Abstract
Based on only a handful of mitochondrial and nuclear loci, the phylogenetic relationships within the genus Acropora have been unclear. However, with the new sequencing technology of massively parallel sequencing (MPS), the inter-specific relationships within Acropora may be resolved. We performed multiplex sequencing of the mitochondrial genome of eleven Acropora species representing different groups using the Illumina Solexa platform. Mitochondrial genomes were sequenced from long PCR-amplified templates ligated with different index sequences (~9 kbp) and analyzed using the mitochondrial genome of Acropora tenuis as a reference. A total of 75 million read outputs in one Illumina lane were obtained, with mapping results having coverage up to 44,000-fold. Assembly results of multiplex samples confirmed with Sanger sequencing produced <0.03 % error. Aligning the eleven mitochondrial genomes with the reference sequence revealed only 110 phylogenetically informative sites over the mitochondrial genome. The largest pairwise genetic distance observed was in the putative control region (0.022). A comparison of two phylogenetic trees based on the whole mitochondrial genome and control region showed that the former tree produces a higher resolution of phylogenetic relationships. In this study, we demonstrated the first case of sequencing cnidarian mitochondrial genomes by using multiplex MPS and applying it in a phylogenomic analyses.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the past, only one or a few loci could be utilized to reconstruct evolutionary patterns due to the high cost and low efficiency of the Sanger DNA sequencing method. The development of high-throughput DNA sequencing technology in recent years has provided researchers with efficient ways to obtain large amounts of the information embedded in genomes (Shendure and Ji 2008). Application of this technology has generated comparative data from large numbers of independent loci in different species (Faircloth et al. 2013). Massively parallel sequencing (MPS) technology, also known as next-generation sequencing, has become an effective tool to conduct genomic studies of non-model organisms in recent years (Hudson 2008; reviewed in Ekblom and Galindo 2011). MPS has been used for sequencing the mitochondrial genomes of nematodes for medical parasitology (Jex et al. 2008, 2009a) and mollusk taxonomy (Feldmeyer et al. 2010; Williams et al. 2014), and for sequencing chloroplast genomes for population and phylogenetic studies (Cronn et al. 2008; Doorduin et al. 2011). With a suitable barcoding system and bioinformatics tools, multiplex MPS is an efficient method for obtaining mitochondrial genome sequences from multiple specimens simultaneously (Jex et al. 2009b; Timmermans et al. 2010). However, recent published cnidarian mitochondrial genomes were obtained using Sanger sequencing coupled with a standard primer-walking protocol (Wang et al. 2013; Figueroa and Baco 2014; Lin et al. 2014; Kitahara et al. 2014), which is an expensive and time-consuming process (Ansorge et al. 1997).
Among cnidarian genera, Acropora is the most diverse of reef-building corals, representing about 30 % of the total number of scleractinian species (Wallace 1999; Veron 2000). Skeletal form and structure are crucial morphological traits in the systematics of this genus, and its species are categorized into 19 groups (Veron and Wallace 1984; Wallace and Wolstenholme 1998; Wallace 1999) with designations for identification purposes only that do not reflect taxonomic affinity (Veron and Wallace 1984). Sympatric ranges (Wallace 1999) and synchronous spawning (Harrison et al. 1984; Babcock et al. 1986) in many of these species offer high potential for hybridization in this genus (Babcock 1995; Willis et al. 1997; Hatta et al. 1999), which has been highlighted as a major contributor to the reticulate evolution of Acropora species (Hatta et al. 1999; reviewed in Willis et al. 2006). Acropora species boundaries have been studied intensively from morphological traits but rarely from a molecular perspective (van Oppen et al. 2000b, 2002; Wolstenholme et al. 2003), and efforts have been focused on reconstructing a molecular phylogeny of Acropora species using several potential genetic markers (Odorico and Miller 1997; van Oppen et al. 2001; Richards et al. 2013). Even so, their phylogeny remains unresolved due to the introgression of loci from ongoing hybridization (reviewed in Wallace and Willis 1994; reviewed in Willis et al. 2006), and accurate phylogenetic discrimination of species still represents a challenge for coral reef biologists.
Mitochondrial genes have been commonly used to study molecular relationships among scleractinian corals (Romano and Palumbi 1996; Chen et al. 2002; van Oppen et al. 2001; Fukami et al. 2000, 2008; Kitahara et al. 2010; Huang et al. 2011). However, the low evolutionary rates of mitochondrial genes and regions limit the determination of Acropora phylogeny (van Oppen et al. 1999; Chen et al. 2009). So far, the mitochondrial genome of A. tenuis is the only one been published and annotated; the length of the genome is nearly 18,300 bp, contains thirteen protein-coding genes and two ribosomal RNA genes (van Oppen et al. 2000a), and may contain valuable phylogenetic information for solving intraspecific relationships. Thus, multiplex MPS could provide a convenient and effective approach to study the molecular phylogenetics of this genus. The large amount of genomic data acquired from MPS could provide great insight into understanding the evolution and diversity of Acropora.
