Abstract
Comprising approximately 7500 living species of various corals and sea anemones, Anthozoa ranks among the most ecologically and economically valuable marine taxa. However, the taxonomy and systematics of anthozoans remain in flux as several facets of their biology (e.g. cryptic speciation, hybridisation and introgression, morphological plasticity and convergence) confound taxonomists even today. Rapid advancements in molecular sequencing and analyses have made available vast quantities of genomic data on an increasing number of species across the anthozoan tree of life. While whole genome assemblies are expected to result in the most robust phylogenetic trees, reduced-representation techniques such as genome skimming, RAD-seq, phylotranscriptomics and hybrid capture have led to well-supported inferences at various taxonomic levels and may still be favoured at this stage due to the high cost associated with even a single genome assembly. Here, we examine the different genotyping and analytical approaches used in anthozoan phylogeny reconstructions, their applicability across different divergences, and the coverage of studies among anthozoan clades to date. Based on our review of 82 phylogenomic studies, we describe the suitability of methods employed relative to their aims, highlight the imbalanced coverage of taxonomic groups studied and assess immediate and long-term needs where consolidation and streamlining of approaches would further advance the field. Overall, we find that Scleractinia (Anthozoa) is the most phylogenetically sampled group and studies on Octocorallia (Anthozoa) and its subclades are emerging. Nevertheless, we emphasise the need for more phylotranscriptomic, hybrid capture and whole genome sequencing across all anthozoans to increase topological support and generate more precise divergence time estimates. The enhanced phylogenetic understanding of Anthozoa is expected to provide insights into the evolution of genes and adaptations to environmental stressors amidst the current climate and mass extinction crises.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
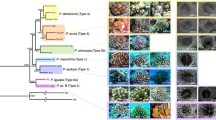
Class Anthozoa (Cnidaria) forms the foundations of a diversity of ecosystems with ~ 7500 extant species spread across shallow waters to deep reefs (Daly et al. 2007), including many of the most ecologically and economically important marine organisms (Moberg and Folke 1999; White et al. 2000; Komyakova et al. 2013). On tropical shallow-water reefs, for example, reef-building stony corals (order Scleractinia) build the underlying structural framework which serves as habitats for an extraordinary diversity of marine taxa (Small et al. 1998; Hughes et al. 2002; Plaisance et al. 2011). In deeper waters, the distribution of anthozoans continues from mesophotic zones to the deepest ocean trenches, contributing to biodiversity and ecological functioning at various spatial scales (Bongiorni et al. 2010; Watling et al. 2011). Anthozoans also provide an array of goods and services such as pharmaceutical products, fisheries, shore protection and tourism (Newton et al. 2007; Chao et al. 2012; Villanoy et al. 2012; Spalding et al. 2017; Morlighem et al. 2018). Despite their importance, the taxonomy and phylogeny of most anthozoan groups (Fig. 1), except for certain scleractinians, remain poorly studied to date.
Phylogeny of orders in Anthozoa, adapted from Quattrini et al. (2020) but excluding Relicanthus daphneae
Traditionally, a lack of consistent and informative markers has stymied progress in the classification of anthozoans based on morphology alone (Budd et al. 2010; McFadden et al. 2010). From the mid-1990s, the application of polymerase chain reaction (PCR) and Sanger sequencing—primarily of mitochondrial and ribosomal RNA (rRNA) genes—profoundly revolutionised the systematics of anthozoans (Chen et al. 1995; Romano and Palumbi 1996; Berntson et al. 1999). The most commonly used molecular markers targeted have been generally similar among anthozoan taxa, with the DNA mismatch repair protein (MutS) in octocorals (subclass Octocorallia) being a remarkable exception (Pont-Kingdon et al. 1995). This gene likely originated from a horizontal gene transfer event in the last common octocorallian ancestor (Bilewitch and Degnan 2011). Phylogenies reconstructed from these mitochondrial and rRNA genes provided unprecedented insights into anthozoan evolution, such as the distinct lineages that arose between the Atlantic and Pacific Oceans (Fukami et al. 2004; Silvestri et al. 2019), the monophyly of subclasses Hexacorallia and Octocorallia (Chen et al. 1995; Berntson et al. 1999; Fig. 1), as well as the role of hybridisation in anthozoan evolution (Odorico and Miller 1997; McFadden and Hutchinson 2004; Reimer et al. 2007).
Despite the use of multiple directly sequenced markers in most recent studies, the taxonomy and systematics of many anthozoans remain in flux today owing to poor phylogenetic resolution of these markers. Uncertainties limiting more comprehensive taxonomic revisions can be attributed chiefly to the poor phylogenetic signal of mitochondrial genes due to slow substitution rates, sequence compositional bias and substitution saturation (Shearer et al. 2008; Huang et al. 2008; Kitahara et al. 2014; Figueroa and Baco 2015; Pratlong et al. 2017a), as well as intragenomic variation in nuclear markers such as the internal transcribed spacers of rRNA (Odorico and Miller 1997; Reimer et al. 2007; Sánchez and Dorado 2008). These issues encountered with the most commonly targeted loci are compounded by anthozoans’ morphological variation, plasticity and convergence that hamper accurate species delimitation and stable taxonomic classification more generally (Todd 2008; Ament-Velásquez et al. 2016; Santos et al. 2016).
Within just the last decade, advances in DNA sequencing technologies have firmly rooted phylogenetic biology in the genomic era. Driven by developments in high-throughput sequencing (HTS), phylogeny reconstructions using genome-wide markers, broadly termed phylogenomic analyses (Philippe et al. 2005), are being churned out at an unprecedented pace and taxonomic scale across the tree of life (Galindo et al. 2019; Laumer et al. 2019; One Thousand Plant Transcriptomes Initiative 2019; Prasanna et al. 2020; Williams et al. 2020). For example, over 700 bird species have been robustly placed on the evolutionary tree by combining multiple phylogenomic studies using either ultraconserved elements (UCEs) or whole genome sequencing (WGS) into a supertree (Kimball et al. 2019).
For Anthozoa, phylogenetic analyses of HTS data primarily employ reduced-representation methods such as genome skimming, restriction site-associated DNA (RAD) sequencing, phylotranscriptomics and hybrid capture (Table 1). Nevertheless, the most comprehensive data available for phylogenomics—whole genome assemblies—have seen an uptick in recent years (Table 1; Fig. 2). Indeed, whole genome sequencing enables the extraction of orthologous loci even for traditional markers (e.g. mitochondrial and rRNA genes; see section on ‘Genome skimming’ below), so that the immense number of Sanger sequences deposited on GenBank can also be integrated for large multi-locus analyses.
There are several key considerations prior to any phylogenomic study. These include the temporal scale of divergence, sample size, the research question, bioinformatic pipeline adopted and costs involved (for a review, see Young and Gillung 2020). For most anthozoans, there exists another layer of consideration: endosymbionts. Due to the simple body plan and often small size of the polyps, digestion of samples typically encompasses tissue from the entire holobiont, defined as the organism along with their endosymbiont microbiota—including the prokaryotic partners and Symbiodiniaceae, plus viruses, fungi and other microorganisms where present (Thompson et al. 2015). As a result, any host genomic study must adopt measures for ensuring non-target DNA from contaminant organisms are discarded to avoid erroneous inferences (e.g. Artamonova and Mushegian 2013). Fortunately, there exist a variety of laboratory and bioinformatic methods to minimise the leakage of contaminant sequences into downstream analyses, such as extracting DNA from gametes (e.g. Shinzato et al. 2011, 2020), reducing the symbiont load prior to extraction (e.g. Cunning et al. 2018), and filtering non-target reads by mapping to appropriate references (e.g. Barshis et al. 2013; Forsman et al. 2017; Quek and Huang 2019; Buitrago-López et al. 2020; Quek et al. 2020).
Phylogenomics is a powerful tool in the arsenal of an evolutionary biologist. As climate change accelerates, the urgency to understand the evolutionary basis of species’ adaptation capacity and resilience has become more pressing than ever. The destruction of large swathes of shallow, mesophotic and deep-sea reef habitats around the world (Roberts et al. 2006; Hughes et al. 2017, 2018; Cunning et al. 2019), compounded by rising water temperatures and ocean acidification, translates into devastating effects projected for the environment and global economy in the next few decades (Speers et al. 2016; Dey et al. 2016; Stuart-Smith et al. 2018). To mitigate impacts on marine populations and initiate conservation policies, genomic data on anthozoans are critical as analyses can help identify cryptic species for biodiversity estimation and protection, improve population connectivity, and evaluate phylogenetic diversity as a key biodiversity measure for prioritisation (Faith 1992; Huang and Roy 2013, 2015; Winter et al. 2013; Huang et al. 2015, 2016b; Balbar and Metaxas 2019; Titus et al. 2019a, 2019b). Furthermore, the study of genomes and transcriptomes in the context of the anthozoan phylogeny can provide insights into the evolution of genes and adaptations in this diverse and ancient lineage (~ 800 Ma; Quattrini et al. 2020; McFadden et al. 2021) to environmental stressors amidst the current climate crisis (Moya et al. 2012; Takeuchi et al. 2016; Huang et al. 2017; Lin et al. 2017a, 2017b; Conci et al. 2019).
This review consolidates phylogenomic studies performed to date on Anthozoa in order to assess the taxonomic coverage and phylogenetic utility of the wide variety of tools available, as well as to identify gaps and opportunities for advancing a comprehensive, resolved and accurate representation of the Anthozoa tree of life. To compile published papers on anthozoan phylogenomic inferences, we searched Google Scholar with associated keywords, each preceded by the term ‘Anthozoa’. We trawled through the first ten pages of results, and selected studies that either employed HTS on anthozoans and/or conducted phylogenomic/mitogenomic reconstructions with anthozoans included as a focal taxon (Table 1, 2, S1; Fig. 2). Studies that investigated gene evolution or population-level phylogeography without reference to species relationships, cryptic or otherwise (e.g. Reitzel et al. 2013; Poole and Weis 2014; Takeuchi et al. 2016; Klompen et al. 2020; Sturm et al. 2020) were excluded. Only reconstructions with a broad sampling of orthologous loci were retained (Table 1, 2; Fig. 2). We note that based on the resolution of search terms, some phylogenomic studies might inadvertently be excluded in this manner of searching. For instance, a RAD-seq study by Spano et al. (2018) on sea anemones (order Actiniaria) does not include “Anthozoa” in the title or as keyword, similar to a number of mitogenome announcements (e.g. Niu et al. 2016a, 2016b, 2017a, 2017b; Teranneo et al. 2018b; Wang et al. 2018; Yu et al. 2019). Therefore, additional studies were identified from the reading of other papers and their references (i.e. snowballing; Greenhalgh and Peacock 2005), in conjunction with our previous research experiences and publications shared by colleagues (Table 1, S1). Together, the publications assembled here encompass a wide variety of studies and methods on a considerable diversity of corals and sea anemones for a comprehensive review of the phylogenomic advances in Anthozoa.
Genome skimming
Genome skimming involves extracting data from shallow HTS of whole genomes, whereby high-copy fractions (e.g. plastid, mitochondrial and rDNA genes) are ‘skimmed’ from the sequencing data for phylogenetic analysis. First established in plant phylogenetic studies, this method has been successfully applied on herbarium samples (Straub et al. 2012; Ripma et al. 2014; Nevill et al. 2020), and is especially useful for non-model organisms with no reference genomes or transcriptomes available for hybrid capture and sequencing of genome-wide markers (see section on ‘Hybrid capture and target enrichment sequencing’ below). Importantly, genome skimming enables analyses to take advantage of the large quantity of Sanger-sequenced data that can be combined with HTS data on the naturally-enriched markers for phylogenetic reconstruction, and may also help identify low-copy nuclear loci to be targeted in follow-up studies (Cronn et al. 2012; see also Zhang et al. 2019a).
Prior to HTS, whole mitochondrial genomes were typically sequenced through long-range polymerase chain reaction or primer walking (Uda et al. 2013; Lin et al. 2014; Figueroa and Baco 2015), both potentially costly and time-consuming methods. With HTS, mitogenomes across a number of samples can be obtained via multiplexing samples. One of the earliest studies investigating the mitogenomic phylogeny of anthozoans using HTS was on 11 Acropora (Scleractinia) species sequenced on an Illumina Solexa platform (Liu et al. 2015). Since then, HTS of mitogenomes has been applied widely on anthozoans, such as pennatulaceans (order Pennatulacea; Hogan et al. 2019), actiniarians (Foox et al. 2016; Xiao et al. 2019), ceriantharians (order Ceriantharia; Stampar et al. 2019) and the poorly studied antipatharians (order Antipatharia; Barrett et al. 2020). In addition to mitogenomes, other studies have skimmed mitochondrial, rRNA and histone genes from alternative sequencing data, including ezRAD (see section on ‘Restriction site-associated DNA sequencing (RAD-seq)’ below), hybrid capture markers (Poliseno et al. 2020; Quek et al. 2020) and transcriptomes (Chi and Johansen 2017).
