Abstract
Objectives
The differential between high-grade glioma (HGG) and metastasis remains challenging in common radiological practice. We compare different natural language processing (NLP)–based deep learning models to assist radiologists based on data contained in radiology reports.
Methods
This retrospective study included 185 MRI reports between 2010 and 2022 from two different institutions. A total of 117 reports were used for the training and 21 were reserved for the validation set, while the rest were used as a test set. A comparison of the performance of different deep learning models for HGG and metastasis classification has been carried out. Specifically, Convolutional Neural Network (CNN), Bidirectional Long Short-Term Memory (BiLSTM), a hybrid version of BiLSTM and CNN, and a radiology-specific Bidirectional Encoder Representations from Transformers (RadBERT) model were used.
Results
For the classification of MRI reports, the CNN network provided the best results among all tested, showing a macro-avg precision of 87.32%, a sensitivity of 87.45%, and an F1 score of 87.23%. In addition, our NLP algorithm detected keywords such as tumor, temporal, and lobe to positively classify a radiological report as HGG or metastasis group.
Conclusions
A deep learning model based on CNN enables radiologists to discriminate between HGG and metastasis based on MRI reports with high-precision values. This approach should be considered an additional tool in diagnosing these central nervous system lesions.
Clinical relevance statement
The use of our NLP model enables radiologists to differentiate between patients with high-grade glioma and metastasis based on their MRI reports and can be used as an additional tool to the conventional image-based approach for this challenging task.
Key Points
• Differential between high-grade glioma and metastasis is still challenging in common radiological practice.
• Natural language processing (NLP)–based deep learning models can assist radiologists based on data contained in radiology reports.
• We have developed and tested a natural language processing model for discriminating between high-grade glioma and metastasis based on MRI reports that show high precision for this task.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The differential diagnosis between central nervous system (CNS) solitary-enhancing lesions, including high-grade gliomas (HGG) and brain metastasis, is still a challenge in common radiological practice [1]. Since both lesions may show similar morphological features on conventional MRI related to enhancement, necrosis, or vasogenic edema, the differential between HGG and solitary metastasis usually needs advanced MRI approaches [2]. In the last two decades, hundreds of papers have addressed the capability of advanced MRI sequences such as diffusion-weighted imaging (DWI), perfusion-weighted imaging (PWI), including dynamic susceptibility contrast (DSC) and dynamic contrast-enhanced (DCE), MR spectroscopy, arterial spin labeling (ASL), or amide proton transfer (APT) among others for this task [3,4,5]. These advanced modalities have provided new radiological features, including quantifiable parameters, for improving the differential diagnosis between both lesions. Moreover, in the last decade, artificial intelligence (AI) solutions based on images derived from conventional or advanced MRI sequences are providing new insights and relevant information for increasing the accuracy, sensitivity, and specificity of MRI in this specific scenario [6,7,8].
At this point, other potential sources of information for feeding AI algorithms are electronic health records (EHR) and, in our case, radiology reports [9]. Radiology reports contain all the information related to the patient’s demographics, clinical history, and, most importantly, the description of radiological findings (including conclusion or report summary), in other words, all the signs and features that radiologists identify during their reporting process [10]. In this scenario, natural language processing (NLP), a division of AI dedicated to giving computers the ability to interpret and understand human language, primarily based on machine learning (ML), has emerged as a promising tool to extract information from radiology reports and establish relationships between them from a general to a word-based level, usually hidden from the human eye [11, 12]. Moreover, NLP tools can manage large datasets in ways humans cannot. In our experience, this scenario is the breeding ground for applying this NLP technology to help radiologists face specific radiological questions[13, 14].
In this paper, we analyzed different NLP-based deep learning systems to distinguish between HGG and metastasis based solely on the information in radiological reports to develop the best automatic decision support system.
