Abstract
Studies of gene expression are common in toxicology and provide important clues to mechanistic understanding of adverse effects of chemicals. Most prior studies have been performed in a single strain or cell line; however, gene expression is heavily influenced by the genetic background, and these genotype-expression differences may be key drivers of inter-individual variation in response to chemical toxicity. In this study, we hypothesized that the genetically diverse Collaborative Cross mouse population can be used to gain insight and suggest mechanistic hypotheses for the dose- and genetic background-dependent effects of chemical exposure. This hypothesis was tested using a model liver toxicant trichloroethylene (TCE). Liver transcriptional responses to TCE exposure were evaluated 24 h after dosing. Transcriptomic dose–responses were examined for both TCE and its major oxidative metabolite trichloroacetic acid (TCA). As expected, peroxisome- and fatty acid metabolism-related pathways were among the most dose–responsive enriched pathways in all strains. However, nearly half of the TCE-induced liver transcriptional perturbation was strain-dependent, with abundant evidence of strain/dose interaction, including in the peroxisomal signaling-associated pathways. These effects were highly concordant between the administered TCE dose and liver levels of TCA. Dose–response analysis of gene expression at the pathway level yielded points of departure similar to those derived from the traditional toxicology studies for both non-cancer and cancer effects. Mapping of expression–genotype–dose relationships revealed some significant associations; however, the effects of TCE on gene expression in liver appear to be highly polygenic traits that are challenging to positionally map. This study highlights the usefulness of mouse population-based studies in assessing inter-individual variation in toxicological responses, but cautions that genetic mapping may be challenging because of the complexity in gene exposure–dose relationships.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gene expression profiling is widely used in the field of toxicology and provides important insights into molecular changes and potential mechanisms of adverse health effects from chemical exposure (Burczynski et al. 2000; Harrill and Rusyn 2008). Transcriptomic signatures of chemical exposures have been widely used in chemical safety evaluation and large reference databases exist to enable comparative analyses and predictive modeling (Fostel 2008; Ganter et al. 2005; Uehara et al. 2010). In addition, many studies demonstrate the value of transcriptomic data-derived dose–response information for both hazard identification and quantitative risk assessment (Andersen et al. 2008; Thomas et al. 2013a) and pathway-based dose–response analysis of transcriptomic data displayed concordance with traditional apical endpoints (Thomas et al. 2007, 2011, 2013a; Zhou et al. 2017). For instance, a key step in deriving exposure limits in traditional risk assessments involves estimating a “point of departure” (POD) from a chronic experimental animal study—e.g., a lifetime dose rate in mg/kg-day estimated to be without a biologically significant, adverse toxic effect. These recent studies have demonstrated that “transcriptional” PODs—i.e., acute dose levels at which gene expression pathways begin to be perturbed—can serve as surrogates for traditional PODs with an accuracy within an order of magnitude. Indeed, more broadly, toxicogenomics data are making their way into decision-making for both drugs and environmental chemicals (Sauer et al. 2017; Xu et al. 2016).
The majority of available gene expression data in toxicology have been collected in genetically homogenous (e.g., a single strain or cell line) or undefined (e.g., outbred strains or mixtures of cells from multiple individuals) model systems. However, it is well established that gene expression, both basal and disease/treatment-perturbed, is heavily impacted by genetic variation among individuals (Bradford et al. 2011; Bystrykh et al. 2005; Chesler et al. 2003; Gatti et al. 2007; Harrill et al. 2009a; Schadt et al. 2008, 2003). Early studies of expression quantitative trait loci (eQTL) were conducted in panels of inbred mouse strains or human cell lines where population stratification was a challenge (Gatti et al. 2009) and recent studies in more heterogeneous mouse models such as Diversity Outbred (DO) and Collaborative Cross (CC) show greater promise of characterizing and replicating eQTLs (Crowley et al. 2015).
Studies of chemical effects using population-based models, both in vivo and in vitro, are increasingly common (Abdo et al. 2015; Church et al. 2015; Cichocki et al. 2017; Eduati et al. 2015; French et al. 2015; Luo et al. 2018; Mosedale et al. 2017; Venkatratnam et al. 2017). Powered by the balanced genetic diversity represented in the DO (Churchill et al. 2012) and CC (Threadgill et al. 2011) populations, these mouse models enable exploration of causal variants driving molecular variation that result in phenotypic differences (Harrill and McAllister 2017). Furthermore, characterization of the genetics-dependent and -independent transcriptional responses to chemical exposure is valuable for elucidating the extent of and potential mechanisms underlying such variation (Church et al. 2015; Harrill et al. 2009a; Mosedale et al. 2017).
The ensuing challenge in toxicology and environmental health is to characterize dose–response relationships and gene expression in studies of chemical exposure while harnessing the power of genetic variation. The complexity and cost of population-based studies that include multiple doses and parallel characterization of multiple dimensions of toxicokinetics (absorption, distribution, metabolism, and excretion of toxic substances), toxicodynamics (biological responses elicited at the target tissue site), and inter-individual variation present unique challenges and may be best addressed using case studies of well-characterized toxicants (Rusyn et al. 2010). Thus, we chose trichloroethylene (TCE), a ubiquitous contaminant and a known carcinogen in both humans and rodents (Rusyn et al. 2014). TCE is a chemical with known variation in both toxicokinetics (Chiu et al. 2014) and toxicodynamics (Bradford et al. 2011). The objective of this study was to evaluate and characterize genetic background-, dose-, and interaction effects of TCE on liver gene expression, and to determine variation in dose–pathway relationships in a large genetically diverse mouse population. Liver transcriptomic data from 50 strains of CC mice that were treated acutely with one of the four doses of TCE were modeled to identify genes and pathways that exhibited significant strain, dose, or strain by dose interaction effects. In addition, we conducted genome-wide linkage mapping to identify loci associated with variation in liver TCA levels.
Materials and methods
Animals and treatments
The in-life portion of the study and tissue collection has been detailed in (Venkatratnam et al. 2017). In brief, adult male mice (8–12 weeks old) from 50 CC strains were orally administered a single dose of 0, 24, 80, 240, or 800 mg/kg TCE (Sigma Aldrich, St. Louis, MO) in 5% Alkamuls EL-620 vehicle (Solvay, Deptford, NJ). Each strain had 5 mice, with one mouse randomly assigned to each dose group to maximize the statistical power of the study by maximizing the number of strains tested (Kaeppler 1997). Mice were sacrificed 24 h after treatment, and tissues were flash frozen in liquid nitrogen and stored at − 80 °C until analysis.
TCA level in liver
Analyses were performed by a modification of US EPA method 552.2 (Domino et al. 2003) as detailed in (Venkatratnam et al. 2017). Data on TCA levels in liver across all CC strains were reported in (Venkatratnam et al. 2017).