In this study, eleven Acropora species representing different species groups were sequenced using the Illumina Solexa platform for mitochondrial genomes. The low taxonomic sampling scheme and maternally inherited locus used in this study may not be good enough to draw a clear picture of the phylogeny of Acropora. However, our aim was to demonstrate the feasibility of sequencing scleractinian mitochondrial genomes using multiplex MPS technology for phylogenomic analysis. Our mitochondrial phylogenomic tree resolved relationships between most species except A. humilis, A. digitifera, and A. florida. We expect that this sequencing approach could be successfully applied to future cnidarian genomic studies.
Materials and methods
Acropora samples and DNA extraction
Eleven Acropora species representing ten morphological species groups (Wallace 1999) were collected from Kenting National Park and Green Island, Taiwan, and the Great Barrier Reef, Australia (Table 1). A fragment (2–3 cm) from an individual from each species was collected and preserved in a modified guanidine solution as described in Fukami et al. (2004). Total genomic DNA was extracted from the solution by standard phenol/chloroform (1:1) extraction followed by ethanol precipitation and resuspended in nuclease-free double-distilled water.
Long PCR-coupled mitochondrial genome sequencing
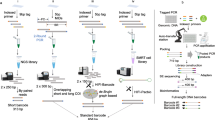
To obtain the complete mitochondrial genome from DNA samples, two pairs of long PCR primers designed for amplifying Acropora coral mitochondrial genomes were used (Monti12Slong-A/Monti16Slong-B and Monti16Slong-A/Monti12Slong-B; Tseng et al. 2005). The mitochondrial genome was amplified as two 9-Kb amplicons: Amplicon 1 was between 10,792–1538 and amplicon 2 between 1852–11,131 (the position in the reference sequence), forming an overlap in the putative control region and a gap in the rnl region. Long PCR was carried out in a 50-μl reaction mixture containing 1-μl DNA template (30–50 ng/μl), 0.2 μM of each primer, 20 mM of each dNTP, 10X LA PCR™ buffer, 2.5 mM MgCl2, and 2.5 U TaKaRa LA Taq™ (Takara Bio Inc.). PCR was carried out with a Thermo PX2 thermal cycler under the following conditions: one cycle of denaturing at 94 °C (1 min), annealing at 98 °C (10 s), and extension at 68 °C (15 min) for 30 cycles, followed by one cycle of final extension at 72 °C (10 min). PCR products were checked with a 1 % agarose gel to verify size and single-band yield. Gel purification was applied to remove nonspecific fragments, and final products were evaluated with a NanoDrop® Spectrophotometer. For library construction, amplicons of eleven samples were sheared and bar-coded for multiplexing purposes. Multiple libraries were pooled in a single-flow cell lane and sequenced on an Illumina Solexa Hi-Seq 2000 to generate 100-bp paired-end reads.
Sequence assembly and verification of consensus sequences
Paired reads were trimmed for adaptors and indexes by using Trimmomatic (Bolger et al. 2014). For the MPS dataset, sequence assembly was performed on CLC Genomics Workbench v4.5 software (CLC Bio, Denmark), an integrated platform for visualizing and analyzing MPS data. Sequence reads of eleven individuals were trimmed and mapped to the reference genome of Acropora tenuis (Genbank Accession No.: AF338425). Prior to mapping, the reference sequence was manipulated by removing the gap at the rnl region and shuffled to generate a continuous sequence. The assembled draft sequences were edited manually by adding ‘N’ to the gap and reshuffled into the correct order. The gap was then filled by conventional sequencing methods in order to generate complete mitochondrial genome sequences. Two datasets were used for mapping the Acropora mitochondrial genomes, including total output reads and reduced reads (i.e., 0.1 million reads of each species) to test the redundancy of output reads. Mapping results were verified by Sanger sequencing of the entire mitochondrial genomes of A. yongei and A. muricata. A total of 36 primers covering the entire mitochondrial genome were modified from Lin et al. (2011). PCR products were sequenced by an Applied Biosystems 3730 DNA analyzer and examined using SeqMan and MegAlign programs from Lasergene software (DNAstar Inc., USA). The differences between MPS and Sanger sequences were identified and used for verifying final consensus sequences. Sanger sequences were chosen when there were ambiguous sites observed between the two methods.