Methods
Data collection dataset
Ethical approval was waived by our local ethics committee because of the retrospective nature of the study, based on radiology reports, and all the procedures being performed were part of the routine radiology practice. A retrospective review of brain MRI reports performed at two different radiology departments between June 2010 and June 2022 was completed. These reports were exported as anonymized text files from each radiology department’s radiology information system (RIS). Inclusion criteria contained MRI reports with diagnosis of HGG or metastasis (proved after biopsy or surgery). Exclusion criteria comprised MRI reports with formal defects (i.e., absence of clinical information or conclusion section). The dataset was reviewed and annotated by consensus by two radiologists with more than 10 years of experience in a binary way: HGG or metastasis.
The corpus comprised 185 reports (99 from institution A and 86 from institution B), including the findings description and conclusions sections (Fig. 1). A total of 11 annotated reports were excluded due to formal defects. Reports were created in Spanish language; however, for better and potential reproducibility of our NLP algorithm, they were translated into English language and revised by an expert in medical English language for ensuring the accuracy of the translation. Maximum and median report lengths (measured in number of words) were 499 and 252 for institution A, and 416 and 186 for institution B, respectively (Fig. 2).
Model training and validation
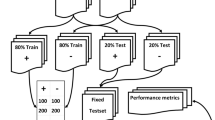
For training and testing the ML models, 117 reports were used as the training set and 21 reports constituted the validation set, while the rest of the data (47) were considered an independent test dataset.
Reports were pre-processed using tokenization based on whitespace (punctuation and other special characters, such as parentheses, were considered separate tokens that contain helpful semantic content within reports). For this purpose, we use the NLTK library and the Python v3.8 programming language [15]. Moreover, to avoid biases in the algorithm, keywords considered highly representative of both HGG and metastasis were eliminated from the texts (Table 1).
Deep learning models
Diverse deep learning models were trained and tested to differentiate between HGG and metastasis using the manually annotated radiology reports as the ground truth. Four different deep learning architectures were evaluated: a simple Convolutional Neural Network (CNN), a Bidirectional Long Short-Term Memory (BiLSTM) network, and a hybrid model comprising a bidirectional LSTM followed by a CNN and a fine-tuned pre-trained model of BERT adapted to radiology as a classifier (RadBERT).
Our proposed CNN used a convolutional layer and a global max-pooling layer to identify the text’s most salient location for each learned feature (Fig. 1 supplementary material). The bidirectional LSTM (BiLSTM) approach processes the input text storing the semantics in two directions, one for positive time direction and another for negative time direction. This type of recurrent network can capture contextual information and long-term dependencies (Fig. 2 supplementary material). A hybrid of bidirectional LSTM and CNN architecture shown in Fig. 3 of the supplementary material (BiLSTM-CNN) was also used to differentiate between HGG and metastasis. The recurrent BiLSTM layer can serve as a language feature encoder from sequences of semantic word embeddings. Then, the convolution layers can encode the category-related features provided by the BiLSTM, while the latter dense layers tune the model for the classification task. For all of these deep learning models used and described so far (CNN, BiLSTM, and BiLSTM-CNN), the first input layer consists of FastText (https://fasttext.cc/docs/en/english-vectors.html) word embeddings with 2-million-word vectors trained with sub-word information in Common Crawl (600B tokens). Because of the number of tokens in these word embeddings, they can accurately represent the textual information of radiological reports. The report tokens were embedded in a vector space using pre-trained FastText.
Finally, we also explore the capability of BERT as a language model to detect the presence of HGG and metastasis. In our case, we fine-tuned the BERT model adapted to radiology named RadBERT (Fig. 4 supplementary material). RadBERT was pre-trained with millions of radiological reports from the US Department of Veterans Affairs healthcare system across the country on various linguistic models [16]. The pre-processed texts belonging to our dataset were tokenized with WordPiece as sub-word tokens and entered into the model.