RNA extraction and sequencing
Left-lobe of livers was pulverized with pestle and mortar pre-chilled in liquid nitrogen. RNA was extracted from ~ 20 mg of pulverized tissue using RNeasy Qiagen mini Kit (Qiagen, Valencia, CA). Concentration of extracted RNA was measured using NanoDrop 2000 (Thermo Fisher Scientific, Wilmington, DE). RNA libraries were prepared using Illumina TruSeq Stranded Total RNA kit (Illumina, San Diego, CA) with Ribo-Zero Gold rRNA removal kit (Illumina). The quality of extracted RNA was tested using Fragment Analyzer (Advanced Analytical Technologies, Ankeny, IA). RNA Quality Number (RQN) equal or greater than 7 was used as a cutoff for RNA quality to further proceed with the preparation of the sequencing libraries. RNA-Seq of 50 base-pair single-end reads was conducted using Illumina HiSeq 2500 instrument (Illumina) at the depth of ~ 50 million reads/sample. Raw reads were trimmed for any sequencing adaptors and low-quality bases using Trimmomatic (Bolger et al. 2014). Reference genomes and inferred marker founder origin for CC mice were downloaded from University of North Carolina Systems Genetics Core Facility (http://csbio.unc.edu/CCstatus/index.py?run=Pseudo). Filtered reads were mapped to each of the corresponding CC reference genome using TopHat version 2.0.3 (Kim et al. 2015). Resulting alignments were re-mapped to reference mm10 assembly coordinates using lapels (https://github.com/shunping/lapels). HTSeq (Anders et al. 2015) was used to generate raw read counts per gene using intersection-non-empty parameter to account for ambiguous read mappings. Differential gene expression tests were then performed with DESeq2 (Love et al. 2014).
Statistical analyses
Normalized count data on 23,948 genes for 246 combinations for four TCE and one vehicle treatments from CC strains were generated with R (version 3.3) package DESeq2 as detailed above. Genes with < 2 counts across all treatments were removed, leaving 19,870 genes for analysis. The “dose” vector was linearized by first replacing the vehicle (i.e., zero) TCE dose with 8 (the average distance between doses was 1/3 the prior dose) followed by a natural log transformation. DESeq2 was used to derive p values and adjusted p values for the main effects of dose and strain, as well as their interaction. In brief, counts were first modeled as counts ~ strain + ln(dose). Genes exhibiting a linear dose effect were identified with a likelihood ratio test comparing a reduced model of counts ~ strain. Similarly, genes exhibiting a strain effect were identified with a likelihood ratio test comparing a reduced model of counts ~ ln(dose) to the full model. Lastly, genes with an interaction effect were identified comparing a reduced model of counts ~ strain + ln(dose) to a full model of counts ~ strain + ln(dose) + strain × ln(dose). In a similar fashion, these analyses were repeated using the natural log of measured liver TCA metabolite concentration in nmol/g (with a + 1 offset to guard against taking logarithm of zero) as a predictor instead of TCE dose. Differential expression gene results were subjected to multiple comparison adjustments by computing Benjamini–Hochberg false discovery q values as implement in DESeq2. Due to the large number of significant findings, stringent significance criteria (q < 0.001) and absolute counts (read counts > 10) were used as criteria for further analysis.
Pathway analyses
Pathway enrichment analysis was conducted using the Database for Annotation, Visualization, and Integrated Discovery (DAVID) (Dennis et al. 2003). The top 3000 or fewer genes with Bonferroni-corrected p values (q < 0.001), ordered based on the q value significance, were uploaded to DAVID to identify pathways influenced by dose-, strain-, or interaction effects.
Comparison of transcriptomic points of departure doses with apical data
For dose–response expression studies, the BMDExpress software (Yang et al. 2007) has been used to evaluate PODs for expression changes of individual genes with exposure. Specifically, this software estimates the “benchmark dose” (BMD) at which gene expression changes relative to controls occur by fitting a parametric model to the dose–expression data. Although the current study has a large sample size, the sample size of 5 per strain presents a challenge in computing strain-specific benchmark doses, and in fact many of the models provided in BMDExpress would suffer from extreme overfitting with such small sample sizes. To best accord with our differential expression analyses and to benefit from the variance shrinkage models used in expression analyses, we devised the following approach to compute a strain-specific pathway BMD analogue. First, we used the strain-specific linear modeling routine from DESeq2 as described to obtain slope (\({\hat {\beta }_1}\)) and intercept (\({\hat {\beta }_0}\)) estimates from the software’s modeling of logarithmic gene expression, as well as the Wald-like t statistic (which utilizes variance shrinkage estimation). Next, an artificial model standard deviation \(\hat {\sigma }\) was computed from the model in order to be consistent with the reported p value. Specifically, if t* represents the shrunken t statistic, \(\hat {\sigma }=\frac{{{{\hat {\beta }}_1}}}{{{t^*}}}\sqrt {\mathop \sum \limits_{i} {{({x_i} - \bar {x})}^2}}\) for x = ln(dose). Finally, we have the per-gene BMD = ln(control dose)+\({\text{sign}}\left( {{{\hat {\beta }}_1}} \right)~\hat {\sigma }/{\hat {\beta }_1}\). The final BMD reflects the point at which the dose–response fit is estimated to exhibit a 1 standard-deviation departure from control expression, using shrunken estimates of variation that are obtained from the DESeq2 modeling. To compute a pathway (gene set) BMD, we used the median BMD of all genes assigned to the pathway.
As described in the introduction, previous studies have suggested that transcriptomic PODs correlated with those for apical endpoints, and that therefore transcriptional BMD values have the potential to serve as POD for quantitative risk assessment (Thomas et al. 2011). We therefore compared transcriptomic BMDs with apical BMDLs used for liver effects in the U.S. EPA’s Toxicological Review for TCE (U.S. EPA 2011a, b). Specifically, U.S. EPA, in their quantitative evaluation of the effects of TCE on the liver, selected increased liver/body weight ratio BMDLs in mice and rats for the PODs for liver non-cancer effects, and the BMDL for increased carcinomas in mice for the POD for liver cancer effects. Because apical endpoint PODs were derived from both rats and mice, each with differing toxicokinetics, we standardized all dose units to human equivalent doses (HEDs) based on equivalent liver oxidative metabolism, using the most up-to-date multi-species physiologically based pharmacokinetic (PBPK) model (Chiu and Ginsberg 2011; Chiu et al. 2009). Median estimates of each internal dose metric from Chiu et al. (2009) were used. An additional reason for this standardization is that margins of exposure can be readily computed and compared based on the human equivalent dose. Apical endpoint HEDs were then compared to median transcriptional BMD values.
Genetic mapping and eQTL analyses
In principle, mapping of traits in CC lines can be performed by analysis of variance, using for each locus the ancestral origin from each of the 8 founding CC lines as a categorical predictor. In practice, for many loci slight uncertainties of CC line origin remain, due to incomplete information on crossover events in the CC breeding outcomes. Hidden Markov modeling enables probabilistic statements of the parental line of origins for each locus, expressed as probabilities (summing to 1.0) for the eight founder lines. For the vast majority of loci and CC lines, a very large probability (greater than 0.95) is placed on the most likely parental origin for the locus. We performed regression modeling using trait as a response, and the probability vector as a predictor for a model with 8 degrees of freedom. Each trait, whether a phenotype such as TCA level, or an expression trait, was first transformed using rank-inverse normal transformation (GTEx Consortium 2015) to ensure robustness to outliers. eQTL analysis was performed separately for each dosage group using the R package DOQTL1, and results were compared to direct likelihood ratios computed using regression F statistics as described below to ensure correct computation. Expression traits were used only if they had both a mean number of reads ≥ 5 and a non-zero read proportion of at least 10%.