Multiple sequence alignment and phylogenetic analysis
The final consensus sequences of the eleven samples were aligned to the A. tenuis mitochondrial genome using MEGA 5.0 (Tamura et al. 2011). The repetitive sequences of the putative control region among Acropora species were trimmed. Annotated sequences were examined for informative and private sites using the Web server DIVEIN (Deng et al. 2010), which estimated genetic variation along the mitochondrial genome sequences. The proportion of nucleotide differences (p-distance) among genes and whole mitochondrial genomes was calculated. The model of nucleotide substitution that best fitted the sequence data was first estimated using Modeltest (Posada and Crandall 1998), followed by tree construction via maximum likelihood and neighbor-joining methods using MEGA 5.0 and a Bayesian method using MrBayes (Huelsenbeck and Ronquist 2001; Ronquist and Huelsenbeck 2003). Each analysis was run with 1000 bootstrap pseudoreplicates to estimate statistical support.
Results and discussion
Massively parallel sequencing and sequence assembly
Eleven Acropora mitochondrial genomes were sequenced using multiplex MPS technology on the Illumina Solexa platform and deposited in GenBank (NC_022823-NC_022833). Our approach was comprised of long-range PCR followed a species-specific indexing system to sequencing multiple mt genomes simultaneously. The four long PCR primers designed for amplifying the mitochondrial genomes of Montipora and Anacropora are applicable in Acropora species, as the rnl and rns region are generally conserved among the family Acroporidae (Tseng et al. 2005). Thus, bridging the gap in rnl regions posted to the sequence assembly was successful and less complicated than generating three amplicons for sequencing.
In total, 75,426,014 sequence reads were obtained from one Illumina lane (Table 2). Each bar-coded sample was successfully identified from total sequence reads with pipeline analysis software. Read number per sample varied from 3.0 to 9.3 million, with an average read length of 94 bases. Sequence trimming by removing low-quality sequences, ambiguous nucleotides, and sequences of specific length generated 74,393,870 high-quality reads, with average read length reduced to 81.07–89.12 bases. Sequence assembly greatly reduced the complexity of the assembling process with the A. tenuis reference genome. In the end, there were 73,142,848 reads with an average read length of 81.66–89.24 bases mapped to the A. tenuis reference sequence, representing 98.3 % of the trimmed reads. The high volume of sequence reads contributed 276,652,018–821,944,202 total bases to the sequence assembly, resulting in 96.9–99.2 % matching, and the mean depth of coverage (DOC) for the eleven mitochondrial genomes was approximately 12,000- to 44,000-fold. The overall genome mapping percentage can be generally considered high. Nevertheless, 0.8–3.1 % sequence reads could not be mapped to the reference genome. These singletons could be artifacts of the multiplex PCR due to the possible multi-template in the total genomic DNA samples (Takahiro 2003), but did not affect sequence mapping.
The sequencing output of each species from the pooled sample had various read counts that ranged from 4 to 14.4 %. The different proportion of output sequence reads was probably due to the unequal molarity of the DNA template. However, each sample produced abundant sequence reads with high genome coverage, providing strong statistical support for the sequence assembly. Even when we reduced the dataset to 0.1 million reads, coverage was still 500-fold and achieved high DOC values for sequence assembly, indicating the great sequencing output that can be analyzed by the Hi-seq 2000 platform. This could be optimized by increasing the sample size across multiple taxa using a large-scale indexing system (Smith et al. 2010; Tu et al. 2012), which would also increase the efficiency for individual sequencing runs.
Sequence analysis
Sanger sequencing of sample A.yongei_KTAcr4 successfully generated a complete A. yongei mitochondrial genome. However, two primers failed to amplify the putative control region of sample A.muricata_KTAcr14, so primers were redesigned to solve this (Table S1). The alignment of the MPS and Sanger sequences exhibited two main discrepancies in both samples: position 1443–1459 and 2358–2382 in A.yongei_KTAcr4 and position 1445–1457 and 2356–2380 in A.muricata_KTAcr14. The discrepancies are probably due to the noise of the sequence end in MPS as the two positions are approximately at the primer-binding sites where there was no sequence coverage. We also discovered the inconsistent length of three poly-G sites and two insertion sequences in the putative control region of A.muricata_KTAcr14. Guanine nucleotide deletions were found at positions 4564–4565, 5788, and 12,631–12,632, while insertions occurred between 12,283–12,396 and 12,593–12,614. The first insertion has been confirmed as a repeat sequence 114 bases long and the second is a short insert of 22 bases (van Oppen et al. 2002). These could be the consequence of assembly errors cause by repeated sequences (Miller et al. 2010). Because of inconsistencies between the two sequencing approaches, we Sanger-sequenced the putative control regions of twelve Acropora species to check repetitive structures and insert sequences, among which only the reference sequence A.florida_KTAcr2 and A.yongei_KTAcr4 did not contain repeat sequences at position 12,283–12,396 and only the reference sequence, and A.yongei_KTAcr4 did not have an insert sequence at position 12,593–12,614. However, another insertion of an 87-base repeat sequence was observed between 12,631 and 12,717 in A.hyacinthus_KTAcr39.