Different model parameters, including network depth, units per layer, optimizers, or activation functions, were evaluated and compared using a grid search to identify optimal architecture parameters. Table 2 summarizes the hyperparameters selected for each model. Occurrence rates of the most common words for HGG and metastasis categories are shown in Fig. 3. The output of all the deep learning models employed was projected through dense connections to a layer of size 2, one unit for each finding (HGG and metastasis). A SoftMax activation function with the multi-class target was applied to the output.
For the development of the deep learning methods, the Python v3.8 programming language was used along with packages such as keras, tensorflow, torch, and transformers.
Statistical analysis
The primary evaluation metrics used consisted of standard measures from the NLP community, namely precision, sensitivity, F1 score, and area under the ROC curve (AUC).
Results
Patients’ demographics and dataset features
Patient’s age included in the study ranged between 32 and 86 years old, (mean 62 years old). Regarding sex, 59% of patients are male and 41% female.
Algorithms trained with NLP were used with our test dataset consistent on 47 MRI brain reports. Twenty-five of the 47 reports were classified as HGG, while the rest (22 reports) were annotated as metastasis. In addition, in order to have consistent variability in the test set, this set also contained a diversity of reports from each institution: 27 reports from institution A and 20 reports from institution B.
Performance evaluation
Table 3 shows the models’ relative performance evaluated in the HGG and metastasis classification task on the test set. Regarding the HGG category, we obtained values above 76% in the F1 score. BiLSTM offers the lowest F1 and precision (76.36% and 70%, respectively), and RadBERT provides a sensibility of 76%. The best results for HGG detection, and taking into account the F1 score, were achieved using the CNN, specifically, over 91% precision, 84 sensitivity, and 87.5% F1 score.
For the detection of metastasis, results similar to the previous ones occur. In this case, the F1 score is in the range of 66% and 87%. The BiLSTM network offers a lower value (66.67% of F1) and a 59.09% sensitivity. In terms of precision, the result obtained by the hybrid BiLSTM-CNN network stands out with a 92.86%. Overall, the CNN network achieved the best results for metastasis classification, with 83.33%, 90.91%, and 86.96% precision, sensitivity, and F1, respectively.
Finally, the AUC metric has been reported to evaluate the true and false positive rates. In this scenario, CNN achieves 87.45%, while the BiLSTM network obtains 71.54%.
CNN model results analysis
The CNN neural network provided the best performance, and Table 4 shows the results in detail, including the macro-average and weighted average metrics. Concerning the macro-avg metric, the overall precision achieved by the system is 87.32%, while the sensitivity is 87.45%, and for the F1 metric, it obtains 87.23%. The weighted average also obtained similar results, 87.57%, 87.23%, and 87.25% of precision, sensitivity, and F1, respectively.
Our CNN model was used to classify all the corpus. Figure 4 shows the matrix confusion analysis with the number of true positives (TP), true negatives (TN), false positives (FP), and false negatives (FN). Among 47 radiological reports, the CNN network does not classify 12% correctly (6 documents). Instead, the system correctly labels 41 documents. For detecting HGG, CNN correctly predicts 21 cases (TP), obtaining 4 FN, 2 FP, and 20 TN. On the other hand, for the automatic detection of metastasis, CNN offers 20 TP, 2 FN, 4 FP, and 21 TN.
Our NLP algorithm detected keywords for positively classifying a radiology report as HGG or metastasis group. Terms such as tumor, temporal, lobe, foci, corpus, callosum, necrotic, or temporal showed the highest positive significance for determining radiology reports as HGG. Terms like CT (computed tomography), DTI (diffusion tensor imaging), or LV (lateral ventricles) showed the highest positive significance value for determining radiology reports as metastasis. In this line, the exact words are negative terms for classifying radiology reports into the opposite group (Fig. 5).