To accord with standard linkage mapping, the log10 likelihood ratio (LOD score) for the fitted model vs. the null was used to represent mapping evidence. At each locus, a p value can be obtained from LOD scores via Chi-square testing and standard likelihood ratio theory. However, initial investigation by permutation showed that p values based on normal theory regression F statistics were superior, i.e., were more nearly uniform under the null, and so were used for multiple comparisons as described below.
To facilitate multiple comparisons and to acknowledge that cis-eQTLs are more common than trans-eQTLs, we obtained separate cis- and trans p values for each expression trait as follows. First, 1000 permutations of a normally distributed “phenotype” were performed, and the linkage F statistics were computed across the genome. As the rank-inverse normal transformation produces identical transformed values for any trait (provided there are no ties), the permutations formed a generic set of sampling outcomes that are applicable to any quantitative trait. For each permutation, the maximum F statistic was recorded for each chromosome. Thus, for each chromosome we obtained a set of 1000 maxima on the chromosome (cis) and 1000 maxima on the remaining chromosomes (trans). For each of these 40 sets of permutations (20 chromosomes, both cis and trans), the maximum F statistic was modeled via maximum likelihood fitting to a Gompertz extreme value distribution, providing the basis for cis- and trans p values for each expression trait. By construction, these p values control for multiple comparisons across different loci. In order to control for multiple comparisons across different genes, each of the sets of cis p values and trans p values were adjusted using the q value package in R, resulting in false discovery q values for both cis and trans. For non-expression traits such as TCA levels, the overall maximum F statistic distribution was used in the extreme value modeling to obtain a genome-wide p value.
Results
Dose-, strain-, and interaction-related effects of TCE on liver gene expression
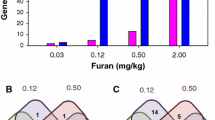
Previous studies of inter-strain variation of TCE-induced responses demonstrated that liver transcriptomic responses are strongly dependent on genetic background and that peroxisome proliferator-activated receptor-associated pathways represented some of the most pronounced genetic background-dependent molecular effects of TCE treatment in mouse liver (Bradford et al. 2011). In this study, to elucidate how TCE responses differ by genetic background- and dose–expression relationships, we used liver gene expression data from 50 CC strains and 4 dose groups (plus vehicle). Dose, strain, and interaction effects were modeled for each transcript for both TCE dose and liver levels of TCA in each mouse. Exemplar plots of genes that were significant for one or several of these relationships, and are relevant to the effects of TCE, are displayed in Fig. 1. Panel A shows the dose–response relationships for the administered dose of TCE and panel B for a correlation with liver TCA level in each individual mouse. We observed that expression of UDP-glucuronosyltransferase family 2 member A3 (Ugt2a3) was down-regulated with TCE dose, but this effect on gene expression was strain independent, with no dose by strain interaction effect. The effects of TCE on UDP-glucuronosyltransferase enzymes is not well characterized, but it is known that glucuronidation of trichloroethanol, a major oxidative metabolite of TCE, is a detoxification mechanism (Chiu et al. 2007). Baseline expression levels of alcohol dehydrogenase 1 (Adh1) varied dramatically by strain but did not exhibit a significant dose–response to TCE or a dose by strain effect. Alcohol dehydrogenases are involved in the biotransformation of TCE metabolites chloral and chloral hydrate to trichloroethanol (Lash et al. 2014) and our finding of a high degree of inter-individual variation is consistent with previous observations in both humans and mice (Bronley-DeLancey et al. 2006; Venkatratnam et al. 2017). In contrast to the Ugt2a3 and Adh1 examples, the expression of acyl-CoA thioesterase 7 (Acot7) exhibited not only a strong baseline strain effect, but also a strain by dose interaction effect. Acot7 is a PPARα-responsive gene (Rakhshandehroo et al. 2010). Indeed, PPARα-signaling plays a critical role in the effects of TCE in rodent liver (Rusyn et al. 2014). Of note, all three example genes depicted in Fig. 1 exhibited highly significant dose, strain, and interaction effects regardless whether TCE dose, or liver TCA concentrations, were used as the “dose,” demonstrating that at least for these exemplars, that TCA plays a key role in transcriptional responses to TCE in mouse liver, likely through its agonism to PPARα (Maloney and Waxman 1999).
Examples of genes (Ugt2a3, Adh1, and Acot7) that were affected by exposure to TCE in mouse liver in genetic background-, dose- or interaction-dependent manner. Top panel a shows correlation with administered TCE dose; bottom panel b is correlation with liver TCA levels at 24 h after dosing. Each circle represents gene expression in a CC mouse. Each line represents a linear dose–response fit for each CC strain. The y axis represents normalized counts of the expression. The x axis in the top panel represents administered TCE dose (mg/kg) and in the bottom panel represents liver TCA levels (nmol/g) in the CC population. False discovery q values (q ≤ 0.001) for dose and strain main effects, as well as their interaction, are displayed in each box
Overall, we found that 5285 transcripts were significant (after multiple testing correction as described) for the effect of TCE dose, 11,820 for the effect of strain, and 2140 for the interaction between the two (Fig. 2, left column of Venn diagrams). When liver TCA was used as an input into the model, 4769 transcripts were significant for TCA, 13,920 for strain, and 2242 for interaction (Fig. 2, right column). Interestingly, a very small number of transcripts were purely dose-dependent, without an effect of strain or interaction, which is less than 1% of the transcriptome. In contrast, the effect of strain on the transcriptome was a major factor, which is greater than 50% of all transcripts that were mapped. This observation is consistent with the dominant effect of genetic variation on transcription in the liver (Gatti et al. 2007; Schadt et al. 2008), and general findings on the impact of genetic variation in gene expression regulation (GTEx Consortium 2013, 2015).
Venn diagrams representing total number of transcripts from the transcriptome that are strongly influenced by genetic background-, dose-, or interaction effects with administered TCE dose (left panel) or liver TCA (right panel) as dose inputs. Numbers within each sector of circles represent either unique or common transcripts. Percentages represent the total percent of transcripts from the liver transcriptome that are common between TCE dose and TCA level analyses
Whether analyzing the effects of TCE dose on expression in liver, or the relationship between liver TCA concentration and liver expression, we found significant overlap in expression signatures. This finding is not completely unexpected, as there is strong correlation (r = 0.78, ρ = 0.86) between TCE dose and liver TCA levels (Fig. 3a). The importance of including the dose–response considerations in the analysis of the population-wide transcriptional response to toxicity is illustrated in Fig. 3b. While there is a significant positive correlation (r = 0.49, p < 0.001) between the number of significantly perturbed transcripts and liver TCA at the highest TCE dose level, this relationship is highly strain-variable whereby many strains with the highest liver TCA were not the most “responsive” transcriptionally.