The consensus sequence verified by the Sanger method shows that the repetitive structures of the putative control region appears to be variable in Acropora species. Consequently, some samples with repeat sequences could not be assembled correctly as the repeat regions did not exist in A. tenuis. Therefore, we replaced the control region sequences generated by MPS with those generated by Sanger sequencing. With the exception of the sequence noise in primer-binding regions, the rough sequencing accuracy rates for A. muricata and A. yongei were 99.97 and 100 %, respectively. The comparison of the two sequencing approaches shows that the limitations of MPS were mainly associated with the assembly of repeat elements (i.e., homopolymers) and regions that may not exist in the reference genome.
General features of the Acropora mt genome
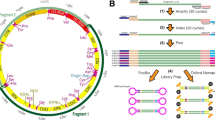
The circular mitochondrial genome sequences of eleven Acropora species ranged between 18,342 and 18,566 bases (Table 2), depending on the presence of the insertion sequence in the putative control region. Acropora hyacinthus had the largest mitochondrial genome due to three insertion sequences found in the putative control region. A. yongei had the smallest mitochondrial genome as no insert was observed in the putative control region. The base compositions of A, C, G, and T were similar in the twelve mitochondrial genomes: 24.9–25.1, 13.7–13.8, 24.2–24.4, and 36.8–37.0 %, respectively. There was no significant difference in codon usage for the thirteen protein-coding genes across the twelve Acropora species (Chi-square test, p > 0.05). The most frequently used codon was UUU (Phenylalanine), followed by UUA (Leucine) and GUU (Valine). The multiple sequence alignment of the twelve Acropora species displayed well-aligned sequences over most of the mitochondrial genome, except for the putative control region where repetitive structures and short insert sequences appeared. The twelve genomes have identical gene arrangement after annotation, indicating that the gene arrangement among Acropora species may be conserved. In the Scleractinia, 27 of the 29 reported complete mitochondrial genomes have identical gene order, but two cases of rearrangement are known in the genera Lophelia and Madrepora. Thus, changes in the gene arrangement of Scleractinia are relatively uncommon (Lin et al. 2014). The gene arrangements that we found in Acropora were conserved within the genus and have been categorized as Type SII gene arrangement, which is the dominant type within scleractinians (Lin et al. 2014).
There were 110 informative sites and 50 private sites in the Acropora mitochondrial genome, and the most variable region was the putative control region with 60 informative sites and 19 private sites. Other informative sites were located in cob (12 sites), nad3 (8 sites), igr3 (6 sites), nad5 (6 sites), rnl (4 sites), nad1 (3 sites), igr2 (2 sites), nad6 (2 sites), and igr1, nad2, nad4, rns, cox3, cox2, and igr13 genes, all of the latter having single sites (Table 3). A pairwise distance analysis of the mitochondrial genome gave p-distance values for the 32 loci (in decreasing order): trimmed putative control region (2.19 %), nad3 (0.61 %), igr1 (0.46 %), igr7 (0.32 %), igr2 (0.31 %), igr3 (0.24 %), cob (0.18 %), igr13 (0.16 %), nad5 (0.14 %), nad6 (0.14 %), cox2 (0.13 %), nad1 (0.11 %), igr6 (0.11 %), rns (0.09 %), nad2 (0.07 %), rnl (0.05 %), nad4 (0.05 %), cox3 (0.04 %), and cox1 (0.02 %); and the remaining thirteen loci (trnM, group I intron5, igr4, igr5, atp6, igr8, igr9, nad4L, igr10, igr11, trnW, igr12, and atp8) have 0 % of the p-distance values (Fig. 1). The overall p-distance estimated from the Acropora mitochondrial genome was 0.22 %, and the evolutionary rate based on the predicted divergence time of two geographically distinct Acropora species from Chen et al. (2009) was calculated to be approximately 0.011–0.063 % Mya−1, indicating a slow evolutionary rate of Acropora species. These results are consistent with previous studies based on a single gene or region comparisons (van Oppen et al. 1999; Fukami et al. 2000; van Oppen et al. 2001; Chen et al. 2009). Several mechanisms have been proposed to explain the extremely low substitution rate found in anthozoans, including a highly efficient DNA repair system, selection on mt DNA genes, and short divergence times (Shearer et al. 2002). These factors could be influence apparent substitution rates within anthozoan mitochondrial genomes at different levels and genetic divergences among species.
