Abstract
In rats, direct exposure to TCDD causes myriad toxicities. Exposed rats experience hepatotoxicity, wasting syndrome and immune suppression, amongst others. “Inherited exposure”, as occurs in the F3 generation of directly exposed F0 animals, has also been shown to cause toxicity: both male and female F3 rats demonstrate an increased incidence of adult onset disease, females also display reproductive abnormalities and increased incidence of ovarian diseases while males show increased incidence of kidney disease and an altered sperm epigenome. Here, we explore the hepatic transcriptomic profile of male and female F3 Sprague–Dawley rats bred through the paternal germ line from F0 dams exposed to a single dose of TCDD (0, 30, 100, 300 or 1000 ng/kg body weight) by oral gavage. We hypothesize that RNA transcripts with altered abundance in livers of unexposed F3 progeny of treated F0 Sprague–Dawley rats may result from epigenetic modifications to the genome. We further survey patterns of differential methylation within male F3 rat testis. Female F3 rats demonstrated more TCDD-mediated hepatic transcriptomic changes than males, with differences primarily in the lowest dose group. In testis from male F3 rats, multiple olfactory receptors displayed patterns of differential methylation. Hypermethylation of Egfr and Mc5r among testes from TCDD lineage rats was observed, but without corresponding changes in hepatic mRNA abundance. Further studies examining these differences in other tissue types are warranted.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
2,3,7,8-Tetrachlorodibenzo-p-dioxin (TCDD) is a persistent, highly lipophilic, environmental contaminant produced in herbicide and pesticide manufacture, as well as a by-product of various industrial products (Schecter et al. 2006; Shen et al. 2009; Von Burg 1988). In rats, TCDD has a half-life of ~ 19 days (Geyer et al. 2002) and exposure to even small amounts (less than 50 µg/kg in sensitive strains) can cause a myriad of toxic outcomes including hepatotoxicity, reproductive and developmental toxicity, thymic atrophy, wasting syndrome, immune suppression and acute lethality (Pohjanvirta and Tuomisto 1994). In humans, the half-life of TCDD is considerably longer, approximately 8 years (Geyer et al. 2002). Direct exposure to TCDD in humans can cause chloracne (Baccarelli et al. 2005), tumourigenesis, possibility including prostate cancer in men (Ansbaugh et al. 2013; Leng et al. 2014), and has been implicated in development of chronic diseases (Yi et al. 2014). Much of what is known about TCDD exposure in humans comes primarily from two sources: exposure from Agent Orange used in the Vietnam War and an industrial accident in Seveso, Italy in 1979. In both cases, a wide range of toxic outcomes have been documented in those individuals exposed directly as well as subsequent generations (Baccarelli et al. 2008; Mocarelli et al. 2011).
Due to its long half-life, effects of TCDD that are observed up to and including the F2 generation can be attributed to direct, multigenerational exposure; TCDD exposure of females (F0) can persist in the body leading to direct exposure of any offspring (F1), through the placenta and breast milk (Nau et al. 1986). This further exposes the developing germ cells (F2) within these offspring. In zebrafish, transgenerational effects, such as skeletal and reproductive abnormalities, have been documented up to the F2 generation (equivalent to rodent F3 generation) (Baker et al. 2014). In rats, repeated exposure to TCDD of F0 animals has been shown to alter the sex ratio of the F2 generation (Ikeda et al. 2005). Various TCDD-mediated effects have been documented in the F3 generation that have been linked to heritable changes of the epigenome of exposed individuals. In particular, Manikkam et al. described an increased prevalence of kidney disease (Manikkam et al. 2012b) in male F3 rats following exposure of the F0 generation to TCDD, as well as reduced serum testosterone levels (Manikkam et al. 2012a). In F3 females increased prevalence of ovarian disease, decline in ovarian follicle numbers and early onset of puberty have all been reported previously in relation to ancestral TCDD exposure (Manikkam et al. 2012a, b; Nilsson et al. 2012; Yu et al. 2019).
In rodents, the liver is a primary site of TCDD-mediated toxicities (Pohjanvirta et al. 1990). Numerous studies have identified substantial transcriptomic (Boutros et al. 2011; Boverhof et al. 2006; Fletcher et al. 2005; Franc et al. 2008; Moffat et al. 2010; Nault et al. 2013; Yao et al. 2012) and proteomic (Forgacs et al. 2012; Lee et al. 2005; Pastorelli et al. 2006; Prokopec et al. 2017) hepatic effects following direct exposure to TCDD in various strains of rats, including the TCDD-sensitive Long–Evans (L–E; oral LD50 of 9.8–17.7 µg/kg TCDD) and TCDD-resistant Han/Wistar (H/W; oral LD50 of > 9600 µg/kg TCDD) strains (Pohjanvirta and Tuomisto 1994). Sprague–Dawley (SD) rats have an oral LD50 of 20 µg/kg TCDD (Bickel 1982) and are considered to be sensitive to the effects of TCDD. Here we examine the hepatic transcriptome of male and female SD rats (F3) from TCDD-exposed lineages to identify genes showing altered mRNA abundance profiles. Furthermore, the promoter-region methylation patterns of genes demonstrating altered RNA abundance are assessed to identify candidate mechanisms for these transgenerational effects. Finally, as transgenerational epigenetic effects have been previously detected in sperm (Manikkam et al. 2012a), and an enhanced testicular inflammation phenotype has been described in F3 rats (Bruner-Tran et al. 2014), we additionally contrast genomic methylation patterns within testes of F3 rats from TCDD-exposed and control lineages.
Results
Experimental design
We evaluated the transgenerational hepatic transcriptomic or testicular epigenetic effects caused by TCDD exposure of pregnant Sprague–Dawley (SD) rats through the paternal germline. Twenty-five pregnant SD rats were treated with a single dose of TCDD (0, 30, 100, 300, 1000 ng/kg). Each dam initiated either a control (control lineage) or TCDD-exposed lineage (TCDD lineage), terminating with the F3 generation (Fig. 1). Hepatic tissue from male and female F3 animals from each treatment group (n = 5–8 each male and female; Table 1) was collected and transcriptomic profiling performed. A total of 67 F3 animals were initially included in this study; complete lineage information is available in Supplementary Table 1. A single array was identified as an outlier and was removed from downstream analyses (Supplementary Figures 1–2), reducing the total number of animals examined to 66. Concurrently, testicular tissues were collected from male F3 rats from the 1000 ng/kg TCDD lineage (n = 4) and control lineage (n = 4) groups and targeted bisulfite sequencing performed to identify differentially methylated regions (DMRs).