In the past decade, mitogenomes have provided valuable insights into the genome and organismal evolution of anthozoans, challenging the conventional understanding of genome organisation in anthozoans and phylogenetic relationships among certain clades. For example, bipartite circular mitochondrial genomes have been discovered in Protanthea simplex (Actiniaria; Dubin et al. 2019) and Umbellula sp. (Pennatulacea; Hogan et al. 2019), contrary to the typical single circular mitogenome in most anthozoans. Relatedly, ceriantharians have been suggested to possess a linear mitochondrial genome (Stampar et al. 2019; but see Smith 2020), a trait characteristic of Medusozoa (Kayal et al. 2013) but not Anthozoa. With respect to phylogenetic relationships, Kayal et al. (2013) doubted the monophyly of Anthozoa, with Octocorallia recovered as sister to Medusozoa on their mitochondrial tree even after accounting for biases that may result because of amino acid composition biases and limited taxon sampling. However, recent studies based on phylotranscriptomics supported the monophyly of Anthozoa (Zapata et al. 2015; Pratlong et al. 2017a; Kayal et al. 2018; see also Fig. 1). More pertinently, one of the greatest controversies first raised with mitogenome phylogenies was the “naked coral hypothesis” (Medina et al. 2006), which posited that corallimorpharians (order Corallimorpharia) are scleractinians that have lost their skeleton. Examining amino acid trees that nested corallimorpharians within scleractinians, Kitahara et al. (2014) surmised that the unexpected relationship was likely an artefact of amino acid sequence biases, and this conclusion was later supported by broad-scale genomic analyses (Lin et al. 2016; Wang et al. 2017a). Given the slow rate of evolution in mitochondrial genes in anthozoans, substitution saturation and amino acid composition biases in some clades (Shearer et al. 2008; Huang et al. 2008; Kitahara et al. 2014; Pratlong et al. 2017a), we caution against over-reliance on mitochondrial analyses in studying anthozoan evolution, especially given the numerous high-throughput methods that have been established (as reviewed here). Instead, mitogenome sequencing would be best targeted at evaluating mitochondrial evolution and gene rearrangements (e.g. Chen et al. 2009; Lin et al. 2014; Dubin et al. 2019; Hogan et al. 2019).
With an increasing number of available genomes, the range of phylogenetic markers for genome skimming is likely to expand beyond mitochondrial and rRNA genes. Increasingly, seascape genomic analyses are taking advantage of genome data to recover phylogeographic patterns. For example, Bellis et al. (2016) identified several locality-specific lineages in the model anemone Exaiptasia (Actiniaria) based on a genome skimming approach, specifically by mapping their sequencing reads to a reference genome from Baumgarten et al. (2015). Likewise, Kitchen et al. (2019) utilised the Acropora digitifera reference genome (Shinzato et al. 2011) to identify single nucleotide variants between the two Caribbean acroporids (A. cervicornis and A. palmata) and their hybrid (A. prolifera).
However, relative to the genome-wide approaches below, the shallow sequencing effort generally results in low probability of obtaining data for low-copy genes that are orthologous across a number of samples. Therefore, the applicability of genome skimming across divergent clades and as a broad-based phylogenomic tool is likely to remain circumscribed (but see Zhang et al. 2019a).
Restriction site-associated DNA sequencing (RAD-seq)
RAD-seq (Baird et al. 2008) and its predecessor, RAD marker identification and analysis (Miller et al. 2007), are traditionally used as tools for genotyping and discovering variable genetic markers. Generally, this approach utilises specific restriction enzymes to digest genomic DNA, and the flanking regions of each restriction site are used to identify homologous loci bioinformatically with software packages such as Stacks (Catchen et al. 2013; Rochette et al. 2019) and PyRAD (Eaton 2014). Alignments of homologous loci enable the identification of single nucleotide polymorphisms (SNPs), which can be used to infer population structure and phylogeny. Analyses of RAD-seq data for phylogenomics are typically carried out with one of two methods: in the form of concatenated supermatrix analysis (Iguchi et al. 2019) or species tree approaches (Herrera and Shank 2016; Johnston et al. 2017), although there are alternatives such as the assembly- and alignment-free method (Fan et al. 2017). Detailed laboratory and experimental protocols, along with variations of RAD-seq, have been covered thoroughly in a review by Andrews et al. (2016).
First tested on Nematostella (Actiniaria) by Reitzel et al. (2013), RAD-seq has since proved to be an invaluable tool for resolving the taxonomy, phylogeny and population structure of anthozoans. The benefits of RAD-seq in phylogenomics stem from several facets: the relatively low investment required in time and cost; applicability for non-model taxa without a reference genome; and suitability for resolving both relatively deep and shallow divergences (Peterson et al. 2012; Toonen et al. 2013; Eaton et al. 2017). There are several variations of RAD-seq commonly used in anthozoans, varying mainly in the number of restriction enzymes used and the sequencing approach. They are: (1) conventional RAD-seq using a single restriction enzyme and single-end reads in Nematostella (Reitzel et al. 2013) and Chrysogorgia (order Alcyonacea) (Pante et al. 2015); (2) RAD-PE with paired-end sequencing of RAD fragments (Etter et al. 2011) in Pocillopora (Scleractinia) (Combosch and Vollmer 2015) and Anemonia viridis (Actiniaria) (Porro et al. 2020); (3) double-digest RAD-seq with two restriction enzymes (Peterson et al. 2012), which have been used on Bartholomea annulata (Actiniaria) (Titus et al. 2019a); (4) DArTSeq, a genome complexity reduction method similar to RAD-seq as tested in Acropora (Scleractinia) (Rosser et al. 2017) and Pocillopora (Smith et al. 2017); and (5) ezRAD (Toonen et al. 2013), a popular method which uses either a single enzyme or a combination of enzymes in non-model organisms, as applied on Porites (Forsman et al. 2017), Montipora (Scleractinia) (Cunha et al. 2019) and Ovabunda (Alcyonacea) (McFadden et al. 2017), for example.
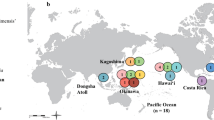
At relatively shallow divergences (< 100 Ma), RAD-seq is a powerful tool that is able to reconstruct robust phylogenies and characterise cryptic species. For example, the species boundaries and phylogeny of the scleractinian genera Pocillopora and Leptastrea have been clarified considerably by Johnston et al. (2017) and Arrigoni et al. (2020), respectively. Herrera and Shank (2016) tested a species delimitation hypothesis with RAD-seq phylogeny for the morphologically enigmatic octocoral Paragorgia (Alcyonacea). In resolving specific cryptic species problems, Iguchi et al. (2019) demonstrated that the blue coral Heliopora coerulea (order Helioporacea) around Okinawa actually comprises two distinct species; and Quattrini et al. (2019) not only discovered several cryptic species within the genus Sinularia (Alcyonacea) at Dongsha Atoll, they also highlighted the importance of hybrid speciation in the evolution of anthozoans (see also Mao 2020).
As with other reduced-representation sequencing techniques (e.g. hybrid capture; Allio et al. 2020), certain genomic features such as mitochondrial genes or genomes can be extracted and assembled (Table 1). Consequently, RAD-seq data have been used to assemble mitogenomes for several Porites (Scleractinia) species to date (Tisthammer et al. 2016; Teranneo et al. 2018a, 2018b). Similarly, Capel et al. (2020) were able to skim mitochondrial and rRNA genes from RAD data, thereafter describing the genus Atlantia (Scleractinia). Beyond phylogeny reconstruction, RAD-seq data analyses guided Pratlong et al. (2017b) to propose an XX/XY sex-determination system in Corallium rubrum (Alcyonacea). Such data were also used to investigate DNA methylation in Porites astreoides resulting from environmental changes (Dimond and Roberts 2020). While Dimond and Roberts (2020) were able to annotate several genes experiencing changes in DNA methylation involved in cell signalling and proliferation, mRNA splicing and apoptosis, they lamented the lack of well-annotated genomes for further functional inferences in other methylated loci (see Neri et al. 2017).
Despite the increasing popularity of RAD-seq in anthozoan phylogenomics (Fig. 2a), its limitations are apparent for deep divergences. In particular, mutations at the restriction sites accumulate with evolutionary time, resulting in an increase in missing data the more evolutionary divergent the focal species are. More importantly, this pattern reduces the ability to infer orthology in RAD loci based on sequence similarity (Rubin et al. 2012). Recent studies (e.g. Herrera and Shank 2016; Eaton et al. 2017) have demonstrated that RAD-seq is able to infer robust phylogenies for divergences between 50 and 80 Ma, but given that the origin of Anthozoa is estimated to be ~ 800 Ma (Quattrini et al. 2020; McFadden et al. 2021), the applicability of RAD-seq for broad-scale phylogenies is limited. Therefore, the use of RAD-seq will likely remain confined to phylogenetic analyses of closely-related species.
Reductively-amplified DNA sequencing (ReAD-seq)
Leveraging the principles of RAD-seq, reductively-amplified DNA sequencing (ReAD-seq) circumvents the need for restriction enzyme digestion by substituting it with PCR-based reactions. Principally, two types of ReAD-seq are employed: Nextera-tagmented reductively-amplified DNA (NextRAD) (Russello et al. 2015) and multiplexed inter-simple sequence repeat (ISSR) genotyping (MIG-seq) (Suyama and Matsuki 2015). In the former, selective primers are used to amplify consistent loci across samples, whereas the latter amplifies anonymous non-repetitive regions using inter-simple sequence repeat (ISSR) primers which anchor on simple sequence repeats (SSRs) (e.g. microsatellites). Similar to RAD-seq, homologous loci are identified, with subsequent genotyping and SNP identification. Applications of this method include cryptic species identification and population genetics.
Despite the high efficiency and utility, the popularity of ReAD-seq pales in comparison with RAD-seq for anthozoan phylogenomics (Fig. 2). Nevertheless, a recent NextRAD study by Bongaerts et al. (2020; see also Bongaerts et al. 2017) clearly demonstrated that, contrary to popular belief, the widespread Indo-Pacific coral Pachyseris speciosa (Scleractinia) is an ancient cryptic species complex with three morphologically indistinguishable lineages. Using MIG-seq, Richards et al. (2018) described a blue coral species Heliopora hiberniana (Helioporacea) from north Western Australia as distinct from the widespread H. coerulea. Despite identical cytochrome c oxidase subunit I (COI) and mutS sequences among all individuals analysed, principal coordinate analysis (PCoA) and STRUCTURE analysis (Pritchard et al. 2000) on MIG-seq SNPs confirmed the two distinct lineages, H. coerulea and H. hiberniana, which were also morphologically distinguishable. Similarly, Pleurocorallium elatius and P. konojoi (Alcyonacea) from Japan were inseparable with two mitochondrial loci, but both morphology and MIG-seq data clearly delineated the two species with almost no demonstrable gene flow (Takata et al. 2019, see also Tu et al. 2015). Notably, these MIG-seq studies did not reconstruct a phylogeny and have been excluded from Fig. 2 and Table 1.
For deeper divergences, the power of both RAD- and ReAD-seq trails off precipitously (see above), so we now turn our attention to the more versatile phylogenomic tools: phylotranscriptomics, hybrid capture and whole genome sequencing.
Phylotranscriptomics
The transcriptome of an organism refers to all RNA transcripts of both coding and non-coding regions. Broadly, data generated from RNA sequencing (or RNA-seq) are typically used for investigating developmental biology (Okubo et al. 2016; Rentzsch and Technau 2016), phylotranscriptomics (Lin et al. 2016; Quek and Huang 2019; Richards et al. 2020), gene evolution (Gacesa et al. 2015; Huang et al. 2016a; Lin et al. 2017a) and comparative genomics (Bhattacharya et al. 2016). Similar to the aforementioned methods, phylotranscriptomics bypasses the need for assembled genome data, as single-copy orthologous genes are identified bioinformatically following sequencing and assembly. Phylogenetic reconstructions tend to be equivalent to inferences made from whole genome sequencing but can be achieved at a fraction of the cost (see Cheon et al. 2020).
Phylotranscriptomics is an ideal tool for analysing the phylogeny of early-diverging metazoans such as sponges (Simion et al. 2017; Feuda et al. 2017), ctenophores (Whelan et al. 2015; Borowiec et al. 2015) and cnidarians (Zapata et al. 2015; Kayal et al. 2018). In Anthozoa, phylotranscriptomics has clarified perplexing phylogenies reconstructed from mitochondrial data. In particular, phylotranscriptomics cemented the monophyly of both Anthozoa (Zapata et al. 2015; Pratlong et al. 2017a; Kayal et al. 2018) and Scleractinia (Lin et al. 2016). To date, the largest phylotranscriptomic study at the order level included only 44 scleractinians (Quek and Huang 2019), despite an ever-increasing number of transcriptomes available. This approach also remains underutilised for shallower timescales. A recent paper on Acroporidae (Scleractinia) determined that Alveopora is sister to Montipora rather than Astreopora (Richards et al. 2020; see Ryu et al. 2019a), but such studies are few and far between.
We note that there has been much research on the transcriptomes of anthozoans such as sea anemones (Urbarova et al. 2012; Mitchell et al. 2020), scleractinians (Pootakham et al. 2018; Ryu et al. 2019a), octocorals (Ryu et al. 2019b) and zoantharians (order Zoantharia; Huang et al. 2016a, 2017), but with specific objectives other than phylogeny reconstruction. Primarily, these studies focused on identifying toxins and bioactive compounds, testing gene responses under a variety of environmental stressors and reconstructing gene evolution. Consequently, there are multiple well-annotated transcriptomes and associated databases (Liew et al. 2016; Zhang et al. 2019b) that serve as foundations for subsequent phylotranscriptomic and gene evolution studies.
Unlike with RAD-seq where anonymous loci are sequenced, annotation of gene transcripts opens up opportunities for investigating gene and genome evolution (Bhattacharya et al. 2016; Lin et al. 2017a; Koch and Grimmelikhuijzen 2019). When genetic changes are traced on a robust phylogeny, specific hypotheses about the evolutionary history of anthozoans can be tested. For example, Bessho-Uehara et al. (2020) demonstrated that the last common ancestor of all octocorals was likely bioluminescent. In the same vein, the origin and evolution of the aragonite skeleton in scleractinians are of particular interest for reef-building corals (Quattrini et al. 2020). Bhattacharya et al. (2016) examined the transcriptomes of 20 different stony corals and identified genes involved in accretion of the coral skeleton, whereas Lin et al. (2017a) focused primarily on differences in carbonic anhydrases between Scleractinia and Corallimorpharia, concluding that the greater number of scleractinian genes involved in secreted and membrane-associated carbonic anhydrases were critical for the evolution of calcification. Threatened by accelerating climate change, more comparative genomic studies across broader phylogenetic scales (e.g. Guzman et al. 2018; Conci et al. 2019; Quattrini et al. 2020) will be needed to determine the fate of reef ecosystems.