Phylogenetic relationships among the twelve Acropora species
Previous studies suggested a much stronger phylogenetic structure in datasets, excluding repeat regions (Van Oppen et al. 2001), so we excluded repeat regions within the control region and mitochondrial genome for phylogenetic analyses. After alignment, the length of the putative control region and mitochondrial genome was 1086 nucleotides and 18,569 nucleotides, respectively. Each tree was constructed under different substitute models: the HKY+G model for the putative control region dataset and HKY+G+I model for the whole mitochondrial genome dataset. The maximum likelihood (ML), Bayesian, and neighbor-joining trees constructed based only on the control region yielded topology similar to the tree constructed from the whole mitochondrial genome, but the latter displayed higher levels of resolution (Fig. 2a, b). This indicates that the data for the entire mitochondrial genome may provide slightly more phylogenetic information than the putative control region alone. Shearer et al. (2002) first revealed slow mitochondrial sequence evolution in Anthozoa by comparing three mitochondrial genes across different taxa. Recent work suggests that the mitochondrial genomes of some scleractinians evolve 5× slower than their nuclear genomes (Chen et al. 2009) and 50–100× slower than the mitochondrial genomes of most other animals (Hellberg 2006). This feature makes it difficult to apply genetic bar coding to anthozoans (Huang et al. 2008; McFadden et al. 2011) for resolving inter-species relationships within the genus Acropora (van Oppen et al. 2001; Richards et al. 2013).
a Maximum likelihood tree analyzed based on the whole mt genomes of twelve Acropora species. Branch values indicate the bootstrap value of 1000 replicates for maximum likelihood, Bayesian inference, and neighbor-joining, respectively. Clade nomenclature in blue on the right reflects Richards et al. 2013. Clade nomenclature in black on the left reflects van Oppen et al. 2001. Species groups are given after each species’ name. b Maximum likelihood tree analyzed based on the control region of twelve Acropora species. Bootstrap values higher than 50 are shown at the nodes
The twelve mitochondrial genomes used in this study belong to ten species groups. The ML tree revealed nine lineages among the twelve Acropora species, with eight out of nine lineages corresponding to their species groups except for the humilis and florida groups (Fig. 2a). A. tenuis and A. yongei, both belonging to the selago group, formed a monophyletic clade at the base with high supporting values concordant with previous studies (van Oppen et al. 2001; Richards et al. 2013). Acropora nasuta, A. divaricata, and A. muricata were included in clade III in the phylogenetic tree published by van Oppen et al. (2001), but our tree revealed a different pattern where A. murica clustered with A. aspera, A. florida, and A. humilis, which were in clade IV of van Oppen et al. (2001). In addition, the position of A. horrida differed from previous studies (Richards et al. 2013), instead being a basal lineage that is sister to A. hyacinthus in the present study. In addition, the mt genome does not have enough resolution to resolve the relationships among A. aspera, A. florida, and A. humilis. These three species belong to two species groups, which have close phylogenetic relationships to each other (Fukami et al. 2000; van Oppen et al. 2001; Wolstenholme 2004; Richards et al. 2013). This lack of resolution may indicate introgression or incomplete lineage sorting (van Oppen et al. 2001; Vollmer and Palumbi 2002; Wolstenholme et al. 2003; Nakajima et al. 2012; Richards et al. 2013). In our case, we cannot exclude these two evolutionary mechanisms, which may have acted on some Acropora species either simultaneously or at different times to contribute to the high similarities that we observed across Acropora mitochondrial genomes.
Conclusion
This study was not tending to resolve the phylogentic relationships among the genus Acropora which known to have reticulate evolution with low number of species represented in each morph groups. Instead, we demonstrated the feasibility of using MPS next-generation sequencing technology on scleractinian mitochondrial genomes. Comparing to traditional long PCRs with primer walking to obtain whole mitochondrial sequence, high-throughput sequencing method does not need to design species-specific primers (~20 primers) for primer walking during sequencing process, and it is more cost-efficient than traditional method. The barcoding system allowed the sequencing of multiple individuals simultaneously at lower processing costs and times while producing highly accurate sequence assemblies. Sequence errors and the difficulty in genome assembly were mainly due to homopolymeric regions and repetitive structures in the putative control region, and ambiguous sites still need to be verified by conventional Sanger sequencing. Our genome comparison and genetic analysis showed similar characteristics in mitochondrial genomes and low evolutionary rates in Acropora species. The relationships among closely related species remain unresolved because of the insufficient phylogenetic information presented in their mitochondrial genomes. Nonetheless, with more cnidarian mitochondrial genomes released, multiplex next-generation sequencing technology can provide an effective platform for molecular systematics and is practical for other cnidarian phylogenomic studies.