Experimental design. Pregnant Sprague–Dawley (SD) rats were treated with a single dose of TCDD (0, 30, 100, 300, 1000 ng/kg) dissolved in corn oil on gestational day 11 by oral gavage. Adult F1 males were mated with untreated female rats to produce the F2 generation. Similarly, adult F2 males were mated with untreated females to produce the F3 generation. Tissues were collected from F3 animals from each lineage (liver from male and female animals; testis from males). Hepatic transcriptome was evaluated using Affymetrix Rat Gene 2.0 ST arrays and testicular tissues used for bisulfite sequencing
TCDD has minimal impact on the F3 hepatic transcriptome
The hepatic transcriptome demonstrated minimal differences between the TCDD- and control lineage, regardless of dose or sex (Supplementary Figure 3). After filtering, we identified 280 genes that demonstrated moderate differences, with a variance in normalized intensity values > 0.5 across the cohort (Supplementary Figure 3b). To evaluate these clustering patterns, we employed the Adjusted Rand Index (ARI). Values of the ARI approaching one indicates that the clustering by RNA abundance data exactly matches the variable of interest, while ARI values near zero suggest the variable has no influence on RNA abundance. From this analysis, the greatest influence was attributed to sex (ARIsex = 1), rather than treatment (ARItreatment = − 0.008) or specific dose (ARIdose = 0.027). Within each sex, however, moderate associations were observed with treatment (ARIsex:treatment = 0.567; green vs. white in the first covariate) but not by specific dose (ARIsex:dose = 0.188; all levels within this first covariate). This highlights the presence of TCDD-mediated effects on RNA abundance that are not dependent on any specific dose and may in fact differ between the males and females. To confirm this, linear modelling was performed to identify specific changes in each treatment group relative to control animals of the same sex. Consistently, minimal changes were detected in any group, with 23 and one gene respectively altered in livers of female F3 rats descending from dams exposed to the lowest (30 ng/kg) and highest (1000 ng/kg) dose of TCDD (FDR < 0.1; Supplementary Figure 3c, Supplementary Table 2). Interestingly, this did not include any typical “AHR-core” response genes (Supplementary Figure 3d). While not statistically significant, we did observe a small increase in RNA abundance for Ahr in female rat liver, particularly at the lower doses of TCDD (foldchanges = 0.3 and 0.05 relative to controls, for 30 and 1000 ng/kg dose groups respectively), while the opposite was observed in males (small decrease in RNA abundance for Ahr, particularly at the higher doses; foldchanges = 0.08 and − 0.25 for 30 and 1000 ng/kg dose groups respectively, relative to controls). Female rats have been shown to be more sensitive to TCDD-induced toxicities than male rats (Pohjanvirta et al. 1993; Silkworth et al. 2008), however, the larger number of TCDD-responsive genes in females of the lowest dose group is unusual based on previous studies. This may represent hormesis, as proposed previously (Kociba et al. 1978; Tuomisto et al. 2006), albeit with specific reference to tumour promotion rather than molecular abundance, or be secondary to a female hormonal change with a non-monotonous dose–response relationship (Karman et al. 2012).
Those genes identified as having differentially abundant transcripts (q < 0.1) in any treatment group were examined further (Fig. 2). Of these, carboxypeptidase A4 (Cpa4), a secreted zinc-dependent metallocarboxypeptidase, showed the largest magnitude change—repressed nearly two-fold (log2|fold-change|= − 0.99) in female F3 rat liver. Curiously, we observed decreasing effect size with increasing TCDD dose, again consistent with hormesis. In previous studies of liver from rats directly exposed to TCDD, only one (L–E treated with 100 μg/kg for 10 days) (Boutros et al. 2011; Moffat et al. 2010) showed significantly reduced mRNA abundance of Cpa4. However, C57BL/6 mouse liver often showed TCDD-mediated effects on mRNA abundance of this gene: in female mice treated with a single dose of 500 μg/kg TCDD for 6 h or treated with 125 or 500 μg/kg TCDD for 4 days, and in male mice treated with 500 μg/kg TCDD for 6 days (Lee et al. 2015; Prokopec et al. 2015). In humans, CPA4 is a maternally imprinted gene (Kayashima et al. 2003) shown to be upregulated in prostate cancer (Huang et al. 1999); it has been suggested to regulate the extracellular environment (Tanco et al. 2010) and adipogenesis (He et al. 2016).
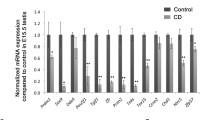
Hormesis-like response within female F3 hepatic transcriptome. Twenty-four genes demonstrated altered mRNA abundance in TCDD-exposed lineages relative to controls. Dot size indicates magnitude of change (log2|foldchange|) while dot colour indicates direction. Background shading represents significance level (FDR adjusted p values). Right heatmap indicates significant differences at FDR < 0.1 (where orange indicates an increase and blue a decrease in abundance relative to controls) in previous studies of TCDD-mediated transcriptomic changes among various species, strains, sexes and tissues exposed to various doses of TCDD for multiple timepoints (Prokopec et al. 2017); here grey indicates that the gene was not tested
Alternatively, DNA-damage inducible transcript 3 (Ddit3) showed increased mRNA abundance in female liver originating from the highest dose lineage. Transgenic mice expressing the DEL variant showed similar increases to mRNA abundance for this gene (Prokopec et al. 2017), as did livers of male C57BL/6 mice (Lee et al. 2015; Prokopec et al. 2015). In contrast, male rat liver often showed reduced expression: L–E (100 μg/kg TCDD for 3 h and 4 days) and H/W (100 μg/kg TCDD for 4 and 10 days) (Boutros et al. 2011; Moffat et al. 2010); female mouse liver also demonstrated significantly reduced mRNA abundance of Ddit3; however, this was short lived (6 h following treatment with 500 μg/kg TCDD) (Lee et al. 2015; Prokopec et al. 2015). Ddit3 has been shown to mediate insulin resistance in adipocytes, with increased expression correlating with systemic insulin resistance (Suzuki et al. 2017).
Physiological transgenerational effects have been previously reported, along with a subset of molecular characteristics (Manikkam et al. 2012a, b). These studies identified 50 significantly DMRs in sperm from F3 progeny of TCDD-exposed F0 rats. We sought to determine whether these genes demonstrated altered transcription in livers of our male F3 rats. However, there was no overlap between these studies, possibly due to differences in sample types. Finally, due to the small number of differentially abundant transcripts observed, we next repeated the above modelling, however pooling the TCDD lineages (Supplementary Figure 3e). We observed clear differences in the mRNA profile between male and female rat livers, however these effects were not associated with exposure to TCDD.
Exposure to TCDD leads to differential methylation patterns in F3 testis
TCDD-exposure of F0 dams did not have any statistically significant influence on global DNA methylation status in the F3 generation, as assessed in the livers of both sexes and in the testes of male rats (Supplementary Table 3).
