuuren DPV, Willems J, Rufino MC, Stehfest E (2013) Exploring global changes in nitrogen and phosphorus cycles in agriculture induced by livestock production over the 1900–2050 period. P Natl Acad Sci USA 110(52):20882–20887. https://doi.org/10.1073/pnas.1012878108
Brookes PC, Powlsen DS, Jenkinson DS (1982) Measurement of microbial biomass phosphorus in soil. Soil Biol Biochem 14(4):319–329. https://doi.org/10.1016/0038-0717(82)90001-3
Brussaard L, Ruiter PC, Brown GG (2007) Soil biodiversity for agricultural sustainability. Agric Ecosyst Environ 121(3):233–244. https://doi.org/10.1016/j.agee.2006.12.013
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Tanya Y, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7(5):335–336. https://doi.org/10.1038/nmeth.f.303
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. P Natl Acad Sci USA 108(Supplement_1):4516–4522. https://doi.org/10.1073/pnas.1000080107
Ceulemans T, Stevens CJ, Duchateau L, Jacquemyn H, Gowing DJG, Merckx R, Wallace H, Rooijen NV, Goethem T, Bobbink R, Dorland E, Gaudnik C, Alard D, Corcket E, Muller S, Dise NB, Dupre C, Diekmann M, Honnay O (2014) Soil phosphorus constrains biodiversity across European grasslands. Glob Chang Biol 20(12):3814–3822. https://doi.org/10.1111/gcb.12650
Chen C, Zhang JN, Lu M, Qin C, Chen YH, Yang L, Huang QW, Wang JC, Shen ZG, Shen QR (2016) Microbial communities of an arable soil treated for 8 years with organic and inorganic fertilizers. Biol Fert Soils 52(4):455–467. https://doi.org/10.1007/s00374-016-1089-5
Chhabra S, Brazil D, Morrissey J, Burke J, O’Gara F, Dowling DN (2013) Fertilization management affects the alkaline phosphatase bacterial community in barley rhizosphere soil. Biol Fert Soils 49(1):31–39. https://doi.org/10.1007/s00374-012-0693-2
Cleveland C, Townsend AR, Taylor P, Alvarez-Clare S, Bustamante MMC, Chuyong G, Dobrowski SZ, Grierson P, Harms KE, Houlton BZ, Marklein A, Parton W, Porder S, Reed SC, Sierra CA, Silver WL, Tanner EV, Wieder WR (2011) Relationships among net primary productivity, nutrients and climate in tropical rain forest: a pan-tropical analysis. Ecol Lett 14(9):939–947. https://doi.org/10.1111/j.1461-0248.2011.01658.x
Crowther TW, Boddy L, Jones TH (2011) Outcomes of fungal interactions are determined by soil invertebrate grazers. Ecol Lett 14(11):1134–1142. https://doi.org/10.1111/j.1461-0248.2011.01682.x
Daynes CM, Mcgee PA, Midgley DJ (2008) Utilisation of plant cell-wall polysaccharides and organic phosphorus substrates by isolates of Aspergillus and Penicillium isolated from soil. Fungal Ecol 1(2-3):94–98. https://doi.org/10.1016/j.funeco.2008.09.001
De’Ath G (2002) Multivariate regression trees: a new technique for modeling species-environment relationships. Ecology 83:1105–1117
Docampo R, Ulrich P, Moreno SNJ (2010) Evolution of acidocalcisomes and their role in polyphosphate storage and osmoregulation in eukaryotic microbes. Philos T R Soc B 365(1541):775–784. https://doi.org/10.1098/rstb.2009.0179
Ezawa T, Cavagnaro TR, Smith SE, Smith FA, Ohtomo R (2003) Rapid accumulation of polyphosphate in extraradical hyphae of an arbuscular mycorrhizal fungus as revealed by histochemistry and a polyphosphate kinase/luciferase system. New Phytol 161:387–392
Ezawa T, Smith SE, Smith FA (2002) P metabolism and transport in AM fungi. Plant Soil 244(1/2):221–230. https://doi.org/10.1023/A:1020258325010
Gómez-Muñoz B, Pittroff SM, Ad N, Jensen LS, Nicolaisen MH, Magid J (2017) Penicillium bilaii effects on maize growth and P uptake from soil and localized sewage sludge in a rhizobox experiment. Biol Fert Soils 53(1):23–35. https://doi.org/10.1007/s00374-016-1149-x
Hammel K (1997) Fungal degradation of lignin. In: Cadisch G (ed) Driven by nature: plant litter quality and decomposition. CAB International, Wallingford, pp 33–45
Hamel C, Hanson K, Selles F, Cruz AF, Lemke R, McConkey B, Zentner R (2006) Seasonal and long-term resource-related variations in soil microbial communities in wheat-based rotations of the Canadian prairie. Soil Biol Biochem 38(8):2104–2116. https://doi.org/10.1016/j.soilbio.2006.01.011