References
Ansorge W, Voss H, Zimmermann J (1997) DNA sequencing strategies. Wiley, New York
Babcock R (1995) Synchronous multispecific spawning on coral reefs: potential for hybridization and roles of gamete recognition. Reprod Fert Develop 7:943–950
Babcock RC, Bull GD, Harrison PL, Heyward AJ, Oliver JK, Wallace CC, Willis BL (1986) Synchronous spawnings of 105 scleractinian coral species on the Great Barrier Reef. Mar Biol 90:379–394
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics: btu170
Chen CA, Wallace CC, Wolstenholme J (2002) Analysis of the mitochondrial 12S rRNA gene supports a two-clade hypothesis of the evolutionary history of scleractinian corals. Mol Phylogenet Evol 23:137–149
Chen IP, Tang CY, Chiou CY, Hsu JH, Wei NV, Wallace CC, Muir P, Wu H, Chen CA (2009) Comparative analyzes of coding and noncoding DNA regions indicate that Acropora (Anthozoa: Scleractina) possesses a similar evolutionary tempo of nuclear vs. mitochondrial genomes as in plants. Mar Biotechnol 11:141–152
Cronn R, Liston A, Parks M, Gernandt DS, Shen R, Mockler T (2008) Multiplex sequencing of plant chloroplast genomes using Solexa sequencing-by-synthesis technology. Nucleic Acids Res 36:e122
Deng W, Maust BS, Nickle DC, Learn GH, Liu Y, Heath L, Sergei L, Pond K, Mullins JI (2010) DIVEIN: a web server to analyze phylogenies, sequence divergence, diversity, and informative sites. Biotechniques 48:405–408
Doorduin L, Gravendee B, Lammers Y, Ariyurek Y, Woeng TCA, Vrieling K (2011) The complete chloroplast genome of 17 individuals of pest species Jacobaea vulgaris: SNPs, microsatellites and barcoding markers for population and phylogenetic studies. DNA Res 18:93–105
Ekblom R, Galindo J (2011) Applications of next generation sequencing in molecular ecology of non-model organisms. Heredity 107:1–15
Faircloth BC, Sorenson L, Santini F, Alfaro ME (2013) A phylogenomic perspective on the radiation of ray-finned fishes based upon targeted sequencing of ultraconserved elements (UCEs). PLoS One 8:e65923
Feldmeyer B, Hoffmeier K, Pfenninger M (2010) The complete mitochondrial genome of Radix balthica (Pulmonata, Basommatophora), obtained by low coverage shot gun next generation sequencing. Mol Phylogenet Evol 57:1329–1333
Figueroa DF, Baco AR (2014) Complete mitochondrial genomes elucidate phylogenetic relationships of the deep-sea octocoral families Coralliidae and Paragorgiidae. Deep-Sea Res Part II 99:83–91
Fukami H, Omori M, Hatta M (2000) Phylogenetic relationships in the coral family Acroporidae, reassessed by inference from mitochondrial genes. Zool Sci 17:689–696
Fukami H, Budd AF, Levitan DR, Jara J, Kersanach R, Knowlton N (2004) Geographic differences in species boundaries among members of the Montastraea Annularis complex based on molecular and morphological markers. Evolution 58:324–337
Fukami H, Chen CA, Budd AF, Collins A, Wallace CC, Chuang YY, Chen C, Dai CF, Iwao K, Sheppard C, Knowlton N (2008) Mitochondrial and nuclear genes suggest that stony corals are monophyletic but most families of stony corals are not (Order Scleractinia, Class Anthozoa, Phylum Cnidaria). PLoS One 3:e3222
Harrison PL, Babcock RC, Bull GD, Oliver JK, Wallace CC, Willis BL (1984) Mass spawning in tropical reef corals. Science 223:1186–1189
Hatta M, Fukami H, Wang W, Omori M, Shimoike K, Hayashibara T, Ina Y, Sugiyama T (1999) Reproductive and genetic evidence for a reticulate evolutionary history of mass-spawning corals. Mol Biol Evol 16:1607–1613
Hellberg ME (2006) No variation and low synonymous substitution rates in coral mtDNA despite high nuclear variation. BMC Evol Biol 6:24
Huang D, Meier R, Todd PA, Chou LM (2008) Slow mitochondrial COI sequence evolution at the base of the metazoan tree and its implications for DNA barcoding. J Mol Evol 66:167–174
Huang D, Licuanan WY, Baird AH, Fukami H (2011) Cleaning up the “Bigmessidae”: molecular phylogeny of scleractinian corals from Faviidae, Merulinidae, Pectiniidae and Trachyphylliidae. BMC Evol Biol 11:37
Hudson ME (2008) Sequencing breakthroughs for genomic ecology and evolutionary biology. Mol Ecol Resour 8:3–17