We next sought to explain the testicular inflammation phenotype previously observed in F3 progeny from F0 mice exposed to TCDD (Bruner-Tran et al. 2014). We performed bisulfite sequencing to identify any DMRs between the TCDD and control lineages. A genome-wide approach was first applied to discover DMRs outside of the gene-body (Supplementary Table 4). Multiple genes contained separate regions of hyper- and hypo-methylation in TCDD lineage when compared to control lineage (for example, Atrnl1 and Ehbp1). Additionally, four separate DMRs were found adjacent (up or downstream) to olfactory receptor genes (Fig. 3a–c). While such receptors may not be expressed in testicular tissue, these results are intriguing if such patterns hold throughout other tissues—aversion to novel foodstuffs is a well-documented response following exposure to AHR agonists (Lensu et al. 2011a, b; Manikkam et al. 2012b).
Signal receptor genes are often differentially methylated. DMRs were identified adjacent to four olfactory receptor genes. DMRs were identified a upstream of Olr137, and downstream of bOlr120 and cOlr150. d Hyper-methylation within the gene-body of Egfr among the TCDD lineage rat testes. Points (and lines) represent group (colour) and coverage (intensity) for each CG site, with position indicating genomic position (x-axis) and proportion of reads demonstrating methylation (y-axis) (colour figure online)
EGFR binding has previously been shown to be reduced in a sustained manner in various tissues of laboratory animals after TCDD exposure, including testes in Sprague–Dawley rats (el-Sabeawy et al. 1998). Although this phenomenon may largely be attributable to receptor internalization (Campion et al. 2016), the pronounced hyper-methylation of Egfr in this study in the TCDD-exposed lineage (62% of reads with methylation vs. 0% in the control animals) is of interest and would warrant analyzing its mRNA expression levels (Fig. 3d). This hyper-methylation is particularly interesting as it occurs within the gene body of Egfr—a mechanism proposed to contribute to transgenerational plasticity to environmental stimuli (Dixon et al. 2018) which has also been linked to tumourigenesis through elevated oncogene expression in liver cancer (Arechederra et al. 2018).
To improve confidence in our calls, we next limited our search to targeted regions (n = 1462; median 4346 reads per region). We identified 43 statistically significant DMRs (q < 0.01) within coding regions (Supplementary Table 5). In particular, three genes demonstrated large increases in the proportion of methylated reads between groups: Mc5r had 21% more reads with methylation in the TCDD lineage compared to the control lineage; Ppp1r27 and Fam109a also had increases of 11.5% and 10.3% methylated reads. Alternatively, Hspa8 and Olr1108 exhibited 10.0 and 8.6 percentage point decreases in methylated reads in testes from the TCDD lineage in comparison to controls. Hyper-methylation of Mc5r (melanocortin 5 receptor; Fig. 4a) may lead to repressed expression of the Hspa8 receptor, a deficiency of which has been implicated in reduced pheromone signalling (Morgan et al. 2004) and obesity (Chagnon et al. 1997; Shukla et al. 2012). Two additional genes previously shown to harbour DMRs in promoter regions in sperm, Pi16 and Rasal3 (Manikkam et al. 2012b), also contained gene-body DMRs in testes (increased methylation in the TCDD-lineage). No DMRs were detected in or around Ahr or any “AHR-core” genes.
Gene-body differential (hyper) methylation. aMc5r demonstrated differential (hyper) methylation in testis tissue obtained from the TCDD-exposed lineage relative to control samples. Points (and lines) represent group (colour) and coverage (intensity) for each CG site, with position indicating genomic position (x-axis) and proportion of reads demonstrating methylation (y-axis). b Pathway analyses were performed to identify areas of enrichment for hyper- or hypo- methylation. Dot size indicates number of hyper- or hypo- methylated genes overlapping the indicated pathway while background shading indicates statistical significance (FDR). Represented domains include biological processes (BP), cellular components (CC) and molecular function (MF) (colour figure online)
Finally, as differences among methylation levels for individual genes were generally small between our treatment and control groups, we next sought to combine genes to identify effects of ancestral TCDD-exposure on broader biological pathways. We performed pathway analysis separately for all of the hyper- and hypo-methylated gene sets using a standard bioinformatic strategy (Reimand et al. 2007). This identified unique sets of enriched pathways depending on DMR direction (hyper- or hypo-methylated in the TCDD-lineage; Fig. 4b). Specifically, genes demonstrating hyper-methylation are frequently involved in biosynthetic processes (including 154 unique genes; FDR = 0.07), cell–cell adhesion molecules (n = 18 genes including many protocadherins; FDR = 0.002) and cell–cell signalling (n = 56 genes; FDR = 0.01). We also detected enrichment of voltage-gated potassium channel activity by a small subset of these hyper-methylated genes (n = 10; FDR = 0.06). As hyper-methylation can lead to reduced mRNA levels, these pathways may all have impaired functionality; this may explain some of the physiological effects observed by Manikkam et al. (2012a; b). Interestingly, many of these hyper-methylated genes produce proteins that are enriched within intracellular, membrane-bound organelles (particularly mitochondria; Fig. 4b), suggesting a long-lasting heritable impact on cellular respiration and perhaps ROS homeostasis. Concurrently, no significant pathway enrichment was observed for genes affected by hypo-methylation (Supplementary Table 6).
Discussion
The degree of hepatotoxicity observed following exposure to TCDD differs dramatically between TCDD-sensitive and TCDD-resistant strains of rat (Niittynen et al. 2007; Viluksela et al. 2000) and is further evidenced by the myriad transcriptomic changes observed within the liver between these animals (Boutros et al. 2008; Yao et al. 2012). The liver is the major site of accumulation of TCDD in rats at doses that result in induction of xenobiotic-metabolizing enzymes and this accumulation persists for a considerable duration after exposure (Pohjanvirta et al. 1990). Previous studies on physiological effects of transgenerational exposure to TCDD did not disclose any hepatic toxicities (Manikkam et al. 2012a, b); however it seemed reasonable to assess the transcriptomic effects of transgenerational exposure within this tissue. Here, few transcriptional changes were detected in livers of male offspring of the F3 generation. In contrast, a number of changes were observed in female rat liver, possibly demonstrating an increased sensitivity, as female rats have been previously shown to be more sensitive to TCDD-induced toxicities than their male counterparts (Pohjanvirta et al. 1993; Silkworth et al. 2008). The heightened response occurring here at such a low dose, and not the higher doses, is intriguing; however, these effects were relatively weak with low statistical significance. Despite this, the alleged impacts of low-dose exposure to endocrine disruptors (such as TCDD) are known to exist (reviewed in Vandenberg et al. 2012). Tuomisto et al. observed an inverse relationship between risk for soft-tissue sarcomas and dioxin exposure (Tuomisto et al. 2004). Since exposure to high levels of dioxin has long been linked to tumour development, they proposed a hormesis-like effect for dioxin: tumorigenic at both high and low doses (Tuomisto et al. 2006). It is possible that a similar pattern would occur at the molecular level, with a large effect induced by both extremely high and low dose. Therefore, our findings warrant further study. Similarly, as previous studies have revealed an increased rate of kidney disease and pubertal and reproductive abnormalities within ancestrally-exposed animals (Manikkam et al. 2012a, b; Nilsson et al. 2012; Yu et al. 2019), transcriptomic analyses of these tissues may produce more exciting results.