While phylotranscriptomic data are cheaper to obtain and more readily available than whole genome data, there are some drawbacks. Crucially, due to the instability of RNA, samples have to be carefully handled and stored prior to extraction (Seelenfreund et al. 2014). This constraint excludes most museum samples and increases the cost of sample preservation. Furthermore, due to uneven expression levels among genes and differences in sequencing depth, missing data would be expected among orthologs. As a result, orthologs may be misidentified by algorithms that use gene-tree-free methods (Cheon et al. 2020). Missing data and data type (i.e. amino acid vs. DNA) may also impact phylogenetic inference, although Quek and Huang (2019) found limited effects of missing data on tree topologies inferred for scleractinians, instead emphasising the importance of meticulously checking for irregularities among gene trees (see also Xi et al. 2016). The authors also noted that DNA sequences are superior to amino acid sequences in achieving higher branch supports and congruence among loci (Quek and Huang 2019; see also Ying et al. 2018). To overcome some of the limitations with phylotranscriptomic analysis, more studies are beginning to opt for hybrid capture instead.
Hybrid capture and target enrichment sequencing
A relatively novel approach, hybrid capture coupled with target enrichment has been gaining traction in phylogenomic studies across the tree of life (Faircloth et al. 2012; Lemmon et al. 2012; Karin et al. 2020). Briefly, specific DNA or RNA ‘probes’ (also known as ‘baits’) that are complementary to target loci are designed which are then hybridised to DNA libraries. Orthologous targets are typically conserved regions of DNA, with variable flanking regions that may be phylogenetically informative. Unbound DNA is then discarded, and the targeted loci are enriched and sequenced accordingly. Hybrid capture has been used in identification of infectious diseases (Gaudin and Desnues 2018), assembling genomes from ancient remains and museum samples (Gasc et al. 2016), and phylogenomics (for a review, see Andermann et al. 2020). This approach is particularly versatile as researchers have the flexibility to design probes based on the intended level of taxonomic inclusivity. Hybrid capture baits have been used for a number of marine taxa based on transcriptomes, genomes and RAD data, including ophiuroids (Hugall et al. 2016), mobulids (White et al. 2017) and zooplankton (Choquet et al. 2019), respectively.
There are currently three sets of anthozoan baits designed for different taxonomic levels and groups: Hexacorallia (Cowman et al. 2020; see also Quattrini et al. 2018), Octocorallia (Erickson et al. 2020; see also Quattrini et al. 2018) and Scleractinia (Quek et al. 2020). The former two consist of a combined set of loci targeting all anthozoans, optimised from Quattrini et al. (2018) based on both ultraconserved elements (UCEs) and exons as references, whereas the latter design is based on exons. These bait sets have shown great promise in phylogenomic reconstructions. Quattrini et al. (2018) first demonstrated the feasibility of their baits across almost all major clades of anthozoans (n = 33). A greatly expanded sampling in Quattrini et al. (2020), sequenced using two bait sets (Quattrini et al. 2018; Cowman et al. 2020), resulted in a robust, time-calibrated phylogeny of 234 taxa from all anthozoan orders. This study found that the evolution of skeletal traits and diversification rates of anthozoans closely mirrored that of palaeoclimatic changes through geologic time. For example, high levels of carbon dioxide in the atmosphere during the early to mid-Paleozoic likely resulted in ocean acidification that impacted aragonitic skeleton production among scleractinians but drove the cladogenesis of actiniarians, ceriantharians and zoantharians. Accordingly, it has been suggested that contemporary climate change and ocean acidification could lead to the replacement of reef-building corals by their more resilient octocoral relatives (Quattrini et al. 2020), as exemplified by contemporary ecological trends (Inoue et al. 2013; Tsounis and Edmunds 2017).
Hybrid capture has further demonstrated applicability on more recent anthozoan divergences. Both Cowman et al. (2020) and Quek et al. (2020) tested their bait sets on scleractinians, with the former focusing on Acroporidae, Agariciidae and Fungiidae, and the latter on a broad sampling across 12 different families. Notably, Quek et al. (2020) highlighted the impact of sequence specificity on bait design, as they captured considerably fewer loci from sister clade Corallimorpharia compared to Scleractinia. At the genus level, Erickson et al. (2020) utilised hybrid capture to study species delimitation and population genomics in soft corals Alcyonium and Sinularia (Alcyonacea). Leveraging variable sequences that flank UCEs or exons (i.e. introns), their analysis broadly corroborated the RAD-seq phylogeny of Sinularia in Quattrini et al. (2019) while resolving the phylogeny and population genomics of the two genera.
Despite the manifold advantages of hybrid capture-based phylogenomics, such as its affordability, applicability for old museum samples and capacity to capture homologous loci at various taxonomic scales (see McKain et al. 2018), it still falls short compared to whole genome methods. For example, bait design and downstream analyses can be confounded by paralogy stemming from whole genome duplications such as in Acropora (Scleractinia) (Mao and Satoh 2019). Furthermore, genome characterisation may generate broader insights beyond phylogenetics and population genomics, such as the evolution of body plans (Putnam et al. 2007; DuBuc et al. 2018), novel metabolic pathways and gene evolution (Ying et al. 2018; Cunning et al. 2018; Surm et al. 2019), sexual reproduction (Wilding et al. 2020), eco-evolutionary processes (Ledoux et al. 2020) and adaptive radiation (Mao et al. 2018). Therefore, whole genome analysis is likely the ultimate frontier for advancing phylogenomic research.
Whole genomes
The undertaking of sequencing a genome is no small feat. Challenges can arise from cross-contamination by endosymbionts, difficulties in genome assembly and annotation as well as the overall investment required (Dominguez Del Angel et al. 2018). The genomes of many anthozoans are particularly difficult to assemble due to cross-contamination from a mixture of Symbiodiniaceae and other microorganisms (Artamonova and Mushegian 2013). To reduce non-target sequence load, DNA can be isolated from gametes (Shinzato et al. 2011, 2020; Robbins et al. 2019), symbionts separated prior to DNA extraction (Cunning et al. 2018) or non-target reads bioinformatically filtered post-sequencing (Buitrago-López et al. 2020).
Presently, whole genomes of anthozoans are still sorely lacking as compared to several other metazoan groups, which have researchers aiming for 5000 to 10,000 genomes (Zhang 2015; i5K Consortium 2013; Fan et al. 2020). In stark contrast, one of the largest anthozoan genome sequencing projects underway involves the sequencing of only 10 scleractinians (ReFuGe 2020 Consortium 2015). Nevertheless, there has been substantial progress since the emergence of the first sea anemone genome (Putnam et al. 2007), with a recent bumper crop of 18 Acropora (Scleractinia) genomes assembled within a single study (Shinzato et al. 2020). Unlike hybrid capture studies where baits can be designed across different scales of divergence, whole genomes are taxon-specific. As a result, there exists an imbalance of whole genomes assembled among the major anthozoan clades. There are now sequenced genomes for scleractinians, actiniarians, corallimorpharians and two octocoral classes Alcyonacea and Pennatulacea (Jiang et al. 2019; Ledoux et al. 2020; Table 1), but not for antipatharians, ceriantharians, helioporaceans and zoantharians.
Similar to phylotranscriptomics, genomic phylogenetic analysis is typically based on ortholog assignments of protein-coding genes (but see Cunning et al. 2018; Sims et al. 2009), which enable combining of transcriptomic and genomic data for phylogeny reconstruction (e.g. Lin et al. 2016; Kayal et al. 2018; Richards et al. 2020). Genome assemblies can generate critical insight into gene and genome evolution. For instance, Ying et al. (2018) identified an ancestral, complete fungal-like histidine biosynthesis pathway that is present only in “Robust” corals, but not in any other metazoan. In the “Complex” scleractinian Acropora, whole genome analyses have suggested that there was likely a whole genome duplication event that occurred ~ 31 Ma in the most recent common ancestor of the genus (Mao and Satoh 2019), and the contemporary diversity likely resulted from introgression and adaptations to past climate conditions (Mao et al. 2018; Shinzato et al. 2020).
Fundamentally, genome-based phylogenies can be applied across nearly all taxonomic levels (Kapli et al. 2020). However, we emphasise that whole genomes do not necessarily lead to more accurate inferences. For example, the earliest branch of metazoans was the subject of great contention even with analyses of large phylogenomic supermatrices (reviewed in Philippe et al. 2017). Rather than maximising the number of loci analysed, it is critical to ensure that taxon sampling is sufficiently broad (Philippe et al. 2011), robust model selection parameters are adopted (Simon et al. 2017; but see Abadi et al. 2019 cf. Gerth 2019), accurate orthologs are inferred (Cheon et al. 2020), and other systematic errors are avoided (see Philippe et al. 2017; Young and Gillung 2019). Unfortunately, whole genome assemblies in the majority of anthozoans remain a distant possibility due to the astronomical costs and challenges of such an undertaking. All of the alternatives reviewed above are therefore likely to be instrumental for anthozoan phylogenetics in the near future.
Considerations in anthozoan phylogenomics
The development of HTS technologies over the last decade has been mirrored by the uptake of genomic approaches for phylogenetic studies (see Fig. 2). The scale of analysis has also increased as a result. Genome skimming is the most popular method to date, but downstream phylogenetic analyses have largely relied on mitochondrial genes (Fig. 2a; Table 1). In contrast, other reduced-representation approaches that are increasingly being used can include hundreds to thousands of loci, with some methods allowing for the merging of loci for phylogeny reconstruction (e.g. Bossert et al. 2019). This trend towards greater gene sampling will continue with the increasing availability of newer sequencing technologies, particularly the third-generation sequencing platforms produced by Oxford Nanopore Technologies and Pacific Biosciences (PacBio). The nanopore and single-molecule real-time (SMRT) sequencing technologies generate longer reads, offering a number of improvements over short-read sequencing, albeit with lower accuracy (Pollard et al. 2018; Burgess 2018). Studies adopting either of these methods thus typically include Illumina sequencing for error correction to generate hybrid assemblies (Jeon et al. 2019; Jiang et al. 2019; Shumaker et al. 2019; Shinzato et al. 2020). Nevertheless, long-read sequencing remains expensive relative to their output and require computationally intensive analyses (for a review, see Amarasinghe et al. 2020).
Hybrid assemblies are the gold standard for reference assemblies and downstream phylogenomic analyses (Philippe et al. 2005; Burgess 2018). Current short-read sequencing technologies face some restrictions, including missing genes in the final assembled genomes (Alkan et al. 2011), technical and computational challenges in de novo assemblies owing to repetitive sequences and high likelihood of misassemblies (for a review, see Liao et al. 2019). Accurate annotations of well-assembled genomes—currently limited for Anthozoa (Table 1; Artamonova and Mushegian 2013; Shinzato et al. 2020)—are also necessary for numerous downstream phylogenomic analyses. For example, they enable cross-validation of sequence similarity-based read filtering which is helpful for removing endosymbiont sequences (e.g. Forsman et al. 2020). Furthermore, adaptive loci under selection sequenced via reduced-representation methods can be identified based on genome annotations and excluded from phylogenomic studies or separately analysed for their associated functions (Titus et al. 2019a; Forsman et al. 2020; see also Ahrens et al. 2017).
Despite the best efforts of research teams around the world, whole genome assemblies for numerous anthozoans remain unlikely even in the next few decades. Realistically, hybrid capture and target enrichment sequencing may hold the greatest promise for building a comprehensive anthozoan tree of life. The flexibility of the approach, combined with the applicability across different scales of divergence, makes it a forerunner among all reduced-representation methods (Faircloth et al. 2012; Lemmon et al. 2012; Andermann et al. 2019; Erickson et al. 2020). However, there are currently only three available anthozoan bait sets (see section on ‘Hybrid capture and target enrichment sequencing’ above), and the baits designed by Quek et al. (2020) for Scleractinia could not capture a large proportion of targeted loci for the sister clade of Corallimorpharia. To this end, we encourage the development of more taxon-specific baits as more targeted baits would result in greater evenness in coverage and higher capture efficacy (Andermann et al. 2020). Where different bait sets are used, the results can be weaved together into a single supertree (e.g. Kimball et al. 2019). Alternatively, multiple bait sets can be combined and used for hybrid capture experiments, and orthologous loci identified post-capture alongside genome and transcriptome data (e.g. Bossert et al. 2019). We expect more bait sets to emerge in the near future, each differing in its targeted taxonomic group, but all contributing towards a comprehensive, robust phylogeny of anthozoans.
Genome-wide data do not shield phylogenomic analyses from incongruence (see Philippe et al. 2011, 2017; Jeffroy et al. 2006; Roure et al. 2013; Lambert et al. 2015). For instance, erroneous phylogenetic inferences can arise from misalignments, errors in ortholog assignments or violations of model assumptions. Furthermore, incongruent gene trees due to incomplete lineage sorting and cross-contamination will affect downstream analyses. At the sequence level, substitution saturation, inappropriate evolutionary model assignments and, to a limited extent, missing data all have negative impacts on phylogenetic accuracy. One key consideration for anthozoan phylogenetic research is introgression, which is an important process in the evolution of scleractinian corals and sea anemones but can confound typical tree reconstruction methods (Mao et al. 2018; Cunha et al. 2019; Porro et al. 2019; Quattrini et al. 2019; Bongaerts et al. 2020). Specifically, introgressive hybridisation gives rise to a reticulated pattern of evolution that cannot be captured in the typical bifurcating model (Knowles 2009; Townsend et al. 2011; Lambert et al. 2015; Tonini et al. 2015; Quattrini et al. 2019). Phylogenetic network approaches can help to represent this pattern (Huson and Bryant 2006; Morrison 2014; Solís-Lemus et al. 2017), but introgression events first need to be characterised with genomic data (see Morrison 2014). More generally, better understanding recent gene flow and introgressive hybridisation would clarify species boundaries and improve phylogenetic estimates for closely-related species (Mao et al. 2018; Quattrini et al. 2019; Erickson et al. 2020).