Concordance between TCE dose and liver TCA levels based on gene- and pathway-based analyses. a Scatter plot showing individual animal’s liver TCA (nmol/g) as compared to the administered TCE dose (mg/kg). The inset shows the results of the correlation analysis of these data. b A relationship between liver TCA levels and the number of significantly (q < 0.05) perturbed transcripts by the TCE (800 mg/kg) in each CC strain. The inset shows the results of the correlation analysis of these data. c Concordance in pathways that were significantly (q < 0.01) associated with dose (top), strain (middle), or interaction (bottom) for the analyses where TCE dose (x axis) or liver TCA (y axis) were used as dose inputs. Each dot is a KEGG or GO pathway/category. Pathways that were significant only for TCE are colored black, only for liver TCA are colored red, and pathways that were significant for both are colored green. (Color figure online)
Dose-, strain-, and interaction-related effects of TCE on liver pathways
Next, we examined the molecular pathways perturbed by TCE in mouse liver in a dose-, strain-, or interaction-dependent manner. To examine concordance in pathways among TCE dose and liver TCA, significant Gene Ontology and KEGG pathways/category enrichment was examined using DAVID/EASE (q value < 0.001), for lists of transcripts with significant dose (Fig. 3c), strain (Fig. 3d), or interaction (Fig. 3e) effects. Most pathways for the dose and strain effects were shared between TCE dose and liver TCA analyses (marked in green), and their level of significance was highly concordant (slopes close to 1).
A complete list of pathways from significant dose-related transcripts is provided in Supplemental Tables 1–5. Pathways with strong TCE dose–response relationships included lipid and fatty acid metabolism. Most of these were also significant for the strain effect and they are closely related to PPARα signaling, a finding consistent with our understanding of the major molecular effects of TCE in the rodent liver (Rusyn et al. 2014). Another prominent group of dose–responsive pathways was the effect on cell–cell adhesion and gap junctional intracellular communication, also consistent with previous findings that exposure to TCE and TCA inhibits gap junctional communications in mouse hepatocytes (Klaunig et al. 1989).
While there was a greater number of genes that exhibited strain-specific changes in gene expression, enrichment analysis yielded fewer discernable significantly enriched pathways. Two translation-related pathways were significant for strain alone and not dose or interaction. Pathways that were both strain- and dose-dependent showed that the large part of TCE dose–response response is highly dependent on the genetic background.
Comparison of PODs for transcriptional and apical effects of TCE in mouse liver
Next, we sought to compare transcriptomics-derived dose–response effects of TCE in this acute exposure study in genetically diverse CC mice to the traditional apical endpoint-derived PODs for the same tissue but in other animal models. Specifically, we based these comparisons on the effects of TCE on liver transcriptome in B6C3F1 strain (Zhou et al. 2017) and the liver non-cancer and cancer endpoints used by U.S. EPA to derive toxicity values (U.S. EPA 2011a, b). For this comparison, we converted both types of PODs to human equivalent doses using a multi-species PBPK models, as described in “Materials and methods”, and the results are shown in Fig. 4a. Because of the lack of PBPK models specifically for CC mice, we used the median estimates for mice from the previous multi-species PBPK model, which was calibrated using data from Swiss and B6C3F1 mice (Chiu et al. 2014; Evans et al. 2009), along with the corresponding human PBPK model. The transcriptional PODs covered the same range of human equivalent doses as the apical endpoints, with the most sensitive median BMDL (KEGG_mmu00071, fatty acid degradation) being nearly the same as the most sensitive apical endpoint-derived BMDL (B6C3F1 mouse liver carcinomas). Overall, the median transcriptional BMDLs across all pathways in CC mice were within 10-fold of the apical PODs for TCE effects in the liver.
Comparison of point of departures (PODs) across significantly (q < 0.001) perturbed pathways due to genetic background-, dose-, and interaction effects in the CC model with apical endpoints from sub-chronic or chronic TCE studies in B6C3F1. Box plots represent PODs, converted to human equivalent dose (mg/kg-day) using mouse and human physiologically based pharmacokinetic models, for the following: apical endpoints (black); transcriptional PODs for B6C3F1 mice from Zhou et al. (2017) (green); transcriptional PODs for CC mice aggregated across pathways and strains (blue, panel a only); transcriptional PODs for individual pathways, aggregated across CC strains (red, panel a); or transcriptional PODs for individual CC strains, aggregated across pathways (red, panel b). See Supplemental Table 6 for full listing of abbreviations used. Vertical gray lines identify human equivalent doses corresponding to each administered dose. (Color figure online)
A corollary of this analysis is a question of whether data from the CC population are more informative as compared to the analysis of the dose–response gene and pathway effects of TCE in B6C3F1 hybrid strain. Thus, we constructed strain-specific distributions using the data for the same pathways (Fig. 4b). We find that B6C3F1 strain-derived data fall into the upper tertile of the overall population variation distribution, above the apical data-derived PODs. Furthermore, the analysis of strain-specific effects of TCE on mouse liver transcriptome that is afforded by the CC population shows that certain strains are more sensitive (e.g., CC004/TauUnc) or resistant (e.g., CC039/Unc, CC023/GeniUnc, and CC018/Unc) and may be selected for further studies in sub-chronic and chronic exposure scenarios. Interestingly, although the most sensitive strain CC004/TauUnc had transcriptional PODs that were 10-fold lower than those of B6C3F1 mice, the median transcriptional POD for this strain was within twofold of the most sensitive apical POD.
Genetic mapping of the transcriptional effects of TCE in mouse liver
To further elucidate whether the CC model provides sufficient resolution to dissect the genetic underpinnings of TCE susceptibility, we conducted genome-wide linkage mapping to identify loci associated with variation in liver TCA levels for the highest TCE dose group. As reported previously (Venkatratnam et al. 2017), we identified a significant QTL on distal chromosome 2 (Fig. 5a) associated with variation in liver TCA levels, although the robust mapping methods used here differ from the previous report. The previous study also reported that expression of PPARα-response gene fat storage inducing transmembrane protein-2 (Fitm2) which resides in this locus (red arrowhead) was positively correlated with liver TCA levels for the highest TCE dose exposure group.
a A genome-wide linkage scan for liver TCA levels at the highest TCE dose (800 mg/kg) in 50 CC lines identifies a significant QTL on chromosome 2. Location of a candidate gene Fitm2 is marked with a red arrowhead. b Genome-wide linkage scan of Fitm2 gene expression shows a cis-eQTL localizing in the same region as for (a). c Scatter plot representing normalized liver TCA levels vs. normalized Fitm2 expression, with dots representing CC founder alleles in the peak region. d Venn diagrams displaying unique local eQTLs by administered TCE dose. (Color figure online)
In the current study, the availability of whole-transcriptomic expression data enabled a more comprehensive examination of this locus for the potential association between genetic polymorphisms and gene expression. A linkage scan for expression of Fitm2 co-localizes with the TCA peak region (Fig. 5b) and indicates a strong eQTL. Among the 128 genes in a 1.5-LOD support interval for this locus from 157.7 to 180.1 Mb, Fitm2 (r = 0.475, p = 8.4 × 10−8) had the highest correlation with TCA levels (Fig. 5c). Further, effects of CC founder alleles in this region revealed that presence of M.m.castaneous alleles in this region was associated with higher expression of Fitm2 (Fig. 5c). However, TCA levels and Fitm2 expression were substantially correlated within sets of CC strains sharing regional diplotypes, indicating possible additional sources of positive correlation and signaling potential additional complexities with confidently pointing to the genetic underpinnings of inter-individual variation in liver TCA levels.