Explainability CNN model
For a better explanation of why our NLP solution misclassified these six cases, we applied the LIME explainability system [17, 18]. In four of these six cases, the algorithm incorrectly classified as metastases four HGG (in two of these cases, a plausible explicability could be related to “multifocal HGG” described in the report). In the other cases, the system misclassified as HGG two metastases (in one of these cases, probably because of the displacement of “corpus callosum” by the mass effect while in the other case, the use of words like “tumor necrosis” conditioned the misclassification as HGG instead of metastasis) (Fig. 5 supplementary material).
Discussion
After analyzing different models, our CNN has achieved an AUC of 87.45% based on how HGG and metastasis are described in radiology reports. Models that involve convolution layers such as CNN and BiLSTM-CNN have achieved the best results, probably because the convolutional architecture using the pre-trained word embeddings can represent the corpus more accurately. For example, the word embeddings selected for the evaluated neural networks were trained on 600 billion tokens, while RadBERT was trained on 466 million tokens [19]. Moreover, the CNN operates locally and does not rely on positional encodings as an order signal to the model network identifying the words that are most meaningful to the task by detecting and establishing that words such as “corpus” or “callosum” which are related to HGG since HGG usually involves the corpus callosum [20]. In the same line, terms such as “tumor,” “necrotic,” or “cyst” appear linked to the HGG category, probably since HGG usually show hypoenhancing necrotic and cystic areas on post-contrast sequences and are lesions usually straightly named as tumor rather than unspecific lesions by radiologists in their reports. Terms such as “CT” have been identified by our NLP algorithm to classify a radiology report into the metastasis category since it is not uncommon to recommend by radiologist’s further exams (like whole body CT) to rule out primary malignancies when there is high suspicion of metastatic brain disease. In the same line, the term “DTI” appears frequently linked to the “metastasis” group, probably due to the recommendations made by radiologist regarding the further performance of this advanced MRI sequence to surgical resection of single metastatic lesion planification. Other words such as “edema” or “vasogenic” have more weight linked to metastasis rather than HGG, probably because of a higher vasogenic edema/lesion ratio linked to metastasis compared with HGG, which usually shows non-enhancing infiltrative areas [5, 21].
The differential between HGG and metastasis is a common challenge in radiological practice. Despite several efforts based on conventional and advanced MRI sequences for improving this differential, nowadays, in some cases, there are still doubts about the nature of solitary-enhancing lesions in MRI studies [22, 23]. In this scenario, AI solutions may help radiologists as a clinical support decision tool for this task. To the best of our knowledge, this is the first paper to attempt to address differences between both lesions based on how they are described in radiology reports using NLP.
One of the critical points in the design of the algorithm was to remove all the keywords that may solely identify a lesion as HGG or metastasis to improve our tool’s clinical, radiological, and statistical value. In this manner, we ensured that the system does not get influenced in its final decision by the detection of terms such as “high grade,” “glioblastoma,” or “metastatic,” among others.
Several authors have recently developed NLP-based tools for extracting relevant information from radiology reports [12, 24]. Sensitive information such as unexpected or relevant findings can be extracted automatically from radiology reports to notify in a preferent manner these relevant findings to referring clinicians. López-Úbeda et al explored this topic, obtaining an F1 score for identifying unexpected findings at free-text radiology reports of 90% using CNN [25]. Regarding glioma evaluation, Di Noto et al developed a weakly supervised learning algorithm with automated labels and transfer learning techniques to detect glioma changes related to progression or response [26, 27]. Senders et al evaluated the role of NLP for automated quantification of brain metastasis reported in unstructured radiology reports finding that the bag-or-words approach combined with a least absolute shrinkage and selection operator (LASSO) provided the better overall accuracy with an AUC of 0.92 for binary classification of patients with single or multiple metastases in MRI brain studies [28]. NLP has also been applied in the CNS for other clinical scenarios, such as predicting stroke outcomes based on brain MRI radiology reports performed during admission. Heo et al obtained specific tokens (MCA, “territori,” “complet,” etc.) that could be used as digital markers of a patient’s prognosis in brain MRI reports linked to poor outcomes of patients with acute ischemic stroke using deep learning and CNN [29].