Along these lines, and consistent with these previous studies (Manikkam et al. 2012b), targeted bisulfite sequencing was performed on testicular tissue from F3 rats. Despite the absence of physiological changes in testes from previous studies (Manikkam et al. 2012a), we detected numerous DMRs within this tissue. In our study, a conspicuous finding was that olfactory receptors seemed to be frequently affected by differential methylation; this in itself is quite interesting as a highly sensitive behavioural response to AHR agonists is aversion to novel foodstuffs (Lensu et al. 2011a; Mahiout and Pohjanvirta 2016), and this may involve the sense of smell. Differential methylation of Olr60 was also reported by Manikkam et al., albeit in a different tissue type (sperm rather than testis) (Manikkam et al. 2012b). Similarly, both studies identified DMRs within regulatory subunits of protein phosphatase 1 (Ppp1r27 in testis and Ppp1r14a in sperm) (Manikkam et al. 2012b); however, these are typically expressed in different tissue types (muscle and lung respectively) (Yu et al. 2014).
In conclusion, a conspicuous sex-based difference was identified in hepatic transcriptomes of unexposed F3 rats derived from TCDD-treated F0 dams through paternal germ line. In female rats, numerous genes exhibited altered expression at the lowest dose tested (30 ng/kg), whereas there were no significant changes at any dose level in male rats. Hyper- and hypo-methylated regions both inside and outside of the gene coding regions were detected in the testes of the TCDD lineage, with genes demonstrating hyper-methylation being enriched in pathways of biosynthetic processes, cell adhesion and cell–cell signalling. These findings warrant further transgenerational studies focusing on other tissues, especially in female rats after low TCDD doses.
Methods
Chemicals
2,3,7,8-Tetrachlorodibenzo-p-dioxin (TCDD) was purchased from the UFA Oil Institute (Ufa, Russia) and was determined to be over 99% pure as analyzed by gas chromatography-mass spectrometry. TCDD was dissolved in corn oil (Sigma Chemicals, St. Louis, MO) and thoroughly mixed using a magnetic stirrer. Solutions were ultra-sonicated for 20 min prior to dosing.
Animal handling
Outbred male and female Sprague–Dawley rats were obtained from Harlan Netherlands (Zeist, The Netherlands). Throughout the study, animals were subjected to regular health surveys to ensure animals were free of typical rodent pathogens (Rehbinder et al. 1996). Animals were acclimatized to the experimental conditions for one week prior to commencing the study. Rats were mated overnight and pregnancy day 0 assigned when the presence of sperm in a vaginal smear was confirmed. On gestational day 11, groups of pregnant females (n = 6–8) were treated with a single dose of TCDD (0, 30, 100, 300 and 1000 ng/kg bodyweight) dissolved in corn oil by oral gavage. Litter sizes were adjusted to 4 males and 4 females on postnatal day 1 whenever possible.
Upon weaning, same-sex littermates were housed in stainless steel, wire-mesh bottom cages with pelleted rat feed (R36, Lactamin, Stockholm, Sweden) and tap water available ad libitum. The housing environment was maintained at 21 ± 0.2 °C and relative humidity at 50 ± 3%, with a 12/12 h artificial light/dark cycle. Adult males were mated with untreated female rats to achieve F2 generations. The above procedure was repeated to further achieve F3 generations. At the end of the examination period, rats were euthanized by carbon dioxide exposure and subjected to tissue sampling. Hepatic and testicular tissue was shipped on dry ice to the analytical laboratory for processing. The study protocols were approved by the Finnish National Animal Experiment Board (Eläinkoelautakunta, ELLA; permit code: ESLH-2008–07,223/Ym-23) and all animal handling and reporting comply with ARRIVE guidelines (Kilkenny et al. 2010).
Microarray hybridization
A total of 67 animals from the F3 generation were examined (Table 1; Supplementary Table 1). Total hepatic RNA was isolated for analysis on microarrays. Briefly, 20 mg hepatic tissue was ground to a powder in liquid nitrogen using a mortar and pestle, followed by rapid homogenization using a Brinkmann Polytron (Polytron PT1600E with a PT-DA 1607 generator). RNA was extracted using an RNeasy Mini Plus Kit following the manufacturer’s recommended protocol (Qiagen, Mississauga, Canada). Total RNA was quantified using a NanoDrop UV Spectrophotometer (Thermo Scientific, Mississauga, ON) and RNA quality was verified by electrophoresis using RNA 6000 Nano kits on an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA). All samples demonstrated RNA integrity number (RIN) greater than 8. Aliquots of RNA were transported to The Centre for Applied Genomics (TCAG) at The Hospital for Sick Children (Toronto, Canada) and assayed on Affymetrix GeneChip Rat Gene 2.0 ST arrays using the manufacturer's protocols.
Statistical analysis of hepatic transcriptome
Raw microarray data (CEL files) were loaded into the R statistical environment (v3.4.0) using the affy package (v1.48.0) of the BioConductor library (Gentleman et al. 2004). All samples were processed together using the RMA algorithm (Irizarry et al. 2003). Mapping of probes to Entrez Gene IDs was performed using the custom cdf ragene20strnentrezgcdf (v21.0.0) package for R (Dai et al. 2005). Distributional and spatial homogeneity was assessed (Supplementary Figure 1) and a single female animal was identified as an outlier. This sample was removed and the remaining arrays were re-processed as described above (Supplementary Figure 2). Filtering was performed using a threshold determined by examining the normalized intensity levels of genes on chromosome Y within the female samples (removed 3007 probesets; Supplementary Figure 3a). Linear modelling was performed to identify those transcripts with altered abundance in rats from TCDD-exposed lineages relative to rats from control lineages. Models were applied to determine coefficients based on the interaction between the sex and dose variables, with familial lineage included as a blocking variable. Contrasts were fit to compare each TCDD-treated lineage with controls. The standard error of the coefficients was adjusted using an empirical Bayes moderation of the standard error (Smyth 2004) and model-based t-tests were applied to determine whether a coefficient was significantly different from zero. A false-discovery rate (FDR) adjustment for multiple testing was applied (Storey and Tibshirani 2003). Linear modelling and subsequent adjustments were performed using the limma (v3.32.5) package for R. For downstream analyses, a significance threshold of padj < 0.1 was used to define genes with significantly altered mRNA abundance (Supplementary Figure 3c). Alternate models were applied as above to identify differentially abundant transcripts based on sex and TCDD (alone or in combination; Supplementary Figure 3e). Visualizations were generated using the BPG package (v5.7.6) (P'ng et al. 2017), with the lattice (v0.20-35) and latticeExtra (v0.6-28) packages for R. Results were contrasted with previously published datasets using the TCDD.Transcriptomics (Prokopec et al. 2017) package for R.