We acknowledge that phylogenomic analyses can seem daunting, especially for the inexperienced. Fortunately, there are a number of software programs designed to ease the entire reads-to-phylogeny workflow for a variety of sequencing approaches. For example, RADIS (Cruaud et al. 2016) automates the analysis of RAD-seq data from raw sequences to phylogenetic inference, and ParGenes (Morel et al. 2019; Darriba et al. 2020) performs model selection and gene/species tree reconstruction from multiple sequence alignments. Similarly, TREEasy (Mao et al. 2020) can automate the processing of sequence alignments to build a phylogenetic network or infer the species tree given a set of FASTA files, each corresponding to an orthologous locus. The degree to which each procedure in the reads-to-phylogeny process is put through these tools depends on the scope, method and expected output of each study, as well as the experience level of the user. More experienced researchers may opt for running each step separately in order to fine-tune specific parameters or compare different analyses. Overall, we recommend that, as far as possible, phylogenomic studies should apply multiple optimality criteria (e.g. maximum likelihood, Bayesian and species tree) to ensure congruence and accuracy of the inferred phylogeny (Table 2) (Mao et al. 2018; Arrigoni et al. 2020; Quek et al. 2020; Quattrini et al. 2020; Shinzato et al. 2020).
Future of anthozoan phylogenomics
Phylogenomics has become an important tool for guiding and supporting research across a wide range of biological phenomena. Indeed, well-sampled and robust phylogenomic trees are powerful for addressing complex evolutionary questions. As anthropogenic climate change is currently driving the “sixth mass extinction” (Barnosky et al. 2011; Ceballos et al. 2015), with marine taxa not spared from this seemingly inexorable decimation (Carpenter et al. 2008; McGill et al. 2015; Johnson et al. 2017; Luypaert et al. 2019), studies focusing on gene evolution, diversification trends, biogeography and trait evolution based on a reliable time-calibrated phylogeny are crucial for shaping and adapting conservation strategies (Drake et al. 2014; Bhattacharyra et al. 2016; Mouillot et al. 2016; Hartmann et al. 2017; Kayal et al. 2018; Mao et al. 2018; Palmer and Traylor-Knowles 2018; Huang et al. 2017; Dishon et al. 2020; Quattrini et al. 2020).
Overall, phylogenomic approaches are revolutionising our understanding of anthozoan evolution, driven by sustained sequencing efforts to churn out unprecedented amounts of data for more species. However, there is clearly an imbalance in taxon coverage among anthozoan clades, with groups such as ceriantharians and zoantharians remaining under-sampled for phylotranscriptomic, hybrid capture and whole genome studies (Fig. 2b). It is imperative that while efforts on actiniarians and scleractinians continue, more attention should be placed on these overlooked taxa which are consequently more uncertain phylogenetically (Kushida and Reimer 2019; Stampar et al. 2019; Xiao et al. 2019; Mejia et al. 2020; Poliseno et al. 2020). Furthermore, while whole genome assemblies are expected to result in the most robust trees, reduced-representation techniques such genome skimming, RAD-seq and hybrid capture should continue to be adopted given the research objectives and resources at hand. Finally, we caution that as phylogenomics becomes increasingly accessible both financially and bioinformatically, tried and tested methods that are appropriate for addressing specific biological questions should not be cast aside. Phylogenetic research using genomic data remains well complemented by established approaches focusing on the morphology, physiology, behaviour and ecology of species (Valdecasas et al. 2007; Padial and De la Riva 2010; Padial et al. 2010).
References
Abadi S, Azouri D, Pupko T, Mayrose I (2019) Model selection may not be a mandatory step for phylogeny reconstruction. Nat Commun 10:934
Aberer AJ, Kobert K, Stamatakis A (2014) ExaBayes: massively parallel Bayesian tree inference for the whole-genome era. Mol Biol Evol 31:2553–2556
Ahrens CW, Rymer PD, Stow A, Bragg J, Dillon S, Umbers KDL, Dudaniec RY (2018) The search for loci under selection: trends, biases and progress. Mol Ecol 27:1342–1356
Alkan C, Sajjadian S, Epichler EE (2011) Limitations of next-generation genome sequence assembly. Nat Methods 8:61–65
Allio R, Schomaker-Bastos A, Romiguier J, Prosdocimi F, Nabholz B, Delsuc F (2020) MitoFinder: efficient automated large-scale extraction of mitogenomic data in target enrichment phylogenomics. Mol Ecol Res 20:892–905
Amarasinghe SL, Su S, Dong X, Zappia L, Ritchie ME, Gouil Q (2020) Opportunities and challenges in long-read sequencing data analysis. Genome Biol 21:30
Ament-Velásquez SL, Breedy O, Cortés J, Guzman HM, Wörheide G, Vargas S (2016) Homoplasious colony morphology and mito-nuclear phylogenetic discordance among Eastern Pacific octocorals. Mol Phylogenet Evol 98:373–381
Andermann T, Fernandes AM, Olsson U, Töpel M, Pfeil B, Oxelman B, Aleixo A, Faircloth BC, Antonelli A (2019) Allele phasing greatly improves the phylogenetic utility of ultraconserved elements. Syst Biol 68:32–46
Andermann T, Torres Jiménez MF, Matos-Maraví P, Batista R, Blanco-Pastor JL, Gustafsson ALS, Kistler L, Liberal IM, Oxelman B, Bacon CD, Antonelli A (2020) A guide to carrying out a phylogenomic target sequence capture project. Front Genet 10:1407
Andrews K, Good J, Miller M, Luikart G, Hohenlohe PA (2016) Harnessing the power of RADseq for ecological and evolutionary genomics. Nat Rev Genet 17:81–92
Arrigoni R, Berumen ML, Mariappan KG, Beck PSA, Hulver AM, Montano S, Pichon M, Strona G, Terraneo TI, Benzoni F (2020) Towards a rigorous species delimitation framework for scleractinian corals based on RAD sequencing: the case study of Leptastrea from the Indo-Pacific. Coral Reefs 39:1001–1025
Artamonova II, Mushegian AR (2013) Genome sequence analysis indicates that the model eukaryote Nematostella vectensis harbors bacterial consorts. Appl Environ Microbiol 79:6868–6873
Baird NA, Etter PD, Atwood TS, Currey MC, Shiver AL, Lewis ZA, Selker EU, Cresko WA, Johnson EA (2008) Rapid SNP discovery and genetic mapping using sequenced RAD markers. PLoS ONE 3:e3376
Balbar AC, Metaxas A (2019) The current application of ecological connectivity in the design of marine protected areas. Glob Ecol Conserv 17:e00569
Barnosky AD, Matzke N, Tomiya S, Wogan GOU, Swartz B, Quental TB, Marshall C, McGuire JL, Lindsey EL, Maguire KC, Mersey B, Ferrer EA (2011) Has the Earth’s sixth mass extinction already arrived? Nature 471:51–57
Barrett NJ, Hogan RI, Allcock AL, Molodtsova T, Hopkins K, Wheeler AJ, Yesson C (2020) Phylogenetics and mitogenome organisation in black corals (Anthozoa: Hexacorallia: Antipatharia): an order-wide survey inferred from complete mitochondrial genomes. Front Mar Sci 7:440
Barshis DJ, Ladner JT, Oliver TA, Seneca FO, Traylor-Knowles N, Palumbi SR (2013) Genomic basis for coral resilience to climate change. Proc Natl Acad Sci USA 110:1387–1392
Baumgarten S, Simakov O, Esherick LY, Liew YJ, Lehnert EM, Michell CT, Li Y, Hambleton EA, Guse A, Oates ME, Gough J, Weis VM, Aranda M, Pringle JR, Voolstra CR (2015) The genome of Aiptasia, a sea anemone model for coral symbiosis. Proc Natl Acad Sci USA 112:11893–11898
Bellis ES, Howe DK, Denver DR (2016) Genome-wide polymorphism and signatures of selection in the symbiotic sea anemone Aiptasia. BMC Genom 17:160
Berntson EA, France SC, Mullineaux LS (1999) Phylogenetic relationships within the class Anthozoa (phylum Cnidaria) based on nuclear 18S rDNA sequences. Mol Phylogenet Evol 13:417–433
Bessho-Uehara M, Francis WR, Haddock SHD (2020) Biochemical characterization of diverse deep-sea anthozoan bioluminescence systems. Mar Biol 167:114
Bhattacharya D, Agrawal S, Aranda M, Baumgarten S, Belcaid M, Drake JL, Erwin D, Forêt S, Gates RD, Gruber DF, Kamel B, Lesser MP, Levy O, Liew YJ, MacManes M, Mass T, Medina M, Mehr S, Meyer E, Price DC, Putnam HM, Qiu H, Shinzato C, Shoguchi E, Stokes AJ, Tambutté S, Tchernov D, Voolstra CR, Wagner N, Walker CW, Weber APM, Weis V, Zelzion E, Zoccola D, Falkowski PG (2016) Comparative genomics explains the evolutionary success of reef-forming corals. eLife 5:e13288
Bilewitch JP, Degnan SM (2011) A unique horizontal gene transfer event has provided the octocoral mitochondrial genome with an active mismatch repair gene that has potential for an unusual self-contained function. BMC Evol Biol 11:228
Bongaerts P, Riginos C, Brunner R, Englebert N, Smith SR, Hoegh-Guldberg O (2017) Deep reefs are not universal refuges: reseeding potential varies among coral species. Sci Adv 3:e1602373
Bongaerts P, Cooke IR, Ying H, Wels D, Haan den S, Hernandez-Agreda A, Brunner CA, Dove S, Englebert N, Eyal G, Forêt S, Grinblat M, Hay KB, Harii S, Hayward DC, Lin Y, Mihaljević M, Moya A, Muir P, Sinniger F, Smallhorn-West P, Torda G, Ragan MA, van Oppen MJH, Hoegh-Guldberg O (2020) Cryptic diversity masks ecologically distinct coral species on tropical reefs. bioRxiv doi: https://doi.org/10.1101/2020.09.04.260208
Bongiorni L, Mea M, Gambi C, Pusceddu A, Taviani M, Danovaro R (2010) Deep-water scleractinian corals promote higher biodiversity in deep-sea meiofaunal assemblages along continental margins. Biol Conserv 143:1687–1700
Borowiec ML, Lee EK, Chiu JC, Plachetzki DC (2015) Extracting phylogenetic signal and accounting for bias in whole-genome data sets supports the Ctenophora as sister to remaining Metazoa. BMC Genom 16:987
Bossert S, Danforth BN (2018) On the universality of target-enrichment baits for phylogenomic research. Methods Ecol Evol 9:1453–1460
Bossert S, Murray EA, Almeida EAB, Brady SG, Blaimer BB, Danforth BN (2019) Combining transcriptomes and ultraconserved elements to illuminate the phylogeny of Apidae. Mol Phylogenet Evol 130:121–131
Bouckaert R, Vaughan TG, Barido-Sottani J, Duchêne S, Fourment M, Gavryushkina A, Heled J, Jones G, Kühnert D, De Maio N, Matschiner M, Mendes FK, Müller NF, Ogilvie HA, du Plessis L, Popinga A, Rambaut A, Rasmussen D, Siveroni I, Suchard MA, Wu C, Xie D, Zhang C, Stadler T, Drummond AJ (2019) BEAST 2.5: an advanced software platform for Bayesian evolutionary analysis. PLoS Comput Biol 15:e1006650
Bryant D, Bouckaert R, Felsenstein J, Rosenberg NA, RoyChoudhury A (2012) Inferring species trees directly from biallelic genetic markers: bypassing gene trees in a full coalescent analysis. Mol Biol Evol 29:1917–1932
Budd AF, Romano SL, Smith ND, Barbeitos MS (2010) Rethinking the phylogeny of scleractinian corals: a review of morphological and molecular data. Integr Comp Biol 50:411–427
Buitrago-López C, Mariappan KG, Cardenas A, Gegner HM, Voolstra CR (2020) The genome of the cauliflower coral Pocillopora verrucosa. Genome Biol Evol 12:1911–1017
Burgess DJ (2018) Next regeneration sequencing for reference genomes. Nat Rev Genet 19:125
Capel KCC, López C, Moltó-Martín I, Zilberberg C, Creed JC, Knapp ISS, Hernández M, Forsman ZH, Toonen RJ, Kitahara MV (2020) Atlantia, a new genus of Dendrophylliidae (Cnidaria, Anthozoa, Scleractinia) from the eastern Atlantic. PeerJ 8:e8633
Carpenter KE, Abrar M, Aeby G, Aronson RB, Banks S, Bruckner A, Chiriboga A, Cortés J, Delbeek JC, DeVantier L, Edgar GJ, Edwards AJ, Fenner D, Guzmán HM, Hoeksema BW, Hodgson G, Johan O, Licuanan WY, Livingstone SR, Lovell ER, Moore JA, Obura DO, Ochavillo D, Polidoro BA, Precht WF, Quibilan MC, Reboton C, Richards ZT, Rogers AD, Sanciangco J, Sheppard A, Sheppard C, Smith J, Stuart S, Turak E, Veron JEN, Wallace C, Weil E, Wood E (2008) One-third of reef-building corals face elevated extinction risk from climate change and local impacts. Science 321:560–563
Catchen J, Hohenlohe PA, Bassham S, Amores A, Cresko WA (2013) Stacks: an analysis tool set for population genomics. Mol Ecol 22:3124–3140
Ceballos G, Ehrlich PR, Barnosky AD, García A, Pringle RM, Palmer TM (2015) Accelerated modern human-induced species losses: entering the sixth mass extinction. Sci Adv 1:e1400253