Our NLP tool can compile all the information in the free-text report and offer the radiologist the likelihood of suggesting HGG or metastasis based on the NLP analysis of finding details. We believe this tool has some potential applications in the standard radiological workflow. First, to serve, especially in the case of less experienced radiologists, as a clinical assistant tool before finalizing their reports, this kind of NLP solution may help them reach a correct final diagnosis on the basis of the findings described. In this line, a deep analysis of terms applied by an expert neuroradiologist can be done to use them as an example of how these reports must be performed or, on the opposite side, to detect poor-quality reports with a non-specific description of HGG or metastasis features and encourage and teach these radiologists to use more precise lexicon. Another potential application could be related to extracting information from radiology reports performed outside our radiology department. It is not uncommon to admit patients with MRI studies performed at other institutions, having only access to their radiology reports. In this manner, avoiding duplication of new MRI studies or improving the interpretation of external MRI reports may be achieved using these NLP solutions. Of course, the most logical and practical approach should be to integrate the NLP outcome with features derived from images (regardless of whether conventional, advanced, or based on AI or radiomics) to provide a final diagnosis using an AI multimodal approach that merges information from image and text. Other approaches may include automatic retrospective searching and labeling radiology reports from past years present at any RIS to ensure reporting quality and recruit patients for research and clinical trials [24].
Our study has some limitations. The insufficient number of radiology reports selected for the training and testing of the NLP tools may impact the absolute accuracy of the differential diagnosis between HGG and metastasis. In our opinion, an increase in the number of labeled reports with more cases of HGG and metastasis will undoubtedly improve our NLP tool’s capability for suggesting radiologist HGG or metastasis. Regarding the language used, probably the translation from Spanish to English language of our reports would have some kind of impact on the outcome of our NLP tool as linguistic nuances are probably being missed during the translation process. Regarding the type of CNS lesions included as part of the differential diagnosis, other solitary-enhancing lesions such as primary central nervous system lymphoma (PCNSL), brain abscesses, or tuberculomas may be potentially included in further studies to encompass a broader range of differential based on the description of radiological features of these additional lesions on the radiology reports.
Conclusions
Differentiation between HGG and brain metastasis remains nowadays a challenge for radiologists. We developed an NLP-based algorithm to extract information from radiology reports and accurately classify them as HGG or metastasis. This NLP-based algorithm could be used as an assistant tool together with imaging features to help radiologists in this challenging task.
Abbreviations
- AI:
-
Artificial intelligence
- BERT:
-
Bidirectional Encoder Representations from Transformers
- BiLSTM:
-
Bidirectional Long Short-Term Memory
- CNN:
-
Convolutional Neural Network
- CNS:
-
Central nervous system
- EHR:
-
Electronic health records
- HGG:
-
High-grade glioma
- ML:
-
Machine learning
- MRI:
-
Magnetic resonance imaging
- NLP:
-
Natural language processing
- RIS:
-
Radiology information system
References
Muccio CF, Tedeschi E, Ugga L et al (2019) Solitary cerebral metastases vs. high-grade gliomas: usefulness of two MRI signs in the differential diagnosis. Anticancer Res 39:4905-4909. https://doi.org/10.21873/anticanres.13677