Bisulfite sequencing
Library preparation, quality control, target capture and sequencing was performed at the Biomedicum Functional Genomics Unit (FuGU) at the University of Helsinki. Target capture was carried out using the Agilent SureSelect XT Rat Methyl-Seq kit. Sequencing was performed using Illumina NextSeq 500 with the NextSeq 500/550 v2 kit (High Output) to produce single-end directional reads (75 bp read length).
Alignment and data processing
Raw sequence data were aligned with Bismark (v0.15.0) (Krueger and Andrews 2011), using Bowtie2 (v2.2.6), to the rattus norvegicus (rn6) reference genome. Bismark was run using default settings, using single-end, unidirectional fastq files. Aligned files were coordinate sorted using samtools (v1.3) and library-level merged with duplicate reads marked using picard tools (v1.141). Finally, Bismark’s methylation extractor tool, with –comprehensive, –merge_non_CpG, –cytosine_report, –CX, –split_by_chromosome, and –bedGraph settings, was used to extract the number of methylated and unmethylated reads at each CpG position in each sample.
The resulting data files for each sample were loaded into the R statistical environment (v3.3.1) and pooled across treatment groups using the DMRcaller package (v1.6.0) on a per-chromosome basis (due to computational intensity). Target regions were first converted to rn6 coordinates from rn4 using liftover (v20111127) and target coverage was estimated across these regions for each chromosome using the compute Methylation Profile function for each CX (CG, CHG, CHH) context. The overall methylation profiles for each group were visualized using the DMRcaller function plot Methylation Profile from data using a window size of 1000 bp for each CX context (Supplementary Figures 4–5).
Differentially methylated regions (DMRs) were identified using a chromosome-wide or gene-wise approach. Gene-wise DMRs were identified using the filterDMRs function in DMRcaller, for each chromosome independently. Specifically, regions evaluated were restricted to genes (Rnor 6.0.88) that had some overlap with the Methyl-Seq target regions (following liftOver conversion from rn4 to rn6 coordinates). Two chromosomes (chrY and chrM) had no targeted regions, therefore all genes on these chromosomes were evaluated. Default methods (noise filter method with a window size of 100 bp with a triangular kernel function) were used, with the score test for statistical significance, followed by (default) adjustment for multiple testing using the Benjamini and Hochberg method. Genes were examined if it contained at least 1 CG site, with minimum of 10 reads per treatment group and a minimum difference in methylation of 1% (the proportion of reads with a methylated CG differed between groups by at least 1%). DMRs less than 200 bp apart were then merged, with respect given to direction of change. Statistically significant DMRs (padj < 0.01) were then visualized using the plot Local Methylation Profile function, using a region ± 5 Kbp around the DMR. Again, default methods were used, however with a window size of 1000 for smoothing. Similarly, chromosome-wide DMRs were identified using the compute DMRs function, for each chromosome independently. Default methods (noise filter method with a window size of 100 bp with a triangular kernel function) were used, again with the score test for statistical significance and correction for multiple testing as above. Statistically significant DMRs (padj < 0.001; DMRs with gene overlap within 1000 bp up/downstream of the DMR) were then visualized as above.
Pathway analysis was performed using those genes with either increased or decreased methylation in the TCDD-exposed lineage relative to the control lineage (up- and down- methylated genes analysed separately). Pathway analysis was performed using gProfileR (v0.6.1) for R, using the rnorvegicus ‘GO’ database, with a minimum overlap of 1 gene, and application of false discovery rate adjustment for multiple testing.
Global DNA methylation status determination
Global DNA methylation status was determined in the liver and testis of F3 generation rats by colorimetric MethylFlash™ Methylated DNA Quantification Kit (Epigentek, Farmingdale, NY) according to the manufacturer’s instructions.
Data availability
All data have been deposited in the Gene Expression Omnibus (GSE126216) and are publically available. Raw, normalized, and modelled data are also available in the TCDD Transcriptomics package for R (v2.2.5).
References
Ansbaugh N, Shannon J, Mori M, Farris PE, Garzotto M (2013) Agent Orange as a risk factor for high-grade prostate cancer. Cancer 119(13):2399–2404. https://doi.org/10.1002/cncr.27941
Arechederra M, Daian F, Yim A et al (2018) Hypermethylation of gene body CpG islands predicts high dosage of functional oncogenes in liver cancer. Nat Commun 9(1):3164. https://doi.org/10.1038/s41467-018-05550-5
Baccarelli A, Giacomini SM, Corbetta C et al (2008) Neonatal thyroid function in Seveso 25 years after maternal exposure to dioxin. PLoS Med 5(7):e161. https://doi.org/10.1371/journal.pmed.005016107-PLME-RA-2317
Baccarelli A, Pesatori AC, Consonni D et al (2005) Health status and plasma dioxin levels in chloracne cases 20 years after the Seveso Italy accident. Br J Dermatol 152(3):459–465. https://doi.org/10.1111/j.1365-2133.2005.06444.x
Baker TR, Peterson RE, Heideman W (2014) Using zebrafish as a model system for studying the transgenerational effects of dioxin. Toxicol Sci 138(2):403–411. https://doi.org/10.1093/toxsci/kfu006
Bickel MH (1982) Polychlorinated persistent compounds. Experientia 38(8):879–882
Boutros PC, Yan R, Moffat ID, Pohjanvirta R, Okey AB (2008) Transcriptomic responses to 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD) in liver: comparison of rat and mouse. BMC Genomics 9:419. https://doi.org/10.1186/1471-2164-9-419
Boutros PC, Yao CQ, Watson JD et al (2011) Hepatic transcriptomic responses to TCDD in dioxin-sensitive and dioxin-resistant rats during the onset of toxicity. Toxicol Appl Pharmacol 251(2):119–129. https://doi.org/10.1016/j.taap.2010.12.010