Chen CA, Odorico DM, Tenlohuis M, Veron JEN, Miller DJ (1995) Systematic relationships within the Anthozoa (Cnidaria: Anthozoa) using the 5′-end of the 28S rDNA. Mol Phylogenet Evol 4:175–183
Chao C, Chou K, Huang C, Wen Z, Hsu C, Wu Y, Dai C, Sheu J (2012) Steroids from the soft coral Sinularia crassa. Mar Drugs 10:439–450
Chen IP, Tang CY, Chiou C, Hsu J, Wei NV, Wallace CC, Muir P, Wu H, Chen CA (2009) Comparative analyses of coding and noncoding DNA regions indicate that Acropora (Anthozoa: Scleractina) possesses a similar evolutionary tempo of nuclear vs. mitochondrial genomes as in plants. Mar Biotechnol 11:141
Cheon S, Zhang J, Park C (2020) Is phylotranscriptomics as reliable as phylogenomics? Mol Biol Evol 37(12):3672. https://doi.org/10.1093/molbev/msaa181
Chi SI, Johansen SD (2017) Zoantharian mitochondrial genomes contain unique complex group I introns and highly conserved intergenic regions. Gene 628:24–31
Chifman J, Kubatko L (2014) Quartet inference from SNP data under the coalescent. Bioinformatics 30:3317–3324
Choquet M, Smolina I, Dhanasiri AKS, Blanco-Bercial L, Kopp M, Jueterbock A, Sundaram AYM, Hoarau G (2019) Towards population genomics in non-model species with large genomes: a case study of the marine zooplankton Calanus finmarchicus. R Soc Open Sci 6:180608
Combosch DJ, Vollmer SV (2015) Trans-Pacific RAD-Seq population genomics confirms introgressive hybridization in Eastern Pacific Pocillopora corals. Mol Phylogenet Evol 88:154–162
Conci N, Wörheide G, Vargas S (2019) New non-bilaterian transcriptomes provide novel insights into the evolution of coral skeletomes. Genome Biol Evol 11:3068–3081
Cowman PF, Quattrini AM, Bridge TCL, Watkins-Colwell GJ, Fadli N, Grinblat M, Roberts TE, McFadden CS, Miller DJ, Baird AH (2020) An enhanced target-enrichment bait set for Hexacorallia provides phylogenomic resolution of the staghorn corals (Acroporidae) and close relatives. Mol Phyloget Evol 153:106944
Cronn R, Knaus BJ, Liston A, Maughan PJ, Parks M, Syring JV, Udall J (2012) Targeted enrichment strategies for next-generation plant biology. Am J Bot 99:291–311
Cruaud A, Gautier M, Rossi J, Rasplus J, Gouzy J (2016) RADIS: analysis of RAD-seq data for interspecific phylogeny. Bioinformatics 32:3027–3028
Cunha RL, Forsman ZH, Belderok R, Knapp ISS, Castilho R, Tooen RJ (2019) Rare coral under the genomic microscope: timing and relationships among Hawaiian Montipora. BMC Evol Biol 19:253
Cunning R, Bay RA, Gillette P, Baker AC, Traylor-Knowles N (2018) Comparative analysis of the Pocillopora damicornis genome highlights role of immune system in coral evolution. Sci Rep 8:16134
Cunning R, Silverstein RN, Barnes BB, Baker AC (2019) Extensive coral mortality and critical habitat loss following dredging and their association with remotely-sensed sediment plumes. Mar Pollut Bull 145:185–199
Daly M, Brugler MR, Cartwright P, Collins AG, Dawson MN, Fautin DG, France SC, McFadden CS, Opresko DM, Rodríguez E, Romano SL, Stake JL (2007) The phylum Cnidaria: a review of phylogenetic patterns and diversity 300 years after Linnaeus. Zootaxa 1668:1–766
Darriba D, Posada D, Kozlov AM, Stamatakis A, Morel B, Flouri T (2019) ModelTest-NG: a new and scalable tool for the selection of DNA and protein evolutionary models. Mol Biol Evol 37:291–294
Dimond JL, Roberts SB (2020) Convergence of DNA methylation profiles of the reef coral Porites astreoides in a novel environment. Front Mar Sci 6:792
Dey MM, Rosegrant MW, Gosh K, Chen OL, Valmonte-Santos R (2016) Analysis of the economic impact of climate change and climate change adaptation strategies for fisheries sector in Pacific coral triangle countries: model, estimation strategy, and baseline results. Mar Policy 67:156–163
Dishon G, Grossowicz M, Krom M, Guy G, Gruber DF, Tchernov D (2020) Evolutionary traits that enable scleractinian corals to survive mass extinction events. Sci Rep 10:3903
Dominguez Del Angel V, Hjerde E, Sterck L, Capella-Gutierrez S, Notredame C, Pettersson OV, Amselem J, Bouri L, Bocs S, Klopp C, Gibrat J, Vlasova A, Leskosek BL, Soler L, Binzer-Panchal M, Lantz H (2018) Ten steps to get started in genome assembly and annotation. F1000Res 7:148
Drake JL, Mass T, Falkowski PG (2014) The evolution and future of carbonate precipitation in marine invertebrates: witnessing extinction or documenting resilience in the Anthropocene? Elem Sci Anth 2:000026
Dubin A, Chi SI, Emblem A, Moum T, Johansen SD (2019) Deep-water sea anemone with a two-chromosome mitochondrial genome. Gene 692:195–200
DuBuc TQ, Stephenson TB, Rock AQ, Martindale MQ (2018) Hox and Wnt pattern the primary body axis of an anthozoan cnidarian before gastrulation. Nat Commun 9:2007
Easton EE, Hicks D (2018) Complete mitochondrial genome of Callogorgia cf. gracilis (Octocorallia: Calcaxonia: Primnoidae). Mitochondrial DNA B 4:361–362
Eaton DAR (2014) PyRAD: assembly of de novo RADseq loci for phylogenetic analyses. Bioinformatics 30:1844–1849
Eaton DAR, Spriggs EL, Park B, Donoghue MJ (2017) Misconceptions on missing data in RAD-seq phylogenetics with a deep-scale example from flowering plants. Syst Biol 66:399–412
Erickson KL, Pentico A, Quattrini AM, McFadden CS (2020) New approaches to species delimitation and population structure of anthozoans: two case studies of octocorals using ultraconserved elements and exons. Mol Ecol Res. https://doi.org/10.1111/1755-0998.13241
Etter PD, Preston JL, Bassham S, Cresko WA, Johnson EA (2011) Local de novo assembly of RAD paired-end contigs using short sequencing reads. PLoS ONE 6:e18561
Faircloth BC, McCormack JE, Crawford NG, Harvey MG, Brumfield RT, Glenn TC (2012) Ultraconserved elements anchor thousands of genetic markers spanning multiple evolutionary timescales. Syst Biol 61:717–726
Faith DP (1992) Conservation evaluation and phylogenetic diversity. Biol Conserv 61:1–10
Fan H, Ives AR, Surget-Groba Y, Cannon CH (2015) An assembly and alignment-free method of phylogeny reconstruction from next-generation sequencing data. BMC Genom 16:522
Fan G, Song Y, Yang L, Huang X, Zhang S, Zhang M, Yang X, Chang Y, Zhang H, Li Y, Liu S, Yu L, Chu J, Seim I, Feng C, Near TJ, Wing RA, Wang W, Wang K, Wang J, Xu X, Yang H, Liu X, Chen N, He S (2020) Initial data release and announcement of the 10,000 fish genomes project (Fish10K). GigaSci 9:giaa080
Felsenstein J (1989) PHYLIP — phylogeny inference package (Version 3.2). Cladistics 5:164–166
Feuda R, Dohrmann M, Pett W, Philippe H, Rota-Stabelli O, Lartillot N, Wörheide G, Pisani D (2017) Improved modeling of compositional heterogeneity supports sponges as sister to all other animals. Curr Biol 27:3864-3870.e4
Figueroa DF, Baco AR (2014) Complete mitochondrial genomes elucidate phylogenetic relationships of the deep-sea octocoral families Coralliidae and Paragorgiidae. Deep-Sea Res Pt II 99:83–91
Figueroa DF, Baco AR (2015) Octocoral mitochondrial genomes provide insights into the phylogenetic history of gene order rearrangements, order reversals, and cnidarian phylogenetics. Genome Biol Evol 7:391–409
Figueroa DF, Hicks D, Figueroa NJ (2019) The complete mitochondrial genome of Tanacetipathes thamnea Warner, 1981 (Antipatharia: Myriopathidae). Mitochondrial DNA B 4:4109–4110
Foox J, Brugler M, Siddall ME, Rodríguez E (2016) Multiplexed pyrosequencing of nine sea anemone (Cnidaria: Anthozoa: Hexacorallia: Actiniaria) mitochondrial genomes. Mitochondrial DNA A 27:2826–2832
Forsman ZH, Ritson-Williams R, Tisthammer KH, Knapp ISS, Toonen RJ (2020) Host-symbiont coevolution, cryptic structure, and bleaching susceptibility, in a coral species complex (Scleractinia; Poritidae). Sci Rep 10:16995
Forsman ZH, Knapp ISS, Tisthammer K, Eaton DAR, Belcaid M, Toonen RJ (2017) Coral hybridization or phenotypic variation? genomic data reveal gene flow between Porites lobata and P. compressa. Mol Phylogenet Evol 111:132–148
Fukami H, Budd AF, Paulay G, Solé-Cava A, Chen CA, Iwao K, Knowlton N (2004) Conventional taxonomy obscures deep divergence between Pacific and Atlantic corals. Nature 427:832–835
Gacesa R, Chung R, Dunn SR, Weston AJ, Jaimes-Becerra A, Marques AC, Morandini AC, Hranueli D, Starcevic A, Ward M, Long PF (2015) Gene duplications are extensive and contribute significantly to the toxic proteome of nematocysts isolated from Acropora digitifera (Cnidaria: Anthozoa: Scleractinia). BMC Genom 16:774
Galindo LJ, Torruella G, Moreira D, Eglit Y, Simpson AGB, Völcker E, Clauß S, López-García P (2019) Combined cultivation and single-cell approaches to the phylogenomics of nucleariid amoebae, close relatives of fungi. Phil Trans R Soc B 374:20190094
Gasc C, Peyretaillade E, Peyret P (2016) Sequence capture by hybridization to explore modern and ancient genomic diversity in model and nonmodel organisms. Nucleic Acids Res 44:4504–4518
Gaudin M, Desnues C (2018) Hybrid capture-based next generation sequencing and its application to human infectious diseases. Front Microbiol 9:2924
Gerth M (2019) Neglecting model selection alters phylogenetic inference. bioRxiv. doi: https://doi.org/10.1101/849018
Gouy M, Guindon S, Gascuel O (2010) SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol Biol Evol 27:221–224
Greenhalgh T, Peacock R (2005) Effectiveness and efficiency of search methods in systematic reviews of complex evidence: audit of primary sources. BMJ 331:1064
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321
Guzman C, Shinzato C, Lu T, Conaco C (2018) Transcriptome analysis of the reef-building octocoral, Heliopora coerulea. Sci Rep 8:8397
Hartmann AC, Baird AH, Knowlton N, Huang D (2017) The paradox of environmental symbiont acquisition in obligate mutualisms. Curr Biol 27:3711–3716
Herrera S, Shank TM (2016) RAD sequencing enables unprecedented phylogenetic resolution and objective species delimitation in recalcitrant divergent taxa. Mol Phylogenet Evol 100:70–79
Hogan IR, Hopkins K, Wheeler AJ, Allcock AL, Yesson C (2019) Novel diversity in mitochondrial genomes of deep-sea Pennatulacea (Cnidaria: Anthozoa: Octocorallia). Mitochondrial DNA A 30:764–777
Horowitz J, Brugler MR, Bridge TCL, Cowman PF (2020) Morphological and molecular description of a new genus and species of black coral (Cnidaria: Anthozoa: Hexacorallia: Antipatharia: Antipathidae: Blastopathes) from Papua New Guinea. Zootaxa 4821:553–569
Huang C, Morlighem JRL, Zhou H, Lima EP, Gomes PB, Cai J, Lou I, Pérez CD, Lee SM, Rádis-Baptista G (2016a) The transcriptome of the zoanthid Protopalythoa variabilis (Cnidaria, Anthozoa) predicts a basal repertoire of toxin-like and venom-auxiliary polypeptides. Genome Biol Evol 8:3045–3064
Huang C, Morlighem JRL, Cai J, Liao Q, Pérez CD, Gomes PB, Guo M, Rádis-Baptista G, Lee SM (2017) Identification of long non-coding RNAs in two anthozoan species and their possible implications for coral bleaching. Sci Rep 7:5333
Huang D, Meier R, Todd P, Chou LM (2008) Slow mitochondrial COI sequence evolution at the base of the metazoan tree and its implications for DNA barcoding. J Mol Evol 66:167–174
Huang D, Roy K (2013) Anthropogenic extinction threats and future loss of evolutionary history in reef corals. Ecol Evol 3:1184–1193
Huang D, Roy K (2015) The future of evolutionary diversity in reef corals. Phil Trans R Soc B 370:20140010
Huang D, Goldberg EE, Roy K (2015) Fossils, phylogenies, and the challenge of preserving evolutionary history in the face of anthropogenic extinctions. Proc Natl Acad Sci USA 112:4909–4914
Huang D, Hoeksema BW, Affendi YA, Ang PO, Chen CA, Huang H, Lane DJW, Licuanan WY, Vibol O, Vo ST, Yeemin T, Chou LM (2016b) Conservation of reef corals in the South China Sea based on species and evolutionary diversity. Biodivers Conserv 25:331–344
Hugall AF, O’Hara TD, Hunjan S, Nilsen R, Moussalli A (2016) An exon–capture system for the entire Class Ophiuroidea. Mol Biol Evol 33:281–294
Hughes TP, Bellwood DR, Connolly SR (2002) Biodiversity hotspots, centres of endemicity, and the conservation of coral reefs. Ecol Lett 5:775–784