He JZ, Zheng Y, Chen CR, He YQ, Zhang LM (2008) Microbial composition and diversity of an upland red soil under long-term fertilization treatments as revealed by culture-dependent and culture-independent approaches. J Soils Sediments 8(5):349–358. https://doi.org/10.1007/s11368-008-0025-1
He D, Xiang XJ, He JS, Wang C, Cao GM, Adams J, Chu HY (2016) Composition of the soil fungal community is more sensitive to phosphorus than nitrogen addition in the alpine meadow on the Qinghai-Tibetan Plateau. Biol Fert Soils 52(8):1059–1072. https://doi.org/10.1007/s00374-016-1142-4
Huang J, Hu B, Qi K, Chen W, Pang X, Bao W, Tian G (2016) Effects of phosphorus addition on soil microbial biomass and community composition in a subalpine spruce plantation. Eur J Soil Biol 72:35–41. https://doi.org/10.1016/j.ejsobi.2015.12.007
Illmer P, Barbato A, Schinner F (1995) Solubilization of hardly-soluble AlPO4 with P-solubilizing microorganisms. Soil Biol Biochem 27(3):265–270. https://doi.org/10.1016/0038-0717(94)00205-F
Jia JX, Li ZP, Liu M, Che YP (2010) Effects of glucose addition on N transformations in paddy soils with a gradient of organic C content in subtropical China. Agr Sci China 9(9):1309–1316. https://doi.org/10.1016/S1671-2927(09)60222-4
Khan KS, Joergensen RG (2012) Relationships between P fractions and the microbial biomass in soils under different land use management. Geoderma 173-174:274–281. https://doi.org/10.1016/j.geoderma.2011.12.022
Kuramae E, Gamper H, Jv V, Kowalchuk G (2011) Soil and plant factors driving the community of soil-borne microorganisms across chronosequences of secondary succession of chalk grasslands with a neutral pH. FEMS Microbiol Ecol 77(2):285–294. https://doi.org/10.1111/j.1574-6941.2011.01110.x
Kwasna H (2001) Fungi in the rhizosphere of common oak and its stumps and their possible effect on infection by Armillaria. Appl Soil Ecol 17(3):215–227. https://doi.org/10.1016/S0929-1393(01)00137-8
Kwasna H (2004) Natural shifts in communities of rhizosphere fungi of common oak after felling. Plant Soil 264(1/2):209–218. https://doi.org/10.1023/B:PLSO.0000047752.41575.c7
Lagos LM, Acuña JJ, Maruyama F, Ogram A, Mora ML, Jorquera MA (2016) Effect of phosphorus addition on total and alkaline phosphomonoesterase-harboring bacterial populations in ryegrass rhizosphere microsites. Biol Fert Soils 52(7):1007–1019. https://doi.org/10.1007/s00374-016-1137-1
Lauber CL, Strickland MS, Bradford MA, Fierer N (2008) The influence of soil properties on the structure of bacterial and fungal communities across land-use types. Soil Biol Biochem 40(9):2407–2415. https://doi.org/10.1016/j.soilbio.2008.05.021
Li J, Li Z, Wang F, Zou B, Chen Y, Zhao J, Mo Q, Li Y, Li X, Xia H (2015) Effects of nitrogen and phosphorus addition on soil microbial community in a secondary tropical forest of China. Biol Fert Soils 51(2):207–215. https://doi.org/10.1007/s00374-014-0964-1
Liu L, Gundersen P, Zhang T, Mo J (2012) Effects of phosphorus addition on soil microbial biomass and community composition in three forest types in tropical China. Soil Biol Biochem 44(1):31–38. https://doi.org/10.1016/j.soilbio.2011.08.017
Liu L, Zhang T, Gilliam FS, Gundersen P, Zhang W, Chen H, Mo J (2013) Interactive effects of nitrogen and phosphorus on soil microbial communities in a tropical forest. PLoS One 8(4):e61188. https://doi.org/10.1371/journal.pone.0061188
Lin XG, Feng YZ, Zhang HY, Chen RR, Wang JH, Zhang JB, Chu HY (2012) Long-term balanced fertilization decreases arbuscular mycorrhizal fungal diversity in an arable soil in North China revealed by 454 pyrosequencing. Environ Sci Technol 46(11):5764–5771. https://doi.org/10.1021/es3001695
Lu R, Shi Z, Shi J (2000) Nutrient balance of agroecosystem in six provinces in southern China. Sci Agr Sin 33:63–67
Malý S, Královec J, Hampel D (2009) Effects of long-term mineral fertilization on microbial biomass, microbial activity, and the presence of r- and K-strategists in soil. Biol Fert Soils 45(7):753–760. https://doi.org/10.1007/s00374-009-0388-5