Boverhof DR, Burgoon LD, Tashiro C et al (2006) Comparative toxicogenomic analysis of the hepatotoxic effects of TCDD in Sprague Dawley rats and C57BL/6 mice. Toxicol Sci 94(2):398–416. https://doi.org/10.1093/toxsci/kfl100
Bruner-Tran KL, Ding T, Yeoman KB, Archibong A, Arosh JA, Osteen KG (2014) Developmental exposure of mice to dioxin promotes transgenerational testicular inflammation and an increased risk of preterm birth in unexposed mating partners. PLoS ONE 9(8):e105084. https://doi.org/10.1371/journal.pone.0105084
Campion CM, Leon Carrion S, Mamidanna G, Sutter CH, Sutter TR, Cole JA (2016) Role of EGF receptor ligands in TCDD-induced EGFR down-regulation and cellular proliferation. Chem Biol Interact 253:38–47. https://doi.org/10.1016/j.cbi.2016.04.031
Chagnon YC, Chen WJ, Perusse L et al (1997) Linkage and association studies between the melanocortin receptors 4 and 5 genes and obesity-related phenotypes in the Quebec Family Study. Mol Med 3(10):663–673
Dai M, Wang P, Boyd AD et al (2005) Evolving gene/transcript definitions significantly alter the interpretation of GeneChip data. Nucleic Acids Res 33(20):e175. https://doi.org/10.1093/nar/gni179
Dixon G, Liao Y, Bay LK, Matz MV (2018) Role of gene body methylation in acclimatization and adaptation in a basal metazoan. Proc Natl Acad Sci USA 115(52):13342–13346. https://doi.org/10.1073/pnas.1813749115
el-Sabeawy F, Wang S, Overstreet J, Miller M, Lasley B, Enan E (1998) Treatment of rats during pubertal development with 2,3,7,8-tetrachlorodibenzo-p-dioxin alters both signaling kinase activities and epidermal growth factor receptor binding in the testis and the motility and acrosomal reaction of sperm. Toxicol Appl Pharmacol 150(2):427–442. https://doi.org/10.1006/taap.1998.8426
Fletcher N, Wahlstrom D, Lundberg R et al (2005) 2,3,7,8-Tetrachlorodibenzo-p-dioxin (TCDD) alters the mRNA expression of critical genes associated with cholesterol metabolism, bile acid biosynthesis, and bile transport in rat liver: a microarray study. Toxicol Appl Pharmacol 207(1):1–24. https://doi.org/10.1016/j.taap.2004.12.003
Forgacs AL, Kent MN, Makley MK et al (2012) Comparative metabolomic and genomic analyses of TCDD-elicited metabolic disruption in mouse and rat liver. Toxicol Sci 125(1):41–55. https://doi.org/10.1093/toxsci/kfr262
Franc MA, Moffat ID, Boutros PC et al (2008) Patterns of dioxin-altered mRNA expression in livers of dioxin-sensitive versus dioxin-resistant rats. Arch Toxicol 82(11):809–830. https://doi.org/10.1007/s00204-008-0303-0
Gentleman RC, Carey VJ, Bates DM et al (2004) Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5(10):R80. https://doi.org/10.1186/gb-2004-5-10-r80
Geyer HJ, Schramm KW, Feicht EA et al (2002) Half-lives of tetra-, penta-, hexa-, hepta-, and octachlorodibenzo-p-dioxin in rats, monkeys, and humans—a critical review. Chemosphere 48(6):631–644
He J, Chen DL, Samocha-Bonet D et al (2016) Fibroblast growth factor-1 (FGF-1) promotes adipogenesis by downregulation of carboxypeptidase A4 (CPA4) - a negative regulator of adipogenesis implicated in the modulation of local and systemic insulin sensitivity. Growth Factors 34(5–6):210–216. https://doi.org/10.1080/08977194.2017.1285764
Huang H, Reed CP, Zhang JS, Shridhar V, Wang L, Smith DI (1999) Carboxypeptidase A3 (CPA3): a novel gene highly induced by histone deacetylase inhibitors during differentiation of prostate epithelial cancer cells. Cancer Res 59(12):2981–2988
Ikeda M, Tamura M, Yamashita J, Suzuki C, Tomita T (2005) Repeated in utero and lactational 2,3,7,8-tetrachlorodibenzo-p-dioxin exposure affects male gonads in offspring, leading to sex ratio changes in F2 progeny. Toxicol Appl Pharmacol 206(3):351–355. https://doi.org/10.1016/j.taap.2004.11.019
Irizarry RA, Hobbs B, Collin F et al (2003) Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4(2):249–264. https://doi.org/10.1093/biostatistics/4.2.2494/2/249
Karman BN, Basavarajappa MS, Craig ZR, Flaws JA (2012) 2,3,7,8-Tetrachlorodibenzo-p-dioxin activates the aryl hydrocarbon receptor and alters sex steroid hormone secretion without affecting growth of mouse antral follicles in vitro. Toxicol Appl Pharmacol 261(1):88–96. https://doi.org/10.1016/j.taap.2012.03.015
Kayashima T, Yamasaki K, Yamada T et al (2003) The novel imprinted carboxypeptidase A4 gene ( CPA4) in the 7q32 imprinting domain. Hum Genet 112(3):220–226. https://doi.org/10.1007/s00439-002-0891-3
Kilkenny C, Browne WJ, Cuthill IC, Emerson M, Altman DG (2010) Improving bioscience research reporting: the ARRIVE guidelines for reporting animal research. PLoS Biol 8(6):e1000412. https://doi.org/10.1371/journal.pbio.1000412
Kociba RJ, Keyes DG, Beyer JE et al (1978) Results of a two-year chronic toxicity and oncogenicity study of 2,3,7,8-tetrachlorodibenzo-p-dioxin in rats. Toxicol Appl Pharmacol 46(2):279–303. https://doi.org/10.1016/0041-008x(78)90075-3
Krueger F, Andrews SR (2011) Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27(11):1571–1572. https://doi.org/10.1093/bioinformatics/btr167
Lee J, Prokopec SD, Watson JD, Sun RX, Pohjanvirta R, Boutros PC (2015) Male and female mice show significant differences in hepatic transcriptomic response to 2,3,7,8-tetrachlorodibenzo-p-dioxin. BMC Genomics 16:625. https://doi.org/10.1186/s12864-015-1840-6
Lee SH, Lee DY, Son WK, Joo WA, Kim CW (2005) Proteomic characterization of rat liver exposed to 2,3,7,8-tetrachlorobenzo-p-dioxin. J Proteome Res 4(2):335–343. https://doi.org/10.1021/pr049830s
Leng L, Chen X, Li CP, Luo XY, Tang NJ (2014) 2,3,7,8-Tetrachlorodibezo-p-dioxin exposure and prostate cancer: a meta-analysis of cohort studies. Public Health 128(3):207–213. https://doi.org/10.1016/j.puhe.2013.10.006
Lensu S, Tuomisto JT, Tuomisto J, Pohjanvirta R (2011a) Characterization of the 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD)-provoked strong and rapid aversion to unfamiliar foodstuffs in rats. Toxicology 283(2–3):140–150. https://doi.org/10.1016/j.tox.2011.03.007