Hughes TP, Barnes ML, Bellwood DR, Cinner JE, Cumming GS, Jackson JBC, Kleypas J, van de Leemput IA, Lough JM, Morrison TH, Palumbi SR, van Nes EH, Scheffer M (2017) Coral reefs in the Anthropocene. Nature 546:82–90
Hughes TP, Kerry JT, Baird AH, Connolly SR, Dietzel A, Eakin M, Heron SF, Hoey AS, Hoogenboom MO, Liu G, McWilliam MJ, Pears RJ, Pratchett MS, Skirving WJ, Stella JS, Torda G (2018) Global warming transforms coral reef assemblages. Nature 556:492–496
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23:254–267
i5K Consortium (2013) The i5K initiative: advancing arthropod genomics for knowledge, human health, agriculture, and the environment. J Hered 104:595–600
Iguchi A, Yoshioka Y, Forsman ZH, Knapp ISS, Toonen RJ, Hongo Y, Nagai S, Yasuda N (2019) RADseq population genomics confirms divergence across closely related species in blue coral (Heliopora coerulea). BMC Evol Biol 19:187
Inoue S, Kayanne H, Yamamoto S, Kurihara H (2013) Spatial community shift from hard to soft corals in acidified water. Nat Clim Change 3:683–687
Jeffroy O, Brinkmann H, Delsuc F, Philippe H (2006) Phylogenomics: the beginning of incongruence? Trends Genet 22:225–231
Jeon Y, Park SG, Lee N, Weber JA, Kim H, Hwang S, Woo S, Kim H, Bhak Y, Jeon S, Lee N, Jo Y, Blazyte A, Ryu T, Cho YS, Kim H, Lee J, Yim H, Bhak J, Yum S (2019) The draft genome of an octocoral, Dendronephthya gigantea. Genome Biol Evol 11:949–953
Jiang JB, Quattrini AM, Francis WR, Ryan JF, Rodríguez E, McFadden CS (2019) A hybrid de novo assembly of the sea pansy (Renilla muelleri) genome. GigaSci 8:giz026
Johnson CN, Balmford A, Brook BW, Buettel JC, Galetti M, Guangchun L, Wilmshurst JM (2017) Biodiversity losses and conservation responses in the Anthropocene. Science 356:270–275
Johnston EC, Forsman ZH, Flot J, Schmidt-Roach S, Pinzón JH, Knapp ISS, Toonen RJ (2017) A genomic glance through the fog of plasticity and diversification in Pocillopora. Sci Rep 7:5991
Kapli P, Yang Z, Telford MJ (2020) Phylogenetic tree building in the genomic age. Nat Rev Genet 21:428–444
Karin BR, Gamble T, Jackman TR (2020) Optimizing phylogenomics with rapidly evolving long exons: comparison with anchored hybrid enrichment and ultraconserved elements. Mol Biol Evol 37:904–922
Kayal E, Roure B, Philippe H, Collins AG, Lavrov DV (2013) Cnidarian phylogenetic relationships as revealed by mitogenomics. BMC Evol Biol 13:5
Kayal E, Bentlage B, Pankey SM, Ohdera AH, Medina M, Plachetzki DC, Collins AG, Ryan JF (2018) Phylogenomics provides a robust topology of the major cnidarian lineages and insights on the origins of key organismal traits. BMC Evol Biol 18:68
Kaehler BD, Yap VB, Zhang R, Huttley GA (2015) Genetic distance for a general non-stationary Markov substitution process. Syst Biol 64:281–293
Kimball RT, Oliveros CH, Wang N, White ND, Barker FK, Field DJ, Ksepka DT, Chesser RT, Moyle RG, Braun MJ, Brumfield RT, Faircloth BC, Smith BT, Braun EL (2019) A phylogenomic supertree of birds. Diversity 11:109
Kitahara MV, Lin MF, Forêt S, Huttley G, Miller DJ, Chen CA (2014) The “naked coral” hypothesis revisited – evidence for and against scleractinian monophyly. PLoS ONE 9:e94774
Kitchen SA, Crowder CM, Poole AZ, Weis VM, Meyer E (2015) De novo assembly and characterization of four anthozoan (Phylum Cnidaria) transcriptomes. G3 5:2441–2452
Kitchen SA, Ratan A, Bedoya-Reina OC, Burhans R, Fogarty ND, Miller W, Baums IB (2019) Genomic variants among threatened Acropora corals. G3 9:1633–1646
Klompen AML, Macrander J, Reitzel AM, Stampar SN (2020) Transcriptomic analysis of four cerianthid (Cnidaria, Ceriantharia) venoms. Mar Drugs 18:413
Knight R, Maxwell P, Birmingham A, Carnes J, Caporaso JG, Easton BC, Eaton M, Hamady M, Lindsay H, Liu Z, Lozupone C, McDonald D, Robeson M, Sammut R, Smit S, Wakefield MJ, Widmann J, Wikman S, Wilson S, Ying H, Huttley GA (2007) PyCogent: a toolkit for making sense from sequence. Genome Biol 8:R171
Knowles LL (2009) Estimating species trees: methods of phylogenetic analysis when there is incongruence across genes. Syst Biol 58:463–467
Koch TL, Grimmelikhuijzen CJP (2019) Global neuropeptide annotations from the genomes and transcriptomes of Cubozoa, Scyphozoa, Staurozoa (Cnidaria: Medusozoa), and Octocorallia (Cnidaria: Anthozoa). Front Endocrinol 10:831
Komyakova V, Munday PL, Jones GP (2013) Relative importance of coral cover, habitat complexity and diversity in determining the structure of reef fish communities. PLoS ONE 8:e83178
Kozlov AM, Aberer AJ, Stamatakis A (2015) ExaML version 3: a tool for phylogenomic analyses on supercomputers. Bioinformatics 31:2577–2579
Kozlov AM, Darriba D, Flouri T, Morel B, Stamatakis A (2019) RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35:4453–4455
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549
Kushida Y, Reimer JD (2019) Molecular phylogeny and diversity of sea pens (Cnidaria: Octocorallia: Pennatulacea) with a focus on shallow water species of the northwestern Pacific Ocean. Mol Phylogenet Evol 131:233–244
Lambert SM, Reeder TW, Wiens JJ (2015) When do species-tree and concatenated estimates disagree? An empirical analysis with higher-level scincid lizard phylogeny. Mol Phylogenet Evol 82:146–155
Lartillot N, Lepage T, Blanquart S (2009) PhyloBayes 3: a Bayesian software package for phylogenetic reconstruction and molecular dating. Bioinformatics 25:2286–2288
Laumer CE, Fernández R, Lemer S, Combosch D, Kocot KM, Riesgo A, Andrade SCS, Sterrer W, Sørensen MV, Giribet G (2019) Revisiting metazoan phylogeny with genomic sampling of all phyla. Proc R Soc B 286:20190831
Ledoux J, Cruz F, Gomez-Garrido J, Antoni R, Blanc J, Gómez-Gras D, Kipson S, López-Sendino P, Antunes A, Linares C, Gut M, Alioto T, Garrabou J (2020) The genome sequence of the octocoral Paramuricea clavata – a key resource to study the impact of climate change in the Mediterranean. G3 10:2941–2952
Lemmon AR, Emme SA, Lemmon EM (2012) Anchored hybrid enrichment for massively high-throughput phylogenomics. Syst Biol 61:727–744
Liao X, Li M, Zou Y, Wu F, Yi-Pan WJ (2019) Current challenges and solutions of de novo assembly. Quant Biol 7:90–109
Liew YJ, Aranda M, Voolstra CR (2016) Reefgenomics. Org – a repository for marine genomics data. Database 2016:baw152
Lin MF, Kitahara MV, Luo H, Tracey D, Geller J, Fukami H, Miller DJ, Chen CA (2014) Mitochondrial genome rearrangements in the Scleractinia/Corallimorpharia complex: implications for coral phylogeny. Genome Biol Evol 6:1086–1095
Lin MF, Chou WH, Kitahara MV, Chen CLA, Miller DJ, Forêt S (2016) Corallimorpharians are not “naked corals”: insights into relationships between Scleractinia and Corallimorpharia from phylogenomic analyses. PeerJ 4:e2463
Lin MF, Moya A, Ying H, Chen CA, Cooke I, Ball EE, Forêt S, Miller DJ (2017a) Analyses of corallimorpharian transcriptomes provide new perspectives on the evolution of calcification in the Scleractinia (corals). Genome Biol Evol 1:150–160
Lin Z, Chen M, Dong X (2017b) Transcriptome profiling of Galaxea fascicularis and its endosymbiont Symbiodinium reveals chronic eutrophication tolerance pathways and metabolic mutualism between partners. Sci Rep 7:42100
Liu SYV, Chan CLC, Hsieh HJ, Fontana S, Wallace CC, Chen CA (2015) Massively parallel sequencing (MPS) assays for sequencing mitochondrial genomes: the phylogenomic implications for Acropora staghorn corals (Scleractinia; Acroporidae). Mar Biol 162:1383–1392
Locatelli NS, Drew JA (2019) Population structure and clonal prevalence of scleractinian corals (Montipora capitata and Porites compressa) in Kaneohe Bay, Oahu. bioRxiv doi: https://doi.org/10.1101/2019.12.11.860585
Luypaert T, Hagan JG, McCarthy ML, Poti M (2020) Status of marine biodiversity in the Anthropocene. In: Jungblut S, Liebich V, Bode-Dalby M (eds) YOUMARES 9 - the oceans: our research, our future. Springer, Cham, pp 57–82
Mao Y, Economo EP, Satoh N (2018) The roles of introgression and climate change in the rise to dominance of Acropora corals. Curr Biol 28:3373–3382
Mao Y, Satoh N (2019) A likely ancient genome duplication in the speciose reef-building coral genus, Acropora. iScience 13:20–32
Mao Y (2020) Genomic insights into hybridization of reef corals. Coral Reefs 39:61–67
Mao Y, Hou S, Shi J, Economo EP (2020) TREEasy: an automated workflow to infer gene trees, species trees, and phylogenetic networks from multilocus data. Mol Ecol Res 20:832–840
McFadden CS, Hutchinson MB (2004) Molecular evidence for the hybrid origin of species in the soft coral genus Alcyonium (Cnidaria: Anthozoa: Octocorallia). Mol Ecol 13:1496–1505
McFadden CS, Sánchez JA, France SC (2010) Molecular phylogenetic insights into the evolution of Octocorallia: a review. Integr Comp Biol 50:389–410
McFadden CS, Haverkort-Yeh R, Reynolds AM, Halàsz A, Quattrini AM, Forsman ZH, Benayahu Y, Toonen RJ (2017) Species boundaries in the absence of morphological, ecological or geographical differentiation in the Red Sea octocoral genus Ovabunda (Alcyonacea: Xeniidae). Mol Phylogenet Evol 112:174–184
McFadden CS, Quattrini AM, Brugler MR, Cowman PF, Dueñas LF, Kitahara MV, Paz-García DA, Reimer JD, Rodríguez E (2021) Phylogenomics, origin and diversification of anthozoans (Phylum Cnidaria). Syst Biol
McGill BJ, Dornelas M, Gotelli NJ, Magurran AE (2015) Fifteen forms of biodiversity trend in the Anthropocene. Trends Ecol Evol 30:104–113
McKain MR, Johnson MG, Uribe-Convers S, Eaton D, Yang Y (2018) Practical considerations for plant phylogenomics. Appl Plant Sci 6:e1038
Medina M, Collins AG, Takoka TL, Kuehl JV, Boore JL (2006) Naked corals: skeleton loss in Scleractinia. Proc Natl Acad Sci USA 103:9096–9100
Mejia ACF, Molodtsova T, Östman C, Bavestrello G, Rouse GW (2020) Molecular phylogeny of Ceriantharia (Cnidaria: Anthozoa) reveals non-monophyly of traditionally accepted families. Zool J Linn Soc 190:397–416
Miller MR, Dunham JP, Amores A, Cresko WA, Johnson EA (2007) Rapid and cost-effective polymorphism identification and genotyping using restriction site associated DNA (RAD) markers. Genome Res 17:240–248
Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, von Haeseler A, Lanfear R (2020) IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol 37:1530–1534
Mitchell ML, Tonkin-Hill GQ, Morales RAV, Purcell AW, Papenfuss AT, Norton RS (2020) Tentacle transcriptomes of the speckled anemone (Actiniaria: Actiniidae: Oulactis sp.): venom-related components and their domain structure. Mar Biotechnol 22:207–219
Moberg F, Folke C (1999) Ecological goods and services of coral reef ecosystems. Ecol Econ 29:215–233
Morel B, Kozlov AM, Stamatakis A (2019) ParGenes: a tool for massively parallel model selection and phylogenetic tree inference on thousands of genes. Bioinformatics 15:1771–1773
Morlighem JRL, Huang C, Liao Q, Gones PB, Pérez CD, Prieto-da-Silva ARDB, Lee SM, Rádis-Baptista G (2018) The holo-transcriptome of the zoantharian Protopalythoa variabilis (Cnidaria: Anthozoa): a plentiful source of enzymes for potential application in green chemistry, industrial and pharmaceutical biotechnology. Mar Drugs 16:207
Morrison DA (2014) Phylogenetic networks: a review of methods to display evolutionary history. Annu Res Rev Biol 4:1518–1543
Mouillot D, Parravicini V, Bellwood D, Leprieur F, Huang D, Cowman PF, Albouy C, Hughes TP, Thuiller W, Guilhaumon F (2016) Global marine protected areas do not secure the evolutionary history of tropical corals and fishes. Nat Commun 7:10359
Moya A, Ganot P, Furla P, Sabourault C (2012) The transcriptomic response to thermal stress is immediate, transient and potentiated by ultraviolet radiation in the sea anemone Anemonia viridis. Mol Ecol 21:1158–1174
Neri F, Rapelli S, Krepelova A, Incarnato D, Parlato C, Basile G, Maldotti M, Anselmi F, Oliviero S (2017) Intragenic DNA methylation prevents spurious transcription initiation. Nature 543:72–77
Nevill PG, Zhong X, Tonti-Filippini J, Byrne M, Hislop M, Thiele K, van Leeuwen S, Boykin LM, Small I (2020) Large scale genome skimming from herbarium material for accurate plant identification and phylogenomics. Plant Methods 16:1