To conduct a more comprehensive analysis of the genetic underpinnings of variation in response to TCE, we additionally performed an analysis using, for each gene and CC strain, the slope of the expression response (β1 in a dose–response linear model) to TCE as a trait for linkage mapping. To be comprehensive, we performed the analysis using β1 values from both simple linear regression of expression vs. ln(dose), as well as β1 values from the DESeq2 analyses. Separate cis and trans p values and corresponding false discovery q values were obtained as described in “Materials and methods”. In terms of q values (which are corrected for multiple comparisons), none of the results were significant.
Finally, to identify genetic loci driving variation in transcriptomic responses, we performed expression quantitative trait locus (eQTL) analysis at each dose group. We observed suggestive trans-bands (or instances where expression of several genes is driven by a common locus) in chromosomes 3, 11, and 12 in the vehicle treatment group. Here “suggestive” indicates that nominally significant (trans p < 0.05) findings were observed, before multiple testing corrections. No trans-bands were observed at lower dose groups, but few trans-bands were observed at higher dose groups (data not shown). Figure 5d reports the numbers of significant local (cis) eQTLs at each dose group in a Venn diagram, illustrating variation by dose, and also considerable commonality, with 2285 genes significant at q < 0.05 for all doses.
Discussion
It is widely acknowledged in the fields of toxicology and risk assessment that population variation is one of the key challenges that begets uncertainty in human health assessments of environmental chemicals (Zeise et al. 2013). Drug safety evaluation is usually more informed through studies in humans at various phases of the clinical trials, still the challenge of idiosyncratic adverse drug reactions is also prominent and subject to active investigations (Atienzar et al. 2016). Solutions to these challenges are currently few, despite the abundance of experimental models from cells, to animals, to human studies. For instance, the tools for studies of genetics in experimental model systems have been originally developed by geneticists (Churchill et al. 2012; The International HapMap Consortium 2003; Threadgill et al. 2011) and only recently have these models been used in studies of acute and repeat-dose exposure to drugs and chemicals (French et al. 2015; Harrill and McAllister 2017). The potential for how these new animal models can inform risk assessment is great, though example applications of incorporating these data into decision-making remain small in number (Chiu and Rusyn 2018). For instance, evaluation of toxicity using population-based in vitro and in vivo models can potentially reduce both false positive and false negative signals and improve hazard identification. Enhanced ability to perform genetic mapping allows for the identification of key biological pathways and mechanisms that may be involved in toxicity and/or susceptibility. In addition, population-based experimental data can serve as a surrogate for human variation, and thus be used to quantitatively estimate the degree of human toxicokinetic/toxicodynamic variation and thereby increase confidence in the dose–response step of risk assessment that sets health-protective exposure limits.
The difficulty in translating the data from studies in population-wide experimental models to real decisions is due not only to the complexities of the relationships between genotypes and phenotypes, but also because of impediments resulting from “cultural” differences between the research questions in genetics, decades-old “standard practices” in toxicology studies, and the needs of decision-makers. Specifically, there appears to be a chasm in what constitutes the most valuable outcome(s) of a toxicology study in a population model. Is it a susceptibly locus, the molecular determinant(s) of inter-individual variation that may be used as a biomarker, a quantitative estimate of the extent of inter-individual variation, a better “model” (i.e., strain or cell line) for susceptible humans, or all of the above?
This study takes the “all of the above” point of view. It adds to the body of knowledge on the utility of the mouse population-based experimental models in toxicology and risk assessment by examining transcriptomic data obtained from a study in CC mice for characterization of strain-dependent and strain-independent mechanisms of TCE toxicity, discovery of the potential susceptibility loci, as well as dose–response assessment and derivation of POD values. This study also provides further evidence of the relative impact of dose and strain variation on transcription, and is among the largest studies to date that have combined large populations, transcriptomics, and toxicity phenotyping.
We found that all known pathways of liver toxicity of TCE (Cichocki et al. 2016; Rusyn et al. 2014) are perturbed in both strain- and dose-dependent manner. Even though strain effects were predominant in terms of liver transcriptome among CC strains, TCE effects were prominent and largely dependent on the formation of TCA. However, despite high concordance in dose-, strain-, and interaction effects between TCE dose and liver TCA levels, inter-individual variation likely depends on factors other than metabolism to TCA. This finding is highly informative with respect to not only the interpretation of the strain differences in the mouse, but also more generally to the extrapolation of experimental animal data to humans. Specifically, the diversity of the pathways involved and the complexity of the signaling mechanisms that were largely strain-dependent caution against assuming that studies in knockout and/or transgenic animals are any more informative, or human-like, models than traditional rodent models. This study also supports the utility of the information on the molecular pathways, rather than individual genes, for cross-species translation and biomarker discovery, similar to the conclusions of the study of tolvaptan-induced liver injury in CC population (Mosedale et al. 2017).
Additionally, studies in genetically defined population-wide models enable discovery of the susceptibility loci through genetic mapping. There are a number of published examples when susceptibility loci and candidate genes were successfully identified for drug and chemical-associated toxicity phenotypes (French et al. 2015; Harrill et al. 2009b; Mosedale et al. 2017; Venkatratnam et al. 2017). Despite some success, these studies have pointed out that chemical-induced toxicities are highly complex traits and thus are polygenic in nature. Our study confirms this sentiment by also exploring the gene expression dimension. We found that TCE-mediated transcriptional responses in mouse liver may be highly polygenic in nature, so that mapping multiple susceptibility loci may be difficult with the sample size of 50 CC strains. One possible solution is to increase the number of strains (Kaeppler 1997), or replicates per strain in future studies; however, the cost and complexity of these studies is likely prohibitive and not proportionate to the value of information that may be obtained. While the knowledge of the exact susceptibility genes/loci may be of use for drugs in the context of “precision medicine,” even if such are discovered they are likely to be less informative in the context of human health assessment of TCE and other chemicals for which genetic testing prior to exposure is highly improbable.