Lensu S, Tuomisto JT, Tuomisto J, Viluksela M, Niittynen M, Pohjanvirta R (2011b) Immediate and highly sensitive aversion response to a novel food item linked to AH receptor stimulation. Toxicol Lett 203(3):252–257. https://doi.org/10.1016/j.toxlet.2011.03.025
Mahiout S, Pohjanvirta R (2016) Aryl hydrocarbon receptor agonists trigger avoidance of novel food in rats. Physiol Behav 167:49–59. https://doi.org/10.1016/j.physbeh.2016.08.033
Manikkam M, Guerrero-Bosagna C, Tracey R, Haque MM, Skinner MK (2012a) Transgenerational actions of environmental compounds on reproductive disease and identification of epigenetic biomarkers of ancestral exposures. PLoS ONE 7(2):e31901. https://doi.org/10.1371/journal.pone.0031901
Manikkam M, Tracey R, Guerrero-Bosagna C, Skinner MK (2012b) Dioxin (TCDD) induces epigenetic transgenerational inheritance of adult onset disease and sperm epimutations. PLoS ONE 7(9):e46249. https://doi.org/10.1371/journal.pone.0046249
Mocarelli P, Gerthoux PM, Needham LL et al (2011) Perinatal exposure to low doses of dioxin can permanently impair human semen quality. Environ Health Perspect 119(5):713–718. https://doi.org/10.1289/ehp.1002134
Moffat ID, Boutros PC, Chen H, Okey AB, Pohjanvirta R (2010) Aryl hydrocarbon receptor (AHR)-regulated transcriptomic changes in rats sensitive or resistant to major dioxin toxicities. BMC Genomics 11:263. https://doi.org/10.1186/1471-2164-11-263
Morgan C, Thomas RE, Ma W, Novotny MV, Cone RD (2004) Melanocortin-5 receptor deficiency reduces a pheromonal signal for aggression in male mice. Chem Senses 29(2):111–115
Nau H, Bass R, Neubert D (1986) Transfer of 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD) via placenta and milk, and postnatal toxicity in the mouse. Arch Toxicol 59(1):36–40
Nault R, Kim S, Zacharewski TR (2013) Comparison of TCDD-elicited genome-wide hepatic gene expression in Sprague–Dawley rats and C57BL/6 mice. Toxicol Appl Pharmacol 267(2):184–191. https://doi.org/10.1016/j.taap.2012.11.028
Niittynen M, Simanainen U, Syrjala P et al (2007) Differences in acute toxicity syndromes of 2,3,7,8-tetrachlorodibenzo-p-dioxin and 1,2,3,4,7,8-hexachlorodibenzo-p-dioxin in rats. Toxicology 235(1–2):39–51. https://doi.org/10.1016/j.tox.2007.03.012
Nilsson E, Larsen G, Manikkam M, Guerrero-Bosagna C, Savenkova MI, Skinner MK (2012) Environmentally induced epigenetic transgenerational inheritance of ovarian disease. PLoS ONE 7(5):e36129. https://doi.org/10.1371/journal.pone.0036129
P'ng C, Green J, Chong LC, et al. (2017) BPG: seamless, automated and interactive visualization of scientific data. bioRxiv doi:10.1101/156067
Pastorelli R, Carpi D, Campagna R et al (2006) Differential expression profiling of the hepatic proteome in a rat model of dioxin resistance: correlation with genomic and transcriptomic analyses. Mol Cell Proteom 5(5):882–894. https://doi.org/10.1074/mcp.M500415-MCP200
Pohjanvirta R, Tuomisto J (1994) Short-term toxicity of 2,3,7,8-tetrachlorodibenzo-p-dioxin in laboratory animals: effects, mechanisms, and animal models. Pharmacol Rev 46(4):483–549
Pohjanvirta R, Unkila M, Tuomisto J (1993) Comparative acute lethality of 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD), 1,2,3,7,8-pentachlorodibenzo-p-dioxin and 1,2,3,4,7,8-hexachlorodibenzo-p-dioxin in the most TCDD-susceptible and the most TCDD-resistant rat strain. Pharmacol Toxicol 73(1):52–56
Pohjanvirta R, Vartiainen T, Uusi-Rauva A, Monkkonen J, Tuomisto J (1990) Tissue distribution, metabolism, and excretion of 14C-TCDD in a TCDD-susceptible and a TCDD-resistant rat strain. Pharmacol Toxicol 66(2):93–100
Prokopec SD, Houlahan KE, Sun RX et al (2017) Compendium of TCDD-mediated transcriptomic response datasets in mammalian model systems. BMC Genomics 18(1):78. https://doi.org/10.1186/s12864-016-3446-z
Prokopec SD, Watson JD, Lee J, Pohjanvirta R, Boutros PC (2015) Sex-related differences in murine hepatic transcriptional and proteomic responses to TCDD. Toxicol Appl Pharmacol 284(2):188–196. https://doi.org/10.1016/j.taap.2015.02.012
Rehbinder C, Baneux P, Forbes D et al (1996) FELASA recommendations for the health monitoring of mouse, rat, hamster, gerbil, guinea pig and rabbit experimental units. Report of the Federation of European Laboratory Animal Science Associations (FELASA) Working Group on Animal Health accepted by the FELASA Board of Management. Lab Anim 30(3):193–208
Reimand J, Kull M, Peterson H, Hansen J, Vilo J (2007) g:Profiler—a web-based toolset for functional profiling of gene lists from large-scale experiments. Nucleic Acids Res 35(Web server issue):W193–W200
Schecter A, Birnbaum L, Ryan JJ, Constable JD (2006) Dioxins: an overview. Environ Res 101(3):419–428. https://doi.org/10.1016/j.envres.2005.12.003
Shen C, Chen Y, Huang S et al (2009) Dioxin-like compounds in agricultural soils near e-waste recycling sites from Taizhou area, China: chemical and bioanalytical characterization. Environ Int 35(1):50–55. https://doi.org/10.1016/j.envint.2008.07.005
Shukla C, Britton SL, Koch LG, Novak CM (2012) Region-specific differences in brain melanocortin receptors in rats of the lean phenotype. NeuroReport 23(10):596–600. https://doi.org/10.1097/WNR.0b013e328354f5c1
Silkworth JB, Carlson EA, McCulloch C, Illouz K, Goodwin S, Sutter TR (2008) Toxicogenomic analysis of gender, chemical, and dose effects in livers of TCDD- or aroclor 1254-exposed rats using a multifactor linear model. Toxicol Sci 102(2):291–309. https://doi.org/10.1093/toxsci/kfm313
Smyth GK (2004) Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3:3. https://doi.org/10.2202/1544-6115.1027
Storey JD, Tibshirani R (2003) Statistical significance for genomewide studies. Proc Natl Acad Sci USA 100(16):9440–9445. https://doi.org/10.1073/pnas.1530509100
Suzuki T, Gao J, Ishigaki Y et al (2017) ER stress protein CHOP mediates insulin resistance by modulating adipose tissue macrophage polarity. Cell Rep 18(8):2045–2057. https://doi.org/10.1016/j.celrep.2017.01.076