Newton K, Côté IM, Pilling GM, Jennings S, Dulvy NK (2007) Current and future sustainability of island coral reef fisheries. Curr Biol 17:655–658
Niu W, Huang H, Lin R, Chen C, Shen K, Hsiao C (2016a) The complete mitogenome of the galaxy coral, Galaxea fascicularis (Cnidaria: Oculinidae). Mitochondrial DNA B 1:10–11
Niu W, Lin R, Shi X, Chen C, Shen K, Hsiao C (2016b) Next-generation sequencing yields the complete mitogenome of massive coral, Porites lutea (Cnidaria: Poritidae). Mitochondrial DNA B 1:8–9
Niu W, Huang H, Yu S (2017a) Next-generation sequencing yields the complete mitogenome of stony coral (Favites abdita). Mitochondrial DNA B 2:426–427
Niu W, Yu S, Chen B, Suharsono S (2017b) The complete mitochondrial genome of the Cyphastrea serailia. Mitochondrial DNA B 2:415–416
Niu W, Yu S, Tian P, Xiao J (2018) Complete mitochondrial genome of Echinophyllia aspera (Scleractinia, Lobophylliidae): mitogenome characterization and phylogenetic positioning. ZooKeys 793:1–14
Niu W, Xiao J, Tian P, Guo F (2020a) The complete mitochondrial genome of Plesiastrea versipora (Scleractinia, Plesiastreidae) sheds light on its phylogeny and taxonomy of the family Plesiastreidae. Saudi J Biol Sci 27:1830–1834
Niu W, Xiao J, Tian P, Yu S, Guo F, Wang J, Huang D (2020b) Characterization of the complete mitochondrial genome sequences of three Merulinidae corals and novel insights into the phylogenetics. PeerJ 8:e8455
Odorico DM, Miller DJ (1997) Variation in the ribosomal internal transcribed spacers and 5.8S rDNA among five species of Acropora (Cnidaria; Scleractinia): patterns of variation consistent with reticulate evolution. Mol Biol Evol 14:465–473
Okubo N, Hayward DC, Forêt S, Ball EE (2016) A comparative view of early development in the corals Favia lizardensis, Ctenactis echinata, and Acropora millepora – morphology, transcriptome, and developmental gene expression. BMC Evol Biol 16:48
One Thousand Plant Transcriptomes Initiative (2019) One thousand plant transcriptomes and the phylogenomics of green plants. Nature 574:679–685
Padial JM, De la Riva I (2010) A response to recent proposals of integrative taxonomy. Biol J Linn Soc 101:747–756
Padial JM, Miralles A, De la Riva I, Vences M (2010) The integrative future of taxonomy. Front Zool 7:16
Palmer CV, Traylor-Knowles NG (2018) Cnidaria: anthozoans in the hot seat. In: Cooper EL (ed) Advances in comparative immunology. Springer, Cham, pp 51–93
Pante E, Abdelkrim J, Viricel A, Gey D, France SC, Boisselier MC, Samadi S (2015) Use of RAD sequencing for delimiting species. Heredity 114:450–459
Peterson BK, Weber JN, Kay EH, Fisher HS, Hoekstra HE (2012) Double digest RADseq: an inexpensive method for de novo SNP discovery and genotyping in model and non-model species. PLoS ONE 7:e37135
Philippe H, Delsuc F, Brinkmann H, Lartillot N (2005) Phylogenomics. Annu Rev Ecol Evol Syst 36:541–562
Philippe H, Brinkmann H, Lavrov DV, Littlewood DT, Manuel M, Worheide G, Baurain D (2011) Resolving difficult phylogenetic questions: why more sequences are not enough. PLoS Biol 9:e1000602
Philippe H, de Vienne DM, Ranwez V, Roure B, Baurain D, Delsuc F (2017) Pitfalls in supermatrix phylogenomics. Eur J Taxon 283:1–25
Plaisance L, Caley MJ, Brainard RE, Knowlton N (2011) The diversity of coral reefs: what are we missing? PLoS ONE 6:e25026
Poliseno A, Breedy O, Eitel M, Wöerheide G, Guzman HM, Krebs S, Blum H, Vargas S (2016) Complete mitochondrial genome of Muricea crassa and Muricea purpurea (Anthozoa: Octocorallia) from the eastern tropical Pacific. bioRxiv doi: https://doi.org/10.1101/042945
Poliseno A, Santos MEA, Kise H, Macdonald B, Quattrini AM, McFadden CS, Reimer JD (2020) Evolutionary implications of analyses of complete mitochondrial genomes across order Zoantharia (Cnidaria: Hexacorallia). J Zool Syst Evol Res 58:858–868
Pollard MO, Gurdasani D, Mentzer AJ, Porter T, Sandhu MS (2018) Long reads: their purpose and place. Hum Mol Genet 27:R234–R241
Pont-Kingdon GA, Okada NA, Macfarlane JL, Beagley CT, Wolstenholme DR, Cavelier-Smith T, Clark-Walker D (1995) A coral mitochondrial mutS gene. Nature 375:109–111
Poole AZ, Weis VM (2014) TIR-domain-containing protein repertoire of nine anthozoan species reveals coral–specific expansions and uncharacterized proteins. Dev Comp Immunol 46:480–488
Pootakham W, Naktang C, Sonthirod C, Yoocha T, Sangsrakru D, Jomchai N, Putchim L, Tangphatsornruang S (2018) Development of a novel reference transcriptome for scleractinian coral Porites lutea using single-molecule long-read isoform sequencing (iso-seq). Front Mar Sci 5:122
Porro B, Mallien C, Hume BCC, Pey A, Aubin E, Christen R, Voolstra CR, Furla P, Forcioli D (2020) The many faced symbiotic snakelocks anemone (Anemonia viridis, Anthozoa): host and symbiont genetic differentiation among colour morphs. Heredity 124:351–366
Prasanna AN, Gerber D, Kijpornyongpan T, Aime MC, Doyle VP, Nagy LG (2020) Model choice, missing data, and taxon sampling impact phylogenomic inference of deep basidiomycota relationships. Syst Biol 69:17–37
Pratlong M, Rancurel C, Pontarotti P, Aurelle D (2017) Monophyly of Anthozoa (Cnidaria): why do nuclear and mitochondrial phylogenies disagree? Zool Scr 46:363–371
Pratlong M, Haguenauer A, Chenesseau S, Brener K, Mitta G, Toulza E, Bonabaud M, Rialle S, Aurelle D, Pontarotti P (2017) Evidence for a genetic sex determination in Cnidaria, the Mediterranean red coral (Corallium rubrum). R Soc Open Sci 4:160880
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Putnam NH, Srivastava M, Hellsten U, Dirks B, Chapman J, Salamov A, Terry A, Shapiro H, Lindquist E, Kapitonov VV, Jurka J, Genikhovich G, Grigoriev IV, Lucas SM, Steele RE, Finnerty JR, Technau U, Martindale MQ, Rokhsar DS (2007) Sea anemone genome reveals ancestral eumetazoan gene repertoire and genomic organization. Science 317:86–94
Quattrini AM, Wu T, Soong K, Jeng M, Benayahu Y, McFadden CS (2019) A next generation approach to species delimitation reveals the role of hybridization in a cryptic species complex of corals. BMC Evol Biol 19:116
Quattrini AM, Faircloth BC, Dueñas LF, Bridge TCL, Brugler MR, Calixto-Botía IF, DeLeo DM, Forêt S, Herrera S, Lee SMY, Miller DJ, Prada C, Rádis-Baptista G, Ramírez-Portilla C, Sánchez JA, Rodríguez E, McFadden CS (2018) Universal target-enrichment baits for anthozoan (Cnidaria) phylogenomics: new approaches to long-standing problems. Mol Ecol Res 18:281–295
Quattrini AM, Rodríguez E, Faircloth BC, Cowman PF, Brugler MR, Farfan GA, Hellberg ME, Kitahara MV, Morrison CL, Paz-García DA, Reimer JD, McFadden CS (2020) Palaeoclimate ocean conditions shaped the evolution of corals and their skeletons through deep time. Nat Ecol Evol 4:1531–1538
Quek ZBR, Huang D (2019) Effects of missing data and data type on phylotranscriptomic analysis of stony corals (Cnidaria: Anthozoa: Scleractinia). Mol Phylogenet Evol 134:12–23
Quek RZB, Jain SS, Neo ML, Rouse GW, Huang D (2020) Transcriptome-based target-enrichment baits for stony corals (Cnidaria: Anthozoa: Scleractinia). Mol Ecol Res 20:807–818
ReFuGe 2020 Consortium (2015) The ReFuGe 2020 Consortium – using “omics” approaches to explore the adaptability and resilience of coral holobionts to environmental change. Front Mar Sci 2:68
Reimer JD, Takishita K, Ono S, Tsukahara J, Maruyama T (2007) Molecular evidence suggesting interspecific hybridization in Zoanthus spp. (Anthozoa: Hexacorallia). Zool Sci 24:346–359
Reitzel AM, Herrera S, Layden MJ, Martindale MQ, Shank TM (2013) Going where traditional markers have not gone before: utility of and promise for RAD sequencing in marine invertebrate phylogeography and population genomics. Mol Ecol 22:2953–2970
Rentzsch F, Technau U (2016) Genomics and development of Nematostella vectensis and other anthozoans. Curr Opin Genet Dev 39:63–70
Richards ZT, Yasuda N, Kikuchi T, Foster T, Mitsuyuki C, Stat M, Suyama Y, Wilson NG (2018) Integrated evidence reveals a new species in the ancient blue coral genus Heliopora (Octocorallia). Sci Rep 8:15875
Richards ZT, Carvajal JI, Wallace CC, Wilson NG (2020) Phylotranscriptomics confirms Alveopora is sister to Montipora within the family Acroporidae. Mar Genomics 50:100703
Ripma LA, Simpson MG, Hasenstab-Lehman K (2014) Geneious! simplified genome skimming methods for phylogenetic systematic studies: a case study in Oreocarya (Boraginaceae). Appl Plant Sci 2:1400062
Rochette NC, Rivera-Colón AG, Catchen JM (2019) Stacks 2: analytical methods for paired-end sequencing improve RADseq-based population genomics. Mol Ecol 28:4737–4754
Robbins SJ, Singleton CM, Chan CX, Messer LF, Geers AU, Ying H, Baker A, Bell SC, Morrow KM, Ragan MA, Miller DJ, Forêt S, Voolstra CR, Tyson GW, Bourne DG (2019) A genomic view of the reef-building coral Porites lutea and its microbial symbionts. Nat Microbiol 4:2090–2100
Roberts JM, Wheeler AJ, Freiwald A (2006) Reefs of the deep: the biology and geology of cold-water coral ecosystems. Science 312:543–547
Romano SL, Palumbi SR (1996) Evolution of scleractinian corals inferred from molecular systematics. Science 271:640–642
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542
Rosser NL, Thomas L, Stankowski S, Richards ZT, Kennington WJ, Johnson MS (2017) Phylogenomics provides new insight into evolutionary relationships and genealogical discordance in the reef-building coral genus Acropora. Proc R Soc B 284:20162182
Roure B, Baurain D, Philippe H (2012) Impact of missing data on phylogenies inferred from empirical phylogenomic data sets. Mol Biol Evol 30:197–214
Rubin BER, Ree RH, Moreau CS (2012) Inferring Phylogenies from RAD Sequence Data. PLoS ONE 7:e33394
Russello MA, Waterhouse MD, Etter PD, Johnson EA (2015) From promise to practice: pairing non-invasive sampling with genomics in conservation. PeerJ 3:e1106
Ryu T, Cho I, Hwang S, Yum S, Kim M, Woo S (2019) Holobiont transcriptome of colonial scleractinian coral Alveopora japonica. Mar Genomics 43:68–71
Ryu T, Cho I, Hwang S, Yum S, Kim M, Woo S (2019) First transcriptome assembly of the temperate azooxanthellate octocoral Eleutherobia rubra. Mar Genom 48:100682
Sánchez JA, Dorado D (2008) Intragenomic ITS2 variation in Caribbean seafans. Proceedings of the 11th International Coral Reef Symposium 1:1383–1387
Santos MEA, Kitahara MV, Linder A, Reimer JD (2015) Overview of the order Zoantharia (Cnidaria: Anthozoa) in Brazil. Mar Biodivers 46:547–559
Seelenfreund E, Robinson WA, Amato CM, Tan A, Kim J, Robinson SE (2014) Long term storage of dry versus frozen RNA for next generation molecular studies. PLoS ONE 9:e111827
Seiblitz IGL, Capel KCC, Stolarski J, Quek ZBR, Huang D, Kitahara MV (2020) The earliest diverging extant scleractinian corals recovered by mitochondrial genomes. Sci Rep 10:20714
Shearer TL, van Oppen MJH, Romano SL, Wörheide G (2008) Slow mitochondrial DNA sequence evolution in the Anthozoa (Cnidaria). Mol Ecol 11:2475–2487
Shen C, Dan Y, Asem A, Wang P, Xue W, Tong X, Li W (2019) The complete mitochondrial genome of soft coral Sarcophyton trocheliophorum (Cnidaria: Anthozoa) using next-generation sequencing. Mitochondrial DNA B 4:3734–3735
Shinzato C, Shogichi E, Kawashima T, Hamada M, Hisata K, Tanaka M, Fujie M, Fujiwara M, Koyanagi R, Ikuta T, Fujiyama A, Miller DJ, Satoh N (2011) Using the Acropora digitifera genome to understand coral responses to environmental change. Nature 476:320–323
Shinzato C, Khalturin K, Inoue J, Zayasu Y, Kanda M, Kawamitsu M, Yoshioka Y, Yamashita H, Suzuki G, Satoh N (2020) Eighteen coral genomes reveal the evolutionary origin of Acropora strategies to accommodate environmental changes. Mol Biol Evol. https://doi.org/10.1093/molbev/msaa216
Shumaker A, Putnam HM, Qiu H, Price DC, Zelzion E, Harel A, Wagner NE, Gates RD, Yoon HS, Bhattacharya D (2019) Genome analysis of the rice coral Montipora capitata. Sci Rep 9:2571
Silvestri S, Figueroa DF, Hicks D, Figueroa NJ (2019) Mitogenomic phylogenetic analyses of Leptogorgia virgulata and Leptogorgia hebes (Anthozoa: Octocorallia) from the Gulf of Mexico provides insight on Gorgoniidae divergence between Pacific and Atlantic lineages. Ecol Evol 9:14114–14129