Pansu M, Gautheyrou J (2006) Handbook of soil analysis: mineralogical, organic and inorganic methods. Springer, Berlin Heidelberg, Berlin. https://doi.org/10.1007/978-3-540-31211-6
Rice WA, Olsen PE, Leggett ME (1994) Co-culture of rhizobium meliloti and a phosphorus-solubilizing fungus (Penicillium bilaii) in sterile peat. Soil Biol Biochem 27:703–705
Ryazanova LP, Suzina NE, Kulakovskaya TV (2009) Phosphate accumulation of Acetobacter xylinum. Arch Microbiol 191(5):467–471. https://doi.org/10.1007/s00203-009-0470-2
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS, Huttenhower C (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12(6):R60. https://doi.org/10.1186/gb-2011-12-6-r60
Schachtman DP, Reid RJ, Ayling SM (1998) Phosphorus uptake by plants: from soil to cell. Plant Physiol 116(2):447–453. https://doi.org/10.1104/pp.116.2.447
Schmidt P, Bálint M, Greshake B, Bandow C, Römbke J, Schmitt I (2013) Illumina metabarcoding of a soil fungal community. Soil Biol Biochem 65:128–132. https://doi.org/10.1016/j.soilbio.2013.05.014
Shi Y, Lalande R, Ziadi N, Sheng M, Hu Z (2012) An assessment of the soil microbial status after 17 years of tillage and mineral P fertilization management. Appl Soil Ecol 62:14–23. https://doi.org/10.1016/j.apsoil.2012.07.004
Siciliano SD, Palmer AS, Winsley T, Lamb E, Bissett A, Brown MV, Dorst J, Ji M, Ferrari BC, Grogan P, Chu H, Snape I (2014) Soil fertility is associated with fungal and bacterial richness, whereas pH is associated with community composition in polar soil microbial communities. Soil Biol Biochem 78:10–20. https://doi.org/10.1016/j.soilbio.2014.07.005
Sigler L, Lumley TC, Currah RS (2000) New species and records of saprophytic ascomycetes (Myxotrichaceae) from decaying logs in the boreal forest. Mycoscience 41(5):495–502. https://doi.org/10.1007/BF02461670
Tan H, Barret M, Mooij MJ, Rice O, Morrissey JP, Dobson A, Griffiths B, O’Gara F (2013) Long-term phosphorus fertilisation increased the diversity of the total bacterial community and the phoD phosphorus mineraliser group in pasture soils. Biol Fert Soils 49(6):661–672. https://doi.org/10.1007/s00374-012-0755-5
Thirukkumaran CM, Parkinson D (2000) Microbial respiration, biomass, metabolic quotient and litter decomposition in a lodgepole pine forest floor amended with nitrogen and phosphorous fertilizers. Soil Biol Biochem 32(1):59–66. https://doi.org/10.1016/S0038-0717(99)00129-7
Wakelin SA, Condron LM, Gerard E, Dignam BEA, Black A, O’Callaghan M (2017) Long-term P fertilisation of pasture soil did not increase soil organic matter stocks but increased microbial biomass and activity. Biol Fert Soils 53(5):511–521. https://doi.org/10.1007/s00374-017-1212-2
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microb 73(16):5261–5267. https://doi.org/10.1128/AEM.00062-07
White T, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis M, Gelfand D, Shinsky J, White T (eds) PCR protocols: a guide to methods and applications. Academic Press, San Diego, pp 315–322
Zhang B, Li G (1998) Roles of soil organisms on the enhancement of plant availability of soil phosphorus. Acta Pedol Sin 35:104–111
Zhong WH, Cai ZC (2007) Long-term effects of inorganic fertilizers on microbial biomass and community functional diversity in a paddy soil derived from quaternary red clay. Appl Soil Ecol 36(2-3):84–91. https://doi.org/10.1016/j.apsoil.2006.12.001
Acknowledgements
The present study was funded by the National Basic Research Program of China [grant 2014CB441003], the National Nature Science Foundation of China [grant 41201242], and the “135” plan and field frontier project [grant ISSASIP1642].
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
ESM 1
(DOCX 13 kb)
Rights and permissions
About this article
Cite this article
Liu, M., Liu, J., Chen, X. et al. Shifts in bacterial and fungal diversity in a paddy soil faced with phosphorus surplus. Biol Fertil Soils 54, 259–267 (2018). https://doi.org/10.1007/s00374-017-1258-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00374-017-1258-1