Tanco S, Zhang X, Morano C, Aviles FX, Lorenzo J, Fricker LD (2010) Characterization of the substrate specificity of human carboxypeptidase A4 and implications for a role in extracellular peptide processing. J Biol Chem 285(24):18385–18396. https://doi.org/10.1074/jbc.M109.060350
Tuomisto J, Pekkanen J, Kiviranta H et al (2004) Soft-tissue sarcoma and dioxin: a case–control study. Int J Cancer 108(6):893–900. https://doi.org/10.1002/ijc.11635
Tuomisto J, Pekkanen J, Kiviranta H et al (2006) Dioxin cancer risk—example of hormesis? Dose Response 3(3):332–341. https://doi.org/10.2203/dose-response.003.03.004
Vandenberg LN, Colborn T, Hayes TB et al (2012) Hormones and endocrine-disrupting chemicals: low-dose effects and nonmonotonic dose responses. Endocr Rev 33(3):378–455. https://doi.org/10.1210/er.2011-1050
Viluksela M, Bager Y, Tuomisto JT et al (2000) Liver tumor-promoting activity of 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD) in TCDD-sensitive and TCDD-resistant rat strains. Cancer Res 60(24):6911–6920
Von Burg R (1988) A rat RNA-Seq transcriptomic BodyMap across 11 organs and 4 developmental stages. J Appl Toxicol 8(2):145–148
Yao CQ, Prokopec SD, Watson JD et al (2012) Inter-strain heterogeneity in rat hepatic transcriptomic responses to 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD). Toxicol Appl Pharmacol 260(2):135–145. https://doi.org/10.1016/j.taap.2012.02.001
Yi SW, Ryu SY, Ohrr H, Hong JS (2014) Agent Orange exposure and risk of death in Korean Vietnam veterans: Korean Veterans Health Study. Int J Epidemiol 43(6):1825–1834. https://doi.org/10.1093/ije/dyu183
Yu K, Zhang X, Tan X et al (2019) Transgenerational impairment of ovarian induced by 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD) associated with Igf2 and H19 in adult female rat. Toxicology 428:152311. https://doi.org/10.1016/j.tox.2019.152311
Yu Y, Fuscoe JC, Zhao C et al (2014) A rat RNA-Seq transcriptomic BodyMap across 11 organs and 4 developmental stages. Nat Commun 5:3230. https://doi.org/10.1038/ncomms4230
Acknowledgements
The authors thank all members of the Boutros lab for helpful suggestions, and Dr. Selma Mahiout and Ms. Susanna Lukkarinen for practical aid.
Funding
This study was conducted with the support of the Ontario Institute for Cancer Research to PCB through funding provided by the government of Ontario. It was also financially supported by the Academy of Finland (Grant n# 261232 for RP). This work was supported by the NIH/NCI under award number P30CA016042.
Author information
Authors and Affiliations
Contributions
Sample preparation: MV, HMM. Performed statistical and bioinformatics analyses: SDP. Wrote the first draft of the manuscript: SDP. Initiated the project: MV, RP, PCB. Supervised research: MV, RP, PCB. Generated tools and reagents: SDP. Approved the manuscript: all authors.
Corresponding authors
Ethics declarations
Conflict of interest
All authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
204_2020_2730_MOESM1_ESM.tif
Supplementary Figure 1: Quality Assessment of Microarrays. All arrays were normalized simultaneously (male and female together). Distribution of intensity values a) pre- and b) post- RMA processing. c) Average intensity across probesets per sample to demonstrate RNA degradation and d) Heatmap showing inter-array correlation; clustering was performed using the DIANA algorithm with a Pearson’s correlation similarity metric (TIF 2001 kb)
204_2020_2730_MOESM2_ESM.tif
Supplementary Figure 2: Quality Assessment of Microarrays with outliers removed. A single female F3 animal was identified as an outlier (Supplementary Figure 1). This sample was excluded and remaining arrays were normalized simultaneously (male and female together). Distribution of intensity values a) pre- and b) post- RMA processing. c) Average intensity across probesets per sample to demonstrate RNA degradation and d) Heatmap showing inter-array correlation; clustering was performed using the DIANA algorithm with a Pearson’s correlation similarity metric (TIF 2003 kb)
204_2020_2730_MOESM3_ESM.tif
Supplementary Figure 3: a) Determination of background intensity threshold. Normalized intensity scores were compared across probes targeting genes on chromosome X, Y or all autosomes for male and female samples; background intensity threshold was determined using the intensity of chromosome Y probes within female rats. b) Following preprocessing, gene-wise variance was calculated across all samples and the normalized intensity values of the top most variant genes (207 genes with a variance > 0.5) visualized. DIANA hierarchical clustering was performed using Pearson’s correlation as a similarity estimate. Samples clustered perfectly according to sex (ARI = 1) with additional grouping by treatment (ARI = 0.58). c) Linear modelling provided coefficients and FDRadjusted p-values giving the effect of TCDD exposure on each gene based on sex and dose. d) Effect sizes and significance for typical “AHR-core” genes are shown. Dot size indicates coefficient (fold-change) and dot colour indicates direction of change while background shading indicates FDR-adjusted p-value. e) Coefficients and FDR-adjusted p-values of alternate models, to identify sex-specific, TCDD-specific or interaction effects (TIF 3668 kb)
204_2020_2730_MOESM4_ESM.pdf
Supplementary Figure 4: Coverage estimates for chromosomes 1-20 and X when considering only targeted regions and each CX context separately (PDF 1607 kb)
204_2020_2730_MOESM5_ESM.pdf
Supplementary Figure 5: Methylation profiles for chromosomes 1-20 and X when considering only targeted regions and each CX context separately (PDF 5376 kb)
Rights and permissions
About this article
Cite this article
Prokopec, S.D., Viluksela, M., Miettinen, H.M. et al. Transgenerational epigenetic and transcriptomic effects of 2,3,7,8-tetrachlorodibenzo-p-dioxin exposure in rat. Arch Toxicol 94, 1613–1624 (2020). https://doi.org/10.1007/s00204-020-02730-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00204-020-02730-5