Suh CH, Kim HS, Jung SC, Kim SJ (2018) Diffusion-weighted imaging and diffusion tensor imaging for differentiating high-grade glioma from solitary brain metastasis: a systematic review and meta-analysis. AJNR Am J Neuroradiol 39:1208–1214. https://doi.org/10.3174/ajnr.A5650
Pons-Escoda A, Garcia-Ruiz A, Naval-Baudin P et al (2022) Voxel-level analysis of normalized DSC-PWI time-intensity curves: a potential generalizable approach and its proof of concept in discriminating glioblastoma and metastasis. Eur Radiol 32:3705–3715. https://doi.org/10.1007/s00330-021-08498-1
Liu J, Han H, Xu Y et al (2021) A comparison of the multimodal magnetic resonance imaging features of brain metastases vs. High-grade gliomas Am J Transl Res 13:3543–3548
Martín-Noguerol T, Mohan S, Santos-Armentia E, Cabrera-Zubizarreta A, Luna A. (2021) Advanced MRI assessment of non-enhancing peritumoral signal abnormality in brain lesions. Eur J Radiol. 143:109900. https://doi.org/10.1016/j.ejrad.2021.109900
Qian Z, Li Y, Wang Y et al (2019) Differentiation of glioblastoma from solitary brain metastases using radiomic machine-learning classifiers. Cancer Lett 451:128–135. https://doi.org/10.1016/j.canlet.2019.02.054
Koh D-M, Papanikolaou N, Bick U et al (2022) Artificial intelligence and machine learning in cancer imaging. Commun Med 2:1–14. https://doi.org/10.1038/s43856-022-00199-0
Kalasauskas D, Kosterhon M, Keric N et al (2022) Beyond glioma: the utility of radiomic analysis for non-glial intracranial tumors. Cancers (Basel). 14:836
Pw C (2016) What can we learn from EHR developments? Int J Comput Assist Radiol Surg 11:S156–S157
Krupinski EA (2019) Artificial intelligence: lessons learned from radiology. Healthc Transform 5–10. https://doi.org/10.1089/heat.2019.0008
Pons E, Braun LMM, Hunink MGM, Kors JA (2016) Natural language processing in radiology: a systematic review. Radiology 279:329–343. https://doi.org/10.1148/radiol.16142770
López-Úbeda P, Martín-Noguerol T, Juluru K, Luna A (2022) Natural language processing in radiology: update on clinical applications. J Am Coll Radiol S1546:1440–7. https://doi.org/10.1016/j.jacr.2022.06.016
Wheater E, Mair G, Sudlow C et al (2019) A validated natural language processing algorithm for brain imaging phenotypes from radiology reports in UK electronic health records. BMC Med Inform Decis Mak 19:1–11. https://doi.org/10.1186/s12911-019-0908-7
Groot OQ, Bongers MER, Karhade AV et al (2020) Natural language processing for automated quantification of bone metastases reported in free-text bone scintigraphy reports. Acta Oncol 59:1455–1460. https://doi.org/10.1080/0284186X.2020.1819563
Bird S, Loper E (2016) The natural language toolkit NLTK: the Natural Language Toolkit. Proc ACL-02 Work Eff tools Methodol Teach Nat Lang Process Comput Linguist 63–70
Yan A, McAuley J, Lu X et al (2022) RadBERT: adapting transformer-based language models to radiology. Radiol Artif Intell 4:e210258. https://doi.org/10.1148/ryai.210258
Ribeiro MT, Singh S, Guestrin C (2016) “Why should I trust you?” explaining the predictions of any classifier. NAACL-HLT 2016 - 2016 Conf North Am Chapter Assoc Comput Linguist Hum Lang Technol Proc Demonstr Sess 97–101. https://doi.org/10.18653/v1/n16-3020
López-Úbeda, P., Martín-Noguerol, T. & Luna, A. (2023) Radiology, explicability and AI: closing the gap. Eur Radiol. https://doi.org/10.1007/s00330-023-09902-8
Yan An, McAuley J, Lu X, Du JCE, Gentili A-N (2022) RadBERT : adapting transformer-based language models to radiology. Radiol Artif Intell 4:e210258
Tay Y, Dehghani M, Gupta J, et al (2021) Are pre-trained convolutions better than pre-trained transformers? In: ACL-IJCNLP 2021 - 59th Annual Meeting of the Association for Computational Linguistics and the 11th International Joint Conference on Natural Language Processing, Proceedings of the Conference. pp 4349–4359. https://doi.org/10.48550/arXiv.2105.03322