Simion P, Philippe H, Baurain D, Jager M, Richter DJ, Franco AD, Roure B, Satoh N, Quéinnec E, Ereskovsky A, Lapébie P, Corre E, Delsuc F, King N, Wörheide G, Manuel M (2017) A large and consistent phylogenomic dataset supports sponges as the sister group to all other animals. Curr Biol 7:958–967
Sims GE, Jun S, Wu GA, Kim S (2009) Alignment-free genome comparison with feature frequency profiles (FFP) and optimal resolutions. Proc Natl Acad Sci USA 106:2677–2682
Small AM, Adey WH, Spoon D (1998) Are current estimates of coral reef biodiversity too low? The view through the window of a microcosm. Atoll Res Bull 458:1–20
Smith DR (2020) Revisiting ceriantharian (Anthozoa) mitochondrial genomes: casting doubts about their structure and size. Genome Biol Evol 12:1440–1443
Smith H, Epstein H, Torda G (2017) The molecular basis of differential morphology and bleaching thresholds in two morphs of the coral Pocillopora acuta. Sci Rep 7:10066
Solís-Lemus C, Ané C (2016) Inferring phylogenetic networks with maximum pseudolikelihood under incomplete lineage sorting. PLoS Genet 12:e1005896
Solís-Lemus C, Bastide P, Ané C (2017) PhyloNetworks: a package for phylogenetic networks. Mol Biol Evol 34:3292–3298
Spalding M, Burke L, Wood SA, Ashpole J, Hutchison J, zu Ermgassen P (2017) Mapping the global value and distribution of coral reef tourism. Mar Policy 82:104–113
Spano CA, Häussermann V, Acuñad FH, Griffiths C, Seeb LW, Gomez-Uchida D (2018) Hierarchical biogeographical processes largely explain the genomic divergence pattern in a species complex of sea anemones (Metridioidea: Sagartiidae: Anthothoe). Mol Phylogenet Evol 127:217–228
Speers AE, Besedin EY, Palardy JE, Moore C (2016) Impacts of climate change and ocean acidification on coral reef fisheries: an integrated ecological–economic model. Ecol Econ 128:33–43
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313
Stampar SN, Broe MB, Macrander J, Reitzel AM, Brugler MR, Daly M (2019) Linear mitochondrial genome in Anthozoa (Cnidaria): a case study in Ceriantharia. Sci Rep 9:6094
Stolarski J, Coronado I, Murphy JG, Kitahara MV, Janiszewska K, Mazur M, Gothmann AM, Bouvier AS, Marin-Carbonne J, Taylor ML, Quattrini AM, McFadden CS, Higgins JA, Robinson LF, Meibom A (2021) A modern scleractinian coral with a two-component calcite–aragonite skeleton. Proc Natl Acad Sci USA 118:e2013316117
Straub SCK, Parks M, Weitemier K, Fishbein M, Cronn RC, Liston A (2012) Navigating the tip of the genomic iceberg: next-generation sequencing for plant systematics. Am J Bot 99:349–364
Stuart-Smith RD, Brown CJ, Ceccarelli DM, Edgar GJ (2018) Ecosystem restructuring along the Great Barrier Reef following mass coral bleaching. Nature 560:92–96
Sturm AB, Eckert RJ, Méndez JG, González-Díaz P, Voss JD (2020) Population genetic structure of the great star coral, Montastraea cavernosa, across the Cuban archipelago with comparisons between microsatellite and SNP markers. Sci Rep 10:15432
Surm JM, Stewart ZK, Papanicalaou A, Pavasovic A, Prentis PJ (2019) The draft genome of Actinia tenebrosa reveals insights into toxin evolution. Ecol Evol 9:11314–11328
Suyama Y, Matsuki Y (2015) MIG-seq: an effective PCR-based method for genome-wide single-nucleotide polymorphism genotyping using the next-generation sequencing platform. Sci Rep 5:16963
Swofford DL (2003) PAUP*. Phylogenetic analysis using parsimony (*and other methods). Sinauer Associates, Sunderland, Massachusetts
Takata K, Taninaka H, Nonaka M, Iwase F, Kikuchi T, Suyama Y, Nagai S, Yasuda N (2019) Multiplexed ISSR genotyping by sequencing distinguishes two precious coral species (Anthozoa: Octocorallia: Coralliidae) that share a mitochondrial haplotype. PeerJ 7:e7769
Takeuchi T, Yamada L, Shinzato C, Sawada H, Satoh N (2016) Stepwise evolution of coral biomineralization revealed with genome-wide proteomics and transcriptomics. PLoS ONE 11:e0156424
Talevich E, Invergo BM, Cock PJ, Chapman BA (2012) Bio.Phylo: a unified toolkit for processing, analyzing and visualizing phylogenetic trees in Biopython. BMC Bioinform 13:209
Terraneo TI, Arrigoni R, Benzoni F, Forsman ZH, Berumen ML (2018a) Using ezRAD to reconstruct the complete mitochondrial genome of Porites fontanesii (Cnidaria: Scleractinia). Mitochondrial DNA B 3:173–174
Terraneo TI, Arrigoni R, Benzoni F, Forsman ZH, Berumen ML (2018b) The complete mitochondrial genome of Porites harrisoni (Cnidaria: Scleractinia) obtained using next-generation sequencing. Mitochondrial DNA B 3:286–287
Tisthammer KH, Forsman ZH, Sindorf VL, Massey TL, Bielecki CR, Toonen RJ (2016) The complete mitochondrial genome of the lobe coral Porites lobata (Anthozoa: Scleractinia) sequenced using ezRAD. Mitochondrial DNA B 1:247–249
Titus BM, Blischak PD, Daly M (2019) Genomic signatures of sympatric speciation with historical and contemporary gene flow in a tropical anthozoan (Hexacorallia: Actiniaria). Mol Ecol 28:3572–3586
Titus BM, Benedict C, Laroche R, Gusmão LC, Van Deusen V, Chiodo T, Meyer CP, Berumen ML, Bartholomew A, Yanagi K, Reimer JD, Fujii T, Daly M, Rodríguez E (2019) Phylogenetic relationships among the clownfish-hosting sea anemones. Mol Phylogenet Evol 139:106526
Thompson JR, Rivera HE, Closek CJ, Medina M (2015) Microbes in the coral holobiont: partners through evolution, development, and ecological interactions. Front Cell Infect Microbiol 4:176
Todd PA (2008) Morphological plasticity in scleractinian corals. Biol Rev 83:315–337
Tonini J, Moore A, Stern D, Shcheglovitova M, Ortí G (2015) Concatenation and species tree methods exhibit statistically indistinguishable accuracy under a range of simulated conditions. PLos Curr. https://doi.org/10.1371/currents.tol.34260cc27551a527b124ec5f6334b6be
Toonen RJ, Puritz JB, Forsman ZH, Whitney JL, Fernandez-Silva I, Andrews KR, Bird CE (2013) ezRAD: a simplified method for genomic genotyping in non-model organisms. PeerJ 1:e203
Townsend TM, Mulcahy DG, Noonan BP, Sites JW Jr, Kuczynski CA, Weins JJ, Reeder TW (2011) Phylogeny of iguanian lizards inferred from 29 nuclear loci, and a comparison of concatenated and species-tree approaches for an ancient, rapid radiation. Mol Phylogenet Evol 61:363–380
Tsounis G, Edmunds PJ (2017) Three decades of coral reef community dynamics in St. John, USVI: a contrast of scleractinians and octocorals. Ecosphere 8:e01646
Tu T, Dai C, Jeng M (2015) Phylogeny and systematics of deep-sea precious corals (Anthozoa: Octocorallia: Coralliidae). Mol Phylogenet Evol 84:173–184
Uda K, Komeda Y, Fujita T, Iwasaki N, Bavestrello G, Giovine M, Cattaneo-Vietti R, Suzuki T (2013) Complete mitochondrial genomes of the Japanese pink coral (Corallium elatius) and the Mediterranean red coral (Corallium rubrum): a reevaluation of the phylogeny of the family Coralliidae based on molecular data. Comp Biochem Phys D 8:209–219
Untiedt CB, Quattrini AM, McFadden CS, Alderslade PA, Pante E, Burridge CP (2021) Phylogenetic relationships within Chrysogorgia (Alcyonacea: Octocorallia), a morphologically diverse genus of octocoral, revealed using a target enrichment approach. Front Mar Sci 7:599984
Urbarova I, Karlsen BO, Okkenhaug S, Seternes OM, Johansen SD, Emblem A (2012) Digital marine bioprospecting: mining new neurotoxin drug candidates from the transcriptomes of cold-water sea anemones. Mar Drugs 10:2265–2279
Valdecasas AG, Williams D, Wheeler QD (2007) ‘Integrative taxonomy’ then and now: a response to Dayrat (2005). Biol J Linn Soc 93:211–216
Villanoy C, David L, Cabrera O, Atrigenio M, Siringan F, Aliño P, Villaluz M (2012) Coral reef ecosystems protect shore from high-energy waves under climate change scenarios. Clim Change 112:493–505
Wang X, Drillon G, Ryu T, Voolstra CR, Aranda M (2017a) Genome-based analyses of six hexacorallian species reject the “naked coral” hypothesis. Genome Biol Evol 9:2626–2634
Wang X, Liew YJ, Li Y, Zoccola D, Tambutte S, Aranda M (2017b) Draft genomes of the corallimorpharians Amplexidiscus fenestrafer and Discosoma sp. Mol Ecol Res 17:e187–e195
Wang X, Tian P, Niu W, Yu S (2018) The complete mitochondrial genome of the Montipora peltiformi (Scleractinia: Acroporidae). Mitochondrial DNA B 3:99–100
Watling L, France SC, Pante E, Simpson A (2011) Chapter two – biology of deep-water octocorals. Adv Mar Biol 60:41–122
Wepfer PH, Nakajima Y, Sutthacheep M, Radice VZ, Richards Z, Ang P, Terraneo T, Sudek M, Fujimura A, Toonen RJ, Mikheyev AS, Economo EP, Mitarai S (2020) Evolutionary biogeography of the reef-building coral genus Galaxea across the Indo-Pacific ocean. Mol Phylogenet Evol 151:106905
Whelan NV, Kocot KM, Moroz LL, Halanych KM (2015) Error, signal, and the placement of Ctenophora sister to all other animals. Proc Natl Acad Sci USA 112:5773–5778
White AT, Vogt HP, Arin T (2000) Philippine coral reefs under threat: the economic losses caused by reef destruction. Mar Pollut Bull 40:598–605
White WT, Corrigan S, Yang L, Henderson AC, Bazinet AL, Swofford DL, Naylor GJP (2017) Phylogeny of the manta and devilrays (Chondrichthyes: Mobulidae), with an updated taxonomic arrangement for the family. Zool J Linn Soc 182:50–75
Wilding CS, Fletcher N, Smith EK, Prentis P, Weedall GD, Stewart Z (2020) The genome of the sea anemone Actinia equina (L.): meiotic toolkit genes and the question of sexual reproduction. Mar Genom 53:100753
Williams TA, Cox CJ, Foster PG, Szöllősi GJ, Embley TM (2020) Phylogenomics provides robust support for a two-domains tree of life. Nat Ecol Evol 4:138–147
Winter M, Devictor V, Schweiger O (2013) Phylogenetic diversity and nature conservation: where are we? Trends Ecol Evol 28:199–204
Xi Z, Liu L, Davis CC (2016) The impact of missing data on species tree estimation. Mol Biol Evol 33:838–860
Xiao M, Brugler MR, Broe MB, Gusmão LC, Daly M, Rodríguez E (2019) Mitogenomics suggests a sister relationship of Relicanthus daphneae (Cnidaria: Anthozoa: Hexacorallia: incerti ordinis) with Actiniaria. Sci Rep 9:18182
Ying H, Cooke I, Sprungala S, Wang W, Hayward DC, Tang Y, Miller DJ (2018) Comparative genomics reveals the distinct evolutionary trajectories of the robust and complex coral lineages. Genome Biol 19:175
Young AD, Gillung JP (2020) Phylogenomics – principles, opportunities and pitfalls of big-data phylogenetics. Syst Entomol 45:225–247
Yu Y, Dong J, Liu KJ, Nakhleh L (2014) Maximum likelihood inference of reticulate evolutionary histories. Proc Natl Acad Sci USA 111:16448–16453
Yu S, Guo F, Tian P, Huang D, Wang J, Xiao J, Niu W (2019) Complete mitochondrial genome of Acropora valida (Scleractinia, Acroporidae): mitogenome characterization and phylogenetic positioning. Mitochondrial DNA B 4:303–304
Zapata F, Goetz FE, Smith SA, Howison M, Siebert S, Church SH, Sanders SM, Ames CL, McFadden CS, France SC, Daly M, Collins AG, Haddock SHD, Dunn CW, Cartwright P (2015) Phylogenomic analyses support traditional relationships within Cnidaria. PLoS ONE 10:e0139068
Zhang G (2015) Bird sequencing project takes off. Nature 522:34
Zhang F, Ding Y, Zhu C, Zhou X, Orr MC, Scheu S, Luan Y (2019) Phylogenomics from low-coverage whole-genome sequencing. Methods Ecol Evol 10:507–517
Zhang Y, Chen Q, Xie JY, Yeung YH, Xiao B, Liao B, Xu J, Qiu J (2019) Development of a transcriptomic database for 14 species of scleractinian corals. BMC Genom 20:387
Zhang C, Rabiee M, Sayyari E, Mirarab S (2020) ASTRAL-Pro: quartet-based species-tree inference despite paralogy. Mol Biol Evol. https://doi.org/10.1093/molbev/msaa139
Acknowledgements
This research was funded by the National Research Foundation, Prime Minister’s Office, Singapore, under its Marine Science R&D Programme (MSRDP-P03).
Author information
Authors and Affiliations
Corresponding author
Additional information
Topic Editor Carly Kenkel
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
About this article
Cite this article
Quek, Z.B.R., Huang, D. Application of phylogenomic tools to unravel anthozoan evolution. Coral Reefs 41, 475–495 (2022). https://doi.org/10.1007/s00338-021-02072-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00338-021-02072-3