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogeny. Bioinformatics 17:754–755
Jex AR, Hu M, Littlewood DTJ, Waeschenbach A, Gasser RB (2008) Using 454 technology for long-PCR based sequencing of the complete mitochondrial genome from single Haemonchus contortus (Nematoda). BMC Genomics 9:11
Jex AR, Waeschenbach A, Hu M, van Wyk JA, Beveridge I, Littlewood DTJ, Gasser RB (2009a) The mitochondrial genomes of Ancylostoma caninum and Bunostomum phlebotomum—two hookworms of animal health and zoonotic importance. BMC Genomics 10:79
Jex AR, Hall RS, Littlewood DTJ, Gasser RB (2009b) An integrated pipeline for next-generation sequencing and annotation of mitochondrial genomes. Nucleic Acids Res 382:522–533
Kitahara MV, Cairns SD, Stolarski J, Blair D, Miller DJ (2010) A comprehensive phylogenetic analysis of the Scleractinia (Cnidaria, Anthozoa) based on mitochondrial CO1 sequence data. PLoS One 5:e11490
Kitahara MV et al (2014) The “naked coral” hypothesis revisited—evidence for and against scleractinian monophyly. PLoS One 9:e94774
Lin MF, Luzon KS, Licuanan WY, Ablan-Lagman MC, Chen CA (2011) Seventy-four universal primers for characterizing the complete mitochondrial genomes of scleractinian corals (Cnidaria; Anthozoa). Zool Stud 50:513–524
Lin MF, Kitahara MV, Luo H, Tracey D, Geller J, Fukami H, Miller DJ, Chen CA (2014) Mitochondrial genome rearrangements in the scleractinia/corallimorpharia complex: implications for coral phylogeny. Genome Biol Evol 6:1086–1095
McFadden CS, Benayahu E, Thoma JN, Nevarez PA, France SC (2011) Limitations of mitochondrial gene barcoding in Octocorallia. Mol Ecol Resour 11:19–31
Miller JR, Koren S, Sutton G (2010) Assembly algorithms for next-generation sequencing data. Genomics 95:315–327
Nakajima Y, Nishikawa A, Iguchi A, Sakai K (2012) The population genetic approach delineates the species boundary of reproductively isolated corymbose acroporid corals. Mol Phylogent Evol 63:527–531
Odorico DM, Miller DJ (1997) Variation in the ribosomal internal transcribed spacers and 5.8 s rDNA among five species of Acropora (Cnidaria; Scleractinia): patterns of variation consistent with reticulate evolution. Mol Biol Evol 14:465–473
Posada D, Crandall KA (1998) Modeltest: testing the model of DNA substitution. Bioinformatics 14:817–818
Richards Z, Miller DJ, Wallaces CC (2013) Molecular phylogenetics of geographically restricted Acropora species: implications for threatened species conservation. Mol Phylogenet Evol 69:837–851
Romano SL, Palumbi SR (1996) Evolution of scleractinian corals inferred from molecular systematics. Science 271:640–642
Ronquist F, Huelsenbeck JP (2003) MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Shearer TL, van Oppen MJH, Romano SL, Wörheide G (2002) Slow mitochondrial DNA sequence evolution in the Anthozoa (Cnidaria). Mol Ecol 11:2475–2487
Shendure J, Ji H (2008) Next-generation DNA sequencing. Nature biotechnol 26:1135–1145
Smith AM, Heisler LE, St Onge RP, Farias-Hesson E, Wallace IM, Bodeau J, Harris AN, Perry KM, Giaever G, Pourmand N, Nislow C (2010) Highly-multiplexed barcode sequencing: an efficient method for parallel analysis of pooled samples. Nucleic Acids Res 38:e142
Takahiro K (2003) Bias and artifacts in multitemplate polymerase chain reactions (PCR). J Biosci Bioeng 96:317–323
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Timmermans M, Dodsworth S, Culverwell CL, Bocak L, Ahrens D, Littlewood DT, Pons J, Vogler AP (2010) Why barcode? High-throughput multiplex sequencing of mitochondrial genomes for molecular systematics. Nucleic Acids Res 38:e197
Tseng CC, Wallace CC, Chen CA (2005) Mitogenomic analysis of Montipora cactus and Anacropora matthai (Cnidaria; Scleractinia; Acroporidae) indicates an unequal rate of mitochondrial evolution among Acroporidae corals. Coral Reefs 24:502–508
Tu J, Ge Q, Wang S, Wang L, Sun B, Yang Q, Bai Y, Lu Z (2012) Pair-barcode high-throughput sequencing for large-scale multiplexed sample analysis. BMC Genomics 13:43
van Oppen MJH, Willis BL, Miller DJ (1999) Atypically low rate of cytochrome b evolution in the scleractinian coral genus Acropora. Proc R Soc Lond B 266:179–183