An outcome of this study that is most likely to be of use for human health risk assessment is the exploration of dose–response relationships in response to TCE at the transcriptomic and population levels. The paper by French and co-workers (French et al. 2015) was the first to demonstrate the value of mouse population studies for quantitative dose–response modeling that is directly applicable for risk assessment. Similarly, we have showed previously for TCE that population-based estimates of toxicokinetic parameters from a study in mice are concordant to those for data from humans (Chiu et al. 2014). Hence, our study explored the quantitative aspects of molecular sequelae of exposure to TCE in a mouse population and used gene expression to derive PODs for various pathways and strains. Thomas and co-workers have demonstrated that pathway-based PODs based on gene expression data from short-term exposure studies are well correlated with the POD on the apical endpoints derived from traditional 90-day and 2-year animal studies (Farmahin et al. 2017; Thomas et al. 2011, 2013b).
We recently reported that in B6C3F1 mice, transcriptional PODs for TCE correlated well with PODs for apical endpoints, after correcting for toxicokinetics (Zhou et al. 2017). Here, we found a similar correspondence to apical endpoint PODs using transcriptomic data from a genetically diverse mouse population. Previously, it was found that the transcriptional PODs were more conservative, generally within one order of magnitude (Thomas et al. 2011). In our earlier study in B6C3F1 mice, transcriptional PODs for TCE were also within 10-fold of apical PODs, but the differences were in both directions, i.e., not consistently conservative. Here, in CC mice, the transcriptional and apical endpoint PODs for TCE substantially overlapped, a large number of transcriptional pathways, including the most sensitive falling within the range of the apical endpoint PODs. This greater apparent correlation suggests that using CC mice may provide a more robust transcriptional POD because of the incorporation of genetic diversity that reduces the potential impact of outliers, but this hypothesis needs to be tested for additional chemicals and target tissues. Moreover, as was the case with Zhou et al. (2017), conversion to human equivalent doses has the additional utility of being directly comparable to human exposure estimates and derivation of the margins of exposure. These results provide further evidence that transcriptomic data can be used as surrogates for in vivo PODs, and suggest that a population-based approach might be more robust than using a single strain.
In sum, our study is among the first to explore the linkages between gene expression and genetic polymorphisms in a toxicological context. This innovative approach extends the common method to analyzing toxicity pathway perturbations to the population level, allowing for an exploration of gene–environment interactions, which are thought to be the basis of phenotypic variation across the population. Using the CC population and TCE liver effects as a prototypical example, we have demonstrated that adding the dimension of genetic diversity has multiple potential benefits. First, by identifying pathways that are dependent on strain, treatment, or their interaction, we obtain deeper insights into toxicological mechanisms. Second, it enables the possibility of genetic mapping to identify susceptibility loci, although this may be challenging for polygenic traits such as TCE-induced liver effects. Finally, at least in this case example, conducting gene expression dose–response analysis across a population appears to be more robust than using a single strain in terms of the correlation between transcriptional and apical PODs. Overall, our study demonstrates the utility of mouse population-based studies in addressing the key issue of inter-individual variation in the human health risk assessment of chemical exposures.
References
Abdo N, Xia M, Brown CC, Kosyk O, Huang R, Sakamuru S, Zhou YH, Jack JR, Gallins P, Xia K, Li Y, Chiu WA, Motsinger-Reif AA, Austin CP, Tice RR, Rusyn I, Wright FA (2015) Population-based in vitro hazard and concentration-response assessment of chemicals: the 1000 genomes high-throughput screening study. Environ Health Perspect 123:458–466
Anders S, Pyl PT, Huber W (2015) HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31:166–169
Andersen ME, Clewell HJ 3rd, Bermudez E, Willson GA, Thomas RS (2008) Genomic signatures and dose-dependent transitions in nasal epithelial responses to inhaled formaldehyde in the rat. Toxicol Sci 105:368–383
Atienzar FA, Blomme EA, Chen M, Hewitt P, Kenna JG, Labbe G, Moulin F, Pognan F, Roth AB, Suter-Dick L, Ukairo O, Weaver RJ, Will Y, Dambach DM (2016) Key challenges and opportunities associated with the use of in vitro models to detect human DILI: integrated risk assessment and mitigation plans. Biomed Res Int 2016:9737920
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120
Bradford BU, Lock EF, Kosyk O, Kim S, Uehara T, Harbourt D, DeSimone M, Threadgill DW, Tryndyak V, Pogribny IP, Bleyle L, Koop DR, Rusyn I (2011) Interstrain differences in the liver effects of trichloroethylene in a multistrain panel of inbred mice. Toxicol Sci 120:206–217
Bronley-DeLancey A, McMillan DC, McMillan JM, Jollow DJ, Mohr LC, Hoel DG (2006) Application of cryopreserved human hepatocytes in trichloroethylene risk assessment: relative disposition of chloral hydrate to trichloroacetate and trichloroethanol. Environ Health Perspect 114:1237–1242
Burczynski ME, McMillian M, Ciervo J, Li L, Parker JB, Dunn RT, Hicken S, Farr S, Johnson MD (2000) Toxicogenomics-based discrimination of toxic mechanism in HepG2 human hepatoma cells. Toxicol Sci 58:399–415
Bystrykh L, Weersing E, Dontje B, Sutton S, Pletcher MT, Wiltshire T, Su AI, Vellenga E, Wang J, Manly KF, Lu L, Chesler EJ, Alberts R, Jansen RC, Williams RW, Cooke MP, de Haan G (2005) Uncovering regulatory pathways that affect hematopoietic stem cell function using ‘genetical genomics’. Nat Genet 37:225–232
Chesler EJ, Wang J, Lu L, Qu Y, Manly KF, Williams RW (2003) Genetic correlates of gene expression in recombinant inbred strains: a relational model system to explore neurobehavioral phenotypes. Neuroinformatics 1:343–357
Chiu WA, Ginsberg GL (2011) Development and evaluation of a harmonized physiologically based pharmacokinetic (PBPK) model for perchloroethylene toxicokinetics in mice, rats, and humans. Toxicol Appl Pharmacol 253:203–234