Voicu IP, Pravatà E, Panara V et al (2022) Differentiating solitary brain metastases from high-grade gliomas with MR: comparing qualitative versus quantitative diagnostic strategies. Radiol Med 127:891–898. https://doi.org/10.1007/s11547-022-01516-2
Cindil E, Sendur HN, Cerit MN et al (2021) Validation of combined use of DWI and percentage signal recovery-optimized protocol of DSC-MRI in differentiation of high-grade glioma, metastasis, and lymphoma. Neuroradiology 63:331–342. https://doi.org/10.1007/s00234-020-02522-9
Fu M, Han F, Feng C et al (2019) Based on arterial spin labeling helps to differentiate high-grade gliomas from brain solitary metastasis: a systematic review and meta-analysis. Medicine (Baltimore) 98:e15580
Casey A, Davidson E, Poon M et al (2021) A systematic review of natural language processing applied to radiology reports. BMC Med Inform Decis Mak 21:1–18. https://doi.org/10.1186/s12911-021-01533-7
López-Úbeda P, Díaz-Galiano MC, Martín-Noguerol T et al (2020) Detection of unexpected findings in radiology reports: a comparative study of machine learning approaches. Expert Syst Appl 160:113647. https://doi.org/10.1016/j.eswa.2020.113647
Di Noto T, Atat C, Teiga EG, et al (2021) Diagnostic surveillance of high-grade gliomas: towards automated change detection using radiology report classification. Commun Comput Inf Sci 1525 CCIS:423–436. https://doi.org/10.1101/2021.09.24.21264002
Di Noto T, Bach Cuadra M, Atat C, et al (2023) Weakly supervised learning with automated labels from radiology reports for glioma change detection. https://doi.org/10.48550/arXiv.2210.09698
Senders JT, Karhade AV, Cote DJ et al (2019) Natural language processing for automated quantification of brain metastases reported in free-text radiology reports. JCO Clin Cancer Inform 3:1–9. https://doi.org/10.1200/cci.18.00138
Heo TS, Kim YS, Choi JM et al (2020) Prediction of stroke outcome using natural language processing-based machine learning of radiology report of brain MRI. J Pers Med 10:1–11. https://doi.org/10.3390/jpm10040286
Funding
This paper was partially funded by the Ministry of Science and Innovation (MCIN/AEI/https://doi.org/10.13039/501100011033), grant number PTQ2021-012120.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Dr. Antonio Luna.
Conflict of interest
The authors of this manuscript declare relationships with the following companies: Antonio Luna, MD, PhD, is an occasional lecturer of Philips, Siemens Healthineers, Bracco, and Canon and receives royalties as a book editor from Springer-Verlag. Albert Pons-Escoda is a member of European Radiology Scientific Editorial Board and has therefore not taken part in review and selection of this article. The rest of the authors of this manuscript declare no relationships with any companies whose products or services may be related to the subject matter of the article.
Statistics and biometry
One of the authors (Pilar López-Úbeda) has significant statistical expertise.
Informed consent
Written informed consent was not required in view of the retrospective nature of the study, based on radiology reports, and all the procedures being performed were part of the routine radiology practice.
Ethical approval
Institutional Review Board approval was not required because of the retrospective nature of the study, based on radiology reports, and all the procedures being performed were part of the routine radiology practice.
Study subjects or cohorts overlap
None
Methodology
• retrospective
• diagnostic or prognostic study
• multicenter study
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Martín-Noguerol, T., López-Úbeda, P., Pons-Escoda, A. et al. Natural language processing deep learning models for the differential between high-grade gliomas and metastasis: what if the key is how we report them?. Eur Radiol 34, 2113–2120 (2024). https://doi.org/10.1007/s00330-023-10202-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-023-10202-4