van Oppen MJH, Catmull J, McDonald BJ, Hislop NR, Hagerman PJ, Miller DJ (2000a) The mitochondrial genome of Acropora tenuis (Cnidaria; Scleractinia) contains a large group I intron and a candidate control region. J Mol Evol 55:1–13
van Oppen MJH, Willis BL, Vugt HWJAV, Miller DJ (2000b) Examination of species boundaries in the Acropora cervicornis group (Scleractinia, Cnidaria) using nuclear DNA sequence analyzes. Mol Ecol 9:1363–1373
van Oppen MJH, McDonald BJ, Willis BL, Miller DJ (2001) The evolutionary history of the coral genus Acropora (Scleractinia, Cnidaria) based on a mitochondrial and a nuclear marker: reticulation, incomplete lineage sorting, or morphological convergence? Mol Biol Evol 18:1315–1329
van Oppen MJH, Willis BL, Van RT, Miller DJ (2002) Spawning times, reproductive compatibilities and genetic structuring in the Acropora aspera group: evidence for natural hybridization and semi-permeable species boundaries in corals. Mol Ecol 11:1363–1376
Veron JEN (2000) Corals of the world. Australian Institute of Marine Science, Townsville
Veron JEN, Wallace CC (1984). Scleractinia of eastern Australia, Part V. Family Acroporidae AIMS Monogr Ser 6: 485
Vollmer SV, Palumbi SR (2002) Hybridization and the evolution of coral reef diversity. Science 296:2023–2025
Wallace CC (1999) Staghorn corals of the world: a revision of the genus Acropora. CSIRO Publishing, Collingwood
Wallace CC, Willis BL (1994) Systematics of the coral genus Acropora: implications of new biological findings for species concepts. Annu Rev Ecol Syst 25:237–262
Wallace CC, Wolstenholme J (1998) Revision of the coral genus Acropora (Scleractinia: Astrocoeniina: Acroporidae) in Indonesia. Zool J Linn Soc 123:199–384
Wang M, Sun J, Li J, Qiu JW (2013) Complete mitochondrial genome of the brain coral Platygyra carnosus. Mitochondrial DNA 24:194–195
Williams ST, Foster PG, Littlewood DTJ (2014) The complete mitochondrial genome of a turbinid vetigastropod from MiSeq Illumina sequencing of genomic DNA and steps towards a resolved gastropod phylogeny. Gene 533:38–47
Willis BL, Babcock RC, Harrison PL, Wallace CC (1997) Experimental hybridization and breeding incompatibilities within the mating systems of mass spawning reef corals. Coral Reefs 16:S53–S65
Willis BL, van Oppen MJH, Miller DJ, Vollmer SV, Ayre DJ (2006) The role of hybridization in the evolution of reef corals. Ann Rev Ecol Evol Syst 37:489–517
Wolstenholme JK (2004) Temporal reproductive isolation and gametic compatibility are evolutionary mechanisms in the Acropora humilis species group (Cnidaria; Scleractinia). Mar Biol 144:567–582
Wolstenholme JK, Wallace CC, Chen CA (2003) Species boundaries within the Acropora humilis species group (Cnidaria; Scleractinia): a morphological and molecular interpretation of evolution. Coral Reefs 22:155–166
Acknowledgments
This project was funded by the National Science Council (2010–2012), Academia Sinica Thematic Grant (2006–2010), and the Biodiversity Research Centre, Academia Sinica (BRCAS) to CAC. We would like to thank the staffs of Penghu Marine Biological Research Centre (PMBRC)-BRCAS joint Marine Laboratory, BRCAS Green Island Reef Laboratory (GIRL) Green Island Marine Station, and Kenting National Park for assistance with fieldwork. Special thanks go to CREEG members and the David Miller Group for intellectual inputs to this paper, and to Tyler McCraney and Buford Pruitt, Jr. for English language editing. We also greatly thank three anonymous reviewers and the communicating editor for useful comments and suggestions to improve the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by C. Riginos.
Shang-Yin Vanson Liu and Chia-Ling Carynn Chan are co-first authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, SY.V., Chan, CL.C., Hsieh, H.J. et al. Massively parallel sequencing (MPS) assays for sequencing mitochondrial genomes: the phylogenomic implications for Acropora staghorn corals (Scleractinia; Acroporidae). Mar Biol 162, 1383–1392 (2015). https://doi.org/10.1007/s00227-015-2657-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00227-015-2657-1