Chiu WA, Rusyn I (2018) Advancing chemical risk assessment decision-making with population variability data: challenges and opportunities. Mamm Genome. https://doi.org/10.1007/s00335-017-9731-6
Chiu WA, Micallef S, Monster AC, Bois FY (2007) Toxicokinetics of inhaled trichloroethylene and tetrachloroethylene in humans at 1 ppm: empirical results and comparisons with previous studies. Toxicol Sci 95:23–36
Chiu WA, Okino MS, Evans MV (2009) Characterizing uncertainty and population variability in the toxicokinetics of trichloroethylene and metabolites in mice, rats, and humans using an updated database, physiologically based pharmacokinetic (PBPK) model, and Bayesian approach. Toxicol Appl Pharmacol 241:36–60
Chiu WA, Campbell JL, Clewell HJ, Zhou YH, Wright FA, Guyton KZ, Rusyn I (2014) Physiologically-based pharmacokinetic (PBPK) modeling of inter-strain variability in trichloroethylene metabolism in the mouse. Environ Health Perspect 122:456–463
Church RJ, Gatti DM, Urban TJ, Long N, Yang X, Shi Q, Eaddy JS, Mosedale M, Ballard S, Churchill GA, Navarro V, Watkins PB, Threadgill DW, Harrill AH (2015) Sensitivity to hepatotoxicity due to epigallocatechin gallate is affected by genetic background in diversity outbred mice. Food Chem Toxicol 76:19–26
Churchill GA, Gatti DM, Munger SC, Svenson KL (2012) The diversity outbred mouse population. Mamm Genome 23:713–718
Cichocki JA, Guyton KZ, Guha N, Chiu WA, Rusyn I, Lash LH (2016) Target organ metabolism, toxicity, and mechanisms of trichloroethylene and perchloroethylene: key similarities, differences, and data gaps. J Pharmacol Exp Ther 359:110–123
Cichocki JA, Furuya S, Venkatratnam A, McDonald TJ, Knap AH, Wade T, Sweet S, Chiu WA, Threadgill DW, Rusyn I (2017) Characterization of variability in toxicokinetics and toxicodynamics of tetrachloroethylene using the collaborative cross mouse population. Environ Health Perspect 125:057006
Crowley JJ, Zhabotynsky V, Sun W, Huang S, Pakatci IK, Kim Y, Wang JR, Morgan AP, Calaway JD, Aylor DL, Yun Z, Bell TA, Buus RJ, Calaway ME, Didion JP, Gooch TJ, Hansen SD, Robinson NN, Shaw GD, Spence JS, Quackenbush CR, Barrick CJ, Nonneman RJ, Kim K, Xenakis J, Xie Y, Valdar W, Lenarcic AB, Wang W, Welsh CE, Fu CP, Zhang Z, Holt J, Guo Z, Threadgill DW, Tarantino LM, Miller DR, Zou F, McMillan L, Sullivan PF, Pardo-Manuel de Villena F (2015) Analyses of allele-specific gene expression in highly divergent mouse crosses identifies pervasive allelic imbalance. Nat Genet 47:353–360
Dennis G Jr, Sherman BT, Hosack DA, Yang J, Gao W, Lane HC, Lempicki RA (2003) DAVID: database for annotation, visualization, and integrated discovery. Genome Biol 4:P3
Domino MM, Pepich BV, Munch DJ, Fair PS, Xie Y (2003) Methd 552.3: determination of haloacetic acids and dalapon in drinking water by liquid-liquid microextraction, derivatization, and gas chromatography with electron capture detection. Office of Ground Water and Drinking Water, Cincinnati
Eduati F, Mangravite LM, Wang T, Tang H, Bare JC, Huang R, Norman T, Kellen M, Menden MP, Yang J, Zhan X, Zhong R, Xiao G, Xia M, Abdo N, Kosyk O, Collaboration N-N-UDT., Friend S, Dearry A, Simeonov A, Tice RR, Rusyn I, Wright FA, Stolovitzky G, Xie Y, Saez-Rodriguez J (2015) Prediction of human population responses to toxic compounds by a collaborative competition. Nat Biotechnol 33:933–940
Evans MV, Chiu WA, Okino MS, Caldwell JC (2009) Development of an updated PBPK model for trichloroethylene and metabolites in mice, and its application to discern the role of oxidative metabolism in TCE-induced hepatomegaly. Toxicol Appl Pharmacol 236:329–340
Farmahin R, Williams A, Kuo B, Chepelev NL, Thomas RS, Barton-Maclaren TS, Curran IH, Nong A, Wade MG, Yauk CL (2017) Recommended approaches in the application of toxicogenomics to derive points of departure for chemical risk assessment. Arch Toxicol 91:2045–2065
Fostel JM (2008) Towards standards for data exchange and integration and their impact on a public database such as CEBS (Chemical Effects in Biological Systems). Toxicol Appl Pharmacol 233:54–62
French JE, Gatti DM, Morgan DL, Kissling GE, Shockley KR, Knudsen GA, Shepard KG, Price HC, King D, Witt KL, Pedersen LC, Munger SC, Svenson KL, Churchill GA (2015) Diversity outbred mice identify population-based exposure thresholds and genetic factors that influence benzene-induced genotoxicity. Environ Health Perspect 123:237–245
Ganter B, Tugendreich S, Pearson CI, Ayanoglu E, Baumhueter S, Bostian KA, Brady L, Browne LJ, Calvin JT, Day GJ, Breckenridge N, Dunlea S, Eynon BP, Furness LM, Ferng J, Fielden MR, Fujimoto SY, Gong L, Hu C, Idury R, Judo MS, Kolaja KL, Lee MD, McSorley C, Minor JM, Nair RV, Natsoulis G, Nguyen P, Nicholson SM, Pham H, Roter AH, Sun D, Tan S, Thode S, Tolley AM, Vladimirova A, Yang J, Zhou Z, Jarnagin K (2005) Development of a large-scale chemogenomics database to improve drug candidate selection and to understand mechanisms of chemical toxicity and action. J Biotechnol 119:219–244
Gatti D, Maki A, Chesler EJ, Kirova R, Kosyk O, Lu L, Manly KF, Williams RW, Perkins A, Langston MA, Threadgill DW, Rusyn I (2007) Genome-level analysis of genetic regulation of liver gene expression networks. Hepatology 46:548–557
Gatti DM, Harrill AH, Wright FA, Threadgill DW, Rusyn I (2009) Replication and narrowing of gene expression quantitative trait loci using inbred mice. Mamm Genome 20:437–446
GTEx Consortium (2013) The genotype-tissue expression (GTEx) project. Nat Genet 45:580–585
GTEx Consortium (2015) Human genomics. The genotype-tissue expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348:648–660
Harrill AH, McAllister KA (2017) New rodent population models may inform human health risk assessment and identification of genetic susceptibility to environmental exposures. Environ Health Perspect 125:086002
Harrill AH, Rusyn I (2008) Systems biology and functional genomics approaches for the identification of cellular responses to drug toxicity. Expert Opin Drug Metab Toxicol 4:1379–1389
Harrill AH, Ross PK, Gatti DM, Threadgill DW, Rusyn I (2009a) Population-based discovery of toxicogenomics biomarkers for hepatotoxicity using a laboratory strain diversity panel. Toxicol Sci 110:235–243
Harrill AH, Watkins PB, Su S, Ross PK, Harbourt DE, Stylianou IM, Boorman GA, Russo MW, Sackler RS, Harris SC, Smith PC, Tennant R, Bogue M, Paigen K, Harris C, Contractor T, Wiltshire T, Rusyn I, Threadgill DW (2009b) Mouse population-guided resequencing reveals that variants in CD44 contribute to acetaminophen-induced liver injury in humans. Genome Res 19:1507–1515
Kaeppler SM (1997) Quantitative trait locus mapping using sets of near-isogenic lines: relative power comparisons and technical considerations. Theor Appl Genet 95:384–392
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357–360
Klaunig JE, Ruch RJ, Lin EL (1989) Effects of trichloroethylene and its metabolites on rodent hepatocyte intercellular communication. Toxicol Appl Pharmacol 99:454–465
Lash LH, Chiu WA, Guyton KZ, Rusyn I (2014) Trichloroethylene biotransformation and its role in mutagenicity, carcinogenicity and target organ toxicity. Mutat Res Rev Mutat Res 762:22–36
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550
Luo YS, Furuya S, Chiu W, Rusyn I (2018) Characterization of inter-tissue and inter-strain variability of TCE glutathione conjugation metabolites DCVG, DCVC, and NAcDCVC in the mouse. J Toxicol Environ Health A 81:37–52
Maloney EK, Waxman DJ (1999) Trans-activation of PPARalpha and PPARgamma by structurally diverse environmental chemicals. Toxicol Appl Pharmacol 161:209–218
Mosedale M, Kim Y, Brock WJ, Roth SE, Wiltshire T, Eaddy JS, Keele GR, Corty RW, Xie Y, Valdar W, Watkins PB (2017) Candidate risk factors and mechanisms for tolvaptan-induced liver injury are identified using a collaborative cross approach. Toxicol Sci 156:438–454
Rakhshandehroo M, Knoch B, Muller M, Kersten S (2010) Peroxisome proliferator-activated receptor alpha target genes. PPAR Res 2010:612089
Rusyn I, Gatti DM, Wiltshire T, Kleeberger SR, Threadgill DW (2010) Toxicogenetics: population-based testing of drug and chemical safety in mouse models. Pharmacogenomics 11:1127–1136
Rusyn I, Chiu WA, Lash LH, Kromhout H, Hansen J, Guyton KZ (2014) Trichloroethylene: mechanistic, epidemiologic and other supporting evidence of carcinogenic hazard. Pharmacol Ther 141:55–68
Sauer UG, Deferme L, Gribaldo L, Hackermuller J, Tralau T, van Ravenzwaay B, Yauk C, Poole A, Tong W, Gant TW (2017) The challenge of the application of ‘omics technologies in chemicals risk assessment: background and outlook. Regul Toxicol Pharmacol 91(Suppl 1):S14–S26
Schadt EE, Monks SA, Drake TA, Lusis AJ, Che N, Colinayo V, Ruff TG, Milligan SB, Lamb JR, Cavet G, Linsley PS, Mao M, Stoughton RB, Friend SH (2003) Genetics of gene expression surveyed in maize, mouse and man. Nature 422:297–302
Schadt EE, Molony C, Chudin E, Hao K, Yang X, Lum PY, Kasarskis A, Zhang B, Wang S, Suver C, Zhu J, Millstein J, Sieberts S, Lamb J, GuhaThakurta D, Derry J, Storey JD, Avila-Campillo I, Kruger MJ, Johnson JM, Rohl CA, van Nas A, Mehrabian M, Drake TA, Lusis AJ, Smith RC, Guengerich FP, Strom SC, Schuetz E, Rushmore TH, Ulrich R (2008) Mapping the genetic architecture of gene expression in human liver. PLoS Biol 6:e107
The International HapMap Consortium (2003) The international HapMap project. Nature 426:789–796
Thomas RS, Allen BC, Nong A, Yang L, Bermudez E, Clewell HJ III, Andersen ME (2007) A method to integrate benchmark dose estimates with genomic data to assess the functional effects of chemical exposure. Toxicol Sci 98:240–248
Thomas RS, Clewell HJ, Allen BC, Wesselkamper SC, Wang NC, Lambert JC, Hess-Wilson JK, Zhao QJ, Andersen ME (2011) Application of transcriptional benchmark dose values in quantitative cancer and noncancer risk assessment. Toxicol Sci 120:194–205
Thomas RS, Philbert MA, Auerbach SS, Wetmore BA, Devito MJ, Cote I, Rowlands JC, Whelan MP, Hays SM, Andersen ME, Meek ME, Reiter LW, Lambert JC, Clewell HJ 3rd, Stephens ML, Zhao QJ, Wesselkamper SC, Flowers L, Carney EW, Pastoor TP, Petersen DD, Yauk CL, Nong A (2013a) Incorporating new technologies into toxicity testing and risk assessment: moving from 21st century vision to a data-driven framework. Toxicol Sci 136:4–18
Thomas RS, Wesselkamper SC, Wang NC, Zhao QJ, Petersen DD, Lambert JC, Cote I, Yang L, Healy E, Black MB, Clewell HJ 3rd, Allen BC, Andersen ME (2013b) Temporal concordance between apical and transcriptional points of departure for chemical risk assessment. Toxicol Sci 134:180–194
Threadgill DW, Miller DR, Churchill GA, de Villena FPM (2011) The collaborative cross: a recombinant inbred mouse population for the systems genetic era. ILAR J 52:24–31
U.S. EPA (2011a) Toxicological review of tetrachloroethylene (CAS No. 127-18-4): in support of summary information on the integrated risk information system (IRIS). U.S. Environmental Protection Agency, Washington, DC
U.S. EPA (2011b) Toxicological review of trichloroethylene (CAS No. 79-01-6): in support of summary information on the integrated risk information system (IRIS). U.S. Environmental Protection Agency, Washington, DC
Uehara T, Ono A, Maruyama T, Kato I, Yamada H, Ohno Y, Urushidani T (2010) The Japanese toxicogenomics project: application of toxicogenomics. Mol Nutr Food Res 54:218–227
Venkatratnam A, Furuya S, Kosyk O, Gold A, Bodnar W, Konganti K, Threadgill DW, Gillespie KM, Aylor DL, Wright FA, Chiu WA, Rusyn I (2017) Collaborative cross mouse population enables refinements to characterization of the variability in toxicokinetics of trichloroethylene and provides genetic evidence for the role of PPAR pathway in its oxidative metabolism. Toxicol Sci 158:48–62
Xu J, Thakkar S, Gong B, Tong W (2016) The FDA’s experience with emerging genomics technologies-past, present, and future. AAPS J 18:814–818
Yang L, Allen BC, Thomas RS (2007) BMDExpress: a software tool for the benchmark dose analyses of genomic data. BMC Genom 8:387
Zeise L, Bois FY, Chiu WA, Hattis D, Rusyn I, Guyton KZ (2013) Addressing human variability in next-generation human health risk assessments of environmental chemicals. Environ Health Perspect 121:23–31
Zhou YH, Cichocki JA, Soldatow VY, Scholl EH, Gallins PJ, Jima D, Yoo HS, Chiu WA, Wright FA, Rusyn I (2017) Comparative dose-response analysis of liver and kidney transcriptomic effects of trichloroethylene and tetrachloroethylene in B6C3F1 mouse. Toxicol Sci 160:95–110
Acknowledgements
This work was supported, in part, by a cooperative agreement (STAR RD83561202) from U.S. EPA to Texas A&M University, a Barry Goldwater scholarship, and grants from the National Institutes of Health (U01 ES026717, R00 ES021535, P42 ES027704, and P30 ES025128). The views expressed in this article are those of the authors and do not necessarily reflect the views of NIH or policies of EPA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there are no conflicts of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Venkatratnam, A., House, J.S., Konganti, K. et al. Population-based dose–response analysis of liver transcriptional response to trichloroethylene in mouse. Mamm Genome 29, 168–181 (2018). https://doi.org/10.1007/s00335-018-9734-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00335-018-9734-y